INTRODUCTION
Colorectal cancer is one of the most common malignant tumors in the digestive tract[1]. There are approximately 1.2 million new cases of colorectal cancer every year worldwide, and its mortality rate accounts for approximately 8% of all malignant tumors[2]. Current primary treatments include radical surgery and radiotherapy; however, these approaches are associated with high postoperative recurrence and metastasis rates[3]. Increasing amounts of clinical evidence indicate that tumor cells secrete intracellular non-coding RNAs[4] with specific functions into the tumor microenvironment via exosome[5] pathways, thereby influencing tumor cell proliferation, apoptosis, and metastasis[6,7]. Moreover, exosomal miR-191 plays a crucial role in the development of various cancers, including prostate cancer[8,9], hepatocellular carcinoma[10], colorectal cancer[11], and endometrial cancer[12]. Previous studies have also demonstrated that tumor-associated macrophages (TAMs) affect the prognosis and efficacy of chemotherapy and immunotherapy[13,14]. In most cases, TAMs even directly promote tumor progression, suggesting their close association with colorectal cancer progression[15]. However, there is currently no reported evidence regarding whether exosomal miR-191 originating from colorectal cancer influences tumor progression by altering macrophage polarization.
INHIBITING MIR-191 SIGNIFICANTLY REDUCES ITS EXPRESSION IN COLORECTAL CANCER CELLS AND THEIR EXOSOMES, THEREBY PREVENTING M2 MACROPHAGE POLARIZATION
A recent study by Zhao et al[16] revealed the role of miR-191 in colorectal cancer cell exosomes. Researchers have transfected miR-191 inhibitors into colorectal cancer cells and isolated exosomes. Quantitative real-time polymerase chain reaction (qRT-PCR) analysis revealed that inhibiting miR-191 significantly reduced its expression in both cancer cells and their exosomes, indicating effective knockdown of miR-191.
The research team then cocultured colorectal cancer cells transfected with miR-191 inhibitors with M0 macrophages. Flow cytometry[17] was used to analyze the polarization status of M2 macrophages[18] and detect the expression levels of their markers cluster of differentiation (CD) 68 and CD163[19,20]. The results revealed that the expression of CD68 and CD163 in the miR-191 inhibitor group was significantly lower than that in the control group. Further analysis of the levels of the intracellular inflammatory factors interleukin (IL)-4 and IL-10[21] revealed a marked decrease in these levels in the miR-191 inhibitor group compared with those in the negative control (NC) group. Western blot experiments also demonstrated that miR-191 inhibition downregulated the expression of anti-chitinase-like protein 3 (YM1) and arginase-1 (Arg-1) in macrophages.
This study employed 1,1’-dioctadecyl-3,3,3’,3’-tetramethylindocarbocyanine perchlorate (PKH26/DiI)[22] for exosome labeling and 4’,6-diamidino-2-phenylindole a nucleic acid-specific fluorescent dye for nuclear staining[23] for macrophage nuclei. Immunofluorescence imaging[24] revealed red fluorescence around the blue-labeled nuclei, confirming successful exosome uptake by the macrophages. Notably, exosomes from colorectal cancer cells treated with miR-191 inhibitors presented significantly reduced intracellular miR-191 levels after macrophage-mediated uptake. These findings collectively demonstrate that inhibiting miR-191 significantly reduces its expression in colorectal cancer cells and their exosomes, thereby effectively preventing M2 polarization in macrophages.
DOWNREGULATION OF MIR-191 ACCELERATES FERROPTOSIS TO SUPPRESS INFLAMMATORY RESPONSES AND M2 POLARIZATION OF MACROPHAGES IN COLORECTAL CANCER CELLS
Colorectal cancer cells transfected with miR-191 inhibitors were cocultured with M0 macrophages. qRT-PCR analysis revealed significant downregulation of miR-191 expression in cancer cells after transfection. Flow cytometry revealed markedly higher levels of reactive oxygen species in the inhibitor group than in the control group. Western blotting further confirmed significantly lower expression levels of the ferroptosis markers solute carrier family 7 member 11 (SLC7A11)[25] and glutathione peroxidase 4 (GPX4)[26] in the inhibitor group than in the control group. SLC7A11, a cysteine-glutamate antiporter, is inhibited, leading to intracellular cysteine depletion, reduced glutathione (GSH) synthesis, and the intracellular accumulation of lipid peroxides[27]. As a key inhibitor of lipid peroxides, GPX4 oxidizes GSH to glutathione disulfide. The inhibition of GPX4 activity accelerates intracellular GSH depletion, ultimately triggering ferroptosis[28]. TdT-mediated dUTP nick-end labeling (TUNEL) staining[29] confirmed the observed changes in apoptosis, indicating that the miR-191 inhibitor significantly increased the apoptosis rate in colorectal cancer cells.
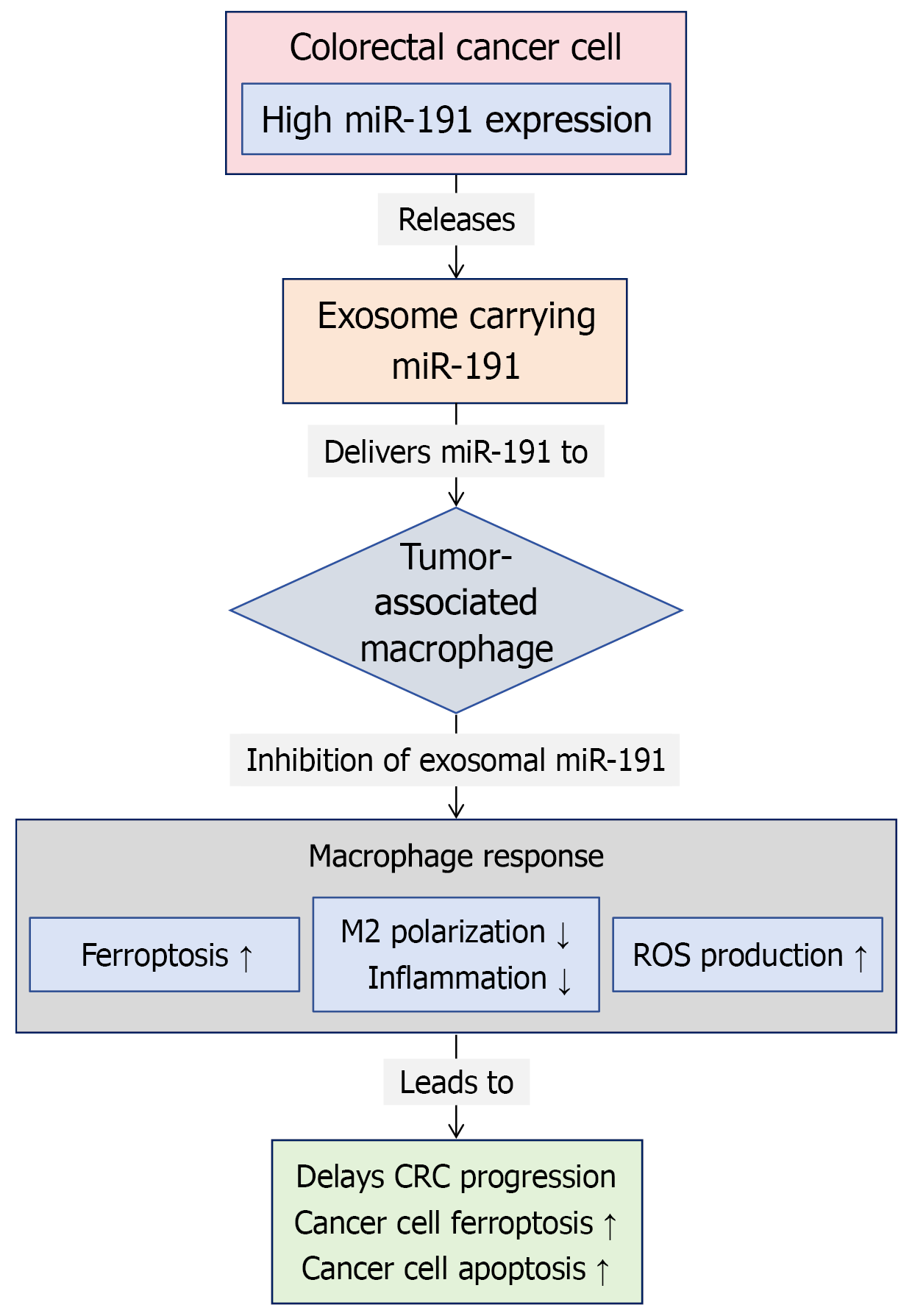
This study further investigated whether miR-191 in colorectal cancer cell exosomes regulates ferroptosis to influence M2 macrophage polarization (Figure 1). In the present study, macrophages treated with colorectal cancer exosomes were subsequently exposed to miR-191 inhibitors and/or ferroptosis inhibitors (ferrostatin-1)[30]. Flow cytometry revealed that the CD68 and CD163 marker expression levels in the miR-191 inhibitor group were significantly lower than those in the control group, with ferrostatin-1 reversing this decrease in CD68/CD163 positivity. Enzyme-linked immunosorbent assay (ELISAs) demonstrated that exosome-derived miR-191 suppression reduced IL-4 and IL-10 concentrations in macrophages, an effect that was partially reversed by ferrostatin-1. Western blotting revealed that exosome miR-191 inhibition downregulated YM1 and Arg-1 expression in macrophages, and ferrostatin-1 treatment significantly alleviated this downregulation. qRT-PCR further confirmed that exosomal miR-191 suppression decreased IL-4 and IL-10 levels in macrophages by inducing ferroptosis.
Figure 1 Mechanism of action of exosomal miR-191 in colorectal cancer progression.
This flowchart illustrates the role of exosomal miR-191 derived from colorectal cancer cells in promoting tumor progression by modulating macrophage function and the therapeutic effects of its inhibition. ROS: Reactive oxygen species; CRC: Colorectal cancer.
Furthermore, by coincubating colorectal cancer cells with exosomes, miR-191 inhibitors, or macrophages treated with hepcidin-1, we investigated whether macrophage ferroptosis is associated with the effects of miR-191 from colorectal cancer-derived exosomes on cancer-related functions. The qRT-PCR results revealed that inhibiting exosomal miR-191 downregulated miR-191 expression in colorectal cancer cells, whereas hepcidin-1 reversed this phenomenon. Inhibiting exosomal miR-191 increased reactive oxygen species production in colorectal cancer cells, which was partially mitigated by hepcidin-1. Hepcidin-1 treatment also significantly reversed the reduced expression of SLC7A11 and GPX4 in colorectal cancer cells caused by miR-191 inhibitors. TUNEL staining demonstrated that hepcidin-1 weakened the increase in apoptosis in the miR-191 inhibitor group. This study ultimately confirmed that inhibiting exosomal miR-191 induces ferroptosis in macrophages, thereby delaying colorectal cancer progression.
EXOSOMES CARRYING MIR-191 FROM COLORECTAL CANCER CELLS PROMOTE DISEASE PROGRESSION BY INDUCING M2 MACROPHAGE POLARIZATION
In this study, exosomes containing miR-191 and/or erastin were injected into a colon cancer model in nude mice, with subcutaneous tumors harvested for observation on day 28. Compared with the exosome control group (ExoNC), the exosome miR-191 group (ExomiR-191) presented significantly increased tumor weight and volume, a phenomenon partially inhibited by erastin. Hematoxylin-eosin staining revealed that ExoNC group tumors consisted mainly of oval, oblong, or polygonal epithelioid cells with tight arrangements, occasionally showing clustered or band-like distributions. In contrast, the tumors in the ExomiR-191 group presented distinct clustered lesions with disordered cell arrangement and reduced interstitial polarity. The combined ExomiR-191 and erastin group presented tumor characteristics similar to those of the ExoNC group. ELISAs demonstrated that ExomiR-191 significantly elevated the serum concentrations of IL-4 and IL-10 in mice, and this effect was reversed by erastin. Additionally, erastin inhibited the upregulation of miR-191 and the increase in four proteins (SLC7A11, GPX4, YM1, and Arg-1) in ExomiR-191 group tumors. Further research confirmed that exosomal miR-191 in colon cancer promotes subcutaneous tumor growth and alters pathological structure by inducing M2 macrophage polarization.
LIMITATIONS AND FUTURE PERSPECTIVES
Although this study preliminarily revealed the functional role of miR-191 in colorectal cancer, certain limitations must be acknowledged. To date, no definitive studies have confirmed that miR-191 directly targets and regulates core genes involved in macrophage ferroptosis. Additionally, the limited patient recruitment period in this study prevented a sufficient sample size for robust statistical analysis, and the single-center design of the experiment hindered further evaluation of the associations between miR-191 expression and clinicopathological parameters or survival outcomes. Given the unexplored direct downstream target genes of miR-191 identified in this study, future research could employ RNA sequencing and dual-luciferase reporter gene assays to identify the direct targets of miR-191 in macrophage ferroptosis, thereby elucidating the molecular mechanism involved. To comprehensively assess the clinical significance of miR-191, future multicenter, large-sample clinical studies are needed to validate its translational value. Furthermore, miR-191 also has protumorigenic effects on prostate cancer and hepatocellular carcinoma. Whether it regulates the progression of other cancers through a similar exosome macrophage ferroptosis mechanism remains to be further investigated. In line with the current research hotspot on ‘tumor immunotherapy combined with ferroptosis induction’, this study provides a novel target for combination therapy strategies. Future research could explore the synergistic effects of miR-191 inhibitors with programmed death-1/programmed cell death ligand 1 antibodies.
CONCLUSION
In CRC cells, inhibiting miR-191 effectively suppresses macrophage M2 polarization by inducing the production of reactive oxygen species in CRC cells, triggering ferroptosis and subsequent apoptosis. Furthermore, blocking miR-191 in CRC cell exosomes prevents macrophage M2 polarization and effectively inhibits CRC progression by inducing macrophage ferroptosis. Therefore, this research provides a robust experimental basis for developing immunotherapeutic strategies against colorectal cancer.
ACKNOWLEDGEMENTS
Hereby, I sincerely thank the university that nurtured me for providing a good scientific research platform and learning environment; I express my deepest gratitude to my supervisor for his careful guidance, patient enlightenment and selfless care during the research process; at the same time, I thank the students in the same group for their joint collaboration and mutual support in the research, as well as all relevant personnel who have provided help for this paper.
Peer review: Externally peer reviewed.
Peer-review model: Single blind
Specialty type: Gastroenterology and hepatology
Country of origin: China
Peer-review report’s classification
Scientific quality: Grade A
Novelty: Grade B
Creativity or innovation: Grade B
Scientific significance: Grade A
P-Reviewer: Liu Y, PhD, Senior Researcher, China S-Editor: Fan M L-Editor: A P-Editor: Zhang L