Published online Mar 26, 2026. doi: 10.4252/wjsc.v18.i3.118401
Revised: January 22, 2026
Accepted: March 9, 2026
Published online: March 26, 2026
Processing time: 84 Days and 15.7 Hours
Male infertility affects approximately 17.5% of the global reproductive-age po
Core Tip: This review integrates single-cell transcriptomics data from 2018 to 2025 to reconstruct the pathological landscape of male infertility, identifying the spermatogonial stem cell niche as a core therapeutic target. We elucidate a novel me
- Citation: Ying ZK, Xu XY, Hu JW, Cui Y, Jiang DW. Single-cell transcriptomics reveals a spermatogonial stem cell-centered spermatogenic microenvironment: Pathophysiological mechanisms and therapeutic targets in male infertility. World J Stem Cells 2026; 18(3): 118401
- URL: https://www.wjgnet.com/1948-0210/full/v18/i3/118401.htm
- DOI: https://dx.doi.org/10.4252/wjsc.v18.i3.118401
Male infertility affects approximately 15%-20% of couples worldwide[1,2] and accounts for nearly half of all cases of reproductive failure[3], posing a substantial yet often underestimated public health burden. In addition to genetic and endocrine factors, increasing evidence indicates that disruption of the testicular spermatogenic microenvironment is a decisive driver of impaired spermatogenesis. In particular, the spermatogonial stem cell (SSC) niche plays a central role in maintaining continuous sperm production, as SSC self-renewal and differentiation are tightly regulated by surrounding Sertoli cells, Leydig cells, immune components, vascular structures, and extracellular matrix-mediated signaling[4-6].
The advent of single-cell RNA sequencing (scRNA-seq) has revolutionized the study of human spermatogenesis[7-10] by enabling high-resolution characterization of cellular heterogeneity and developmental trajectories that cannot be resolved by conventional bulk transcriptomic approaches. Large-scale single-cell atlases of the human testis have de
However, several critical gaps remain insufficiently integrated. Most existing reviews primarily emphasize atlas construction or germ cell differentiation programs, whereas the mechanistic consequences of microenvironmental re
This review was conducted as a narrative and integrative synthesis of recent advances in human testicular single-cell research, rather than a formal systematic review[39]. We focused on original studies published between January 2018 and September 2025 that applied scRNA-seq and, where available, spatial transcriptomic technologies to characterize the cellular composition and microenvironmental regulation of human spermatogenesis. Literature searches were performed primarily in PubMed and Web of Science using combinations of keywords including “single-cell RNA sequencing”, “testis”, “spermatogenesis”, “spermatogonial stem cell”, and “male infertility”.
Studies were included if they: (1) Analyzed human testicular tissue; (2) Generated original scRNA-seq or spatial transcriptomic datasets; and (3) Provided biologically interpretable results relevant to germ cell development, somatic cell function, or microenvironmental regulation, with particular attention to SSCs and NOA. We excluded studies based exclusively on animal models, technical reports without substantive biological analysis, conference abstracts, and narrative commentaries. Reviews were used only to contextualize findings but were not considered primary sources of evidence.
To address population-related bias, we recorded the reported geographic origin of study cohorts whenever available and considered this factor when interpreting conserved vs population-specific molecular signatures. Given the substantial heterogeneity across sequencing platforms, tissue dissociation protocols, and computational pipelines, no quantitative meta-analysis was attempted. Instead, we adopted a qualitative comparative approach, emphasizing reproducible cellular patterns, convergent signaling pathways, and mechanistic consistency across independent datasets[40,41].
scRNA-seq has become the principal technology for dissecting cellular heterogeneity within the human testis, enabling unbiased identification of rare cell populations, developmental intermediates, and disease-associated transcriptional states. In contrast to bulk transcriptomics, which averages signals across mixed cell types, scRNA-seq resolves cell type-specific gene expression patterns and reconstructs differentiation trajectories, thereby providing a powerful framework for studying SSCs, somatic support cells, and their coordinated regulation within the spermatogenic niche[42,43].
Despite these advantages, several technical limitations should be acknowledged when interpreting scRNA-seq-based findings. Tissue dissociation can introduce transcriptional stress responses and preferentially affect fragile cell popu
More recently, spatial transcriptomic technologies have complemented scRNA-seq by preserving the anatomical context of gene expression, enabling mapping of cellular interactions within seminiferous tubules and interstitial compartments[48]. This spatial information is particularly valuable for characterizing the organization of the SSC niche and its interactions with supporting somatic and immune cell populations[49]. However, current platforms remain constrained by limited spatial resolution, signal mixing within capture spots, and substantial computational complexity during data integration. Consequently, most available studies adopt a combined strategy in which scRNA-seq defines cell identities and spatial transcriptomics provides positional information[50,51]. Together, these technologies form the methodological foundation for investigating microenvironmental remodeling in male infertility, while their technical constraints highlight the need for continued methodological refinement[52].
Single-cell transcriptomic profiling has delineated the major cellular constituents of the adult human testis, including SSCs and differentiating germ cells, Sertoli cells, Leydig cells, peritubular myoid cells, endothelial cells, and diverse immune populations. Within the germline compartment, scRNA-seq has resolved discrete transcriptional states spanning undifferentiated spermatogonia, differentiating spermatogonia, spermatocytes, and post-meiotic spermatids, enabling reconstruction of spermatogenic trajectories with unprecedented resolution[11,13,14]. Importantly, SSCs are now re
Beyond germ cells, single-cell atlases have uncovered substantial heterogeneity among somatic and immune cell com
Comparative analyses between fertile controls and patients with NOA have consistently demonstrated profound microenvironmental remodeling, characterized by altered Sertoli cell maturation states, increased inflammatory signaling, endothelial dysfunction, and depletion or transcriptional reprogramming of SSC populations[19-21]. Dis
| Ref. | Journal | Sample size & type | Platform (cells analyzed) | Key findings | Clinical relevance | Geographic location |
| Wang et al[23], 2018 | Cell Stem Cell | n = 3 normal; adult donors | 10X Genomics (approximately 6490 cells) | First comprehensive atlas of human testis; identified 5 germ cell states; characterized niche-specific gene signatures; discovered PLPPR3+ spermatogonia subtype; state 0 SSC defined | Established baseline reference for normal spermatogenesis; identified SSC markers for potential therapeutic targeting | China (Beijing) |
| Hermann et al[24], 2018 | Cell Reports | n = 7 normal; adult testis (18-36 years) | Fluidigm C1 + 10X (approximately 62000 total cells, 182 SSCs) | Focused analysis of SSC heterogeneity; defined ID4+/UTF1+ SSC population; revealed FGFR3 as functional SSC marker; complete spermatogenesis transcriptome mapping | Potential for SSC isolation and expansion for fertility preservation | United States (Pittsburgh) |
| Guo et al[25], 2018 | Cell Research | n = 3 normal; adult males | 10X Genomics (approximately 6500 cells) | Adult human testis transcriptional cell atlas; state 0 SSC identification; RNA velocity reveals spermatogonial plasticity; epigenetic landscape analysis | Understanding SSC self-renewal and differentiation balance; SSC-specific marker identification | United States (UT) |
| Sohni et al[26], 2019 | Cell Reports | Neonatal (n = 2) and adult (n = 3) human testis | 10X Genomics (approximately 17000 cells) | Defined neonatal and adult human testis at single-cell level; identified four undifferentiated spermatogonia clusters; characterized protein markers for primitive SPG state; mapped timeline from PGCs to adult SPG | SSC-enriched cell subset purification; understanding developmental trajectories | United States (CA) |
| Guo et al[27], 2020 | Cell Stem Cell | n = 4 normal; juvenile males (7-14 years) | 10X Genomics (approximately 10000 cells) | Pubertal testis development atlas; identified common pre-pubertal progenitor for Leydig and myoid cells; two distinct pre-pubertal Sertoli cell states; testosterone roles via transfemale testis analysis | Insights into pubertal developmental disorders; critical windows for testicular maturation | United States (UT) |
| Guo et al[28], 2021 | Cell Stem Cell | n = 10 normal; embryonic to infant (6 weeks to 5 months) | 10X Genomics (approximately 32500 cells) | Fetal testis development atlas; sertoli and interstitial cells from common progenitor at 6-7 weeks; PGCs transition to state 0-like cells at 14-16 weeks; somatic niche specification precedes germline transition | Understanding fetal gonadal development; insights into congenital testicular disorders | United States (UT/UCLA) |
| Shami et al[29], 2020 | Developmental Cell | n = 8 normal; young adults (17-25 years) | 10X Genomics (> 35000 cells) | Comprehensive human-mouse comparison; species-specific gene expression programs; extended meiotic progression in humans; novel markers for meiotic stages | Caution for translating mouse models; human-specific therapeutic targets | United States (MI) |
| Zhao et al[30], 2020 | Nature Communications | n = 10 NOA, n = 10 normal (infant to adult) | 10X Genomics (> 88000 cells) | First large-scale NOA single-cell analysis; three-stage Sertoli cell maturation roadmap; Sertoli cell maturation blockade in iNOA; inflammatory microenvironment signature; loss of SSC niche factors (GDNF, FGF2) | Identified targetable pathways in NOA; potential biomarkers for diagnosis; Sertoli cell-centered therapeutic approach | China (Nanjing) |
| Di Persio et al[31], 2021 | Cell Reports Medicine | n = 5 NOA, n = 5 cryptozoospermia, n = 5 normal | 10X Genomics (> 24000 cells) | Comparative analysis of impaired spermatogenesis; major alterations in cryptozoospermia SSC compartment; increased PIWIL4+ undifferentiated spermatogonia; transcriptional switch driven by EGR4 overexpression; reduced UTF1+ reserve spermatogonia (Adark) | Distinct pathological mechanisms guide personalized treatment | Germany (Münster) |
| Alfano et al[32], 2021 | Nature Communications | n = 8 infertile, n = 3 normal (TESE samples) | Smart-seq2 (1246 cells) | Aging, inflammation and DNA damage in somatic niche; idiopathic germ cell aplasia pathology; senescence and immune activation in Sertoli cells; testicular M1 macrophage polarization; chronic inflammation signature | Hormonal therapy optimization; testosterone production defects in infertility | Italy (Milan) |
| Chen et al[33], 2021 | Cell Reports | n = 3 normal adult testis (mouse + human) | 10X Genomics + slide-seq spatial transcriptomics | Spatial transcriptomic atlas of mammalian spermatogenesis; near-single-cell resolution spatial gene expression; identified spatially patterned genes along seminiferous tubules; Habp4 as chromatin remodeling regulator; compared WT vs diabetic mouse testis | Understanding spatial organization for targeted therapy; zone-specific molecular signatures | United States (Harvard/Broad Institute) |
| Mahyari et al[34], 2021 | American Journal of Human Genetics | n = 3 Klinefelter syndrome, n = 3 normal controls | 10X Genomics (approximately 13000 cells) | First single-cell analysis of Klinefelter testis; identified immature sertoli and Leydig cells; revealed pro-inflammatory macrophage enrichment; discovered altered microenvironment in KS | Understanding Klinefelter-specific pathology; potential therapeutic targets for KS | Estonia (Tartu) |
| Nie et al[35], 2022 | Developmental Cell | n = 12 normal (young + older), n = 6 with elevated BMI | 10X Genomics (> 44000 cells) | Human testis aging study; age-related changes in spermatogenesis and somatic cells; altered pathways: Inflammation, metabolic signaling in Sertoli cells, hedgehog/testosterone in Leydig cells; BMI correlation with dysregulation in older men; cell-cell communication changes during aging | Age-specific fertility preservation strategies; BMI management for reproductive health in aging men | United States (UT) |
| Di Persio and Neuhaus[36], 2023 | Human Reproduction | Review article covering multiple NOA subtypes | N/A (comprehensive review) | Comprehensive scRNA-seq review; novel concepts on SSC subtypes (state 0, state 1); SSC niche crosstalk mechanisms; transcriptional alterations in NOA, cryptozoospermia, Klinefelter syndrome, AZF deletions | Precision medicine approaches based on genetic etiology; marker genes for SSC subsets | Germany (Münster) |
| Amodio et al[37], 2025 | Nature Communications | NOA and OAT patients vs controls | scRNA-seq + multiparameter phenotyping | Different infertility subtypes correlated with T cell exhaustion/senescence signatures; young infertile men show pro-inflammatory milieu (similar to healthy elderly men); immune alterations in seminal fluid and peripheral blood; interferon-gamma and -alpha response upregulation | Identifies infertility-specific immune signatures; suggests personalized immunomodulatory treatment strategies; links infertility to systemic health | Italy (Milan) |
| Cui et al[38], 2025 | Nature Aging | n = 35 normal donors (21-69 years) | 10X Genomics (214369 cells) | Machine learning reveals somatic cells show stronger aging response than germ cells; two waves of aging-related changes: Age 30-39 years old (peritubular cells, basement membrane thickening), Age 50-59 years old (functional changes in Leydig cells and macrophages); BMI impact on spermatogenic capacity after age 45 | Age-specific fertility preservation; potential diagnostic markers and therapeutic targets; BMI management critical for reproductive health in aging men | China (multi-center) |
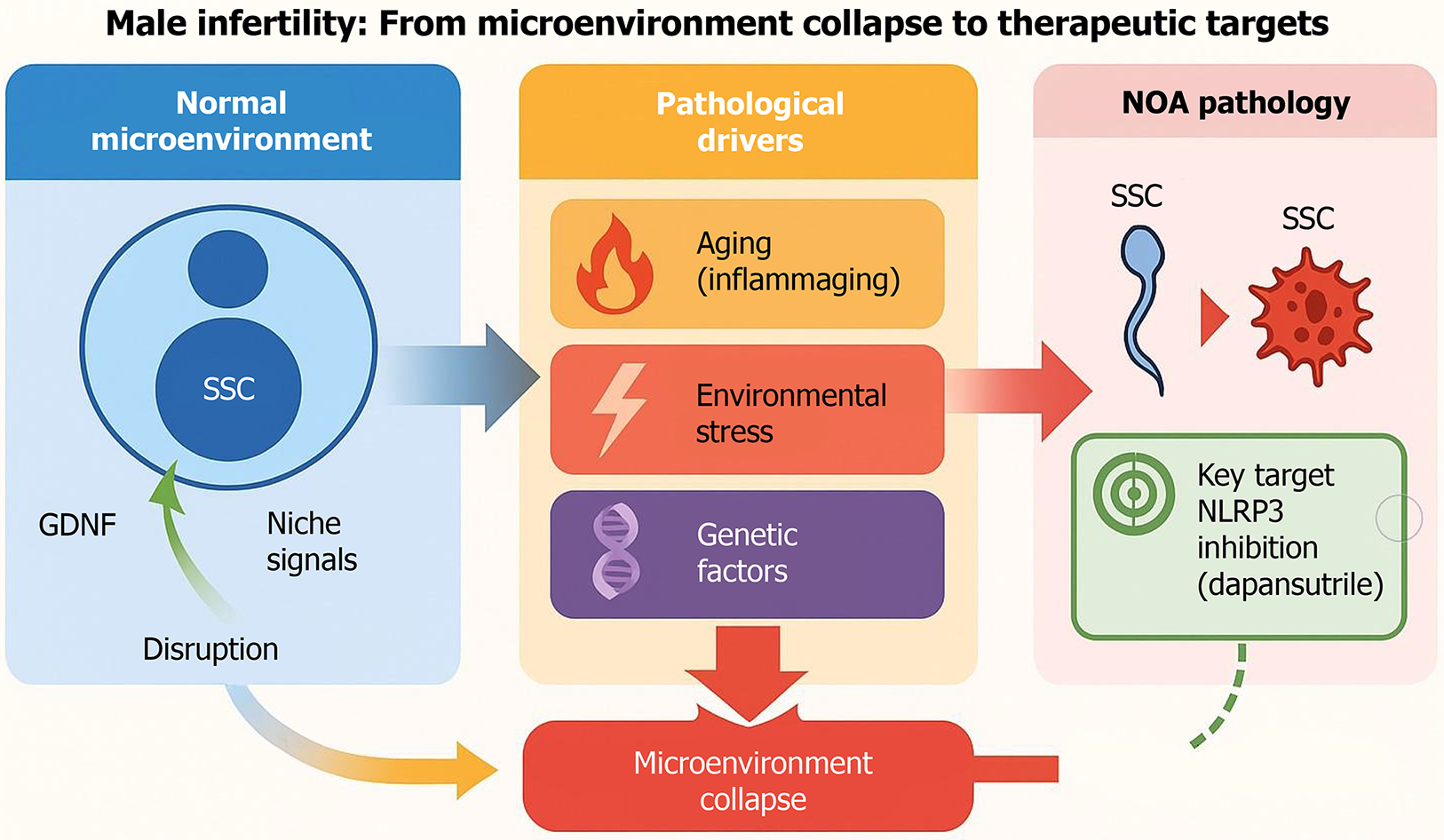
Non-obstructive azoospermia represents the most severe form of spermatogenic failure and is a major cause of male infertility[3]. Single-cell transcriptomic studies have revealed that NOA is not merely characterized by the absence of mature germ cells, but rather by profound reorganization of the entire testicular microenvironment[19-21]. Compared with fertile controls, testes from patients with NOA exhibit marked alterations in Sertoli cell maturation states, impaired metabolic support programs, endothelial dysfunction, and expansion of inflammatory immune cell subsets. A schematic overview of these interconnected pathological mechanisms is provided in Figure 1.
The pathological cascade initiates with testicular niche dysfunction (Sertoli cell maturation arrest, M1-like macrophage polarization, Leydig cell decline), amplified by endogenous aging and environmental exposures, converging on NLRP3 inflammasome and mammalian target of rapamycin pathway activation. This leads to SSC depletion and spermatogenic failure. Genetic variants and transgenerational epigenetic inheritance via sperm small non-coding RNAs further modulate susceptibility. Red denotes pathological states; blue indicates therapeutic intervention points.
Within the germline compartment, SSCs display reduced abundance, transcriptional instability, and impaired differentiation trajectories. These changes are accompanied by disruption of key paracrine signaling pathways, including glial cell line-derived neurotrophic factor, KIT-KITLG, and extracellular matrix-mediated interactions, suggesting that SSC exhaustion in NOA reflects failure of niche support rather than an isolated intrinsic defect.
Accumulating evidence indicates that chronic low-grade inflammation associated with aging, metabolic disorders, and environmental stress - commonly referred to as “inflammaging” - plays a central role in reshaping the testicular microenvironment[56]. Single-cell analyses consistently demonstrate upregulation of pro-inflammatory cytokines, chemokines, and stress-response pathways in Sertoli cells, Leydig cells, and resident immune populations in NOA testes[57,58].
This persistent inflammatory milieu disrupts blood-testis barrier integrity, alters redox homeostasis, and interferes with cellular metabolic coupling between somatic cells and germ cells[59]. Importantly, SSCs are highly sensitive to oxidative stress and inflammatory signaling, which can impair self-renewal capacity and promote premature differentiation or apoptosis. Thus, inflammaging provides a systemic and local context that predisposes the spermatogenic niche to progressive functional decline.
Among inflammatory pathways, activation of the NLRP3 inflammasome has emerged as a key molecular node connecting chronic inflammatory stress to testicular dysfunction. Single-cell and bulk transcriptomic studies have reported increased expression of NLRP3, CASPASE-1, interleukin-1β, and interleukin-18 in Sertoli cells, macrophages, and endothelial cells from NOA samples, indicating inflammasome activation within the niche[60]. Functionally, NLRP3-driven cytokine release amplifies local inflammation, promotes endothelial permeability, and perturbs Sertoli cell-mediated metabolic and structural support[61]. Experimental models further suggest that inflammasome activation directly compromises SSC survival by inducing mitochondrial dysfunction, DNA damage responses, and apoptotic signaling. Together, these findings support a pathogenic cascade in which inflammaging triggers sustained NLRP3 activation, leading to progressive deterioration of niche integrity and depletion of the SSC pool.
Although chronic inflammation represents a central driver of microenvironmental dysfunction in NOA, additional pathological mechanisms also contribute to SSC impairment. Single-cell and bulk transcriptomic studies have consistently reported increased oxidative stress signatures, mitochondrial dysfunction, and activation of DNA damage response pathways in both somatic and germ cell populations[62]. Excessive reactive oxygen species disrupt mitochondrial energy metabolism in Sertoli cells and SSCs, thereby compromising cellular homeostasis and self-renewal capacity[63-65].
Furthermore, mitochondrial injury and genomic instability may sensitize SSCs to inflammatory cytokines and inflammasome-mediated apoptosis, acting synergistically with NLRP3 activation to accelerate niche deterioration. These non-inflammatory pathways likely operate in parallel with chronic immune remodeling, collectively driving progressive collapse of the spermatogenic microenvironment. Recognition of these convergent mechanisms underscores the need for therapeutic strategies that target not only inflammatory signaling but also metabolic and genomic stress responses within the SSC niche.
Beyond direct impairment of spermatogenesis, microenvironmental stress may exert longer-term biological consequences through epigenetic reprogramming of male germ cells. Experimental studies have demonstrated that metabolic disorders, chronic inflammation, and toxic environmental exposures can alter the composition of sperm small non-coding RNAs, including microRNAs and tRNA-derived fragments, which are capable of transmitting phenotypic traits to offspring[66-68].
Although evidence in humans remains limited, emerging data suggest that dysregulation of inflammatory and metabolic pathways within the testicular niche may influence the epigenetic landscape of SSC-derived germ cells[69]. These observations raise the possibility that chronic microenvironmental injury not only compromises fertility but may also contribute to intergenerational disease susceptibility. Genetic factors, including copy number variations such as DAZ gene deletions, further modulate individual vulnerability to microenvironmental stress[70]. Further large-scale human studies integrating single-cell transcriptomics with sperm epigenomic profiling will be required to clarify the clinical relevance and mechanistic basis of this phenomenon.
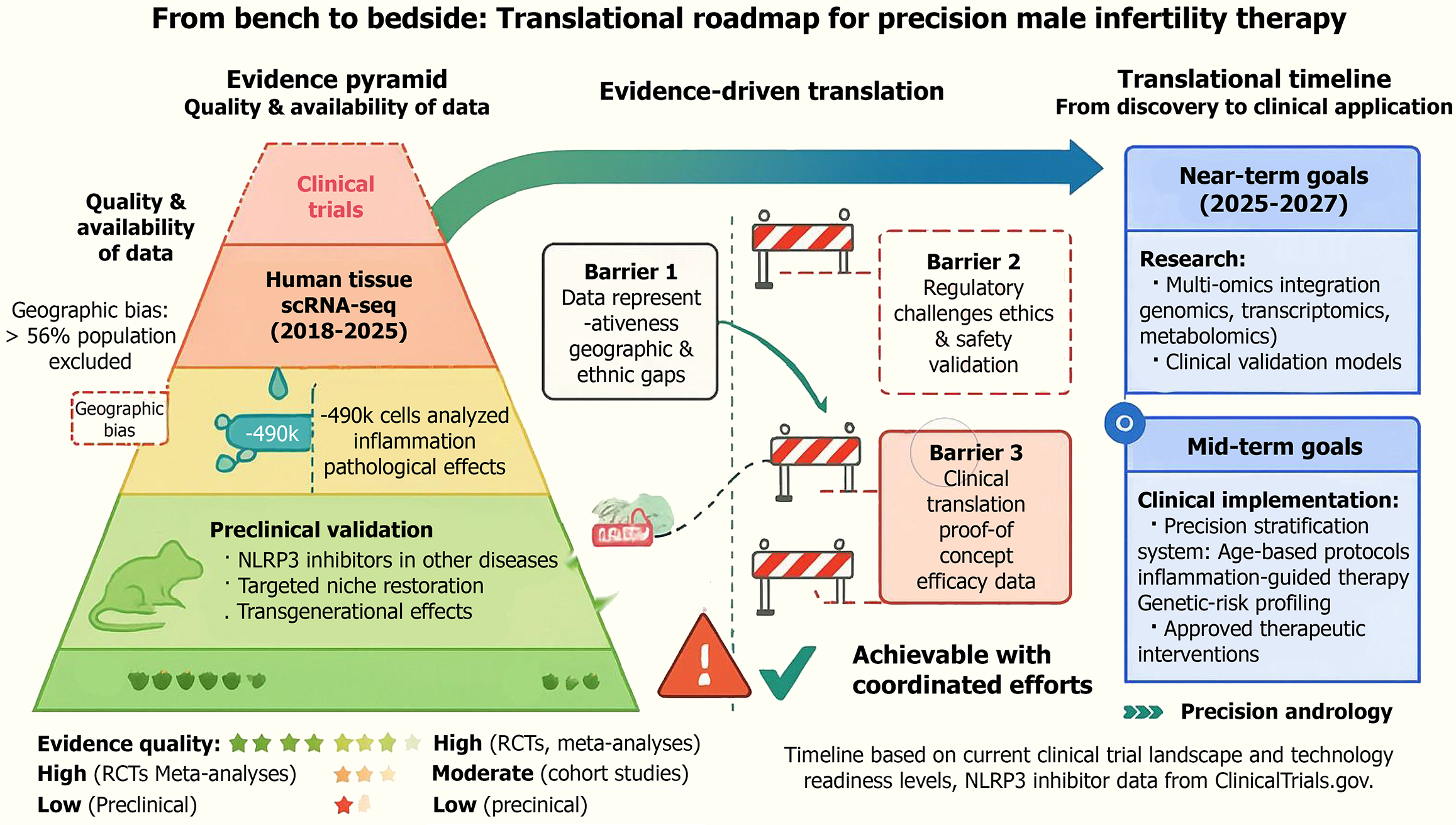
Given the central role of NLRP3 inflammasome activation in linking chronic inflammation to SSC dysfunction, pharmacological targeting of this pathway represents a promising therapeutic direction[60,61]. Preclinical studies in models of metabolic syndrome, testicular inflammation, and systemic inflammatory disorders have demonstrated that genetic or pharmacological inhibition of NLRP3 can attenuate cytokine release, preserve blood-testis barrier integrity, and improve germ cell survival[71]. In parallel, several small-molecule NLRP3 inhibitors, including dapansutrile, have entered clinical trials for inflammatory and metabolic diseases, showing favorable safety profiles and anti-inflammatory efficacy[72,73]. Key translational barriers between single-cell discoveries and clinical implementation are summarized in Figure 2.
The translational gap comprises three interconnected barriers: Technical limitations (batch effects, spatial information loss), data biases (geographic underrepresentation of non-European populations risking therapy failures in neglected groups), and ethical/socioeconomic challenges (germline editing concerns, healthcare inequity, data privacy). Bridging this gap requires globally diverse cohorts, bias-aware artificial intelligence integration, and equitable governance fra
Although direct clinical evidence in male infertility is currently lacking, these convergent findings support the bio
Advances in CRISPR-Cas9 technology and stem cell biology have raised the prospect of directly correcting genetic defects or regenerating impaired germline compartments[74,75]. In theory, genome editing could be applied to patient-derived SSCs followed by autologous transplantation, thereby avoiding systemic genetic modification[76]. However, multiple obstacles currently limit clinical translation, including inefficient delivery to target cells, off-target mutagenesis, genomic mosaicism, and long-term safety concerns[77].
Beyond technical barriers, germline manipulation raises profound ethical and regulatory challenges due to the po
Nanotechnology-based drug delivery systems offer an alternative strategy for modulating the spermatogenic niche without direct genetic intervention[81]. Nanoparticles engineered to cross or accumulate near the blood-testis barrier have shown potential for targeted delivery of anti-inflammatory agents, antioxidants, and small interfering RNAs in preclinical models[82]. Such approaches may enhance local drug concentration while minimizing systemic exposure and adverse effects. Nevertheless, substantial challenges persist, including biocompatibility, long-term tissue accumulation, immune activation, and incomplete understanding of nanoparticle transport dynamics within testicular tissue. At pre
The increasing feasibility of manipulating SSCs and their microenvironment raises important ethical considerations[78]. Although autologous SSC editing and transplantation theoretically avoid systemic genetic modification, any intervention targeting the germline carries the potential for heritable genomic alterations. Off-target mutations, mosaicism, and unpredictable long-term consequences remain substantial unresolved risks, particularly in the context of complex polygenic disorders such as male infertility[79,80].
International scientific and regulatory bodies currently maintain strict limitations on clinical germline editing, em
Another critical consideration is the pronounced geographic and socioeconomic imbalance in current single-cell datasets and translational research pipelines[85,86]. Most available scRNA-seq studies of the human testis originate from a limited number of high-income regions, whereas male infertility imposes a substantial burden in low- and middle-income countries where access to advanced reproductive technologies is often restricted[87].
This disparity raises two major concerns. First, molecular signatures and therapeutic targets derived from homoge
Single-cell transcriptomics has fundamentally reshaped the conceptual framework of male infertility, revealing it to be a disorder driven not only by germ cell defects but by coordinated dysfunction of the spermatogenic microenvironment and its resident SSC compartment. Evidence synthesized in this review supports a pathogenic model in which chronic inflammatory remodeling, particularly through activation of the NLRP3 inflammasome, contributes to progressive niche deterioration and SSC depletion in NOA. By integrating single-cell, spatial transcriptomic, and emerging epigenetic data, this work highlights the central role of microenvironmental regulation in sustaining spermatogenesis and identifies inflammatory signaling pathways as rational targets for therapeutic intervention. Although substantial technical, ethical, and population-related challenges remain, continued integration of multi-omics technologies and broader representation of diverse populations may facilitate the development of stem cell-oriented precision strategies for the diagnosis and treatment of male infertility.
| 1. | World Health Organization. 1 in 6 people globally affected by infertility: WHO. [cited 12 October 2025]. Available from: https://www.who.int/news/item/04-04-2023-1-in-6-people-globally-affected-by-infertility. |
| 2. | Feng J, Wu Q, Liang Y, Liang Y, Bin Q. Epidemiological characteristics of infertility, 1990-2021, and 15-year forecasts: an analysis based on the global burden of disease study 2021. Reprod Health. 2025;22:26. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 48] [Cited by in RCA: 29] [Article Influence: 29.0] [Reference Citation Analysis (1)] |
| 3. | Agarwal A, Baskaran S, Parekh N, Cho CL, Henkel R, Vij S, Arafa M, Panner Selvam MK, Shah R. Male infertility. Lancet. 2021;397:319-333. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1049] [Cited by in RCA: 828] [Article Influence: 165.6] [Reference Citation Analysis (4)] |
| 4. | Meng X, Lindahl M, Hyvönen ME, Parvinen M, de Rooij DG, Hess MW, Raatikainen-Ahokas A, Sainio K, Rauvala H, Lakso M, Pichel JG, Westphal H, Saarma M, Sariola H. Regulation of cell fate decision of undifferentiated spermatogonia by GDNF. Science. 2000;287:1489-1493. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1146] [Cited by in RCA: 972] [Article Influence: 37.4] [Reference Citation Analysis (1)] |
| 5. | Oatley JM, Brinster RL. The germline stem cell niche unit in mammalian testes. Physiol Rev. 2012;92:577-595. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 440] [Cited by in RCA: 388] [Article Influence: 27.7] [Reference Citation Analysis (1)] |
| 6. | Kubota H, Avarbock MR, Brinster RL. Growth factors essential for self-renewal and expansion of mouse spermatogonial stem cells. Proc Natl Acad Sci U S A. 2004;101:16489-16494. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 793] [Cited by in RCA: 709] [Article Influence: 32.2] [Reference Citation Analysis (1)] |
| 7. | Jia H, Wang W, Zhou Z, Chen Z, Lan Z, Bo H, Fan L. Single-cell RNA sequencing technology in human spermatogenesis: Progresses and perspectives. Mol Cell Biochem. 2024;479:2017-2033. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 9] [Article Influence: 4.5] [Reference Citation Analysis (1)] |
| 8. | Green CD, Ma Q, Manske GL, Shami AN, Zheng X, Marini S, Moritz L, Sultan C, Gurczynski SJ, Moore BB, Tallquist MD, Li JZ, Hammoud SS. A Comprehensive Roadmap of Murine Spermatogenesis Defined by Single-Cell RNA-Seq. Dev Cell. 2018;46:651-667.e10. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 220] [Cited by in RCA: 408] [Article Influence: 51.0] [Reference Citation Analysis (1)] |
| 9. | Grive KJ, Hu Y, Shu E, Grimson A, Elemento O, Grenier JK, Cohen PE. Dynamic transcriptome profiles within spermatogonial and spermatocyte populations during postnatal testis maturation revealed by single-cell sequencing. PLoS Genet. 2019;15:e1007810. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 114] [Cited by in RCA: 92] [Article Influence: 13.1] [Reference Citation Analysis (1)] |
| 10. | Tan K, Wilkinson MF. A single-cell view of spermatogonial stem cells. Curr Opin Cell Biol. 2020;67:71-78. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 42] [Article Influence: 7.0] [Reference Citation Analysis (1)] |
| 11. | Rabbani M, Zheng X, Manske GL, Vargo A, Shami AN, Li JZ, Hammoud SS. Decoding the Spermatogenesis Program: New Insights from Transcriptomic Analyses. Annu Rev Genet. 2022;56:339-368. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 36] [Article Influence: 9.0] [Reference Citation Analysis (1)] |
| 12. | Chen Y, Zheng Y, Gao Y, Lin Z, Yang S, Wang T, Wang Q, Xie N, Hua R, Liu M, Sha J, Griswold MD, Li J, Tang F, Tong MH. Single-cell RNA-seq uncovers dynamic processes and critical regulators in mouse spermatogenesis. Cell Res. 2018;28:879-896. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 365] [Cited by in RCA: 306] [Article Influence: 38.3] [Reference Citation Analysis (1)] |
| 13. | Dong F, Ping P, Ma Y, Chen XF. Application of single-cell RNA sequencing on human testicular samples: a comprehensive review. Int J Biol Sci. 2023;19:2167-2197. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 28] [Article Influence: 9.3] [Reference Citation Analysis (1)] |
| 14. | Guo J, Grow EJ, Yi C, Mlcochova H, Maher GJ, Lindskog C, Murphy PJ, Wike CL, Carrell DT, Goriely A, Hotaling JM, Cairns BR. Chromatin and Single-Cell RNA-Seq Profiling Reveal Dynamic Signaling and Metabolic Transitions during Human Spermatogonial Stem Cell Development. Cell Stem Cell. 2017;21:533-546.e6. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 214] [Cited by in RCA: 211] [Article Influence: 23.4] [Reference Citation Analysis (1)] |
| 15. | La Manno G, Soldatov R, Zeisel A, Braun E, Hochgerner H, Petukhov V, Lidschreiber K, Kastriti ME, Lönnerberg P, Furlan A, Fan J, Borm LE, Liu Z, van Bruggen D, Guo J, He X, Barker R, Sundström E, Castelo-Branco G, Cramer P, Adameyko I, Linnarsson S, Kharchenko PV. RNA velocity of single cells. Nature. 2018;560:494-498. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2712] [Cited by in RCA: 2911] [Article Influence: 363.9] [Reference Citation Analysis (5)] |
| 16. | Teerds KJ, Huhtaniemi IT. Morphological and functional maturation of Leydig cells: from rodent models to primates. Hum Reprod Update. 2015;21:310-328. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 95] [Cited by in RCA: 119] [Article Influence: 10.8] [Reference Citation Analysis (1)] |
| 17. | Laurentino S, Heckmann L, Di Persio S, Li X, Meyer Zu Hörste G, Wistuba J, Cremers JF, Gromoll J, Kliesch S, Schlatt S, Neuhaus N. High-resolution analysis of germ cells from men with sex chromosomal aneuploidies reveals normal transcriptome but impaired imprinting. Clin Epigenetics. 2019;11:127. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 39] [Article Influence: 5.6] [Reference Citation Analysis (1)] |
| 18. | Li L, Dong J, Yan L, Yong J, Liu X, Hu Y, Fan X, Wu X, Guo H, Wang X, Zhu X, Li R, Yan J, Wei Y, Zhao Y, Wang W, Ren Y, Yuan P, Yan Z, Hu B, Guo F, Wen L, Tang F, Qiao J. Single-Cell RNA-Seq Analysis Maps Development of Human Germline Cells and Gonadal Niche Interactions. Cell Stem Cell. 2017;20:891-892. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70] [Cited by in RCA: 94] [Article Influence: 10.4] [Reference Citation Analysis (1)] |
| 19. | Tirumalasetty MB, Bhattacharya I, Mohiuddin MS, Baki VB, Choubey M. Understanding testicular single cell transcriptional atlas: from developmental complications to male infertility. Front Endocrinol (Lausanne). 2024;15:1394812. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 10] [Reference Citation Analysis (1)] |
| 20. | Jiang Z, Zhang J, Qiu Z, Zhang Y, Li N, Hu J, Zhu Z. Single-cell sequencing in non-obstructive azoospermia: insights from primary and re-analysis studies. Front Endocrinol (Lausanne). 2025;16:1539063. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 3] [Reference Citation Analysis (1)] |
| 21. | Xia K, Luo P, Yu J, He S, Dong L, Gao F, Chen X, Ye Y, Gao Y, Ma Y, Yang C, Zhang Y, Yang Q, Han D, Feng X, Wan Z, Cai H, Ke Q, Wang T, Li W, Tu X, Sun X, Deng C, Xiang AP. Single-cell RNA sequencing reveals transcriptomic landscape and potential targets for human testicular ageing. Hum Reprod. 2024;39:2189-2209. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 25] [Article Influence: 12.5] [Reference Citation Analysis (1)] |
| 22. | Fatumo S, Chikowore T, Choudhury A, Ayub M, Martin AR, Kuchenbaecker K. A roadmap to increase diversity in genomic studies. Nat Med. 2022;28:243-250. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 256] [Cited by in RCA: 459] [Article Influence: 114.8] [Reference Citation Analysis (0)] |
| 23. | Wang M, Liu X, Chang G, Chen Y, An G, Yan L, Gao S, Xu Y, Cui Y, Dong J, Chen Y, Fan X, Hu Y, Song K, Zhu X, Gao Y, Yao Z, Bian S, Hou Y, Lu J, Wang R, Fan Y, Lian Y, Tang W, Wang Y, Liu J, Zhao L, Wang L, Liu Z, Yuan R, Shi Y, Hu B, Ren X, Tang F, Zhao XY, Qiao J. Single-Cell RNA Sequencing Analysis Reveals Sequential Cell Fate Transition during Human Spermatogenesis. Cell Stem Cell. 2018;23:599-614.e4. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 369] [Cited by in RCA: 357] [Article Influence: 44.6] [Reference Citation Analysis (1)] |
| 24. | Hermann BP, Cheng K, Singh A, Roa-De La Cruz L, Mutoji KN, Chen IC, Gildersleeve H, Lehle JD, Mayo M, Westernströer B, Law NC, Oatley MJ, Velte EK, Niedenberger BA, Fritze D, Silber S, Geyer CB, Oatley JM, McCarrey JR. The Mammalian Spermatogenesis Single-Cell Transcriptome, from Spermatogonial Stem Cells to Spermatids. Cell Rep. 2018;25:1650-1667.e8. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 566] [Cited by in RCA: 449] [Article Influence: 56.1] [Reference Citation Analysis (1)] |
| 25. | Guo J, Grow EJ, Mlcochova H, Maher GJ, Lindskog C, Nie X, Guo Y, Takei Y, Yun J, Cai L, Kim R, Carrell DT, Goriely A, Hotaling JM, Cairns BR. The adult human testis transcriptional cell atlas. Cell Res. 2018;28:1141-1157. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 620] [Cited by in RCA: 501] [Article Influence: 62.6] [Reference Citation Analysis (1)] |
| 26. | Sohni A, Tan K, Song HW, Burow D, de Rooij DG, Laurent L, Hsieh TC, Rabah R, Hammoud SS, Vicini E, Wilkinson MF. The Neonatal and Adult Human Testis Defined at the Single-Cell Level. Cell Rep. 2019;26:1501-1517.e4. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 311] [Cited by in RCA: 256] [Article Influence: 36.6] [Reference Citation Analysis (1)] |
| 27. | Guo J, Nie X, Giebler M, Mlcochova H, Wang Y, Grow EJ; DonorConnect, Kim R, Tharmalingam M, Matilionyte G, Lindskog C, Carrell DT, Mitchell RT, Goriely A, Hotaling JM, Cairns BR. The Dynamic Transcriptional Cell Atlas of Testis Development during Human Puberty. Cell Stem Cell. 2020;26:262-276.e4. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 272] [Cited by in RCA: 225] [Article Influence: 37.5] [Reference Citation Analysis (1)] |
| 28. | Guo J, Sosa E, Chitiashvili T, Nie X, Rojas EJ, Oliver E; DonorConnect, Plath K, Hotaling JM, Stukenborg JB, Clark AT, Cairns BR. Single-cell analysis of the developing human testis reveals somatic niche cell specification and fetal germline stem cell establishment. Cell Stem Cell. 2021;28:764-778.e4. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 103] [Cited by in RCA: 160] [Article Influence: 32.0] [Reference Citation Analysis (1)] |
| 29. | Shami AN, Zheng X, Munyoki SK, Ma Q, Manske GL, Green CD, Sukhwani M, Orwig KE, Li JZ, Hammoud SS. Single-Cell RNA Sequencing of Human, Macaque, and Mouse Testes Uncovers Conserved and Divergent Features of Mammalian Spermatogenesis. Dev Cell. 2020;54:529-547.e12. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 213] [Article Influence: 35.5] [Reference Citation Analysis (4)] |
| 30. | Zhao L, Yao C, Xing X, Jing T, Li P, Zhu Z, Yang C, Zhai J, Tian R, Chen H, Luo J, Liu N, Deng Z, Lin X, Li N, Fang J, Sun J, Wang C, Zhou Z, Li Z. Single-cell analysis of developing and azoospermia human testicles reveals central role of Sertoli cells. Nat Commun. 2020;11:5683. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 39] [Cited by in RCA: 205] [Article Influence: 34.2] [Reference Citation Analysis (1)] |
| 31. | Di Persio S, Tekath T, Siebert-Kuss LM, Cremers JF, Wistuba J, Li X, Meyer Zu Hörste G, Drexler HCA, Wyrwoll MJ, Tüttelmann F, Dugas M, Kliesch S, Schlatt S, Laurentino S, Neuhaus N. Single-cell RNA-seq unravels alterations of the human spermatogonial stem cell compartment in patients with impaired spermatogenesis. Cell Rep Med. 2021;2:100395. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 62] [Article Influence: 12.4] [Reference Citation Analysis (0)] |
| 32. | Alfano M, Tascini AS, Pederzoli F, Locatelli I, Nebuloni M, Giannese F, Garcia-Manteiga JM, Tonon G, Amodio G, Gregori S, Agresti A, Montorsi F, Salonia A. Aging, inflammation and DNA damage in the somatic testicular niche with idiopathic germ cell aplasia. Nat Commun. 2021;12:5205. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 68] [Article Influence: 13.6] [Reference Citation Analysis (1)] |
| 33. | Chen H, Murray E, Sinha A, Laumas A, Li J, Lesman D, Nie X, Hotaling J, Guo J, Cairns BR, Macosko EZ, Cheng CY, Chen F. Dissecting mammalian spermatogenesis using spatial transcriptomics. Cell Rep. 2021;37:109915. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 96] [Article Influence: 19.2] [Reference Citation Analysis (1)] |
| 34. | Mahyari E, Guo J, Lima AC, Lewinsohn DP, Stendahl AM, Vigh-Conrad KA, Nie X, Nagirnaja L, Rockweiler NB, Carrell DT, Hotaling JM, Aston KI, Conrad DF. Comparative single-cell analysis of biopsies clarifies pathogenic mechanisms in Klinefelter syndrome. Am J Hum Genet. 2021;108:1924-1945. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 57] [Article Influence: 11.4] [Reference Citation Analysis (1)] |
| 35. | Nie X, Munyoki SK, Sukhwani M, Schmid N, Missel A, Emery BR, DonorConnect, Stukenborg JB, Mayerhofer A, Orwig KE, Aston KI, Hotaling JM, Cairns BR, Guo J. Single-cell analysis of human testis aging and correlation with elevated body mass index. Dev Cell. 2022;57:1160-1176.e5. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 152] [Article Influence: 38.0] [Reference Citation Analysis (1)] |
| 36. | Di Persio S, Neuhaus N. Human spermatogonial stem cells and their niche in male (in)fertility: novel concepts from single-cell RNA-sequencing. Hum Reprod. 2023;38:1-13. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 42] [Article Influence: 14.0] [Reference Citation Analysis (1)] |
| 37. | Amodio G, Giacomini G, Boeri L, Raffo M, Cilio S, Pozzi E, Belladelli F, Negri F, Ferrara AM, d'Arma A, Santoni de Sio FR, Pagliardini L, Papaleo E, Ventimiglia E, Alfano M, Montorsi F, Salonia A, Gregori S. Specific types of male infertility are correlated with T cell exhaustion or senescence signatures. Nat Commun. 2025;16:971. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 7] [Article Influence: 7.0] [Reference Citation Analysis (1)] |
| 38. | Cui L, Nie X, Guo Y, Ren P, Guo Y, Wang X, Li R, Hotaling JM, Cairns BR, Guo J. Single-cell transcriptomic atlas of the human testis across the reproductive lifespan. Nat Aging. 2025;5:658-674. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 26] [Article Influence: 26.0] [Reference Citation Analysis (1)] |
| 39. | Grant MJ, Booth A. A typology of reviews: an analysis of 14 review types and associated methodologies. Health Info Libr J. 2009;26:91-108. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3665] [Cited by in RCA: 3770] [Article Influence: 221.8] [Reference Citation Analysis (0)] |
| 40. | Büttner M, Miao Z, Wolf FA, Teichmann SA, Theis FJ. A test metric for assessing single-cell RNA-seq batch correction. Nat Methods. 2019;16:43-49. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 182] [Cited by in RCA: 316] [Article Influence: 39.5] [Reference Citation Analysis (2)] |
| 41. | Tran HTN, Ang KS, Chevrier M, Zhang X, Lee NYS, Goh M, Chen J. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 2020;21:12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 559] [Cited by in RCA: 733] [Article Influence: 122.2] [Reference Citation Analysis (5)] |
| 42. | Luecken MD, Theis FJ. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol Syst Biol. 2019;15:e8746. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1396] [Cited by in RCA: 1399] [Article Influence: 199.9] [Reference Citation Analysis (5)] |
| 43. | Hwang B, Lee JH, Bang D. Single-cell RNA sequencing technologies and bioinformatics pipelines. Exp Mol Med. 2018;50:1-14. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1432] [Cited by in RCA: 1161] [Article Influence: 145.1] [Reference Citation Analysis (7)] |
| 44. | Hafemeister C, Satija R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 2019;20:296. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2049] [Cited by in RCA: 3157] [Article Influence: 451.0] [Reference Citation Analysis (6)] |
| 45. | Risso D, Perraudeau F, Gribkova S, Dudoit S, Vert JP. A general and flexible method for signal extraction from single-cell RNA-seq data. Nat Commun. 2018;9:284. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 363] [Cited by in RCA: 441] [Article Influence: 55.1] [Reference Citation Analysis (1)] |
| 46. | Saelens W, Cannoodt R, Todorov H, Saeys Y. A comparison of single-cell trajectory inference methods. Nat Biotechnol. 2019;37:547-554. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 664] [Cited by in RCA: 1054] [Article Influence: 150.6] [Reference Citation Analysis (4)] |
| 47. | Street K, Risso D, Fletcher RB, Das D, Ngai J, Yosef N, Purdom E, Dudoit S. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genomics. 2018;19:477. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2852] [Cited by in RCA: 2106] [Article Influence: 263.3] [Reference Citation Analysis (5)] |
| 48. | Jain S, Eadon MT. Spatial transcriptomics in health and disease. Nat Rev Nephrol. 2024;20:659-671. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 75] [Article Influence: 37.5] [Reference Citation Analysis (1)] |
| 49. | Rajachandran S, Zhang X, Cao Q, Caldeira-Brant AL, Zhang X, Song Y, Evans M, Bukulmez O, Grow EJ, Nagano M, Orwig KE, Chen H. Dissecting the spermatogonial stem cell niche using spatial transcriptomics. Cell Rep. 2023;42:112737. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 31] [Article Influence: 10.3] [Reference Citation Analysis (1)] |
| 50. | Elosua-Bayes M, Nieto P, Mereu E, Gut I, Heyn H. SPOTlight: seeded NMF regression to deconvolute spatial transcriptomics spots with single-cell transcriptomes. Nucleic Acids Res. 2021;49:e50. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 279] [Cited by in RCA: 485] [Article Influence: 97.0] [Reference Citation Analysis (1)] |
| 51. | Cable DM, Murray E, Zou LS, Goeva A, Macosko EZ, Chen F, Irizarry RA. Robust decomposition of cell type mixtures in spatial transcriptomics. Nat Biotechnol. 2022;40:517-526. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 146] [Cited by in RCA: 898] [Article Influence: 224.5] [Reference Citation Analysis (7)] |
| 52. | Lim HJ, Wang Y, Buzdin A, Li X. A practical guide for choosing an optimal spatial transcriptomics technology from seven major commercially available options. BMC Genomics. 2025;26:47. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 46] [Article Influence: 46.0] [Reference Citation Analysis (1)] |
| 53. | Zheng W, Zhang S, Jiang S, Huang Z, Chen X, Guo H, Li M, Zheng S. Evaluation of immune status in testis and macrophage polarization associated with testicular damage in patients with nonobstructive azoospermia. Am J Reprod Immunol. 2021;86:e13481. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 29] [Article Influence: 5.8] [Reference Citation Analysis (1)] |
| 54. | Yang K, Xue J, Feng C, Li W. Testicular macrophages: Sculpting male fertility through immune regulation and tissue interactions. J Reprod Immunol. 2025;170:104543. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 6] [Reference Citation Analysis (1)] |
| 55. | Xia P, Ouyang S, Shen R, Guo Z, Zhang G, Liu X, Yang X, Xie K, Wang D. Macrophage-Related Testicular Inflammation in Individuals with Idiopathic Non-Obstructive Azoospermia: A Single-Cell Analysis. Int J Mol Sci. 2023;24:8819. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 20] [Reference Citation Analysis (1)] |
| 56. | Matzkin ME, Calandra RS, Rossi SP, Bartke A, Frungieri MB. Hallmarks of Testicular Aging: The Challenge of Anti-Inflammatory and Antioxidant Therapies Using Natural and/or Pharmacological Compounds to Improve the Physiopathological Status of the Aged Male Gonad. Cells. 2021;10:3114. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 55] [Article Influence: 11.0] [Reference Citation Analysis (8)] |
| 57. | Dong S, Chen C, Zhang J, Gao Y, Zeng X, Zhang X. Testicular aging, male fertility and beyond. Front Endocrinol (Lausanne). 2022;13:1012119. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 80] [Article Influence: 20.0] [Reference Citation Analysis (1)] |
| 58. | Zhan MW, Zhou MH, Zhao BB, Bao XJ, Chen Y, Zhu J. Testicular immunosenescence: a key player in age-related spermatogenic decline. Front Cell Dev Biol. 2025;13:1669826. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 10] [Reference Citation Analysis (1)] |
| 59. | Dutta S, Sengupta P, Slama P, Roychoudhury S. Oxidative Stress, Testicular Inflammatory Pathways, and Male Reproduction. Int J Mol Sci. 2021;22:10043. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 246] [Article Influence: 49.2] [Reference Citation Analysis (1)] |
| 60. | Mirghanizadeh Bafghi SA, Fesahat F, Zare F, Imani M, Vahidi S, Ansariniya H, ZareHoroki A, Hadinedoushan H. The role of inflammasome dysregulation in obstructive and non-obstructive azoospermia: a comparative molecular analysis of blood, tissue, and seminal plasma. Front Immunol. 2024;15:1507885. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (1)] |
| 61. | Kelley N, Jeltema D, Duan Y, He Y. The NLRP3 Inflammasome: An Overview of Mechanisms of Activation and Regulation. Int J Mol Sci. 2019;20:3328. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3034] [Cited by in RCA: 2753] [Article Influence: 393.3] [Reference Citation Analysis (5)] |
| 62. | Drevet JR, Hallak J, Nasr-Esfahani MH, Aitken RJ. Reactive Oxygen Species and Their Consequences on the Structure and Function of Mammalian Spermatozoa. Antioxid Redox Signal. 2022;37:481-500. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 26] [Article Influence: 6.5] [Reference Citation Analysis (1)] |
| 63. | Meng K, Liu Q, Qin Y, Qin W, Zhu Z, Sun L, Jiang M, Adu-Amankwaah J, Gao F, Tan R, Yuan J. Mechanism of mitochondrial oxidative phosphorylation disorder in male infertility. Chin Med J (Engl). 2025;138:379-388. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 18] [Article Influence: 18.0] [Reference Citation Analysis (1)] |
| 64. | Fomichova O, Oliveira PF, Bernardino RL. Exploring the interplay between inflammation and male fertility. FEBS J. 2025;292:3321-3349. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 21] [Article Influence: 21.0] [Reference Citation Analysis (1)] |
| 65. | Carrageta DF, Pereira SC, Ferreira R, Monteiro MP, Oliveira PF, Alves MG. Signatures of metabolic diseases on spermatogenesis and testicular metabolism. Nat Rev Urol. 2024;21:477-494. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 33] [Article Influence: 16.5] [Reference Citation Analysis (1)] |
| 66. | Chen Q, Yan M, Cao Z, Li X, Zhang Y, Shi J, Feng GH, Peng H, Zhang X, Zhang Y, Qian J, Duan E, Zhai Q, Zhou Q. Sperm tsRNAs contribute to intergenerational inheritance of an acquired metabolic disorder. Science. 2016;351:397-400. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 804] [Cited by in RCA: 1046] [Article Influence: 95.1] [Reference Citation Analysis (4)] |
| 67. | Sharma U, Conine CC, Shea JM, Boskovic A, Derr AG, Bing XY, Belleannee C, Kucukural A, Serra RW, Sun F, Song L, Carone BR, Ricci EP, Li XZ, Fauquier L, Moore MJ, Sullivan R, Mello CC, Garber M, Rando OJ. Biogenesis and function of tRNA fragments during sperm maturation and fertilization in mammals. Science. 2016;351:391-396. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1132] [Cited by in RCA: 929] [Article Influence: 92.9] [Reference Citation Analysis (5)] |
| 68. | Zhang Y, Zhang X, Shi J, Tuorto F, Li X, Liu Y, Liebers R, Zhang L, Qu Y, Qian J, Pahima M, Liu Y, Yan M, Cao Z, Lei X, Cao Y, Peng H, Liu S, Wang Y, Zheng H, Woolsey R, Quilici D, Zhai Q, Li L, Zhou T, Yan W, Lyko F, Zhang Y, Zhou Q, Duan E, Chen Q. Dnmt2 mediates intergenerational transmission of paternally acquired metabolic disorders through sperm small non-coding RNAs. Nat Cell Biol. 2018;20:535-540. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 302] [Cited by in RCA: 358] [Article Influence: 44.8] [Reference Citation Analysis (0)] |
| 69. | Nätt D, Kugelberg U, Casas E, Nedstrand E, Zalavary S, Henriksson P, Nijm C, Jäderquist J, Sandborg J, Flinke E, Ramesh R, Örkenby L, Appelkvist F, Lingg T, Guzzi N, Bellodi C, Löf M, Vavouri T, Öst A. Human sperm displays rapid responses to diet. PLoS Biol. 2019;17:e3000559. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 107] [Cited by in RCA: 145] [Article Influence: 20.7] [Reference Citation Analysis (1)] |
| 70. | Krausz C, Riera-Escamilla A. Genetics of male infertility. Nat Rev Urol. 2018;15:369-384. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 294] [Cited by in RCA: 607] [Article Influence: 86.7] [Reference Citation Analysis (1)] |
| 71. | Marchetti C, Swartzwelter B, Gamboni F, Neff CP, Richter K, Azam T, Carta S, Tengesdal I, Nemkov T, D'Alessandro A, Henry C, Jones GS, Goodrich SA, St Laurent JP, Jones TM, Scribner CL, Barrow RB, Altman RD, Skouras DB, Gattorno M, Grau V, Janciauskiene S, Rubartelli A, Joosten LAB, Dinarello CA. OLT1177, a β-sulfonyl nitrile compound, safe in humans, inhibits the NLRP3 inflammasome and reverses the metabolic cost of inflammation. Proc Natl Acad Sci U S A. 2018;115:E1530-E1539. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 516] [Cited by in RCA: 487] [Article Influence: 60.9] [Reference Citation Analysis (1)] |
| 72. | Klück V, Jansen TLTA, Janssen M, Comarniceanu A, Efdé M, Tengesdal IW, Schraa K, Cleophas MCP, Scribner CL, Skouras DB, Marchetti C, Dinarello CA, Joosten LAB. Dapansutrile, an oral selective NLRP3 inflammasome inhibitor, for treatment of gout flares: an open-label, dose-adaptive, proof-of-concept, phase 2a trial. Lancet Rheumatol. 2020;2:e270-e280. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 150] [Cited by in RCA: 229] [Article Influence: 38.2] [Reference Citation Analysis (1)] |
| 73. | Wohlford GF, Van Tassell BW, Billingsley HE, Kadariya D, Canada JM, Carbone S, Mihalick VL, Bonaventura A, Vecchié A, Chiabrando JG, Bressi E, Thomas G, Ho AC, Marawan AA, Dell M, Trankle CR, Turlington J, Markley R, Abbate A. Phase 1B, Randomized, Double-Blinded, Dose Escalation, Single-Center, Repeat Dose Safety and Pharmacodynamics Study of the Oral NLRP3 Inhibitor Dapansutrile in Subjects With NYHA II-III Systolic Heart Failure. J Cardiovasc Pharmacol. 2020;77:49-60. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 51] [Cited by in RCA: 132] [Article Influence: 22.0] [Reference Citation Analysis (1)] |
| 74. | Wang HQ, Wang T, Gao F, Ren WZ. Application of CRISPR/Cas Technology in Spermatogenesis Research and Male Infertility Treatment. Genes (Basel). 2022;13:1000. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 14] [Reference Citation Analysis (1)] |
| 75. | Kumar N. Genome Editing for Fertility: Unlocking the Promise of CRISPR/Cas9 in Addressing Male Infertility - A Narrative Review. Reprod Sci. 2025;32:3221-3239. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 76. | Mulder CL, Zheng Y, Jan SZ, Struijk RB, Repping S, Hamer G, van Pelt AM. Spermatogonial stem cell autotransplantation and germline genomic editing: a future cure for spermatogenic failure and prevention of transmission of genomic diseases. Hum Reprod Update. 2016;22:561-573. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 44] [Cited by in RCA: 50] [Article Influence: 5.0] [Reference Citation Analysis (1)] |
| 77. | Fang F, Li Z, Zhao Q, Li H, Xiong C. Human induced pluripotent stem cells and male infertility: an overview of current progress and perspectives. Hum Reprod. 2018;33:188-195. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 41] [Cited by in RCA: 48] [Article Influence: 6.9] [Reference Citation Analysis (1)] |
| 78. | Schleidgen S, Dederer HG, Sgodda S, Cravcisin S, Lüneburg L, Cantz T, Heinemann T. Human germline editing in the era of CRISPR-Cas: risk and uncertainty, inter-generational responsibility, therapeutic legitimacy. BMC Med Ethics. 2020;21:87. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 19] [Article Influence: 3.2] [Reference Citation Analysis (1)] |
| 79. | Wiley L, Cheek M, LaFar E, Ma X, Sekowski J, Tanguturi N, Iltis A. The Ethics of Human Embryo Editing via CRISPR-Cas9 Technology: A Systematic Review of Ethical Arguments, Reasons, and Concerns. HEC Forum. 2025;37:267-303. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 20] [Reference Citation Analysis (1)] |
| 80. | Knoppers BM, Kleiderman E. "CRISPR babies": What does this mean for science and Canada? CMAJ. 2019;191:E91-E92. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 9] [Article Influence: 1.3] [Reference Citation Analysis (1)] |
| 81. | Mitchell MJ, Billingsley MM, Haley RM, Wechsler ME, Peppas NA, Langer R. Engineering precision nanoparticles for drug delivery. Nat Rev Drug Discov. 2021;20:101-124. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6637] [Cited by in RCA: 4750] [Article Influence: 950.0] [Reference Citation Analysis (5)] |
| 82. | Iftikhar M, Noureen A, Uzair M, Jabeen F, Abdel Daim M, Cappello T. Perspectives of Nanoparticles in Male Infertility: Evidence for Induced Abnormalities in Sperm Production. Int J Environ Res Public Health. 2021;18:1758. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 47] [Cited by in RCA: 40] [Article Influence: 8.0] [Reference Citation Analysis (1)] |
| 83. | Baltimore D, Berg P, Botchan M, Carroll D, Charo RA, Church G, Corn JE, Daley GQ, Doudna JA, Fenner M, Greely HT, Jinek M, Martin GS, Penhoet E, Puck J, Sternberg SH, Weissman JS, Yamamoto KR. Biotechnology. A prudent path forward for genomic engineering and germline gene modification. Science. 2015;348:36-38. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 402] [Cited by in RCA: 311] [Article Influence: 28.3] [Reference Citation Analysis (1)] |
| 84. | Lander ES, Baylis F, Zhang F, Charpentier E, Berg P, Bourgain C, Friedrich B, Joung JK, Li J, Liu D, Naldini L, Nie JB, Qiu R, Schoene-Seifert B, Shao F, Terry S, Wei W, Winnacker EL. Adopt a moratorium on heritable genome editing. Nature. 2019;567:165-168. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 202] [Cited by in RCA: 198] [Article Influence: 28.3] [Reference Citation Analysis (1)] |
| 85. | Hindorff LA, Bonham VL, Brody LC, Ginoza MEC, Hutter CM, Manolio TA, Green ED. Prioritizing diversity in human genomics research. Nat Rev Genet. 2018;19:175-185. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 253] [Cited by in RCA: 292] [Article Influence: 36.5] [Reference Citation Analysis (1)] |
| 86. | Sirugo G, Williams SM, Tishkoff SA. The Missing Diversity in Human Genetic Studies. Cell. 2019;177:26-31. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 448] [Cited by in RCA: 952] [Article Influence: 158.7] [Reference Citation Analysis (1)] |
| 87. | Mills MC, Rahal C. A scientometric review of genome-wide association studies. Commun Biol. 2019;2:9. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 270] [Cited by in RCA: 274] [Article Influence: 39.1] [Reference Citation Analysis (1)] |
| 88. | Martin AR, Kanai M, Kamatani Y, Okada Y, Neale BM, Daly MJ. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat Genet. 2019;51:584-591. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2493] [Cited by in RCA: 1991] [Article Influence: 284.4] [Reference Citation Analysis (1)] |
| 89. | Wouters OJ, McKee M, Luyten J. Estimated Research and Development Investment Needed to Bring a New Medicine to Market, 2009-2018. JAMA. 2020;323:844-853. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1274] [Cited by in RCA: 890] [Article Influence: 148.3] [Reference Citation Analysis (1)] |
| 90. | Rajkomar A, Hardt M, Howell MD, Corrado G, Chin MH. Ensuring Fairness in Machine Learning to Advance Health Equity. Ann Intern Med. 2018;169:866-872. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 366] [Cited by in RCA: 595] [Article Influence: 74.4] [Reference Citation Analysis (5)] |
| 91. | Kerasidou A. The role of trust in global health research collaborations. Bioethics. 2019;33:495-501. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 25] [Article Influence: 3.6] [Reference Citation Analysis (1)] |