Published online Mar 26, 2026. doi: 10.4252/wjsc.v18.i3.116388
Revised: December 11, 2025
Accepted: February 9, 2026
Published online: March 26, 2026
Processing time: 135 Days and 3.8 Hours
Stromal vascular fraction (SVF) derived from adipose tissue (AT) is a heterogeneous cell mixture. SVF is commonly prepared via enzymatic digestion of AT. Typically, only the initial SVF (SVF1) is harvested, and the remaining AT is discarded. However, residual AT may contain viable regenerative cells.
To explore a novel approach involving a second enzymatic digestion to maximize stem cell yield, particularly relevant for pediatric patients or those with limited adipose reserves.
AT samples obtained via liposuction were digested twice using collagenase type IV to yield SVF1 and SVF2 (after 1st and 2nd digestions, respectively). Both fractions were assessed for cell viability, mononuclear cell count, plating efficiency, and the expression of surface markers via flow cytometry. Gene expression analysis of vascular endothelial growth factor, stromal cell-derived factor, nitric oxide syn
Results demonstrated that SVF2 contained a significant number of mononuclear cells and exhibited a high plating efficiency. Flow cytometry and gene expression analyses further confirmed that SVF2 retained regenerative cells. Furthermore, histological evaluation using hematoxylin and eosin staining revealed the presence of intact cells within the residual tissue post-first digestion.
In conclusion, secondary digestion of AT (typically discarded after 1st digestion), can yield additional regenerative cells. This method enhances stem cell number for stem cell-based therapies and is particularly valuable for pe
Core Tip: This study presents a novel dual-digestion method to isolate stem cells from small adipose tissue samples, optimizing cell yield for regenerative medicine applications. Traditionally, only the first stromal vascular fraction is retained for research or therapeutic purpose, while remaining undigested tissue is discarded. However, a second enzymatic digestion (yielding stromal vascular fraction 2) revealed a significant number of viable regenerative cells. They presence of viable regenerative cells was confirmed by plating efficiency, flow cytometry, gene expression analysis and histological studies. This approach is especially beneficial for pediatric patients or individuals with limited adipose reserves, enhancing the number of autologous stem cell by maximizing the utility of small-volume adipose tissue samples.
- Citation: Mushtaq A, Choudhery MS, Arif T, Niaz A, Pervaiz A. Maximizing pediatric stem cell harvest: A novel two-step digestion approach for adipose tissue. World J Stem Cells 2026; 18(3): 116388
- URL: https://www.wjgnet.com/1948-0210/full/v18/i3/116388.htm
- DOI: https://dx.doi.org/10.4252/wjsc.v18.i3.116388
Adipose tissue (AT), once considered primarily as a passive energy reservoir, is now widely recognized as a dynamic and accessible source of regenerative cells, particularly of adipose-derived stem cells (ASCs). Among its cellular components, the stromal vascular fraction (SVF) holds significant therapeutic potential due to its heterogeneous nature with various types of cells such as vascular endothelial progenitors, ASCs, blood cells, preadipocytes, fibroblasts, immune cells and smooth muscle cells[1]. Recently, the medical community has shown a growing interest in using SVF as a treatment option to circumvent various regulatory challenges. Additionally, SVF can be obtained in a short period of time with the requirement of in vitro culturing, and its preparation is relatively inexpensive. Clinically, SVF has been applied in number of conditions such as non-healing wounds, diabetes, neurodegenerative and malignant diseases, cosmetics, aesthetic surgeries and hair restoration. These therapeutic applications are primarily attributed to the regenerative, angiogenic, and anti-inflammatory properties of the cellular components within SVF[2,3]. In chronic wounds, SVF promotes tissue regeneration and accelerates healing by enhancing vascularization and modulating the local immune response[4]. In aesthetic and cosmetic procedures, SVF contributes to skin rejuvenation and tissue volume restoration, while in hair restoration, it supports follicular regeneration and improves hair density[5-7].
SVF is commonly isolated using enzymatic digestion, typically with collagenase, to release regenerative cells embedded within the extracellular matrix (ECM). It is pertinent to note that for successful stem cell-based therapy, the sufficient number of regenerative cells, particularly ASCs, is very important. In clinical scenarios involving pediatric patients and those with low-AT reserves, maximizing the yield of stem cells from limited tissue volumes becomes critically important[8]. SVF contains a low number of regenerative cells i.e., ASCs. To get sufficient number of rege
During processing, AT is usually discarded after the initial (1st) enzymatic digestion. However, we propose that the residual AT may still contain viable regenerative cells that remain after first digestion (SVF1). It is pertinent to note that different laboratories employ varying enzyme types and concentrations, incubation times, and temperatures. The second digestion (yielding SVF2) of AT which is usually discarded following SVF1 extraction, is based on the hypothesis that a significant population of regenerative cells remains entrapped within the partially digested AT ECM. Given this, a second round of digestion could serve as a practical method to extract additional stem cells, which could be particularly bene
This study developed a novel two-step enzymatic digestion approach aimed at obtaining maximum number of regenerative cells from residual tissue (discarded fat) of small AT samples. To evaluate the efficacy of this approach, we have assessed the SVF2 for cellular viability, total number of mononuclear cells, plating efficiency (PE), and cellular & molecular characterization. In addition, histological analysis with hematoxylin and eosin (H&E) staining was used to assess the structural integrity of the tissue pre- and post-digestion. Results indicated that AT retained after first digestion contained a significant number of viable regenerative cells as indicated by the number of mononucleated cells, colony forming units (CFU), PE, flow cytometry, quantitative reverse transcription polymerase chain reaction (qRT-PCR) and histological analysis. To the best of our knowledge, this is the first such study for the implementation of optimized protocol for isolation of maximum number of stem cells from a small AT samples for use in cell-based therapies. This study highlights a simple yet powerful protocol modification that substantially improves the yield of regenerative cells, including ASCs, from limited AT sources. The findings underscore the potential of this method to improve cell-based therapies, especially in pediatric patients or individuals with limited tissue availability.
This study was carried out in accordance with the recommendations of the Institutional Review Board at the University of Health Sciences, Lahore, Pakistan. The study and protocols were approved by the University of Health Sciences, Lahore (No. UHS/Education/126-24/1062). Informed written consent was obtained from all the participants before collecting AT samples in a local hospital.
In this study, AT was collected from five (n = 5) healthy donors (both male and female genders). All participants were clinically stable and free of metabolic or inflammatory conditions and the aspirated adipose volume per donor ranged from 2 mL to 5 mL. The AT was obtained through syringe liposuction under local anesthesia. The donor site was first sterilized with alcohol swabs. The tumescent fluid [standard tumescent fluid recipe for 1 L (1000 mL): Normal saline or lactated Ringer’s - 1000 mL + lidocaine (0.05% to 0.1%) + epinephrine (1:1000000) - 1 mg (e.g., 1 mL of 1:1000 epinephrine) + sodium bicarbonate 10 mL of 8.4%] was injected using a syringe to facilitate fat removal and reduce pain and dis
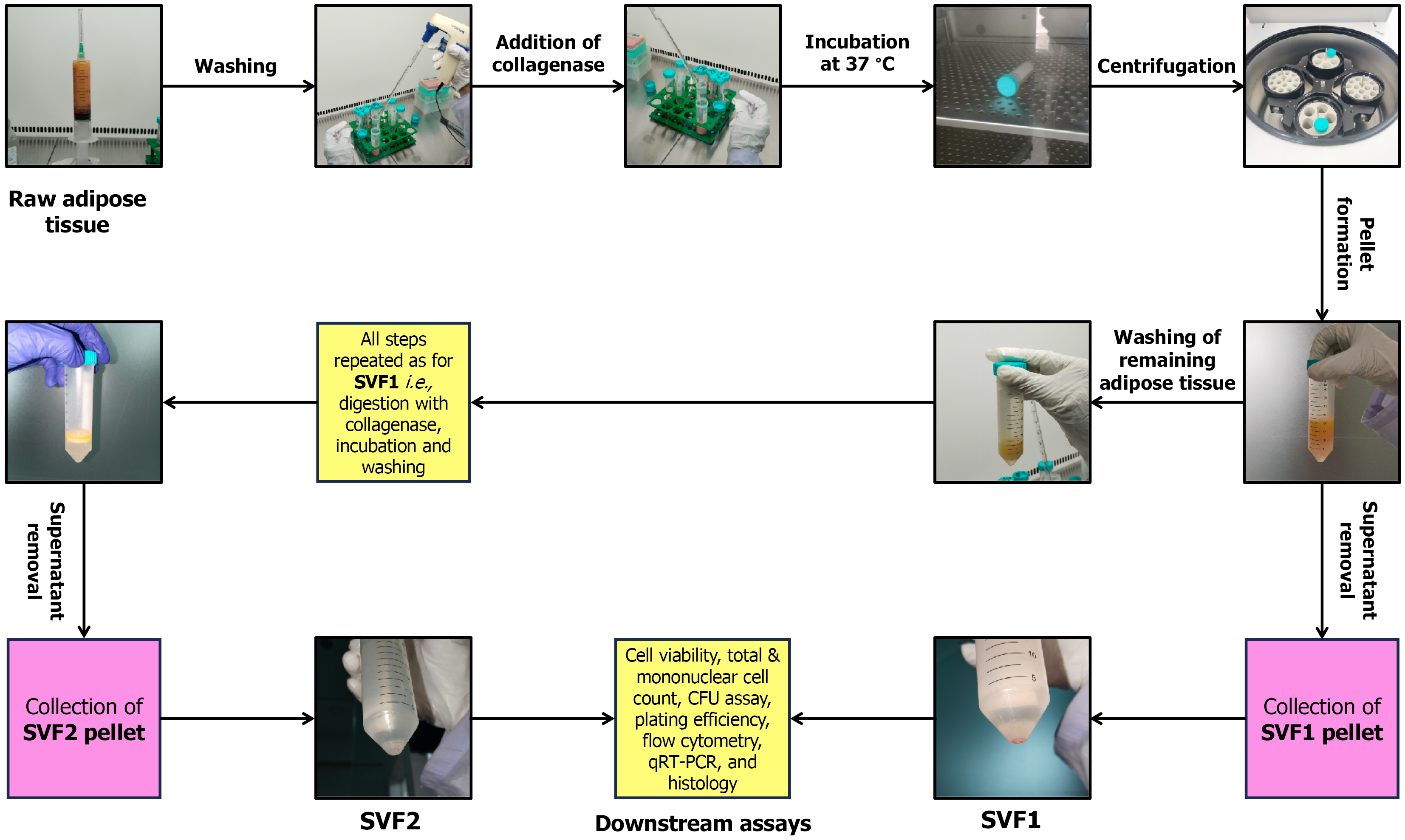
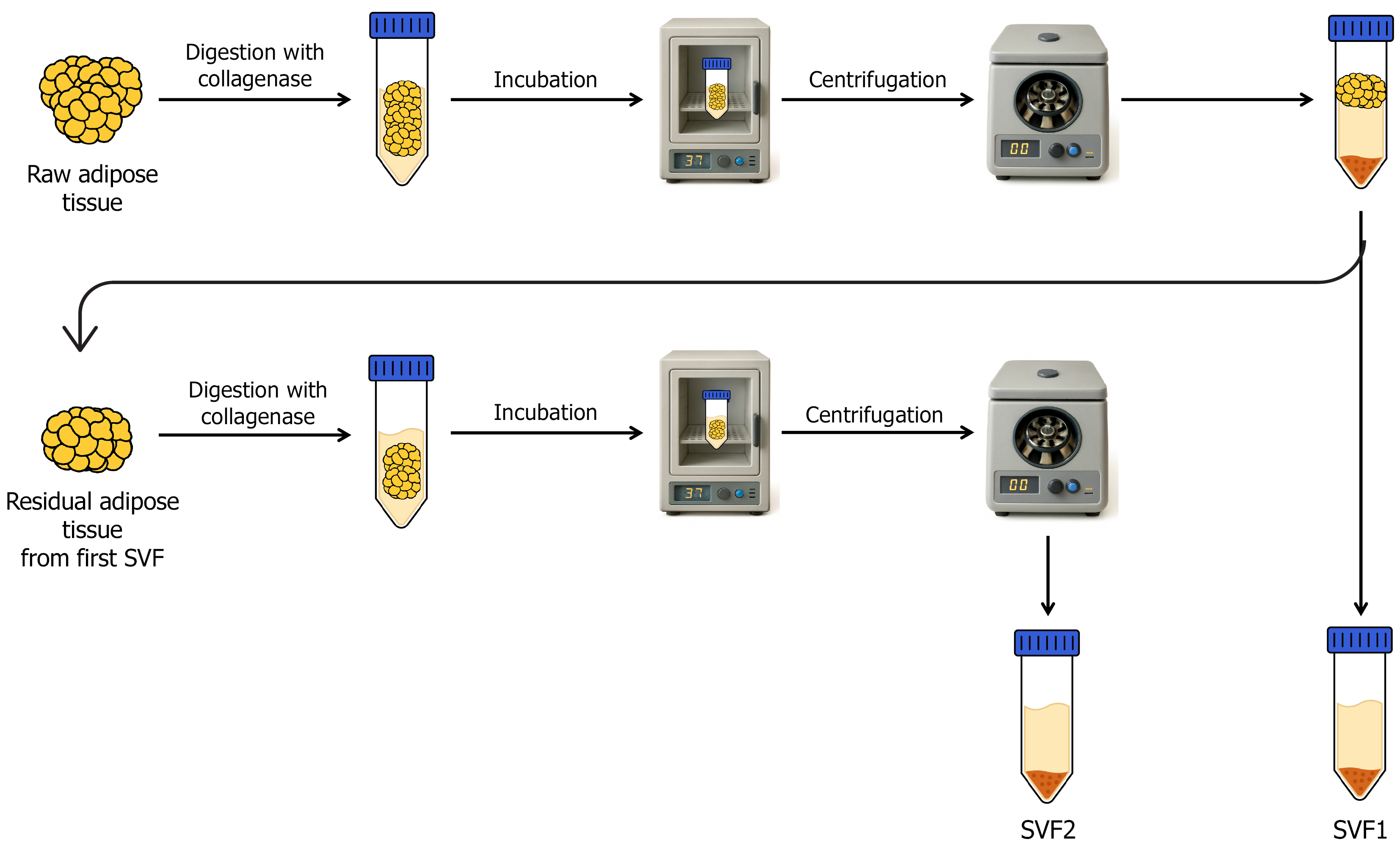
The raw lipoaspirate of AT was processed within 24 hours under sterilized conditions to obtain SVF. Briefly, raw lipoaspirate (approximately 10 cc) was transferred into a sterile 50 mL centrifuge tube. An equal volume of phosphate buffered saline (PBS) was added to wash the AT. After shaking the tubes, they were allowed to settle for few seconds. The liquid portion was discarded while the AT was retained. This washing step was repeated 3-5 times or until blood was removed. An equal volume of collagenase type IV (2.5 mg/mL) was added and mixed properly. Collagenase type IV was used for its milder proteolytic activity, facilitating the efficient release of stromal and perivascular cells while reducing catalytic degradation. The mixture was incubated at 37 °C for 30 minutes with periodic agitation. After the incubation, cell suspension was passed through a 70 μm cell strainer to remove undigested tissue fragments. The cell suspension was centrifuged at 250 × g for 5 minutes. Following centrifugation, supernatant was discarded and resulting cell pellet was washed again with PBS. The final cell pellet was designated as SVF1. The same procedure was repeated with residual AT remained after first enzymatic digestion to obtain SVF2. The two-step digestion workflow used to isolate SVF1 and SVF2 is illustrated in Figure 1.
The total number of cells in SVF1 and SVF2 was determined using a hemocytometer and the trypan blue exclusion assay. Both the stained (dead) and unstained (viable) cells were counted manually to determine the cell viability. Cell concentration (cells/mL) was calculated by multiplying the average cell count by the dilution factor and by hemocytometer factor (104). In addition, percentage cell viability was calculated using the following formula: Percent cell viability = [(number of unstained cells)/(total cell count)] × 100.
Mononucleated cells were counted using a hemocytometer as described above, with the aid of Turk’s solution. Cells with well-defined nuclei were identified and counted as mononucleated cells. Non-nucleated cells either did not stain or exhibited irregular staining and were excluded from the count. The total number of mononucleated cells was recorded and their concentration/number (cells/mL) was calculated by multiplying the average cell count by the dilution factor and the hemocytometer volume factor (i.e., 104).
CFU assay was performed for each SVF fraction (i.e., SVF1 and SVF2) to determine the frequency of cells capable of forming colonies. The assay was performed according to the methodology outlined by Choudhery et al[9]. Briefly, the cells from both SVF1 and SVF2 were counted, and seeded at low density in culture dishes. The culture dishes were incubated under standard culture conditions (5% CO2, 37 °C under humidity) in a complete culture medium containing DMEM supplemented with 5% foetal bovine serum, 1% each of penicillin/streptomycin and L-glutamine (Gibco™, NY, United States). The 5% foetal bovine serum was used to limit serum-induced proliferative effects and preserve the intrinsic clonogenic behavior of SVF-derived cells, enabling more accurate comparison between SVF1 and SVF2.
The cells were allowed to grow and form colonies over a period of 14 days. During this incubation period, the cells proliferated and formed visible colonies. After the 14-day culture period, the colonies were carefully washed with PBS to remove any residual medium or non-adherent cells. The colonies were then fixed with 10% formalin to preserve their morphology. Following fixation, crystal violet dye was used to stain the colonies, facilitating their visualization. Colonies were observed and counted under a microscope. The number of colonies formed in each fraction of AT (i.e., SVF1 and SVF2) was recorded. The data collected from each fraction were used to determine the colony-forming efficiency of cells in SVF1 and SVF2. The PE of both SVF1 and SVF2 was determined using the following formula: PE = [(number of colonies formed)/(number of cells seeded)] × 100.
Cell surface antigen profiling of cells from 1st and 2nd enzymatic digestion was evaluated by flow cytometry. After the cells were harvested and pelleted, they were washed with PBS (3 times). Cell suspension was centrifuged to make the pellet again. The cells were then fixed with 10% formalin. The cells were subsequently washed and re-suspended in PBS. A total of 100000 cells were incubated with 10 μL of primary antibodies targeting CD44 (PE conjugated, Invitrogen, CA, United States), CD73 (FITC conjugated, Invitrogen, CA, United States) and CD146 (PE conjugated, Invitrogen, CA, United States). The antibodies were mixed with the cell suspension and cells were incubated for 60 minutes in dark at 4 °C to allow for optimal binding of the antibodies to the cell surface markers. Samples were analyzed using FACS Chorus software (BD Biosciences, CA, United States). During flow cytometry, debris and auto-fluorescence signals were removed by gating.
RNA extraction and cDNA synthesis: Total RNA was extracted from AT samples [before digestion (AT), after 1st digestion (AT1) and after 2nd digestion (AT2)] as well as from SVF pellets (SVF1 and SVF2) using TRIzol™ reagent (Life Technologies, Carlsbad, CA, United States). Briefly, TRIzol™ reagent was added to AT samples (AT, AT1 and AT2), as well as SVF pellets (SVF1 and SVF2). All samples were thoroughly homogenized by repeated pipetting. The homogenized samples were incubated at room temperature for 5 minutes to completely dissociate nucleoprotein complexes. Chloroform was added at a ratio of 0.2 mL per 1 mL of TRIzol™ reagent. After vigorous shaking, the tubes were incubated at room temperature for 3 minutes. Samples were then centrifuged at 12000 × g for 15 minutes at 4 °C. The upper aqueous phase containing RNA was carefully transferred into new sterilized tubes. RNA was precipitated by adding 500 μL of isopropanol per 1 mL of Trizol™ reagent used. Samples were incubated for 10 minutes at room temperature and centrifuged again at 12000 × g for 10 minutes at 4 °C. The supernatant was removed and RNA pellet was washed once with 75% ethanol, followed by centrifugation at 7500 × g for 5 minutes at 4 °C. RNA pellet was desiccated for 15 minutes. RNA was dissolved in RNase-free water and incubated for 10 minutes at 60 °C. RNA concentration and purity was determined using a Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, MA, United States).
cDNA was synthesized using commercially available cDNA synthesis Kit (Thermo Scientific, MA, United States) according to the manufacturer’s instructions. A total of 500 ng of RNA was reverse-transcribed. The kit reagents were gently combined and subsequently placed in a T100 thermocycler (Bio-Rad, CA, United States) after brief centrifugation. The revere transcription reaction was carried out at 42 °C for 1 hour, followed by an inactivation step at 70 °C for 5 minutes. The cDNA was stored at -20 °C until use for gene expression analysis.
qRT-PCR: The expression of vascular endothelial growth factor (VEGF), stromal cell-derived factor (SDF), nitric oxide synthase-3 (NOS3), CD14, matrix metalloproteinase 3 (MMP3), interleukin-4 receptor (IL4R), P16, and P53 genes was measured at the mRNA level using RT-PCR in AT, AT1 and AT2, as well as in SVF1 and SVF2[10]. The final reaction mixture contained template cDNA, Taq DNA polymerase, SYBR Green and gene-specific primers. Samples were subjected to qRT-PCR with the following conditions: Initial denaturation at 95 °C for 5 minutes and 40 cycles of denaturation at 95 °C for 15 seconds, annealing & extension at 60 °C for 1 minute. A standard melt curve analysis was included at the end of the run to verify amplification specificity. Relative gene expression levels were calculated using the comparative ∆Ct method, with target gene expression normalized to a GAPDH housekeeping gene. Table 1 shows the primer sequences of target genes.
| Genes | Primer sequence 5’ → 3’ |
| VEGF | F: TTGCCTTGCTGCTCTACCTCCA |
| R: GATGGCAGTAGCTGCGCTGATA | |
| SDF | F: CTCAACACTCCAAACTGTGCCC |
| R: CTCCAGGTACTCCTGAATCCAC | |
| NOS3 | F: GAAGGCGACAATCCTGTATGGC |
| R: TGTTCGAGGGACACCACGTCAT | |
| CD14 | F: CTGGAACAGGTGCCTAAAGGAC |
| R: GTCCAGTGTCAGGTTATCCACC | |
| MMP3 | F: CACTCACAGACCTGACTCGGTT |
| R: AAGCAGGATCACAGTTGGCTGG | |
| IL4R | F: CTGCTCATGGATGACGTGGTCA |
| R: GGTGTGAACTGTCAGGTTTCCTG | |
| CDKN2A (P16) | F: CTCGTGCTGATGCTACTGAGGA |
| R: GGTCGGCGCAGTTGGGCTCC | |
| P53 | F: CCTCAGCATCTTATCCGAGTGG |
| R: TGGATGGTGGTACAGTCAGAGC | |
| GAPDH | F: ACGGATTTGGTCGTATTGGG |
| R: CGCTCCTGGAAGATGGTGAT |
To assess the structural integrity and cellular status of AT at different stages of enzymatic digestion, histological analysis was performed. Samples included intact AT before digestion (AT), AT after the first digestion (AT1), and AT after the second digestion (AT2). Each of these tissue samples was fixed in 10% neutral-buffered formalin for 48 hours to preserve cellular morphology. Following fixation, the AT samples were dehydrated with ethanol to remove water, followed by clearing in xylene to eliminate residual alcohol and prepare the tissue for infiltration. The cleared tissues were then infiltrated with molten paraffin wax at 60 °C under vacuum pressure for 1-2 hours to ensure complete penetration of wax into the tissue matrix. Once fully infiltrated, the tissues were embedded in paraffin blocks by orienting them in metal molds filled with molten wax and allowing them to solidify at room temperature. From each paraffin-embedded sample, 4-5 μm thick sections were cut using a rotary microtome (Leica, RM212RT). The tissue sections were placed in warm water to remove the wrinkles. Tissue sections were picked up on glass microscopic slides that were placed in a hot air oven for 15 minutes to help the sections adhere to the slides. These thin sections were then mounted on glass slides and stained using H&E staining. The stained slides were examined under an inverted microscope (LEICA MC170 HD) to evaluate the morphological integrity and cellular components at different digestion stages.
The data was analyzed using GraphPad Prism (version 10; GraphPad Software, Boston, MA, United States), mean ± SD of each variable was calculated. Student’s t test was applied to evaluate the distribution of all experimental group variables in cell-based assays (two groups). Mean and P-values were calculated using the standard one-way ANOVA test for gene expression analysis (more than two groups). A difference was considered statistically significant if the related P-value was ≤ 0.05.
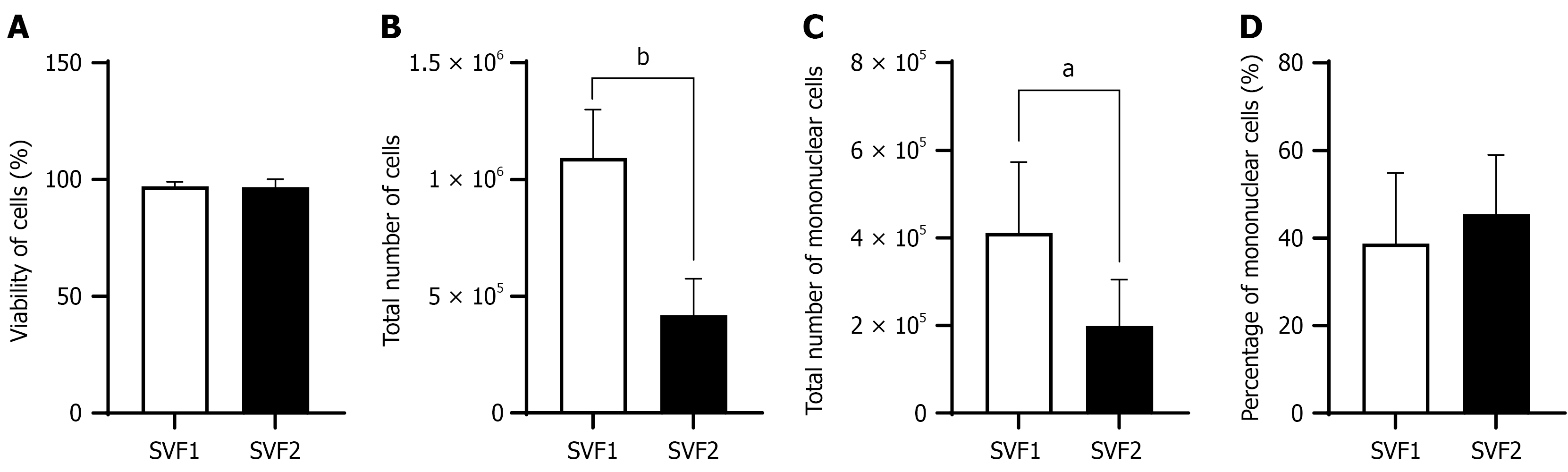
Cell viability in SVF1 and SVF2 was assessed using the trypan blue exclusion assay, a standard technique to differentiate alive (unstained) from dead (blue-stained) cells. Cell viability in both fractions was high with 97.22 ± 1.88 viable cells in SVF1 as compared to 96.84 ± 3.39 viable cells in SVF2 (Figure 2A). The small difference in the number of viable cells between SVF1 and SVF2 was not statistically significant (P = 0.83). Additionally, the total number of cells in SVF1 (1.09 × 106 ± 2.07 × 105 cells) were higher than the total number of cells obtained from SVF2 (4.19 × 105 ± 1.56 × 105 cells). This difference in total cell yields between SVF1 and SVF2 was significant (P = 0.0004) as indicated in Figure 2B. SVF2 contributed an additional 38%-40% to the total cell yield per sample. Although the absolute number of cells in SVF2 was lower than SVF1, the second digestion step recovered a meaningful additional cell population that would otherwise be discarded. The fraction of cells obtained after 1st digestion i.e., SVF1 yielded more mononucleated cells (4.11 × 105 ± 1.62 × 105 cells) as compared to 1.99 × 105 ± 1.05 × 105 cells in SVF2, the fraction of cells obtained after 2nd enzymatic digestion. The total number of mononucleated cells present in SVF1 was significantly higher as compared to the total mononucleated cells yield in SVF2 (P = 0.03). Figure 2C highlight these differences. Interestingly, however, the percentage of mononuclear cells in SVF2 (45.46% ± 13.54%) was higher than the percentage mononucleated cells in SVF1 (38.71% ± 16.11%), although the difference was non-significant with a P value 0.49 as shown in Figure 2D.
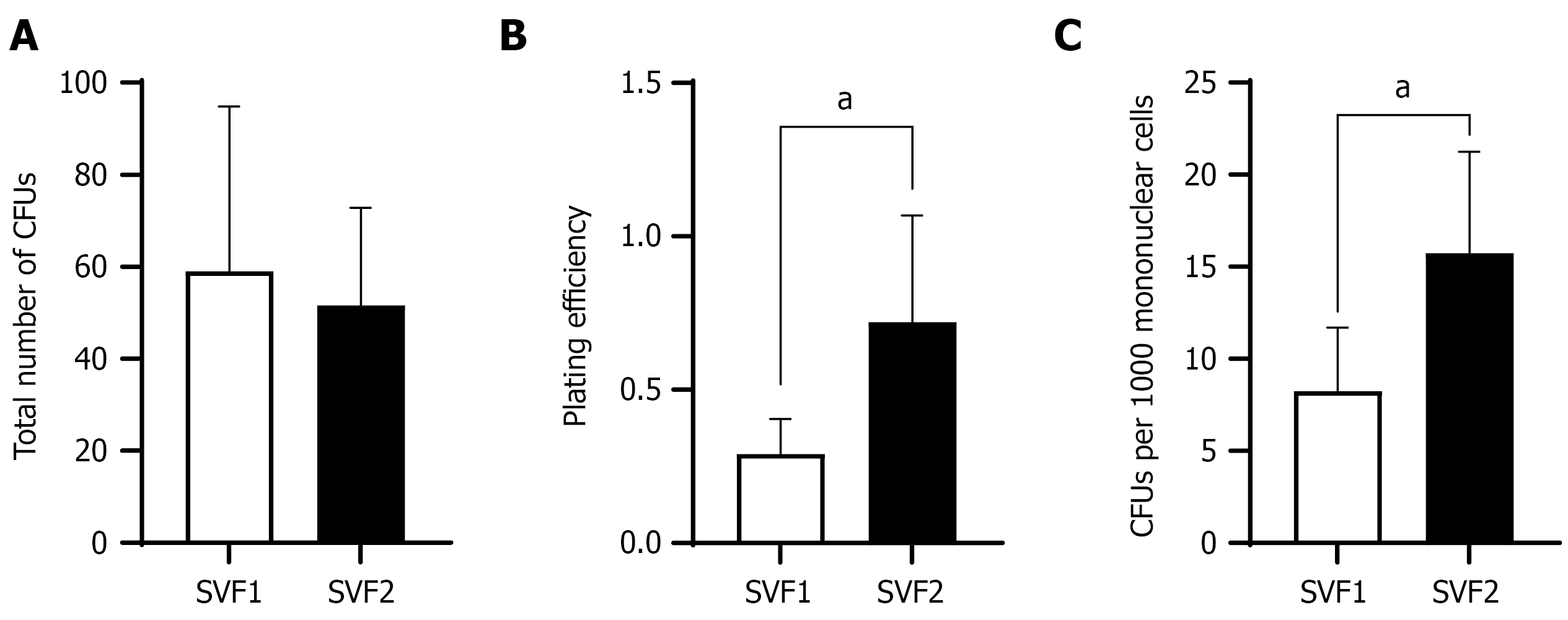
The CFU assay was conducted to assess the clonogenic potential of the cells, specifically, their ability to proliferate and form colonies from single progenitor cells. The total number of CFUs formed by cells in SVF1 was slightly higher than the total number of CFUs formed by cells contained in SVF2. There was no statistically significant difference (P = 0.70) between number of CFUs between SVF1 and SVF2 (Figure 3A). Interestingly, the results of our study indicated that SVF2 had a higher PE (0.72 ± 0.34) than SVF1 (0.29 ± 0.11) with a significant difference (P = 0.03) as shown in Figure 3B. Figure 3C indicated that the number of CFUs per thousand mononucleated cells was also significantly higher in SVF2 (15.75 ± 5.50) than that of SVF1 (8.23 ± 3.47) with a P value of 0.03.
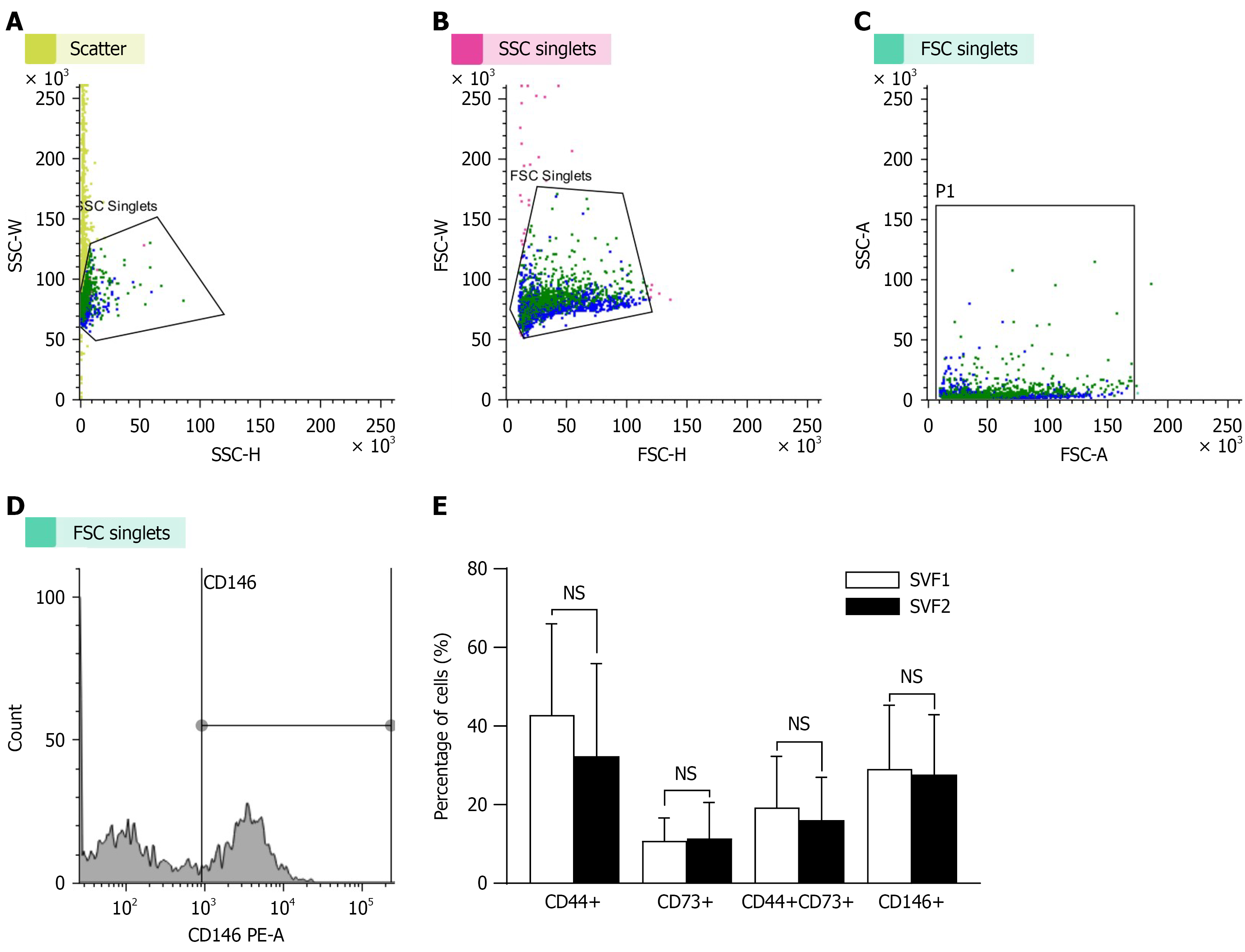
Flow cytometric analysis was used to identify specific surface markers that are characteristic of mesenchymal stem/stromal cells (MSCs). The markers assessed by flow cytometry included CD44, CD73, and CD146. Results showed that even after first enzymatic digestion, the remaining AT, that yielded SVF2, contained numerous regenerative cells (Figure 4) as indicated by the positive expression of CD44, CD73 and CD146. Interestingly, the number of CD44, CD73 and CD146 positive cells in SVF2 was comparable with the cells exhibiting the expression of these markers in SVF1. The percentage of CD44 positive cells was 42.85 ± 23.21 in SVF1 and 32.38 ± 23.55 in SVF2. The SVF1 exhibited 10.86% ± 5.79% CD73 positive cells as compared to 11.41% ± 9.13% CD73 positive cells in SVF2. The cells positive for both CD44 and CD73+ were 19.30 ± 12.97 in SVF1 and 16.11 ± 10.84 in SVF2. Similarly, CD146+ cells in SVF1 were 29.09% ± 16.23% as compared to 27.68% ± 15.21% in SVF2.
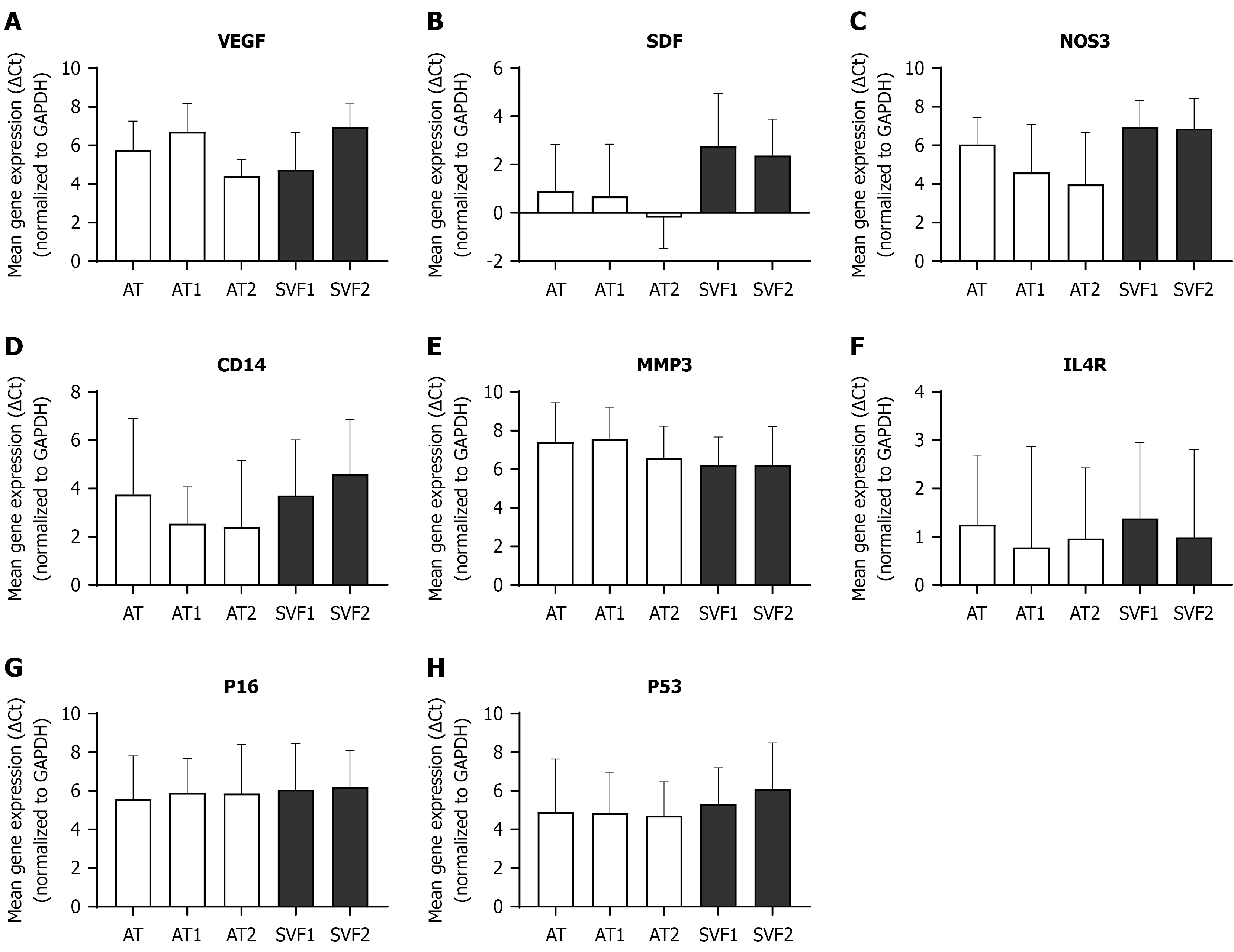
Gene expression profiling was performed using qRT-PCR to evaluate the functional characteristics of cells in both SVF fractions. The expression levels of various genes were measured in both SVF1 and SVF2, as well as in the tissue fractions (AT, AT1, AT2) using qRT-PCR. The target genes expression levels were normalized to a housekeeping gene (GAPDH) to obtain the normalized gene expression (∆Ct). Our results indicated that all target genes were expressed in AT fractions (AT, AT1, AT2) and SVF pellets (SVF1, SVF2). However, statistical analysis showed that the differences in gene expression were not significant (P > 0.05). These results indicate that the expression of these genes is retained after the initial enzymatic digestion of AT, both in the residual AT and in the SVF2 fraction. Gene expression patterns across AT, AT1, AT2, SVF1, SVF2 are shown in Figure 5.
VEGF exhibited slightly higher mean expression in AT1 than in AT (P = 0.83). Although the mean gene expression of SDF, NOS3, CD14, and IL4R was slightly lower in AT1 than in AT (P = 0.99, P = 0.95, P = 0.93, and P = 0.99, respectively), the differences were not statistically significant. The remaining three genes MMP3, P16 and P53 had expressed almost equal mean expression in AT1 and AT (P = 0.99, P > 0.99, P = 0.99 respectively). Conversely, in the case of raw AT and AT following second digestion, there were non-significant variations in the mean expression of VEGF, NOS3, CD14, MMP3, and IL4R between AT2 and AT (P = 0.58, P = 0.70, P = 0.90, P = 0.94, and P = 0.99, respectively). The SDF showed negative mean expression in AT2 when compared to AT (P = 0.87). While P16 and P53 showed almost equal mean expression in both AT2 and AT (P = 0.99, P > 0.99).
In case of SVF pellets, VEGF, CD14 and P53 had shown more mean expression in SVF2 as compared to SVF1 (P = 0.14, P = 0.97, P = 0.97 respectively) with non-significant P value. While SDF and IL4R had shown less mean expression in SVF2 as compared to SVF1 (P = 0.99 in both genes) but the differences are not statistically significant. The remaining three genes NOS3, MMP3 and P16 had shown almost equal mean expression in both SVF1 and SVF2 (P > 0.99 in all genes). These results suggest that both SVF1 and SVF2 maintain similar transcriptional profiles, and although SVF2 may trend toward a more regenerative gene expression pattern, the differences are not robust enough to reach statistical significance.
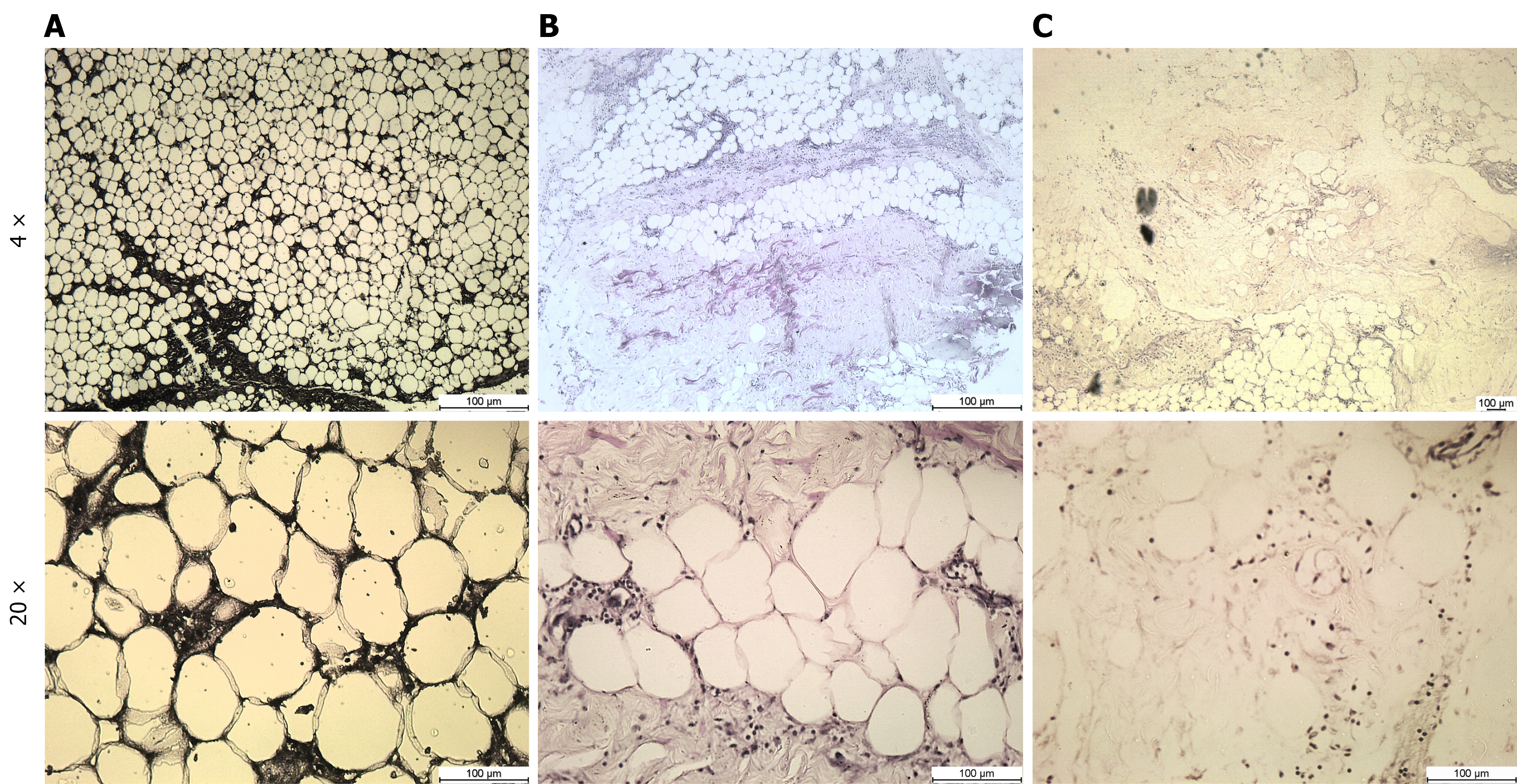
Histological analysis was performed to visualize structural changes in AT across three conditions: Undigested (control), after the first collagenase digestion, and after the second digestion. Key histological features were assessed, including adipocyte morphology, ECM integrity, and the density of stromal vascular cells remaining within the tissue (Table 2). This analysis also aimed to correlate the localization of regenerative cell populations within the tissue matrix and their progressive release during enzymatic digestion. Figure 6 shows the representative images of each digestion stage.
| Feature | Image 1 | Image 2 | Image 3 |
| Adipocyte presence and size | Visible; small and densely packed | Visible; large and sparsely distributed | Not detectable or structurally degraded |
| Adipocyte integrity | Intact membranes | Somewhat deformed | Completely lysed or absent |
| Cell density | Dense regenerative cells | Moderate cell presence | Low regenerative cells (absence of adipocytes, only entrapped nuclei seen) |
| ECM visibility | Moderately apparent | Minimally visible | Highly apparent, rich in collagen content |
Histological evidence supports the functional rationale of sequential digestion to release regenerative cells from AT. In undigested samples, AT served as a cellular and structural reservoir, with multipotent stromal cells such as AT-derived stem cells (ADSCs), pericytes, and endothelial progenitors, embedded in the ECM. These cells remained sparsely distributed and tightly integrated within the matrix. Following the first collagenase digestion, early tissue dissociation was evident. While most adipocytes retained their structure, ECM thinning and wider intercellular spacing indicated partial matrix degradation. Many cells remained associated with residual matrix showing partial release. After the second digestion, AT showed significant structural breakdown. Adipocytes were largely disrupted and ECM components were degraded or absent. Most cells had been released.
The present study investigated a novel approach to increase the number of stem cells for clinical use, particularly, for the patients with limited adipose reserves such as pediatric patients. The study employed a two-step sequential SVF isolation method through enzymatic digestion of AT. The conventional SVF isolation protocols typically rely on a single digestion phase and discard undigested AT. However, the results of the current study confirm that the residual AT contains a significant number of regenerative cells that can be retrieved through further digestion of it. By combining SVF2 with SVF1, the total cell yield can be significantly increased, potentially enhancing the clinical outcomes of SVF-based cell therapies. We selected a sequential two step digestion method because prolonged collagenase digestion was previously found detrimental to the viability of AT derived cells in SVF[11]. Therefore, in our novel two-step enzymatic digestion protocol, we reduced the collagenase exposure time for the released cells to avoid potential damage. Initially SVF was harvested with standard protocol to obtain the first pellet (SVF1) while the remaining undigested AT was further processed to obtain a second pellet (SVF2). Here, we aimed to explore whether SVF2, obtained after a second collagenase digestion step, retains functional, viable cells. Our findings demonstrated that SVF2 contains a significant number of viable cells that could be used to attain the relevant number of cells for stem cell-based therapies, particularly, when AT quantity is limited.
Both SVF1 and SVF2 demonstrated high viability (> 95%), with SVF1 showing a slightly higher percentage of viable cells, although the difference was statistically not significant. The results indicated that SVF1 contained a significantly higher number of total as well as mononuclear cells as compared to SVF2. The significantly higher total and mononuclear cell counts in SVF1 reflects the greater ease of detachment of loosely embedded stromal cells during the initial digestion phase. Similar outcomes were reported in another study that observed maximal release of ASCs and stromal cells during the initial 30 minutes of digestion[11]. Interestingly, although SVF2 yielded fewer total and mononuclear cells, it showed a higher percentage of mononuclear cells relative to the total cell population. The mononuclear cells in SVF include lymphocytes (T cells, B cells), monocytes, macrophages, natural killer cells, endothelial cells, preadipocytes, fibroblasts and MSCs or ASCs etc. Most of which are easy to isolate because these cells are loosely attached and several of them comes from the blood itself. In the second round of AT digestion, the mononuclear cells which are more closely associated with the ECM are released with more enzymatic penetration. Our data support the hypothesis proposed by another study[12], that distinct stromal niches within AT are accessed progressively, and that pericyte-rich fractions may require extended digestion times for effective liberation.
CFU assay provides an indirect measure of stemness and proliferative potential. This assay determines the number of SVF-derived cells that are capable to form colonies. It is pertinent to note that not all types of mononuclear cells are able to adhere to plastic surfaces and form colonies. For examples, lymphocytes, monocytes, macrophages and natural killer cells do not reliably adhere to plastic surfaces, proliferate and form colonies. However, ADSCs i.e., ADSCs have the capacity to adhere to plastic surfaces and form adherent colonies. So, it means that the colonies formed depict the origin from ADSCs. ADSCs are multipotent MSCs derived from AT, capable of self-renewal and differentiation into adipocytes, osteoblasts, and chondrocytes. They also exhibit strong immunomodulatory and regenerative properties. While the total number of colonies are significantly higher in SVF1 as compared to SVF2, SVF2 had a higher PE than SVF1. PE refers to the percentage of cells that are capable of adhering to the culture surface, surviving, and proliferating to form colonies under in vitro conditions. This parameter reflects the functional viability and proliferative capacity of the cell population. The high PE in SVF2 suggests that this fraction holds more ADSCs. We also calculated the number of CFUs per 1000 mononuclear cells and found that SVF2 contains significantly high count. These findings highlight the superior clonogenic density of SVF2, which suggests a higher proportion of progenitor cells capable of expansion under in vitro conditions. Comparable findings were reported by another study[13], who noted that deeper stromal layers harbor cells with higher self-renewal potential, possibly due to their perivascular origin. A key observation is that SVF2, despite being smaller in yield, harbors more functionally potent clonogenic cells. This finding is similar to the work in which AT subjected to prolonged enzymatic digestion yielded SVF populations with enhanced clonogenicity and a better capacity for vascular differentiation[14]. Thus, our study contributes to the growing evidence that total yield alone is not the best indicator of regenerative quality.
Flow cytometry analysis confirmed the presence of classical MSC markers (CD44, CD73, and CD146) in both SVF1 and SVF2. However, the expression of these markers was not significantly different between SVF1 and SVF2. The comparable number of CD44, CD73 and CD146 positive cells indicate that even after the first enzymatic digestion of AT, a significant number of MSC like cells i.e., ADSCs remained in SVF2. CD44 was more prominent in SVF1, which is consistent with reports by Zuk et al[15], they observed strong CD44 expression in early-released SVF cells. In contrast, CD73 and CD146 expression was relatively higher in SVF2. Although the differences were not statistically significant, CD73 and CD146 enrichment in SVF2 may indicate a shift toward a more therapeutically relevant MSC phenotype. CD146, a pericyte-associated marker, was present in both fractions but was particularly notable in SVF2. This supports the findings that CD146+ pericytes are concentrated around small vessels and are often retained in the deeper AT ECM. Their release during the second digestion step suggests that SVF2 could be particularly enriched in perivascular progenitors, which play key roles in angiogenesis and tissue remodeling. The primary aim of this study was to compare stromal and progenitor cell retention between SVF1 and SVF2 rather than to establish full ISCT-compliant MSC identity, we used representative MSC-associated markers (CD44, CD73, CD146). Future investigations may employ the full ISCT panel such as CD73, CD90, CD105 with corresponding negative markers to provide an extended phenotypic profile.
The biological function of both SVF fractions was further evaluated by gene expression profiling. The panel comprised genes that were involved in angiogenesis (VEGF), homing (SDF), mobilization (NOS3), matrix remodeling (MMP3), immunomodulation (IL4R), differentiation (CD14), and cell cycle regulation (P16, P53). Although none of the differences reached statistical significance, SVF2 showed trends towards higher expression of several genes that suggest functional cells in SVF2. The higher expression of these markers in SVF2 suggests that the cells in SVF2 have a repair, immunological and angiogenic potential[16]. The similar expression of tumor suppressor genes P16 and P53 in both fractions is reassuring from a safety standpoint, as it suggests that neither population has aberrant proliferative profiles. NOS3 and MMP3, involved in nitric oxide synthesis and ECM remodeling respectively, showed uniform expression, suggesting that both fractions retain matrix-interactive potential necessary for wound healing applications[17,18]. The gene expression data support the idea that SVF2 retains transcriptional competence and regenerative functionality similar to SVF1. This finding challenges the notion that extended digestion leads to inferior or senescent cell populations, and instead supports the view that sequential enzymatic digestion enhances accessibility to distinct regenerative niches.
Histological assessment further substantiated the sequential release of regenerative cells from AT. The control (undigested) samples showed intact adipocytes and ECM, with regenerative cell populations embedded within perivascular and interstitial zones. After the first digestion, partial dissociation was observed, while AT after the second digestion resulted in substantial structural breakdown and stromal cell release. These results are consistent with previous work which described the gradual release of regenerative cells from vascular and matrix compartments with prolonged collagenase exposure during a single digestion[19]. However, this is the first study to develop a two-step enzymatic digestion procedure, demonstrating that a second round of collagenase digestion yields a higher number of regenerative cells. Importantly, these histological changes correlate with functional outcomes observed in other assays such as CFU assays, qPCR and flow cytometry. The deep structural disintegration observed after the second digestion provides morphological evidence of the release of tightly bound pericytes and matrix-anchored progenitors. Thus, our study highlights the physical and biological rationale for sequential SVF extraction and justifies the added processing step in protocols aiming for maximal regenerative harvest.
From a clinical perspective, the data provide compelling evidence to consider SVF2 not as residual waste but as a valuable biological resource. In clinical applications particularly those involving autologous cell therapy, soft tissue repair, or angiogenic induction, SVF2 can add the functional attributes to SVF1 for better clinical outcomes. The findings suggest that clinicians and researchers should reconsider single-step digestion protocols and explore modified processing strategies to harness the full potential of AT especially when AT is not available in sufficient quantities. Moreover, the concept of tailoring cell-based therapies based on SVF composition is gaining traction. While this study presents important insights, certain limitations should be considered. The relatively small sample size may limit statistical power in some assays, and further validation in larger cohorts is warranted. Additionally, functional assays such as tri-lineage differentiation, angiogenesis assays, or in vivo transplantation models could enhance understanding of the long-term regenerative efficacy of SVF2 cells.
The present study demonstrates that two-step sequential enzymatic digestion of AT significantly enhances SVF yield by recovering a second viable and regenerative cell population (SVF2) typically discarded in conventional protocols. While SVF1 yields higher cell numbers, SVF2 was enriched in CFU forming cells, showed a favorable MSC marker profile and showed supportive gene expression trends. These findings highlight the biological and therapeutic relevance of SVF2 and support the adoption of multi-step AT processing to maximize regenerative potential, especially when tissue availability is limited (Figure 7).
The research was partially supported by a grant from the Higher Education Commission of Pakistan. The authors are grateful to the Department of Immunology and the Department of Histopathology for their valuable assistance in this study.
| 1. | Ramakrishnan VM, Boyd NL. The Adipose Stromal Vascular Fraction as a Complex Cellular Source for Tissue Engineering Applications. Tissue Eng Part B Rev. 2018;24:289-299. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 114] [Cited by in RCA: 128] [Article Influence: 16.0] [Reference Citation Analysis (2)] |
| 2. | Andia I, Maffulli N, Burgos-Alonso N. Stromal vascular fraction technologies and clinical applications. Expert Opin Biol Ther. 2019;19:1289-1305. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 106] [Cited by in RCA: 85] [Article Influence: 12.1] [Reference Citation Analysis (1)] |
| 3. | Choudhery MS, Arif T, Mahmood R. Cancer stem cells: Review of current state and future directions. Biomed Current Insights. 2024;1:1-6. [DOI] [Full Text] |
| 4. | Vuerich R, Groppa E, Vodret S, Ring NAR, Stocco C, Bossi F, Agostinis C, Cauteruccio M, Colliva A, Ramadan M, Simoncello F, Benvenuti F, Agnelli A, Dore F, Mazzarol F, Moretti M, Paulitti A, Palmisano S, De Manzini N, Chiesa M, Casaburo M, Raucci A, Lorizio D, Pompilio G, Bulla R, Papa G, Zacchigna S. Ischemic wound revascularization by the stromal vascular fraction relies on host-donor hybrid vessels. NPJ Regen Med. 2023;8:8. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 13] [Reference Citation Analysis (1)] |
| 5. | Roshdy OH, Abdallah WI, Farid CI, Mehanna RA, Bayoumi NH, Ismail AI. Stromal vascular fraction improves the durability of autologous fat temple augmentation-A split-face randomized study using ultrasound biomicroscopy. J Plast Reconstr Aesthet Surg. 2022;75:1870-1877. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 13] [Article Influence: 3.3] [Reference Citation Analysis (2)] |
| 6. | Mantovani GP, Marra C, De Maria F, Pinto V, De Santis G. Adipose-derived stromal vascular fraction (SVF) for the treatment of androgenic alopecia (AGA): a systematic review. Acta Biomed. 2023;94:e2023236. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (2)] |
| 7. | Butt G, Hussain I, Ahmad FJ, Choudhery MS. Stromal vascular fraction-enriched platelet-rich plasma therapy reverses the effects of androgenetic alopecia. J Cosmet Dermatol. 2020;19:1078-1085. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 33] [Article Influence: 5.5] [Reference Citation Analysis (1)] |
| 8. | Tauer JT, Al-Jallad H, Umebayashi M, Bakhsh D, Rauch D, Tran SD, Rauch F, Hamdy R. Characterization and functional analysis of the adipose tissue-derived stromal vascular fraction of pediatric patients with osteogenesis imperfecta. Sci Rep. 2022;12:2414. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 8] [Reference Citation Analysis (1)] |
| 9. | Choudhery MS, Badowski M, Muise A, Pierce J, Harris DT. Subcutaneous Adipose Tissue-Derived Stem Cell Utility Is Independent of Anatomical Harvest Site. Biores Open Access. 2015;4:131-145. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 28] [Cited by in RCA: 32] [Article Influence: 2.9] [Reference Citation Analysis (3)] |
| 10. | Umer K, Khan A, Pervaiz A. Effects of a ribosome inactivating plant protein (riproximin) on transcriptomic profile of apoptosis pathway genes in breast cancer cells. Biomed Current Insights. 2024;1:29-34. [DOI] [Full Text] |
| 11. | Seaman SA, Tannan SC, Cao Y, Peirce SM, Lin KY. Differential Effects of Processing Time and Duration of Collagenase Digestion on Human and Murine Fat Grafts. Plast Reconstr Surg. 2015;136:189e-199e. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 19] [Article Influence: 1.7] [Reference Citation Analysis (1)] |
| 12. | West CC, Hardy WR, Murray IR, James AW, Corselli M, Pang S, Black C, Lobo SE, Sukhija K, Liang P, Lagishetty V, Hay DC, March KL, Ting K, Soo C, Péault B. Prospective purification of perivascular presumptive mesenchymal stem cells from human adipose tissue: process optimization and cell population metrics across a large cohort of diverse demographics. Stem Cell Res Ther. 2016;7:47. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 40] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 13. | Baer PC. Adipose-derived mesenchymal stromal/stem cells: An update on their phenotype in vivo and in vitro. World J Stem Cells. 2014;6:256-265. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 126] [Cited by in RCA: 140] [Article Influence: 11.7] [Reference Citation Analysis (1)] |
| 14. | Chen SY, Mahabole M, Horesh E, Wester S, Goldberg JL, Tseng SC. Isolation and characterization of mesenchymal progenitor cells from human orbital adipose tissue. Invest Ophthalmol Vis Sci. 2014;55:4842-4852. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 21] [Article Influence: 1.8] [Reference Citation Analysis (1)] |
| 15. | Zuk PA, Zhu M, Mizuno H, Huang J, Futrell JW, Katz AJ, Benhaim P, Lorenz HP, Hedrick MH. Multilineage cells from human adipose tissue: implications for cell-based therapies. Tissue Eng. 2001;7:211-228. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6425] [Cited by in RCA: 5496] [Article Influence: 219.8] [Reference Citation Analysis (15)] |
| 16. | Navarro A, Marín S, Riol N, Carbonell-Uberos F, Miñana MD. Human adipose tissue-resident monocytes exhibit an endothelial-like phenotype and display angiogenic properties. Stem Cell Res Ther. 2014;5:50. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 30] [Cited by in RCA: 27] [Article Influence: 2.3] [Reference Citation Analysis (1)] |
| 17. | Gill SE, Parks WC. Metalloproteinases and their inhibitors: regulators of wound healing. Int J Biochem Cell Biol. 2008;40:1334-1347. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 577] [Cited by in RCA: 517] [Article Influence: 27.2] [Reference Citation Analysis (1)] |
| 18. | Witte MB, Barbul A. Role of nitric oxide in wound repair. Am J Surg. 2002;183:406-412. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 361] [Cited by in RCA: 329] [Article Influence: 13.7] [Reference Citation Analysis (1)] |
| 19. | Cao L, Xiaoming F, Zhang Q, Fang J, Chu C, Lv J, Ma Y, Lu G, Yang K, Pan R. An Optimized Method for Adipose Stromal Vascular Fraction Isolation and its Application in Fat Grafting. Aesthetic Plast Surg. 2022;46:2500-2508. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 10] [Article Influence: 2.5] [Reference Citation Analysis (1)] |