Published online Mar 15, 2026. doi: 10.4251/wjgo.v18.i3.115679
Revised: November 12, 2025
Accepted: December 10, 2025
Published online: March 15, 2026
Processing time: 141 Days and 10.7 Hours
CD44, also known as differentiation antigen 44, is a key marker for cancer stem cells and an indicator of poor prognosis in cancer patients. Currently, there are no cancer therapies that specifically target CD44 in clinical practice. Pancreatic cancer has an extremely high mortality rate, and there are few effective treatment options. The effects of CD44 gene knockout on pancreatic cancer have not been previously reported.
To investigate the impact of CD44 gene knockout on pancreatic tumorigenesis and treatment.
CRISPR/Cas9 technology was used to disrupt CD44 gene exons in PaTu-8988 and Panc-1 pancreatic cancer cells to establish CD44 knockout cell lines. The effects of CD44 gene disruption on tumorigenesis in pancreatic cancer cells were then investigated at the molecular, cellular, and animal levels.
CD44 disruption inhibited tumor formation and pancreatic cancer cell migration and invasion capabilities. It increased DNA damage and sensitivity to the chemotherapeutic agent cisplatin, while also suppressing tumor growth in syngeneic mouse xenografts. Mechanistically, CD44 knockout revealed a novel oncogenic axis in cancer cells marked by a 35-fold reduction in promoter activity of the X-chromosome inactivation-specific transcription factor. Furthermore, CD44 knockout suppressed the expression of multiple other oncogenes, weakened the pro-tumor AKT and ERK signaling pathways, and simultaneously activated the tumor-suppressing p38 and p53 pathways.
Oncogenic CD44 plays a pivotal role in pancreatic cancer cell carcinogenesis, and CD44 knockout presents a promising therapeutic target for pancreatic cancer treatment. These findings demonstrate the central role of CD44 in pancreatic cancer and provide new therapeutic strategies for its management.
Core Tip: This study employed CRISPR/Cas9-mediated gene editing to knock out the CD44 gene, elucidating its critical role in pancreatic cancer progression. CD44 deficiency significantly suppressed cell proliferation, migration, and invasion, while simultaneously enhancing sensitivity to cisplatin treatment. Furthermore, we identified a novel regulatory axis involving CD44 and X-inactive-specific transcript. Mechanistically, CD44 knockout inhibited the AKT/ERK signaling pathway while activating the p38/p53 pathway. Collectively, these findings indicate that targeting CD44 through CRISPR/Cas9-based gene editing may represent a promising therapeutic strategy for pancreatic cancer.
- Citation: Liu YX, Zheng NN, Wang XX, Zhou QS, Meng M. Oncogenic CD44 is essential for pancreatic cancer tumorigenesis: A novel targeted therapeutic strategy. World J Gastrointest Oncol 2026; 18(3): 115679
- URL: https://www.wjgnet.com/1948-5204/full/v18/i3/115679.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v18.i3.115679
Pancreatic cancer is a highly lethal cancer with a 5-year survival rate of only 8%[1]. The incidence of pancreatic cancer is increasing globally, and by 2039, it is projected to be the second leading cause of cancer-related deaths in Western countries and the fourth in developing countries[2]. Despite efforts to combat pancreatic cancer in recent decades, its 5-year survival rate has seen minimal improvement. The unclear mechanisms underlying pancreatic cancer initiation and progression, along with the lack of effective therapies, contribute to its poor prognosis[3,4].
Pancreatic cancer stem cells, identified by the molecular markers CD44, CD133, and CD24, play crucial roles in cancer initiation, progression, metastasis, drug resistance, and recurrence[5]. These cells comprise 36% of all cells in pancreatic tumor tissues, and are correlated with a poor prognosis[6,7]. Therefore, targeting pancreatic cancer stem cells is a logical approach for therapy, yet eliminating these cells remains a challenge in cancer research[3,4].
CD44 is overexpressed in cancer stem/progenitor cells, but absent or barely expressed in normal cells[8]. CD44 overexpression triggers epithelial-mesenchymal transition, promoting tumor cell proliferation and increasing cancer cell stemness, dissemination, and drug resistance[9-12]. Thus, CD44 is a reasonable target for cancer therapy[13]. CD44 is a hyaluronic acid (HA) receptor and is activated in various types of malignant tumors[14,15].
The long noncoding RNA X-inactive-specific transcript (Xist)[16,17] has been implicated in oncogenesis, tumor progression, metastasis, and a worse prognosis, particularly in pancreatic cancer[18-23]. However, a clear understanding of its regulatory mechanisms is currently lacking.
CRISPR/Cas9 technology, a known method for gene knockout and genetic error correction, shows promise in targeted cancer therapy[24-26]. The impact of CD44 gene knockout on pancreatic cancer cells has not yet been explored. Given the presence of a start codon in exon 1, we hypothesized that disrupting CD44 gene exon 1 using CRISPR/Cas9 technology could silence CD44 expression, potentially suppressing tumor growth. This study aimed to establish CD44 knockout pancreatic cancer cell lines, assess their impact on tumor growth, and investigate the potential of CD44 knockout as a targeted therapy. The results suggest that CD44 is essential for pancreatic cancer cell tumor growth and that CD44 knockout could be a viable therapeutic strategy. Additionally, we identified a novel oncogenic CD44-Xist axis in pan
The human pancreatic carcinoma cell line Panc-1 was obtained from the American Tissue Culture Collection (Manassas, VA, United States) and the human pancreatic carcinoma cell line Patu-8988 was sourced from the Cell Resource Centre, Shanghai Academy of Life Sciences (Shanghai, China). Fetal bovine serum (FBS), DMEM, and blasticidin were purchased from Thermo Scientific (Gaithersburg, MD, United States). DNA ligase was obtained from Annoron (Beijing, China). Antibodies against CD44, Erk1/2, phosphorylated Erk1/2 (T202/Y204), AKT, phosphorylated AKT (T308), p53, phosphorylated p53, p38, and phosphorylated γH2AX were acquired from Cell Signaling Technology (Boston, MA, United States). The antibody against phosphorylated p38 (Thr180/Tyr182) was sourced from Proteintech (Wuhan, China). The β-actin antibody was purchased from Sigma (Darmstadt, Germany). An RNA reverse transcription kit was obtained from Vazyme (Nanjing, China). The Cas9-expression-Lenti-Cas9-Blast vector and the Lenti-guide-puro-IRES-GFP vector were sourced from Addgene (#52962, Shanghai, China).
The human pancreatic cancer cell lines Panc-1 and Patu-8988 were cultured in DMEM (high glucose) supplemented with 100 mL/L heat-inactivated FBS at 37 °C in a humidified atmosphere containing 50 mL/L CO2, as previously reported[27,28].
CRISPR/Cas9 technology was employed to knock out the CD44 gene in pancreatic cancer cells. Initially, Patu-8988 and Panc-1 cells were transfected with the Lenti-Cas9 cDNA-Blast vector and stably selected using 10 g/L blasticidin. Single-guide RNA (sgRNA) sequence targeting the CD44 gene were selected from the CRISPR library using a CRISPR design tool (http://www.crisprscan.org/). CD44-sgRNA1 (GCTACTTCAGACAACCACA) and CD44-sgRNA2 (CGCTAC
Equal amounts of protein were loaded into each lane and resolved by SDS-PAGE using Tris-glycine running buffer. The separated proteins were subsequently transferred to nitrocellulose membranes. The membranes were blocked with 50 g/L non-fat milk and incubated with primary antibodies against CD44, H2AX, AKT, ERK, p53, or p38 at 4 °C overnight. Next, the membranes were incubated with HRP-coupled secondary antibody for 1 hour at room temperature. The blots were visualized using enhanced chemiluminescence detection reagents and exposed to X-ray film.
Total RNA was extracted from pancreatic cancer cells using Trizol (Vazyme, China). Subsequently, 1 μg RNA was employed to synthesize cDNA using reverse transcriptase (Vazyme, Nanjing, China). The resulting cDNA was then used for PCR analysis. RT-PCR was conducted in each tube with a 20 µL PCR reaction mixture. The PCR protocol included an initial denaturation step at 94 °C for 3 minutes, followed by 22-35 cycles of denaturation at 94 °C for 30 seconds, annealing at 60 °C for 30 seconds, and extension at 72 °C for 1 minute. The PCR products were analyzed using 10 g/L agarose gel electrophoresis. Gene expression levels were normalized to β-actin. All reactions were conducted in triplicate using ABI QuantStudio 6 (Applied Biosystems, United States).
Pancreatic cancer cells were seeded in six-well plates at a density of 5 × 105 cells per well and incubated with 2 mL DMEM supplemented with 100 mL/L FBS for 48 hours. Cells were then scraped with a 200 μL plastic pipette tip, with the time noted as 0 hour. The wells were subsequently washed with PBS to remove any floating cells. Following this, the cells were incubated in fresh DMEM supplemented with 20 mL/L FBS for 24 hours and imaged. The number of migrated cells was counted under a light microscope.
Pancreatic cancer cell invasion was assessed using a cell Transwell assay. The upper chamber of the Transwell was first coated with 100 mL/L Matrigel and incubated at 37 °C for 30 minutes. Subsequently, 2 × 104 cells were suspended in serum-free medium and seeded into the upper chamber. DMEM medium supplemented with 200 mL/L FBS was then added to the lower chambers of the Transwell. After 24 hours, the cells on the upper membrane surface were gently scraped off with a cotton swab. The invaded cells on the sub-membrane surface of the upper Transwell were stained and quantified under a microscope as described previously[27].
The genomic DNA fragment of the 5' flanking promoter region of the Xist gene was cloned into the pGL4.17 luciferase reporter vector. Pancreatic cancer cells were transfected with the pGL4.17-Xist promoter construct or the control pGL4.17 vector using Lipofectamine® 2000 reagent (Invitrogen). Transfected cells were incubated at 37 °C for 48 hours. Luciferase activity in cell lysates was subsequently measured using a Dual-Luciferase Reporter Assay Kit (Promega, Madison, WI, United States).
Tumor xenograft experiments were conducted in mice in accordance with the protocols approved by the Institutional Animal Care and Use Committee of Soochow University (protocol code 202206A0253 and date of approval 06/10/2022). The methods for the tumor xenograft mouse experiment were the same as previously reported[27,28]. Eight-week-old female NOD-SCID mice (18-22 g) were randomly divided into two groups (n = 8). Each mouse was subcutaneously injected on the back with 1 × 107CD44 knockout or control Panc-1 cells in 200 μL PBS. The weight of each mouse was recorded every other day, and the tumor volume was monitored using a digital caliper. The tumor volume was calculated according to the following formula: Tumor volume = 0.5 × length × width2. After 67 days of xenograft, the mice were imaged, and the tumors and main organs were collected, weighed, and statistically analyzed.
All results are presented as the mean ± SD. The experiments were repeated in triplicate. Differences between the two groups were assessed by one-way ANOVA using GraphPad Prism 5. Pairwise statistical comparisons were performed using a student’s t-test. The significance of differences is indicated as follows: aP < 0.05, bP < 0.01.
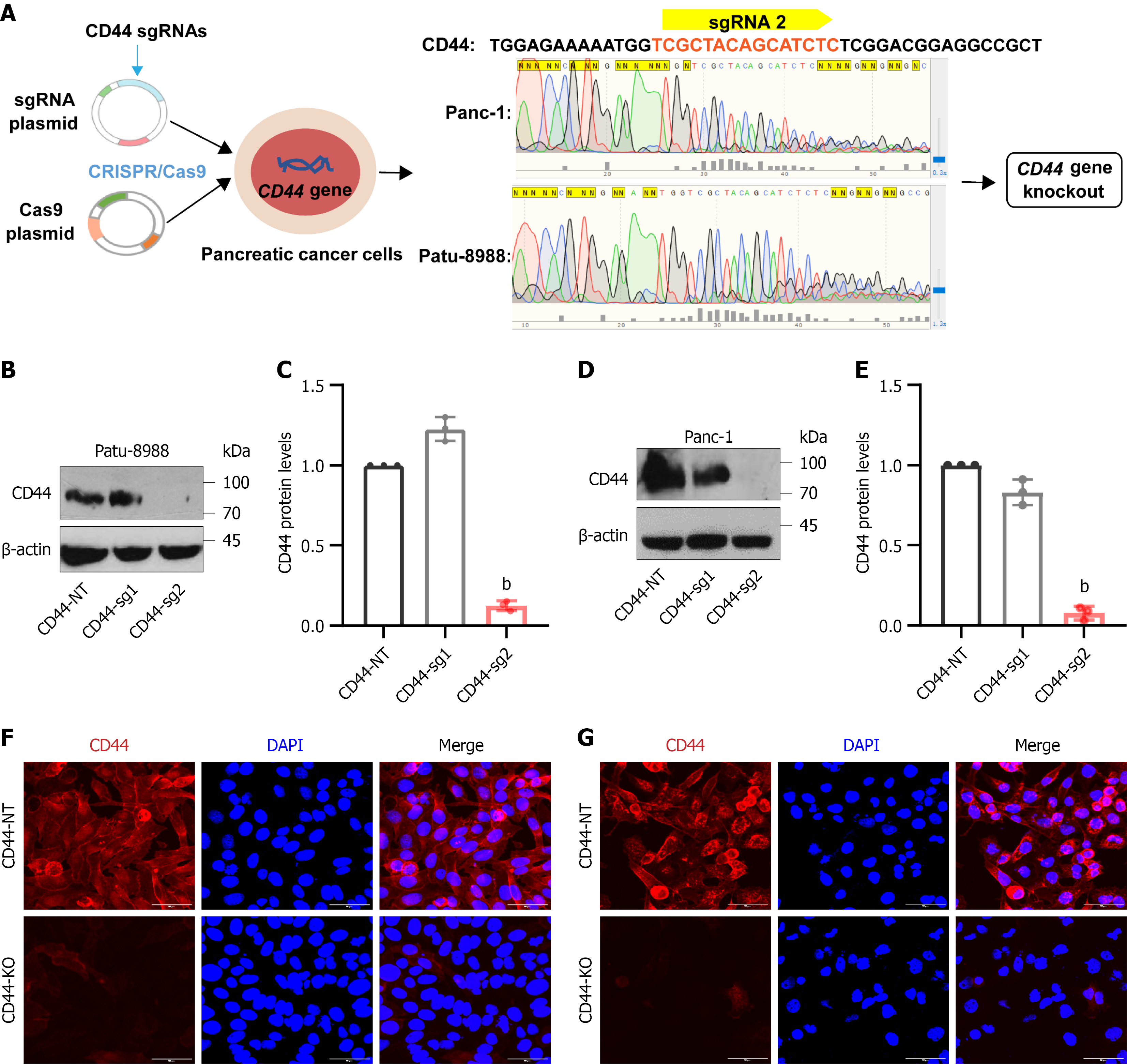
Because CD44 knockout in pancreatic cells has not been reported, we used CRISPR/Cas9 technology to create CD44 knockout pancreatic cancer cell lines (Figure 1A). To accomplish this, we screened a sgRNA library and identified two CD44 sgRNAs with the highest targeting scores for the CD44 gene, named CD44-sgRNA1 (GCTACTTCAGACAA
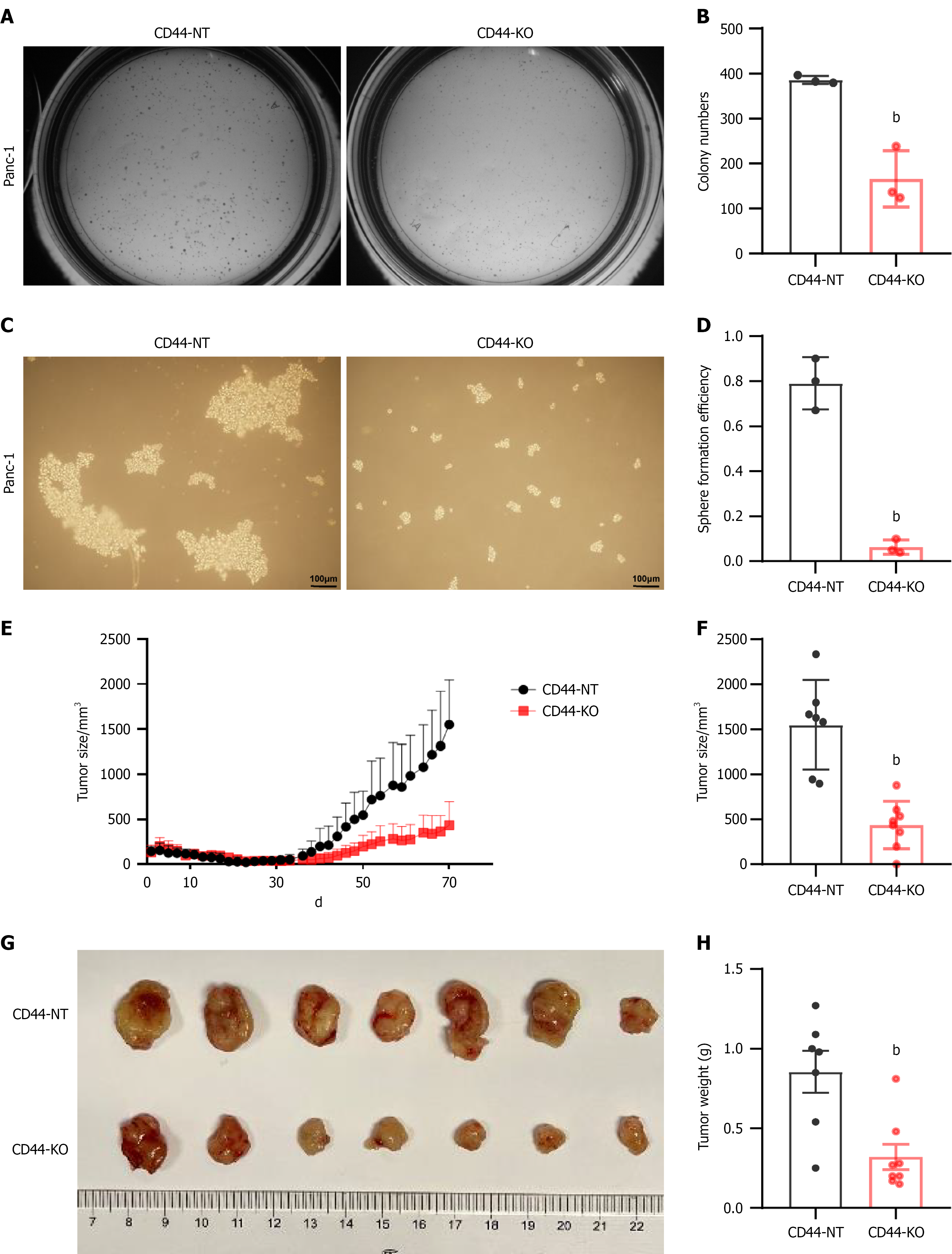
To investigate the impact of CD44 gene knockout on pancreatic cancer cell tumor growth, migration, and invasion, we performed several assays. Soft agar colony formation assays revealed a significant reduction in the colony number of CD44-KO Panc-1 cells compared to control Panc-1 cells (Figure 2A and B), indicating that CD44 knockout diminished tumor growth. In addition, cell sphere formation assays showed that CD44-KO Panc-1 cells formed smaller colonies than CD44-expressing Panc-1 cells (Figure 2C and D), indicating a decrease in cell stemness. In vivo studies showed that the tumor growth rate and weight were significantly lower in mice transplanted with CD44-KO Panc-1 cells compared to control mice transplanted with control Panc-1 cells (Figure 2E-H), further supporting the inhibitory effects of CD44 knockout on tumor growth and stemness in pancreatic cancer cells.
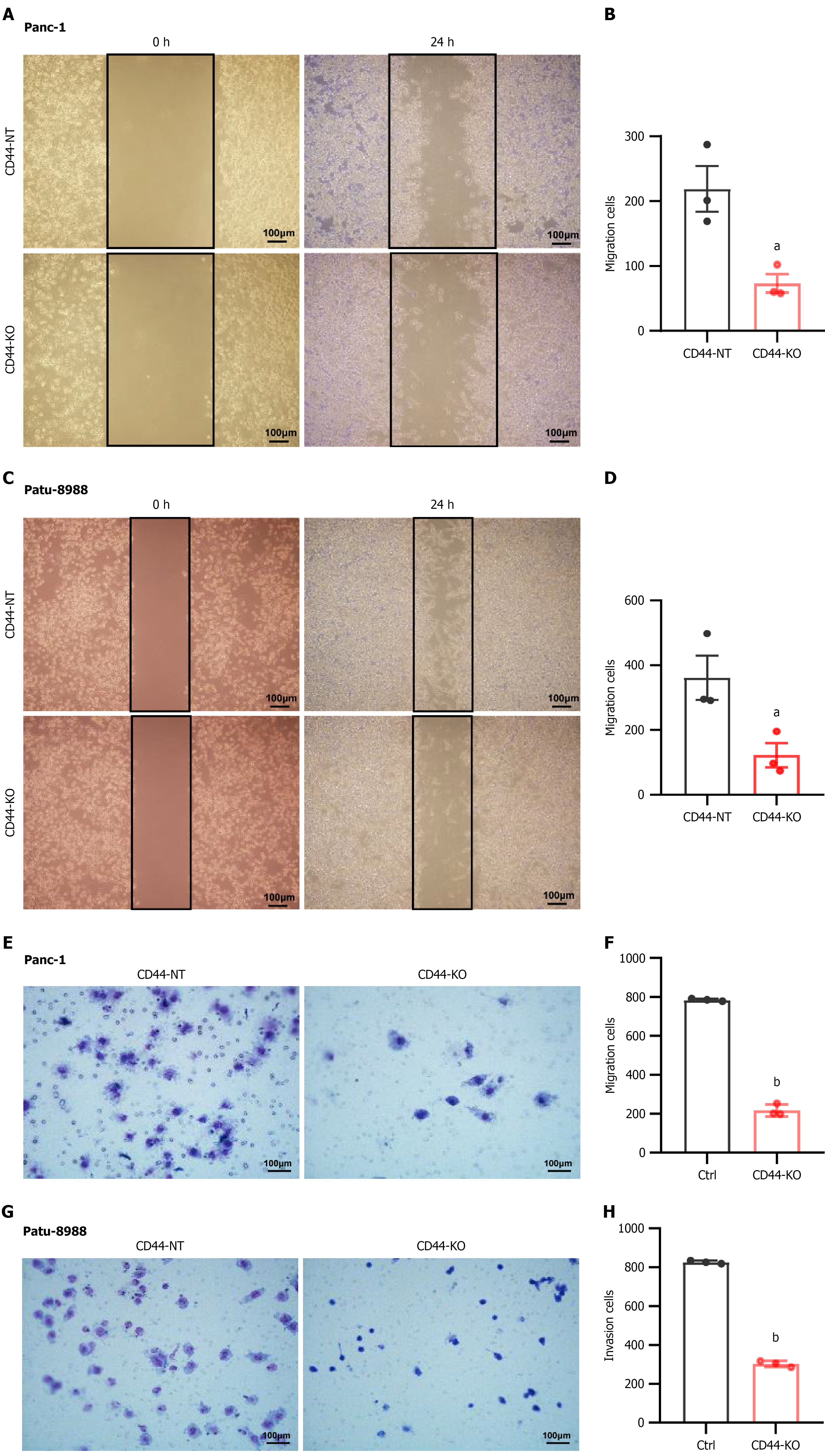
Given the high motility and metastatic potential of pancreatic cancer cells, we also examined the impacts of CD44 knockout on cell migration and invasion. Cell migration assays demonstrated reduced migration in CD44-KO Panc-1 (Figure 3A and B) and Patu-8988 (Figure 3C and D) cells, while cell invasion assays showed notable inhibition of invasion in both CD44-KO Panc-1 (Figure 3E and F) and Patu-8988 (Figure 3G and H) cells. These findings indicate that CD44 knockout effectively inhibited cancer cell migration and invasion.
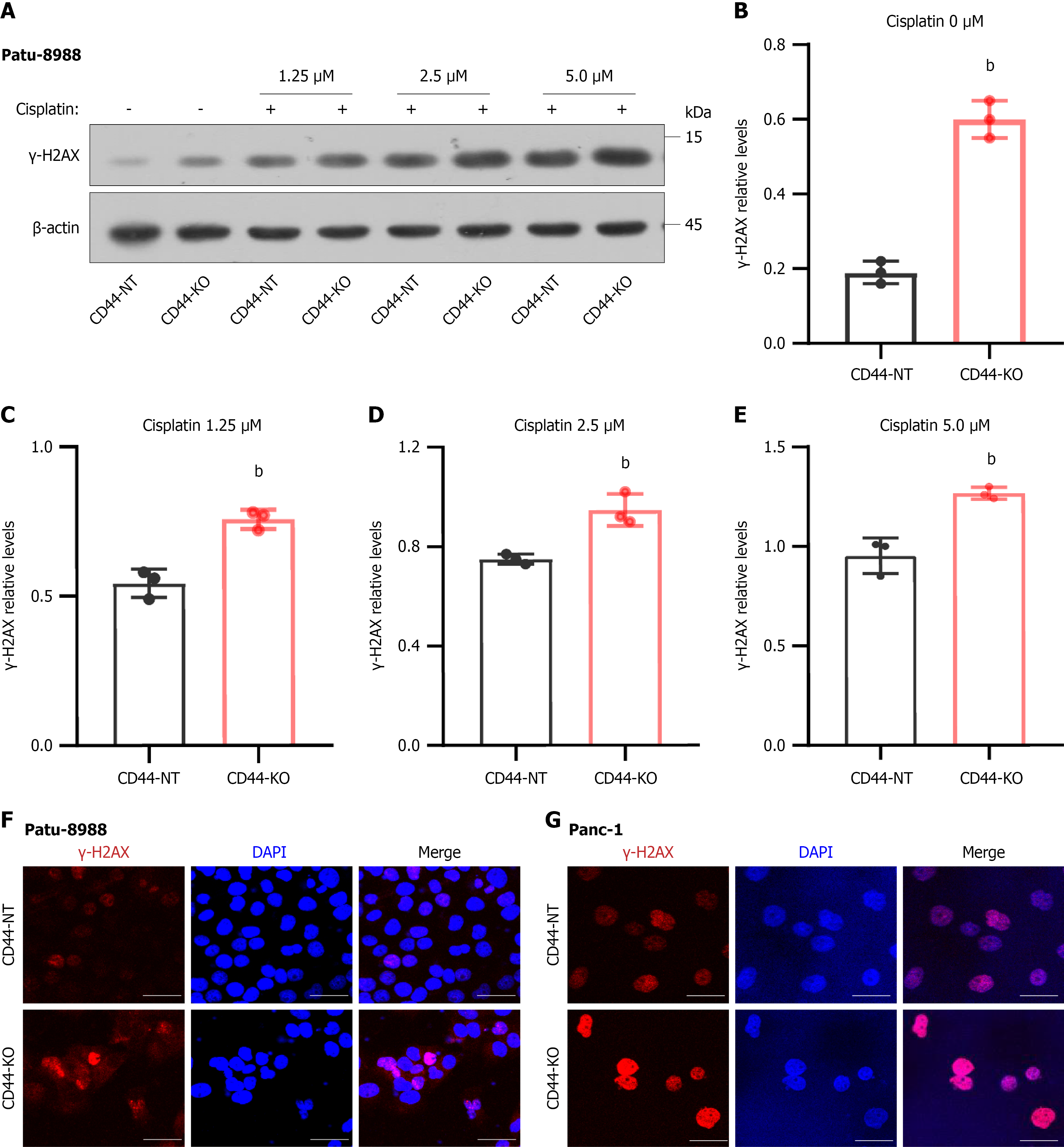
Cisplatin is a widely used anti-cancer drug that induces DNA damage, but it can have toxic effects on normal cells and can lead to drug resistance in pancreatic cancer. To address this issue, we investigated whether CD44 knockout could sensitize pancreatic cancer cells to cisplatin-induced DNA damage. Western blotting and immunofluorescence analysis revealed significantly higher levels of the DNA damage marker γH2AX in CD44-KO Panc-1 cells compared to CD44-NT, even without cisplatin treatment (Figure 4A-F). Moreover, CD44 knockout further increased γH2AX levels in Panc-1 cells in the presence of cisplatin (Figure 4G), indicating that CD44 knockout enhanced the sensitivity of pancreatic cancer cells to cisplatin.
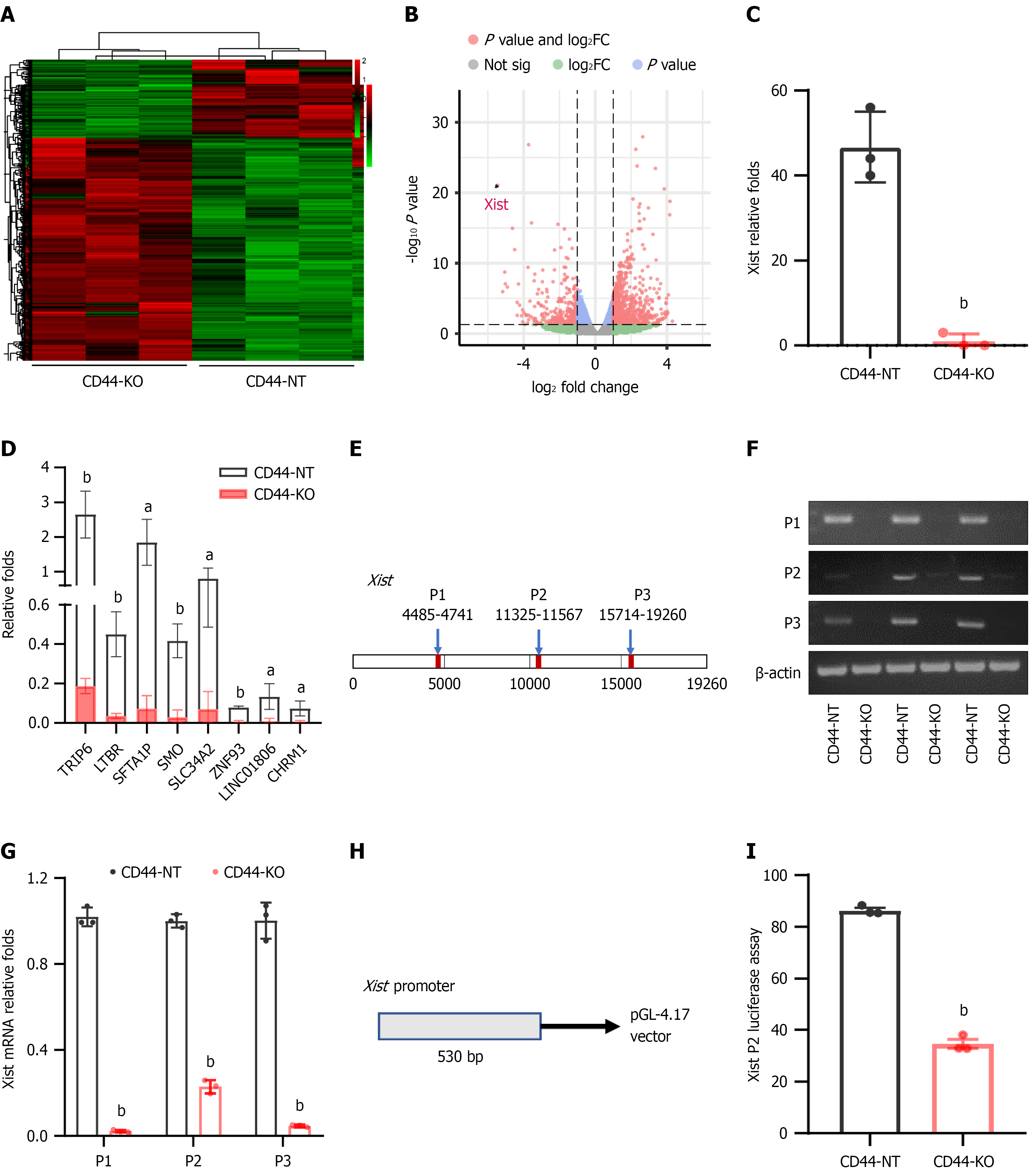
CD44 genetic disruption markedly reduces X-inactive specific transcript expression in pancreatic cancer cells: We performed RNA sequencing (RNA-Seq) to analyze the gene expression profiles of CD44-KO and control Panc-1 cells. The results showed that CD44 gene knockout significantly altered the expression levels of 734 genes (> 2-fold, P < 0.05); among them, 544 genes were upregulated and 190 genes were downregulated (Figure 5A and B). Subsequent analyses focused on the top differentially expressed oncogenic genes in pancreatic cancer cells. Strikingly, the expression level of Xist, an emerging oncogenic long non-coding RNA consisting of 1290 bp, was reduced by 43.6-fold in CD44-KO compared to CD44-NT Panc-1 cells (Figure 5C). In addition, the expression levels of other oncogenic genes, such as SFTA1P, UGT8, ZNF93, CHRM1, SMO, TRIP6, and LTBR, were also significantly decreased (Figure 5D).
Given that the overexpression of Xist in tumor cells triggers tumorigenesis and cancer progression, we designed three PCR primer pairs to further confirm the CD44 knockout-mediated reduction of Xist expression in pancreatic cancer cells (Figure 5E). RT-PCR showed that CD44 knockout almost completely inhibited Xist expression in pancreatic cancer cells (Figure 5F), and RT-qPCR showed that CD44 knockout markedly reduced the Xist expression by 35-fold in pancreatic cancer cells (Figure 5G). This indicates that CD44 knockout effectively reduced Xist expression in cancer cells.
We next investigated the mechanism underlying the CD44 knockout-mediated reduction of Xist expression by studying Xist gene promoter activity. We initially predicted the putative promoter region of the Xist gene using JASPER software, and then we cloned a genomic DNA fragment of Xist (530 bp) with a predicted high promoter score into the gene transcription reporter vector pGL4.17 (Figure 5H). Next, Panc-1 cells were transfected with the Xist gene promoter DNA fragment-pGL4.17 or control pGL4.17. Luciferase assays showed that the Xist promoter activity was significantly reduced in CD44-KO Panc-1 cells compared with CD44-NT Panc-1 cells (Figure 5I), suggesting that CD44 knockout reduced Xist promoter activity, thereby diminishing its gene transcription.
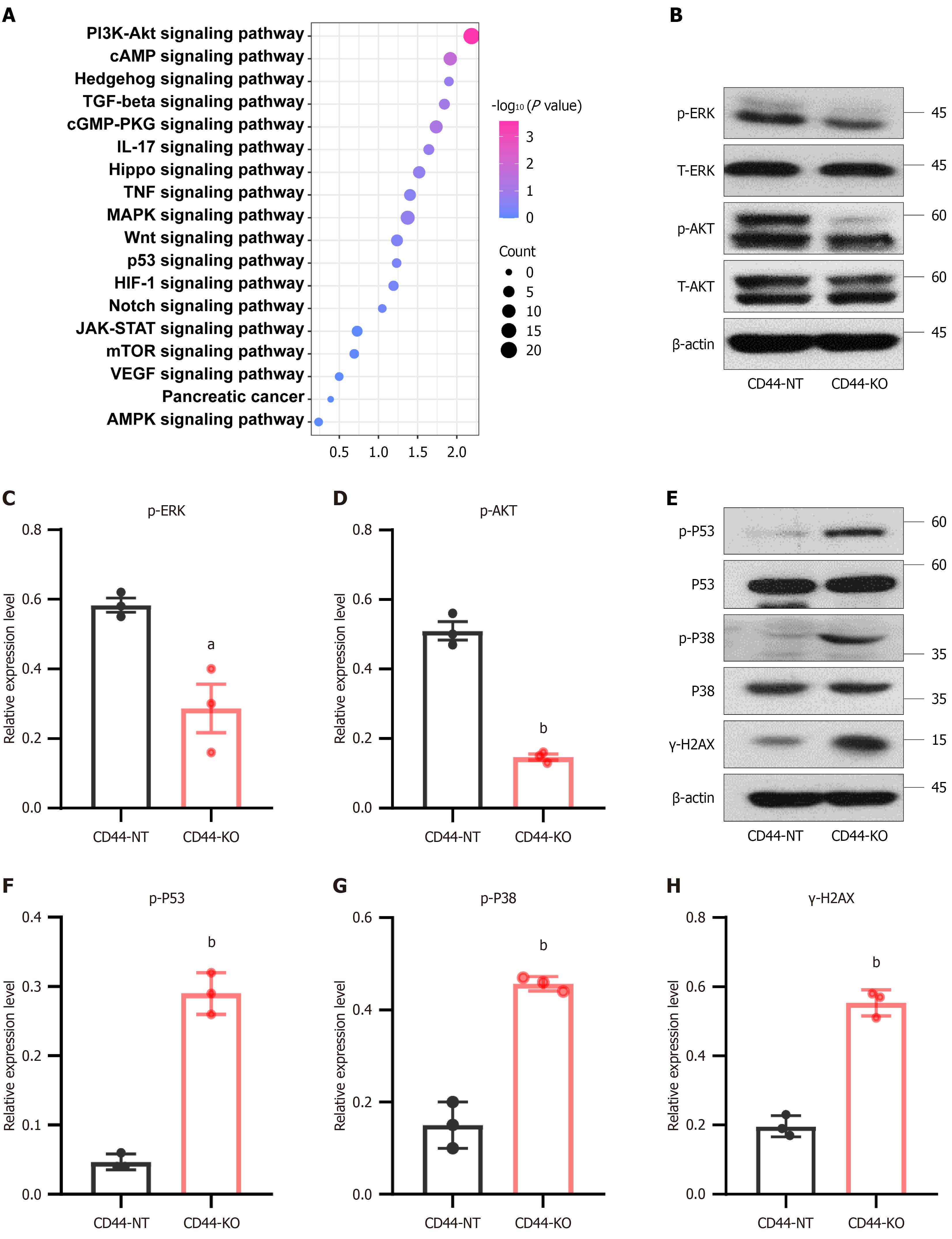
CD44 genetic disruption inhibits tumorigenesis signaling pathways but activates tumor-suppressive signaling pathways in pancreatic cancer cells: The RNA-Seq results indicated that CD44 knockout affected the expression of 734 genes in pancreatic cancer cells. Following this, we performed Kyoto Encyclopedia of Genes and Genomes signaling pathway analysis to assess the effect of CD44 knockout on cell signaling in pancreatic cancer cells. The results showed that CD44 knockout affected multiple signaling pathways, including DNA repair, PI3K-AKT, Hedgehog, Ras, MAPK, and Wnt (Figure 6A). Western blotting showed that CD44 knockout significantly reduced phosphorylated ERK and phosphorylated AKT protein levels (Figure 6B-D); notably, the 60 kDa band of the phosphorylated AKT protein was markedly diminished (Figure 6B). These data suggest that CD44 knockout inhibited tumorigenic AKT and ERK signaling pathways in pancreatic cancer. On the other hand, CD44 knockout increased the levels of the tumor-suppressive phosphorylated p53 (Figure 6E and F) and phosphorylated p38 proteins (Figure 6G) in pancreatic cancer cells. This suggests that CD44 knockout activated the tumor-suppressive p38-p53 signaling pathway in pancreatic cancer cells.
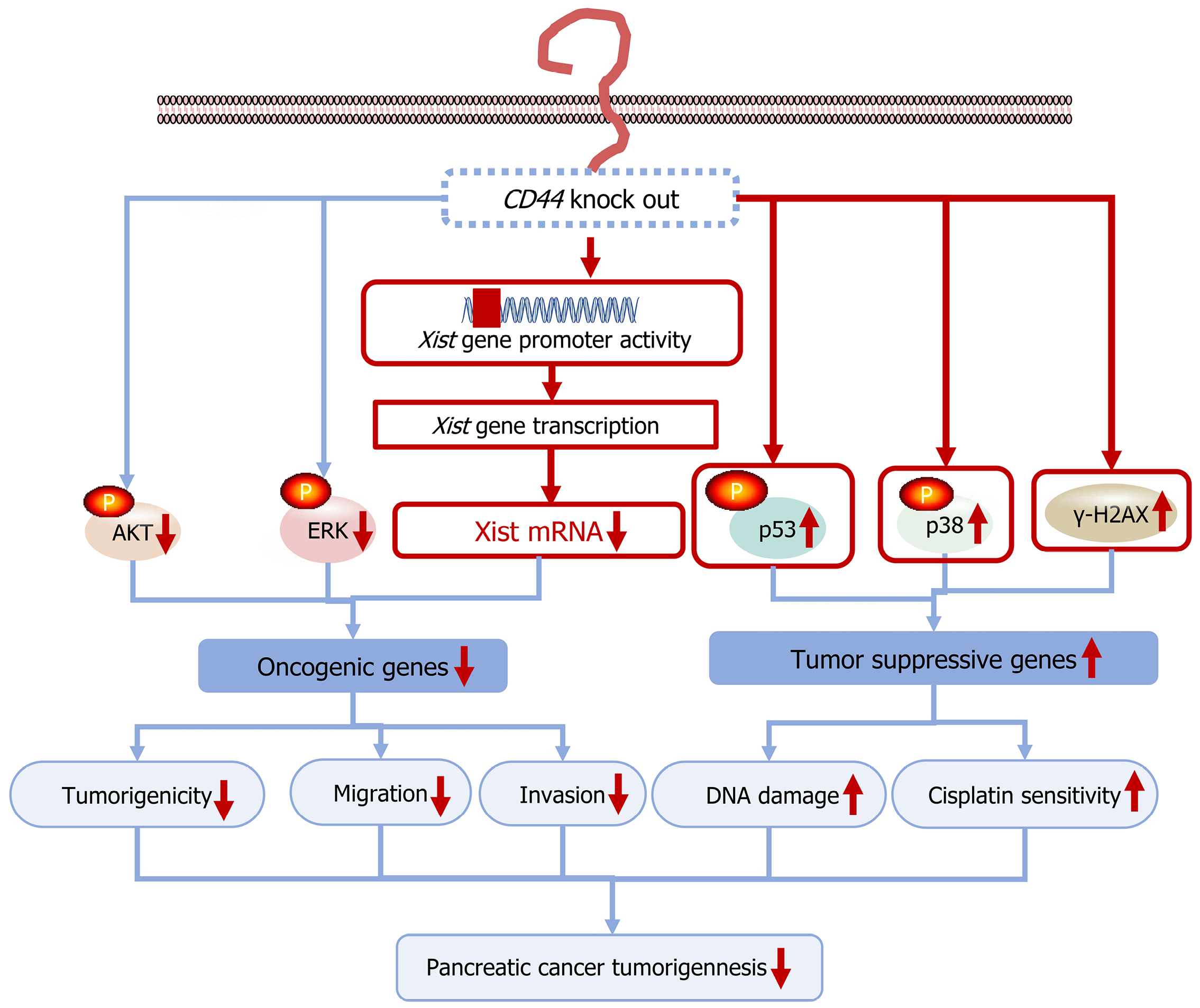
Additionally, CD44 knockout notably elevated the levels of the DNA damage marker γH2AX protein (Figure 6H), suggesting that these cells experienced DNA breakage even in the absence of a DNA damage inducer. In summary, CD44 knockout was found to inhibit pancreatic cancer cell tumor growth, migration, and invasion, and suppress tumor growth in xenograft mice. Additionally, CD44 knockout induced pancreatic cancer cell DNA damage and sensitized cancer cells to cisplatin. Mechanistically, CD44 knockout inhibited the expression of Xist and various other oncogenic genes, suppressed the tumorigenic ERK and AKT signaling pathways, and simultaneously promoted the tumor-suppressive p38-p53 signaling pathway in pancreatic cancer cells (Figure 7). These findings highlight the critical role of CD44 in pancreatic cancer cell tumorigenesis and provide a new approach for targeted pancreatic cancer therapy.
Although great efforts have been made in fighting pancreatic cancer in recent decades, the 5-year survival rate of pancreatic cancer patients is only 8%; thus, pancreatic cancer still poses a serious threat to human life and health[1,2]. The main reasons for these dire statistics include our limited understanding of the mechanisms behind pancreatic cancer growth and the lack of effective therapies currently available in the clinical setting[3,4]. Moreover, pancreatic tumor tissues are rich in cancer stem/progenitor cells that trigger tumorigenesis, metastasis, and drug resistance, and thus overexpress CD44[5-7]. In the current study, we found that CD44 knockout in pancreatic cancer cells effectively inhibited tumor growth, migration, and invasion, primarily through the downregulation of Xist, suppression of the tumorigenic AKT and ERK signaling pathways, and activation of the tumor-suppressive p38-p53 signaling pathway. These findings indicate that CRISPR/Cas9-mediated CD44 knockout is a new option for targeted pancreatic cancer therapy.
Xist is a long non-coding RNA abnormally overexpressed in various types of malignant tumors[16,17]. Overexpression of Xist upregulates the expression of oncogenic EGFR, YAP, ZEB1, TGF-β, Notch1, and Notch2, promotes cancer cell proliferation, migration, and invasion, and triggers the initiation and development of many diseases, including pancreatic cancer[16-23]. As a key driver of tumorigenesis and progression, the EGFR signaling pathway regulates cell proliferation, survival, and metastasis through crucial downstream pathways, such as the RAS-MAPK and PI3K-AKT pathways. Its aberrant activation is common across many solid tumors[19,23,29,30].
While the oncogenic role of Xist and its link to EGFR are known[19,31], the regulatory mechanism of Xist expression is unclear, and a method that effectively inhibits the expression of oncogenic Xist in cancer has not yet been reported in the literature. In this study, CD44 knockout markedly suppressed Xist expression in pancreatic cancer cells through the enhancement of its gene promoter activity in cancer cells. To our knowledge, this is the first report of a novel CD44-Xist axis in malignant tumors. These findings highlight the importance of the CD44-Xist axis in tumorigenesis, providing a new method to effectively reduce oncogenic Xist expression in cancer, and offering a new platform for the development of therapies inhibiting Xist in malignant tumors.
Notably, we found that CD44 knockout concurrently suppressed tumorigenic AKT and ERK signaling while activating the tumor-suppressive p38-p53 pathway. The concurrent inhibition of AKT/ERK and activation of p53 presents a significant therapeutic challenge. Integrating our findings with the literature, we propose that the observed effects on AKT, ERK, and p53 are likely mediated, at least in part, by the CD44-induced downregulation of Xist and subsequent EGFR inhibition. The role of EGFR in activating the AKT and ERK pathways has been extensively described in the literature, supporting the view that an attenuation of EGFR signaling would weaken these pathways. While the direct interaction between Xist/EGFR and the p53 pathway is complex and context-dependent, EGFR activation has been linked to p53 suppression in some cancers[32], suggesting that reducing EGFR signaling could potentially restore p53 function.
Importantly, CD44 knockout increased pancreatic cancer cell DNA damage and cisplatin sensitivity. This is consistent with the role of AKT and ERK in DNA repair and chemoresistance[33], and the pro-apoptotic and DNA repair functions of p53. Therefore, AKT/ERK suppression and p53 activation by CD44 knockout could synergistically contribute to enhanced chemosensitivity.
CRISPR/Cas9 technology has recently been used in preclinical and primary clinical trials for the treatment of diseases and has shown promising therapeutic results[25,26]. In this study, CD44-sgRNA2 (CGCTACAGCATCTCTCGGA) and CRISPR/Cas9-mediated CD44 genomic disruption inhibited pancreatic cancer cell tumorigenesis. These findings demonstrate that CD44 gene knockout is a potential targeted therapy for pancreatic cancer. The combination of CRISPR/Cas9 and nano-micelle technologies is an attractive method for CD44-targeted cancer therapy. CD44-sgRNA2 (CGCTACAGCATCTCTCGGA) and Cas9 could be concurrently packed in HA-liposome nanoparticles, as described in recent studies[25,26]. In this manner, CD44-sgRNA2 and Cas9 could be delivered to cancer tissues in vivo. Here, HA can bind to CD44 on the surface of cancer stem/progenitor cells, and the nanoparticles can enter cancer cells, releasing CD44-sgRNA2 and Cas9, consequently disrupting the CD44 gene in vivo in cancer cells, therefore increasing the efficacy of this therapy.
In conclusion, CD44 knockout inhibited pancreatic cancer cell migration and invasion, and suppressed tumor growth in xenograft mice. Mechanistically, CD44 knockout markedly reduced oncogenic Xist expression, suppressed the tumorigenic AKT and ERK signaling pathways, and concurrently activated key tumor-suppressive p38 and p53 signaling pathways. CD44 knockout also increased pancreatic cancer cell DNA damage and sensitivity to the anti-cancer drug cisplatin. These findings highlight the critical role of CD44 in pancreatic cancer pathogenesis and provide a new approach for targeted pancreatic cancer therapy.
We sincerely thank Meng-Li Zhang, Jun-Tao Liu, Yu Chen, Xu Li, Xiao-Xiao Song, and Peng Xu for their expert guidance and assistance in gene-editing techniques during this study.
| 1. | Siegel RL, Kratzer TB, Giaquinto AN, Sung H, Jemal A. Cancer statistics, 2025. CA Cancer J Clin. 2025;75:10-45. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1828] [Cited by in RCA: 2057] [Article Influence: 2057.0] [Reference Citation Analysis (4)] |
| 2. | Yu J, Yang X, He W, Ye W. Burden of pancreatic cancer along with attributable risk factors in Europe between 1990 and 2019, and projections until 2039. Int J Cancer. 2021;149:993-1001. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 105] [Article Influence: 21.0] [Reference Citation Analysis (0)] |
| 3. | Halbrook CJ, Lyssiotis CA, Pasca di Magliano M, Maitra A. Pancreatic cancer: Advances and challenges. Cell. 2023;186:1729-1754. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1043] [Cited by in RCA: 910] [Article Influence: 303.3] [Reference Citation Analysis (2)] |
| 4. | Laface C, Memeo R, Maselli FM, Santoro AN, Iaia ML, Ambrogio F, Laterza M, Cazzato G, Guarini C, De Santis P, Perrone M, Fedele P. Immunotherapy and Pancreatic Cancer: A Lost Challenge? Life (Basel). 2023;13:1482. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 18] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 5. | Xia P, Liu DH. Cancer stem cell markers for liver cancer and pancreatic cancer. Stem Cell Res. 2022;60:102701. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 22] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 6. | Sharma R, Malviya R. Cancer Stem Cells in Carcinogenesis and Potential Role in Pancreatic Cancer. Curr Stem Cell Res Ther. 2024;19:1185-1194. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 7. | Zhao Y, Qin C, Zhao B, Wang Y, Li Z, Li T, Yang X, Wang W. Pancreatic cancer stemness: dynamic status in malignant progression. J Exp Clin Cancer Res. 2023;42:122. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 22] [Reference Citation Analysis (0)] |
| 8. | Dzobo K, Sinkala M. Cancer Stem Cell Marker CD44 Plays Multiple Key Roles in Human Cancers: Immune Suppression/Evasion, Drug Resistance, Epithelial-Mesenchymal Transition, and Metastasis. OMICS. 2021;25:313-332. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 49] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 9. | Hassn Mesrati M, Syafruddin SE, Mohtar MA, Syahir A. CD44: A Multifunctional Mediator of Cancer Progression. Biomolecules. 2021;11:1850. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 387] [Cited by in RCA: 328] [Article Influence: 65.6] [Reference Citation Analysis (5)] |
| 10. | Li XP, Zhang XW, Zheng LZ, Guo WJ. Expression of CD44 in pancreatic cancer and its significance. Int J Clin Exp Pathol. 2015;8:6724-6731. [PubMed] |
| 11. | Ohara Y, Oda T, Sugano M, Hashimoto S, Enomoto T, Yamada K, Akashi Y, Miyamoto R, Kobayashi A, Fukunaga K, Morishita Y, Ohkohchi N. Histological and prognostic importance of CD44(+) /CD24(+) /EpCAM(+) expression in clinical pancreatic cancer. Cancer Sci. 2013;104:1127-1134. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 50] [Cited by in RCA: 62] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 12. | Xiaoping L, Xiaowei Z, Leizhen Z, Weijian G. Expression and significance of CD44 and p-AKT in pancreatic head cancer. World J Surg Oncol. 2015;13:334. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 22] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 13. | Xu H, Niu M, Yuan X, Wu K, Liu A. CD44 as a tumor biomarker and therapeutic target. Exp Hematol Oncol. 2020;9:36. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 65] [Cited by in RCA: 276] [Article Influence: 46.0] [Reference Citation Analysis (0)] |
| 14. | Sato N, Kohi S, Hirata K, Goggins M. Role of hyaluronan in pancreatic cancer biology and therapy: Once again in the spotlight. Cancer Sci. 2016;107:569-575. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 112] [Cited by in RCA: 118] [Article Influence: 11.8] [Reference Citation Analysis (0)] |
| 15. | Placencio-Hickok VR, Lauzon M, Moshayedi N, Guan M, Kim S, Nissen N, Lo S, Pandol S, Larson BK, Gong J, Hendifar AE, Osipov A. Hyaluronan heterogeneity in pancreatic ductal adenocarcinoma: Primary tumors compared to sites of metastasis. Pancreatology. 2022;22:92-97. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 10] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 16. | Ghafouri-Fard S, Dashti S, Farsi M, Taheri M, Mousavinejad SA. X-Inactive-Specific Transcript: Review of Its Functions in the Carcinogenesis. Front Cell Dev Biol. 2021;9:690522. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 16] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 17. | Zhou J, Chen J, Chen Z, Wu G, Zhou Z, Wu T, Wang W, Luo Y, Liu T. Prognostic significance of long non-coding RNA five prime to XIST in various cancers. BMC Cancer. 2022;22:61. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 5] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 18. | Yang J, Qi M, Fei X, Wang X, Wang K. Long non-coding RNA XIST: a novel oncogene in multiple cancers. Mol Med. 2021;27:159. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 49] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 19. | Wei W, Liu Y, Lu Y, Yang B, Tang L. LncRNA XIST Promotes Pancreatic Cancer Proliferation Through miR-133a/EGFR. J Cell Biochem. 2017;118:3349-3358. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 139] [Cited by in RCA: 170] [Article Influence: 18.9] [Reference Citation Analysis (0)] |
| 20. | Sun Z, Zhang B, Cui T. Long non-coding RNA XIST exerts oncogenic functions in pancreatic cancer via miR-34a-5p. Oncol Rep. 2018;39:1591-1600. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 36] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 21. | Sun J, Zhang Y. LncRNA XIST enhanced TGF-β2 expression by targeting miR-141-3p to promote pancreatic cancer cells invasion. Biosci Rep. 2019;39:BSR20190332. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 42] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 22. | Shen J, Hong L, Yu D, Cao T, Zhou Z, He S. LncRNA XIST promotes pancreatic cancer migration, invasion and EMT by sponging miR-429 to modulate ZEB1 expression. Int J Biochem Cell Biol. 2019;113:17-26. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 68] [Article Influence: 9.7] [Reference Citation Analysis (0)] |
| 23. | Zou L, Chen FR, Xia RP, Wang HW, Xie ZR, Xu Y, Yu JH, Wang KH. Long noncoding RNA XIST regulates the EGF receptor to promote TGF-β1-induced epithelial-mesenchymal transition in pancreatic cancer. Biochem Cell Biol. 2020;98:267-276. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 17] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 24. | Ledford H. Super-precise CRISPR tool enters US clinical trials for the first time. Nature. 2023;621:667-668. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 7] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 25. | Huang K, Zapata D, Tang Y, Teng Y, Li Y. In vivo delivery of CRISPR-Cas9 genome editing components for therapeutic applications. Biomaterials. 2022;291:121876. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 33] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 26. | Noormohamadi H, Soleimani Samarkhazan H, Kargar M, Maroufi F, Servatian N, Davami F. CRISPR/Cas technologies in pancreatic cancer research and therapeutics: recent advances and future outlook. Discov Oncol. 2025;16:1530. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 7] [Reference Citation Analysis (2)] |
| 27. | Yang B, Zhang B, Cao Z, Xu X, Huo Z, Zhang P, Xiang S, Zhao Z, Lv C, Meng M, Zhang G, Dong L, Shi S, Yang L, Zhou Q. The lipogenic LXR-SREBF1 signaling pathway controls cancer cell DNA repair and apoptosis and is a vulnerable point of malignant tumors for cancer therapy. Cell Death Differ. 2020;27:2433-2450. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 32] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 28. | Qi J, Meng M, Liu J, Song X, Chen Y, Liu Y, Li X, Zhou Z, Huang X, Wang X, Zhou Q, Zhao Z. Lycorine inhibits pancreatic cancer cell growth and neovascularization by inducing Notch1 degradation and downregulating key vasculogenic genes. Biochem Pharmacol. 2023;217:115833. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 11] [Reference Citation Analysis (0)] |
| 29. | Mucciolo G, Araos Henríquez J, Jihad M, Pinto Teles S, Manansala JS, Li W, Ashworth S, Lloyd EG, Cheng PSW, Luo W, Anand A, Sawle A, Piskorz A, Biffi G. EGFR-activated myofibroblasts promote metastasis of pancreatic cancer. Cancer Cell. 2024;42:101-118.e11. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 120] [Cited by in RCA: 138] [Article Influence: 69.0] [Reference Citation Analysis (0)] |
| 30. | Lv Y. Research Progress on the Role of EGFR in Tumorigenesis, Development and Treatment. Adv Clin Med. 2024;14:796-803. [DOI] [Full Text] |
| 31. | Hong W, Xiong Z, Wang X, Liao X, Liu M, Jiang Z, Min D, Li J, Guo G, Fu Z. Long noncoding RNA XIST promotes cell proliferation and migration in diabetic foot ulcers through the miR-126-3p/EGFR axis. Diabetol Metab Syndr. 2024;16:35. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 8] [Reference Citation Analysis (0)] |
| 32. | Ding J, Li X, Khan S, Zhang C, Gao F, Sen S, Wasylishen AR, Zhao Y, Lozano G, Koul D, Alfred Yung WK. EGFR suppresses p53 function by promoting p53 binding to DNA-PKcs: a noncanonical regulatory axis between EGFR and wild-type p53 in glioblastoma. Neuro Oncol. 2022;24:1712-1725. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 31] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 33. | Alemi F, Raei Sadigh A, Malakoti F, Elhaei Y, Ghaffari SH, Maleki M, Asemi Z, Yousefi B, Targhazeh N, Majidinia M. Molecular mechanisms involved in DNA repair in human cancers: An overview of PI3k/Akt signaling and PIKKs crosstalk. J Cell Physiol. 2022;237:313-328. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 31] [Article Influence: 6.2] [Reference Citation Analysis (0)] |