Published online Mar 24, 2026. doi: 10.5306/wjco.v17.i3.114744
Revised: October 21, 2025
Accepted: January 28, 2026
Published online: March 24, 2026
Processing time: 177 Days and 19 Hours
Neuroblastoma is the most common extracranial solid tumor in childhood, and its prognosis is strongly influenced by MYCN amplification. Non-invasive imaging biomarkers using deep learning (DL)-based radiomic analysis could capture whole-tumor characteristics to improve preoperative risk stratification and trea
To evaluate the performance of a DL-based strategy on contrast-enhanced com
All 103 eligible patients were included retrospectively between 2008 and 2015, and assigned into the training cohort (n = 72) from one institution and the testing cohort (n = 31) from the other institution. We extracted DL-based features on pretrained convolutional neural networks via transfer learning automatically, which were classified by a support vector machine. A DL-based signature was formed from the DL-based model with the optimal area under the receiver ope
The nomogram model for MYCN amplification represented great prediction and discriminative performance with an AUC of 0.959 and accuracy of 96.7% in the training cohort and an AUC of 0.819 and accuracy of 74.7% in the testing cohort. Predicted probabilities ≤ -1.996 was considered as non-amplified MYCN status, and predicted probabilities > -1.996 was considered as amplified MYCN status. The non-amplified and amplified MYCN status showed a significant survival difference in event-free survival, categorized by both nomogram predicted and histopathological results.
The integrated clinical-DL model achieved accurate abilities in the identification of MYCN amplification before surgery, which might optimize therapeutic strategy and improve patients’ survival in neuroblastomas.
Core Tip: We constructed a deep learning (DL)-based radiomics signature on computed tomography, which had the ability to identify MYCN amplification in neuroblastoma. Integrating the DL-based radiomics signature and clinical predictors, the nomogram model showed improvement in the prediction of MYCN amplification in neuroblastomas. The DL-based radiomics signature was found to be associated with disease-specific events of neuroblastomas significantly after radical resection.
- Citation: Yang YH, Li Y. Deep learning radiomic analysis in the prediction of MYCN status and survival outcome in children with neuroblastoma. World J Clin Oncol 2026; 17(3): 114744
- URL: https://www.wjgnet.com/2218-4333/full/v17/i3/114744.htm
- DOI: https://dx.doi.org/10.5306/wjco.v17.i3.114744
Neuroblastoma represents the highest incidence among extracranial solid tumors in childhood, deriving from the developing sympathetic nervous system[1]. The prognosis of neuroblastoma depends on various factors, such as age at diagnosis, tumor stage, and histopathological characteristics, especially MYCN amplification[2]. The presence of MYCN, in about 20% of neuroblastomas, implies high aggressiveness and poor prognosis with the necessity of systematic trea
Imaging materials have significant potential in representativeness of tumor characterization, and computed tomo
We recruited 103 patients retrospectively diagnosed with neuroblastoma who underwent radical resection between January 2008 and December 2015 in two institutions. The inclusion criteria were as follows: (1) Age less than 18 years old at diagnosis; (2) Confirmation of MYCN amplification detection by histopathological specimens; (3) Receiving radical resection; and (4) Presence of contrast-enhanced CT images before interventions. The inclusion patients from two institutions were assigned to the training cohort (72 patients) and testing cohort (31 patients), respectively. The institutional review boards of two institutions approved this retrospective study and waived the requirement to obtain written informed consent from all participants. Demographic and clinical data were collected for each patient and comprised age at diagnosis, gender, tumor stage, and histopathological results. The tumor stage was defined according to the International Neuroblastoma Risk Group Staging System (INRGSS)[10]. The differentiation results were divided into two groups: Poorly differentiated or undifferentiated neuroblastoma and differentiated neuroblastoma[11]. The CT imaging protocol parameters in neuroblastomas were listed in the Supplementary material.
The primary outcome was to identify whether there was the presence of MYCN amplification in histopathological specimens. MYCN status was measured by fluorescence in situ hybridization technique, using a dual-color probe and a chromosome 2 centromere enumeration probe as a reference (Vysis LSI N-MYC SpectrumGreen/CEP 2 SpectrumOrange Probes, Abbott Laboratories, IL, United States). The presence of MYCN amplification was defined as a fourfold increase in the MYCN signal number compared with the reference probe[4]. Event-free survival (EFS), the secondary outcome, was defined as the time from the date of operation to the date of any events, such as recurrence, metastasis, death, or the last follow-up. The follow-up schedule was performed every 3 months for the first year, then every 6 months from the second year to the fifth year, and annually thereafter. After surgery, each follow-up consisted of CT scanning of primary lesions, chest X-ray, and blood tests. The last follow-up was censored in December, 2020.
The regions of interest (ROIs) of the primary tumor were delineated by two experienced radiologists in contrast-enhanced CT images separately using ITK-SNAP software[12]. For patients absent of MYCN amplification with more than one primary lesion in CT images, the largest lesion was depicted with the consensus of two radiologists. For patients present of MYCN amplification with more than one primary lesion in CT images, the lesion found with MYCN amplification was selected for ROI delineation. The current study used expert manual ROI delineation because pre-existing, multi-institutional test-retest data were not available. With the masked delineation of ROIs for primary tumor, we adjust the size of consecutive axial slices of primary lesions into 224 mm × 224 mm for the input layer of the pre-trained CNN models using a bounding box covering the whole tumor area. The resized images with one axial slice of the maximum area as image channels would be candidate of the CNNs for feature extraction of 2-dimensional (2D) DL features. And the resized images of the whole tumor volume as image channels would be candidates for the pre-trained DL algorithms for feature extraction of 3-dimensional (3D) DL features.
Pretrained CNN architectures were applied to contrast-enhanced CT images for the extraction of 2D and 3D DL-based features, respectively. There were six pre-trained CNN algorithms used in this study, including Xception[13], VGG16[14], VGG19[14], ResNet50[15], InceptionV3[16], and InceptionResNetV2[17]. These pre-trained CNNs were commonly trained and applied to the large-scale and well-annotated ImageNet database[18]. These extracted DL-based features from different pre-trained CNNs would be used for the construction of DL-machine learning approaches to identify MYCN amplification at the next step. A detailed description of DL features extraction was shown in the Supplementary material. For ResNet50, we extracted features from several stages, including early, mid, and the last convolutional block, Res5c, and constructed DL-support vector machine (SVM) models for each layer’s features to empirically assess which layer provided the best discriminative information. Since the mechanism for the prognostication of DL-based features was not clear, considering the complexity of pre-trained CNNs, we interpreted ROIs recognized by DL-CNNs into heatmaps. Guided Gradient-weighted Class Activation Mapping visualized the output of DL-CNNs in the last convolutional layer[19], which highlighted specific subregions CNNs are concerned with for the generation of DL-based features.
We constructed DL-machine learning models for discrimination between the absence and presence of MYCN amp
All clinical variables were evaluated by univariate and multivariate logistic regression in the training cohort. The uni
All clinical variables for EFS in the training cohort were evaluated by Cox proportional hazard analysis, where the univariate analysis selected the variables with a significant P-value, entered the multivariate regression model, and the multivariate Cox regression identified the prognostic factors with significant statistical results. The Pi of all patients was categorized into the low-score and high-score groups according to the optimal cut-off points from the training cohort on EFS, respectively, via the X-tile software (version 3.6.1; Yale University School of Medicine, New Haven, CT, United States)[21]. The category with low-score Pi was considered as the absence of MYCN amplification group, and the category with high-score Pi was considered as the presence of MYCN amplification group. Survival analysis was performed between the absent and present MYCN amplification groups by Pi-predicted and histopathological results, respectively.
As for the distribution of clinicopathological parameters, we applied Fisher’s exact test or χ2 test for categorical data and non-parametric Mann-Whitney test for continuous data to identify the difference between the training and testing cohorts. For the evaluation of predictive performance of DL-SVM models, we applied the ROC curves and their value of AUC, accuracy, sensitivity, and specificity in the training and testing cohorts. The calibration performance was measured by the Hosmer-Lemeshow test and visualized by calibration plots[22]. Discrimination ability was quantified by Harrell’s concordance indices (C-index)[23]. The nomogram of the MYCN amplification-related clinical-DL model was applied, evaluated by ROC curves, and calibration plots. Survival curves with the Kaplan-Meier (KM) method were generated and compared by a 2-sided log-rank test. Variables that reached statistical significance in the univariate analysis were considered for the multivariate model. A two-tailed P-value less than 0.05 was considered statistically significant. The whole statistical analysis and graphic production were completed by Python (version 3.8) and R (version 3.6.1). All inv
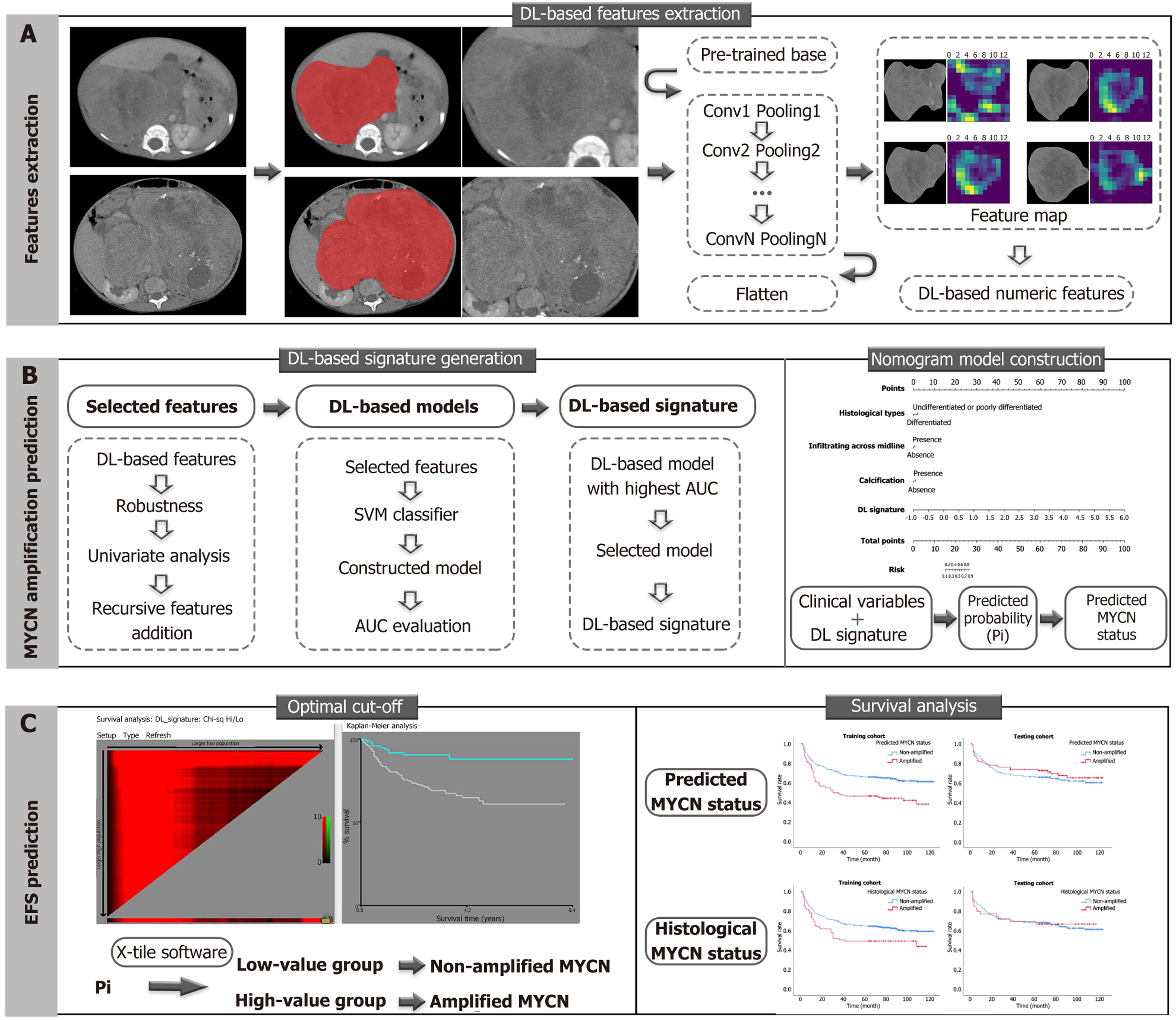
The whole scheme was depicted, including feature extraction, MYCN amplification prediction, and EFS prediction (Figure 1). A total of 103 patients were included in this study that 72 patients from one institution were assigned to the training cohort, and 31 patients from the other institution were assigned to the testing cohort. There were no significant differences found in baseline variables and clinical outcomes between the training and testing groups (P > 0.05). Demo
| Characteristic | All subjects (n = 103) | Training cohort (n = 72) | Testing cohort (n = 31) | P value |
| Age, median (IQR), months | 24 (17) | 24 (17) | 25 (18) | 0.701 |
| Gender | 0.905 | |||
| Male | 81 (78.6) | 57 (79.2) | 24 (77.4) | |
| Female | 22 (21.4) | 15 (20.8) | 7 (22.6) | |
| MYCN amplification | 0.518 | |||
| Non-amplified | 91 (88.3) | 64 (88.9) | 27 (87.1) | |
| Amplified | 12 (11.7) | 8 (11.1) | 4 (12.9) | |
| INRGSS stage | 0.706 | |||
| L1 | 47 (45.6) | 33 (45.8) | 14 (45.2) | |
| L2 | 56 (54.4) | 39 (54.2) | 17 (54.8) | |
| Histological differentiation | 0.288 | |||
| Differentiated | 38 (36.9) | 26 (36.1) | 12 (38.7) | |
| Undifferentiated or poorly differentiated | 65 (63.1) | 46 (63.9) | 19 (61.3) | |
| Infiltrating across the midline | 0.880 | |||
| Absence | 54 (52.4) | 38 (52.8) | 16 (51.6) | |
| Presence | 49 (47.6) | 34 (47.2) | 15 (48.4) | |
| Calcification | 0.960 | |||
| Absence | 32 (31.1) | 22 (30.6) | 10 (32.3) | |
| Presence | 71 (68.9) | 50 (69.4) | 21 (67.7) | |
| DL-based signature, median (IQR) | -0.243 (0.154) | -0.244 (0.153) | -0.243 (0.229) | 0.311 |
| EFS, median (IQR), month | 72 (78) | 71 (78) | 72 (78) | 0.437 |
Three significant clinical variables were selected as independent predictors for discrimination between amplified and non-amplified MYCN status in the training cohort, including histological differentiation, presence of infiltration across midline, and calcification (Supplementary Table 1). Their AUC and corresponding 95% confidence interval (CI) were calculated with 0.433 (95%CI: 0.337-0.529) for histological differentiation, 0.581 (95%CI: 0.485-0.678) for presence of infiltrating across midline, and 0.505 (95%CI: 0.408-0.603) for presence of calcification in the testing cohort (Table 2).
| Model | Training cohort (n = 72) | Testing cohort (n = 31) | ||||||
| AUC (95%CI) | Accuracy | Sensitivity | Specificity | AUC (95%CI) | Accuracy | Sensitivity | Specificity | |
| Histological differentiation | 0.560 (0.495-0.625) | 47.2 (34/72) | 62.5 (5/8) | 45.3 (29/64) | 0.433 (0.337-0.529) | 48.4 (15/31) | 25.0 (1/4) | 51.9 (14/27) |
| Infiltrating across midline | 0.603 (0.536-0.669) | 65.3 (47/72) | 62.5 (5/8) | 65.6 (42/64) | 0.581 (0.485-0.678) | 61.3 (19/31) | 50.0 (2/4) | 63.0 (17/27) |
| Calcification | 0.518 (0.450-0.587) | 88.9 (64/72) | 1.3 (1/8) | 98.4 (63/64) | 0.505 (0.408-0.603) | 87.1 (27/31) | 0.0 (0/4) | 100.0 (27/27) |
| DL-based signature | 0.958 (0.929-0.987) | 91.7 (66/72) | 87.5 (7/8) | 92.2 (59/64) | 0.803 (0.723-0.883) | 77.4 (24/31) | 75.0 (3/4) | 77.8 (21/27) |
| Nomogram model1 | 0.959 (0.930-0.988) | 95.8 (69/72) | 87.5 (7/8) | 96.9 (62/64) | 0.819 (0.740-0.898) | 74.2 (23/31) | 75.0 (3/4) | 74.1 (20/27) |
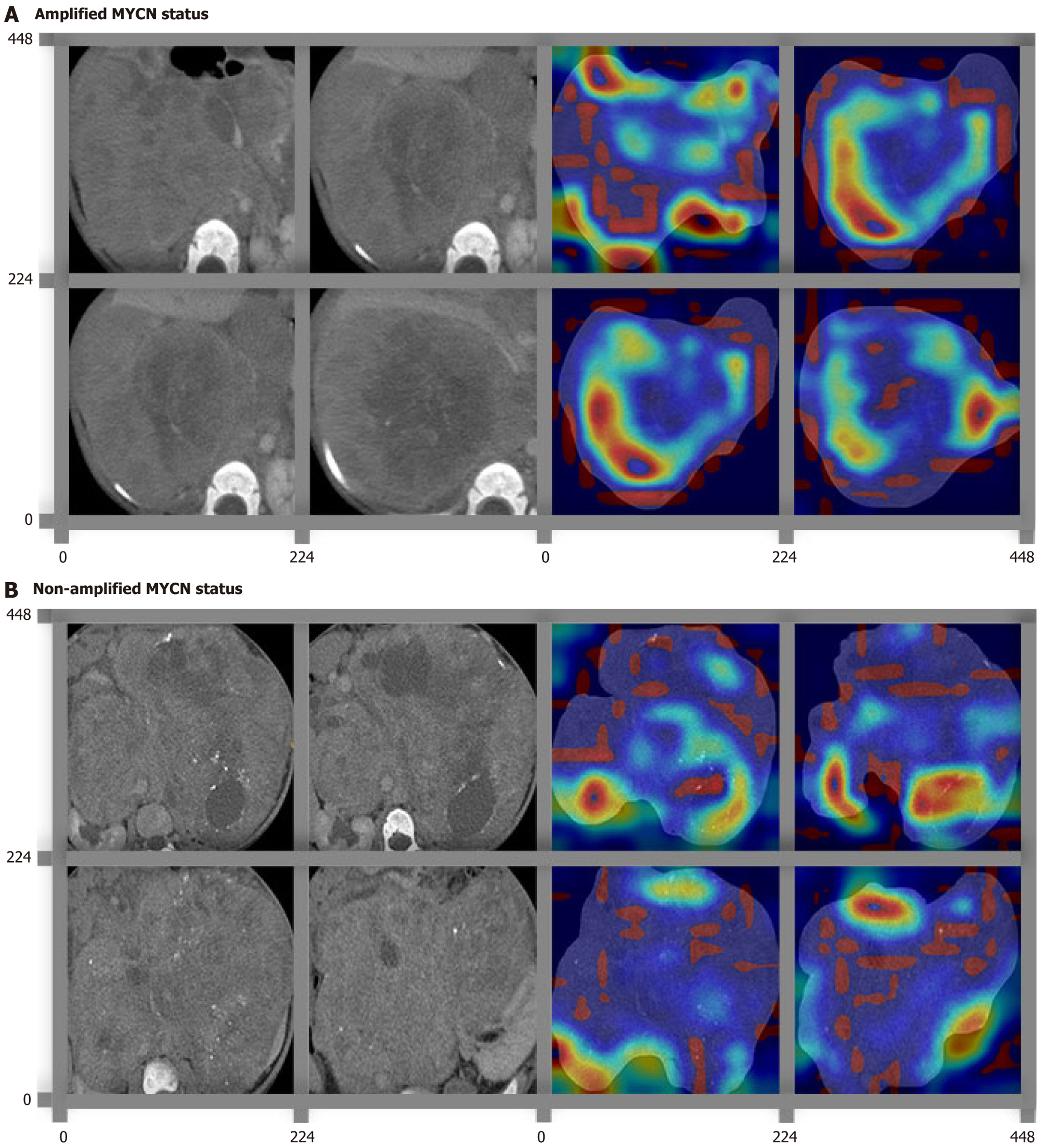
As for the performance of DL-SVM models for differentiation between amplified and non-amplified MYCN, the AUCs ranged from 0.873 to 0.966 for the training group and from 0.553 to 0.819 for the testing group (Supplementary Table 2). A total of 18 DL-SVM models, including 6 2D models, 6 3D models, and 6 combined models, were constructed based on selected features from different pre-trained DL-CNNs (Supplementary Table 3). The combined model with ResNet50 algorithm achieved the best discriminative performance considering the AUC value of the testing cohort among all DL-SVM models, with an AUC of 0.959, a C-index of 0.959, accuracy of 96.53%, sensitivity of 90.12%, and specificity of 97.34% in the training cohort, and an AUC of 0.819 (95%CI: 0.740-0.898), a C-index of 0.819, accuracy of 74.68%, sensitivity of 74.36%, and specificity of 74.72% in the testing cohort (Supplementary Table 2). For the ResNet50 CNN model, DL-based features extracted from different layers might exist differences since its complex DL-CNN architecture. We constructed DL-SVM models on DL features from earlier layers of the ResNet50 algorithm to explore the superiority of DL features from the last layer before the fully connected layer, Res5c, in the classification of amplified and non-amplified MYCN (Supplementary Table 4). The model constructed using features from the Res5c layer outperformed models built from earlier layers, as shown in Supplementary Table 4, where the DL-SVM using Res5c features achieved higher AUC and C-index in both training and testing cohorts than DL-SVMs built from earlier blocks, suggesting Res5c features capture more task-specific, high-level image patterns relevant to MYCN amplification. The results demonstrated the advantages of the current extraction strategy from the layer of Res5c in the ResNet50 algorithm. ResNet50 architecture was applied to generate feature maps highlighting important subregions for the generation of the output (Figure 2). As shown in the feature heatmaps, the valuable areas were different in different MYCN status for the extraction of specific feature pat
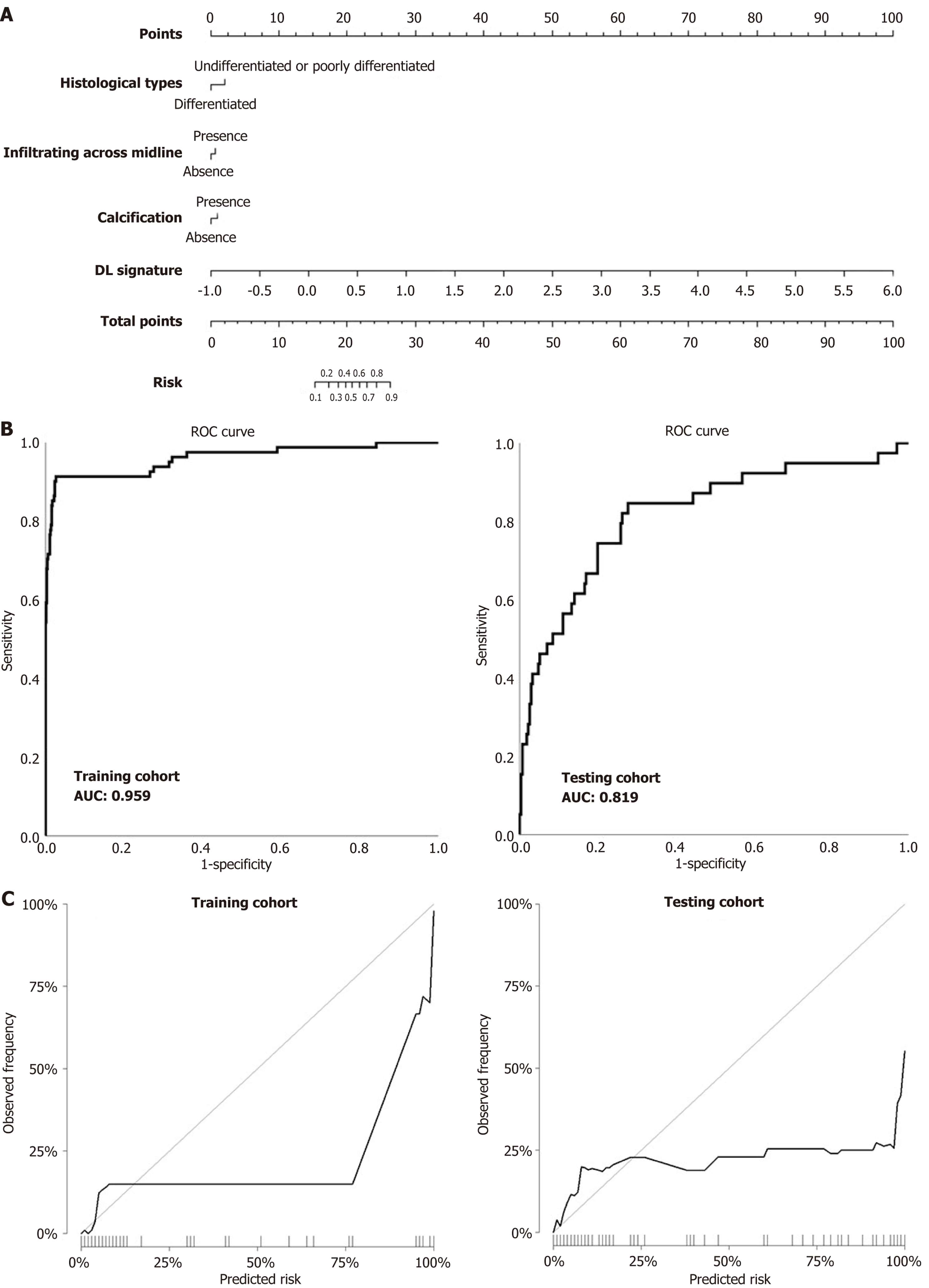
The DL-based signature was generated based on the combined ResNet50 DL-SVM model. Integrating independent clinical predictors and the DL-based signature (Figure 3A), an integrated nomogram model for prediction of histological patterns was constructed, including significant clinical variables and the DL-based signature (Figure 3A) with an AUC of 0.959 and accuracy of 96.7% in the training cohort, and an AUC of 0.819, accuracy of 74.7%, sensitivity of 74.4%, and specificity of 74.7% in the testing cohort (Table 2). This nomogram model represented great classification performance by ROC curves (Figure 3B) and consistent calibration capacities (Figure 3C) in both training and testing cohorts. The Pi derived from the nomogram model was calculated with the formula: Pi = 0.823 × Histological differentiation + 0.280 × presence of infiltrating across midline + 0.368 × presence of calcification + 5.684 × DL-based signature - 2.577.
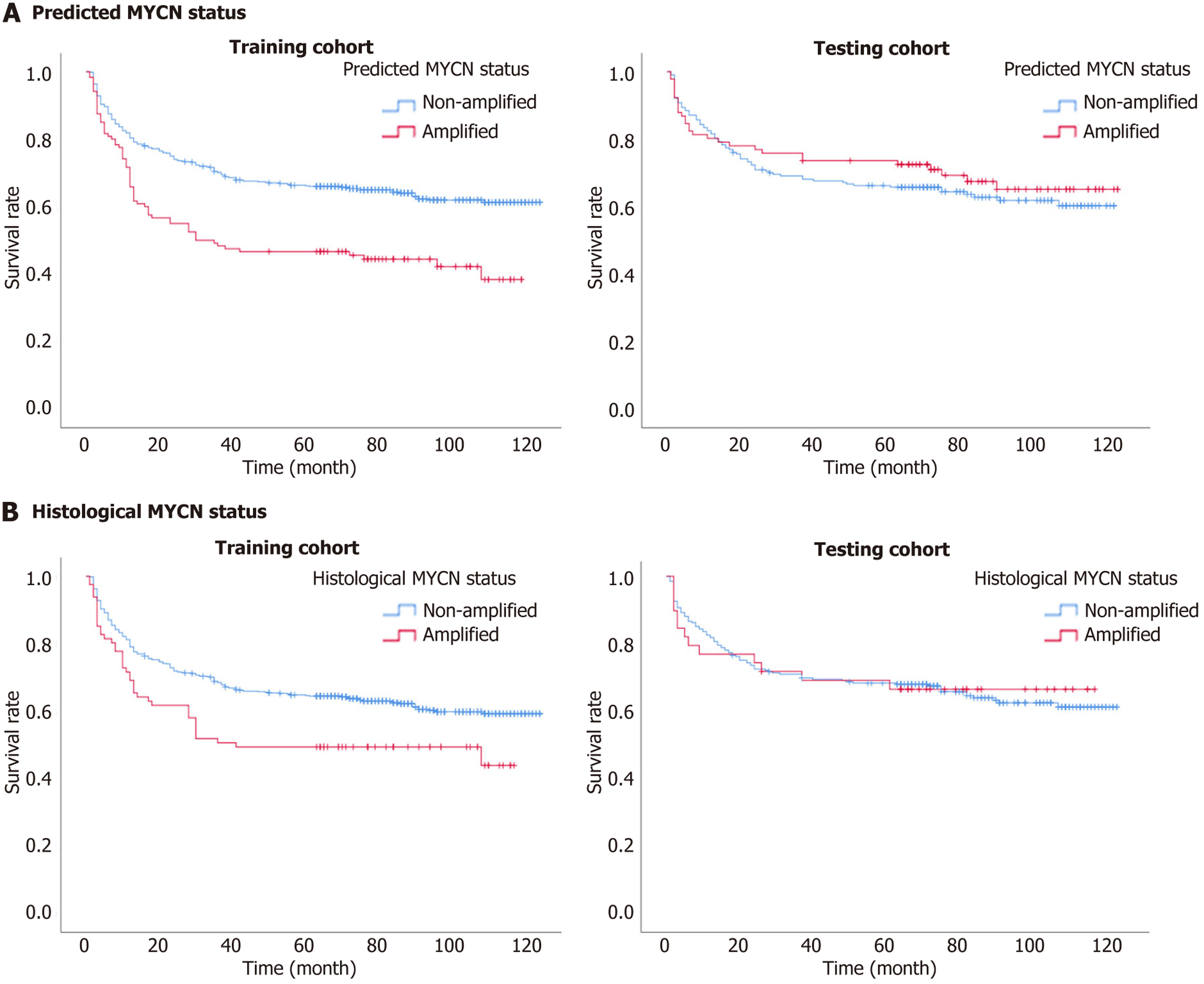
According to the results of multivariate Cox regression, age at diagnosis, INRGSS stage, histological differentiation, presence of infiltrating across midline, presence of MYCN amplification, and nomogram model-predicted probability were considered as independent predictors for EFS (Supplementary Table 5). The hazard ratio and corresponding 95%CI were calculated with 1.011 (95%CI: 1.002-1.021) for age at diagnosis, 2.674 (95%CI: 2.117-3.377) for INRGSS stage, 1.319 (95%CI: 1.046-1.665) for histological differentiation, and 1.541 (95%CI: 1.210-1.963) for presence of Infiltrating across midline (Supplementary Table 5). The nomogram model-predicted probability was stratified with the optimal cut-off points of -1.996 for recurrence-free survival via X-tile software from the training cohort. Pi ≤ -1.996 was considered as non-amplified MYCN status, and Pi > -1.996 was considered as amplified MYCN status. The KM survival analysis for the histopathological non-amplified and amplified MYCN groups showed the mean survival of 83.476 months (95%CI: 79.401-87.550 months) vs 63.938 months (52.568-75.308 months) in the training cohort, and the mean survival of 85.458 months (95%CI: 79.308-91.609 months) vs 82.538 months (66.945-98.130 months) in the testing cohort, with the P-value of 0.031 in the log-rank test (Table 3). The KM survival analysis for the model-predicted non-amplified and amplified MYCN groups showed the mean survival of 85.504 months (95%CI: 81.349-89.659 months) vs 60.239 months (50.833-69.644 months) in the training cohort, and the mean survival of 83.366 months (95%CI: 76.510-90.222 months) vs 89.230 months (78.847-99.614 months) in the testing cohort, with the P-value of 0.002 in the log-rank test (Figure 4 and Table 3).
| Variable | Subgroup | Training database | Testing database | Log rank test | ||
| Mean survival time | 95%CI | Mean survival time | 95%CI | |||
| Histological MYCN amplification1 | Non-amplified | 83.476 | 79.401-87.550 | 85.458 | 79.308-91.609 | 0.031 |
| Amplified | 63.938 | 52.568-75.308 | 82.538 | 66.945-98.130 | ||
| Predicted MYCN amplification2 | Non-amplified | 85.504 | 81.349-89.659 | 83.366 | 76.510-90.222 | 0.002 |
| Amplified | 60.239 | 50.833-69.644 | 89.230 | 78.847-99.614 | ||
In this study, we aimed to develop predictive models for MYCN amplification and survival outcomes in patients with neuroblastomas. We converted CT images into quantitative signatures via DL-based methods and constructed an inte
The correct identification of MYCN amplification before surgery is pivotal in order to establish an appropriate therapeutic strategy and to improve survival outcomes of patients with neuroblastomas. Indeed, accurate preoperative diagnosis of patients with amplified MYCN status needs multi-model approaches on clinicopathological information, which guides individualized therapeutic regimens. In the present study, the DL-based radiomics signature on contrast-enhanced CT images provided useful information for detecting MYCN amplification. Nowadays, the acquisition of histopathological specimens from needle core biopsy is the mainstay for the determination of MYCN status to assist clinicians in establishing subsequent treatment[24]. The specimens from the open or needle biopsies lack representativeness of the entire bladder cancer lesion due to the small amount of the specimen and sampling bias[5]. Previous literature showed different MYCN amplification status between needle-biopsy and resection specimens[25]. These studies reported that MYCN amplification status defined on needle core-biopsy samples was slightly concordant with evidence determined by resection specimens of the whole tumor due to the small specimen size hard to capture important and specific morphological features such as high vascularity induced by MYCN amplification[26]. The needle-biopsy samples represented little feasibility for the identification of MYCN amplification, whereas the surgical specimens might result in too much invasive damage for children[27]. Because of the inaccurate reflection of tumor characterization on insufficient samples, we need other tools to evaluate extensive regions for evidence about histological inclination on neuroblastomas, as the preoperative CT images have the ability to provide useful data on the whole tumor lesions.
The correct identification of MYCN amplification could improve the patients’ survival and quality of life. In cases of amplified MYCN neuroblastomas, patients show strong tendencies for rapid progression in local infiltration, metastasis, and relapse[8]. If any preoperative histological evidence indicated the presence of MYCN amplification, the subsequent treatment would be inclined to neoadjuvant chemotherapy rather than surgical resection for better control of local lesions[28]. The computer-aided diagnosis (CAD) on imaging data has the potential to provide information about MYCN amplification based on CT images. The application of DL-based techniques was expected to achieve great prediction performance with well concordance with findings on surgical resection of the whole tumor. In our main results, the DL-based signature in the present study achieved great discrimination abilities in the classification of non-amplified and amplified MYCN status with an AUC of 0.958 (95%CI: 0.929-0.987) in the training cohort and 0.803 (95%CI: 0.723-0.883) in the testing cohort. The integration of the DL-based signature and prognostic clinical variables had a slight impact on discriminative abilities with an AUC of 0.959 (95%CI: 0.930-0.988) in the training cohort and 0.819 (95%CI: 0.740-0.898) in the testing cohort. The DL-based imaging analysis indeed yielded strong classification capacities to differentiate patients with MYCN amplification from patient absent of MYCN amplification on preoperative CT images, which might assist in establishing individualized treatment before operation.
In the present study, we found that the presence of MYCN amplification was significantly associated with poor EFS in both nomogram-predicted and histopathological results. According to the previous studies, the impact of MYCN status was not consistent on survival outcomes of patients with neuroblastoma. The findings in early-stage neuroblastomas showed agreement with our study that VHs were significantly associated with poor EFS[29]. On the contrary, some research indicated that there existed little robust association between the presence of MYCN amplification and survival outcomes in neuroblastomas in advanced neuroblastomas[30]. The effects of MYCN amplification might have a strong association with treatments based on tumor stage. These conflicting findings underline the importance of exploring the actual impact of MYCN amplification on survival outcomes in patients with neuroblastoma. These controversial con
In this study, we expected to explore further evidence to classify MYCN amplification by our non-invasive CAD systems. The diagnostic performance of our CAD systems was reliable and cost-effective, which might be an effective complementary measure before surgery. The practicability of our DL-based CAD systems was demonstrated in solving one specific medical task with respect to the satisfactory predictive results. The actual procedure of these CAD systems was hard to interpret directly for understanding. Class activation mapping heatmaps were used to translate the important subregions recognized by CAD systems. According to Figure 2, a different area was recognized for differentiation between amplified and non-amplified MYCN status. These differences demonstrated the potential of CAD systems in the detection of spatial heterogeneity of tumoral and peritumoral microenvironment.
Some limitations existed in the present study. First and foremost, the retrospective design of the present study was companied with the presence of inherent selection bias and the absence of standardization of therapeutic protocols. Since the small testing sample, an external validation plan based on a multi-center prospective study is being conducted with pre-specified endpoints, sample size calculation, model update, and performance stability measures. Second, the ROI contour-based method might be insufficient in assurance of DL-based feature robustness and repeatability compared with the test-retest method[31]. Since no test-retest scans were available in this retrospective cohort, the robustness of DL-derived features could be affected by contour variability and acquisition differences. Our research team is trying to apply the test-retest method in a prospective study as the next step. In our planned prospective study, we will obtain dedicated test-retest CT scans on a subset of patients and use phantom experiments to quantify repeatability and derive robust DL-features using the intraclass correlation coefficient and concordance metrics, and exclude unstable features prior to model development. Third, the heterogeneity between the original pre-trained CNN bases and target databases was hard to avoid, even though we adopted transfer learning to mitigate it. A DL-based model from scratch, constructed on a large database, is an ideal solution for excellent generalizability and clinical practicability of DL-based CAD methods. Last but not least, we considered the mixed cases as amplified MYCN status once the appearance of morphological MYCN amplification occurred without quantifying the amount and percentage. In this retrospective study, mixed or focal amplification was classified as positive due to limitations of available pathology reports. An “all-or-nothing” dichotomy might misrepresent intratumoral heterogeneity and could bias model predictions and survival associations. A further comprehensive evaluation of image-to-genetic associations of neuroblastomas will be conducted for prediction of survival outcomes. In future work, we will collect detailed pathology reports with quantitative measures of amplification, including percentage of amplified cells and copy-number estimates, perform sensitivity analyses excluding focal/mixed cases to assess robustness, and explore multi-class outcomes or continuous target labels like copy-number or ampli
The DL-based nomogram has been envisioned as a preoperative decision-support and triage tool rather than an immediate replacement for histopathology. In practice the workflow would be: (1) All children undergo routine preo
In conclusion, we demonstrated that the DL-based signature showed notable advantages in the identification of MYCN amplification over existing clinical variables for neuroblastomas. The integrated nomogram model achieved a great differentiation performance for the classification of non-amplified and amplified MYCN status. The integrated diagnostic tool represented a novel manner with high efficiency and without manual tumor delineation to assist the preoperative findings and enhance the correct diagnostic rate of MYCN amplification before surgery. MYCN amplification identification could facilitate predicting tumor-associated events and optimizing treatment for patients with neuroblastoma.
| 1. | Ahmed AA, Zhang L, Reddivalla N, Hetherington M. Neuroblastoma in children: Update on clinicopathologic and genetic prognostic factors. Pediatr Hematol Oncol. 2017;34:165-185. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 61] [Cited by in RCA: 76] [Article Influence: 8.4] [Reference Citation Analysis (10)] |
| 2. | Campbell K, Gastier-Foster JM, Mann M, Naranjo AH, Van Ryn C, Bagatell R, Matthay KK, London WB, Irwin MS, Shimada H, Granger MM, Hogarty MD, Park JR, DuBois SG. Association of MYCN copy number with clinical features, tumor biology, and outcomes in neuroblastoma: A report from the Children's Oncology Group. Cancer. 2017;123:4224-4235. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 107] [Article Influence: 11.9] [Reference Citation Analysis (7)] |
| 3. | Seeger RC, Brodeur GM, Sather H, Dalton A, Siegel SE, Wong KY, Hammond D. Association of multiple copies of the N-myc oncogene with rapid progression of neuroblastomas. N Engl J Med. 1985;313:1111-1116. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1458] [Cited by in RCA: 1432] [Article Influence: 34.9] [Reference Citation Analysis (0)] |
| 4. | Ambros PF, Ambros IM, Brodeur GM, Haber M, Khan J, Nakagawara A, Schleiermacher G, Speleman F, Spitz R, London WB, Cohn SL, Pearson AD, Maris JM. International consensus for neuroblastoma molecular diagnostics: report from the International Neuroblastoma Risk Group (INRG) Biology Committee. Br J Cancer. 2009;100:1471-1482. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 345] [Cited by in RCA: 276] [Article Influence: 16.2] [Reference Citation Analysis (0)] |
| 5. | Marrano P, Irwin MS, Thorner PS. Heterogeneity of MYCN amplification in neuroblastoma at diagnosis, treatment, relapse, and metastasis. Genes Chromosomes Cancer. 2017;56:28-41. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 38] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 6. | Mullassery D, Sharma V, Salim A, Jawaid WB, Pizer BL, Abernethy LJ, Losty PD. Open versus needle biopsy in diagnosing neuroblastoma. J Pediatr Surg. 2014;49:1505-1507. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 28] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 7. | Gillies RJ, Kinahan PE, Hricak H. Radiomics: Images Are More than Pictures, They Are Data. Radiology. 2016;278:563-577. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6963] [Cited by in RCA: 6221] [Article Influence: 622.1] [Reference Citation Analysis (5)] |
| 8. | LeCun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015;521:436-444. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 70666] [Cited by in RCA: 21028] [Article Influence: 1911.6] [Reference Citation Analysis (10)] |
| 9. | Raghu S, Sriraam N, Temel Y, Rao SV, Kubben PL. EEG based multi-class seizure type classification using convolutional neural network and transfer learning. Neural Netw. 2020;124:202-212. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 142] [Cited by in RCA: 133] [Article Influence: 22.2] [Reference Citation Analysis (0)] |
| 10. | Cohn SL, Pearson AD, London WB, Monclair T, Ambros PF, Brodeur GM, Faldum A, Hero B, Iehara T, Machin D, Mosseri V, Simon T, Garaventa A, Castel V, Matthay KK; INRG Task Force. The International Neuroblastoma Risk Group (INRG) classification system: an INRG Task Force report. J Clin Oncol. 2009;27:289-297. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1625] [Cited by in RCA: 1378] [Article Influence: 81.1] [Reference Citation Analysis (0)] |
| 11. | Shimada H, Ambros IM, Dehner LP, Hata J, Joshi VV, Roald B. Terminology and morphologic criteria of neuroblastic tumors: recommendations by the International Neuroblastoma Pathology Committee. Cancer. 1999;86:349-363. [PubMed] [DOI] [Full Text] |
| 12. | Yushkevich PA, Piven J, Hazlett HC, Smith RG, Ho S, Gee JC, Gerig G. User-guided 3D active contour segmentation of anatomical structures: significantly improved efficiency and reliability. Neuroimage. 2006;31:1116-1128. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5192] [Reference Citation Analysis (0)] |
| 13. | Chollet F. Xception: Deep Learning with Depthwise Separable Convolutions. 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR): 2017. [DOI] [Full Text] |
| 14. | Simonyan K, Zisserman A. Very Deep Convolutional Networks for Large-Scale Image Recognition. Available from: arXiv:1409.1556. [DOI] [Full Text] |
| 15. | He KM, Zhang XY, Ren SQ, Sun J. Deep Residual Learning for Image Recognition. Available from: arXiv:1512.03385. [DOI] [Full Text] |
| 16. | Szegedy C, Vanhoucke V, Ioffe S, Shlens J, Wojna Z. Rethinking the Inception Architecture for Computer Vision. 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR): 2016. [DOI] [Full Text] |
| 17. | Szegedy C, Ioffe S, Vanhoucke V, Alemi A. Inception-v4, Inception-ResNet and the Impact of Residual Connections on Learning. Available from: arXiv:1602.07261. [DOI] [Full Text] |
| 18. | Russakovsky O, Deng J, Su H, Krause J, Satheesh S, Ma S, Huang Z, Karpathy A, Khosla A, Bernstein M, Berg AC, Li FF. ImageNet Large Scale Visual Recognition Challenge. Int J Comput Vis. 2015;115:211-252. [RCA] [DOI] [Full Text] [Cited by in Crossref: 16940] [Cited by in RCA: 6594] [Article Influence: 599.5] [Reference Citation Analysis (0)] |
| 19. | Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D. Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization. Int J Comput Vis. 2020;128:336-359. [RCA] [DOI] [Full Text] [Cited by in Crossref: 977] [Cited by in RCA: 1709] [Article Influence: 244.1] [Reference Citation Analysis (0)] |
| 20. | Amari S, Wu S. Improving support vector machine classifiers by modifying kernel functions. Neural Netw. 1999;12:783-789. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 659] [Cited by in RCA: 223] [Article Influence: 8.3] [Reference Citation Analysis (0)] |
| 21. | Camp RL, Dolled-Filhart M, Rimm DL. X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin Cancer Res. 2004;10:7252-7259. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3094] [Cited by in RCA: 3114] [Article Influence: 141.5] [Reference Citation Analysis (3)] |
| 22. | Vickers AJ, Cronin AM, Elkin EB, Gonen M. Extensions to decision curve analysis, a novel method for evaluating diagnostic tests, prediction models and molecular markers. BMC Med Inform Decis Mak. 2008;8:53. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1046] [Cited by in RCA: 1056] [Article Influence: 58.7] [Reference Citation Analysis (4)] |
| 23. | Schoop R, Beyersmann J, Schumacher M, Binder H. Quantifying the predictive accuracy of time-to-event models in the presence of competing risks. Biom J. 2011;53:88-112. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 52] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 24. | Hassan SF, Mathur S, Magliaro TJ, Larimer EL, Ferrell LB, Vasudevan SA, Patterson DM, Louis CU, Russell HV, Nuchtern JG, Kim ES. Needle core vs open biopsy for diagnosis of intermediate- and high-risk neuroblastoma in children. J Pediatr Surg. 2012;47:1261-1266. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 45] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 25. | Campagna G, Rosenfeld E, Foster J, Vasudevan S, Nuchtern J, Kim E, Commander S, Naik-Mathuria B. Evolving biopsy techniques for the diagnosis of neuroblastoma in children. J Pediatr Surg. 2018;53:2235-2239. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 29] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 26. | Huang M, Weiss WA. Neuroblastoma and MYCN. Cold Spring Harb Perspect Med. 2013;3:a014415. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 341] [Cited by in RCA: 526] [Article Influence: 40.5] [Reference Citation Analysis (0)] |
| 27. | Mohamed H, Pastor MC, Langlois S, Cowan KN. Comparing safety and adequacy between surgical biopsy versus core needle biopsy in diagnosing neuroblastoma. J Pediatr Surg. 2022;57:866-870. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 8] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 28. | Yanishevski D, McCarville MB, Doubrovin M, Spiegl HR, Zhao X, Lu Z, Federico SM, Furman WL, Murphy AJ, Davidoff AM. Impact of MYCN status on response of high-risk neuroblastoma to neoadjuvant chemotherapy. J Pediatr Surg. 2020;55:130-134. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 16] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 29. | Bagatell R, Beck-Popovic M, London WB, Zhang Y, Pearson AD, Matthay KK, Monclair T, Ambros PF, Cohn SL; International Neuroblastoma Risk Group. Significance of MYCN amplification in international neuroblastoma staging system stage 1 and 2 neuroblastoma: a report from the International Neuroblastoma Risk Group database. J Clin Oncol. 2009;27:365-370. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 93] [Cited by in RCA: 101] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 30. | Cohn SL, London WB, Huang D, Katzenstein HM, Salwen HR, Reinhart T, Madafiglio J, Marshall GM, Norris MD, Haber M. MYCN expression is not prognostic of adverse outcome in advanced-stage neuroblastoma with nonamplified MYCN. J Clin Oncol. 2000;18:3604-3613. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 78] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 31. | Zwanenburg A, Leger S, Agolli L, Pilz K, Troost EGC, Richter C, Löck S. Assessing robustness of radiomic features by image perturbation. Sci Rep. 2019;9:614. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 97] [Cited by in RCA: 189] [Article Influence: 27.0] [Reference Citation Analysis (3)] |