Published online Mar 25, 2026. doi: 10.5501/wjv.v15.i1.114375
Revised: October 11, 2025
Accepted: December 24, 2025
Published online: March 25, 2026
Processing time: 177 Days and 6.7 Hours
Low-level viremia (LLV) defined as [human immunodeficiency virus (HIV)-RNA 51-999 copies/mL] has been associated with an increased risk of drug resistance and treatment failure. Advances in next-generation sequencing enabled the detection of drug resistance mutations (DRM) among people with LLV. However, evidences remain limited in low-income and middle-income countries (LMIC) where surveillance is most needed to inform global epidemic control strategies.
To determine the prevalence of HIV DRM among people living with HIV who have LLV in low- and middle-income countries.
PubMed, Cochrane Library, and EMBASE were systematically searched for articles published between January 2015 and May 2025. Studies were included if they reported DRM among adolescents and/or adults with LLV in LMIC. Mutations were interpreted using Stanford University HIV Drug Resistance Database. Prevalence of DRM was computed as the proportion of resistance mutations among successfully sequenced samples. Pooled estimates of resistance mutation and 95%CI were calculated using random-effects models with stratified analyses comparing mutations by geographic location (Africa vs Asia).
Twenty studies including 7613 people with LLV were included. Of these, 5252 (73.9%) had their samples suc
Most individuals with LLV have resistance mutations and remain on a failed regimen over an extended period. Because resistance testing is not routinely performed in LMIC, lowering the viral failure threshold may improve timely switching to effective regimens, preserve treatment options, and reduce resistance accumulation in high HIV burden regions.
Core Tip: Next-generation sequencing enabled successful sequencing and determined drug resistance mutations (DRM) among people with low-level viremia (LLV). Data on the DRM among people with LLV in low-income and middle-income countries (LMIC) is limited and may help achieve human immunodeficiency virus epidemic control. This systematic review and meta-analysis analyzed 7613 people living with human immunodeficiency virus who had LLV from 20 studies conducted in LMIC between 2015 and 2025. Overall, the pooled prevalence of DRM was 50.4%. Pooled prevalence of nucleoside reverse transcriptase inhibitor, non-nucleoside reverse transcriptase inhibitor and protease inhibitor-associated mutations were 44.6%, 50.9% and 5.1% respectively. In LMIC, most patients with LLV have resistance mutations and remain on a failed regimen over an extended period. Because resistance testing is not routinely performed in LMIC, lowering the viral failure threshold may hasten patients switch to effective drugs.
- Citation: El-Imam IA, Chenya UK, Mbishi JV, Peter TA, Mpimo BK, Ndembi N, Bakari HM, Mbwana MS, Fussi HF, Ally HM, Dababneh SW, Ramadhani HO. Characterization of human immunodeficiency virus drug-resistance mutations among individuals with low-level-viremia in low-income and middle-income countries: A meta-analysis. World J Virol 2026; 15(1): 114375
- URL: https://www.wjgnet.com/2220-3249/full/v15/i1/114375.htm
- DOI: https://dx.doi.org/10.5501/wjv.v15.i1.114375
Over the past three decades, the global response to human immunodeficiency virus (HIV) has yielded remarkable progress, with expanded access to antiretroviral therapy (ART), widespread adoption of routine viral load (VL) monitoring, and substantial reduction in acquired immunodeficiency syndrome-related mortality. As of 2024, nearly 77% of the 40.8 million people living with HIV (PLWH) were receiving ART, and 73% had achieved viral suppression, a major global health milestone[1]. Despite this gain of expanded ART uptake and VL suppression, progress remains uneven across regions, particularly in low-income and middle-income countries (LMIC), which account for over two-thirds of all PLWH and face persistent threats from geographical instability, funding disruption, and weak health system[1,2].
Within this context, low-level viremia (LLV) has emerged as a clinically significant challenge. Although the World Health Organization (WHO) does not operationalize LLV as a formal treatment category in its current guidelines, it recognizes LLV as detectable plasma VL between 50 copies/mL and 999 copies/mL in individuals on ART[2]. Several studies and national programs similarly define LLV within this range with varying sub-categories[3,4]. In contrast, guidelines from high-income countries often define virologic failure (VF) at lower VL thresholds ranging from > 50 copies/mL to > 200 copies/mL[5-7]. This lack of consensus complicates cross-study comparisons and global surveillance. LLV, once considered biologically benign, has been linked with higher risk of subsequent VF, immunologic deterioration, onward HIV transmission and increased mortality, particularly when persistent[8,9]. Persistent LLV unlike transient “blips”, reflects ongoing viral replication under drug pressure and may signal suboptimal treatment efficacy[10]. Crucially, persistent LLV creates a selective environment for the emergence of HIV drug resistance mutations (DRM), particularly in individuals with sub-optimal adherence or exposure to partially suppressive ART regimen[11,12]. These DRMs often remain undetected in LMICs, where genotypic resistance testing is typically reserved for individuals with VL < 1000 copies/mL due to cost, infrastructure limitations, and policy constraints[2].
This diagnostic gap has important implications for treatment outcomes and population level transmission. Emerging and multi drug resistant strains not only drive VF, but also increase the risk of disease progression and mortality in resource limited settings where resistance testing and VL monitoring is lacking[13,14]. The predominance of non-B HIV subtypes in these regions adds further complexity, as such subtypes may follow distinct mutational pathway and exhibits varying susceptibility to antiretroviral agent, complicating resistance interpretation[15-17].
Despite emerging observational data, no systematic review has synthesized the burden and mutational patterns of HIV DRM during LLV in LMICs. This gap hinders efforts to refine VL thresholds for resistance testing and guide ART policy. Advanced technologies like next generation sequencing could improve detection at lower viremia levels but remain inaccessible in most LMICs due to costs and infrastructure gaps. Addressing this knowledge gap is essential to improve clinical decision-making and sustain global HIV control in high-burden, resource limited settings.
This meta-analysis aims to characterize the prevalence, mutational profiles, and clinical implications of HIV DRMs among PLWH experiencing LLV in LMIC. By synthesizing available evidence from real-world, resource-constrained contexts. This meta-analysis evaluated the appropriateness of current VL monitoring thresholds and explore the potential utility of resistance testing during LLV in informing timely decision-making and global policy recommendations.
This systematic review protocol was registered in the International Prospective Registry of Systematic Review with registration number CRD420251081973.
Ethical approval was not required as the review utilized only published data.
PubMed, Cochrane CENTRAL, EMBASE and clinicaltrials.gov were systematically searched for articles published between January 2015 and May 2025. Search terms were used to capture information on LLV, DRM, among PLWH in LMIC. The searches were restricted to papers published in English. Search results were uploaded to Covidence Systematic Review Software (Melbourne, Australia), where deduplication and screening was performed.
Eligibility criteria were guided by the guidelines of systematic reviews and meta-analysis with prevalence approach (CoCoPop)[18]. The CoCoPoP acronym stands for condition/problem, context and study population. In this review, under condition, we included studies that reported prevalence of DRM among PLWH who had low level viremia; context, we explored components of different studies that could explain variation in the reported prevalence of DRM among PLWH who had low level viremia such as study designs, number of study sites (single vs multiple sites), country of origin; population, we included PLWH who had low level viremia.
Studies that involved PLWH who had LLV, underwent DRM testing in LMIC and written in English were included in the study.
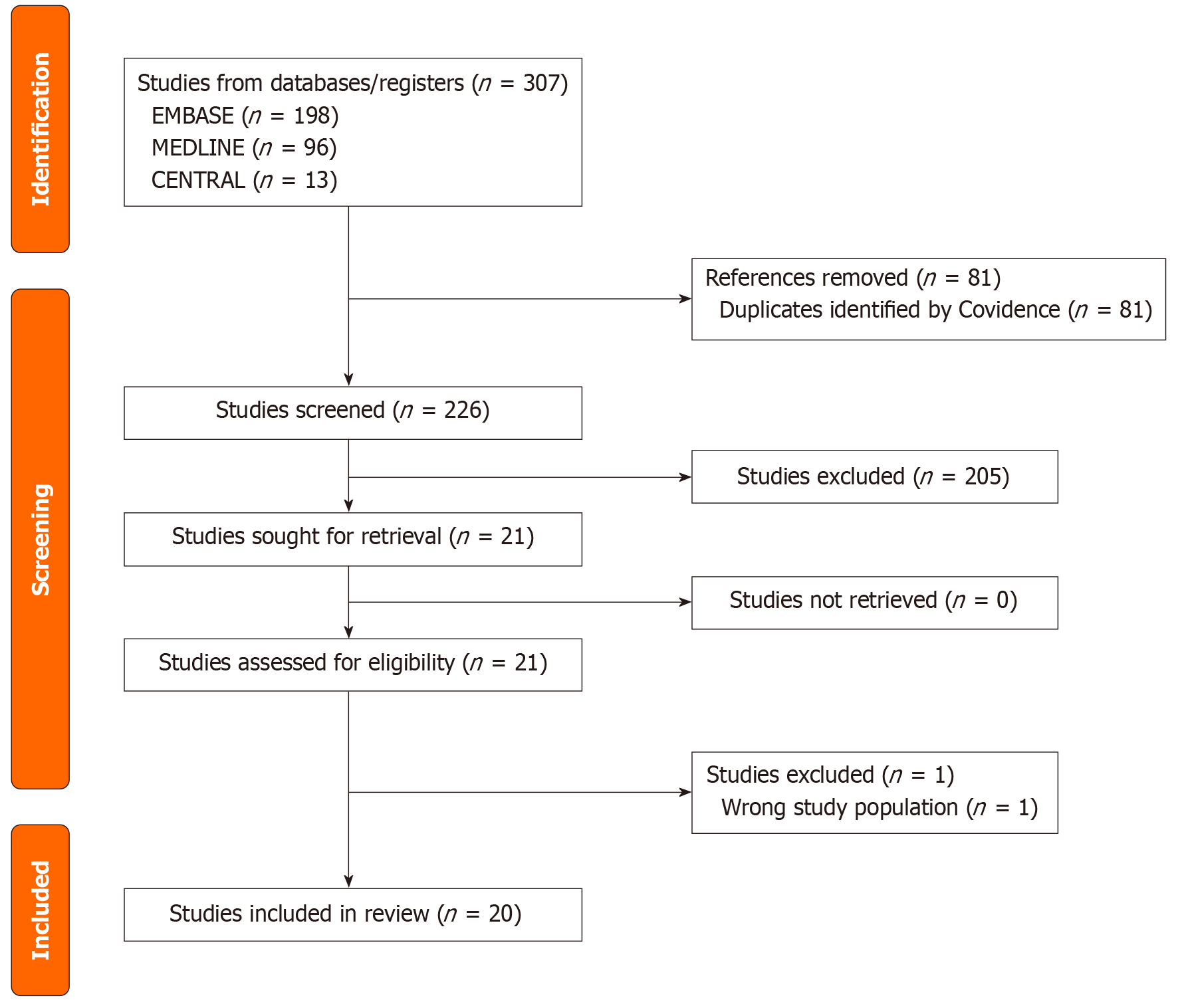
Studies that reported the percentage of individuals who underwent DRM testing without actual numerators and denominators used to compute those percentages. Studies that were not written in English. Figure 1 describes the literature search process of all included studies as shown below. The reporting of this systematic review and meta-analysis was done according to the Preferred Reporting Items for Systematic Reviews and Meta-Analyses guidelines[19].
After the duplication in Covidence, two pairs of reviewers (El-Imam IA, Mpimo BK, Ally HM, and Dababneh SW) independently screened titles/abstracts and assessed full texts for eligibility. Disagreements were resolved through discussion or adjudicated by a third reviewer pair (Fussi HF and Ramadhani HO). Using a pre-specified excel spreadsheet template, the same team independently abstracted the following data from the final list of manuscripts; authors, year of publication, the country in which the study was conducted, the year(s) in which data was collected, study design, study sites (single vs multi sites), number of people with low level viremia, number of people who were successfully sequenced, number of people who had DRM to nucleoside reverse transcriptase inhibitor (NRTI), non-NRTI (NNRTI), protease inhibitor (PI) and integrase strand transfer inhibitor (INSTI). Thereafter, extracted data was compared from two pairs of data abstractor, any discrepancies between the two pairs of reviewers were handled by consensus; and the third pair of reviewers (Fussi HF and Ramadhani HO) was consulted when necessary. Strategies in the Cochrane Handbook for Systematic Reviews of Interventions for data management were followed[20].
Study quality was assessed using the Joanna Briggs Institute checklists for cross-sectional and cohort studies[21]. Each study was rated on nine questions with responses coded as: (1) Yes; (2) No; (3) Not clear; and (4) Not applicable. A score of 1 was assigned to a “yes” response and 0 to a “no”, or “not clear” response. The total score was summed up and cate
Outcome variables: The main outcome of interest was the prevalence of DRM defined as percentage of PLWH with LLV who had confirmed DRMs among those who were successfully sequenced for resistance testing. The secondary outcomes were the prevalence of DRM to NRTI, NNRTI, PI and INSTI.
Exposure variables: This was a prevalent study and therefore no main exposure of interest. Although there was no main exposure of interest, the prevalence of DRM was compared between studies conducted from Africa vs Asia and those conducted from single vs multiple sites to understand disparities of DRM by continent and site.
Random effects models were used to compute pooled prevalence of DRM and Freeman-Tukey double arcsine transformation was used to stabilize variance of proportions prior to the computation of pooled estimates. Stabilization of variance adjusts for potential biases and safeguard the robustness of our meta-analysis. In addition, stabilization of variance allows for the inclusion of all studies, including those with proportions at the boundaries, facilitating the computation of confidence intervals that remain within permissible bounds. Subgroup analysis on the pooled estimates of DRM was performed to compare studies done in Africa vs Asia and those conducted from single vs multiple sites. Two sample tests for proportions were used for comparison in these subgroup analyses. We evaluated heterogeneity across studies using the I2 statistic and Cochran’s Q test. The I2 statistics explain the variance attributable to study heterogeneity with scores of 75%, 50% and 25% indicating high, moderate and low heterogeneity, respectively[22]. Publication bias was assessed through funnel plots and the Egger regression asymmetry test (P < 0.05 implies publication bias and heterogeneity). Furthermore, proactive comparison of the study protocol and final results was done to ensure all that was deemed to be reported from the protocol has been reported in the manuscripts. An influential analysis using leave-one-out method was used to identify potential sources of heterogeneity[23]. Studies with missing information, such as those that reported proportions of outcomes without actual numerators and/or denominators, were excluded from the analysis. All statistical tests were performed using STATA version 18 (Stata Corporation, College Station, TX, United States).
A total of 307 articles were identified through databases searches (EMBASE: 198, MEDLINE: 96, and CENTRAL: 13). Of these, 81 articles were duplicates and deleted. The remaining 226 articles were eligible for title and abstract screening. Of the 21 articles eligible for full text review, 20 articles met inclusion criteria and were finally included in our analysis (Figure 1).
All included studies were assessed as high quality using the Joanna Briggs Institute critical appraisal tool (Table 1)[3,12,21-38]. Studies with scores less than 9 typically had smaller sample sizes but met all other methodological standards.
| Ref. | Study design | Continent | Number of sites | Low level viremia definition | Quality assessment |
| Bangalee et al[24], 2021 | - | Africa | Single | 51-999 | 8 |
| Bareng et al[3], 2022 | Randomized clinical trial | Africa | Single | 51-999 | 9 |
| Brown et al[26], 2021 | Randomized clinical trial | Africa | Single | 100-999 | 7 |
| Djiyou et al[20], 2023 | Cross sectional | Africa | Single | 200-999 | 7 |
| Kao et al[21], 2021 | Cross sectional | Asia | Single | 20-999 | 7 |
| Liu et al[12], 2024 | - | Asia | Single | 50-999 | 8 |
| Kantor et al[27], 2018 | Cross sectional | Africa | Single | 40-999 | 7 |
| Mundo et al[28], 2024 | Cross sectional | Africa | Multiple | < 1000 | 8 |
| Yuan et al[23], 2022 | Cross sectional | Asia | Single | < 1000 | 9 |
| Lan et al[29], 2023 | Cross sectional | Asia | Single | 200-999 | 9 |
| Li et al[30], 2022 | Cohort | Asia | Single | 50-999 | 7 |
| Rupérez et al[31], 2015 | Cross sectional | Africa | Single | 150-999 | 7 |
| Liu et al[32], 2018 | Cross sectional | Asia | Single | 50-999 | 8 |
| Bareng et al[25], 2022 | Randomized control trial | Africa | Single | 400-999 | 7 |
| Chenwi et al[33], 2024 | Cross sectional | Africa | Single | < 1000 | 8 |
| Choga et al[34], 2025 | Cross sectional | Asia | Multiple | 200-999 | 7 |
| Shu et al[35], 2025 | Cross sectional | Asia | Single | 50-999 | 9 |
| Cao et al[36], 2023 | Cross sectional | Asia | Single | 50-999 | 7 |
| Liu et al[37], 2022 | Cross sectional | Asia | Multiple | 50-999 | 9 |
| Labhardt et al[38], 2015 | Cohort | Africa | Single | 80-999 | 7 |
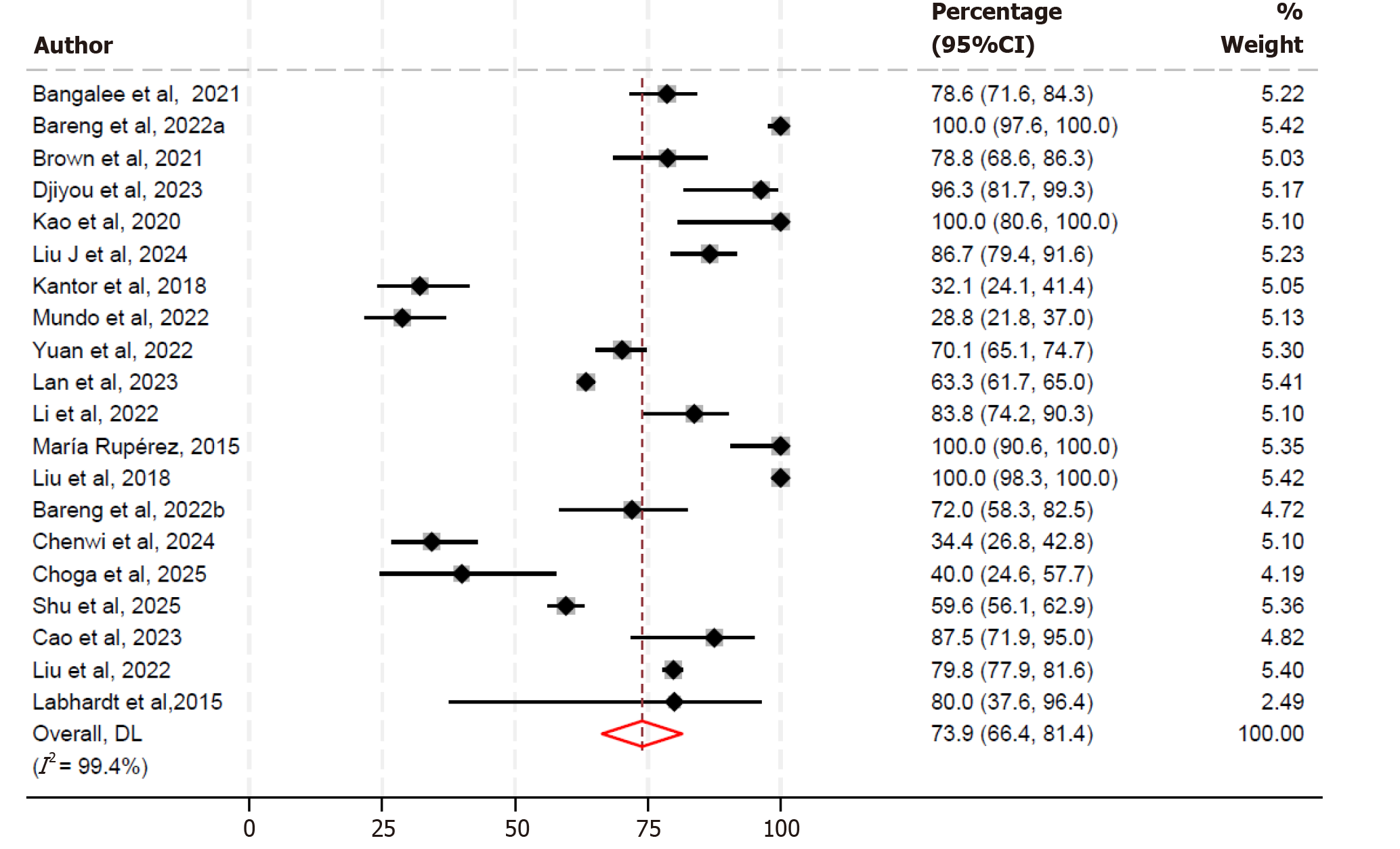
Sequencing success rate: A total of 20 studies reported on sequencing outcomes among PLWH experiencing LLV[3,12,20,21,23-38]. The random effects model estimated that 73.9% (95%CI: 66.4-81.4) of them were successfully sequenced (Figure 2)[3,12,20,21,23-38]. This indicates that, on average, nearly three-quarters of people with LLV had their VL samples successfully sequenced. Heterogeneity was observed (I² = 99.4%), reflecting substantial variability in sequencing success rates across studies.
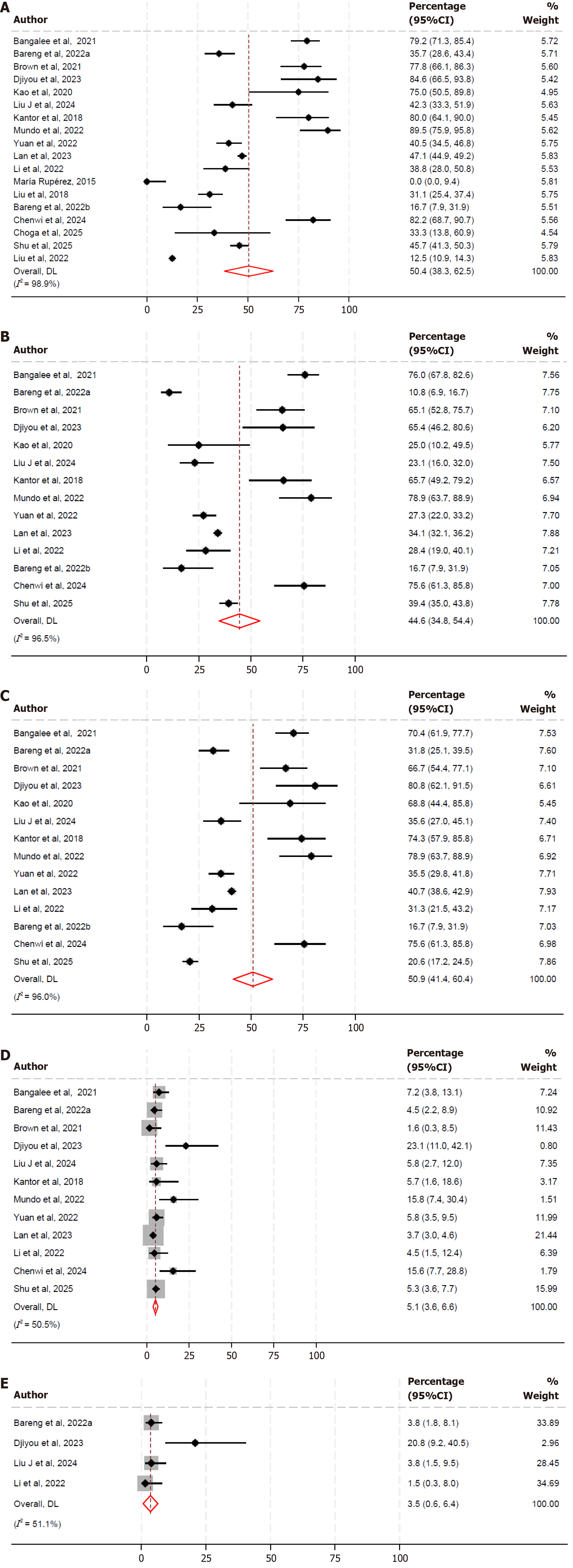
Pooled prevalence of DRM: The pooled prevalence of DRM among people with LLV was estimated at 50.4% (95%CI: 38.3-62.5) using a random-effects model presented in Figure 3A[3,12,20,21,23-35,37]. Subgroup analysis by site showed a pooled prevalence of 41.7% (95%CI: 11.0-72.3) in multi-site studies and 52.8% (95%CI: 39.3-66.3) in single-site studies (Table 2). When stratified by continent, the pooled prevalence was 58.0% (95%CI: 30.8-85.2) in studies conducted in Africa and 40.7% (95%CI: 26.0-55.5) in studies conducted in Asia (Table 2). Heterogeneity was considerable across studies (I² = 98.9%) and remained considerable even after subgroup analyses by site and continent. Leave-one-out sensitivity analysis showed that no single study substantially influenced the overall pooled estimate, as all P values remained statistically significant and pooled effect sizes were consistent across iterations (Supplementary Figure 1)[3,12,20,21,23-35,37].
| Outcome | Continent | Number of sites | ||||
| Africa | Asia | P value | Single | Multiple | P value | |
| Overall | 58.0 (30.8-85.2) | 40.7 (26.0-55.5) | < 0.0001 | 52.8 (39.3-66.3) | 41.7 (11.0-72.3) | < 0.0001 |
| Nucleoside reverse transcriptase inhibitor mutation | 57.9 (53.9-61.9) | 34.7 (33.3-36.1) | < 0.0001 | 47.3 (42.2-52.4) | 16.7 (7.4-30.4) | < 0.0001 |
| Non-nucleoside reverse transcriptase inhibitor mutation | 61.6 (44.5-78.8) | 36.2 (26.4-46.1) | < 0.0001 | 48.7 (39.3-58.1) | 78.9 (63.7-88.9) | < 0.0001 |
| Protease inhibitors mutation | 7.3 (3.4-11.2) | 4.1 (3.4-4.9) | < 0.0001 | 4.9 (3.5-6.3) | 15.8 (7.4-30.4) | < 0.0001 |
Pooled prevalence of DRM due to NRTIs: The random effects model estimated the pooled prevalence of DRM due to NRTIs at 44.6% (95%CI: 34.8-54.4) as shown in Figure 3B[3,12,20,21,23-30,33,35]. Subgroup analysis by study setting revealed a significantly higher prevalence in single-site studies (47.3%, 95%CI: 42.2-52.4) compared to those conducted across multiple sites (16.7%, 95%CI: 7.4-30.4; P < 0.0001; Table 2). Similarly, subgrouping by geographical region showed a higher prevalence in studies from Africa (57.9%, 95%CI: 53.9-61.9) than in Asia (34.7%, 95%CI: 33.3-36.1; P < 0.0001) as presented in (Table 2). Heterogeneity was initially substantial (I² = 96.5%) but decreased to 50.5% following subgroup analysis, indicating reduced variability between studies. Leave-one-out sensitivity analysis confirmed the stability of the findings, as no single study disproportionately influenced the overall estimate. The overall prevalence estimate was stable irrespective of sequential exclusion of each single study (Supplementary Figure 2)[3,12,20,21,23-30,33,35].
Pooled prevalence of DRM due to NNRTIs:Figure 3C presents the random effects model which estimated pooled prevalence of DRM due to NNRTIs at 50.9% (95%CI: 41.4-60.4)[3,12,20,21,23-30,33,35]. Subgroup analysis by study setting revealed a significantly lower prevalence in single-site studies (48.7%, 95%CI: 39.3-58.1) compared to multiple-site studies (78.9%, 95%CI: 63.7-88.9; P < 0.0001), as shown in (Table 2). Similarly, regional subgrouping showed a higher pooled prevalence in studies conducted in Africa (61.6%, 95%CI: 44.5-78.8) than in Asia (36.2%, 95%CI: 26.4-46.1; P < 0.0001; Table 2). Heterogeneity was initially substantial (I² = 96.0%) but decreased in the stratified analyses (I² = 0.0% for multiple-site and I² = 95.9% for single-site studies; I² = 94.8% for both regions). Leave-one-out sensitivity analysis confirmed the stability of the findings, as no single study substantially influenced the overall pooled estimate; all P values remained statistically significant and the effect sizes consistent across iterations (Supplementary Figure 3)[3,12,20,21,23-30,33,35].
Pooled prevalence of DRM due to PI drugs: As illustrated in Figure 3D, the pooled prevalence of DRM associated with PIs was estimated at 5.1% (95%CI: 3.6-6.6) using a random-effects model[3,12,20,23,24,26-30,33,35]. When stratified by study setting, prevalence was significantly higher in studies conducted across multiple sites (15.8%, 95%CI: 7.4-30.4) compared to those conducted at a single site (4.9%, 95%CI: 3.5-6.3; P < 0.0001; Table 2). A similar pattern was observed across regions, with studies from Africa reporting a higher pooled prevalence (7.3%, 95%CI: 3.4-11.2) than those from Asia (4.1%, 95%CI: 3.4-4.9; P < 0.0001; Table 2). Between-study heterogeneity was moderate overall (I² = 50.5%) and was further reduced within the subgroups, particularly among multiple-site studies (I² = 0.0%) and studies from Asia (I² = 1.8%). Finally, leave-one-out sensitivity analysis confirmed the robustness of the pooled estimate, as exclusion of individual studies did not substantially alter the effect size or statistical significance (Supplementary Figure 4)[3,12,20,23,24,26-30,33,35].
Pooled prevalence of DRM due to INSTI drugs: The pooled prevalence of DRM associated with INSTIs was estimated to be 3.5% (95%CI: 0.6-6.4) using a random-effects model (Figure 3E)[3,12,20,30]. Although moderate heterogeneity was observed (I² = 51.1%), the leave-one-out sensitivity analysis demonstrated that the overall effect estimate was stable across iterations, with each omitted study yielding consistently significant pooled estimates (Supplementary Figure 5)[3,12,20,30].
To assess the influence of studies reporting no DRM, a sensitivity analysis was conducted excluding Rupérez et al[31], which reported zero cases. Upon its exclusion, the pooled prevalence slightly increased to 53.5% (95%CI: 41.3%-65.7%) compared to 50.4% (95%CI: 38.3-65.5) when it was included. The two prevalences are comparable with considerable overlapping confidence intervals, indicating that the exclusion of this study did not substantially affect the overall estimate. Heterogeneity was similar prior and after its exclusion (98.9% vs 98.8%), indicating similar variation in true effects size across different studies prior and after its exclusion.
Egger’s regression test for small-study effects was used and it detected no significant publication bias for the pooled prevalence of successfully sequenced individuals (P = 0.1487), or overall DRM (P = 0.2338). However, significant small-study effects were detected in the analyses of DRM due to NRTIs (P = 0.0157), non-nucleoside NNRTIs (P = 0.0014), and PIs (P = 0.0245). These findings suggest the presence of publication bias in these subgroup analyses, which may influence the interpretation of the pooled estimates and should be interpreted with caution.
This meta-analysis synthesized global evidence on the prevalence of HIV DRM among PLWH who had LLV in LMIC. Approximately 74% of the VL samples from these patients were successfully sequenced. Overall, half (50.4%) of the sequenced individuals harbored at least one resistant mutation. Stratified prevalence by antiretroviral drug class revealed striking differences in DRM prevalence. DRM due to NNRTI was highest at 50.9%, followed closely by NRTI resistance at 44.6%. In contrast, PI-resistance and NSTI-resistance remained low at 5.1% and 3.5% respectively. This pattern reflects historical ART prescribing patterns in LMICs, where NNRTI-based regimens (e.g., efavirenz, nevirapine) were widely used, often in combination with NRTIs like lamivudine or tenofovir. Regional disparities were evident, with studies from Africa showing higher pooled DRM prevalence than those from Asia. Single-site studies also reported higher DRM rates than multi-sites, possibly reflecting patient complexity or differences in sequencing thresholds. Further investigation into study setting and participant characteristics is warranted.
These findings reinforce growing clinical and public health concerns that the current WHO VL threshold of < 1000 copies/mL to define virologic suppression may be insufficient to detect clinically meaningful resistance. Under the current WHO guidelines, PLWH who had LLV are regarded as virally suppressed and therefore remain in their current ART regimen without resistance testing or regimen adjustment. However, this analysis demonstrates that half of those with LLV had evidence of resistance mutations indicating that these patients remained on failing regimens. Continued low level viremia in the context of ineffective therapy increases the risk of acquiring additional mutations[37], pro
The findings of this analysis are consistent with Swenson et al[11], who reported that 30% of patients had detectable resistance during their first episode of LLV, with NRTI resistance in 28%, NNRTI resistance in 16%, and PI resistance in 7% of patients. In their study, 50% of LLV patients eventually failed treatment, and the risk of failure increased as resistance intensified. Similarly, finding from this review also reflect similar patterns reported in China by Liu et al[12], where DRMs among PLWH with LLV were identified in 42.3% of patients. NNRTI resistance was again dominant (35.6%), followed by NRTIs (23.8%), PIs (5.8%), and INSTIs (3.9%). Their identification of M184V and K65R mutations, both common in our reviewed papers, further validates the widespread circulation of key mutations in LMICs, particularly in regions with long-standing reliance on lamivudine-based and tenofovir-based regimens.
The apparent differences in prevalence DRM between African and Asian studies may reflect underlying epidemiologic and programmatic factors including poor adherence, HIV subtypes, genetic diversity, duration on treatment, and type of regimen)[39,40]. Gupta-Wright et al[14] observed that 88% of hospitalized patients in Malawi with VLS between 400-999 copies/mL had multi-class resistance, and nearly all harbored resistance to NNRTIs and at least one NRTI. Despite low prevalence of PI and INSTI resistance, their cohort experienced high mortality (25% in those with VF vs 18% in those without), reinforcing the life-threatening implication of under-detected resistance. While INSTI resistance is uncommon, it is emerging. Only four studies in this review reported INSTI resistance, likely due to more recent introduction of dolutegravir-based regimens in LMICs. Still, as seen in Swenson et al[11], and Liu et al[12], resistance to INSTIs was documented, particularly in patients with prior treatment failure or incomplete adherence. This highlights the need for prospective surveillance in LLV populations now receiving INSTI-based regimens. The disproportionately high resistance found in single-site studies (particularly for NRTIs and NNRTIs) may be attributable to their location in tertiary or referral centers, where more complex or treatment experienced patients are typically managed. However, caution is warranted in over-interpreting this trend, as few studies explicitly reported facility type or patient ART history. Future research should report study setting, ART duration, and patient-level factors to improve contextual interpretation of DRM estimates.
Due to its effectiveness, improved side effects profile and high genetic barrier to resistance mutations, between 2018 and 2019, the WHO recommended the use of dolutegravir based regimen as first line treatment in PLWH[41]. Subsequently, many countries in Asia and Africa adopted its use. Current data showed that dolutegravir based regimen uptake ranging from 66%-77%[42]. Of the 175 HIV clinics surveyed in LMIC, 90% reported adoption of dolutegravir based regimen as first line treatment[43]. Expanded use of dolutegravir based regimen would prevent development of DRM, however, increased adherence support and frequent viral monitoring should still be emphasized to minimize development of DRM and preserve limited treatment options.
Several limitations were acknowledged in this study. First, although this analysis reported DRM among PLWH who had LLV; it was not possible to distinguish whether these mutations were accumulated prior to LLV periods. The temporal relationship between DRM emergence and LLV remains unclear due to limited longitudinal data in the included studies. Second, the geographical distribution of included studies was uneven. While the review focused on LMICs, most eligible studies were from sub-Saharan Africa and Asia with noticeable gaps from Latin America, Eastern Europe, and francophone countries. Additionally, all included studies were limited to English-language publications, which may have introduced language bias. Thirdly, heterogeneity in study design, patient populations, and sequencing methodologies may have influenced the pooled estimates. For example, some studies were single-site investigations conducted at tertiary referral centers that typically manage more complex or treatment-experienced cases. These sites may report higher DRM prevalence limiting generalizability. Additionally, small-study effects were observed in some subgroup analyses, particularly for some drug-class subgroup analyses (NRTI, NNRTI, PI); therefore, those subgroup pooled estimates should be interpreted with caution. Despite these limitations, the inclusion of 20 studies from different LMICs countries resulted in a large sample size and strengthened results observed in this study. This review is important because it summarizes DRM among PLWH who had LLV and provides recommendations to consider lowering VL suppression threshold to prevent keeping people on failed regimen.
This meta-analysis demonstrates that a substantial proportion of PLWH with LLV harbors DRM particularly to NRTIs and NNRTIs regimens, placing them at risk of silent VF. Routine genotyping is largely unavailable in LMICs, and WHO current VL threshold of ≥ 1000 copies/mL may delay identification of failing regimens. These findings support a global reconsideration of suppression thresholds, particularly in resource-limited settings, to prevent accumulation of resistance and preserve remaining treatment options. Enhance adherence support, periodic VL monitoring, and surveillance of emerging dolutegravir resistance should be prioritized to safeguard ART effectiveness in low level viremia population.
We acknowledged Emilie Ludeman for systematic search of the manuscripts and their compilation into the Covidence software.
| 1. | UNAIDS. UNAIDS Global AIDS Update Geneva; Joint United Nation Programme on HIV/AIDS. Available from: https://www.unaids.org/sites/default/files/media_asset/2024-unaids-global-aids-update-summary_en.pdf. |
| 2. | World Health Organization. Consolidated guidelines on HIV prevention, testing, treatment, service delivery and monitoring: recommendations for a public health approach. Available from: https://www.who.int/publications/i/item/9789240031593. |
| 3. | Bareng OT, Moyo S, Zahralban-Steele M, Maruapula D, Ditlhako T, Mokaleng B, Mokgethi P, Choga WT, Moraka NO, Pretorius-Holme M, Mine MO, Raizes E, Molebatsi K, Motswaledi MS, Gobe I, Mohammed T, Gaolathe T, Shapiro R, Mmalane M, Makhema JM, Lockman S, Essex M, Novitsky V, Gaseitsiwe S. HIV-1 drug resistance mutations among individuals with low-level viraemia while taking combination ART in Botswana. J Antimicrob Chemother. 2022;77:1385-1395. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 21] [Article Influence: 5.3] [Reference Citation Analysis (1)] |
| 4. | Mbishi JV, Koola A, Ally HM, Ayalew BD, Sileshi RM, Hundisa MI, Rodoshi ZN, Htoo SW, Bakari HM, Ally ZM, Fussi HF, Ludeman E, Lascko T, Buyu CA, Ramadhani HO. Impact of low-level viremia on HIV non-viral load suppression in low and middle-income countries. Ann Med Surg (Lond). 2025;87:3777-3785. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 3] [Reference Citation Analysis (1)] |
| 5. | Abdulahi IJ, Björkman P, Abdissa A, Medstrand P, Reepalu A, Elvstam O. Low-level viremia in people with HIV in Ethiopia is associated with subsequent lack of viral suppression and attrition from care. Glob Health Action. 2025;18:2464342. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (1)] |
| 6. | European AIDS Clinical Society. European AIDS Clinical Society Guidelines. Available from: https://www.eacsociety.org/media/guidelines-12.0.pdf. |
| 7. | Clinicalinfo. hiv.gov. Guidelines for the Use of Antiretroviral Agents in Adults and Adolescents With HIV. Available from: https://clinicalinfo.hiv.gov/en/guidelines/adult-and-adolescent-arv. |
| 8. | McKenzie KP, Nguyen DT, Komba LB, Ketang'enyi EW, Kipiki NE, Mgeyi EN, Mwita LF. Low-level viraemia as a risk factor for virologic failure in children and adolescents living with HIV on antiretroviral therapy in Tanzania: a multicentre, retrospective cohort study. J Int AIDS Soc. 2025;28:e26474. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 4] [Reference Citation Analysis (1)] |
| 9. | Grennan JT, Loutfy MR, Su D, Harrigan PR, Cooper C, Klein M, Machouf N, Montaner JS, Rourke S, Tsoukas C, Hogg B, Raboud J; CANOC Collaboration. Magnitude of virologic blips is associated with a higher risk for virologic rebound in HIV-infected individuals: a recurrent events analysis. J Infect Dis. 2012;205:1230-1238. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 108] [Cited by in RCA: 102] [Article Influence: 7.3] [Reference Citation Analysis (1)] |
| 10. | Bernal E, Martínez-Rodríguez R, Gómez JM, Tomás C, García-Villalba E, Valero S, Muñoz Á, Alcaraz A, Díez C, García-Fraile LJ, Gómez-García T, Navarro-Marcotegui M, Alemán-Valls MR, Olalla J, Masiá M, Gutiérrez F; and Cohort of the Spanish HIV/AIDS Research Network (CoRIS). Low-level viremia linked to virological failure but not clinical events. AIDS. 2025;39:1545-1557. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 11. | Swenson LC, Min JE, Woods CK, Cai E, Li JZ, Montaner JS, Harrigan PR, Gonzalez-Serna A. HIV drug resistance detected during low-level viraemia is associated with subsequent virologic failure. AIDS. 2014;28:1125-1134. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 67] [Cited by in RCA: 82] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 12. | Liu J, Li C, Sun Y, Fu C, Wei S, Zhang X, Ma J, Zhao Q, Huo Y. Characteristics of drug resistance mutations in ART-experienced HIV-1 patients with low-level viremia in Zhengzhou City, China. Sci Rep. 2024;14:10620. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 15] [Reference Citation Analysis (1)] |
| 13. | Nyandiko W, Holland S, Vreeman R, DeLong AK, Manne A, Novitsky V, Sang F, Ashimosi C, Ngeresa A, Chory A, Aluoch J, Orido M, Jepkemboi E, Sam SS, Caliendo AM, Ayaya S, Hogan JW, Kantor R; Resistance in a Pediatric Cohort (RESPECT) Study. HIV-1 Treatment Failure, Drug Resistance, and Clinical Outcomes in Perinatally Infected Children and Adolescents Failing First-Line Antiretroviral Therapy in Western Kenya. J Acquir Immune Defic Syndr. 2022;89:231-239. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 21] [Article Influence: 5.3] [Reference Citation Analysis (1)] |
| 14. | Gupta-Wright A, Fielding K, van Oosterhout JJ, Alufandika M, Grint DJ, Chimbayo E, Heaney J, Byott M, Nastouli E, Mwandumba HC, Corbett EL, Gupta RK. Virological failure, HIV-1 drug resistance, and early mortality in adults admitted to hospital in Malawi: an observational cohort study. Lancet HIV. 2020;7:e620-e628. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 45] [Cited by in RCA: 67] [Article Influence: 11.2] [Reference Citation Analysis (1)] |
| 15. | Taylor BS, Hammer SM. The challenge of HIV-1 subtype diversity. N Engl J Med. 2008;359:1965-1966. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 78] [Cited by in RCA: 81] [Article Influence: 4.5] [Reference Citation Analysis (1)] |
| 16. | Santos AF, Soares MA. HIV Genetic Diversity and Drug Resistance. Viruses. 2010;2:503-531. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 44] [Article Influence: 2.8] [Reference Citation Analysis (1)] |
| 17. | Bhargava M, Cajas JM, Wainberg MA, Klein MB, Pant Pai N. Do HIV-1 non-B subtypes differentially impact resistance mutations and clinical disease progression in treated populations? Evidence from a systematic review. J Int AIDS Soc. 2014;17:18944. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 35] [Article Influence: 2.9] [Reference Citation Analysis (1)] |
| 18. | Munn Z, Stern C, Aromataris E, Lockwood C, Jordan Z. What kind of systematic review should I conduct? A proposed typology and guidance for systematic reviewers in the medical and health sciences. BMC Med Res Methodol. 2018;18:5. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 876] [Cited by in RCA: 660] [Article Influence: 82.5] [Reference Citation Analysis (1)] |
| 19. | Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, Shamseer L, Tetzlaff JM, Akl EA, Brennan SE, Chou R, Glanville J, Grimshaw JM, Hróbjartsson A, Lalu MM, Li T, Loder EW, Mayo-Wilson E, McDonald S, McGuinness LA, Stewart LA, Thomas J, Tricco AC, Welch VA, Whiting P, Moher D. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. 2021;372:n71. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9803] [Reference Citation Analysis (0)] |
| 20. | Djiyou ABD, Penda CI, Madec Y, Ngondi GD, Moukoko A, Eboumbou CE, Aghokeng AF. Prevalence of HIV drug resistance among adolescents receiving ART in Cameroon with low- or high-level viraemia. J Antimicrob Chemother. 2023;78:2938-2942. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 8] [Reference Citation Analysis (0)] |
| 21. | Kao SW, Liu ZH, Wu TS, Ku SW, Tsai CL, Shie SS, Huang PY, Wu YM, Hsiao YH, Chen NY. Prevalence of drug resistance mutations in HIV-infected individuals with low-level viraemia under combination antiretroviral therapy: an observational study in a tertiary hospital in Northern Taiwan, 2017-19. J Antimicrob Chemother. 2021;76:722-728. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 19] [Article Influence: 3.8] [Reference Citation Analysis (1)] |
| 22. | Higgins JP, Thompson SG, Deeks JJ, Altman DG. Measuring inconsistency in meta-analyses. BMJ. 2003;327:557-560. [RCA] [PubMed] [DOI] [Full Text] [Reference Citation Analysis (0)] |
| 23. | Yuan D, Zhou Y, Shi L, Liu Y, Lu J, Chen J, Fu G, Wang B. HIV-1 Drug Resistance Profiles of Low-Level Viremia Patients and Factors Associated With the Treatment Effect of ART-Treated Patients: A Cross-Sectional Study in Jiangsu, China. Front Public Health. 2022;10:944990. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 11] [Reference Citation Analysis (1)] |
| 24. | Bangalee A, Hans L, Steegen K. Feasibility and clinical relevance of HIV-1 drug resistance testing in patients with low-level viraemia in South Africa. J Antimicrob Chemother. 2021;76:2659-2665. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 15] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 25. | Bareng OT, Choga WT, Maphorisa ST, Seselamarumo S, Seatla KK, Mokgethi PT, Maruapula D, Mogwele ML, Ditshwanelo D, Moraka NO, Gobe I, Motswaledi MS, Makhema JM, Musonda R, Shapiro R, Essex M, Novitsky V, Moyo S, Gaseitsiwe S. HIV-1C in-House RNA-Based Genotyping Assay for Detection of Drug Resistance Mutations in Samples with Low-Level Viral Loads. Infect Drug Resist. 2022;15:7565-7576. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 10] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 26. | Brown JA, Amstutz A, Nsakala BL, Seeburg U, Vanobberghen F, Muhairwe J, Klimkait T, Labhardt ND. Extensive drug resistance during low-level HIV viraemia while taking NNRTI-based ART supports lowering the viral load threshold for regimen switch in resource-limited settings: a pre-planned analysis from the SESOTHO trial. J Antimicrob Chemother. 2021;76:1294-1298. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 16] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 27. | Kantor R, DeLong A, Schreier L, Reitsma M, Kemboi E, Orido M, Obonge S, Boinett R, Rono M, Emonyi W, Brooks K, Coetzer M, Buziba N, Hogan J, Diero L. HIV-1 second-line failure and drug resistance at high-level and low-level viremia in Western Kenya. AIDS. 2018;32:2485-2496. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 35] [Article Influence: 4.4] [Reference Citation Analysis (1)] |
| 28. | Mundo RAN, Abba A, Angong G, Christelle KA, Chenwi CA, Djupsa S, Fokam J, Ndjolo A, Semengue EN, Durand A, Togna WLP, Perno CF, Takou D. HIV drug resistance at low-level viremia: An appeal for revision of the viral suppression threshold in cameroon. In: Sexually transmitted diseases. Philadelphia: Mundo, 2024: S169. |
| 29. | Lan Y, Ling X, Deng X, Lin Y, Li J, Li L, He R, Cai W, Li F, Li L, Hu F. Drug Resistance Profile Among HIV-1 Infections Experiencing ART with Low-Level Viral Load in Guangdong China During 2011-2022: A Retrospective Study. Infect Drug Resist. 2023;16:4953-4964. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 16] [Reference Citation Analysis (1)] |
| 30. | Li Q, Yu F, Song C, Zhao H, Xiao Q, Lao X, Yang S, Tang Y, Zhang F. HIV-1 Genotypic Resistance Testing Using Sanger and Next-Generation Sequencing in Adults with Low-Level Viremia in China. Infect Drug Resist. 2022;15:6711-6722. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 12] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 31. | Rupérez M, Pou C, Maculuve S, Cedeño S, Luis L, Rodríguez J, Letang E, Moltó J, Macete E, Clotet B, Alonso P, Menéndez C, Naniche D, Paredes R. Determinants of virological failure and antiretroviral drug resistance in Mozambique. J Antimicrob Chemother. 2015;70:2639-2647. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 48] [Article Influence: 4.4] [Reference Citation Analysis (1)] |
| 32. | Liu PT, Xing H, Liao LJ, Leng XB, Wang J, Kan W, Yan J, Zuo ZB, Ruan YH, Shao YM. [Study on the relationship between HIV drug resistance and CD4(+)T cell counts among antiretroviral therapy patients with low viral load]. Zhonghua Yu Fang Yi Xue Za Zhi. 2018;52:277-281. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (1)] |
| 33. | Chenwi CA, Nayang Mundo RA, Nka AD, Semengue ENJ, Beloumou GA, Ka'e AC, Togna Pabo WL, Takou D, Abba A, Djupsa SC, Molimbou E, Etame NK, Kengni Ngueko AM, Same DK, Bouba Pamen JN, Abah Abah AS, Billong SC, Ajeh Awoh R, Halle-Ekane GE, Cappelli G, Njom-Nlend AE, Zk Bissek AC, Temfack E, Santoro MM, Ceccherini-Silberstein F, Colizzi V, Kaseya J, Ndembi N, Ndjolo A, Perno CF, Fokam J. Plasma Viral Load of 200 Copies/mL is a Suitable Threshold to Define Viral Suppression and HIV Drug Resistance Testing in Low- and Middle-Income Countries: Evidence From a Facility-Based Study in Cameroon. J Int Assoc Provid AIDS Care. 2024;23:23259582241306484. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 34. | Choga OT, Lemogang GM, Choga WT, Muzanywa G, Shadreck TM, Ralegoreng C, Maruapula D, Moraka NO, Koofhethile CK, Mokgethi PT, Seru K, Zuze BJL, Montshosi P, Gobe I, Motswaledi MS, Musonda R, Mbulawa MB, Makhema J, Shapiro R, Lockman S, Chebani T, Nawa J, Bochena L, Moyo S, Gaseitsiwe S. High prevalence of reverse transcriptase inhibitors associated resistance mutations among people living with HIV on dolutegravir-based antiretroviral therapy in Francistown, Botswana. J Antimicrob Chemother. 2025;80:767-776. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 4] [Reference Citation Analysis (0)] |
| 35. | Shu Y, Liu J, Yang C, Li J, Zhang M, Li Y, Deng X, Dong X. Prevalence of drug resistance mutations in low-level viremia patients under antiretroviral therapy in Southwestern China: a cross-sectional study. J Antimicrob Chemother. 2025;80:947-954. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 36. | Cao B, Liu M, Jiang T, Yu Q, Yuan T, Ding P, Zhou X, Huang F, Huang Y, Jiang J. HIV-1 RNA and DNA Genotyping Drug Resistance Detection in Patients with Low-Level Viremia in Liangshan, China. AIDS Res Hum Retroviruses. 2023;39:429-435. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 6] [Reference Citation Analysis (1)] |
| 37. | Liu P, You Y, Liao L, Feng Y, Shao Y, Xing H, Lan G, Li J, Ruan Y, Li D. Impact of low-level viremia with drug resistance on CD4 cell counts among people living with HIV on antiretroviral treatment in China. BMC Infect Dis. 2022;22:426. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 15] [Reference Citation Analysis (0)] |
| 38. | Abstracts of the 9th European Congress on Tropical Medicine and International Health, 6-10 September 2015, Basel, Switzerland. Trop Med Int Health. 2015;20 Suppl 1:1-441. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 4] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 39. | Lu X, Zhao H, Zhang Y, Wang W, Zhao C, Li Y, Ma L, Cui Z, Chen S. HIV-1 drug-resistant mutations and related risk factors among HIV-1-positive individuals experiencing treatment failure in Hebei Province, China. AIDS Res Ther. 2017;14:4. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 27] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 40. | Khodadad N, Hashempour A, Nazar MMKA, Ghasabi F. Evaluating HIV drug resistance in the middle East and North Africa and its associated factors: a systematic review. Virol J. 2025;22:112. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 4] [Reference Citation Analysis (1)] |
| 41. | World Health Organization. Update of recommendations on first- and second-line antiretroviral regimens. 2019. Available from: https://iris.who.int/server/api/core/bitstreams/0d7426a0-bdb1-4b0f-b2ba-fc5cfb130d58/content. |
| 42. | Brazier E, Romo ML, Ciaranello A, Odhiambo F, Pujari S, Murenzi G, Kasozi C, Kiertiburanakul S, Nsonde DM, Muyindike WR, Khol V, Lelo P, Lyamuya R, Lee MP, Nash D. Lingering sex and age disparities in dolutegravir uptake among adults with HIV: A multi-country observational cohort study. medRxiv. 2025;2025.05.23.25325682. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 43. | Zaniewski E, Skrivankova VW, Brazier E, Avihingsanon A, Wagner Cardoso S, Cesar C, Chenal H, Crabtree-Ramírez BE, Ditangco RA, Ebasone PV, Eley B, Euvrard JG, Fatti G, Huwa JM, Lelo P, Machado DM, Messou EK, Minga AK, Muleebwa J, Mundhe S, Murenzi G, Muyindike WR, Nsonde DM, Obatsa SM, Odhiambo J, Prozesky HW, Rungmaitree S, Semeere AS, Seydi M, Sipambo N, Sudjaritruk T, Technau KG, Tiendrebeogo T, Twizere C, Ballif M. Transition to dolutegravir-based ART in 35 low- and middle-income countries: a global survey of HIV care clinics. AIDS. 2024;38:2073-2085. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 8] [Article Influence: 4.0] [Reference Citation Analysis (1)] |