Published online May 7, 2026. doi: 10.3748/wjg.v32.i17.116590
Revised: January 7, 2026
Accepted: February 12, 2026
Published online: May 7, 2026
Processing time: 160 Days and 19 Hours
Ulcerative colitis (UC) is a chronic inflammatory bowel disease for which effective therapies are lacking. Niu Huang (NH) is a traditional Chinese medicine used for inflammatory disorders. However, its protective effect on UC and its underlying mechanisms are unknown.
To uncover the mechanisms underlying the anti-colitis effects of the NH.
Network pharmacology was applied to predict the active ingredients and targets of NH. Experimental validation was conducted in a dextran sulfate sodium-in
Network pharmacology revealed that the bioactive component of NH is bile acid. Our animal experiments de
This study demonstrates the therapeutic effects of NH on UC. Mechanistically, NH acts by activating FXR, which subsequently inhibits the nuclear factor kappa B pathway to reduce C3 accumulation and suppress excessive NLRP3 inflammasome activation in colon tissue.
Core Tip: This study demonstrates that Niu Huang effectively alleviates ulcerative colitis. By identifying its active com
- Citation: Shi J, Ma CY, Zhang XH, Liu KJ, Liu JY, Wang QG, Wang XQ, Cheng FF, Xu T. Niu Huang mitigates dextran sulfate sodium-induced colitis by modulating farnesoid X receptor activation and the complement 3/NLRP3 signaling pathway. World J Gastroenterol 2026; 32(17): 116590
- URL: https://www.wjgnet.com/1007-9327/full/v32/i17/116590.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i17.116590
Ulcerative colitis (UC) is a severe chronic inflammatory bowel disease (IBD) that is often characterized by alternating periods of exacerbation and remission and presents with abdominal pain, diarrhea, and the presence of bloody mucus in stools. It is a lifelong, incurable disorder that substantially impairs quality of life and is increasing in incidence worldwide[1]. Currently, the primary therapeutic choices for UC are mesalamine, corticosteroids, immunosuppressants, biologics, small molecules, and surgery[2]. Although these therapies provide partial symptomatic relief, they are often accompanied by high relapse rates and substantial adverse events. Consequently, identifying optimal therapeutic strategies for UC remains imperative.
The pathogenesis of IBD is centered on immune dysregulation[3]. The key mediators that drive the initiation and perpetuation of chronic intestinal inflammation are immune cells, inflammatory cytokines, the complement system and inflammasomes. The complement system has long been recognized as a crucial element of the innate immune response, and it facilitates both mucosal defense and clearance of debris. However, uncontrolled and sustained complement system activation induces severe inflammatory processes and may result in tissue damage[4]. Many studies have shown that the complement system is involved in intestinal inflammation and may be related to IBD[5]. Research has shown that complement component 3 (C3) is expressed in intestinal tissues, is secreted into the lumen, and is upregulated in the presence of inflammation[6], playing a crucial role in colitis progression and therapeutic response[7]. Clinical data have indicated that significantly higher concentrations of C3 are associated with severe or extensive UC[8]. Complement inhibition reduces inflammatory cytokines, leukocyte infiltration, and tissue damage while promoting wound healing and mucosal repair[9]. In particular, inhibition of the C3 signaling pathway suppresses the level of proinflammatory cytokines and contributes to antioxidant effects[10]. Recent studies have demonstrated that complement activation potentiates NOD-like receptor family pyrin domain containing 3 (NLRP3) inflammasome, thereby enhancing the secretion of the proinflammatory cytokines interleukin (IL)-1β and IL-18[11]. Serving as the primary intracellular receptor for bile acids, the farnesoid X receptor (FXR) regulates the expression of C3[12]. These studies indicate that the comp
Niu Huang (NH, Calculus bovis), a traditional animal drug derived from the dried gallstones of cattle or buffaloes, has been widely used in traditional Chinese medicine for the treatment of cerebrovascular diseases, hepatobiliary diseases, wound healing and various inflammatory diseases[13,14]. The Compendium of Materia Medica documents that NH was traditionally used for treating severe diarrhea with pus and blood. Recent pharmacological studies have demonstrated that NH has anti-inflammatory, antipyretic, sedative, and hepatobiliary-restorative effects[15]. NH has been reported to mitigate gut injury in rats by regulating bile acid profiles, inflammation, oxidative stress, and apoptosis[16]. Previous studies indicated that NH ameliorates dextran sulfate sodium (DSS)-induced colitis, attenuating key pathological features such as colonic edema, mucosal damage, and inflammatory cell infiltration[17]. However, the mechanism through which NH protects against colitis remains unknown.
To elucidate the mechanisms by which NH exerts its effects against colitis, we initially employed a network pharmacology approach to investigate its systemic mechanisms. After predicting the main biological function of NH against colitis, we used a DSS-induced colitis model to explore the pharmacological efficacy and dose-dependent effects of NH. We also used a multiple-omics approach, consisting of transcriptomics and proteomics, to study the underlying mechanisms. After experimental validation, we observed that NH significantly activated FXR signaling and therefore inhibited the expression of nuclear factor kappa B (NF-κB) and its target genes C3 and NLRP3, thereby contributing to the regulation of the dysregulated complement system and inflammation.
We utilized the HERB database (http://herb.ac.cn)[18] to acquire information on the active components of NH (ID: HERB004201). Given that NH is typically administered via the oral route, we applied Lipinski’s rules to evaluate the drug-like properties of NH’s chemical components and to identify potential active ingredients based on these criteria. Lipinski’s rules stipulate the following criteria for screening: Molecular weight ≤ 500, number of hydrogen bond donors ≤ 5, number of hydrogen bond acceptors ≤ 10, and the calculated lipophilicity ≤ 5. The chemical structures of all constituents were retrieved, and their corresponding SMILES identifiers were acquired, from the PubChem database (https://pubchem.ncbi.nlm.nih.gov). SMILES files were imported into the SwissTargetPrediction database (http://www.swisstargetprediction.ch) to obtain the drug targets.
The keyword “ulcerative colitis” was used to retrieve disease-related targets from the GeneCards (https://www.genecards.org/), Therapeutic Target Database (https://db.idrblab.net/ttd/), Online Mendelian Inheritance in Man (https://omim.org/), and Drugbank databases (https://go.drugbank.com/). To identify targets relevant to UC, those collected from individual databases were combined and screened for duplicates. Venn diagrams were constructed using the online tool Venny 2.1 to visualize common targets obtained by intersecting NH component targets with UC disease targets.
The potential targets of the active ingredients in NH were intersected with the disease targets of UC. These intersecting genes were then imported into the STRING database (https://string-db.org/) to generate protein-protein interaction (PPI) networks. The resulting gene network was imported into Cytoscape 3.10.2 for further analysis of the interaction network. An ingredient–target network was constructed in Cytoscape 3.10.2 using the active ingredients of NH and their corresponding targets.
Enrichment analysis of the overlapping genes was conducted via the Metascape platform (https://metascape.org/) for both Gene Ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways, with statistical significance set at P < 0.05. GO enrichment covered the three main domains: Biological processes (BP), cellular components (CC), and molecular functions (MF).
An ultra-high performance liquid chromatography coupled to tandem mass spectrometry (LC-MS/MS) system was used to quantitate bile acids. Separation was performed on a Waters ACQUITY UPLC BEH C18 column (2.1 mm × 100 mm, 1.7 μm).
Six-week-old specific pathogen-free male C57BL/6J mice were purchased from Beijing Sipeifu Laboratory Animal Technology Co., Ltd., Beijing, China. The Fxr-/- mice were constructed using CRISPR/Cas9 gene editing technology on a C57BL/6 background, with the assistance of Cyagen Biotechnology. The animals were raised under pathogen-free conditions (ambient temperature of 22 ± 1 °C; relatively constant humidity of 50% ± 15%; 12-hour light/dark cycle). Water and food were freely available. The study was approved by the Ethics Committee of Capital Medical University, approval No. AEEI-2024-007.
Prior to induction with DSS, all the mice were acclimatized for one week. Fifty mice were divided into five groups according to the random number table method: The control group, the model group (DSS), the low-dose NH group, the high-dose NH group (NHH) and the mesalamine group (Mes). The control group was provided with standard drinking water, while the other experimental groups were given 2.5% DSS (36000-50000 molecular weight; CAS: 216011080; MP Biomedicals) in their drinking water for five days to establish a DSS-induced UC model. Thereafter, all groups were transitioned back to standard drinking water for a subsequent two-day period. During the experimental period (day 0 to day 7), both the control and model groups received a daily intragastric administration of 0.5% CMC-Na solution (IS9000; Solarbio). The low-dose NH group received a gavage of 50 mg/kg body weight of NH (CULTURAL CALCULUS BOVIS; P230601; ShangYaoJu Pharmaceutical Co., Ltd). Previous studies have shown that NH exerts therapeutic effects at this dosage level[17,19]. The NHH group received a gavage of 100 mg/kg body weight NH solution. The Mes group received a gavage of 50 mg/kg body weight mesalamine (HY-15027, MCE). Both NH and Mes were dissolved in 0.5% CMC-Na solution. The body weight of each mouse, the consistency of their feces, and the presence of occult blood in their stool samples were monitored on a daily basis. The disease activity index (DAI) was determined per animal to evaluate disease severity[20].
To examine the role of the FXR in the protective effects of NH against DSS-induced colitis, mice were categorized into two groups: Wild-type mice + DSS and Fxr knockout mice + DSS. Each group was administered 2.5% DSS solution. Subsequently, the mice were sacrificed as described previously.
In the inhibitor experiment, we randomly divided 20 mice into two groups: DSS + guggulsterone (Gug) (HY-107738, MCE) and DSS + Gug + NH. Mice in the DSS + Gug+ NH group received the NH (100 mg/kg) by oral gavage daily for seven consecutive days. Each group had free access to a 2.5% DSS solution in their drinking water. They were also administered Gug (100 mg/kg) by oral gavage daily.
The colon tissues were fixed in 4% paraformaldehyde solution. After fixation, dehydration and paraffin embedding, the sections were stained with hematoxylin and eosin.
Total RNA was prepared from colon tissues using the HiPure Total RNA Plus Kit (R4111, Magen), after which the concentration of RNA was determined by a Nanodrop 2000 spectrometer. cDNA was synthesized using a ReverAid First Standard cDNA Synthesis Kit (K1622; Thermo Fisher Scientific). Quantitative polymerase chain reaction was performed using GoTaq® quantitative polymerase chain reaction Master Mix (A6002; Promega) and a CFX Connect Real-Time system. β-actin was selected as an endogenous control. The relative mRNA expression was determined by the 2-ΔΔCt method. Primer sequences are provided in Table 1.
| Primer | Forward primer 5’-3’ | Reverse primer 5’-3’ |
| Nr0b2 | GCCAAGACAGTAGCCTTCCTCAG | GTACCGCTGCTGGCTTCCTC |
| Fgf15 | CTCTGTGGCTGGCTGTGTCTG | ATTCGGAGGAAGCAGTTGGAGAC |
| Il1b | GGGCTGCTTCCAAACCTTTG | AAGACACAGGTAGCTGCCAC |
| Il18 | ACTTTGGCCGACTTCACTGT | CCTCGAACACAGGCTGTCTT |
| Tnf | GCCTCTTCTCATTCCTGCTTGTGG | GTGGTTTGTGAGTGTGAGGGTCTG |
| Il6 | GAGAGGAGACTTCACAGAGGATACC | TCATTTCCACGATTTCCCAGAGAAC |
| Rela | AGGCTTCTGGGCCTTATGTG | TGCTTCTCTCGCCAGGAATAC |
| β-actin | GTCACGCACGATTTCCCTCTCAG | TATGCTCTCCCTCACGCCATCC |
Proteins were extracted from colon tissue using RIPA lysis buffer (PC101; Epizyme Biotech). The protein concentration was subsequently quantified using a bicinchoninic acid protein assay kit (ZJ102; Epizyme Biotech). The proteins were separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (PG212; Epizyme Biotech) and then transferred onto a polyvinylidene difluoride membrane. The membranes were blocked with 5% skim milk for 2 hours. Afterward, the membranes were incubated with primary antibodies at 4 °C overnight. The primary antibodies used included anti-E-cadherin (1:1000, 3195S, CST), anti-occludin (1:1000, ab216327, Abcam), anti-claudin3 (1:1000, ab317319, Abcam), anti-leucine rich repeat containing G protein-coupled receptor 5 (Lgr5) (1:1000, ab75850, Abcam), anti-FXR (1:1000, 417200, Thermo Fisher Scientific), anti-C3 (1:1000, 21337-1-AP, Proteintech), anti-NF-κB p65 (1:1000, 8242, CST), anti-p-NF-κB p65 (1:1000, SC-136548, Santa), anti-NLRP3 (1:1000, ab263899, Abcam), anti-apoptosis-associated speck-like protein containing a CARD (1:1000, ab283684, Abcam), anti-β-tubulin (1:5000, ab179513, Abcam), anti-β-actin (1:10000, 20536-1-AP, Proteintech), and anti-glyceraldehyde 3-phosphate dehydrogenase (1:5000, 10494-1-AP, Proteintech). Afterward, the membranes were incubated with the secondary antibody for 2 hours at room temperature. Finally, images of the protein bands were captured using an Amersham ImageQuant 800 imaging system (Cytiva) and quantified using ImageJ software.
The levels of IL-1β (MLB00C, Bio-Techne), IL-6 (M6000B, Bio-Techne), and tumor necrosis factor (TNF)-α (MTA00B, Bio-Techne) in the colonic tissue were measured using enzyme-linked immunosorbent assay kits according to the manufacturer's instructions. The concentrations of these cytokines in colon tissue were normalized using the bicinchoninic acid kit (P0012, Beyotime).
The colon tissue sections underwent antigen repair and were blocked with 5% goat serum. The sections were then incubated overnight at 4 °C with anti-CD11b (1:500, ab133357, Abcam), anti-F4/80 (1:500, ab111101, Abcam), and anti-NF-κB p65 (1:500, ab16502, Abcam). After incubation, secondary antibodies were applied and the sections were stained with 3,3’-diaminobenzidine. Finally, the sections were dehydrated, clarified, and mounted with neutral resin.
Total RNA was extracted from colon tissues using TRIzol® Reagent (Invitrogen, Carlsbad, CA, United States) and qua
A proteomics analysis was conducted using data-independent acquisition quantitative proteomics technology. Following protein extraction, the protein concentration was determined using a protein quantification kit (Bradford). The samples underwent proteolysis and were subsequently analyzed by LC-MS/MS analysis in data-independent acquisition mode. The proteins were identified, quantified, and functionally annotated.
All data were analyzed using GraphPad Prism 10.0 and are presented as mean ± SD. Data distribution was first assessed for normality and homogeneity of variance. For comparisons between two groups, an unpaired Student’s t-test was used. For comparisons among three or more groups, one-way analysis of variance was performed, followed by Dunnett’s post hoc test. A P value of less than 0.05 was considered statistically significant.
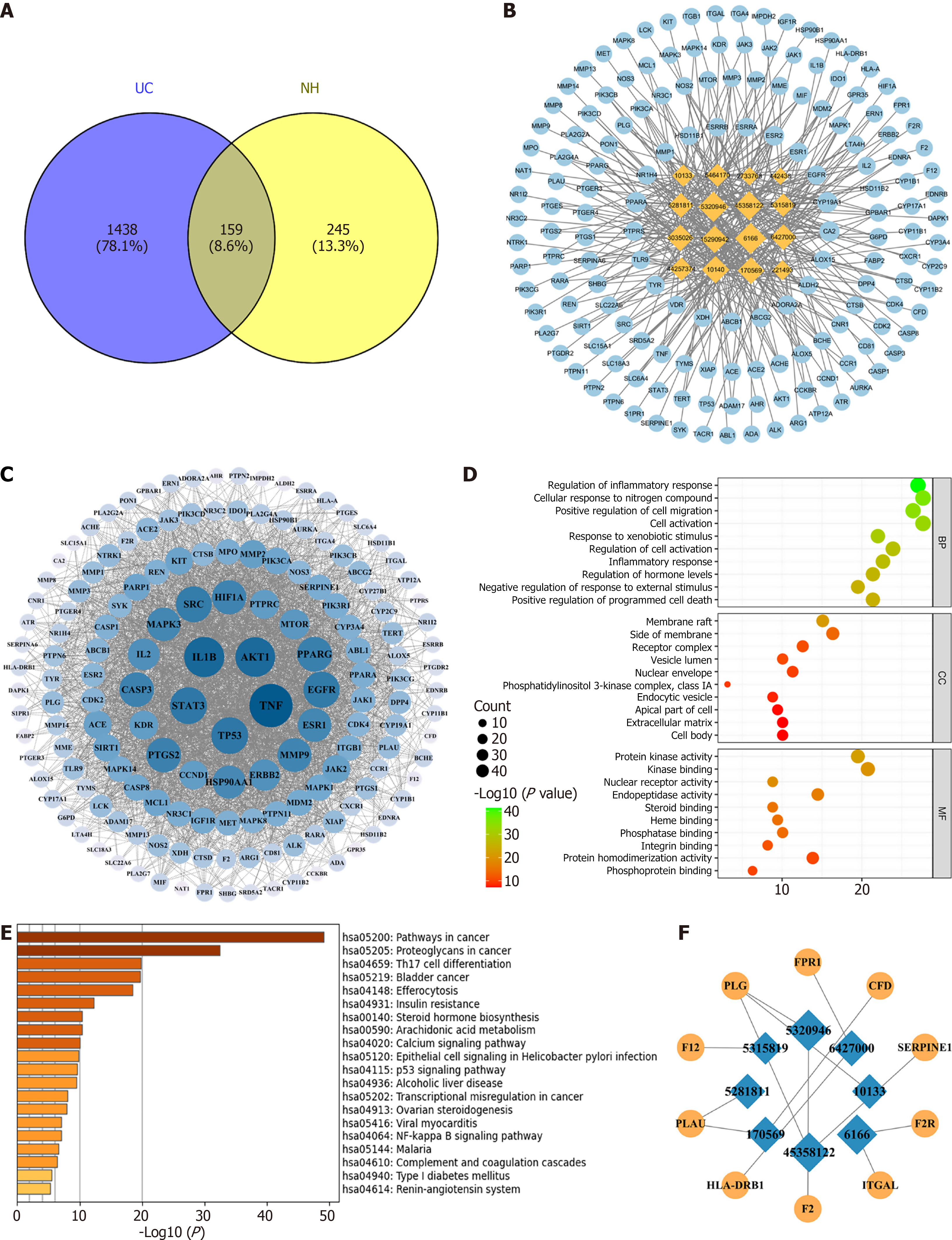
In total, 16 active ingredients of NH were identified through the HERB database and subsequent Linpinski rule screening. According to ultra-high performance LC-MS/MS analyses, cholic acid, glycocholic acid, glycodeoxycholic acid, and taurodeoxycholic acid were identified in NH (Supplementary Figure 1). After removing duplicates, 404 unique targets were identified for NH. We retrieved UC-related target genes from the GeneCards, DrugBank, Online Mendelian Inheritance in Man and Therapeutic Target Database. After merging the data and removing duplicates, we identified 1597 targets related to UC. By intersecting the potential targets of NH with these UC-related targets, we subsequently identified 159 common targets (Figure 1A and B).
We input the 159 NH-UC intersecting targets into the STRING database to construct a PPI network. The data were subsequently imported into Cytoscape 3.10.2 for visualization, generating a new PPI network with 159 nodes and 2656 edges. Network analysis was performed using the Network Analyzer plugin. The size of each node was proportional to its connectivity, with a greater number of connections indicating stronger correlation intensity. Notably, TNF, IL-1β, serine/threonine kinase 1, signal transducer and activator of transcription 3 and TP53 emerged as key nodes within the network (Figure 1C).
The 159 intersecting genes were input into the Metascape database for GO functional enrichment and KEGG pathway enrichment analyses. The enrichment analysis results revealed 1871 BP, 109 CC, 229 MF, and 191 KEGG pathways. The top 10 terms according to the GO functional enrichment results and the top 20 terms according to the KEGG pathway enrichment results were visualized (Figure 1D and E). The biological process-related targets were closely related to the regulation of inflammatory response, positive regulation of cell migration, cell activation, and positive regulation of programmed cell death. With respect to CC, NH primarily affects the receptor complex, nuclear envelope, extracellular matrix and cell body. At the molecular level, the functions of NH were associated mainly with protein kinase activity, nuclear receptor activity, steroid binding and phosphoprotein binding. KEGG pathway analysis revealed that these genes were involved mainly in key pathways, including Th17 cell differentiation, the p53 signaling pathway, the NF-κB signaling pathway and complement and coagulation cascades. The active components and targets related to the complement and coagulation cascades were visualized (Figure 1F). These findings suggest that the bioactive components of NH may target these pathways to treat UC.
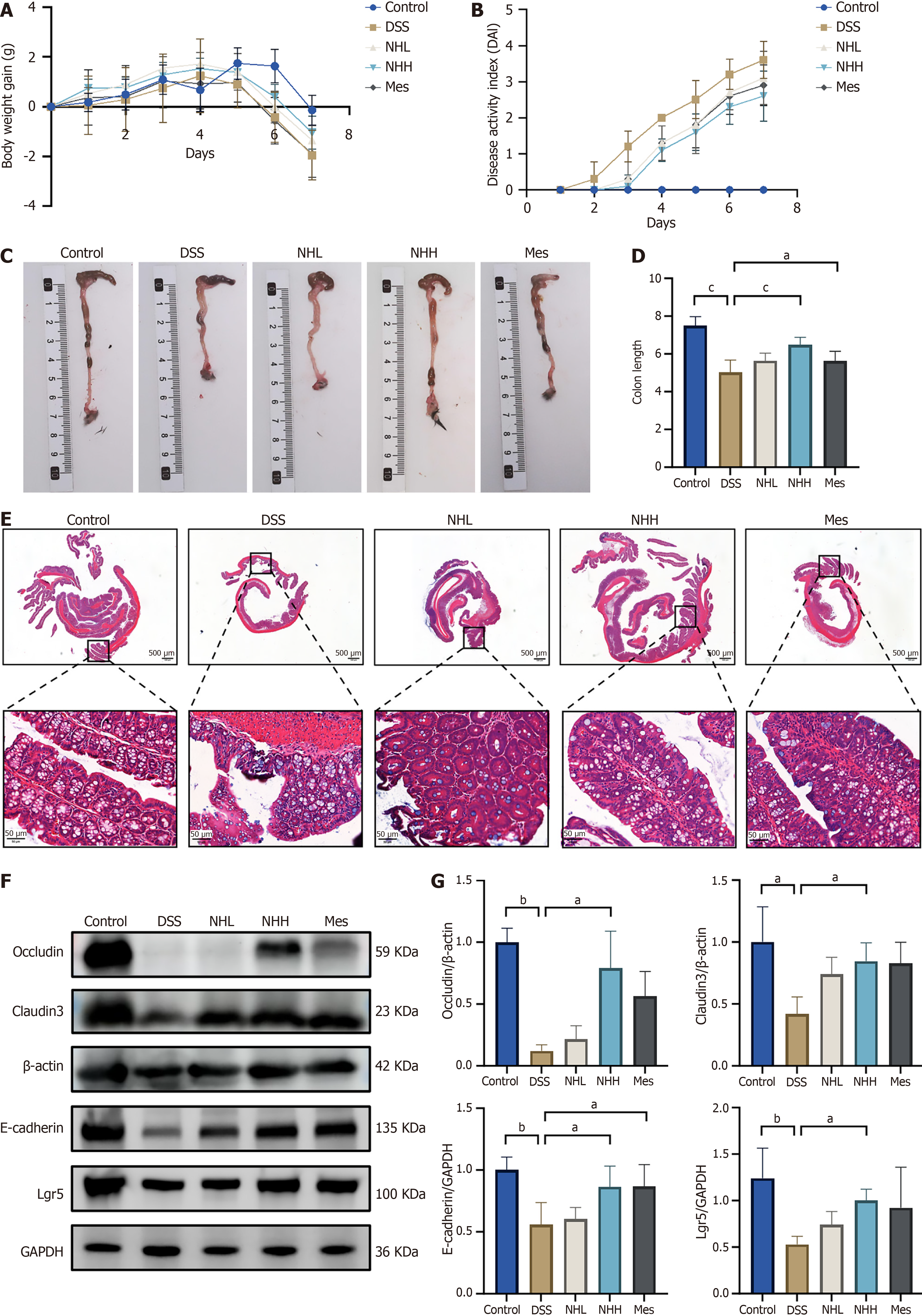
This study investigated the therapeutic effect of NH on UC using a DSS-induced colitis model. The mice were treated with 2.5% DSS in their drinking water, and NH was administered by oral gavage for 7 days. Compared with the normal group of mice, the DSS group exhibited significantly lower body weight, increased diarrhea and blood in the stools, higher disease activity index (DAI) scores and significantly shorter colon length. Compared with the DSS treatment, the administration of NH ameliorated DSS-induced colitis. The weight loss of the mice was significantly lower (Figure 2A), and both diarrhea and blood-related symptoms were significantly suppressed. The DAI scores were lower (Figure 2B), and the colon length was longer (Figure 2C and D). Additionally, hematoxylin and eosin staining was performed to evaluate pathological changes in the mouse colon. After DSS treatment, colonic mucosal damage occurred in the mice, with the disruption of crypt and villus structures and the infiltration of inflammatory cells. However, NH significantly reduced mucosal damage, restored the structural integrity of the crypts and villi, and alleviated inflammatory infiltration (Figure 2E). An impaired gut mucosal barrier is a critical feature of UC. We therefore assessed the protein levels of occludin, claudin3, E-cadherin, and Lgr5, which maintain intestinal barrier function. In the DSS group of mice, a decrease in the expression of these proteins in the colonic tissue was observed. This finding is indicative of a disruption in the intestinal barrier. High-dose NH treatment significantly upregulated the expression of these genes, indicating that it can protect the intestinal barrier (Figure 2F and G). These findings indicate that NH effectively attenuated colonic inflammation and preserved the integrity of the epithelial barrier.
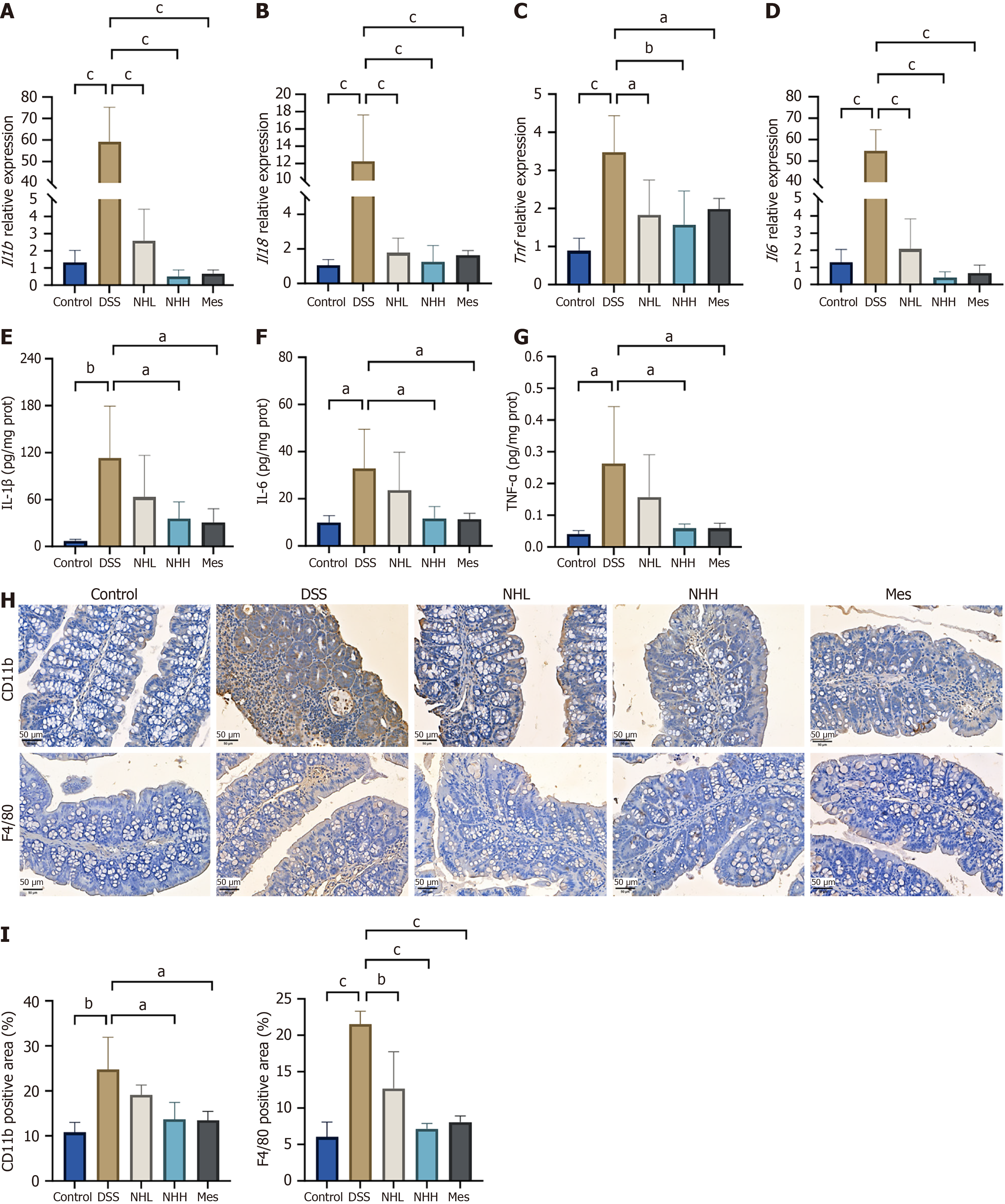
Additionally, we found that the DSS group significantly increased the mRNA levels of Il1β, Il18, Tnf and Il6 in colon tissue. Compared with the DSS group, the low and high dose of NH significantly reduced the expression of Il1β, Il18, Tnf and Il6 (Figure 3A-D). We used an enzyme-linked immunosorbent assay to measure the IL-1β, TNF-α, and IL-6 levels in colon tissue. The results demonstrated that, compared with the control group, the levels of IL-1β,
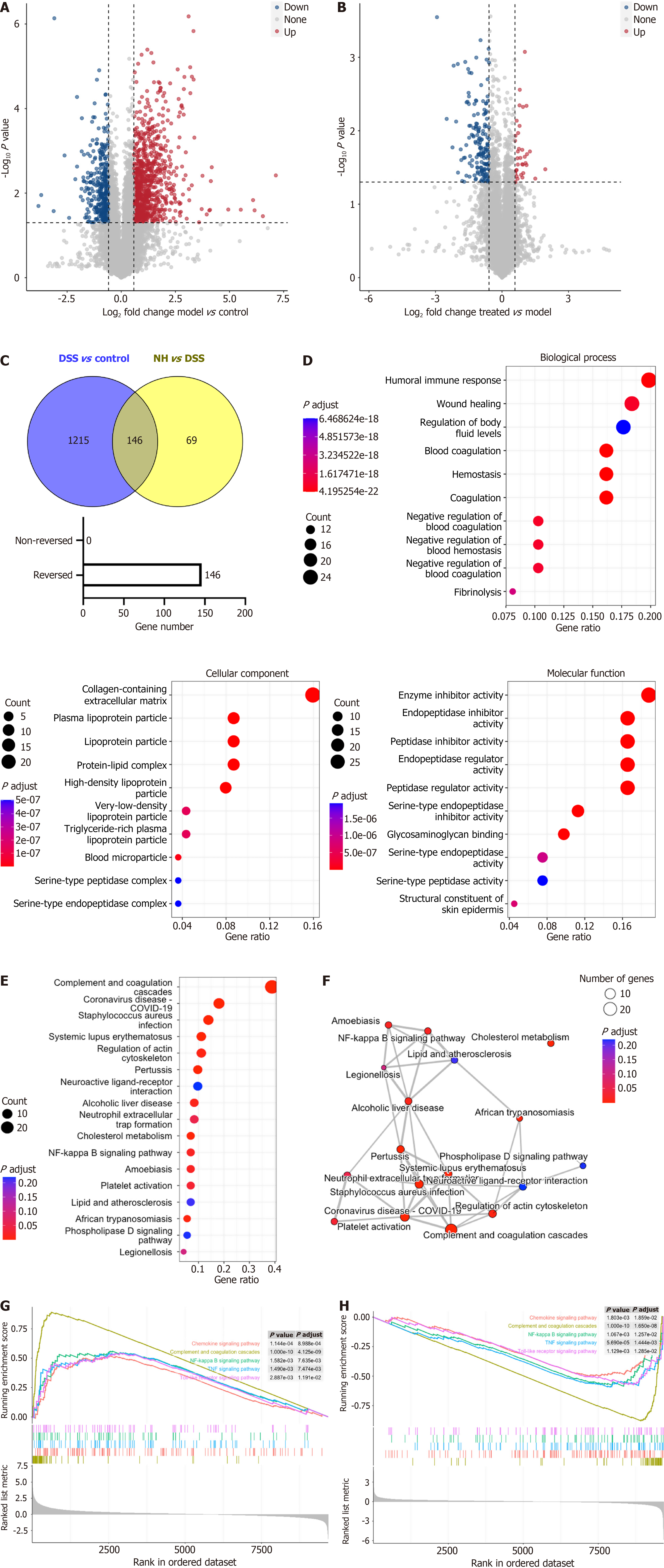
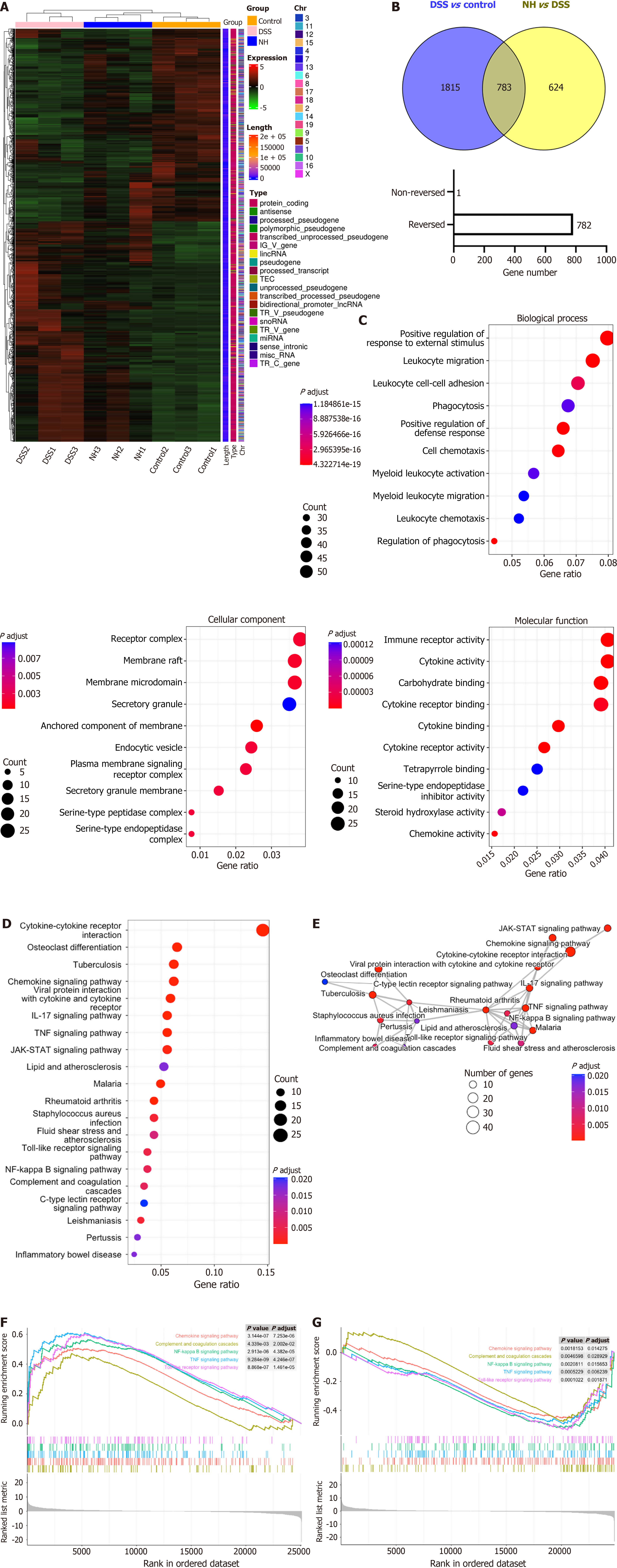
To investigate the molecular targets of NH, transcriptomic analysis was performed on colon tissue samples from three groups: The control group, the model group (DSS), and the high-dose NH group (NHH). A total of 1361 genes were identified as differentially expressed between the DSS group and the control group. Volcano plots were generated to visualize disparities in gene expression between the DSS group and the control group and between the NHH group and the DSS group (Figure 4A and B). Additionally, 215 genes were identified as differentially expressed between the NHH group and the DSS group. Among these genes, 146 genes were differentially expressed in the DSS group and their expression was restored after NHH treatment (Figure 4C).
To further explore the potential mechanisms of NH in the treatment of UC, GO and KEGG enrichment analyses were conducted. The results indicated that NH-regulated genes were associated with BP such as humoral immune response, coagulation, and wound healing; CC such as collagen-containing extracellular matrix, lipoprotein particle, and protein-lipid complex; and MF such as enzyme inhibitor activity, endopeptidase inhibitor activity, and peptidase inhibitor activity (Figure 4D). KEGG pathway analysis revealed that the majority of the differentially expressed genes were significantly enriched in complement and coagulation cascades, cholesterol metabolism, and the NF-κB signaling pathway (Figure 4E). Additionally, the network interactions among the top 20 pathways were visualized (Figure 4F).
Furthermore, gene set enrichment analysis (GSEA) was performed to comprehensively analyze the transcriptomic sequencing results and elucidate the biological changes in gene expression levels. The results demonstrated that genes associated with the chemokine signaling pathway, complement and coagulation cascades, NF-κB signaling pathway, TNF signaling pathway, and toll-like receptor signaling pathway were upregulated in the DSS group compared with the control group. Conversely, the genes associated with the chemokine signaling pathway, complement and coagulation cascades, NF-κB signaling pathway, TNF signaling pathway and toll-like receptor signaling pathway were downregulated in the NHH group compared with the DSS group (Figure 4G and H).
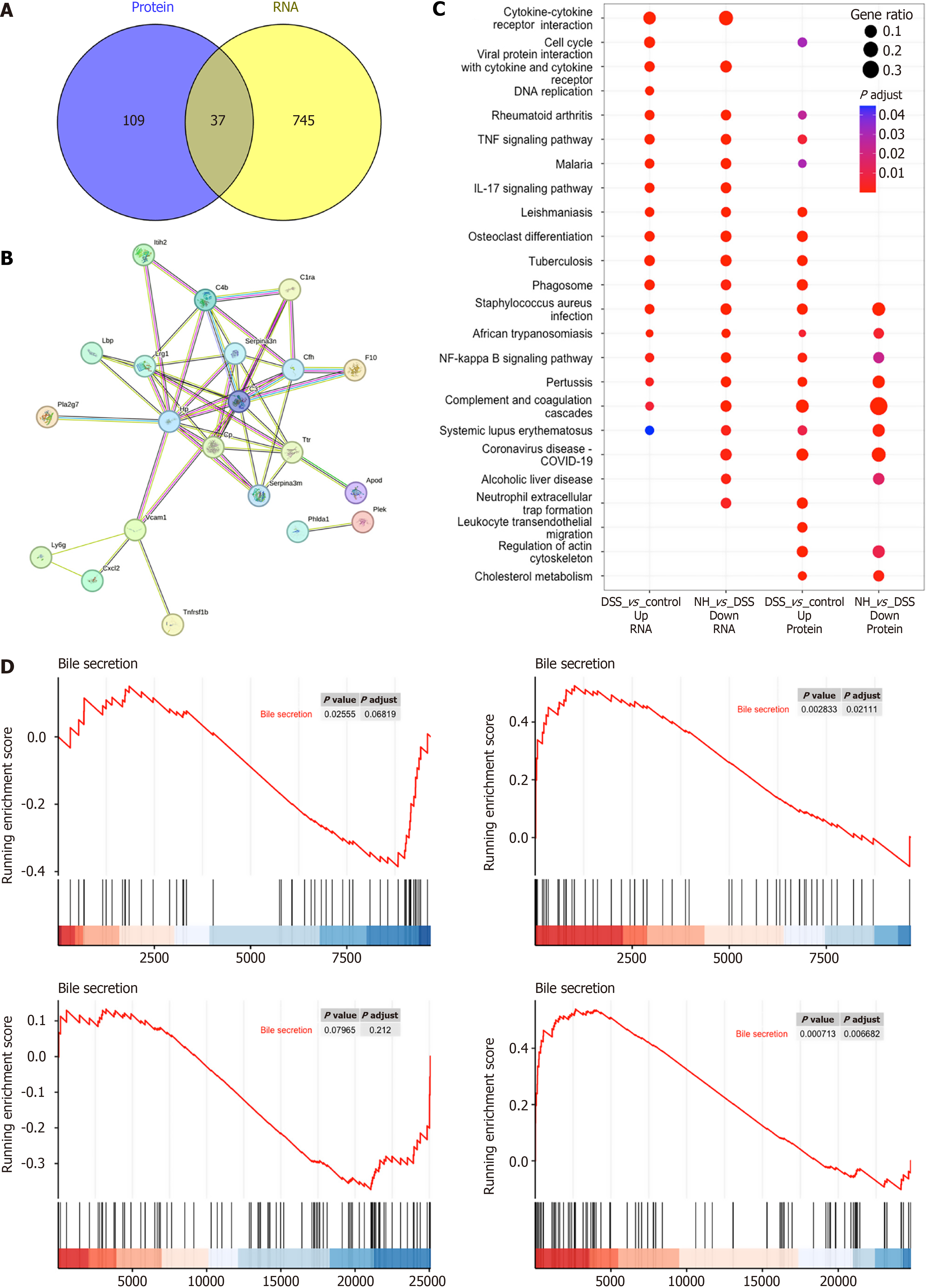
To further investigate the mechanisms underlying the effects of NH treatment on UC mice, we performed proteomics sequencing on colon tissues from the DSS, NHH, and control groups to quantify protein expression. The heatmaps depict the hierarchical clustering of the differentially expressed proteins (Figure 5A). A Venn diagram revealed 2598 differentially expressed proteins between the DSS and control groups and 1407 between the NHH and DSS groups. Among these differentially expressed proteins, 782 were dysregulated in the DSS group and restored to baseline levels after NHH treatment (Figure 5B).
We performed GO and KEGG enrichment analyses on the proteomics sequencing data. The results demonstrated that NH-regulated proteins were associated with BP such as leukocyte migration, phagocytosis, and cell chemotaxis; CC such as receptor complex, membrane raft, and membrane microdomain; and MF such as immune receptor activity, cytokine activity, and carbohydrate binding (Figure 5C). KEGG pathway analysis revealed significant enrichment in pathways such as the chemokine signaling pathway, TNF signaling pathway, toll-like receptor signaling pathway, NF-κB signaling pathway, complement and coagulation cascades, and IBD (Figure 5D). Furthermore, the network interactions among the top 20 enriched pathways were visualized (Figure 5E).
The results from GSEA based on proteomics sequencing were consistent with those from transcriptomics sequencing. Compared with the control group, the DSS group exhibited upregulated expression of genes related to the chemokine signaling pathway, complement and coagulation cascades, the NF-κB signaling pathway, the TNF signaling pathway, and the toll-like receptor signaling pathway. Conversely, NH treatment attenuated the expression of these pathways (Figure 5F and G).
On the basis of the sequencing results, we determined that the differentially expressed genes were dysregulated in the DSS group and subsequently normalized following NHH treatment. These genes exhibited consistent differential expression at both the protein and RNA levels (Figure 6A). In addition, interaction analysis of these genes revealed that C3 served as a central node in the network (Figure 6B). Additionally, a comprehensive analysis of the enriched pathways identified in the KEGG pathway analysis revealed that pathways such as Staphylococcus aureus infection, African trypanosomiasis, the NF-κB signaling pathway, pertussis, complement and coagulation cascades, and systemic lupus erythematosus were upregulated in the DSS group relative to those in the control group and downregulated in the NHH group relative to those in the DSS group. This trend was observed at both the RNA and protein levels (Figure 6C).
Since NH is an animal bile-derived drug, we assessed bile secretion changes via GSEA in DSS vs control and NHH vs DSS. The results revealed that bile secretion was downregulated in the DSS group but upregulated after NH treatment. This trend was observed by both transcriptomics and proteomics (Figure 6D).
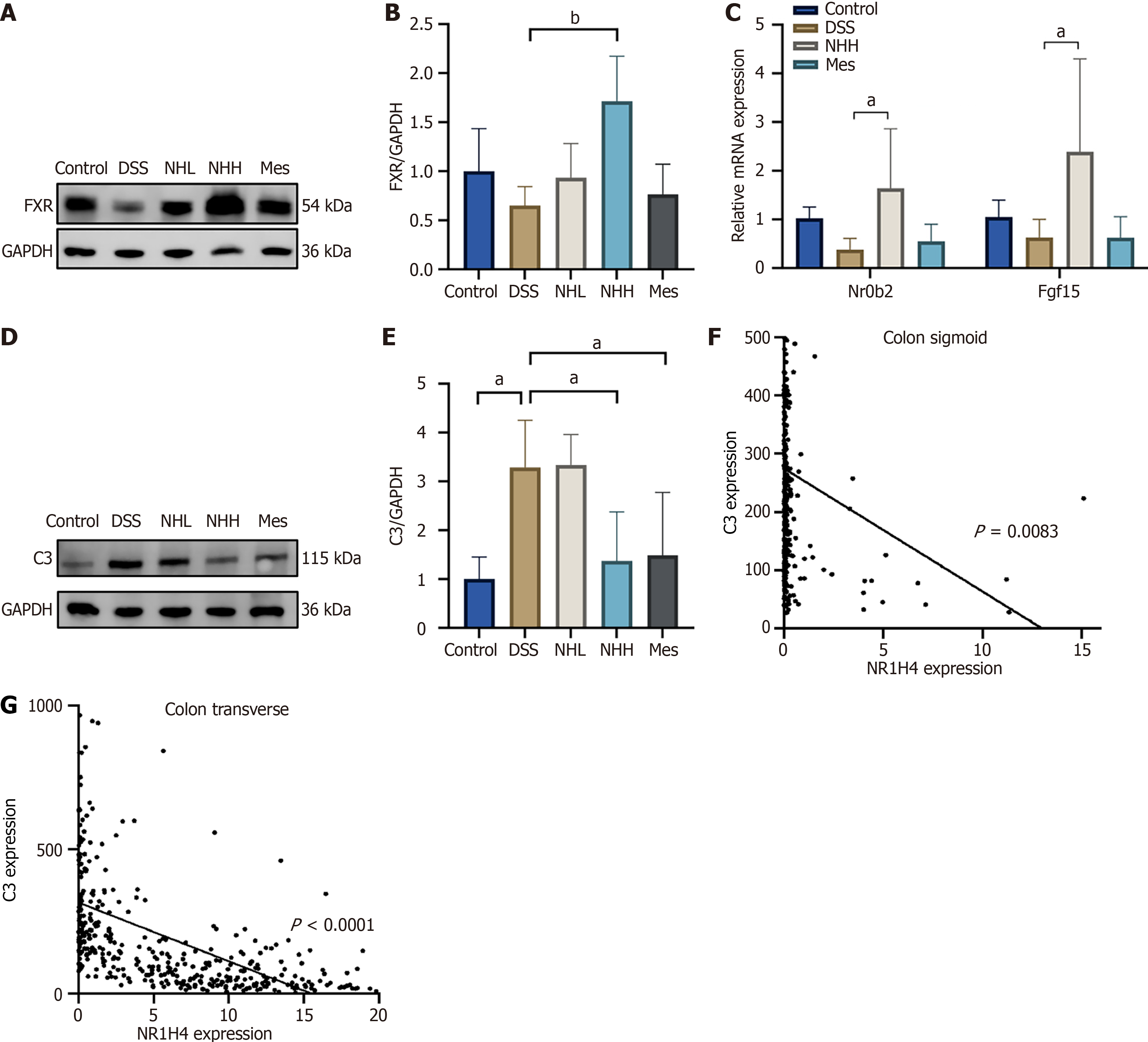
NH, a bile-derived traditional Chinese medicine, has demonstrated clinical efficacy. FXR is a classic bile acid receptor. Activation of FXR has been demonstrated to have anti-inflammatory effects and to play a protective role in chemically induced colitis. To further confirm these results, western blot analysis was performed on colonic tissue. Compared with the control group, DSS treatment significantly decreased FXR expression. Conversely, NH treatment significantly increased FXR expression (Figure 7A and B). The results of the polymerase chain reaction analysis revealed that NH significantly upregulated the expression of the FXR target genes Nr0b2 and Fgf15 (Figure 7C), indicating activation of FXR. Compared with the control group, DSS treatment significantly increased C3 expression. Conversely, NH treatment significantly decreased C3 expression (Figure 7D and E). Pearson correlation analysis revealed that C3 expression was inversely correlated with FXR expression in the sigmoid colon and transverse colon (Figure 7F and G).
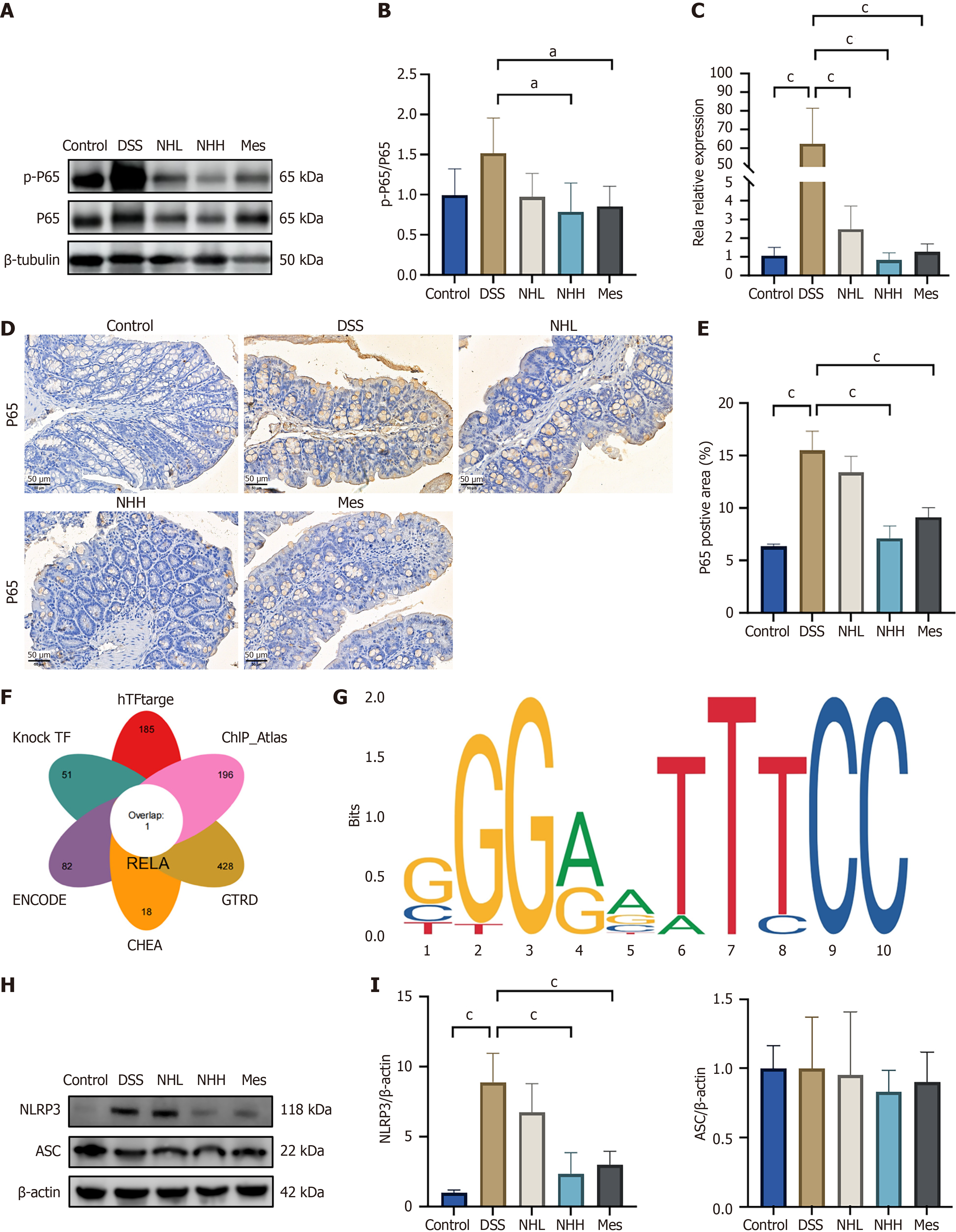
NF-κB is a pivotal transcription factor in inflammatory responses. Compared with the control group, the DSS group exhibited increased NF-κB p-p65 expression. However, following NH treatment, the expression of NF-κB p-p65 expression decreased, particularly at a high dose of NH (Figure 8A and B). The mRNA level of Rela in the DSS group was significantly upregulated (Figure 8C). Compared with the DSS group, the middle and high-dose of NH significantly reduced Rela expression. Additionally, IHC analysis showed that P65 expression was reduced in the high-dose NH group compared with that in the DSS group (Figure 8D and E). These results suggest that the NH significantly affects the transcription of P65, thereby mitigating inflammatory damage following DSS exposure. By screening several transcription factor databases, we identified RELA (NF-κB p65) as an overlapping transcription factor that regulates C3 (Figure 8F). Through bioinformatics analysis, we identified a potential binding site of RELA located in the C3 gene promoter that can be bound and activated by NF-κB p65 (Figure 8G). In addition, we found that the DSS group increased NLRP3 exp
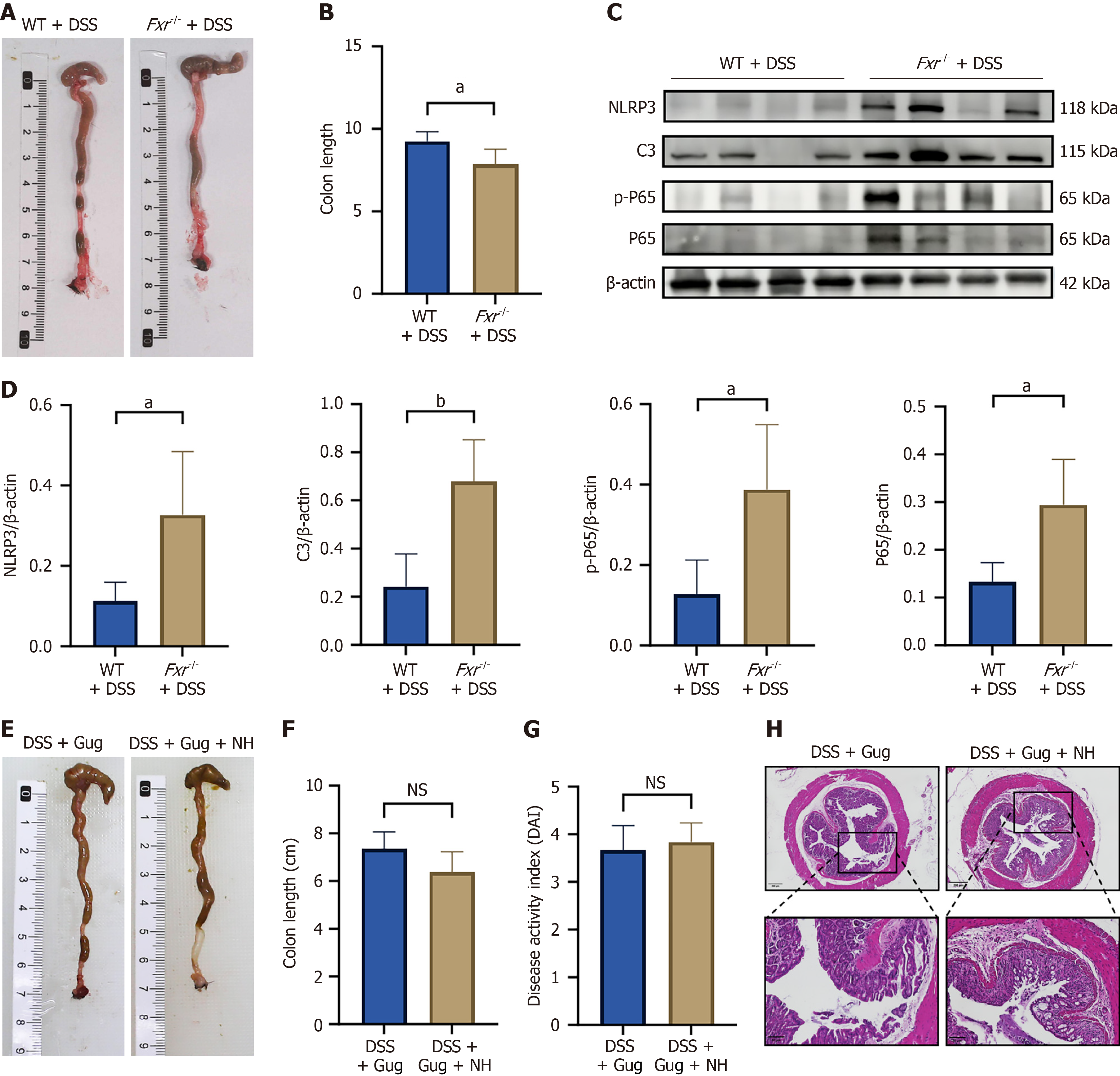
To understand the potential biological mechanism by which NH alleviates colitis, the Fxr-/- mouse models and FXR inhibitor Gug were used. Fxr knockout resulted in a reduction of the colonic length (Figure 9A and B) and increased the expression of NLRP3, C3, p-P65 and P65 (Figure 9C and D). The experimental results demonstrated that Gug significantly impaired the therapeutic efficacy of NH in the colitis model, as evidenced by colon length and DAI scores that were not improved (Figure 9E-G). Moreover, following Gug administration, NH failed to protect against inflammatory cell infiltration and mucosal damage in colonic tissue (Figure 9H). These data suggest that FXR activation is essential for the therapeutic effect of NH on colitis.
The incidence of UC is increasing worldwide. Nevertheless, its pathogenesis remains incompletely understood because of its mechanistic complexity. Current therapeutic interventions primarily serve to alleviate symptoms rather than induce a state of complete recovery. NH, a traditional Chinese medicine derived from animal bile, has demonstrated significant therapeutic efficacy in the treatment of gut injury[16]. We used network pharmacology and multi-omics integrated analysis to explore the active components and targets of NH in the treatment of UC. Through in vivo experiments, we evaluated the therapeutic potential of NH and elucidated its mechanisms in the inhibition of colitis.
NH has been demonstrated to have a protective effect on UC through its antioxidative and anti-inflammatory effects[17]. Additionally, previous studies have indicated that NH also protects the blood-brain barrier, promotes the repair of damaged endothelial cells and tight junction proteins, and reduces inflammatory damage and inflammatory factor levels in the context of cerebrovascular disease[14]. These findings suggest that NH may similarly ameliorate UC by restoring barrier function and reducing inflammatory damage. Our animal experiments demonstrated that NH treatment alleviated DSS-induced colitis in mice, significantly ameliorating colitis-related symptoms, including weight loss, diarrhea, bloody stools, and colon shortening. Furthermore, NH administration reduced inflammatory cell infiltration and mitigated mucosal damage in colonic tissues. Crucially, NH treatment upregulated the expression of intestinal barrier function-related proteins, including occludin, claudin3, E-cadherin, and the stem cell marker Lgr5. These results align with those of prior research and substantiate the therapeutic potential of NH for ameliorating UC-related symptoms, attenuating inflammatory damage, and restoring intestinal barrier integrity.
A characteristic feature of traditional Chinese medicine is its ability to simultaneously target multiple biological pathways, a property that may exceed the efficacy of single-target therapeutic approaches. NH, characterized by multiple components and targets, has significant therapeutic potential in UC. Network pharmacology analysis revealed a total of 16 NH components, including taurodeoxycholic acid, glycocholic acid, glycodeoxycholic acid, cholic acid, and taurine. Collectively, these data indicate that bile acids constitute the predominant class of bioactive constituents within NH. Furthermore, integrated transcriptomic and proteomic analyses demonstrated that NH intervention significantly increased bile secretion. Bile acids play established roles as signaling molecules in metabolic and inflammatory diseases[21-24]. Bile acids have been reported to inhibit NLRP3 inflammasome activation, which endogenously constrains NLRP3 inflammasome-related inflammation[23]. Recent evidence suggests that BA and its derived metabolites play a role in intestinal immunity and are associated with the progression of IBD and colorectal cancer[22]. Taurodeoxycholate attenuates colitis by modulating the balance between M1 and M2 macrophages and suppressing Th1, Th2, and Th17 cells[25]. Taurocholic acid and glycocholic acid were identified as the predominant bile acids in NH and can significantly inhibit macrophage migration and the secretion of proinflammatory cytokines and chemokines. Furthermore, taurocholic acid or glycocholic acid administration markedly increased FXR expression at the mRNA and protein levels[26]. The results of the above studies indicate that bile acids in NH act as significant anti-inflammatory signaling molecules, mediating their effects through FXR activation. FXR is a key regulatory factor in the bile acid signaling pathway. Research has shown that FXR activation is beneficial for alleviating UC[27]. Conversely, knockout or silencing of FXR exacerbates mucosal inflammation in mouse models of colitis[28]. In this study, NH intervention significantly increased the protein expression of FXR and its downstream targets Nr0b2 and Fgf15 in the colon tissue of UC mice, confirming the activation of the FXR pathway. We found that the FXR inhibitor, Gug, significantly diminished the therapeutic action of the NH in the colitis model. These results indicate that the activation of FXR prevents intestinal inflammation, reduces mucosal damage, promotes intestinal healing, and exerts protective effects in colitis[27,29].
FXR has been shown to regulate the expression of component C3[12], which may be related to the development of inflammation in the colon. The complement system is involved in intestinal inflammation and is associated with IBD[5]. C3 is a key mediator of the gut complement system and plays a crucial role in colitis progression and therapeutic response[7]. Integrated network pharmacology, transcriptomic and proteomic analyses revealed that NH has regulatory effects on complement and coagulation cascades. Moreover, interaction analysis of differentially expressed genes identified complement component C3 as a hub gene. In this study, correlation analysis revealed that C3 expression was inversely correlated with FXR expression in the human sigmoid colon and transverse colon, which differs from the findings of a previous rat study in which an FXR agonist was used[12]. We observed elevated C3 expression in DSS-induced colitis, which was significantly attenuated by NH treatment. Complement inhibition reduces inflammatory cytokines, leukocyte infiltration, and tissue damage while promoting wound healing and mucosal repair[9]. These findings suggest that NH activates FXR, resulting in the suppression of C3 expression and anti-inflammatory activity.
NF-κB signaling is antagonized by FXR, which may be an intermediate step in the anti-inflammatory activity of NH[30]. We found that FXR knockout increased the expression of p-P65 and P65. Mechanistically, Fxr suppresses p65 transactivation activity by reducing its DNA binding activity. Moreover, FXR activation represses specific sets of NF-κB target genes[31]. Additionally, integrated transcriptomic and proteomic analyses revealed that NH altered the NF-κB signaling pathway similarly at the RNA and protein levels. The results of western blotting, polymerase chain reaction, and IHC demonstrated that P65 was upregulated in the DSS group compared with that in the control group. In contrast, P65 exhibited decreased expression in the NHH group compared with that in the DSS group. These results suggest that NH significantly affects the transcription of P65, thereby mitigating inflammatory damage following DSS exposure. Bioinformatics analysis identified RELA (P65) as an overlapping transcription factor that regulates C3. Analysis of the human complement C3 gene revealed the presence of functional RELA response elements in the promoter of C3. Collectively, these findings establish NF-κB as a key transcriptional regulator of complement component C3[32]. NF-κB-mediated inflammation represents a central pathogenic mechanism in DSS-induced colitis[33]. Furthermore, the NF-κB pathway is closely linked to the NLRP3 inflammasome, as NF-κB plays a pivotal role in inflammasome activation[34]. Inhibiting the NF-κB/NLRP3 signaling pathway has been reported to ameliorate intestinal inflammation and mucosal barrier alterations in DSS-induced colitis rats[35,36].
Functioning as a pivotal signaling node, complement component C3 transmits upstream initiation signals to engage downstream effectors, which can lead to the potent stimulation and amplification of immune and inflammatory pathways[37]. Complement activation has been shown to serve as a key driver for NLRP3 inflammasome activation, potentiating the subsequent secretion of IL-1β and IL-18[11]. Notably, NLRP3 inflammasome activation is associated with the onset and progression of UC[38]. Complement component C3 and its metabolic or cleavage products engage and activate the NLRP3 inflammasome. For example, C3d, as a cleavage product of C3, triggers IL-1β secretion via activation of the NLRP3 inflammasome[39]. Binding of the C3 metabolic product C3a to its receptor induces oxidative stress and generates reactive oxygen species. The resulting reactive oxygen species overload induces NLRP3 deubiquitination, thereby promoting rapid activation of the NLRP3 inflammasome. In this study, we observed markedly elevated NLRP3 expression in DSS-induced colitis, which was significantly attenuated following NH treatment. These results indicate that therapeutic targeting of the C3/NLRP3 axis may effectively attenuate UC. In summary, our findings establish that NH has significant therapeutic efficacy in UC. Mechanistically, NH activates the bile acid receptor FXR, which in turn suppresses NF-κB signaling. This cascade further inhibits complement C3 and NLRP3 inflammasome overactivation, mediating potent anti-inflammatory effects.
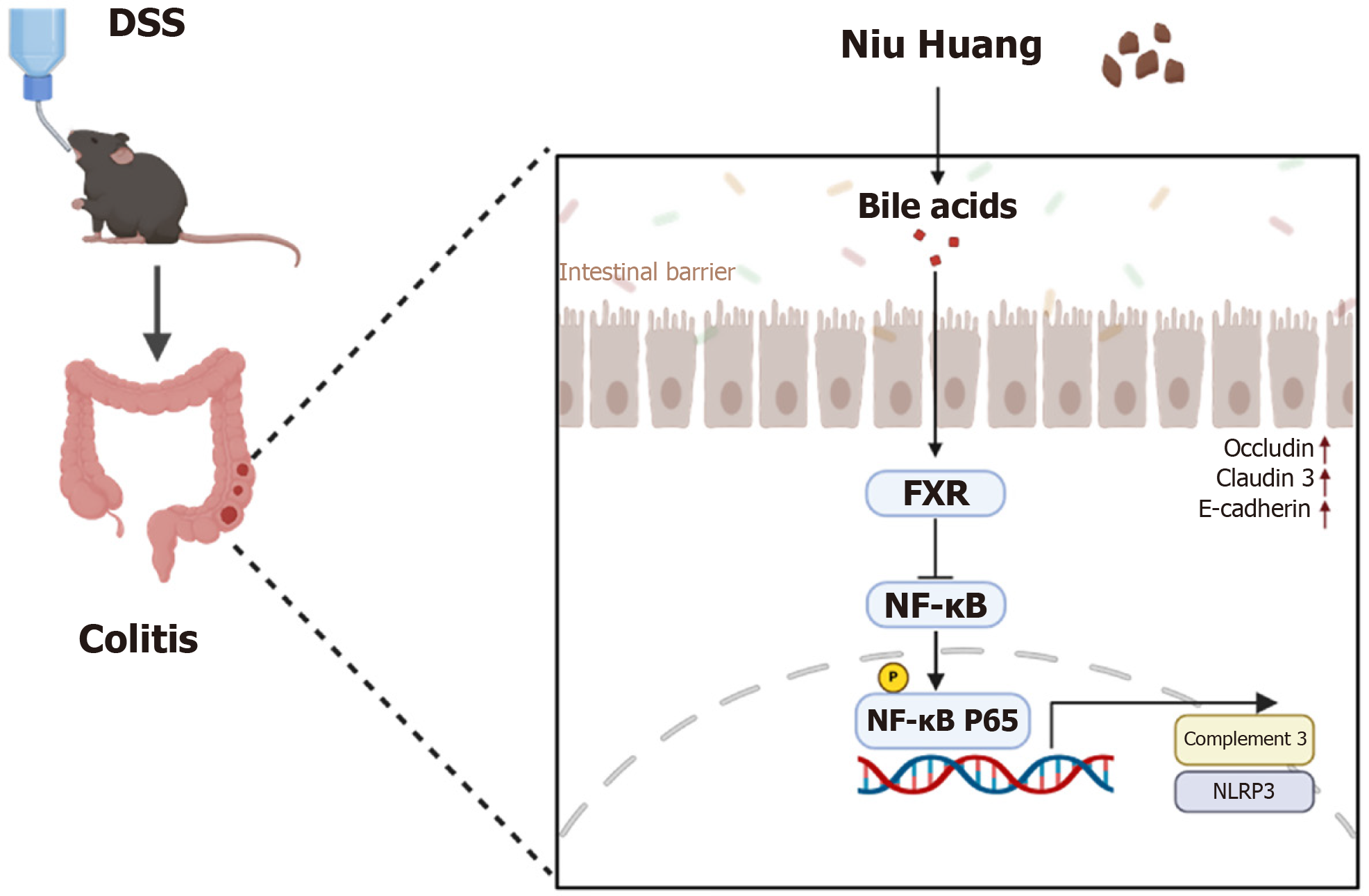
In this study, network pharmacology, transcriptomics, proteomics, and experimental validation were performed to confirm the protective role of NH in colitis and to elucidate its mechanism. Our findings suggest that the potential mechanisms through which NH acts against colitis are associated with the activation of FXR, which reduces the accumulation of complement C3 in colonic tissue and the excessive activation of the NLRP3 inflammasome by inhibiting the NF-κB inflammatory pathway. These results underscore the complex role of NH in UC treatment through FXR activation and C3/NLRP3 signaling (Figure 10). This study provides insights into the anti-inflammatory effects of NH and the underlying mechanisms involved and establishes a basis for its therapeutic application.
| 1. | Le Berre C, Honap S, Peyrin-Biroulet L. Ulcerative colitis. Lancet. 2023;402:571-584. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1251] [Cited by in RCA: 1082] [Article Influence: 360.7] [Reference Citation Analysis (5)] |
| 2. | Gros B, Kaplan GG. Ulcerative Colitis in Adults: A Review. JAMA. 2023;330:951-965. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 562] [Cited by in RCA: 486] [Article Influence: 162.0] [Reference Citation Analysis (1)] |
| 3. | de Souza HS, Fiocchi C. Immunopathogenesis of IBD: current state of the art. Nat Rev Gastroenterol Hepatol. 2016;13:13-27. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1339] [Cited by in RCA: 1203] [Article Influence: 120.3] [Reference Citation Analysis (5)] |
| 4. | Kulkarni DH, Starick M, Aponte Alburquerque R, Kulkarni HS. Local complement activation and modulation in mucosal immunity. Mucosal Immunol. 2024;17:739-751. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 22] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 5. | Jain U, Otley AR, Van Limbergen J, Stadnyk AW. The complement system in inflammatory bowel disease. Inflamm Bowel Dis. 2014;20:1628-1637. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 47] [Article Influence: 3.9] [Reference Citation Analysis (1)] |
| 6. | Sorbara MT, Foerster EG, Tsalikis J, Abdel-Nour M, Mangiapane J, Sirluck-Schroeder I, Tattoli I, van Dalen R, Isenman DE, Rohde JR, Girardin SE, Philpott DJ. Complement C3 Drives Autophagy-Dependent Restriction of Cyto-invasive Bacteria. Cell Host Microbe. 2018;23:644-652.e5. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 103] [Article Influence: 12.9] [Reference Citation Analysis (0)] |
| 7. | Ohno M, Hasegawa M, Hayashi A, Caballero-Flores G, Alteri CJ, Lawley TD, Kamada N, Núñez G, Inohara N. Lipopolysaccharide O structure of adherent and invasive Escherichia coli regulates intestinal inflammation via complement C3. PLoS Pathog. 2020;16:e1008928. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 21] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 8. | Ward M, Eastwood MA. Serum C3 and C4 complement components in ulcerative colitis and Crohn's disease. Digestion. 1975;13:100-103. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 12] [Article Influence: 0.2] [Reference Citation Analysis (0)] |
| 9. | Schepp-Berglind J, Atkinson C, Elvington M, Qiao F, Mannon P, Tomlinson S. Complement-dependent injury and protection in a murine model of acute dextran sulfate sodium-induced colitis. J Immunol. 2012;188:6309-6318. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 40] [Article Influence: 2.9] [Reference Citation Analysis (0)] |
| 10. | Gu D, Wang H, Yan M, Li Y, Yang S, Shi D, Guo S, Wu L, Liu C. Echinacea purpurea (L.) Moench extract suppresses inflammation by inhibition of C3a/C3aR signaling pathway in TNBS-induced ulcerative colitis rats. J Ethnopharmacol. 2023;307:116221. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 17] [Reference Citation Analysis (0)] |
| 11. | Dai L, Chen Y, Wu J, He Z, Zhang Y, Zhang W, Xie Y, Zeng H, Zhong X. A novel complement C3 inhibitor CP40-KK protects against experimental pulmonary arterial hypertension via an inflammasome NLRP3 associated pathway. J Transl Med. 2024;22:164. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 12] [Reference Citation Analysis (0)] |
| 12. | Li J, Pircher PC, Schulman IG, Westin SK. Regulation of complement C3 expression by the bile acid receptor FXR. J Biol Chem. 2005;280:7427-7434. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 78] [Article Influence: 3.7] [Reference Citation Analysis (3)] |
| 13. | Huang Z, Meng FY, Lu LZ, Guo QQ, Lv CJ, Tan NH, Deng Z, Chen JY, Zhang ZS, Zou B, Long HP, Zhou Q, Tian S, Mei S, Tian XF. Calculus bovis inhibits M2 tumor-associated macrophage polarization via Wnt/β-catenin pathway modulation to suppress liver cancer. World J Gastroenterol. 2024;30:3511-3533. [PubMed] [DOI] [Full Text] |
| 14. | Du X, Li C, Zhang S, Sun C, Zhang X, Chen C, Wang X, Cheng F, Wang Q. Exploring the pharmacological mechanism of calculus bovis in cerebral ischaemic stroke using a network pharmacology approach. J Ethnopharmacol. 2022;284:114507. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 15] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 15. | Yu ZJ, Xu Y, Peng W, Liu YJ, Zhang JM, Li JS, Sun T, Wang P. Calculus bovis: A review of the traditional usages, origin, chemistry, pharmacological activities and toxicology. J Ethnopharmacol. 2020;254:112649. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 61] [Article Influence: 10.2] [Reference Citation Analysis (1)] |
| 16. | Xiang D, Liu Y, Zu Y, Yang J, He W, Zhang C, Liu D. Calculus Bovis Sativus alleviates estrogen cholestasis-induced gut and liver injury in rats by regulating inflammation, oxidative stress, apoptosis, and bile acid profiles. J Ethnopharmacol. 2023;302:115854. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 17] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 17. | Li X, Xu Y, Zhang C, Deng L, Chang M, Yu Z, Liu D. Protective Effect of Calculus Bovis Sativus on Dextran Sulphate Sodium-Induced Ulcerative Colitis in Mice. Evid Based Complement Alternat Med. 2015;2015:469506. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 22] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 18. | Fang S, Dong L, Liu L, Guo J, Zhao L, Zhang J, Bu D, Liu X, Huo P, Cao W, Dong Q, Wu J, Zeng X, Wu Y, Zhao Y. HERB: a high-throughput experiment- and reference-guided database of traditional Chinese medicine. Nucleic Acids Res. 2021;49:D1197-D1206. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 581] [Cited by in RCA: 480] [Article Influence: 96.0] [Reference Citation Analysis (4)] |
| 19. | He W, Xu Y, Zhang C, Lu J, Li J, Xiang D, Yang J, Chang M, Liu D. Hepatoprotective effect of calculus bovis sativus on nonalcoholic fatty liver disease in mice by inhibiting oxidative stress and apoptosis of hepatocytes. Drug Des Devel Ther. 2017;11:3449-3460. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 20] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 20. | Wirtz S, Popp V, Kindermann M, Gerlach K, Weigmann B, Fichtner-Feigl S, Neurath MF. Chemically induced mouse models of acute and chronic intestinal inflammation. Nat Protoc. 2017;12:1295-1309. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1333] [Cited by in RCA: 1208] [Article Influence: 134.2] [Reference Citation Analysis (3)] |
| 21. | Chávez-Talavera O, Tailleux A, Lefebvre P, Staels B. Bile Acid Control of Metabolism and Inflammation in Obesity, Type 2 Diabetes, Dyslipidemia, and Nonalcoholic Fatty Liver Disease. Gastroenterology. 2017;152:1679-1694.e3. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 847] [Cited by in RCA: 773] [Article Influence: 85.9] [Reference Citation Analysis (7)] |
| 22. | Cai J, Sun L, Gonzalez FJ. Gut microbiota-derived bile acids in intestinal immunity, inflammation, and tumorigenesis. Cell Host Microbe. 2022;30:289-300. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 706] [Cited by in RCA: 610] [Article Influence: 152.5] [Reference Citation Analysis (8)] |
| 23. | Guo C, Xie S, Chi Z, Zhang J, Liu Y, Zhang L, Zheng M, Zhang X, Xia D, Ke Y, Lu L, Wang D. Bile Acids Control Inflammation and Metabolic Disorder through Inhibition of NLRP3 Inflammasome. Immunity. 2016;45:802-816. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 650] [Cited by in RCA: 596] [Article Influence: 59.6] [Reference Citation Analysis (3)] |
| 24. | Jia W, Li Y, Cheung KCP, Zheng X. Bile acid signaling in the regulation of whole body metabolic and immunological homeostasis. Sci China Life Sci. 2024;67:865-878. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 128] [Article Influence: 64.0] [Reference Citation Analysis (0)] |
| 25. | Zou Y, Ghaderpour A, Munkhbileg B, Seo SU, Seong SY. Taurodeoxycholate ameliorates DSS-induced colitis in mice. Int Immunopharmacol. 2023;122:110628. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 10] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 26. | Ge X, Huang S, Ren C, Zhao L. Taurocholic Acid and Glycocholic Acid Inhibit Inflammation and Activate Farnesoid X Receptor Expression in LPS-Stimulated Zebrafish and Macrophages. Molecules. 2023;28:2005. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 45] [Cited by in RCA: 39] [Article Influence: 13.0] [Reference Citation Analysis (0)] |
| 27. | Gadaleta RM, van Erpecum KJ, Oldenburg B, Willemsen EC, Renooij W, Murzilli S, Klomp LW, Siersema PD, Schipper ME, Danese S, Penna G, Laverny G, Adorini L, Moschetta A, van Mil SW. Farnesoid X receptor activation inhibits inflammation and preserves the intestinal barrier in inflammatory bowel disease. Gut. 2011;60:463-472. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 742] [Cited by in RCA: 698] [Article Influence: 46.5] [Reference Citation Analysis (3)] |
| 28. | Liu F, Yao Y, Wang Q, Zhang F, Wang M, Zhu C, Lin C. Nigakinone alleviates DSS-induced experimental colitis via regulating bile acid profile and FXR/NLRP3 signaling pathways. Phytother Res. 2023;37:15-34. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 22] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 29. | Liu X, Milik A, O’connell R, White J, Wagner B. P029 Activation of FXR restores expression of genes dysregulated in human IBD and suppresses TNFa and IL-6 signaling pathways in adoptive transfer colitis model. J Crohn's Colitis. 2021;15:S145-S146. [DOI] [Full Text] |
| 30. | Pi Y, Wu Y, Zhang X, Lu D, Han D, Zhao J, Zheng X, Zhang S, Ye H, Lian S, Bai Y, Wang Z, Tao S, Ni D, Zou X, Jia W, Zhang G, Li D, Wang J. Gut microbiota-derived ursodeoxycholic acid alleviates low birth weight-induced colonic inflammation by enhancing M2 macrophage polarization. Microbiome. 2023;11:19. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 112] [Cited by in RCA: 96] [Article Influence: 32.0] [Reference Citation Analysis (0)] |
| 31. | Wang YD, Chen WD, Wang M, Yu D, Forman BM, Huang W. Farnesoid X receptor antagonizes nuclear factor kappaB in hepatic inflammatory response. Hepatology. 2008;48:1632-1643. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 566] [Cited by in RCA: 528] [Article Influence: 29.3] [Reference Citation Analysis (4)] |
| 32. | Nitkiewicz J, Borjabad A, Morgello S, Murray J, Chao W, Emdad L, Fisher PB, Potash MJ, Volsky DJ. HIV induces expression of complement component C3 in astrocytes by NF-κB-dependent activation of interleukin-6 synthesis. J Neuroinflammation. 2017;14:23. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 37] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 33. | Zheng J, Zhang J, Zhou Y, Zhang D, Guo H, Li B, Cui S. Taurine Alleviates Experimental Colitis by Enhancing Intestinal Barrier Function and Inhibiting Inflammatory Response through TLR4/NF-κB Signaling. J Agric Food Chem. 2024;72:12119-12129. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 37] [Article Influence: 18.5] [Reference Citation Analysis (0)] |
| 34. | Chen Y, Chen M, Chen H, Zhu JH, Liang C, Wu S, Gu S, Sun W, Yan J. Crosstalk patterns of necroptosis signaling and NLRP3 inflammasome in the colonic epithelium and its initial role in colitis. Int Immunopharmacol. 2025;152:114413. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 6] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 35. | Dai L, Cao X, Miao X, Yang X, Zhang J, Shang X. The chemical composition, protective effect of Rheum officinale leaf juice and its mechanism against dextran sulfate sodium-induced ulcerative colitis. Phytomedicine. 2024;129:155653. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 9] [Reference Citation Analysis (0)] |
| 36. | Han F, Zhao X, Li X, Peng L, Liu W, Han J. Bovine lactoferricin ameliorates intestinal inflammation and mucosal barrier lesions in colitis through NF-κB/NLRP3 signaling pathways. J Funct Foods. 2022;93:105090. [RCA] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 9] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 37. | Xu L, Xu H, Chen S, Jiang W, Afridi SK, Wang Y, Ren X, Zhao Y, Lai S, Qiu X, Alvin Huang YW, Cui Y, Yang H, Qiu W, Tang C. Inhibition of complement C3 signaling ameliorates locomotor and visual dysfunction in autoimmune inflammatory diseases. Mol Ther. 2023;31:2715-2733. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 26] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 38. | Bauer C, Duewell P, Mayer C, Lehr HA, Fitzgerald KA, Dauer M, Tschopp J, Endres S, Latz E, Schnurr M. Colitis induced in mice with dextran sulfate sodium (DSS) is mediated by the NLRP3 inflammasome. Gut. 2010;59:1192-1199. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 810] [Cited by in RCA: 754] [Article Influence: 47.1] [Reference Citation Analysis (0)] |
| 39. | Gogoi D, Yu H, Casey M, Baird R, Yusuf A, Forde L, O' Brien ME, West JR, Flagg T, McElvaney NG, Eden E, Mueller C, Brantly ML, Geraghty P, Reeves EP. Monocyte NLRP3 inflammasome and interleukin-1β activation modulated by alpha-1 antitrypsin therapy in deficient individuals. Thorax. 2024;79:822-833. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 5] [Article Influence: 2.5] [Reference Citation Analysis (0)] |