Published online Mar 15, 2026. doi: 10.4251/wjgo.v18.i3.115722
Revised: December 9, 2025
Accepted: December 30, 2025
Published online: March 15, 2026
Processing time: 140 Days and 9.7 Hours
Colorectal cancer (CRC) is a highly prevalent and lethal malignancy worldwide, marked by a multifaceted pathogenesis that involves various genetic, environmental, and lifestyle factors. Recent studies have increasingly underscored the significant role of gut microbiota in CRC, particularly focusing on how these mi
Core Tip: The treatment of colorectal cancer (CRC) faces challenges of drug resistance mediated by the tumor microenvironment (TME). This review proposes that the gut microbiota is a core regulator in reshaping CRC immune metabolism and TME. We have established an integrated framework of the “microbiota-immune metabolism-TME” axis, systematically elaborating on the mechanisms of CRC occurrence, development, and treatment resistance from the micro level to the macro level. This perspective provides new ideas for deconstructing complexities such as clinical drug resistance. Based on this, we further explore translational strategies targeting the microbiota to reverse immunosuppression, enhance the efficacy of radiotherapy, chemotherapy, and immunotherapy, offering innovative theoretical basis and intervention prospects for the precise treatment of CRC.
- Citation: Luan WY, Zhang SP, Xu KZ, Shang YH, Hu WJ, Sun H, Miao YD. Microbiota-driven immunometabolic regulation in colorectal cancer: Mechanisms and therapeutic opportunities. World J Gastrointest Oncol 2026; 18(3): 115722
- URL: https://www.wjgnet.com/1948-5204/full/v18/i3/115722.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v18.i3.115722
Colorectal cancer (CRC) has the second-highest global incidence and mortality rates of all cancers[1]. A key challenge in treating CRC is that complex immunometabolic changes within tumors can induce immunosuppression, limiting the effectiveness of immunotherapy. Although the 5-year survival rate for CRC has improved over time, rising from 50% in the 1970s to 65% in 2012-2018, a concerning trend has emerged: Incidence rates among individuals aged 15-50 years have increased by 80%-100% in recent years[2]. This rise in early-onset CRC presents significant new challenges for treatment. In the era of precision medicine, immunotherapy has gained widespread attention as a complement to conventional surgery, radiotherapy, and chemotherapy. Its principal advantage is the precise targeting of tumor cells, which minimizes damage to healthy tissues[3]. However, immunotherapy is effective only in a limited subset of patients. A key predictive biomarker is high microsatellite instability, but this status is found in only about 15% of all CRC cases and a mere 4% of metastatic cases[3-7]. The tumor microenvironment (TME) is a critical factor in this limited response. Understanding the dynamic interactions between the cellular and non-cellular components of the CRC TME is essential for improving the clinical efficacy of immunotherapy.
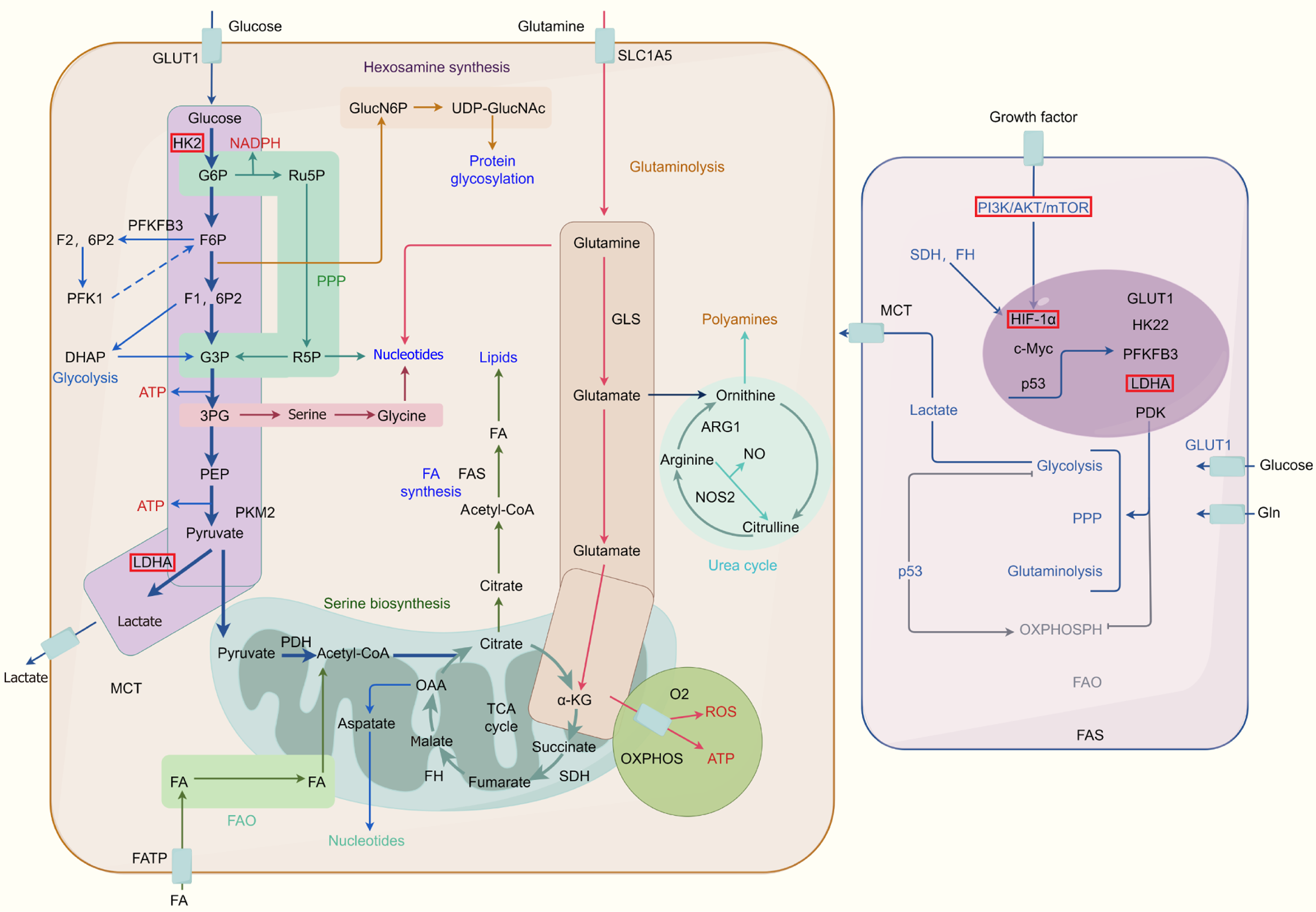
Metabolic reprogramming is a hallmark of CRC[8] (Figure 1) and is closely linked to the low immunotherapy response rates observed in this cancer. For instance, intestinal stem cells, which can initiate CRC, possess unique metabolic traits that confer resistance to conventional therapies[9]. A primary metabolic alteration in CRC cells is a heightened reliance on glycolysis, known as the Warburg effect, even in the presence of oxygen[10]. Concurrently, alterations in lipid meta
Driven by these metabolic alterations, along with genetic and epigenetic changes, CRC evolves robust immune evasion mechanisms. These are typified by the exhaustion of T cells and the polarization of macrophages towards a pro-tumor (M2) phenotype[14-17]. The gut microbiota is now recognized as a key regulator of immunometabolism and a potent modifier of the TME in CRC. For example, in advanced CRC, microbial-derived ammonia can accumulate and trigger metabolic reprogramming in T cells, promoting their exhaustion and contributing to immunotherapy resistance[18]. Given its direct anatomical connection to the colon and its demonstrated role in carcinogenesis, the gut microbiota holds unique significance in CRC, distinguishing it from many other cancer types.
The role of the gut microbiota in the initiation and progression of CRC is now widely acknowledged. As a major symbiotic community, the composition and function of the gut microbiota are critical for host health. An imbalance, known as dysbiosis, is strongly linked to CRC development[19]. Specific pro-oncogenic species include Fusobacterium nucleatum (F. nucleatum), certain strains of Escherichia coli (E. coli), and enterotoxigenic Bacteroides fragilis (B. fragilis)[20]. Furthermore, modulating the gut microbiota shows promise for improving immunotherapy efficacy and reducing radiotherapy-related toxicity[21]. For instance, the compound ginsenoside Rh4 exerts anti-CRC effects in a microbiota-dependent manner by altering bile acid metabolism and increasing levels of ursodeoxycholic acid (UDCA)[22]. Therefore, a deeper understanding of the gut microbiota's interaction with host immunometabolism is crucial for developing novel therapeutic strategies for CRC.
Therefore, investigating the changes in the relationship between CRC, the immune system, and the gut microbiota during the development and progression of CRC is of great clinical significance. The primary objective of this review is to integrate and synthesize recent research on the interplay between the gut microbiota and immunometabolism phe
To ensure a comprehensive and reproducible overview of the current knowledge, a systematic literature search was conducted. Primary searches were performed in the electronic databases PubMed, Web of Science, and Scopus for articles published between January 1984 and October 2025. The search strategy employed a combination of keywords and MeSH terms related to three core themes: (1) Colorectal cancer (“colorectal cancer”, “CRC”, “colon cancer”, “rectal cancer”); (2) Gut microbiota (“gut microbiome”, “intestinal flora”, “microbiota”); and (3) Immunometabolism and tumor microenvironment (“immunometabolism”, “tumor microenvironment”, “TME”, “immunotherapy”, “metabolic reprogramming”). Boolean operators (AND, OR) were used to link these concepts. The initial search results were screened based on their titles and abstracts to identify relevant studies. The full texts of the shortlisted articles were then thoroughly assessed against predefined inclusion criteria (original research or review articles in English focusing on the interplay between gut microbiota, immunometabolism, and CRC) and exclusion criteria (studies not relevant to the core focus, non-English publications, or conference abstracts). Any disagreements during the selection process were resolved through consensus among the authors. Finally, a total of 2010 documents were included in the analysis. In addition, we will outline the current progress in the clinical translation of research related to the gut microbiota and immunometabolism, further elucidating promising innovative therapeutic approaches from a clinical perspective. We hope that our study can cont
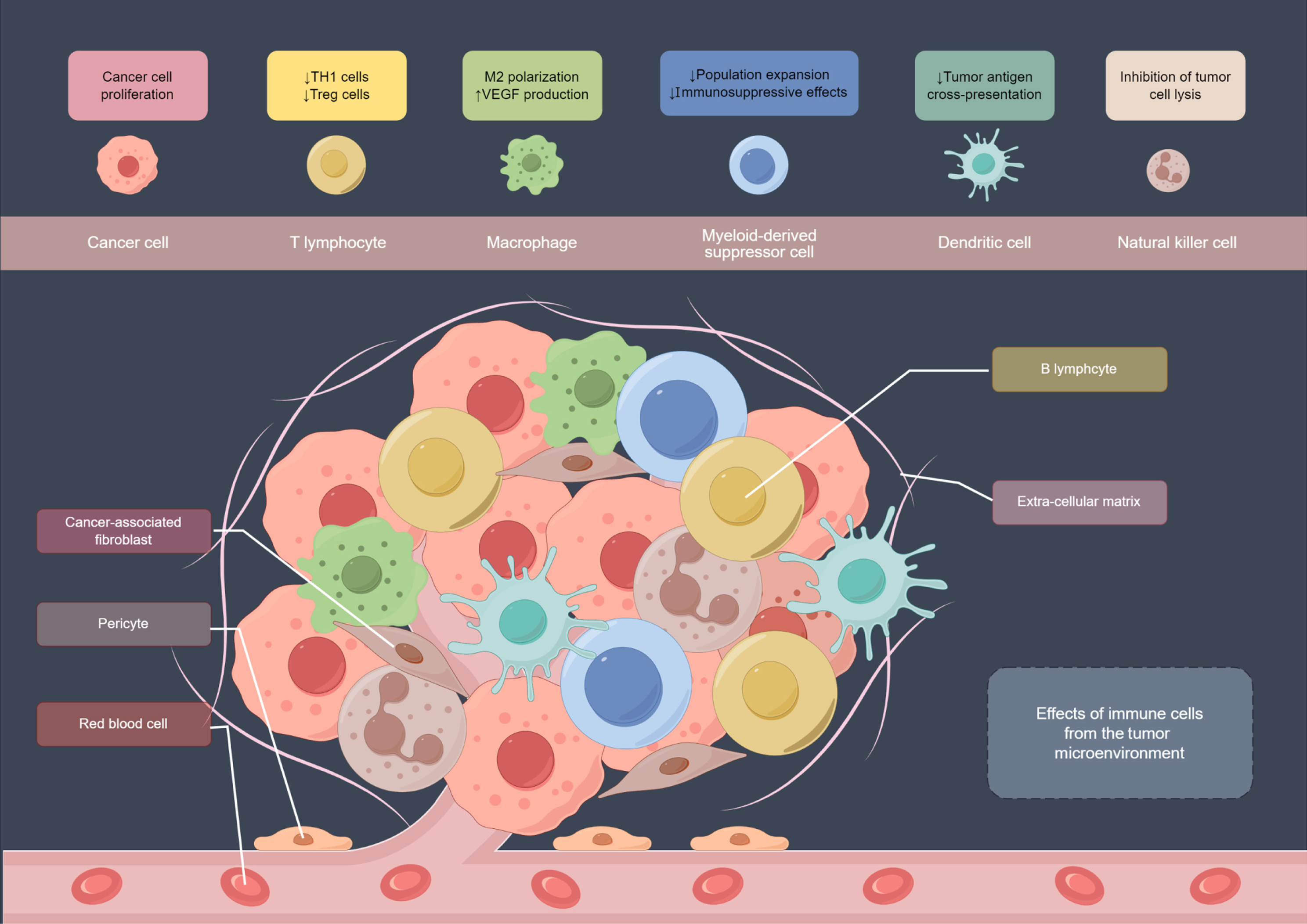
A conspicuous infiltration of T cells stands as a hallmark feature of CRC[23], yet their functional fate is dictated by a complex interplay with other immune constituents within the TME. Staging based on CD3+/CD8+ T cell density can provide better prognostic stratification compared with traditional staging, and it has the potential to become a promising independent prognostic biomarker[24]. The mechanisms that cause immunosuppression can be divided into three main areas: (1) Cell-cell interaction circuits: Intestinal epithelial-derived microvesicles deliver dual apoptotic signals to induce T cell death[25,26], while programmed death 1 (PD-1)-expressing epithelial cells subvert immunity through both CD28 co-stimulation blockade and Treg homeostasis maintenance[27,28]; (2) Metabolic cross-talk: The glycolytic switch in CRC cells elevates lactic acid to 10-30 mmol/L in TME[29]. Creating a paradoxical scenario where lactic acid simultaneously inhibits effector T cells/natural killer (NK) cells while fueling Treg immunosuppressive functions through HIF-1α-mediated metabolic adaptation[30]; and (3) Spatial reconfiguration: Recent spatial transcriptomics reveals M2-polarized tumor-associated macrophages (TAMs; CD163+, CD206+) preferentially accumulate at tumor-stroma interfaces, forming physical barriers that sequester CD8+ T cells from carcinoma nests[31-33]. The effects of immune cells from the TME indicated in Figure 2.
The above examples are intended to illustrate that the antitumor immune response and the immunosuppressive response in CRC TME is a complex process, and that synergistic or antagonistic effects between various immune cells may have a greater impact than those produced by a single immune cell, for example, moving away from an immune cell-centered immunotherapy mindset alone to targeting the entire TME on the basis of metabolic alterations in immune cells may be the key to breaking through immunosuppression. Next, we will start from the metabolic changes of the main immune cells, and gradually reveal the active or passive synergism and antagonism between immune cells, while exploring the game between immune cells and tumor cells in TME, with a view to providing new ideas for CRC immunotherapy.
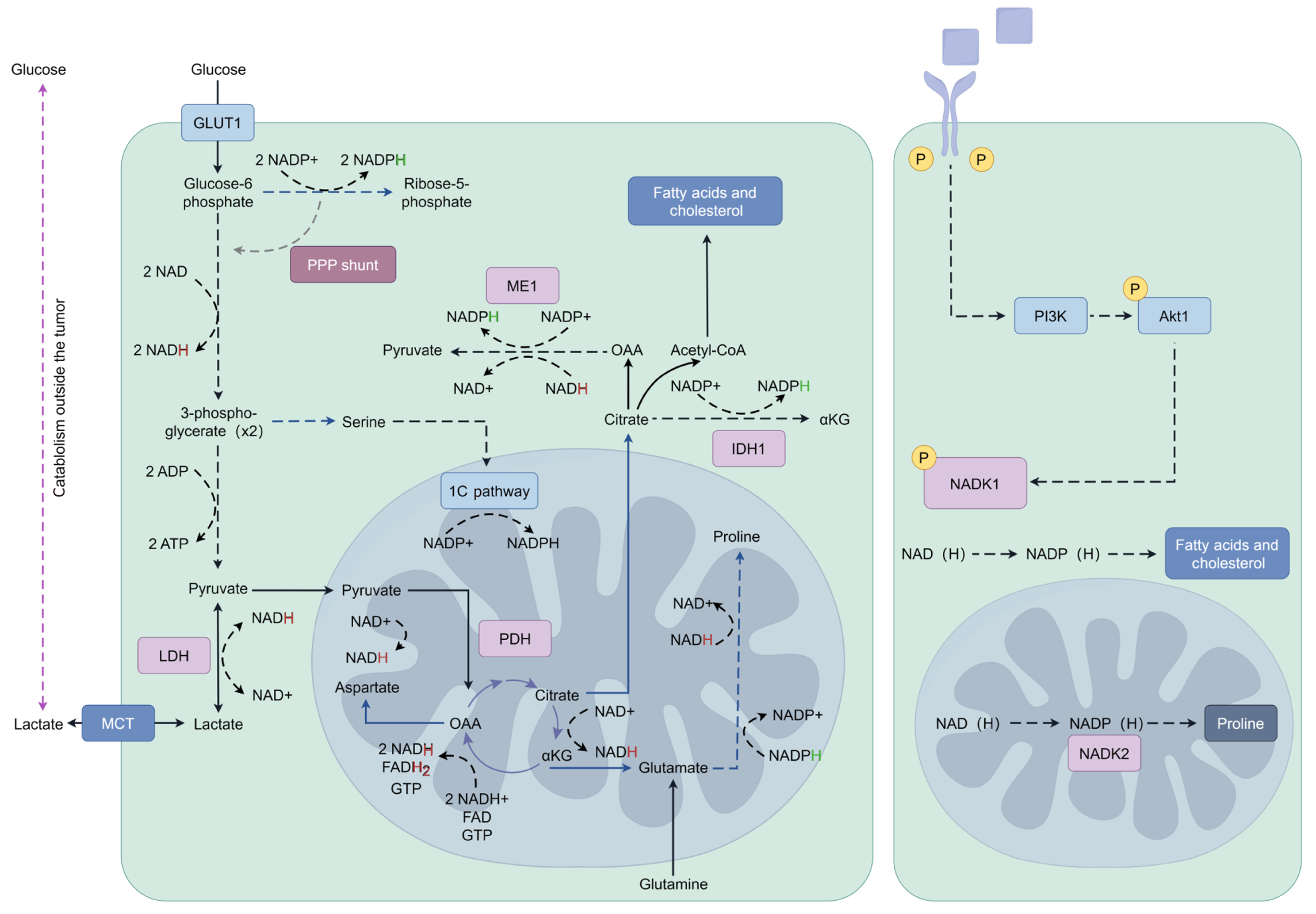
T-cells & B-cells: The metabolic plasticity of tumor-infiltrating CD8+ T cells is hijacked during CRC progression through multiple co-opted pathways. In early tumor stages, activated CD8+ T cells undergo Warburg-like glycolytic shifts[34] (Figure 3), characterized by PDK1-mediated diversion of pyruvate to lactate production rather than mitochondrial CoA generation, and this process, amplified by PI3K/AKT-driven GLUT1 trafficking[35-37]. Consequently, should the gly
Next, when the main energy substance glucose is gradually depleted, T cells shift to OXPHOS and fatty acid (FA) cat
By the discussion in the previous paragraph, we already know the main metabolic changes that occur in T cells in CRC TME, that is, one of the essences of the metabolic reprogramming phenomenon in CRC TME is the competition between CRC cells and immune cells for energy substances. This competition also exists among other nutrients that can provide energy, for example, amino acids. The metabolic competition between tumor-infiltrating lymphocytes (TILs) and CRC cells manifests as reciprocal fuel preferences: TILs predominantly utilize glycolysis, while CRC cells rely on glutamine metabolism—a dichotomy conserved across CRC subtypes[46]. Activated effector T cells require glutamine uptake to sustain proliferation, yet paradoxically, pharmacological glutaminase inhibition triggers compensatory glycolytic hyperactivation, compromising their long-term functionality[47]. While L-arginine supplementation enhances TIL cyt
Importantly, B cells amplify this metabolic warfare—their reprogramming toward NADPH-generating PPP and immunoregulatory lipid metabolism reshapes the TME to favor CRC immune evasion[51]. B cells deploy context-dependent metabolic programs in CRC. Upon activation, B cells increase glycolysis and OXPHOS, fueled by PI3K/mTORC2-mediated glucose/glutamine co-utilization[52,53]. However, chronic antigen exposure drives a metabolic shift towards leucine dependency—LARS B cells utilize LARS2 to stabilize mitochondrial NADH dehydrogenases, sustaining proliferation against tumor-derived oxidative stress[54,55]. This phenomenon existing in B cells further supports our view that paying attention to changes in the entire TME may be more important than focusing on changes in a single im
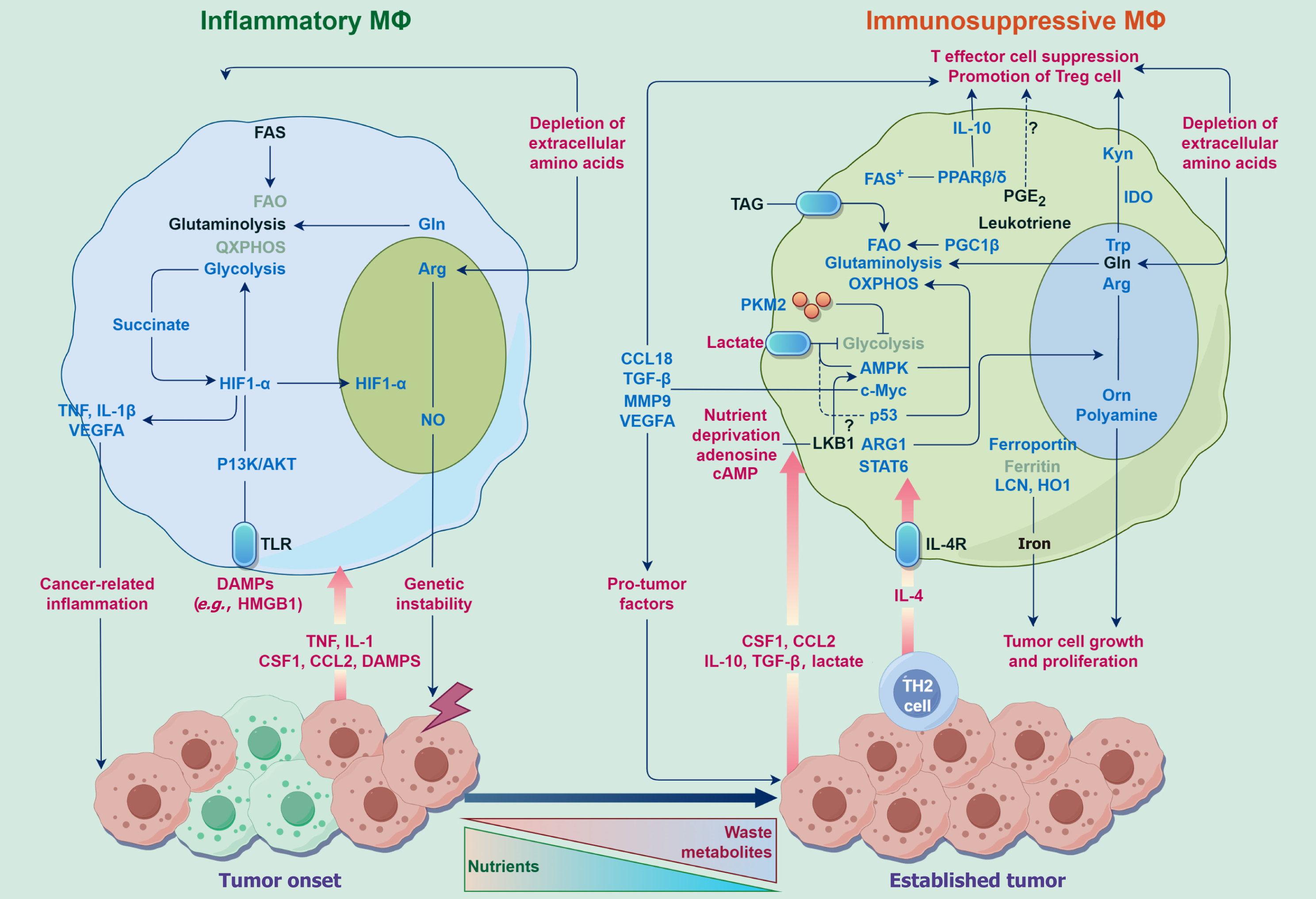
Macrophages: Based on distinct functional characteristics, it is possible to further categorize the predominant mac
When intestinal epithelial cells (IECs) are damaged, macrophages increase the metabolism of various amino acids, such as arginine, through the Tsc2/mTORC1 pathway to generate more energy, but accordingly, polyamine metabolites from amino acid metabolism further promote the proliferation of CRC cells[59], while the key transporter protein amino acid transporter LAT1 (also known as SLC7A5) promotes CRC progression by mediating increased leucine metabolism and inducing polarization of TAMs toward the M2 phenotype[60].
Regarding lipid metabolism reprogramming, the phenomenon of metabolic reprogramming in macrophages manifests in a more diverse manner. A clinical study involving 40 CRC patients demonstrated that GRP78, when incorporated into macrophage lipid droplets, enhances the protein stability of adipose triglyceride lipase by inhibiting ubiquitination. This process promotes the hydrolysis of triglycerides and the production of arachidonic acid and docosahexaenoic acid to maintain a tumor-immunosuppressive TME and also induce the polarization of macrophages towards the M2 phenotype, this phenomenon particularly pronounced in advanced stages of CRC[61]. In addition, contrary to T cells, the accumulation of cholesterol in CRC infiltrating macrophages facilitates their polarization towards the M1 phenotype[62]. This polarization enhances the activation of T cells by macrophages, thereby amplifying the anti-tumor effect. This repro
We have mentioned earlier that of equal interest to the altered metabolism of immune cells is the game between immune cells and tumor cells in the TME, and we would now like to add further to this idea by drawing on the dis
MDSCs: MDSCs renowned for their immunosuppressive effects on T cells. Under physiological conditions, bone marrow-derived multipotent hematopoietic stem cells have the capacity to differentiate into macrophages, dendritic cells, and granulocytes[66]. Conversely, the TME can suppress MDSCs differentiation, leading to the accumulation of a sig
During the differentiation and activation of MDSCs, its metabolic changes have many similarities to those mentioned earlier[69,70]. For example, activation of leucine-rich repeats, nucleotide-binding oligomeric structural domains, and pyrimidine-containing structural domain 3 inflammatory vesicles via the STAT1-NLRP3 axis, which in turn drives CRC progression[71,72]. On the contrary, CRC cells contribute to the generation of MDSCs. These MDSCs induced by CRC can inhibit T cell proliferation and facilitate the growth of CRC cells through cell-cell contact. However, by suppressing oxidative metabolism, including the production of NO and ROS, this suppression of T cells can be mitigated[72].
The upregulation of the scavenger receptor CD36 promotes the uptake of fats by MDSCs, shifting their primary energy source from glycolysis to FAO[73], in contrast, deletion of Mir4435-2hg leads to an increase in polymorphonuclear (PMN)-MDSCs and disrupts their FA metabolism to enhance their immunosuppressive capacity[74]. In light of this process, it is widely acknowledged in preclinical research that FA transport protein 2 acts as a regulatory factor in the immunosuppressive function of PMN-MDSCs, and inhibiting FA transport protein 2 can significantly enhance the efficacy of imm
Finally, there are changes in amino acid metabolism. Amino acid metabolism is included as a separate segment because this metabolic alteration in MDSCs is often accompanied by suppression of T cells. The metabolism of arginine and nitric oxide involved here has been introduced earlier[77,78], but a deficiency in 5-lipoxygenase reduces the activity of arginase 1 in MDSCs, thereby diminishing their capacity to suppress T cell functionality, and this reduction leads to a decrease in the occurrence of colonic polyps and CRC[79]. Similarly, this mechanism of impairing T cell functionality by depleting essential amino acids in T cells also occurs with amino acids such as tryptophan and cysteine[80]. MDSCs diminish tryptophan levels in the TME by relying on indoleamine-2,3-dioxygenase (IDO)-dependent tryptophan metabolism, depleting T cell nutrients, thereby inhibiting their normal immune activity[81]. It is an interesting phenomenon that in addition to the game between immune cells and tumor cells, there is also a mutual game between immune cells, which, although also mentioned in the previous section, appears to be particularly obvious in MDSCs and T cells as two mu
NK cells: NK cells exhibit rapid cytotoxic responses against tumor and virus-infected cells through granzyme/perforin-mediated killing and IFN-γ production[82]. Competitive uptake of cystine/glutamine by CRC cells leads to polyamine depletion in NK cells, and subsequent inhibition of c-Myc expression downregulates key enzymes of glycolysis by approximately 60%, ultimately leading to a reduction in the efficiency of perforin-mediated lysis of tumor cells to 39% ± 6% of baseline levels[83,84]. In patients with liver metastases of CRC, there is a significant increase in lactate levels within the TME. NK cells unable to regulate their internal pH consequently experience mitochondrial metabolic dysfunction, then leads to an excessive accumulation of ROS, ultimately resulting in mitochondrial stress and cell apoptosis[85]. Additionally, NK cell FA metabolism is enhanced to ensure their energy supply too. However, if NK cells continue to be stimulated, such as IL-15, they are eventually induced to generate PPARγ-driven lipid overload and endoplasmic reti
Specific conserved molecules on the surface of gut microbiota can be recognized by pattern recognition receptors (PRRs, such as Toll-like receptors TLRs and NOD-like receptors NLRs) on immune cells, thereby activating downstream signaling pathways and precisely regulating immune responses. Studies have found that certain intestinal bacteria (such as CAG-977) promote the transformation of effector T cells into phenotypes with more anti-tumor effects by activating specific signaling pathways[87]. F. nucleatum adheres to, invades, and induces the occurrence of CRC through its unique FadA adhesin. It can also directly bind to the TIGIT receptor on immune cells, inhibiting the killing function of NK cells and cytotoxic T cells[88,89]. For example, in diarrhea caused by E. coli, some regulatory factors can induce microorga
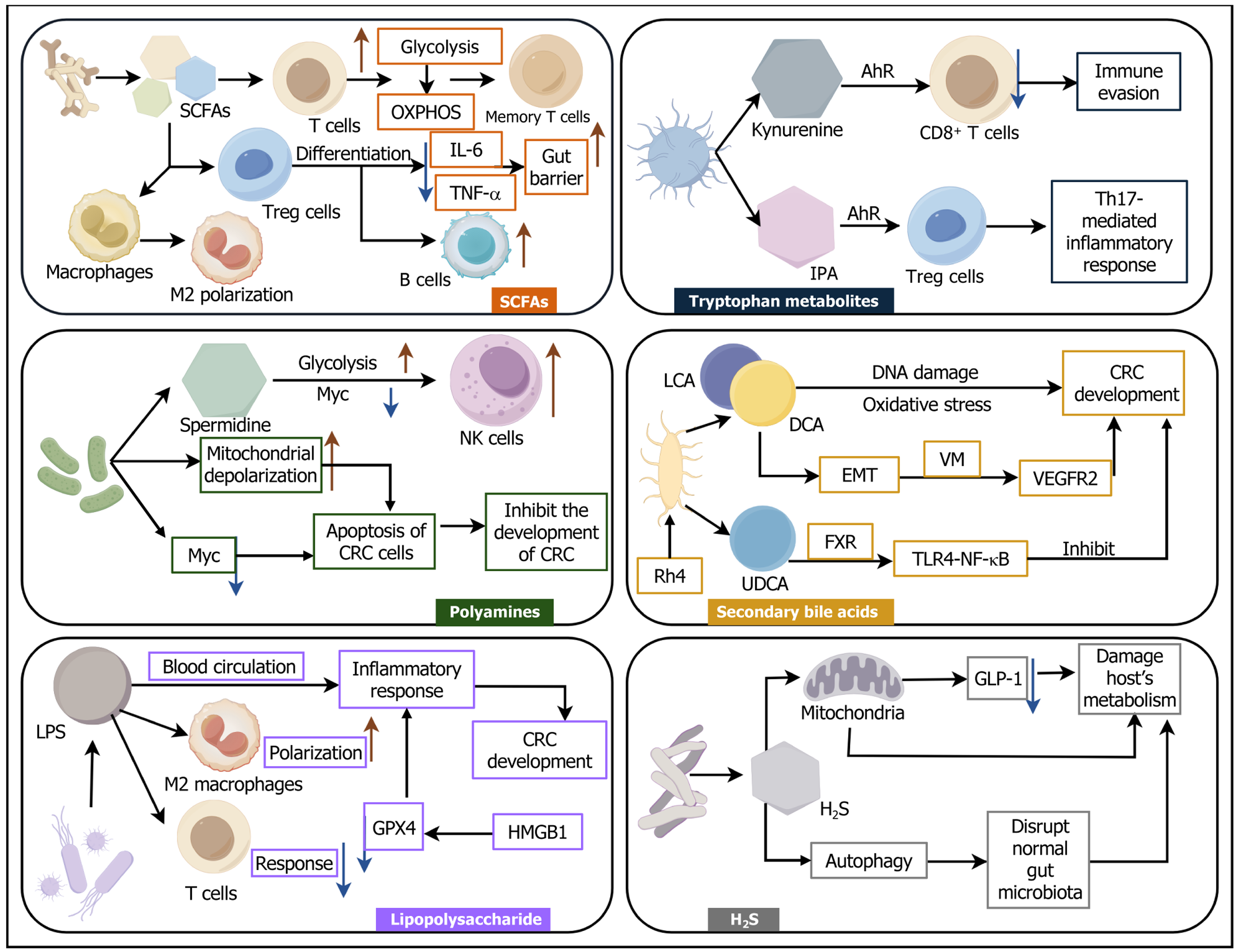
SCFAs: SCFAs, primarily acetate, propionate, and butyrate, are produced by gut bacteria such as Bacteroides, Prevotella, and Clostridium. In CRC, SCFAs modulate immune cell metabolism to exert anti-tumor effects. For instance, SCFAs activate the GPR41 receptor on CD8+ T cells, enhancing both their OXPHOS and glycolytic capacities. This metabolic reprogramming promotes CD8+ T cell proliferation, differentiation, and strengthens their anti-tumor immune response[91,92]. Furthermore, SCFAs promote FA uptake in these cells. Utilizing FA catalysts and glutamine, the cells shift their glucose metabolism towards OXPHOS and differentiate into memory T cells, sustaining long-term immunity[93,94]. Additionally, butyrate acts as a histone deacetylase (HDAC) inhibitor to promote regulatory T cell (Treg) differentiation, suppress pro-inflammatory factors [e.g., IL-6, tumor necrosis factor alpha (TNF-α)], and enhance intestinal barrier fun
Tryptophan metabolites: Tryptophan is metabolized by gut microbiota, including Bacteroides, Clostridium, and E. coli, into indoles and kynurenine[98]. Alterations in gut microbiota-mediated tryptophan metabolism are closely related to intestinal barrier function and may be involved in the occurrence and development of CRC[99]. These metabolites exert divergent immunomodulatory effects via the aryl hydrocarbon receptor (AhR). Indole derivatives, such as indolepropionic acid, activate AhR to promote Treg differentiation and suppress Th17-driven inflammation. In contrast, kyn
Polyamines: Polyamines, including putrescine, cadaverine, and spermidine, are primarily produced by gut microbiota such as Bacteroides, Clostridium, and Streptococcus[101]. Mouse studies indicate that polyamines help maintain gut homeostasis. Spermine oxidase deficiency reduces α-defensin production, leading to microbial dysbiosis and promoting the transition from colitis to CRC. Conversely, spermidine supplementation can restore homeostasis and prevent CRC development[102]. Moreover, in the context of CRC, spermidine supplementation can reactivate NK cells by alleviating c-Myc suppression and restoring glycolytic energy production, thereby restoring their tumor-killing activity[83]. Beyond their role in immune cell function, polyamines can also directly induce apoptosis in CRC cells by mechanisms that include inhibiting MYC synthesis and promoting mitochondrial depolarization.
Secondary bile acids: Secondary bile acids mainly include deoxycholic acid (DCA), lithocholic acid (LCA), etc., and are mainly derived from Firmicutes (such as Clostridium scindens). Studies have shown that the metabolic level of secondary bile acids in the gut microbiota is significantly increased in inflammatory bowel disease and CRC, and DCA and LCA are more carcinogenic than other secondary bile acids due to their properties of causing DNA damage and inducing oxi
Lipopolysaccharide: Lipopolysaccharides (LPS) are mainly derived from Gram-negative bacteria (such as E. coli, F. nucleatum), etc. LPS involved in blood circulation can promote systemic inflammation and affect the coagulation system, thereby leading to chronic inflammation and the occurrence of tumors[105]. The production of this inflammation is related to the negative regulation of glutathione peroxidase 4 by acetylated high mobility group box-1. Further studies have shown that LPS can regulate immune responses in the CRC TME, and this process is related to macrophage M2 polarization and inhibition of T cell responses[106]. In addition, LPS can also increase intestinal muscular inflammation by activating p38-MAPK, increasing the risk of intestinal obstruction in patients after CRC surgery[107]. It can thus be inferred that regulating inflammatory responses is also one of the pathways through which LPS promotes the deve
H2S: Hydrogen sulfide mainly originates from sulfate-reducing bacteria (such as Desulfovibrio) and the like. Studies have found that H2S derived from Desulfovibrio can inhibit mitochondrial respiration and induce the unfolded protein response in intestinal L cells, thereby hindering the secretion and gene expression of GLP-1 and further impairing the host's meta
Branched-chain amino acids: Branched-chain amino acids (BCAAs), namely leucine, isoleucine, and valine, are essential amino acids that serve as crucial nutrient signals and metabolic fuels in the CRC TME. While diet is the primary source, the gut microbiota also contributes to the host's BCAA pool through synthesis and degradation, thereby influencing systemic and local BCAA availability. Parabacteroides merdae can decompose BCAAs to produce three BCAA metabolites-isovaleric acid, 2-methylbutyric acid, and isobutyric acid. It can delay aging in mice through various health benefits such as reducing oxidative stress and inflammation, improving muscle capacity, reversing brain acetylcholine levels, and regulating blood glucose[111]. In addition, BCAA metabolites can also regulate metabolic disorders, thereby maintaining the homeostasis of gut microbiota and effectively preventing the occurrence of precancerous lesions such as inflammatory bowel disease[112].
Gut microbiota represented by F. nucleatum, enterotoxin-producing B. fragilis, and adherent-invasive E. coli can impair intestinal barrier function through various pathways. This causes a large number of bacteria and their metabolites to penetrate into the submucosa, which are then recognized by immune cells and continuously activate inflammatory pathways (such as the NF-κB signaling pathway)[113], and release pro-inflammatory factors, ultimately forming a chronic inflammatory environment that is conducive to the occurrence of CRC[114]. Preclinical studies have shown that F. nucleatum can release outer membrane vesicles, activate TLR4 and NF-κB, and promote the development of intestinal inflammation[115]. In CRC, F. nucleatum activates the TLR4/Myd88/NF-κB signaling pathway, transcriptionally activates the expression of miRNA-155-5p, thereby negatively regulating MSH6 and accelerating the malignant progression of CRC in mice[116]. In addition, enterotoxigenic B. fragilis disrupts the intestinal barrier by stimulating IECs to induce the activation of MMP-7 and transcription factors NF-κB and AP-1, and promoting the release of syndecan-2[117]. However, B. fragilis that can produce capsular polysaccharide A can promote the expression of anti-inflammatory IL-10, inhibit inflammation, and maintain the integrity of the intestinal barrier[118]. In patients with inflammatory bowel disease, the promoter of this type of capsular polysaccharide A is often in a closed state[119], which suggests the potential of targeting intestinal microorganisms for CRC treatment. Another type of microbiota that has been confirmed to play an important pro-inflammatory role is adherent-invasive E. coli. By virtue of its produced FimH adhesin and cell adhesion molecule 6, it activates the NF-κB pathway to promote the expression of a large number of pro-inflammatory cytokines such as TNF-α and chemokines such as IL-8, thereby inducing chronic inflammation. At the same time, it activates inflammasomes to induce the formation of IL-1β and IL-18, aggravating the inflammatory response[120,121], and ultimately promoting the occurrence and development of CRC.
First, the gut microbiota can directly suppress anti-tumor immunity. A representative mechanism is the inhibition of the killing ability of CD8+ T cells through the calcineurin-NFAT axis[122]. For example, the metabolite secondary bile acid inhibits this pathway by targeting the plasma membrane Ca2+ ATPase, thereby achieving immunosuppression[123]. Secondly, the gut microbiota can recruit myeloid-derived immunosuppressive cells or induce M2 polarization of macrophages to achieve immunosuppression. For example, the gut microbiota metabolite 4-hydroxyphenylacetic acid upregulates the transcription of CXCL3 by activating the JAK2/STAT3 pathway, thereby recruiting PMN-MDSCs into the CRC TME, inhibiting the anti-tumor responses of cells such as CD8+ T cells, and promoting the progression of CRC[124], In metastatic CRC, the intestinal microbiota induces the secretion of cathepsin K, and promotes the metastasis of CRC by mediating the polarization of TLR4-dependent M2 macrophages[125]. Third, the gut microbiota competes with immune cells for nutrients, directly or indirectly participating in the shaping of an immunosuppressive TME. For example, as mentioned earlier, the gut microbiota can consume tryptophan and produce kynurenine, which directly or indirectly inhibits T cell proliferation while promoting the function of Tregs cells. In addition, different microbial metabolites can also regulate the direction of T cell differentiation by inhibiting the activation of antigen-presenting cells (APCs). For instance, SCFAs and secondary bile acids inhibit APC activation and reduce the differentiation of inflammatory T cells (Th1/Th17); whereas trimethylamine N-oxide and others promote APC activation and enhance the anti-tumor immunity of T cells[126]. Finally, the gut microbiota can also affect the CRC TME by acting on stromal cells. For example, F. nucleatum attaches to and invades cancer-associated fibroblasts (CAFs) in CRC, causing them to exhibit an inflammatory CAF phenotype. This phenotype is characterized by the secretion of pro-inflammatory cytokines (such as CXCL1, IL-6, and IL-8), the production of ROS, and enhanced metabolic activity. This transformation makes CRC cells more prone to proliferation and metastasis[127]. In addition, in young CRC patients, researchers have observed that actinomycetes abnormally reside in CAFs, thereby activating the TLR2/NF-κB pathway and reducing the infiltration of CD8+ T lym
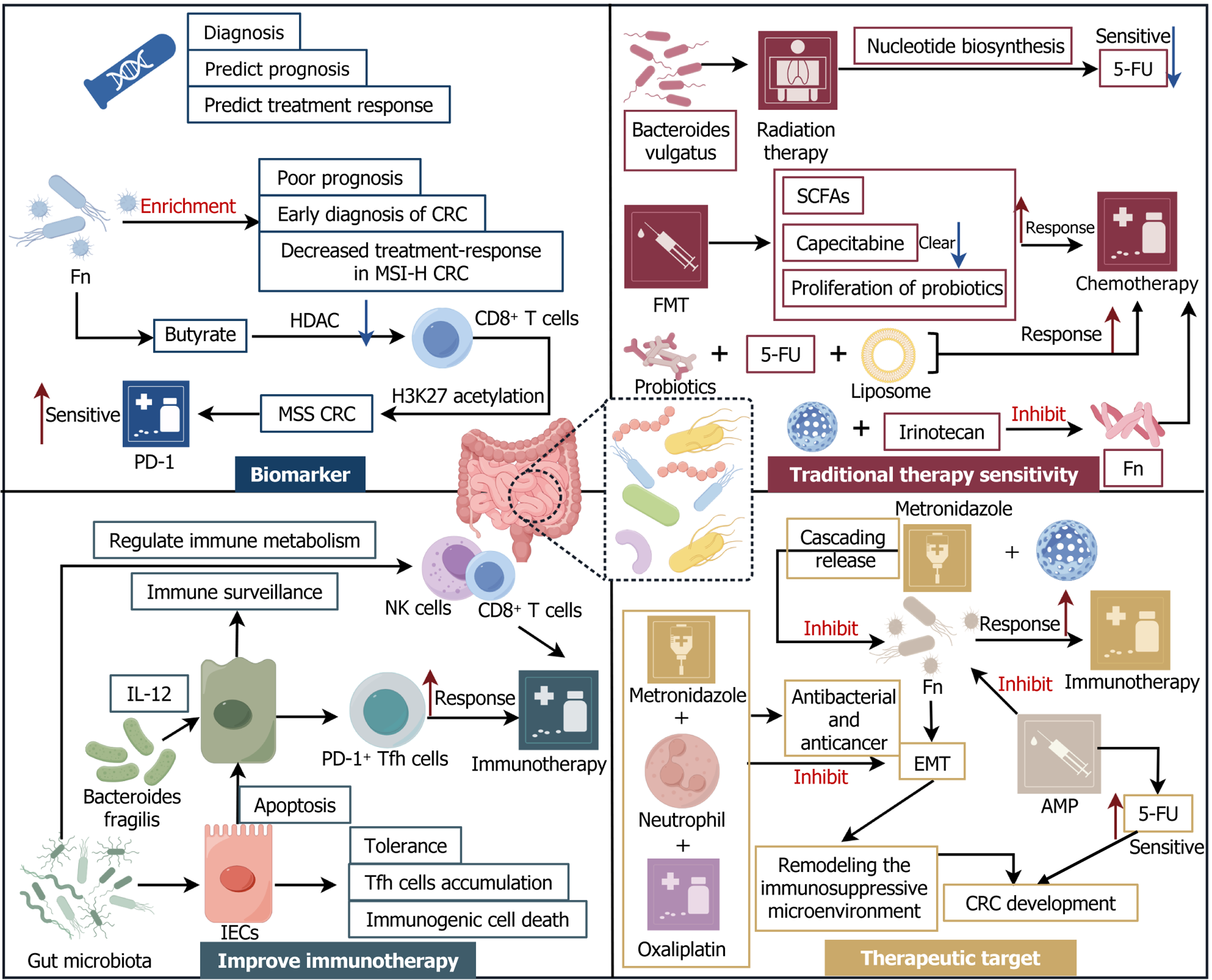
It has been confirmed that the enrichment of certain gut microbiota is significantly associated with the early diagnosis of CRC, treatment response, and prognostic levels. F. nucleatum is one of the representative microbiota[129]. A clinical study has shown that F. nucleatum, especially the F. nucleatum a C2 subgroup, dominates the TME of CRC, and its enrichment level is significantly correlated with the early diagnosis and prognosis of CRC[130]. An increase in the enrichment level often indicates a decrease in the response of CRC to treatment and a poor prognosis. However, in the immunotherapy of patients with microsatellite-stable CRC, researchers have observed the opposite phenomenon. The study found that butyrate, a metabolite of F. nucleatum, inhibited HDAC 3/8 in CD8+ T cells, induced H3K27 acetylation and expression of the Tbx21 promoter, and instead made MSS CRC more sensitive to anti-PD-1 therapy[131]. This indicates that the regulation of gut microbiota on CRC is not static; even the same microbiota can play different roles in different stages of CRC. Regarding the toxic and side effects after chemotherapy, clinical studies on the toxicity and side effects after chemo
Although treatment methods such as immunotherapy and targeted therapy are developing rapidly, radiotherapy and chemotherapy, as traditional therapies, still occupy an important position in the treatment of CRC. This is particularly true for CRC patients who require surgery, where adjuvant radiotherapy and chemotherapy before and after surgery play an undeniable positive role in the surgery and postoperative expectations. Researchers conducted 16S rRNA sequencing on fecal samples from patients with locally advanced rectal cancer and found that the diversity of gut microbiota decreased after preoperative neoadjuvant radiotherapy and chemotherapy. Take Bacteroides vulgatus as an example: The nucleotide biosynthesis mediated by it reduces the sensitivity of CRC cells to 5-fluorouracil or radiotherapy[134]. In addition, the gut microbiota has also been found to be closely related to the abscopal effect in radiotherapy[135], and fecal microbiota transplantation technology has been confirmed by clinical research to reduce radiotherapy-related toxicity and improve its efficacy[136]. Other preclinical studies have shown that, by artificially intervening to regulate the gut microbiota, for example, using nanoparticles constructed with the prebioticxylan-stearic acid conjugate to delay the clearance of capecitabine in the blood, while promoting the proliferation of probiotics and the production of SCFAs, the anti-tumor immune response and chemotherapy response are ultimately enhanced, increasing the tumor inhibition rate from 5.29% to 71.78% and significantly prolonging the survival period of mice with CRC[137]. Similarly, the chemo
As mentioned earlier, researchers have confirmed that the efficacy of drugs such as anti-PD-1/programmed death ligand-1 (PD-L1) can be improved by altering the composition of the gut microbiota in CRC[140]. For example, studies have shown that in CRC patients, Prevotellaceae can regulate the immune TME, and its abundance is significantly positively correlated with the therapeutic effect of anti-PD-L1 therapy[141]. Another preclinical study found that the ileal microbiota determines the tolerance and immunogenic cell death of ileal IECs, as well as the accumulation of follicular helper T cells (TFH). Inhibiting the apoptosis of IECs impairs chemotherapy-induced immune surveillance against colon cancer. However, the colonization of B. fragilis and Erysipelotrichaceae in the ileum enables apoptotic ileal IECs to induce the production of PD-1+ TFH cells in an IL-1R1 and IL-12-dependent manner, thereby enhancing the efficacy of chem
This treatment method includes two directions: Positive and negative. First, it promotes the stability of “good” gut microbiota. For example, ENPP2 inhibitors restore the intestinal barrier by increasing the expression of tight junction proteins claudin-1, occludin, and ZO-1, reduce the polarization of M2 TAMs and colonic inflammation, and alleviate gut microbiota imbalance in mice, thereby inhibiting the proliferation of CRC cells[147,148]. In addition, some clinical studies have confirmed that, gut microbiota imbalance has been confirmed to be associated with the development of colorectal tumors related to a high-fat diet, while berberine significantly improves intestinal barrier damage and inhibits colonic inflammation and related carcinogenic pathways, providing a promising microbiota-regulating therapeutic strategy for the clinical prevention and treatment of Western diet-related CRC. In contrast to the above, for example, as mentioned earlier, F. nucleatum causes immunotherapy resistance through its metabolite succinate. In such cases, the abundance of F. nucleatum can be reduced by directly using antibiotics such as metronidazole to improve immunotherapy[144], nanogels modified by nanotechnology can combat F. nucleatum-infected CRC through the cascade release of metronidazole and chemotherapeutic drugs, further enhancing the therapeutic effect[149]. In addition, neutrophil-mimicking nanomedicines loaded with metronidazole and oxaliplatin can simultaneously achieve antibacterial and anticancer effects. They can reverse the EMT in the process of F. nucleatum-induced tumor cell metastasis, reshape the immunosuppressive TME, thereby inhibiting the development of CRC and liver metastasis, and minimizing damage to the normal symbiotic microbiota[150]. It can be seen from this that the combination of nanotechnology and gut microbiota may become a new therapeutic approach for CRC. Similarly, antimicrobial peptides represent a new class of antibacterial drugs. The halogenated derivative Jelleine-I, Br- Jelleine-I shows significant anti-F. nucleatum activity. At the same time, Br-JI may target membrane-associated FadA to disrupt the F. nucleatum membrane. Compared with metronidazole, it has a stronger ability to kill F. nucleatum and is sensitive to the anticancer effect of the chemotherapy drug 5-fluorouracil, which may become a new strategy for the treatment of CRC[151]. This part is shown in Figure 6 and Table 1 respectively.
| Drug | Targets | Clinical phase | NCT | Sponsored by | Status | Purpose | |
| ICIs | Relatlimab | LAG-3 | Phase 2 | NCT03642067 | Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins | Completed | The purpose of this study is to evaluate the safety and clinical activity of nivolumab and relatlimab in patients with metastatic or locally advanced MSS colorectal cancer |
| MK-4830 | ILT-4 | Phase 2 | NCT04895722 | Merck Sharp & Dohme LLC | Active, not recruiting | Evaluation of co-formulated pembrolizumab/quavonlimab (MK-1308A) vs other treatments in participants with MSI-H or dMMR stage IV CRC (MK-1308A-008/KEYSTEP-008) | |
| Tucatinib | HER2 | Phase 3 | NCT05253651 | Seagen, a wholly owned subsidiary of Pfizer | Recruiting | A study of tucatinib with trastuzumab and mFOLFOX6 vs standard of care treatment in first-line HER2+ metastatic CRC (MOUNTAINEER-03) | |
| B1962 | VEGF/PD-L1 | Phase 2 | NCT04296019 | Tasly Biopharmaceuticals Co., Ltd. | Not yet recruiting | A multicenter, open phase IIb clinical study to evaluate the efficacy and safety of B1962 injection in the treatment of advanced CRC | |
| MGD007 | GPA33 | Phase 1 | NCT02248805 | MacroGenics | Completed | Phase 1 study of MGD007 in relapsed/refractory metastatic colorectal carcinoma | |
| Belzutifan | HIF-2α | Phase 2 | NCT04976634 | Merck Sharp & Dohme LLC | Active, not recruiting | Pembrolizumab plus lenvatinib in combination with belzutifan in solid tumors (MK-6482-016) | |
| PF-07062119 | GUCY2c/CD3 | Phase 1 | NCT04171141 | Pfizer | Terminated | Study to test the safety and tolerability of PF-07062119 in patients with selected advanced or metastatic gastrointestinal tumors | |
| BMS-986340 | CCR8 | Phase 1 and phase 2 | NCT04895709 | Bristol-Myers Squibb | Recruiting | A study of BMS-986340 as monotherapy and in combination with nivolumab or docetaxel in participants with advanced solid tumors | |
| Sintilimab | PD-1 | Phase 3 | NCT05236972 | Sun Yat-sen University | Recruiting | PACE: PD-1 antibody for dMMR/MSI-H stage III CRC | |
| Avelumab | PD-L1 | Phase 2 | NCT03854799 | Gruppo Oncologico del Nord-Ovest | Completed | Immunotherapy in locally advanced rectal cancer (AVANA) | |
| Tislelizumab | PD-1 | Phase 1 and phase 2 | NCT04577963 | Hutchison Medipharma Limited | Terminated | A study of fruquintinib in combination with tislelizumab in advanced solid tumors | |
| Pembrolizumab | PD-1 | Phase 2 | NCT05131919 | The Netherlands Cancer Institute | Active, not recruiting | Pembrolizumab for locally advanced, irresectable, non-metastatic dMMR CRCs (PUMA) | |
| Nivolumab | PD-1 | Phase 2 | NCT03642067 | Sidney Kimmel Comprehensive Cancer Center at Johns Hopkins | Completed | The purpose of this study is to evaluate the safety and clinical activity of nivolumab and relatlimab in patients with metastatic or locally advanced MSS CRC | |
| HFB200603 | BTLA | Phase 1 | NCT05789069 | HiFiBiO Therapeutics | Recruiting | A study of HFB200603 as a single agent and in combination with tislelizumab in adult patients with advanced solid tumors | |
| MK0457 | pan-Aurora | Phase 1 | NCT00099346 | Merck Sharp & Dohme LLC | Terminated | MK0457 (an aurora kinase inhibitor) study in patients with advanced CRC and other advanced solid tumors (0457-001) | |
| HRO761 | WRN | Phase 1 | NCT05838768 | Novartis Pharmaceuticals | Recruiting | Study of HRO761 alone or in combination in cancer patients with specific DNA alterations called microsatellite instability or mismatch repair deficiency | |
| Metabolic inhibitor | Epacadostat | IDO1 | Phase 1 and phase 2 | NCT03516708 | Washington University School of Medicine | Recruiting | Epacadostat (INCB024360) added to preoperative chemoradiation in patients with locally advanced rectal cancer |
| Ivosidenib | IDH1 | Phase 2 | NCT04195555 | National Cancer Institute | Active, not recruiting | Ivosidenib in treating patients with advanced solid tumors, lymphoma, or histiocytic disorders with IDH1 mutations (a pediatric MATCH treatment trial) | |
| Metformin | Mitochondria | Phase 3 | NCT05921942 | Ain Shams University | Completed | The impact of metformin administration on the clinical outcome of stage IV colon cancer | |
| Telaglenastat | GLS | Phase 1 and phase 2 | NCT02861300 | David Bajor, MD | Completed | CB-839 + capecitabine in solid tumors and fluoropyrimidine resistant PIK3CA mutant CRC | |
| Gut microbiota | Gut microbiota | / | Observational | NCT02845973 | Shanghai Jiao Tong University School of Medicine | Completed | Study of fecal bacteria in early diagnosis of CRC |
| Gut microbiota | / | Observational | NCT06738173 | Gianluca Ianiro, Catholic University of the Sacred Heart (Responsible Party) | Recruiting | Microbiome-based diagnostic tool for the screening of CRC (GUILTI) | |
| Gut microbiota | / | Observational | NCT06588166 | Fondazione Policlinico Universitario Agostino Gemelli Irccs | Recruiting | Microbiome testing for the screening of CRC (NI-GUILTI) | |
| Gut microbiota | / | Observational | NCT04291755 | Persephone Biosciences (Responsible Party) | Recruiting | Development and analysis of a stool bank for cancer patients | |
| Gut microbiota | Observational (patient registry) | NCT07154173 | Sixth Affiliated Hospital, Sun Yat-sen University | Recruiting | Comprehensive analysis of gut microbiota signatures in metastatic CRC | ||
| Fusobacterium nucleatum | Interventional | NCT05945082 | Malmö University | Recruiting | Fusobacterium nucleatum at CRC sites | ||
| EPEC | Observational (patient registry) | NCT02373020 | Mansoura University | Recruiting | EPEC: Does it have a role in colorectal tumourigenesis (EPEC)? |
Although we have made every effort to conduct a systematic review on this topic, there are still several limitations. First, due to space constraints, many studies have inevitably not been included in the analysis, which may fail to fully reflect the complexity of human CRC. Second, although we have revealed the association between gut microbiota-mediated immunometabolism and the TME, many of their exact causal mechanisms remain to be elucidated. Finally, many of the possible clinical translation pathways mentioned in the article have not been further verified, and the efficiency and feasibility of their clinical translation still need further research and improvement.
In recent years, the relationship between the intestinal microbiota and CRC has attracted much attention. Studies have confirmed that the gut microbiota significantly regulates the immunometabolism of CRC through its metabolites, thereby affecting the TME and its progression. This finding provides a new perspective for us to understand the role of the gut microbiota in cancer development. Immunometabolic reprogramming is considered an important mechanism of tumor immune escape and therapeutic resistance, and the gut microbiota plays a key role in this process, which provides us with new targets. In addition, by understanding the characteristics of an individual’s gut microbiota, we can develop more targeted prevention and treatment strategies to improve patient prognosis. Research in this direction will promote the application of gut microbiota in cancer and ultimately achieve the goal of personalized medicine. However, there are still many complex issues. The role of specific microorganisms is often context-dependent, and the inconsistencies in human cohort studies - which may stem from dietary, geographical, and host genetic factors - highlight the challenges in identifying universal microbiota biomarkers. Furthermore, some preclinical studies have not been clinically validated, and the feasibility and efficiency of their clinical translation remain unknown. Emerging technologies represented by artificial intelligence and nanotechnology have provided many conveniences for our clinical treatment, but their technical limitations still need to be overcome, so there is still a long way to go before clinical application.
The authors would like to express their sincere gratitude to all colleagues and collaborators who contributed valuable insights and support throughout the development of this manuscript. Special appreciation is extended to the researchers whose studies laid the foundation for our discussion. Finally, we acknowledge the constructive feedback from reviewers, which greatly improved the quality of this paper.
| 1. | Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71:209-249. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76817] [Cited by in RCA: 69604] [Article Influence: 13920.8] [Reference Citation Analysis (45)] |
| 2. | Siegel RL, Wagle NS, Cercek A, Smith RA, Jemal A. Colorectal cancer statistics, 2023. CA Cancer J Clin. 2023;73:233-254. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2171] [Cited by in RCA: 1959] [Article Influence: 653.0] [Reference Citation Analysis (2)] |
| 3. | Weng J, Li S, Zhu Z, Liu Q, Zhang R, Yang Y, Li X. Exploring immunotherapy in colorectal cancer. J Hematol Oncol. 2022;15:95. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 295] [Cited by in RCA: 260] [Article Influence: 65.0] [Reference Citation Analysis (4)] |
| 4. | André T, de Gramont A, Vernerey D, Chibaudel B, Bonnetain F, Tijeras-Raballand A, Scriva A, Hickish T, Tabernero J, Van Laethem JL, Banzi M, Maartense E, Shmueli E, Carlsson GU, Scheithauer W, Papamichael D, Möehler M, Landolfi S, Demetter P, Colote S, Tournigand C, Louvet C, Duval A, Fléjou JF, de Gramont A. Adjuvant Fluorouracil, Leucovorin, and Oxaliplatin in Stage II to III Colon Cancer: Updated 10-Year Survival and Outcomes According to BRAF Mutation and Mismatch Repair Status of the MOSAIC Study. J Clin Oncol. 2015;33:4176-4187. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 581] [Cited by in RCA: 519] [Article Influence: 47.2] [Reference Citation Analysis (3)] |
| 5. | Vesely MD, Zhang T, Chen L. Resistance Mechanisms to Anti-PD Cancer Immunotherapy. Annu Rev Immunol. 2022;40:45-74. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 243] [Cited by in RCA: 347] [Article Influence: 86.8] [Reference Citation Analysis (7)] |
| 6. | Ganesh K. Optimizing immunotherapy for colorectal cancer. Nat Rev Gastroenterol Hepatol. 2022;19:93-94. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 96] [Cited by in RCA: 88] [Article Influence: 22.0] [Reference Citation Analysis (7)] |
| 7. | Endesfelder D, Burrell R, Kanu N, McGranahan N, Howell M, Parker PJ, Downward J, Swanton C, Kschischo M. Chromosomal instability selects gene copy-number variants encoding core regulators of proliferation in ER+ breast cancer. Cancer Res. 2014;74:4853-4863. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 64] [Cited by in RCA: 62] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 8. | Sedlak JC, Yilmaz ÖH, Roper J. Metabolism and Colorectal Cancer. Annu Rev Pathol. 2023;18:467-492. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 115] [Article Influence: 38.3] [Reference Citation Analysis (0)] |
| 9. | Yi M, Li J, Chen S, Cai J, Ban Y, Peng Q, Zhou Y, Zeng Z, Peng S, Li X, Xiong W, Li G, Xiang B. Emerging role of lipid metabolism alterations in Cancer stem cells. J Exp Clin Cancer Res. 2018;37:118. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 178] [Cited by in RCA: 186] [Article Influence: 23.3] [Reference Citation Analysis (5)] |
| 10. | WARBURG O. On the origin of cancer cells. Science. 1956;123:309-314. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11238] [Cited by in RCA: 10100] [Article Influence: 144.3] [Reference Citation Analysis (6)] |
| 11. | Liu Q, Luo Q, Halim A, Song G. Targeting lipid metabolism of cancer cells: A promising therapeutic strategy for cancer. Cancer Lett. 2017;401:39-45. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 322] [Cited by in RCA: 272] [Article Influence: 30.2] [Reference Citation Analysis (0)] |
| 12. | Dai W, Xiang W, Han L, Yuan Z, Wang R, Ma Y, Yang Y, Cai S, Xu Y, Mo S, Li Q, Cai G. PTPRO represses colorectal cancer tumorigenesis and progression by reprogramming fatty acid metabolism. Cancer Commun (Lond). 2022;42:848-867. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 61] [Cited by in RCA: 84] [Article Influence: 21.0] [Reference Citation Analysis (4)] |
| 13. | Leone RD, Powell JD. Metabolism of immune cells in cancer. Nat Rev Cancer. 2020;20:516-531. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 400] [Cited by in RCA: 636] [Article Influence: 106.0] [Reference Citation Analysis (4)] |
| 14. | Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646-674. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55210] [Cited by in RCA: 48556] [Article Influence: 3237.1] [Reference Citation Analysis (11)] |
| 15. | Galon J, Costes A, Sanchez-Cabo F, Kirilovsky A, Mlecnik B, Lagorce-Pagès C, Tosolini M, Camus M, Berger A, Wind P, Zinzindohoué F, Bruneval P, Cugnenc PH, Trajanoski Z, Fridman WH, Pagès F. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science. 2006;313:1960-1964. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5379] [Cited by in RCA: 4984] [Article Influence: 249.2] [Reference Citation Analysis (5)] |
| 16. | Blank CU, Haining WN, Held W, Hogan PG, Kallies A, Lugli E, Lynn RC, Philip M, Rao A, Restifo NP, Schietinger A, Schumacher TN, Schwartzberg PL, Sharpe AH, Speiser DE, Wherry EJ, Youngblood BA, Zehn D. Defining 'T cell exhaustion'. Nat Rev Immunol. 2019;19:665-674. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1395] [Cited by in RCA: 1202] [Article Influence: 171.7] [Reference Citation Analysis (4)] |
| 17. | Miller BC, Sen DR, Al Abosy R, Bi K, Virkud YV, LaFleur MW, Yates KB, Lako A, Felt K, Naik GS, Manos M, Gjini E, Kuchroo JR, Ishizuka JJ, Collier JL, Griffin GK, Maleri S, Comstock DE, Weiss SA, Brown FD, Panda A, Zimmer MD, Manguso RT, Hodi FS, Rodig SJ, Sharpe AH, Haining WN. Subsets of exhausted CD8(+) T cells differentially mediate tumor control and respond to checkpoint blockade. Nat Immunol. 2019;20:326-336. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1839] [Cited by in RCA: 1639] [Article Influence: 234.1] [Reference Citation Analysis (4)] |
| 18. | Bell HN, Huber AK, Singhal R, Korimerla N, Rebernick RJ, Kumar R, El-Derany MO, Sajjakulnukit P, Das NK, Kerk SA, Solanki S, James JG, Kim D, Zhang L, Chen B, Mehra R, Frankel TL, Győrffy B, Fearon ER, Pasca di Magliano M, Gonzalez FJ, Banerjee R, Wahl DR, Lyssiotis CA, Green M, Shah YM. Microenvironmental ammonia enhances T cell exhaustion in colorectal cancer. Cell Metab. 2023;35:134-149. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 109] [Cited by in RCA: 140] [Article Influence: 46.7] [Reference Citation Analysis (0)] |
| 19. | Li J, Guo Y, Liu J, Guo F, Du L, Yang Y, Li X, Ma Y. Depicting the landscape of gut microbial-metabolic interaction and microbial-host immune heterogeneity in deficient and proficient DNA mismatch repair colorectal cancers. J Immunother Cancer. 2023;11:e007420. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 10] [Reference Citation Analysis (0)] |
| 20. | Wong SH, Yu J. Gut microbiota in colorectal cancer: mechanisms of action and clinical applications. Nat Rev Gastroenterol Hepatol. 2019;16:690-704. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1100] [Cited by in RCA: 927] [Article Influence: 132.4] [Reference Citation Analysis (3)] |
| 21. | Shakhpazyan NK, Mikhaleva LM, Bedzhanyan AL, Gioeva ZV, Mikhalev AI, Midiber KY, Pechnikova VV, Biryukov AE. Exploring the Role of the Gut Microbiota in Modulating Colorectal Cancer Immunity. Cells. 2024;13:1437. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 22] [Reference Citation Analysis (0)] |
| 22. | Bai X, Duan Z, Deng J, Zhang Z, Fu R, Zhu C, Fan D. Ginsenoside Rh4 inhibits colorectal cancer via the modulation of gut microbiota-mediated bile acid metabolism. J Adv Res. 2025;72:37-52. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 47] [Cited by in RCA: 39] [Article Influence: 39.0] [Reference Citation Analysis (0)] |
| 23. | Girardin A, McCall J, Black MA, Edwards F, Phillips V, Taylor ES, Reeve AE, Kemp RA. Inflammatory and regulatory T cells contribute to a unique immune microenvironment in tumor tissue of colorectal cancer patients. Int J Cancer. 2013;132:1842-1850. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 29] [Article Influence: 2.1] [Reference Citation Analysis (3)] |
| 24. | Galon J, Mlecnik B, Bindea G, Angell HK, Berger A, Lagorce C, Lugli A, Zlobec I, Hartmann A, Bifulco C, Nagtegaal ID, Palmqvist R, Masucci GV, Botti G, Tatangelo F, Delrio P, Maio M, Laghi L, Grizzi F, Asslaber M, D'Arrigo C, Vidal-Vanaclocha F, Zavadova E, Chouchane L, Ohashi PS, Hafezi-Bakhtiari S, Wouters BG, Roehrl M, Nguyen L, Kawakami Y, Hazama S, Okuno K, Ogino S, Gibbs P, Waring P, Sato N, Torigoe T, Itoh K, Patel PS, Shukla SN, Wang Y, Kopetz S, Sinicrope FA, Scripcariu V, Ascierto PA, Marincola FM, Fox BA, Pagès F. Towards the introduction of the 'Immunoscore' in the classification of malignant tumours. J Pathol. 2014;232:199-209. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1181] [Cited by in RCA: 1097] [Article Influence: 91.4] [Reference Citation Analysis (5)] |
| 25. | O'Connell J, O'Sullivan GC, Collins JK, Shanahan F. The Fas counterattack: Fas-mediated T cell killing by colon cancer cells expressing Fas ligand. J Exp Med. 1996;184:1075-1082. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 652] [Cited by in RCA: 623] [Article Influence: 20.8] [Reference Citation Analysis (3)] |
| 26. | Huber V, Fais S, Iero M, Lugini L, Canese P, Squarcina P, Zaccheddu A, Colone M, Arancia G, Gentile M, Seregni E, Valenti R, Ballabio G, Belli F, Leo E, Parmiani G, Rivoltini L. Human colorectal cancer cells induce T-cell death through release of proapoptotic microvesicles: role in immune escape. Gastroenterology. 2005;128:1796-1804. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 434] [Cited by in RCA: 411] [Article Influence: 19.6] [Reference Citation Analysis (0)] |
| 27. | Kamphorst AO, Wieland A, Nasti T, Yang S, Zhang R, Barber DL, Konieczny BT, Daugherty CZ, Koenig L, Yu K, Sica GL, Sharpe AH, Freeman GJ, Blazar BR, Turka LA, Owonikoko TK, Pillai RN, Ramalingam SS, Araki K, Ahmed R. Rescue of exhausted CD8 T cells by PD-1-targeted therapies is CD28-dependent. Science. 2017;355:1423-1427. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 758] [Cited by in RCA: 817] [Article Influence: 90.8] [Reference Citation Analysis (0)] |
| 28. | Tanaka A, Sakaguchi S. Regulatory T cells in cancer immunotherapy. Cell Res. 2017;27:109-118. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1587] [Cited by in RCA: 1465] [Article Influence: 162.8] [Reference Citation Analysis (6)] |
| 29. | Fischer K, Hoffmann P, Voelkl S, Meidenbauer N, Ammer J, Edinger M, Gottfried E, Schwarz S, Rothe G, Hoves S, Renner K, Timischl B, Mackensen A, Kunz-Schughart L, Andreesen R, Krause SW, Kreutz M. Inhibitory effect of tumor cell-derived lactic acid on human T cells. Blood. 2007;109:3812-3819. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1588] [Cited by in RCA: 1462] [Article Influence: 76.9] [Reference Citation Analysis (0)] |
| 30. | Watson MJ, Vignali PDA, Mullett SJ, Overacre-Delgoffe AE, Peralta RM, Grebinoski S, Menk AV, Rittenhouse NL, DePeaux K, Whetstone RD, Vignali DAA, Hand TW, Poholek AC, Morrison BM, Rothstein JD, Wendell SG, Delgoffe GM. Metabolic support of tumour-infiltrating regulatory T cells by lactic acid. Nature. 2021;591:645-651. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 605] [Cited by in RCA: 952] [Article Influence: 190.4] [Reference Citation Analysis (4)] |
| 31. | Palmqvist R, Wikberg ML, Ling A, Edin S. The Association of Immune Cell Infiltration and Prognosis in Colorectal Cancer. Curr Colorectal Cancer Rep. 2013;9:372-379. [RCA] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 10] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 32. | Heusinkveld M, van der Burg SH. Identification and manipulation of tumor associated macrophages in human cancers. J Transl Med. 2011;9:216. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 358] [Cited by in RCA: 347] [Article Influence: 23.1] [Reference Citation Analysis (4)] |
| 33. | Feng Y, Ma W, Zang Y, Guo Y, Li Y, Zhang Y, Dong X, Liu Y, Zhan X, Pan Z, Luo M, Wu M, Chen A, Kang D, Chen G, Liu L, Zhou J, Zhang R. Spatially organized tumor-stroma boundary determines the efficacy of immunotherapy in colorectal cancer patients. Nat Commun. 2024;15:10259. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 48] [Article Influence: 24.0] [Reference Citation Analysis (0)] |
| 34. | Menk AV, Scharping NE, Moreci RS, Zeng X, Guy C, Salvatore S, Bae H, Xie J, Young HA, Wendell SG, Delgoffe GM. Early TCR Signaling Induces Rapid Aerobic Glycolysis Enabling Distinct Acute T Cell Effector Functions. Cell Rep. 2018;22:1509-1521. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 214] [Cited by in RCA: 393] [Article Influence: 56.1] [Reference Citation Analysis (0)] |
| 35. | Kane LP, Weiss A. The PI-3 kinase/Akt pathway and T cell activation: pleiotropic pathways downstream of PIP3. Immunol Rev. 2003;192:7-20. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 202] [Cited by in RCA: 211] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 36. | Gubser PM, Bantug GR, Razik L, Fischer M, Dimeloe S, Hoenger G, Durovic B, Jauch A, Hess C. Rapid effector function of memory CD8+ T cells requires an immediate-early glycolytic switch. Nat Immunol. 2013;14:1064-1072. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 330] [Cited by in RCA: 423] [Article Influence: 32.5] [Reference Citation Analysis (0)] |
| 37. | Pfeiffer T, Schuster S, Bonhoeffer S. Cooperation and competition in the evolution of ATP-producing pathways. Science. 2001;292:504-507. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 922] [Cited by in RCA: 959] [Article Influence: 38.4] [Reference Citation Analysis (0)] |
| 38. | Shao Q, Wang L, Yuan M, Jin X, Chen Z, Wu C. TIGIT Induces (CD3+) T Cell Dysfunction in Colorectal Cancer by Inhibiting Glucose Metabolism. Front Immunol. 2021;12:688961. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 33] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 39. | Wang G, Wang YZ, Yu Y, Yin PH, Xu K, Zhang H. The Anti-Tumor Effect and Mechanism of Triterpenoids in Rhus chinensis Mill. on Reversing Effector CD8+ T-cells Dysfunction by Targeting Glycolysis Pathways in Colorectal Cancer. Integr Cancer Ther. 2021;20:15347354211017219. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 9] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 40. | Lu Y, Li Y, Liu Q, Tian N, Du P, Zhu F, Han Y, Liu X, Liu X, Peng X, Wang X, Wu Y, Tong L, Li Y, Zhu Y, Wu L, Zhang P, Xu Y, Chen H, Li B, Tong X. MondoA-Thioredoxin-Interacting Protein Axis Maintains Regulatory T-Cell Identity and Function in Colorectal Cancer Microenvironment. Gastroenterology. 2021;161:575-591. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 84] [Cited by in RCA: 82] [Article Influence: 16.4] [Reference Citation Analysis (1)] |
| 41. | Harrington JS, Ryter SW, Plataki M, Price DR, Choi AMK. Mitochondria in health, disease, and aging. Physiol Rev. 2023;103:2349-2422. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 146] [Cited by in RCA: 439] [Article Influence: 146.3] [Reference Citation Analysis (0)] |
| 42. | Ma X, Xiao L, Liu L, Ye L, Su P, Bi E, Wang Q, Yang M, Qian J, Yi Q. CD36-mediated ferroptosis dampens intratumoral CD8(+) T cell effector function and impairs their antitumor ability. Cell Metab. 2021;33:1001-1012. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 186] [Cited by in RCA: 772] [Article Influence: 154.4] [Reference Citation Analysis (4)] |
| 43. | Chen J, Wu Y, Zhou Q, Song Y, Zhuang J, Lu K, Yang X. GPX3 is a key cholesterol-related gene associated with prognosis and tumor-infiltrating T cells in colorectal cancer. Neoplasma. 2023;70:230704N348. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 7] [Reference Citation Analysis (0)] |
| 44. | Zhang Q, Zhao Q, Li T, Lu L, Wang F, Zhang H, Liu Z, Ma H, Zhu Q, Wang J, Zhang X, Pei Y, Liu Q, Xu Y, Qie J, Luan X, Hu Z, Liu X. Lactobacillus plantarum-derived indole-3-lactic acid ameliorates colorectal tumorigenesis via epigenetic regulation of CD8(+) T cell immunity. Cell Metab. 2023;35:943-960. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 368] [Cited by in RCA: 293] [Article Influence: 97.7] [Reference Citation Analysis (4)] |
| 45. | Zhang X, Yu D, Wu D, Gao X, Shao F, Zhao M, Wang J, Ma J, Wang W, Qin X, Chen Y, Xia P, Wang S. Tissue-resident Lachnospiraceae family bacteria protect against colorectal carcinogenesis by promoting tumor immune surveillance. Cell Host Microbe. 2023;31:418-432. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 182] [Reference Citation Analysis (1)] |
| 46. | Reinfeld BI, Madden MZ, Wolf MM, Chytil A, Bader JE, Patterson AR, Sugiura A, Cohen AS, Ali A, Do BT, Muir A, Lewis CA, Hongo RA, Young KL, Brown RE, Todd VM, Huffstater T, Abraham A, O'Neil RT, Wilson MH, Xin F, Tantawy MN, Merryman WD, Johnson RW, Williams CS, Mason EF, Mason FM, Beckermann KE, Vander Heiden MG, Manning HC, Rathmell JC, Rathmell WK. Cell-programmed nutrient partitioning in the tumour microenvironment. Nature. 2021;593:282-288. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 914] [Cited by in RCA: 812] [Article Influence: 162.4] [Reference Citation Analysis (0)] |
| 47. | Madden MZ, Rathmell JC. The Complex Integration of T-cell Metabolism and Immunotherapy. Cancer Discov. 2021;11:1636-1643. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 90] [Cited by in RCA: 117] [Article Influence: 23.4] [Reference Citation Analysis (0)] |
| 48. | Heys SD, Segar A, Payne S, Bruce DM, Kernohan N, Eremin O. Dietary supplementation with L-arginine: modulation of tumour-infiltrating lymphocytes in patients with colorectal cancer. Br J Surg. 1997;84:238-241. [PubMed] |
| 49. | Geiger R, Rieckmann JC, Wolf T, Basso C, Feng Y, Fuhrer T, Kogadeeva M, Picotti P, Meissner F, Mann M, Zamboni N, Sallusto F, Lanzavecchia A. L-Arginine Modulates T Cell Metabolism and Enhances Survival and Anti-tumor Activity. Cell. 2016;167:829-842. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 716] [Cited by in RCA: 1381] [Article Influence: 138.1] [Reference Citation Analysis (0)] |
| 50. | Bronte V, Zanovello P. Regulation of immune responses by L-arginine metabolism. Nat Rev Immunol. 2005;5:641-654. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1489] [Cited by in RCA: 1384] [Article Influence: 65.9] [Reference Citation Analysis (3)] |
| 51. | Helmink BA, Reddy SM, Gao J, Zhang S, Basar R, Thakur R, Yizhak K, Sade-Feldman M, Blando J, Han G, Gopalakrishnan V, Xi Y, Zhao H, Amaria RN, Tawbi HA, Cogdill AP, Liu W, LeBleu VS, Kugeratski FG, Patel S, Davies MA, Hwu P, Lee JE, Gershenwald JE, Lucci A, Arora R, Woodman S, Keung EZ, Gaudreau PO, Reuben A, Spencer CN, Burton EM, Haydu LE, Lazar AJ, Zapassodi R, Hudgens CW, Ledesma DA, Ong S, Bailey M, Warren S, Rao D, Krijgsman O, Rozeman EA, Peeper D, Blank CU, Schumacher TN, Butterfield LH, Zelazowska MA, McBride KM, Kalluri R, Allison J, Petitprez F, Fridman WH, Sautès-Fridman C, Hacohen N, Rezvani K, Sharma P, Tetzlaff MT, Wang L, Wargo JA. B cells and tertiary lymphoid structures promote immunotherapy response. Nature. 2020;577:549-555. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2268] [Cited by in RCA: 2030] [Article Influence: 338.3] [Reference Citation Analysis (0)] |
| 52. | Zhang J, Wu X, Ma J, Long K, Sun J, Li M, Ge L. Hypoxia and hypoxia-inducible factor signals regulate the development, metabolism, and function of B cells. Front Immunol. 2022;13:967576. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 21] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 53. | Doughty CA, Bleiman BF, Wagner DJ, Dufort FJ, Mataraza JM, Roberts MF, Chiles TC. Antigen receptor-mediated changes in glucose metabolism in B lymphocytes: role of phosphatidylinositol 3-kinase signaling in the glycolytic control of growth. Blood. 2006;107:4458-4465. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 230] [Cited by in RCA: 299] [Article Influence: 15.0] [Reference Citation Analysis (0)] |
| 54. | Cabrita R, Lauss M, Sanna A, Donia M, Skaarup Larsen M, Mitra S, Johansson I, Phung B, Harbst K, Vallon-Christersson J, van Schoiack A, Lövgren K, Warren S, Jirström K, Olsson H, Pietras K, Ingvar C, Isaksson K, Schadendorf D, Schmidt H, Bastholt L, Carneiro A, Wargo JA, Svane IM, Jönsson G. Tertiary lymphoid structures improve immunotherapy and survival in melanoma. Nature. 2020;577:561-565. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 660] [Cited by in RCA: 1694] [Article Influence: 282.3] [Reference Citation Analysis (0)] |
| 55. | Wang Z, Lu Z, Lin S, Xia J, Zhong Z, Xie Z, Xing Y, Qie J, Jiao M, Li Y, Wen H, Zhao P, Zhang D, Zhou P, Qian J, Luo F, Wang L, Yu H, Liu J, Gu J, Liu R, Chu Y. Leucine-tRNA-synthase-2-expressing B cells contribute to colorectal cancer immunoevasion. Immunity. 2022;55:1067-1081. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 88] [Article Influence: 22.0] [Reference Citation Analysis (0)] |
| 56. | Murray PJ. Macrophage Polarization. Annu Rev Physiol. 2017;79:541-566. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2560] [Cited by in RCA: 2285] [Article Influence: 253.9] [Reference Citation Analysis (6)] |
| 57. | Li M, Yang Y, Xiong L, Jiang P, Wang J, Li C. Metabolism, metabolites, and macrophages in cancer. J Hematol Oncol. 2023;16:80. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 362] [Cited by in RCA: 365] [Article Influence: 121.7] [Reference Citation Analysis (5)] |
| 58. | Scheurlen KM, Billeter AT, O'Brien SJ, Galandiuk S. Metabolic dysfunction and early-onset colorectal cancer - how macrophages build the bridge. Cancer Med. 2020;9:6679-6693. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 40] [Cited by in RCA: 37] [Article Influence: 6.2] [Reference Citation Analysis (1)] |
| 59. | Fritsch SD, Sukhbaatar N, Gonzales K, Sahu A, Tran L, Vogel A, Mazic M, Wilson JL, Forisch S, Mayr H, Oberle R, Weiszmann J, Brenner M, Vanhoutte R, Hofmann M, Pirnes-Karhu S, Magnes C, Kühnast T, Weckwerth W, Bock C, Klavins K, Hengstschläger M, Moissl-Eichinger C, Schabbauer G, Egger G, Pirinen E, Verhelst SHL, Weichhart T. Metabolic support by macrophages sustains colonic epithelial homeostasis. Cell Metab. 2023;35:1931-1943. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 63] [Article Influence: 21.0] [Reference Citation Analysis (0)] |
| 60. | Yoon BR, Oh YJ, Kang SW, Lee EB, Lee WW. Role of SLC7A5 in Metabolic Reprogramming of Human Monocyte/Macrophage Immune Responses. Front Immunol. 2018;9:53. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 122] [Cited by in RCA: 112] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 61. | Tian J, Zhang L, La X, Fan X, Li A, Wu C, An Y, Yan S, Dong X, Wu H, Li Z. Tumor-secreted GRP78 induces M2 polarization of macrophages by promoting lipid catabolism. Cell Signal. 2023;108:110719. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 13] [Reference Citation Analysis (0)] |
| 62. | Tall AR, Yvan-Charvet L. Cholesterol, inflammation and innate immunity. Nat Rev Immunol. 2015;15:104-116. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1036] [Cited by in RCA: 1157] [Article Influence: 105.2] [Reference Citation Analysis (4)] |
| 63. | Shi R, Zhao K, Wang T, Yuan J, Zhang D, Xiang W, Qian J, Luo N, Zhou Y, Tang B, Li C, Miao H. 5-aza-2'-deoxycytidine potentiates anti-tumor immunity in colorectal peritoneal metastasis by modulating ABC A9-mediated cholesterol accumulation in macrophages. Theranostics. 2022;12:875-890. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 44] [Cited by in RCA: 39] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 64. | Miao H, Ou J, Peng Y, Zhang X, Chen Y, Hao L, Xie G, Wang Z, Pang X, Ruan Z, Li J, Yu L, Xue B, Shi H, Shi C, Liang H. Macrophage ABHD5 promotes colorectal cancer growth by suppressing spermidine production by SRM. Nat Commun. 2016;7:11716. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 50] [Cited by in RCA: 79] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 65. | Ma J, Ma R, Zeng X, Zhang L, Liu J, Zhang W, Li T, Niu H, Bao G, Wang C, Wang PG, Wang J, Li X, Zou T, Xie S. Lysosome blockade induces divergent metabolic programs in macrophages and tumours for cancer immunotherapy. J Exp Clin Cancer Res. 2023;42:192. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 15] [Reference Citation Analysis (0)] |
| 66. | Schulz C, Gomez Perdiguero E, Chorro L, Szabo-Rogers H, Cagnard N, Kierdorf K, Prinz M, Wu B, Jacobsen SE, Pollard JW, Frampton J, Liu KJ, Geissmann F. A lineage of myeloid cells independent of Myb and hematopoietic stem cells. Science. 2012;336:86-90. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1721] [Cited by in RCA: 1997] [Article Influence: 142.6] [Reference Citation Analysis (2)] |
| 67. | Gabrilovich DI, Ostrand-Rosenberg S, Bronte V. Coordinated regulation of myeloid cells by tumours. Nat Rev Immunol. 2012;12:253-268. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3097] [Cited by in RCA: 2934] [Article Influence: 209.6] [Reference Citation Analysis (7)] |
| 68. | Solito S, Falisi E, Diaz-Montero CM, Doni A, Pinton L, Rosato A, Francescato S, Basso G, Zanovello P, Onicescu G, Garrett-Mayer E, Montero AJ, Bronte V, Mandruzzato S. A human promyelocytic-like population is responsible for the immune suppression mediated by myeloid-derived suppressor cells. Blood. 2011;118:2254-2265. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 259] [Cited by in RCA: 272] [Article Influence: 18.1] [Reference Citation Analysis (0)] |
| 69. | Bader JE, Voss K, Rathmell JC. Targeting Metabolism to Improve the Tumor Microenvironment for Cancer Immunotherapy. Mol Cell. 2020;78:1019-1033. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 566] [Cited by in RCA: 751] [Article Influence: 125.2] [Reference Citation Analysis (6)] |
| 70. | Veglia F, Perego M, Gabrilovich D. Myeloid-derived suppressor cells coming of age. Nat Immunol. 2018;19:108-119. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 896] [Cited by in RCA: 1454] [Article Influence: 181.8] [Reference Citation Analysis (5)] |
| 71. | Zhang Z, Chen Y, Yin Y, Chen Y, Chen Q, Bing Z, Zheng Y, Hou Y, Shen S, Chen Y, Wang T. Candida tropicalis induces NLRP3 inflammasome activation via glycogen metabolism-dependent glycolysis and JAK-STAT1 signaling pathway in myeloid-derived suppressor cells to promote colorectal carcinogenesis. Int Immunopharmacol. 2022;113:109430. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 25] [Reference Citation Analysis (0)] |
| 72. | OuYang LY, Wu XJ, Ye SB, Zhang RX, Li ZL, Liao W, Pan ZZ, Zheng LM, Zhang XS, Wang Z, Li Q, Ma G, Li J. Tumor-induced myeloid-derived suppressor cells promote tumor progression through oxidative metabolism in human colorectal cancer. J Transl Med. 2015;13:47. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 113] [Cited by in RCA: 160] [Article Influence: 14.5] [Reference Citation Analysis (4)] |
| 73. | Hossain F, Al-Khami AA, Wyczechowska D, Hernandez C, Zheng L, Reiss K, Valle LD, Trillo-Tinoco J, Maj T, Zou W, Rodriguez PC, Ochoa AC. Inhibition of Fatty Acid Oxidation Modulates Immunosuppressive Functions of Myeloid-Derived Suppressor Cells and Enhances Cancer Therapies. Cancer Immunol Res. 2015;3:1236-1247. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 500] [Cited by in RCA: 490] [Article Influence: 44.5] [Reference Citation Analysis (1)] |
| 74. | Yu H, Chen C, Han F, Tang J, Deng M, Niu Y, Lai M, Zhang H. Long Noncoding RNA MIR4435-2HG Suppresses Colorectal Cancer Initiation and Progression By Reprogramming Neutrophils. Cancer Immunol Res. 2022;10:1095-1110. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 35] [Article Influence: 8.8] [Reference Citation Analysis (0)] |
| 75. | Veglia F, Tyurin VA, Blasi M, De Leo A, Kossenkov AV, Donthireddy L, To TKJ, Schug Z, Basu S, Wang F, Ricciotti E, DiRusso C, Murphy ME, Vonderheide RH, Lieberman PM, Mulligan C, Nam B, Hockstein N, Masters G, Guarino M, Lin C, Nefedova Y, Black P, Kagan VE, Gabrilovich DI. Fatty acid transport protein 2 reprograms neutrophils in cancer. Nature. 2019;569:73-78. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 266] [Cited by in RCA: 601] [Article Influence: 85.9] [Reference Citation Analysis (23)] |
| 76. | Sun HW, Chen J, Wu WC, Yang YY, Xu YT, Yu XJ, Chen HT, Wang Z, Wu XJ, Zheng L. Retinoic Acid Synthesis Deficiency Fosters the Generation of Polymorphonuclear Myeloid-Derived Suppressor Cells in Colorectal Cancer. Cancer Immunol Res. 2021;9:20-33. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 24] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 77. | Rodriguez PC, Quiceno DG, Zabaleta J, Ortiz B, Zea AH, Piazuelo MB, Delgado A, Correa P, Brayer J, Sotomayor EM, Antonia S, Ochoa JB, Ochoa AC. Arginase I production in the tumor microenvironment by mature myeloid cells inhibits T-cell receptor expression and antigen-specific T-cell responses. Cancer Res. 2004;64:5839-5849. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 799] [Cited by in RCA: 986] [Article Influence: 44.8] [Reference Citation Analysis (0)] |
| 78. | Raber PL, Thevenot P, Sierra R, Wyczechowska D, Halle D, Ramirez ME, Ochoa AC, Fletcher M, Velasco C, Wilk A, Reiss K, Rodriguez PC. Subpopulations of myeloid-derived suppressor cells impair T cell responses through independent nitric oxide-related pathways. Int J Cancer. 2014;134:2853-2864. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 239] [Cited by in RCA: 244] [Article Influence: 20.3] [Reference Citation Analysis (0)] |
| 79. | Cheon EC, Khazaie K, Khan MW, Strouch MJ, Krantz SB, Phillips J, Blatner NR, Hix LM, Zhang M, Dennis KL, Salabat MR, Heiferman M, Grippo PJ, Munshi HG, Gounaris E, Bentrem DJ. Mast cell 5-lipoxygenase activity promotes intestinal polyposis in APCDelta468 mice. Cancer Res. 2011;71:1627-1636. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 73] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 80. | Joshi S, Sharabi A. Targeting myeloid-derived suppressor cells to enhance natural killer cell-based immunotherapy. Pharmacol Ther. 2022;235:108114. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 92] [Reference Citation Analysis (4)] |
| 81. | Platten M, Nollen EAA, Röhrig UF, Fallarino F, Opitz CA. Tryptophan metabolism as a common therapeutic target in cancer, neurodegeneration and beyond. Nat Rev Drug Discov. 2019;18:379-401. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 445] [Cited by in RCA: 1171] [Article Influence: 167.3] [Reference Citation Analysis (0)] |
| 82. | Shimasaki N, Jain A, Campana D. NK cells for cancer immunotherapy. Nat Rev Drug Discov. 2020;19:200-218. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 381] [Cited by in RCA: 909] [Article Influence: 151.5] [Reference Citation Analysis (0)] |
| 83. | He H, Song Z, Lin S, Wang Y, Wang G. Exploring the effect of polyamines on NK cell function in colorectal cancer process based on glycolysis. Int Immunopharmacol. 2023;117:109944. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 10] [Reference Citation Analysis (0)] |
| 84. | Poznanski SM, Singh K, Ritchie TM, Aguiar JA, Fan IY, Portillo AL, Rojas EA, Vahedi F, El-Sayes A, Xing S, Butcher M, Lu Y, Doxey AC, Schertzer JD, Hirte HW, Ashkar AA. Metabolic flexibility determines human NK cell functional fate in the tumor microenvironment. Cell Metab. 2021;33:1205-1220. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 237] [Article Influence: 47.4] [Reference Citation Analysis (0)] |
| 85. | Harmon C, Robinson MW, Hand F, Almuaili D, Mentor K, Houlihan DD, Hoti E, Lynch L, Geoghegan J, O'Farrelly C. Lactate-Mediated Acidification of Tumor Microenvironment Induces Apoptosis of Liver-Resident NK Cells in Colorectal Liver Metastasis. Cancer Immunol Res. 2019;7:335-346. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 293] [Cited by in RCA: 274] [Article Influence: 39.1] [Reference Citation Analysis (0)] |
| 86. | Felices M, Lenvik AJ, McElmurry R, Chu S, Hinderlie P, Bendzick L, Geller MA, Tolar J, Blazar BR, Miller JS. Continuous treatment with IL-15 exhausts human NK cells via a metabolic defect. JCI Insight. 2018;3:e96219. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 173] [Cited by in RCA: 214] [Article Influence: 26.8] [Reference Citation Analysis (0)] |
| 87. | Liu Z, Zhou X, Kuang L, Chen Q, Zhao J, Yin H, Zhou Z, Liu X, Liu D, Wu S, Wu L. Novel insights into immune-gut microbiota interactions in colorectal cancer: a Mendelian randomization study. Infect Agent Cancer. 2025;20:27. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 8] [Reference Citation Analysis (0)] |
| 88. | Rubinstein MR, Wang X, Liu W, Hao Y, Cai G, Han YW. Fusobacterium nucleatum promotes colorectal carcinogenesis by modulating E-cadherin/β-catenin signaling via its FadA adhesin. Cell Host Microbe. 2013;14:195-206. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2107] [Cited by in RCA: 1867] [Article Influence: 143.6] [Reference Citation Analysis (13)] |
| 89. | Gur C, Maalouf N, Shhadeh A, Berhani O, Singer BB, Bachrach G, Mandelboim O. Fusobacterium nucleatum supresses anti-tumor immunity by activating CEACAM1. Oncoimmunology. 2019;8:e1581531. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 53] [Cited by in RCA: 130] [Article Influence: 18.6] [Reference Citation Analysis (1)] |
| 90. | Mellies JL, Platenkamp A, Osborn J, Ben-Avi L. PerC Manipulates Metabolism and Surface Antigens in Enteropathogenic Escherichia coli. Front Cell Infect Microbiol. 2017;7:32. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 10] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 91. | Trompette A, Gollwitzer ES, Pattaroni C, Lopez-Mejia IC, Riva E, Pernot J, Ubags N, Fajas L, Nicod LP, Marsland BJ. Dietary Fiber Confers Protection against Flu by Shaping Ly6c(-) Patrolling Monocyte Hematopoiesis and CD8(+) T Cell Metabolism. Immunity. 2018;48:992-1005. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 295] [Cited by in RCA: 536] [Article Influence: 76.6] [Reference Citation Analysis (2)] |
| 92. | Kovtonyuk LV, McCoy KD. Microbial metabolites and immunotherapy: Basic rationale and clinical indications. Semin Immunol. 2023;67:101755. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 21] [Reference Citation Analysis (0)] |
| 93. | Yang Q, Wang B, Zheng Q, Li H, Meng X, Zhou F, Zhang L. A Review of Gut Microbiota-Derived Metabolites in Tumor Progression and Cancer Therapy. Adv Sci (Weinh). 2023;10:e2207366. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 158] [Article Influence: 52.7] [Reference Citation Analysis (0)] |
| 94. | Bachem A, Makhlouf C, Binger KJ, de Souza DP, Tull D, Hochheiser K, Whitney PG, Fernandez-Ruiz D, Dähling S, Kastenmüller W, Jönsson J, Gressier E, Lew AM, Perdomo C, Kupz A, Figgett W, Mackay F, Oleshansky M, Russ BE, Parish IA, Kallies A, McConville MJ, Turner SJ, Gebhardt T, Bedoui S. Microbiota-Derived Short-Chain Fatty Acids Promote the Memory Potential of Antigen-Activated CD8(+) T Cells. Immunity. 2019;51:285-297. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 209] [Cited by in RCA: 534] [Article Influence: 76.3] [Reference Citation Analysis (3)] |
| 95. | Zou F, Qiu Y, Huang Y, Zou H, Cheng X, Niu Q, Luo A, Sun J. Effects of short-chain fatty acids in inhibiting HDAC and activating p38 MAPK are critical for promoting B10 cell generation and function. Cell Death Dis. 2021;12:582. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 100] [Article Influence: 20.0] [Reference Citation Analysis (0)] |
| 96. | Zhang Q, Liu Y, Li Y, Bai G, Pang J, Wu M, Li J, Zhao X, Xia Y. Implications of gut microbiota-mediated epigenetic modifications in intestinal diseases. Gut Microbes. 2025;17:2508426. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 33] [Article Influence: 33.0] [Reference Citation Analysis (0)] |
| 97. | Scott NA, Andrusaite A, Andersen P, Lawson M, Alcon-Giner C, Leclaire C, Caim S, Le Gall G, Shaw T, Connolly JPR, Roe AJ, Wessel H, Bravo-Blas A, Thomson CA, Kästele V, Wang P, Peterson DA, Bancroft A, Li X, Grencis R, Mowat AM, Hall LJ, Travis MA, Milling SWF, Mann ER. Antibiotics induce sustained dysregulation of intestinal T cell immunity by perturbing macrophage homeostasis. Sci Transl Med. 2018;10:eaao4755. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 295] [Cited by in RCA: 264] [Article Influence: 33.0] [Reference Citation Analysis (4)] |
| 98. | Agus A, Planchais J, Sokol H. Gut Microbiota Regulation of Tryptophan Metabolism in Health and Disease. Cell Host Microbe. 2018;23:716-724. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2576] [Cited by in RCA: 2204] [Article Influence: 275.5] [Reference Citation Analysis (7)] |
| 99. | Sun XZ, Zhao DY, Zhou YC, Wang QQ, Qin G, Yao SK. Alteration of fecal tryptophan metabolism correlates with shifted microbiota and may be involved in pathogenesis of colorectal cancer. World J Gastroenterol. 2020;26:7173-7190. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 52] [Cited by in RCA: 59] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 100. | Zhang X, Liu X, Zhou W, Du Q, Yang M, Ding Y, Hu R. Blockade of IDO-Kynurenine-AhR Axis Ameliorated Colitis-Associated Colon Cancer via Inhibiting Immune Tolerance. Cell Mol Gastroenterol Hepatol. 2021;12:1179-1199. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 65] [Article Influence: 13.0] [Reference Citation Analysis (0)] |
| 101. | Zhang C, Zhen Y, Weng Y, Lin J, Xu X, Ma J, Zhong Y, Wang M. Research progress on the microbial metabolism and transport of polyamines and their roles in animal gut homeostasis. J Anim Sci Biotechnol. 2025;16:57. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 20] [Reference Citation Analysis (0)] |
| 102. | Gobert AP, Latour YL, Asim M, Barry DP, Allaman MM, Finley JL, Smith TM, McNamara KM, Singh K, Sierra JC, Delgado AG, Luis PB, Schneider C, Washington MK, Piazuelo MB, Zhao S, Coburn LA, Wilson KT. Protective Role of Spermidine in Colitis and Colon Carcinogenesis. Gastroenterology. 2022;162:813-827. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 110] [Article Influence: 27.5] [Reference Citation Analysis (0)] |
| 103. | Collins SL, Stine JG, Bisanz JE, Okafor CD, Patterson AD. Bile acids and the gut microbiota: metabolic interactions and impacts on disease. Nat Rev Microbiol. 2023;21:236-247. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 922] [Cited by in RCA: 760] [Article Influence: 253.3] [Reference Citation Analysis (5)] |
| 104. | Jia W, Xie G, Jia W. Bile acid-microbiota crosstalk in gastrointestinal inflammation and carcinogenesis. Nat Rev Gastroenterol Hepatol. 2018;15:111-128. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1651] [Cited by in RCA: 1458] [Article Influence: 182.3] [Reference Citation Analysis (5)] |
| 105. | de Waal GM, de Villiers WJS, Forgan T, Roberts T, Pretorius E. Colorectal cancer is associated with increased circulating lipopolysaccharide, inflammation and hypercoagulability. Sci Rep. 2020;10:8777. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 37] [Cited by in RCA: 64] [Article Influence: 10.7] [Reference Citation Analysis (1)] |
| 106. | Sulit AK, Daigneault M, Allen-Vercoe E, Silander OK, Hock B, McKenzie J, Pearson J, Frizelle FA, Schmeier S, Purcell R. Bacterial lipopolysaccharide modulates immune response in the colorectal tumor microenvironment. NPJ Biofilms Microbiomes. 2023;9:59. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 34] [Article Influence: 11.3] [Reference Citation Analysis (0)] |
| 107. | Liu L, Liu Y, Guo X, Jin X, Yan W, Lin B, Cai T, Wei Y. Activation of p38 mitogen-activated protein kinase pathway by lipopolysaccharide aggravates postoperative ileus in colorectal cancer patients. J Gastroenterol Hepatol. 2022;37:518-530. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 9] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 108. | Qi Q, Zhang H, Jin Z, Wang C, Xia M, Chen B, Lv B, Peres Diaz L, Li X, Feng R, Qiu M, Li Y, Meseguer D, Zheng X, Wang W, Song W, Huang H, Wu H, Chen L, Schneeberger M, Yu X. Hydrogen sulfide produced by the gut microbiota impairs host metabolism via reducing GLP-1 levels in male mice. Nat Metab. 2024;6:1601-1615. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 58] [Article Influence: 29.0] [Reference Citation Analysis (0)] |
| 109. | Libiad M, Vitvitsky V, Bostelaar T, Bak DW, Lee HJ, Sakamoto N, Fearon E, Lyssiotis CA, Weerapana E, Banerjee R. Hydrogen sulfide perturbs mitochondrial bioenergetics and triggers metabolic reprogramming in colon cells. J Biol Chem. 2019;294:12077-12090. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 87] [Cited by in RCA: 114] [Article Influence: 16.3] [Reference Citation Analysis (0)] |
| 110. | Wang M, Wang Z, Lessing DJ, Guo M, Chu W. Fusobacterium nucleatum and its metabolite hydrogen sulfide alter gut microbiota composition and autophagy process and promote colorectal cancer progression. Microbiol Spectr. 2023;11:e0229223. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 26] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 111. | Wang H, Feng L, Pei Z, Zhao J, Lu S, Lu W. Gut microbiota metabolism of branched-chain amino acids and their metabolites can improve the physiological function of aging mice. Aging Cell. 2025;24:e14434. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 21] [Article Influence: 21.0] [Reference Citation Analysis (0)] |
| 112. | Agus A, Clément K, Sokol H. Gut microbiota-derived metabolites as central regulators in metabolic disorders. Gut. 2021;70:1174-1182. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1074] [Cited by in RCA: 910] [Article Influence: 182.0] [Reference Citation Analysis (5)] |
| 113. | Karin M, Greten FR. NF-kappaB: linking inflammation and immunity to cancer development and progression. Nat Rev Immunol. 2005;5:749-759. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2568] [Cited by in RCA: 2439] [Article Influence: 116.1] [Reference Citation Analysis (7)] |
| 114. | Brennan CA, Garrett WS. Gut Microbiota, Inflammation, and Colorectal Cancer. Annu Rev Microbiol. 2016;70:395-411. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 497] [Cited by in RCA: 452] [Article Influence: 45.2] [Reference Citation Analysis (0)] |
| 115. | Engevik MA, Danhof HA, Ruan W, Engevik AC, Chang-Graham AL, Engevik KA, Shi Z, Zhao Y, Brand CK, Krystofiak ES, Venable S, Liu X, Hirschi KD, Hyser JM, Spinler JK, Britton RA, Versalovic J. Fusobacterium nucleatum Secretes Outer Membrane Vesicles and Promotes Intestinal Inflammation. mBio. 2021;12:e02706-e02720. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 224] [Cited by in RCA: 205] [Article Influence: 41.0] [Reference Citation Analysis (0)] |
| 116. | Wang P, Huang Q, Zhu Y, Chen L, Ye K. Fusobacterium Nucleatum Promotes Microsatellite Instability in Colorectal Carcinoma Through Up-regulation of miRNA-155-5p-Targeted Inhibition of MSH6 via the TLR4/NF-κB Signaling Pathway. Adv Biol (Weinh). 2024;8:e2400293. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 7] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 117. | Jeon JI, Lee KH, Kim JM. Bacteroides fragilis Enterotoxin Upregulates Matrix Metalloproteinase-7 Expression through MAPK and AP-1 Activation in Intestinal Epithelial Cells, Leading to Syndecan-2 Release. Int J Mol Sci. 2021;22:11817. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 8] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 118. | Zhong Y, Chang X, Zhao Z, Zheng L, Kuang G, Li P, Liu C, Fan Y, Liang Z, Zhuang K, Xie Q, Liu Y. Bacteroides fragilis capsular polysaccharide A ameliorates ulcerative colitis in rat by recovering intestinal barrier integrity and restoring gut microbiota. Front Pharmacol. 2024;15:1402465. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 12] [Reference Citation Analysis (1)] |
| 119. | Blandford LE, Johnston EL, Sanderson JD, Wade WG, Lax AJ. Promoter orientation of the immunomodulatory Bacteroides fragilis capsular polysaccharide A (PSA) is off in individuals with inflammatory bowel disease (IBD). Gut Microbes. 2019;10:569-577. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 38] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 120. | Palmela C, Chevarin C, Xu Z, Torres J, Sevrin G, Hirten R, Barnich N, Ng SC, Colombel JF. Adherent-invasive Escherichia coli in inflammatory bowel disease. Gut. 2018;67:574-587. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 476] [Cited by in RCA: 411] [Article Influence: 51.4] [Reference Citation Analysis (1)] |
| 121. | Buisson A, Sokol H, Hammoudi N, Nancey S, Treton X, Nachury M, Fumery M, Hébuterne X, Rodrigues M, Hugot JP, Boschetti G, Stefanescu C, Wils P, Seksik P, Le Bourhis L, Bezault M, Sauvanet P, Pereira B, Allez M, Barnich N; Remind study group. Role of adherent and invasive Escherichia coli in Crohn's disease: lessons from the postoperative recurrence model. Gut. 2023;72:39-48. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 53] [Cited by in RCA: 46] [Article Influence: 15.3] [Reference Citation Analysis (0)] |
| 122. | Chen L, Zhang L, Hua H, Liu L, Mao Y, Wang R. Interactions between toll-like receptors signaling pathway and gut microbiota in host homeostasis. Immun Inflamm Dis. 2024;12:e1356. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 60] [Cited by in RCA: 42] [Article Influence: 21.0] [Reference Citation Analysis (1)] |
| 123. | Cong J, Liu P, Han Z, Ying W, Li C, Yang Y, Wang S, Yang J, Cao F, Shen J, Zeng Y, Bai Y, Zhou C, Ye L, Zhou R, Guo C, Cang C, Kasper DL, Song X, Dai L, Sun L, Pan W, Zhu S. Bile acids modified by the intestinal microbiota promote colorectal cancer growth by suppressing CD8(+) T cell effector functions. Immunity. 2024;57:876-889. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 198] [Article Influence: 99.0] [Reference Citation Analysis (2)] |
| 124. | Liao Q, Zhou X, Wu L, Yang Y, Zhu X, Liao H, Zhang Y, Lian W, Zhang F, Wang H, Ding Y, Zhao L. Gut microbial metabolite 4-hydroxybenzeneacetic acid drives colorectal cancer progression via accumulation of immunosuppressive PMN-MDSCs. J Clin Invest. 2025;135:e181243. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 24] [Article Influence: 24.0] [Reference Citation Analysis (0)] |
| 125. | Li R, Zhou R, Wang H, Li W, Pan M, Yao X, Zhan W, Yang S, Xu L, Ding Y, Zhao L. Gut microbiota-stimulated cathepsin K secretion mediates TLR4-dependent M2 macrophage polarization and promotes tumor metastasis in colorectal cancer. Cell Death Differ. 2019;26:2447-2463. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 334] [Cited by in RCA: 299] [Article Influence: 42.7] [Reference Citation Analysis (4)] |
| 126. | Tran M, Huh JR, Devlin AS. The role of gut microbial metabolites in the T cell lifecycle. Nat Immunol. 2025;26:1246-1257. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 15] [Article Influence: 15.0] [Reference Citation Analysis (0)] |
| 127. | Karta J, Meyers M, Rodriguez F, Koncina E, Gilson C, Klein E, Gabola M, Benzarti M, Pérez Escriva P, Molina Tijeras JA, Correia Tavares Bernardino C, Ponath F, Carpentier A, Pujabet MA, Schmoetten M, Tsenkova M, Saoud P, Gaigneaux A, Ternes D, Alonso L, Zügel N, Willemssen E, Koppes P, Léonard D, Casanova LP, Haan S, Mittelbronn M, Meiser J, Pozdeev VI, Vogel J, Nuciforo PG, Wilmes P, Letellier E. Fusobacterium nucleatum interacts with cancer-associated fibroblasts to promote colorectal cancer. EMBO J. 2025;44:5375-5393. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 8] [Reference Citation Analysis (0)] |
| 128. | Xu Z, Lv Z, Chen F, Zhang Y, Xu Z, Huo J, Liu W, Yu S, Tuersun A, Zhao J, Zong Y, Shen X, Feng W, Lu A. Dysbiosis of human tumor microbiome and aberrant residence of Actinomyces in tumor-associated fibroblasts in young-onset colorectal cancer. Front Immunol. 2022;13:1008975. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 55] [Cited by in RCA: 44] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 129. | Wang N, Fang JY. Fusobacterium nucleatum, a key pathogenic factor and microbial biomarker for colorectal cancer. Trends Microbiol. 2023;31:159-172. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 321] [Cited by in RCA: 265] [Article Influence: 88.3] [Reference Citation Analysis (4)] |
| 130. | Zepeda-Rivera M, Minot SS, Bouzek H, Wu H, Blanco-Míguez A, Manghi P, Jones DS, LaCourse KD, Wu Y, McMahon EF, Park SN, Lim YK, Kempchinsky AG, Willis AD, Cotton SL, Yost SC, Sicinska E, Kook JK, Dewhirst FE, Segata N, Bullman S, Johnston CD. A distinct Fusobacterium nucleatum clade dominates the colorectal cancer niche. Nature. 2024;628:424-432. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 339] [Cited by in RCA: 293] [Article Influence: 146.5] [Reference Citation Analysis (0)] |
| 131. | Wang X, Fang Y, Liang W, Wong CC, Qin H, Gao Y, Liang M, Song L, Zhang Y, Fan M, Liu C, Lau HC, Xu L, Li X, Song W, Wang J, Wang N, Yang T, Mo M, Zhang X, Fang J, Liao B, Sung JJY, Yu J. Fusobacterium nucleatum facilitates anti-PD-1 therapy in microsatellite stable colorectal cancer. Cancer Cell. 2024;42:1729-1746. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 169] [Cited by in RCA: 136] [Article Influence: 68.0] [Reference Citation Analysis (0)] |
| 132. | Xiaofeng N, Jian C, Jingjing W, Zhanbo Q, Yifei S, Jing Z, Shuwen H. Correlation of gut microbiota with leukopenia after chemotherapy in patients with colorectal cancer. BMC Microbiol. 2023;23:349. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 8] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 133. | Mezerová K, Starý L, Zbořil P, Klementa I, Stašek M, Špička P, Skalický P, Raclavský V. Cyclomodulins and Hemolysis in E. coli as Potential Low-Cost Non-Invasive Biomarkers for Colorectal Cancer Screening. Life (Basel). 2021;11:1165. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 134. | Teng H, Wang Y, Sui X, Fan J, Li S, Lei X, Shi C, Sun W, Song M, Wang H, Dong D, Geng J, Zhang Y, Zhu X, Cai Y, Li Y, Li B, Min Q, Wang W, Zhan Q. Gut microbiota-mediated nucleotide synthesis attenuates the response to neoadjuvant chemoradiotherapy in rectal cancer. Cancer Cell. 2023;41:124-138. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 98] [Article Influence: 32.7] [Reference Citation Analysis (0)] |
| 135. | Zhang S, Wang Q, Zhou C, Chen K, Chang H, Xiao W, Gao Y. Colorectal cancer, radiotherapy and gut microbiota. Chin J Cancer Res. 2019;31:212-222. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 28] [Cited by in RCA: 26] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 136. | Van Dingenen L, Segers C, Wouters S, Mysara M, Leys N, Kumar-Singh S, Malhotra-Kumar S, Van Houdt R. Dissecting the role of the gut microbiome and fecal microbiota transplantation in radio- and immunotherapy treatment of colorectal cancer. Front Cell Infect Microbiol. 2023;13:1298264. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 18] [Reference Citation Analysis (0)] |
| 137. | Lang T, Zhu R, Zhu X, Yan W, Li Y, Zhai Y, Wu T, Huang X, Yin Q, Li Y. Combining gut microbiota modulation and chemotherapy by capecitabine-loaded prebiotic nanoparticle improves colorectal cancer therapy. Nat Commun. 2023;14:4746. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 94] [Reference Citation Analysis (0)] |
| 138. | Sun X, Shan X, Zhu B, Cai Y, He Z, Zhou L, Yin L, Liu Y, Liu K, Zhang T, Yang N, Li Y, Lang T. 5-Fluorouracil Loaded Prebiotic-Probiotic Liposomes Modulating Gut Microbiota for Improving Colorectal Cancer Chemotherapy. Adv Healthc Mater. 2025;14:e2403587. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 11] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 139. | Zheng DW, Dong X, Pan P, Chen KW, Fan JX, Cheng SX, Zhang XZ. Phage-guided modulation of the gut microbiota of mouse models of colorectal cancer augments their responses to chemotherapy. Nat Biomed Eng. 2019;3:717-728. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 417] [Cited by in RCA: 353] [Article Influence: 50.4] [Reference Citation Analysis (3)] |
| 140. | Wang S, Xu B, Zhang Y, Chen G, Zhao P, Gao Q, Yuan L. The role of intestinal flora on tumorigenesis, progression, and the efficacy of PD-1/PD-L1 antibodies in colorectal cancer. Cancer Biol Med. 2023;21:65-82. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 12] [Reference Citation Analysis (0)] |
| 141. | Xu S, Kong J, Dai Y, Li H. Prevotellaceae Modulates Colorectal Cancer Immune Microenvironment to Assist Anti-PD-L1 Immunotherapy. Turk J Gastroenterol. 2024;35:909-921. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 5] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 142. | Roberti MP, Yonekura S, Duong CPM, Picard M, Ferrere G, Tidjani Alou M, Rauber C, Iebba V, Lehmann CHK, Amon L, Dudziak D, Derosa L, Routy B, Flament C, Richard C, Daillère R, Fluckiger A, Van Seuningen I, Chamaillard M, Vincent A, Kourula S, Opolon P, Ly P, Pizzato E, Becharef S, Paillet J, Klein C, Marliot F, Pietrantonio F, Benoist S, Scoazec JY, Dartigues P, Hollebecque A, Malka D, Pagès F, Galon J, Gomperts Boneca I, Lepage P, Ryffel B, Raoult D, Eggermont A, Vanden Berghe T, Ghiringhelli F, Vandenabeele P, Kroemer G, Zitvogel L. Chemotherapy-induced ileal crypt apoptosis and the ileal microbiome shape immunosurveillance and prognosis of proximal colon cancer. Nat Med. 2020;26:919-931. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 181] [Cited by in RCA: 161] [Article Influence: 26.8] [Reference Citation Analysis (0)] |
| 143. | Xu X, Lv J, Guo F, Li J, Jia Y, Jiang D, Wang N, Zhang C, Kong L, Liu Y, Zhang Y, Lv J, Li Z. Gut Microbiome Influences the Efficacy of PD-1 Antibody Immunotherapy on MSS-Type Colorectal Cancer via Metabolic Pathway. Front Microbiol. 2020;11:814. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 92] [Cited by in RCA: 138] [Article Influence: 23.0] [Reference Citation Analysis (4)] |
| 144. | Jiang SS, Xie YL, Xiao XY, Kang ZR, Lin XL, Zhang L, Li CS, Qian Y, Xu PP, Leng XX, Wang LW, Tu SP, Zhong M, Zhao G, Chen JX, Wang Z, Liu Q, Hong J, Chen HY, Chen YX, Fang JY. Fusobacterium nucleatum-derived succinic acid induces tumor resistance to immunotherapy in colorectal cancer. Cell Host Microbe. 2023;31:781-797. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 318] [Cited by in RCA: 285] [Article Influence: 95.0] [Reference Citation Analysis (1)] |
| 145. | Lu S, Xu J, Zhao Z, Guo Y, Zhang H, Jurutka PW, Huang D, Cao C, Cheng S. Dietary Lactobacillus rhamnosus GG extracellular vesicles enhance antiprogrammed cell death 1 (anti-PD-1) immunotherapy efficacy against colorectal cancer. Food Funct. 2023;14:10314-10328. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 34] [Reference Citation Analysis (0)] |
| 146. | Niu M, Pei Y, Jin T, Li J, Bai L, Zheng C, Song Q, Zhao H, Zhang Y, Wang L. Colon-specific controlled release of oral liposomes for enhanced chemo-immunotherapy against colorectal cancer. Acta Pharm Sin B. 2024;14:4977-4993. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 16] [Reference Citation Analysis (0)] |
| 147. | Yan J, Duan W, Gao Q, Mao T, Wang M, Duan J, Li J. ENPP2 inhibitor improves proliferation in AOM/DSS-induced colorectal cancer mice via remodeling the gut barrier function and gut microbiota composition. Pharmacol Res. 2023;195:106877. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 21] [Reference Citation Analysis (0)] |
| 148. | Wang M, Li Y, Yang X, Liu Z, Wang K, Gong D, Li J. Effects of metronidazole on colorectal cancer occurrence and colorectal cancer liver metastases by regulating Fusobacterium nucleatum in mice. Immun Inflamm Dis. 2023;11:e1067. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 14] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 149. | Xie S, Wei L, Liu Y, Meng J, Cao W, Qiu B, Li X. Size-tunable nanogels for cascaded release of metronidazole and chemotherapeutic agents to combat Fusobacterium nucleatum-infected colorectal cancer. J Control Release. 2024;365:16-28. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 24] [Reference Citation Analysis (0)] |
| 150. | Niu Y, Zhao X, Li Y, Ma X, Yang W, Ma J, Li W, Yuan W. Neutrophil-Mimicking Nanomedicine Eliminates Tumor Intracellular Bacteria and Enhances Chemotherapy on Liver Metastasis of Colorectal Cancer. Adv Sci (Weinh). 2025;12:e04188. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 11] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 151. | Jia F, Yu Q, Wang R, Zhao L, Yuan F, Guo H, Shen Y, He F. Optimized Antimicrobial Peptide Jelleine-I Derivative Br-J-I Inhibits Fusobacterium Nucleatum to Suppress Colorectal Cancer Progression. Int J Mol Sci. 2023;24:1469. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 26] [Reference Citation Analysis (0)] |