Published online Apr 27, 2026. doi: 10.4254/wjh.v18.i4.117456
Revised: January 20, 2026
Accepted: March 3, 2026
Published online: April 27, 2026
Processing time: 134 Days and 22.7 Hours
Chronic unconjugated hyperbilirubinemia requires careful differentiation bet
To establish a convenient nomogram model to distinguish chronic unconjugated hyperbilirubinemia associated with UGT1A1 gene mutation from hemolytic diseases.
In this retrospective study, patients diagnosed with chronic unconjugated hy
A total of 429 patients (357 with UGT1A1 mutation-associated group, 72 with hemolytic disease-associated group) were enrolled. Patients diagnosed from January 2022 to December 2024 were randomly divided into training (n = 265) and internal validation (n = 114) cohorts. Patients diagnosed from January 2025 to May 2025 (n = 50) were used for external validation. Four key variables - abnormality of peripheral blood smear, hematocrit, red cell distribution width standard deviation, and reticulocyte percentage - were selected to construct the nomogram. External validation yielded an area under the receiver operating characteristic curve of 0.986, sensitivity of 100%, specificity of 90%, area under the precision-recall curve of 0.938, and F1-score of 0.833. Calibration curves showed good agreement between predicted and actual outcomes. Decision curve analysis confirmed the clinical utility of this nomogram.
We developed an effective nomogram model for the differential diagnosis of UGT1A1 gene mutation-associated and hemolytic disease-associated unconjugated hyperbilirubinemia, which improves clinical preliminary scr
Core Tip: Clinical characteristic overlap and the need for complex specialized testing make it challenging to distinguish uridine diphosphate glucuronosyltransferase 1A1 mutations from hemolytic diseases in patients with chronic unconjugated hyperbilirubinemia. In a genetically confirmed cohort, we applied least absolute shrinkage and selection operator and logistic regression to identify four variables for differential diagnosis: Abnormal peripheral blood smear, hematocrit, red cell distribution width standard deviation, and reticulocyte percentage. These variables were used to establish a nomogram model, which achieved an area under the receiver operating characteristic curve of 0.986 and an area under precision-recall curve of 0.938. This model provides a simple, interpretable, and cost-effective tool for early screening and triage, potentially reducing the need for complex testing.
- Citation: Yu HT, Li MH, Tang S, Liang C, Sheng DC, Jiang H, Dong JX, Hou W, Zheng SJ. Novel nomogram for differential diagnosis of UGT1A1 gene mutation-associated unconjugated hyperbilirubinemia with hemolytic diseases. World J Hepatol 2026; 18(4): 117456
- URL: https://www.wjgnet.com/1948-5182/full/v18/i4/117456.htm
- DOI: https://dx.doi.org/10.4254/wjh.v18.i4.117456
Chronic unconjugated hyperbilirubinemia is a common condition encountered in liver outpatient clinics. After excluding liver diseases, clinicians must differentiate between unconjugated hyperbilirubinemia caused by uridine diphosphate glucuronosyltransferase 1A1 (UGT1A1) gene mutations (Gilbert’s and Crigler-Najjar’s syndromes) and that caused by hemolytic diseases. The management of these two disorders differ markedly. Specially, UGT1A1 gene mutation-asso
Accurately distinguishing between UGT1A1 mutation-associated and hemolytic disease-associated unconjugated hyperbilirubinemia poses several clinical challenges. Firstly, the clinical and biochemical profiles of these two etiologies show significant heterogeneity and partial overlap. For instance, some patients with Gilbert’s syndrome may have mildly elevated reticulocyte count (RET)[5]. In contrast, hemoglobin level in certain well-compensated patients with chronic hemolysis may remain within the normal range, thus blurring diagnostic boundaries. Secondly, comprehensive screening for the etiology of hemolysis is time-consuming and complex, often requiring specialized tests, such as high-performance liquid chromatography, hemoglobin electrophoresis, qualitative and quantitative red blood cell enzyme assays, osmotic fragility testing, and genetic testing. Unfortunately, these tests are challenging to perform routinely in many primary care settings. Moreover, current screening indices lack specificity and sensitivity. Lactate dehydrogenase (LDH) elevation is highly sensitive for suggesting hemolysis but lacks specificity, as levels can also increase in liver disease, malignancy, and muscle injury. Decreased haptoglobin (HP) is specific for intravascular hemolysis but has limited sensitivity because its synthesis increases during infection or inflammation, potentially masking hemolysis-induced consumption[6]. Relying on a single indicator for initial screening can easily lead to missed or incorrect diagnoses. Given these limitations, developing a differential diagnostic model that integrates routine clinical parameters is necessary for early, precise, and convenient screening for hemolytic disease in patients with unconjugated hyperbilirubinemia. This study aims to develop a differential diagnosis model based on routine clinical indicators to facilitate early distinction between UGT1A1 mutation-associated and hemolytic disease-associated unconjugated hyperbilirubinemia, enabling prompt and appropriate patient management.
This retrospective study assessed patients diagnosed with chronic unconjugated hyperbilirubinemia at Beijing YouAn Hospital from January 2022 to May 2025. All participants must meet the following inclusion criteria: Serum total bilirubin (TBiL) > 17.1 μmol/L, and the ratio of indirect bilirubin (IBiL) to TBiL is greater than 50%. Exclusion criteria were: (1) Coexisting acute or chronic liver disease, defined as alanine aminotransferase, aspartate aminotransferase, gamma-glutamyl transpeptidase levels exceeding 2 times the upper limit of normal, alkaline phosphatase exceeding 1.5 times the upper limit of normal, or abdominal ultrasound indicating liver cirrhosis, widened portal vein, or collateral circulation; and (2) Missing key clinical indicators.
The diagnosis of UGT1A1 mutation-associated unconjugated hyperbilirubinemia, encompassing both Gilbert’s syndrome and Crigler-Najjar’s syndrome type II in this study, is primarily based on shared biochemical and genetic criteria. Both conditions require at least two serum TBiL elevations, spaced more than 6 months apart, with each elevation predominantly in indirect (unconjugated) bilirubin. Critically, both diagnoses require excluding hemolytic diseases and the absence of concurrent elevations in liver enzymes (such as alanine aminotransferase, aspartate aminotransferase, alkaline phosphatase, or gamma-glutamyl transpeptidase). The definitive distinction lies in the severity of the hyperbilirubinemia: Gilbert’s syndrome is characterized by mild elevations of TBiL (> 1 time the upper limit of normal, typically 17.1-102.6 μmol/L), while Crigler-Najjar’s syndrome type II presents with significantly higher TBiL levels (102.6-342.0 μmol/L)[7]. Finally, the diagnosis is confirmed by genetic testing, which shows the presence of at least two UGT1A1 gene variants.
In this study, hemolytic disease-associated hyperbilirubinemia etiologies included autoimmune hemolytic anemia, hereditary spherocytosis, hereditary stomatocytosis, α-thalassemia, and hemolysis of unknown etiology. Their respective diagnostic criteria as follows: Autoimmune hemolytic anemia: (1) Hemoglobin < 120 g/L for males and < 110 g/L for females; (2) Reduced HP (< 250 mg/L), elevated TBiL (≥ 17.1 μmol/L, predominantly IBiL), elevated LDH, and reticulocyte percentage (RET%) > 4% or RET > 120 × 109/L; and (3) Detection of red blood cell autoantibodies[8]. Hereditary spherocytosis: (1) Clinical manifestations of anemia, jaundice, splenomegaly, and cholelithiasis; (2) Family history of hereditary spherocytosis; (3) Elevated RET, elevated IBiL, numerous spherocytes on peripheral blood smear, and positive osmotic fragility test or eosin-5’-maleimide binding test; and (4) Genetic testing confirming pathogenic variants in hereditary spherocytosis-associated genes[9]. Hereditary stomatocytosis: (1) Clinical manifestations of anemia, jaundice, splenomegaly, and cholelithiasis; (2) Increased stomatocytes on peripheral blood smear; and (3) Genetic testing confirming pathogenic variants in hereditary stomatocytosis-associated genes[10]. α-thalassemia: (1) Clinical manifestations of anemia, jaundice, splenomegaly, and cholelithiasis; (2) Red blood cell parameter analysis showing microcytic hypochromic changes; (3) Peripheral blood smear showing hypochromic, poikilocytic red blood cells, often increased reticulocytes, and increased mature red blood cells; (4) Hemoglobin composition analysis showing decreased hemoglobin A2, and in some cases, hemoglobin H and hemoglobin Bart’s detectable; and (5) Genetic testing confirmed pathogenic variants in the hemoglobin A1 and hemoglobin A2 genes[11]. In this study, hemolysis of unknown etiology is defined as: Consistent with the clinical diagnosis of hemolysis[12], with the etiology remaining unknown. The detailed criteria are as follows: (1) RET > 4%; (2) Presence of one or more of the following: Clinical manifestations of anemia and jaundice, peripheral blood smear showing increased fragmented or abnormal red blood cells, or reduced HP levels; and (3) The specific etiology of hemolysis remains unclear.
Patients with Gilbert’s syndrome and Crigler-Najjar’s syndrome type II were grouped into the “UGT1A1 gene mutation-associated unconjugated hyperbilirubinemia group” (UGT1A1 mutation group). Patients with the above hemolytic diseases were grouped into the “hemolytic disease-associated unconjugated hyperbilirubinemia group” (hereafter referred to as the hemolysis group).
We collected comprehensive clinical data from participants, including past medical history, family history, and demographic information, alongside detailed laboratory, abdominal ultrasound, and genetic testing results. The laboratory parameters included a complete blood count, RET and percentage, peripheral blood smear, Coombs test, LDH, serum liver and renal function tests, and HP. An abnormal peripheral blood smear result was defined as marked anisocytosis or presence of numerous abnormal red cells, including nucleated red blood cells, spherocytes, elliptocytes, stomatocytes, target cells, teardrop cells, and schistocytes. UGT1A1 variants were confirmed by mass spectrometry-based nucleic acid detection, Sanger sequencing, or whole-exome sequencing. Hereditary hemolysis diagnoses were confirmed by whole-exome sequencing.
Patients diagnosed from January 2022 to December 2024 were randomly divided into training and internal validation groups in a 7:3 ratio. Patients diagnosed from January 2025 to May 2025 were enrolled in the external validation cohort. Missing indicators (< 30% missing rate) were imputed using a random forest model. Non-normally distributed continuous variables were described using the median and interquartile range and compared using the Wilcoxon rank-sum test. Categorical variables were described using n (%) and compared using the χ2 test.
Cutoff values derived from receiver operating characteristic curves were utilized to convert nonlinear continuous variables into binary categorical variables to enhance model interpretability. The least absolute shrinkage and selection operator regression was used to screen for differential indicators, with the optimal λ value selected as the one standard error above the minimum on the 10-fold cross-validation plot. Variance inflation factor < 5 confirmed no multicollinearity. Multivariate logistic regression with backward stepwise selection was used to build a nomogram.
The discrimination, calibration, and clinical applicability of the nomogram were evaluated using the receiver operating characteristic curve, precision-recall (PR) curve, calibration curve, Hosmer-Lemeshow test, and decision curve analysis. Statistical analysis was performed using SPSS 29.0 and R 4.5.0, with P < 0.05 considered statistically significant.
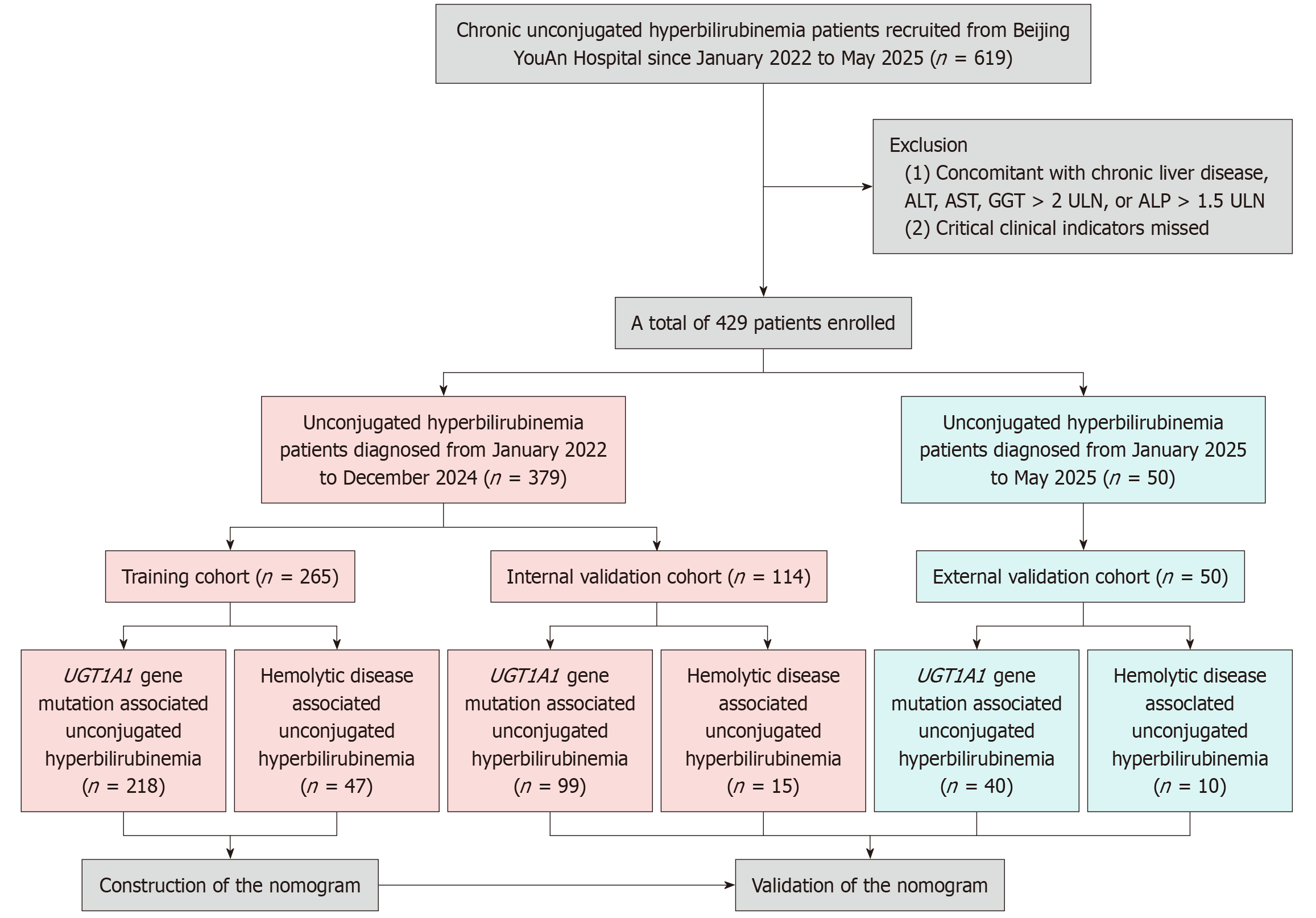
A total of 429 patients were ultimately enrolled in this study. The training and internal validation cohorts consisted 317 patients in the UGT1A1 mutation group (306 with Gilbert’s syndrome and 11 with Crigler-Najjar’s syndrome type II) and 62 patients in the hemolysis group. The hemolysis group comprised 36 cases of hereditary spherocytosis, 8 cases of autoimmune hemolytic anemia, 4 cases of hereditary stomatocytosis, 1 case of thalassemia, 1 case of BPGM gene mutation-associated erythrocytosis, and 12 cases of hemolysis of unknown etiology. The patients were randomly divided into a training cohort (n = 265, including 218 UGT1A1 mutation group cases and 47 hemolysis group cases) and an internal validation cohort (n = 114, including 99 UGT1A1 mutation group cases and 15 hemolysis group cases). The external validation cohort included 40 patients in the UGT1A1 mutation group (all with Gilbert’s syndrome) and 10 patients in the hemolysis group (3 with hereditary spherocytosis and 7 with hemolysis of unknown etiology) (Figure 1).
UGT1A1 variation information of 346 Gilbert’s syndome and 11 Crigler-Najjar’s syndome type II patients are in Supplementary Table 1. Among the hemolysis group, 39 patients with congenital hemolytic disease provided genetic variation information, as presented in Supplementary Table 2. Eight patients with autoimmune hemolytic anemia were clinically diagnosed, and 6 patients with hereditary spherocytosis were clinically diagnosed based on a positive family history and abnormal hematological parameters. The remaining 19 patients had hemolysis of unknown etiology.
We collected a total of 51 clinical indicators, and 48 of these (with a missing rate of < 30%) were imputed using a random forest method for subsequent analysis (Supplementary Table 3). Table 1 showed differences in clinical characteristics between the UGT1A1 mutation group and the hemolysis group in the training and internal validation cohorts. Both patients groups were predominantly male. The median age of patients in the UGT1A1 mutation group of the training cohort was 39 (32-52) years and 36 (21.25-48) years in the hemolysis group (P = 0.035). The hemolysis group had significantly higher levels of TBiL (94.1 μmol/L vs 35.8 μmol/L, P < 0.001), IBiL/TBiL ratio (0.86 vs 0.69, P < 0.001), higher incidence of cholelithiasis (59.68% vs 6.62%, P < 0.001), and higher incidence of splenomegaly (85.48% vs 7.26%, P < 0.001). In terms of red blood cell-associated indicators, the hemolysis group had significantly lower level of red blood cell count, hemoglobin, hematocrit (HCT), and higher level of mean corpuscular volume, mean corpuscular hemoglobin, mean corpuscular hemoglobin concentration, red cell distribution width, red cell distribution width-standard deviation (RDWSD), RET, RET%, and abnormal peripheral blood smear (all P < 0.001). No significant differences were observed in the clinical indicators between the training and internal validation cohorts (Supplementary Table 4). The patient features of the external validation cohort are shown in Supplementary Table 5.
| Variables | Total (n = 379) | UGT1A1 mutation group (n = 317) | Hemolysis group | Statistic | P value |
| Age (years) | 39.00 (31.00, 51.00) | 39.00 (32.00, 52.00) | 36.00 (21.25, 48.00) | Z = -2.11 | 0.035 |
| Sex, male | 260 (68.60) | 219 (69.09) | 41 (66.13) | χ2 = 0.21 | 0.646 |
| Cholelithiasis | 58 (15.30) | 21 (6.62) | 37 (59.68) | χ2 = 112.61 | < 0.001 |
| Splenomegaly | 76 (20.05) | 23 (7.26) | 53 (85.48) | χ2 = 197.95 | < 0.001 |
| WBC (× 109/L) | 5.57 (4.71, 6.53) | 5.56 (4.70, 6.50) | 5.67 (4.76, 6.72) | Z = -0.66 | 0.510 |
| PLT (× 109/L) | 209.00 (172.00, 243.00) | 211.00 (176.00, 244.00) | 185.00 (150.25, 235.00) | Z = -2.63 | 0.009 |
| RBC (× 1012/L) | 4.81 (4.38, 5.21) | 4.93 (4.55, 5.25) | 3.44 (2.96, 4.15) | Z = -9.49 | < 0.001 |
| HGB (g/L) | 147.00 (134.00, 160.00) | 151.00 (139.00, 162.00) | 111.00 (93.25, 135.75) | Z = -9.07 | < 0.001 |
| HCT (%) | 43.30 (39.30, 46.50) | 44.40 (41.40, 46.90) | 31.55 (27.55, 38.15) | Z = -9.92 | < 0.001 |
| MCV (fL) | 90.60 (88.10, 93.20) | 90.30 (87.90, 92.40) | 93.80 (90.08, 99.20) | Z = -5.26 | < 0.001 |
| MCH (pg) | 30.90 (30.00, 31.85) | 30.70 (29.90, 31.60) | 32.80 (31.02, 34.40) | Z = -6.63 | < 0.001 |
| MCHC (g/L) | 340.00 (334.00, 348.00) | 339.00 (333.00, 346.00) | 349.00 (339.00, 358.50) | Z = -4.18 | < 0.001 |
| RDW (%) | 12.40 (12.00, 13.00) | 12.24 (11.90, 12.60) | 17.30 (14.93, 21.32) | Z = -10.98 | < 0.001 |
| RDWSD (fL) | 40.90 (39.40, 43.40) | 40.40 (39.00, 41.90) | 56.30 (49.92, 68.20) | Z = -11.15 | < 0.001 |
| RET (× 109/L) | 68.80 (57.60, 94.45) | 66.60 (55.60, 78.70) | 225.05 (147.75, 364.07) | Z = -11.98 | < 0.001 |
| RET percentage (%) | 1.43 (1.19, 1.96) | 1.37 (1.14, 1.58) | 7.04 (4.19, 11.25) | Z = -12.12 | < 0.001 |
| ALT (U/L) | 20.00 (15.00, 28.00) | 20.00 (15.00, 28.00) | 17.50 (12.00, 37.12) | Z = -1.07 | 0.285 |
| AST (U/L) | 23.00 (18.50, 28.85) | 23.00 (19.00, 28.00) | 21.50 (17.00, 31.00) | Z = -0.10 | 0.924 |
| TBiL (μmol/L) | 40.00 (30.10, 59.10) | 35.80 (28.60, 47.10) | 94.10 (78.95, 131.02) | Z = -10.59 | < 0.001 |
| DBiL (μmol/L) | 11.30 (8.75, 14.75) | 10.80 (8.50, 13.90) | 14.75 (10.00, 20.00) | Z = -4.94 | < 0.001 |
| IBiL (μmol/L) | 27.10 (20.30, 43.15) | 24.90 (19.70, 32.90) | 82.05 (54.78, 115.62) | Z = -10.33 | < 0.001 |
| IBiL/TBiL ratio (%) | 0.70 (0.65, 0.76) | 0.69 (0.65, 0.74) | 0.86 (0.74, 0.91) | Z = -7.11 | < 0.001 |
| ALB (g/L) | 46.50 (44.60, 48.30) | 46.60 (44.90, 48.30) | 45.56 (42.17, 48.20) | Z = -2.61 | 0.009 |
| GLB (g/L) | 27.20 (25.10, 29.20) | 27.30 (25.60, 29.55) | 25.05 (23.07, 27.48) | Z = -4.57 | < 0.001 |
| ALP (U/L) | 70.83 (58.00, 86.65) | 70.00 (58.00, 85.00) | 71.95 (58.00, 96.27) | Z = -0.77 | 0.439 |
| GGT (U/L) | 17.00 (13.00, 24.72) | 17.00 (13.00, 23.23) | 20.50 (12.25, 40.50) | Z = -1.61 | 0.108 |
| TBA (μmol/L) | 3.60 (2.30, 6.00) | 3.31 (2.20, 5.30) | 5.80 (3.95, 10.85) | Z = -5.45 | < 0.001 |
| BUN (mmol/L) | 5.00 (4.25, 5.76) | 5.01 (4.24, 5.76) | 4.86 (4.26, 5.99) | Z = -0.11 | 0.911 |
| CR (μmol/L) | 64.00 (55.00, 72.21) | 66.00 (56.00, 73.00) | 55.50 (49.18, 65.86) | Z = -3.96 | < 0.001 |
| GFR [mL/(minute × 1.73 m2)] | 112.61 (103.25, 121.85) | 111.50 (102.80, 119.60) | 120.70 (109.67, 134.77) | Z = -4.30 | < 0.001 |
| UA (μmol/L) | 348.00 (288.57, 402.50) | 337.00 (285.00, 393.00) | 411.89 (330.75, 485.00) | Z = -5.21 | < 0.001 |
| GLU (mmol/L) | 5.20 (4.96, 5.46) | 5.21 (4.99, 5.48) | 5.10 (4.78, 5.44) | Z = -2.08 | 0.038 |
| TG (mmol/L) | 1.00 (0.78, 1.40) | 0.96 (0.77, 1.32) | 1.27 (0.89, 1.75) | Z = -3.31 | < 0.001 |
| TC (mmol/L) | 4.17 (3.63, 4.88) | 4.32 (3.87, 5.03) | 3.00 (2.66, 3.38) | Z = -9.39 | < 0.001 |
| HDL (mmol/L) | 1.20 (1.00, 1.49) | 1.28 (1.11, 1.52) | 0.82 (0.73, 1.02) | Z = -9.48 | < 0.001 |
| LDL (mmol/L) | 2.44 (1.85, 3.09) | 2.58 (2.12, 3.21) | 1.46 (1.20, 1.69) | Z = -9.28 | < 0.001 |
| Spleen wide (mm) | 33.55 (30.87, 39.00) | 32.57 (30.00, 35.02) | 49.04 (41.48, 59.75) | Z = -10.74 | < 0.001 |
| CAP (dB/m) | 236.02 (206.39, 265.35) | 237.00 (207.00, 263.00) | 235.28 (205.75, 265.68) | Z = -0.27 | 0.789 |
| LSM (kPa) | 5.10 (4.40, 6.10) | 5.00 (4.30, 5.90) | 6.52 (5.06, 8.57) | Z = -5.27 | < 0.001 |
| COOMBS | χ2 = 10.78 | 0.001 | |||
| Negative | 352 (92.88) | 301 (94.95) | 51 (82.26) | ||
| Positive | 27 (7.12) | 16 (5.05) | 11 (17.74) | ||
| Abnormal peripheral blood smear | χ2 = 183.42 | < 0.001 | |||
| Normal | 338 (89.18) | 313 (98.74) | 25 (40.32) | ||
| Abnormal | 41 (10.82) | 4 (1.26) | 37 (59.68) |
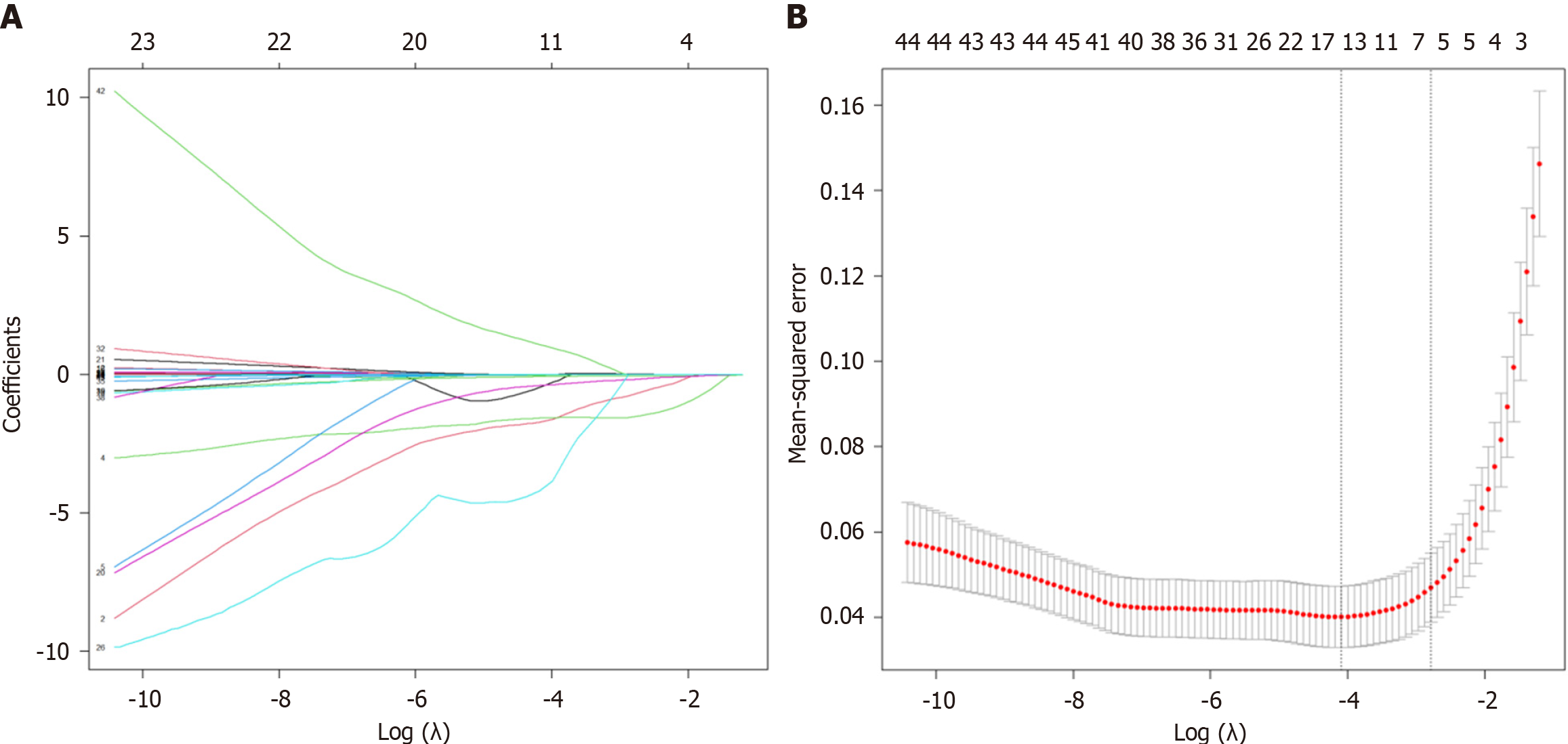
The least absolute shrinkage and selection operator regression and cross-validation initially screened six indicators: Abnormal peripheral blood smear, splenomegaly, HCT, RDWSD, RET%, and IBiL/TBiL ratio (Figure 2). Variance inflation factor analysis confirmed that there was no multicollinearity among these six indicators (Supplementary Table 6). The optimal cutoff values for HCT, RDWSD, and RET% were determined by maximizing the Youden index in receiver operating characteristic curve analysis, as 40.15%, 45.55 fL, and 2.095%, respectively (Table 2). The six indicators were then entered into a multivariate logistic regression with stepwise backward selection, yielding four final indicators: Abnormal peripheral blood smear, HCT, RDWSD, and RET% (all P < 0.05) (Table 3).
| AUC (95%CI) | Accuracy (95%CI) | Sensitivity (95%CI) | Specificity (95%CI) | PPV (95%CI) | NPV (95%CI) | Cut off | |
| HCT | 0.903 (0.839-0.966) | 0.826 (0.776-0.867) | 0.872 (0.748-0.940) | 0.817 (0.760-0.862) | 0.506 (0.400-0.612) | 0.967 (0.931-0.985) | 40.152 |
| RDWSD | 0.953 (0.910-0.997) | 0.955 (0.923-0.974) | 0.872 (0.748-0.940) | 0.972 (0.941-0.987) | 0.872 (0.748-0.940) | 0.972 (0.941-0.987) | 45.55 |
| RET percentage | 0.986 (0.971-1.00) | 0.936 (0.899-0.962) | 0.957 (0.858-0.988) | 0.931 (0.890-0.958) | 0.750 (0.628-0.842) | 0.990 (0.965- 0.997) | 2.095 |
| IBiL/TBiL ratio | 0.776 (0.673-0.857) | 0.830 (0.780-0.871) | 0.660 (0.517-0.778) | 0.867 (0.815- 0.906) | 0.517 (0.393-0.638) | 0.922 (0.877-0.951) | 0.776 |
| Variables | Univariate analysis | Multivariate analysis | ||||||||
| β | SE | Z | P value | OR (95%CI) | β | SE | Z | P value | OR (95%CI) | |
| Splenomegaly | ||||||||||
| No | 1.00 (reference) | |||||||||
| Yes | 4.28 | 0.49 | 8.82 | < 0.001 | 72.14 (27.88-186.68) | |||||
| Peripheral blood smear | ||||||||||
| Normal | 1.00 (reference) | 1.00 (reference) | ||||||||
| Abnormal | 4.66 | 0.65 | 7.14 | < 0.001 | 105.61 (29.37-379.76) | 4.12 | 1.74 | 2.37 | 0.018 | 61.73 (2.05-1854.84) |
| RDWSD | ||||||||||
| < 45.55 fL | 1.00 (reference) | 1.00 (reference) | ||||||||
| ≥ 45.55 fL | 5.49 | 0.6 | 9.11 | < 0.001 | 241.44 (74.19-785.72) | 3.43 | 1.24 | 2.76 | 0.006 | 30.72 (2.71-348.77) |
| RET percentage | ||||||||||
| < 2.1% | 1.00 (reference) | 1.00 (reference) | ||||||||
| ≥ 2.1% | 5.72 | 0.77 | 7.42 | < 0.001 | 304.50 (67.25-1378.84) | 4.43 | 1.21 | 3.65 | < 0.001 | 84.27 (7.82-908.54) |
| HCT | ||||||||||
| < 40.15% | 1.00 (reference) | 1.00 (reference) | ||||||||
| ≥ 40.15% | -3.41 | 0.47 | -7.25 | < 0.001 | 0.03 (0.01-0.08) | -2.16 | 1.07 | -2.02 | 0.043 | 0.11 (0.01-0.94) |
| IBiL/TBiL ratio | 10.2 | 1.67 | 6.09 | < 0.001 | 26851.06 (1009.96-713868.82) | 7.81 | 4.68 | 1.67 | 0.095 | 2457.55 (0.25-23737621.41) |
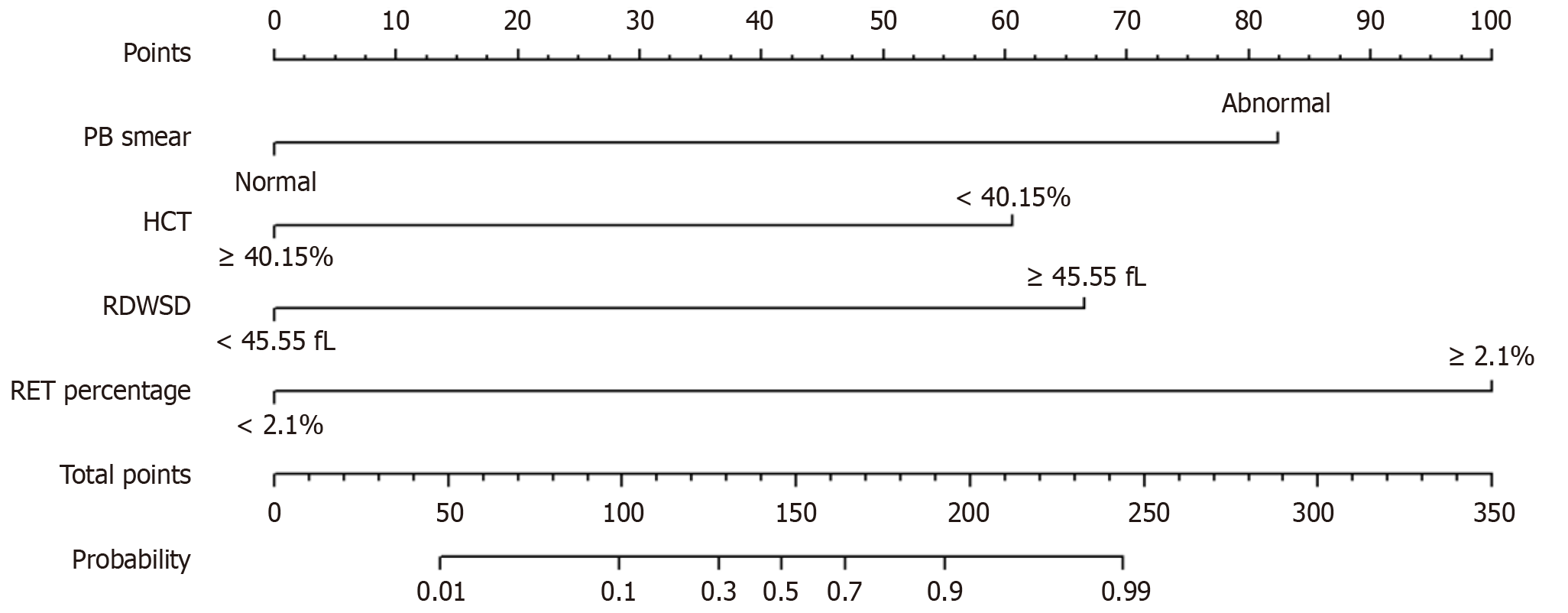
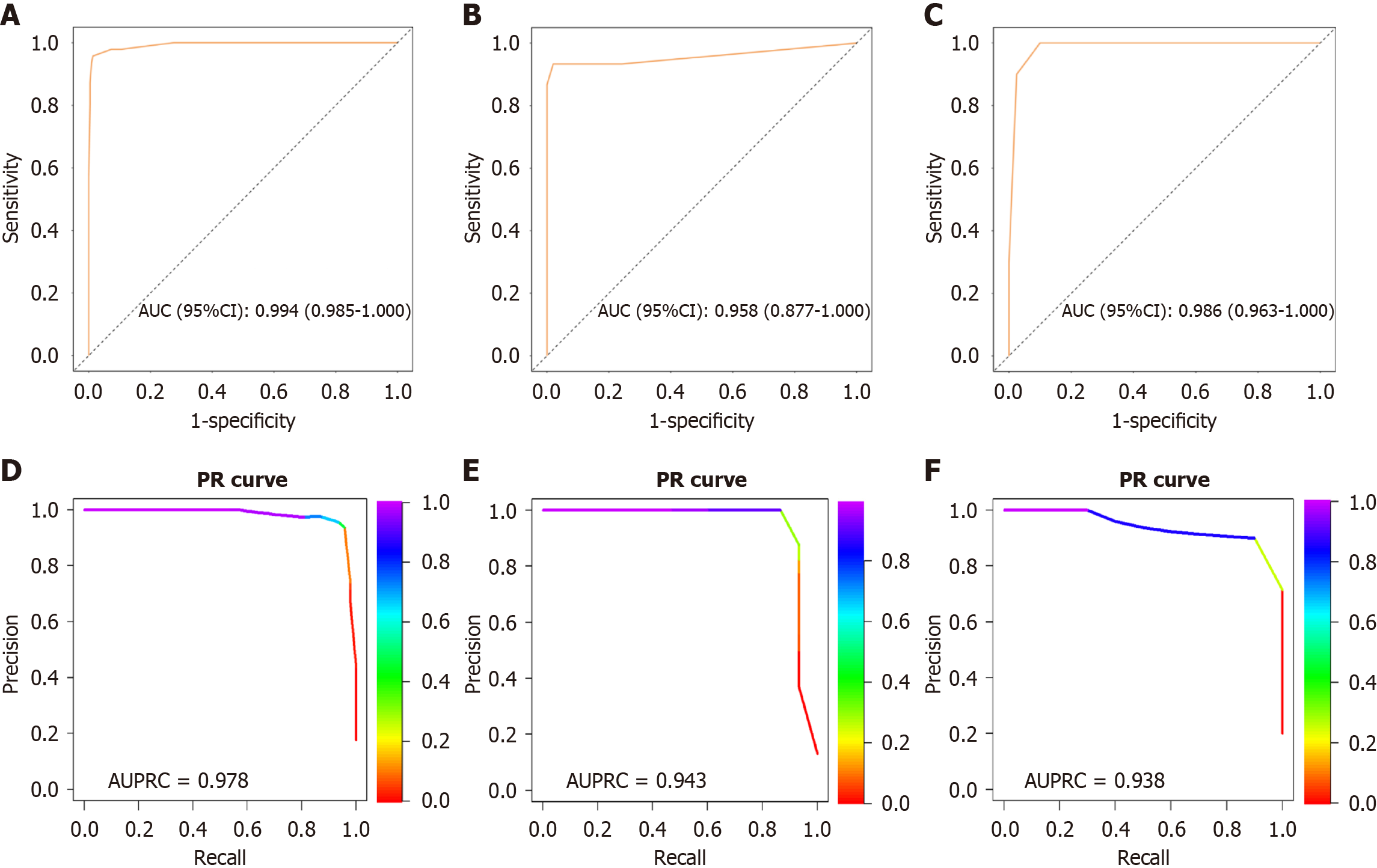
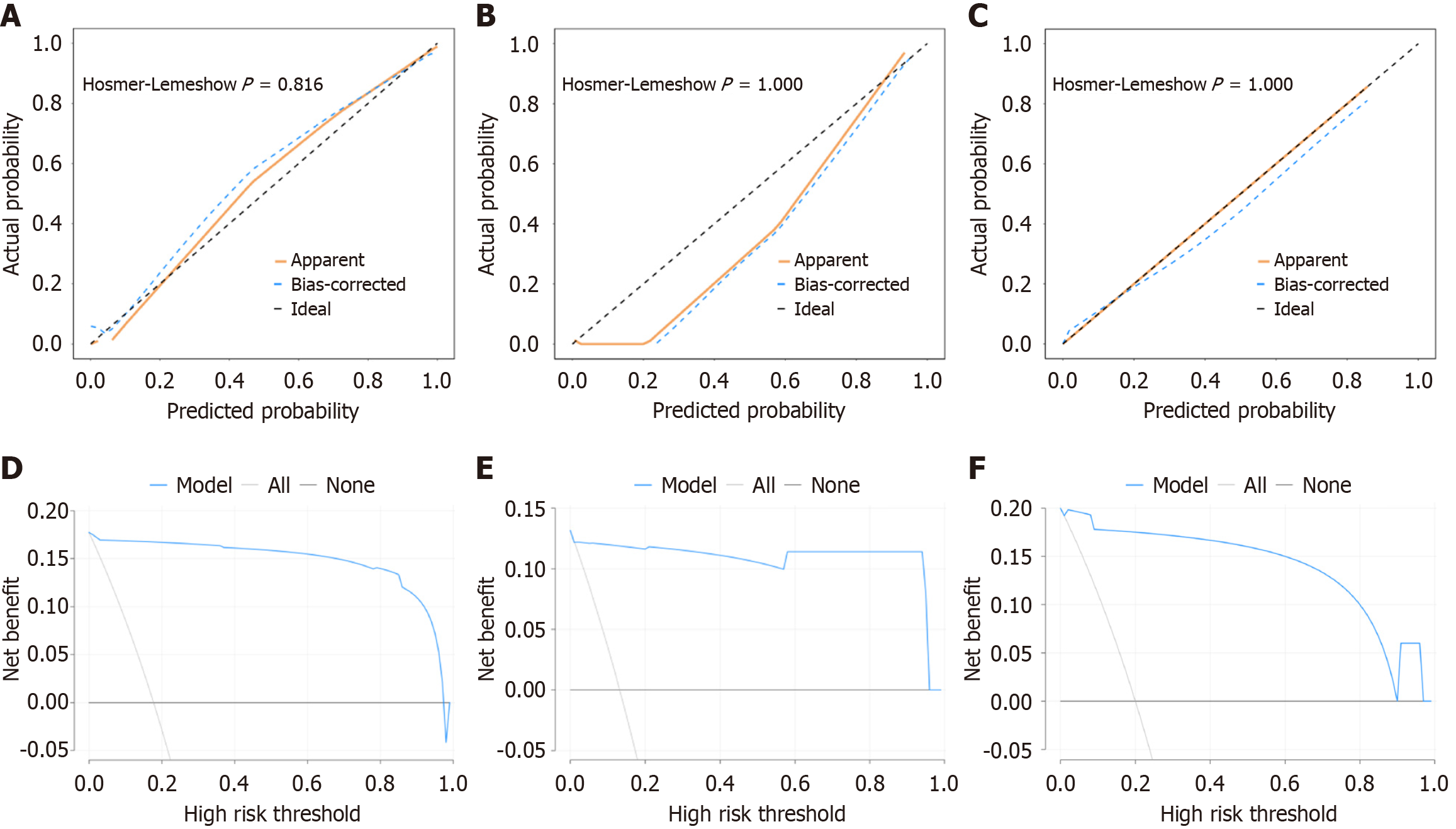
The nomogram was constructed using four independent indicators to differentiate between UGT1A1 gene mutation-associated and hemolytic disease-associated unconjugated hyperbilirubinemia (Figure 3). With a diagnostic threshold of 126 points (probability > 0.285 for hemolytic disease), the model showed excellent discriminative performance. The area under the receiver operating characteristic values were 0.994, 0.958, and 0.986 in the training, internal validation, and external validation cohorts, respectively (Figure 4). Performance metrics also remained consistently high across cohorts, including sensitivity (95.7%, 93.3%, 100%), specificity (98.6%, 98.0%, 90.0%), and accuracy (98.1%, 97.4%, 92.0%) (Supplementary Table 7). In light of potential class imbalance, PR curve analysis further highlighted the model’s robustness in predicting the positive class, showing consistently high area under PR curve (0.978, 0.943, 0.938) and F1-scores (0.947, 0.903, 0.833) across three cohorts (Figure 4 and Supplementary Table 7). Moreover, calibration curves and the Hosmer-Lemeshow test (all P > 0.05) demonstrated good calibration across all three cohorts (Figure 5). Decision curve analysis confirmed the model’s clinical utility, showing a high net benefit in discriminating between the two etiologies within a threshold probability range of 0.05-0.95 (Figure 5).
To facilitate timely management, we developed a differential diagnostic nomogram using four routine indicators to distinguish hemolytic diseases from UGT1A1 mutation associated unconjugated hyperbilirubinemia. The model highlights abnormal peripheral blood smear, elevated RET%, decreased HCT, and elevated RDWSD as key predictors of hemolytic disease. Its simplicity, interpretability, and reliance on accessible tests make it suitable for primary care. Likewise, this model can assist clinicians in early screening and guide patients to appropriate, tailored examinations, thereby avoiding unnecessary tests and reducing the financial burden on patients. The model demonstrated high diagnostic efficacy and clinical benefit in validation cohorts, supporting its clinical applicability.
From the pathogenesis perspective, elevated unconjugated bilirubin can be attributed to overproduction of bilirubin (e.g., hemolytic anemia, large hematoma absorption), impaired bilirubin conjugation (e.g., hereditary UGT1A1 enzyme deficiency like Gilbert’s syndrome and Crigler-Najjar’s syndrome), or impaired hepatic uptake (e.g., conditions that reduce hepatic blood flow, like portosystemic shunts and congestive heart failure, or the administration of drugs or contrast agents that interfere with bilirubin uptake)[13]. We focused on differentiating the two most common causes: UGT1A1 gene mutation and hemolytic disease.
Current diagnosis of Gilbert’s and Crigler-Najjar’s syndrome is based on clinical exclusion, particularly excluding liver and hemolytic diseases[1]. However, excluding hemolytic diseases is complex. Hemolytic diseases can be categorized as congenital (due to intrinsic red cell defects, such as membrane abnormalities like hereditary spherocytosis, enzyme deficiencies like glucose-6-phosphate dehydrogenase deficiency, and hemoglobin disorders like thalassemia) or acquired (due to extrinsic factors like infection, medication, tumor, autoimmune hemolytic anemia, paroxysmal nocturnal hemoglobinuria). Due to the high heterogeneity of clinical manifestations and laboratory findings, diagnosing hemolytic diseases is challenging[14]. Diagnosis often requires combined assessment of history, clinical presentation, red cell morphology, specialized tests, and genetic testing for congenital forms[15]. To our knowledge, only one previous study has focused on differentiating hemolytic disease from Gilbert’s syndrome, utilizing Levitt’s CO breath test to assess red blood cell lifespan. This approach achieved its best diagnostic performance at a cutoff of 60 days, with an area under the receiver operating characteristic curve of 0.982, sensitivity of 94.3%, and specificity of 96.1%[5]. However, the limited avail
Our model highlighted peripheral blood smear, RET%, HCT, and RDWSD as critical indicators for differential diagnosis. RET% reflects the rate of red blood cell turnover. In hemolytic diseases, increased red cell destruction often triggers compensatory erythropoiesis, leading to elevated RET count and percentage. In this study, the calculated cutoff for RET% was 2.1%, slightly above the normal range (0.5%-1.5%). This finding suggests a mildly elevated red blood cell turnover in some patients with UGT1A1 mutation-associated unconjugated hyperbilirubinemia. This was also supported by studies showing a slight reduction in red blood cell lifespan in patients with Gilbert’s syndrome[16]. However, RET% can be normal or even decreased in some hemolytic disorders(e.g., in congenital dyserythropoietic anemia), underscoring the limitations of relying on RET% alone[17].
Peripheral blood smear is essential for diagnosing hemolysis, with unique findings that can be diagnostic of specific hemolytic anemias[18]. For instance, spherocytes suggest acquired autoimmune hemolytic anemia or hereditary spherocytosis, whereas stomatocytes or elliptocytes suggest hereditary stomatocytosis or elliptocytosis, respectively[12]. Red cell fragmentation suggests microangiopathic hemolytic anemia or mechanical hemolysis. In contrast, patients with UGT1A1 mutation-associated hyperbilirubinemia usually have a normal peripheral blood smear result. However, red blood cell destruction during peripheral blood sample collection can yield a false-positive results.
HCT and RDWSD are complete blood count parameters and are of significant value in the differential diagnosis. HCT reflects the percentage of red blood cells in the blood and assesses the severity of anemia. Its decrease in hemolytic disease is due to increased red cell destruction. Conversely, HCT is usually normal or slightly elevated in Gilbert’s syndrome[19]. RDWSD reflects the heterogeneity of red cell volume and is typically elevated in hemolytic diseases because compensatory bone marrow activity produces larger reticulocytes alongside smaller mature red cells, thereby increasing red cell volume variability. In addition, abnormal red blood cell morphology due to membrane structural disorders (e.g., hereditary spherocytosis) also increases red blood cell volume heterogeneity.
The selected variables in this model primarily relate to red cell morphology, quantity, content, and volume heterogeneity. In contrast, bilirubin levels, biochemical indicators, and abdominal ultrasound findings are not included in the final nomogram model. This suggests that clinicians should particularly focus on red blood cell-associated indicators when differentiating the etiology of unconjugated hyperbilirubinemia. In our study, the diagnostic threshold of the nomogram is 126 points, with abnormal peripheral blood smear contributing 82.5 points, HCT < 40.15% contributing 60 points, RDWSD ≥ 45.55 fL contributing 67 points, and RET% ≥ 2.1% contributing 100 points. Therefore, we propose that satisfying any two or more of the above indicators strongly suggests hemolytic disease. For patients flagged by the model as likely having hemolytic disease, subsequent investigations to determine the etiology of hemolysis are warranted, including whole-exome sequencing if necessary. Conversely, for patients predicted to have UGT1A1 mutation-associated unconjugated hyperbilirubinemia, complex testing can often be omitted, and UGT1A1 genetic testing may be discussed with the patient. The diagnosis and management of unconjugated hyperbilirubinemia usually require excellent coordination and interdisciplinary cooperation among primary care physicians, gastroenterologists/hepatologists, hematologists, pediatricians, and other specialists. This model effectively assists with early patient triage and guides referrals to the appropriate specialist.
This is the first nomogram model developed for differential diagnosing UGT1A1 mutation and hemolytic disease-associated unconjugated hyperbilirubinemia. However, this study has some limitations. First, the hemolysis group was relatively small, resulting in class imbalance during model construction. This limitation is due to the data constraints of our specialized liver disease center, where hemolytic cases are rare. Despite this, the model maintained strong diagnostic performance in validation cohorts, with area under PR curve values remaining above 0.93 and F1-score exceeding 0.83, demonstrating robust predictive capability for the positive class. Second, the hemolytic patients in this study predominantly had hereditary spherocytosis, with a lower proportion due to other etiologies and no rare hemolytic types. Although this distribution reflects real-world epidemiological patterns - hereditary spherocytosis is the most common hemolytic anemia, with a prevalence of about 1 in 2000[20] - it may still limit the model’s generalizability to broader hemolytic populations. Third, the subjects were primarily adults, pediatric applicability requires validation. Finally, the external validation performed was temporal - the validation cohort came from the same center as the training cohort but from a distinct and later time period. This design tests the model’s robustness over time but does not establish its generalizability across different institutions or populations. Taken together, further validation in a large-scale, multi-center cohort with balanced sample groups and greater etiological diversity of hemolysis is warranted to confirm the model’s broader clinical applicability.
We developed an effective nomogram model for the differential diagnosis of UGT1A1 gene mutation-associated and hemolytic disease-associated unconjugated hyperbilirubinemia, ultimately improving the clinical utility of preliminary screening.
| 1. | Vítek L, Tiribelli C. Gilbert's syndrome revisited. J Hepatol. 2023;79:1049-1055. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 47] [Article Influence: 15.7] [Reference Citation Analysis (0)] |
| 2. | Sambati V, Laudisio S, Motta M, Esposito S. Therapeutic Options for Crigler-Najjar Syndrome: A Scoping Review. Int J Mol Sci. 2024;25:11006. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 9] [Reference Citation Analysis (0)] |
| 3. | Michel M, Crickx E, Fattizzo B, Barcellini W. Autoimmune haemolytic anaemias. Nat Rev Dis Primers. 2024;10:82. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 27] [Reference Citation Analysis (0)] |
| 4. | Turpaev K, Bovt E, Shakhidzhanov S, Sinauridze E, Smetanina N, Koleva L, Kushnir N, Suvorova A, Ataullakhanov F. An overview of hereditary spherocytosis and the curative effects of splenectomy. Front Physiol. 2025;16:1497588. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 7] [Reference Citation Analysis (0)] |
| 5. | Kang LL, Liu ZL, Han QS, Chen YW, Liu LW, Xie XH, Luo JF, Ji YQ, Zhu GL, Ma YJ, Ji KM, Zhang HD. Levitt's CO breath test in the differential diagnosis of chronic isolated hyperbilirubinemia. J Breath Res. 2022;16. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 6. | di Masi A, De Simone G, Ciaccio C, D'Orso S, Coletta M, Ascenzi P. Haptoglobin: From hemoglobin scavenging to human health. Mol Aspects Med. 2020;73:100851. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 124] [Cited by in RCA: 107] [Article Influence: 17.8] [Reference Citation Analysis (0)] |
| 7. | Inherited Metabolic Liver Disease Collaboration Group; Chinese Society of Hepatology; Chinese Medical Association. [Expert consensus on the diagnosis and therapy of inherited hyperbilirubinemia (version 2025)]. Zhonghua Gan Zang Bing Za Zhi. 2025;33:747-759. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 8. | Red Blood Cell Disease (Anemia) Group; Chinese Society of Hematology; Chinese Medical Association. [Chinese guideline for the diagnosis and treatment of adult autoimmune hemolytic anemia (2023)]. Zhonghua Xue Ye Xue Za Zhi. 2023;44:12-18. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 4] [Reference Citation Analysis (0)] |
| 9. | Polizzi A, Dicembre LP, Failla C, Matola TD, Moretti M, Ranieri SC, Papa F, Cenci AM, Buttarello M. Overview on Hereditary Spherocytosis Diagnosis. Int J Lab Hematol. 2025;47:18-25. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 7] [Reference Citation Analysis (0)] |
| 10. | Andolfo I, Russo R, Gambale A, Iolascon A. Hereditary stomatocytosis: An underdiagnosed condition. Am J Hematol. 2018;93:107-121. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 60] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 11. | Writing Group for Practice Guidelines for Diagnosis and Treatment of Genetic Diseases Medical Genetics Branch of Chinese Medical Association; Shang X, Zhang X, Yang F, Xu X. [Clinical practice guidelines for alpha-thalassemia]. Zhonghua Yi Xue Yi Chuan Xue Za Zhi. 2020;37:235-242. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 6] [Reference Citation Analysis (0)] |
| 12. | Ruivard M, Michel M, Garçon L. [Diagnosis of non-autoimmune hemolysis in the adult]. Rev Med Interne. 2025;46:725-732. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 13. | Nelson M, Mulani SR, Saguil A. Evaluation of Jaundice in Adults. Am Fam Physician. 2025;111:25-30. [PubMed] |
| 14. | Fattizzo B, Giannotta JA, Cecchi N, Barcellini W. Confounding factors in the diagnosis and clinical course of rare congenital hemolytic anemias. Orphanet J Rare Dis. 2021;16:415. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 8] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 15. | Jamwal M, Sharma P, Das R. Laboratory Approach to Hemolytic Anemia. Indian J Pediatr. 2020;87:66-74. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 33] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 16. | Kang LL, Ma YJ, Zhang HD. Carbon monoxide breath test assessment of mild hemolysis in Gilbert's syndrome. Medicine (Baltimore). 2020;99:e19109. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 6] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 17. | Iolascon A, Andolfo I, Russo R. Congenital dyserythropoietic anemias. Blood. 2020;136:1274-1283. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 78] [Article Influence: 15.6] [Reference Citation Analysis (0)] |
| 18. | Baldwin C, Pandey J, Olarewaju O. Hemolytic Anemia. 2023 Jul 24. In: StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing; 2025. [PubMed] |
| 19. | Buyukasik Y, Akman U, Buyukasik NS, Goker H, Kilicarslan A, Shorbagi AI, Hascelik G, Haznedaroglu IC. Evidence for higher red blood cell mass in persons with unconjugated hyperbilirubinemia and Gilbert's syndrome. Am J Med Sci. 2008;335:115-119. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 16] [Article Influence: 0.9] [Reference Citation Analysis (4)] |
| 20. | Perrotta S, Gallagher PG, Mohandas N. Hereditary spherocytosis. Lancet. 2008;372:1411-1426. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 401] [Cited by in RCA: 430] [Article Influence: 23.9] [Reference Citation Analysis (1)] |