Copyright: ©Author(s) 2026.
World J Hepatol. Apr 27, 2026; 18(4): 114804
Published online Apr 27, 2026. doi: 10.4254/wjh.v18.i4.114804
Published online Apr 27, 2026. doi: 10.4254/wjh.v18.i4.114804
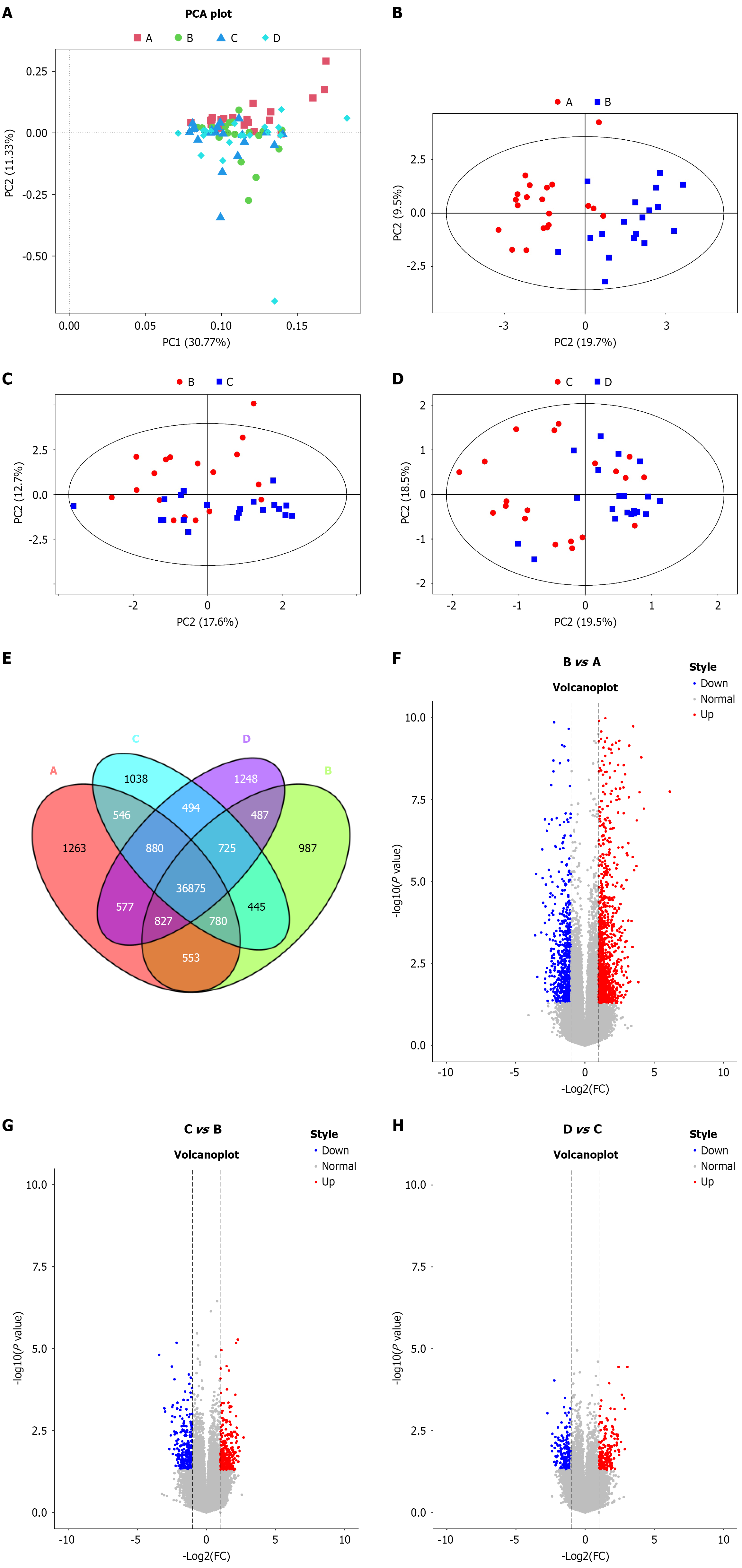
Figure 1 Identification of differentially expressed genes after interferon treatment for chronic hepatitis B through transcriptomic analysis.
A: Principal component analysis (PCA) of all differentially expressed genes (DEGs) across the four groups: Untreated (group A); 1-month interferon (IFN) treatment (group B); 3-month IFN treatment (group C); 6-month IFN treatment (group D); B: PCA comparing (B vs A); C: PCA comparing (B vs C); D: PCA comparing (C vs D); E: Venn diagram showing the overlap of all DEGs across the four groups; F: Volcano plots of DEGs in the comparisons (B vs A); G: Volcano plots of DEGs in the comparisons (C vs B); H: Volcano plots of DEGs in the comparisons (D vs C). PCA: Principal component analysis.
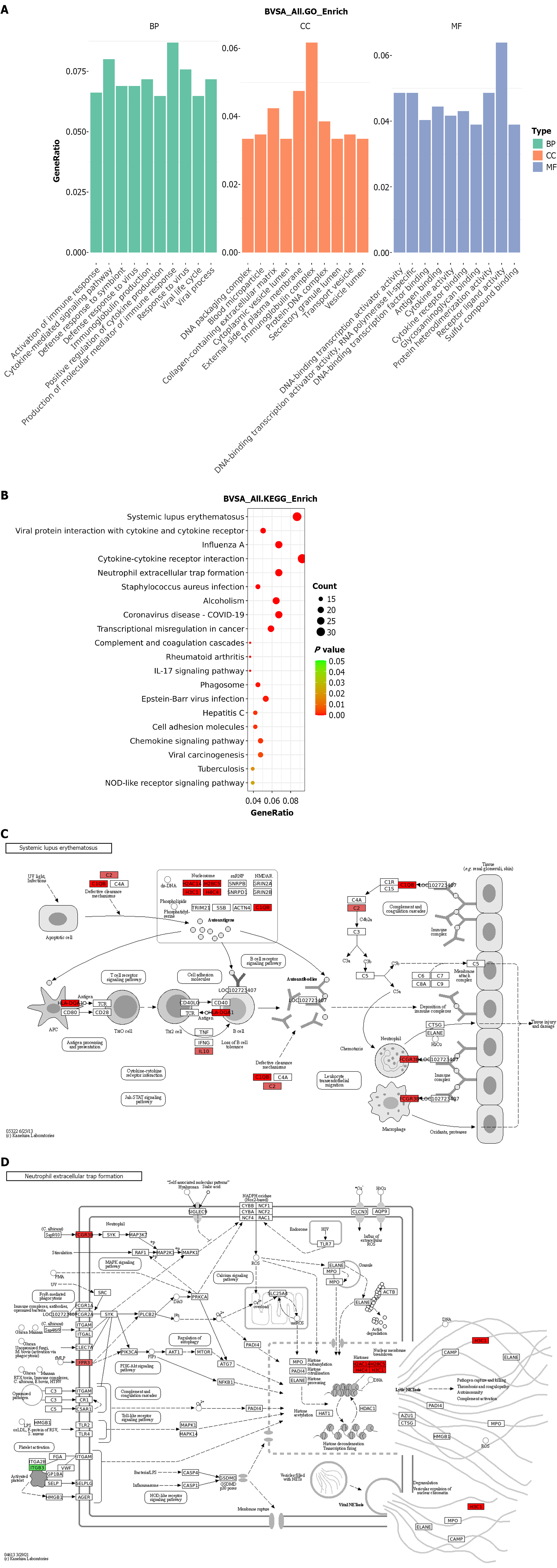
Figure 2 Enrichment analysis of differentially expressed genes after interferon treatment.
A: Bar chart of Gene Ontology classification enrichment, including the three ontologies: Cellular component, molecular function, and biological process, for the B vs A group; B: Bar chart showing Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis of the top 20 pathways for differentially expressed genes in the B vs A group; C: KEGG pathway maps for “systemic lupus erythematosus” (KEGG map 05322) and “neutrophil extracellular trap” (KEGG map 04613); D: KEGG pathway maps for “systemic lupus erythematosus” (KEGG map 05322) for the B vs A group [permission from KEGG (12)]. GO: Gene Ontology; KEGG: Kyoto Encyclopedia of Genes and Genomes; BP: Biological process; CC: Cellular component; MF: Molecular function.
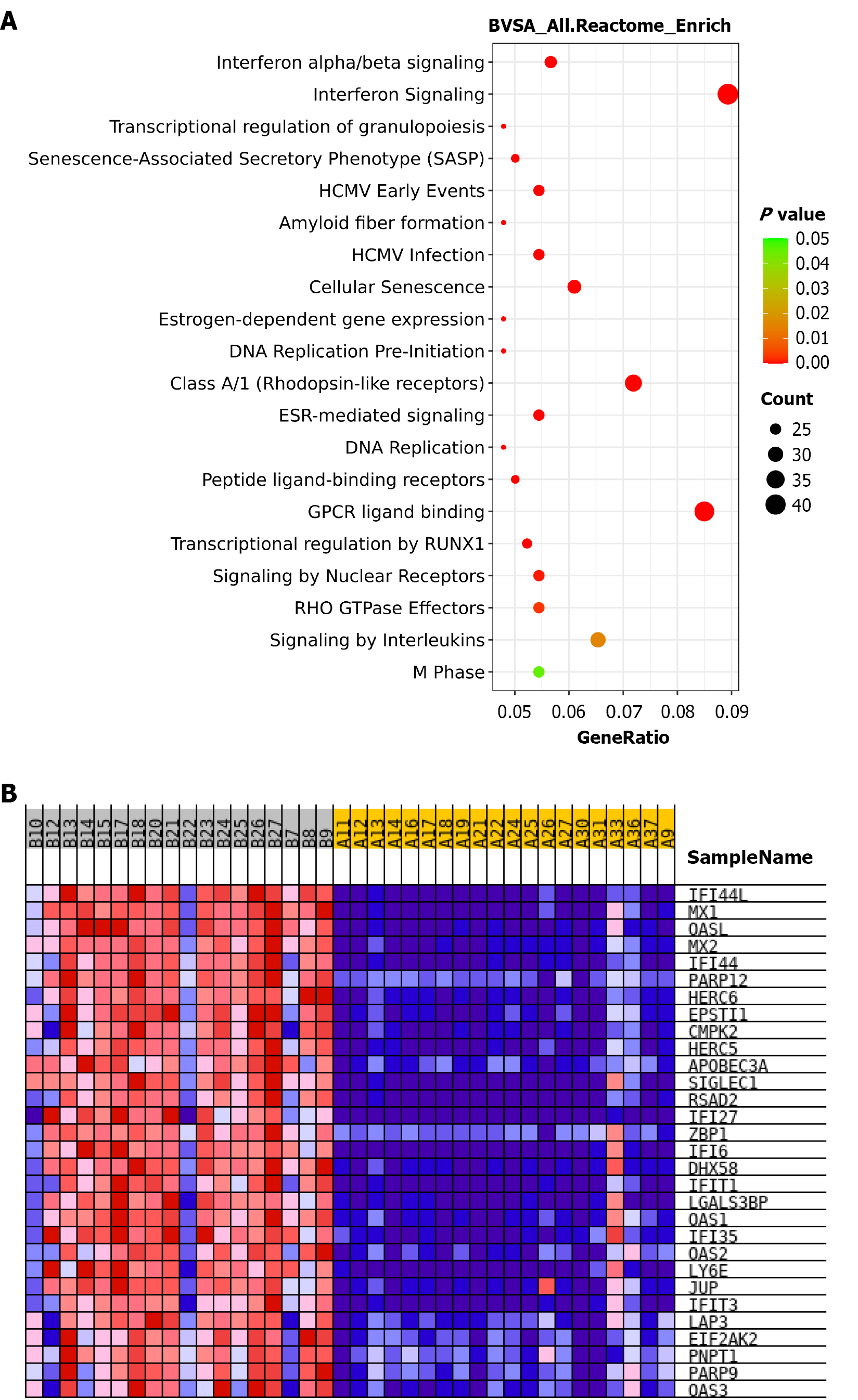
Figure 3 Kyoto Encyclopedia of Genes and Genomes enrichment analysis of differentially expressed genes after interferon treatment.
A: Rectangular tree diagram showing the top enriched differentially expressed genes pathways in the B vs A group; B: Heatmap of differentially expressed genes in the “IFNbeta targets” gene set, based on gene set enrichment analysis for the B vs A group.
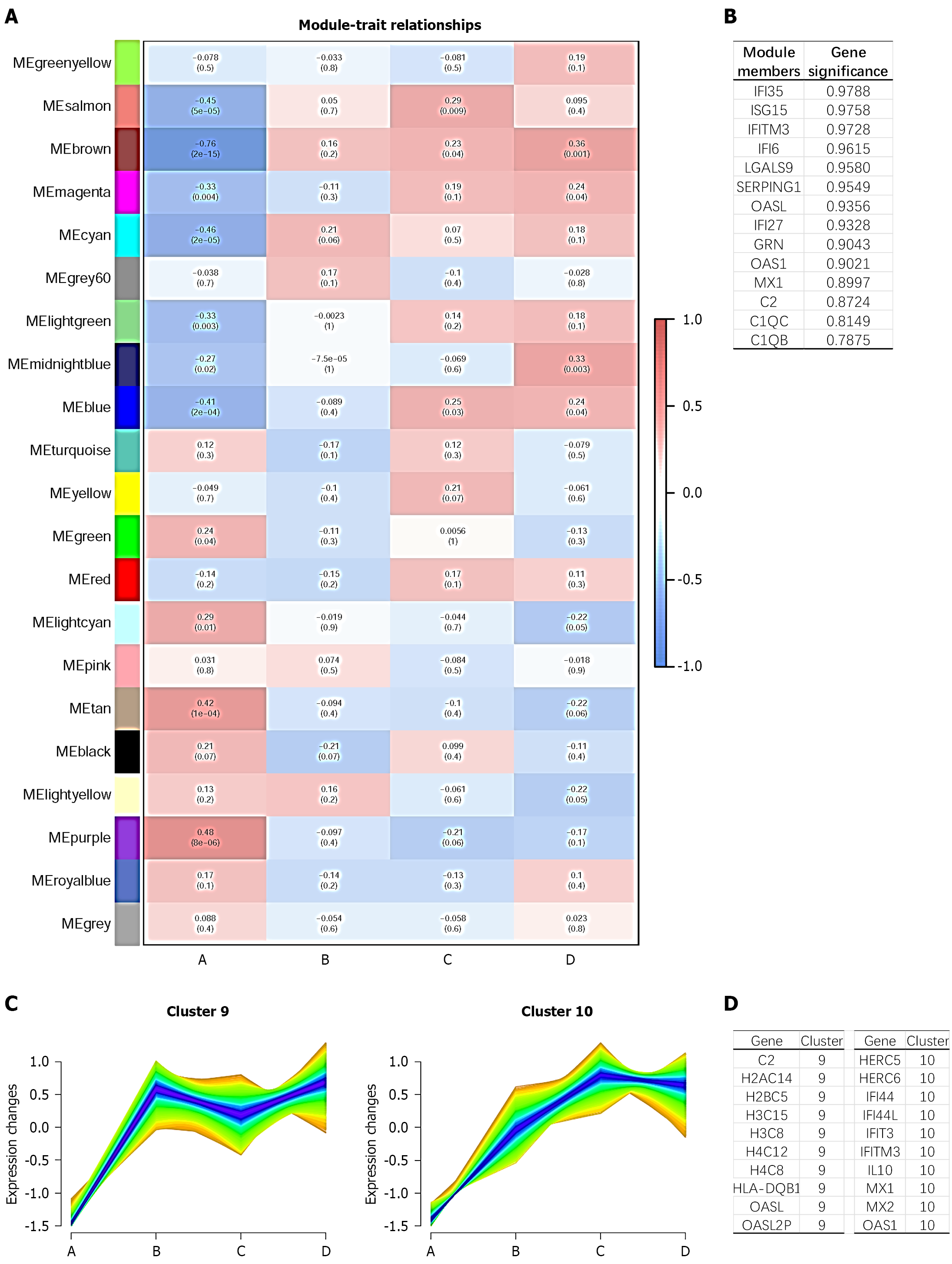
Figure 4 Weighted gene co-expression network analysis and Mfuzz analysis of differentially expressed genes after interferon treatment.
A: Weighted gene co-expression network analysis showing the correlation between modules and the four groups; B: List of module members with the top gene significance; C: Mfuzz pseudo-time analysis clustering of differentially expressed genes in clusters 9 and 10 based on metabolic network pathways; D: List of genes related to clusters 9 and 10.
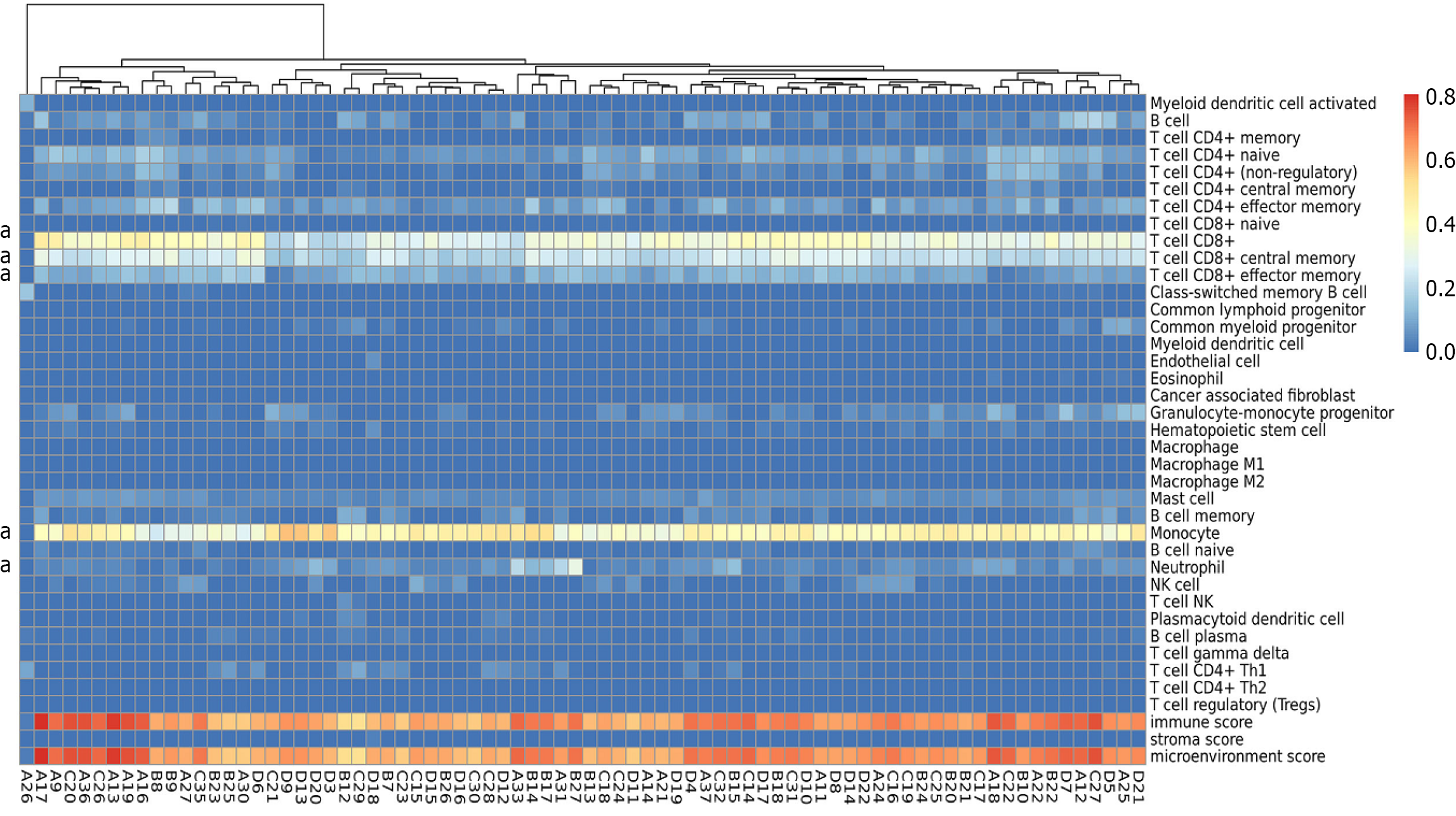
Figure 5 Immune infiltration analysis of blood samples across all four groups after interferon treatment for chronic hepatitis B.
a: Indicates the significantly altered immune cell subtypes.
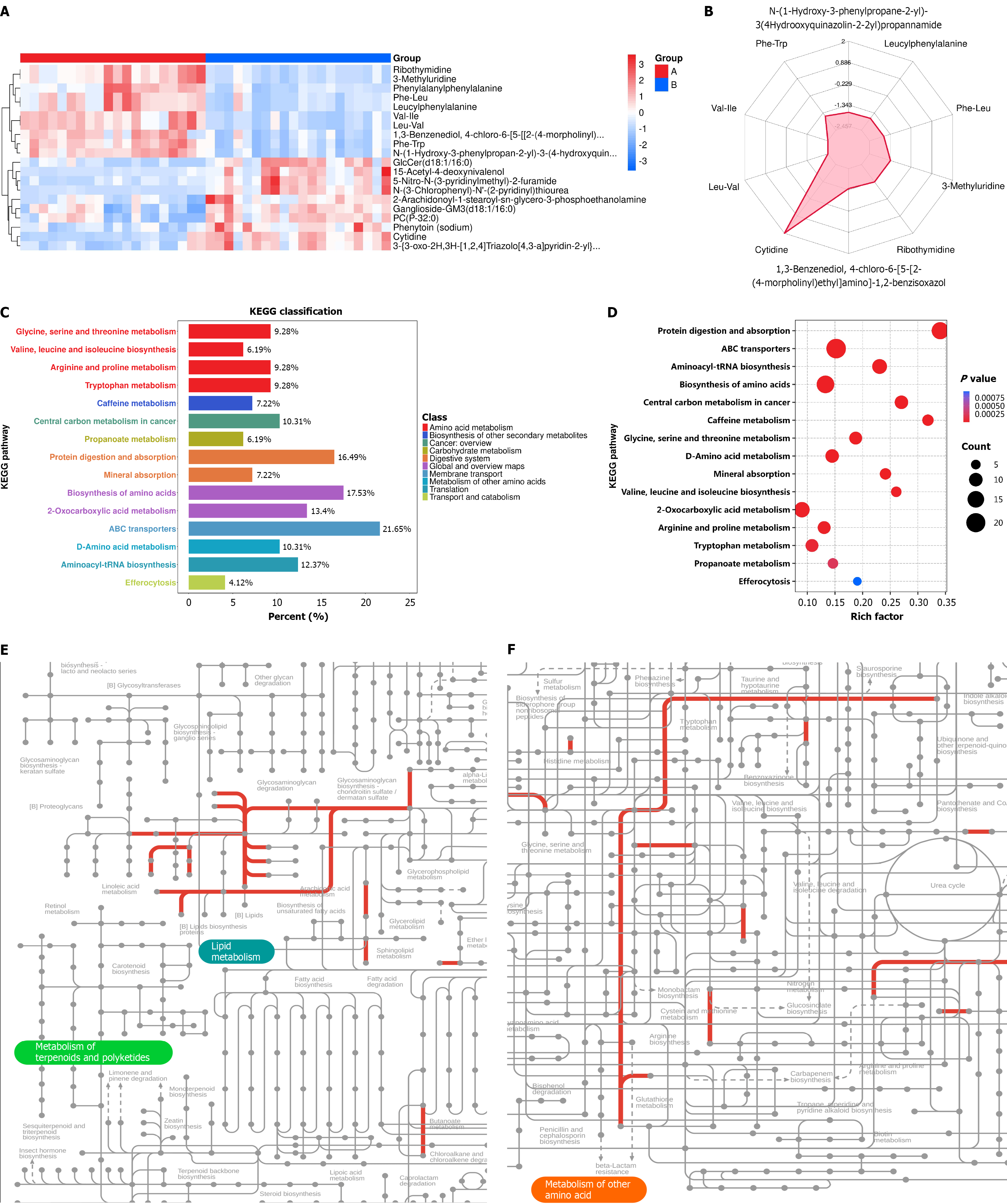
Figure 6 Metabolomics analysis after interferon treatment.
A: Hierarchical clustering analysis of the top 20 differential metabolites in the B vs A group; red represents upregulation, and blue represents downregulation; B: Radar plot showing the top 10 differential metabolites in the B vs A group; C: Enrichment analysis of differential metabolites in the B vs A group; Kyoto Encyclopedia of Genes and Genomes classification chart; D: Enrichment analysis of differential metabolites in the B vs A group; bar chart of the top 15 enriched Kyoto Encyclopedia of Genes and Genomes pathways; E: Parts of the iPath analysis for metabolites in the B vs A group, including lipid metabolism and metabolism of other amino acids; F: Parts of the iPath analysis for metabolites in the B vs A group, including lipid metabolism; red lines indicate upregulated pathways.
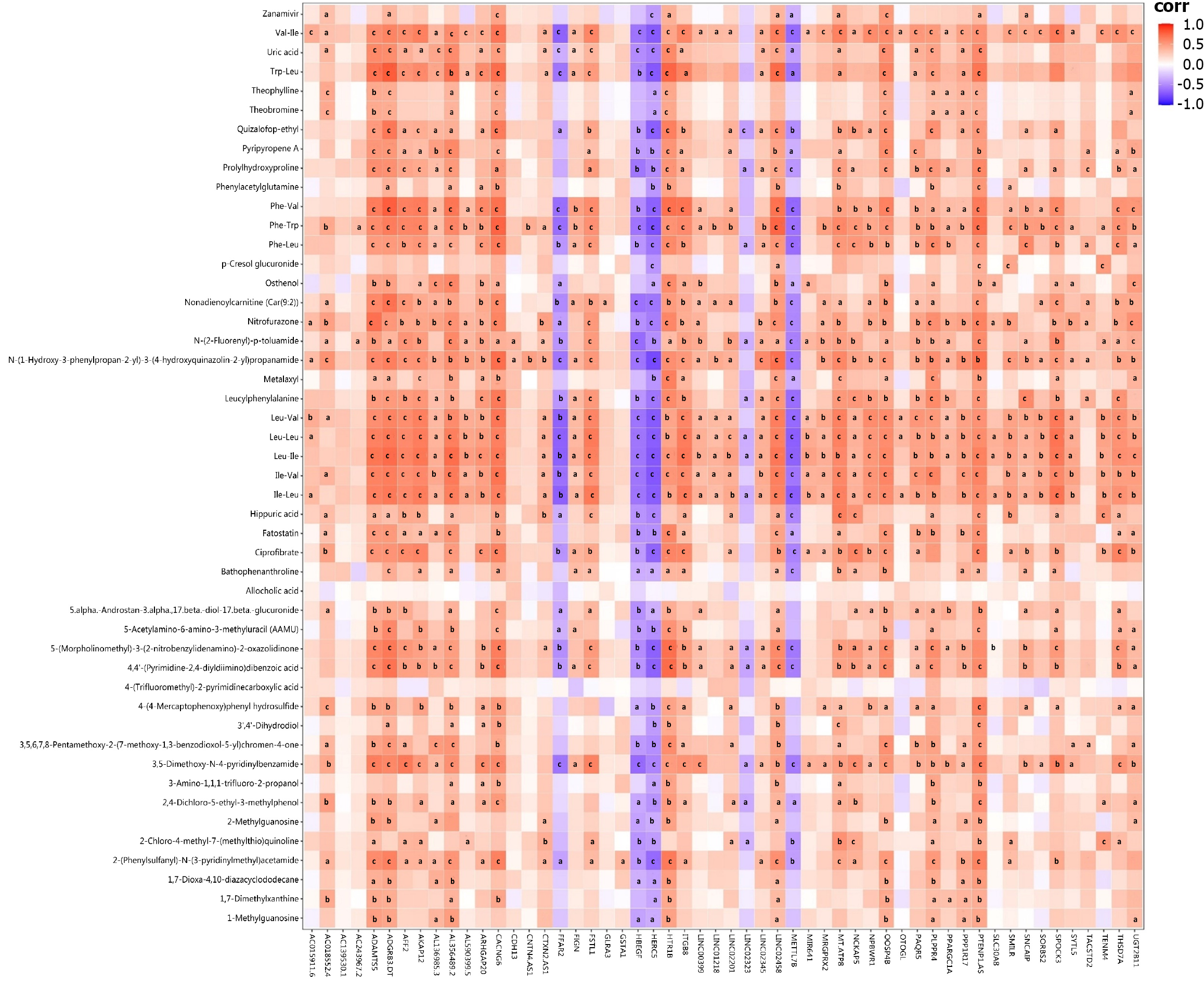
Figure 7 Correlation between differentially expressed genes and differential metabolites after interferon alpha treatment.
Pearson correlation analysis of the datasets from transcriptomics and metabolomics in the B vs A group. aP < 0.05, bP < 0.01, cP < 0.001.
- Citation: Ye XY, He XZ, Hu ZT, Zheng FF, Huang XG, Xie XM, Chen FH, Ou HB, Qiu RX. Integrated transcriptomics and metabolomics reveal neutrophil extracellular trap associated with interferon treatment for chronic hepatitis B. World J Hepatol 2026; 18(4): 114804
- URL: https://www.wjgnet.com/1948-5182/full/v18/i4/114804.htm
- DOI: https://dx.doi.org/10.4254/wjh.v18.i4.114804