Copyright: ©Author(s) 2026.
World J Clin Oncol. Mar 24, 2026; 17(3): 114744
Published online Mar 24, 2026. doi: 10.5306/wjco.v17.i3.114744
Published online Mar 24, 2026. doi: 10.5306/wjco.v17.i3.114744
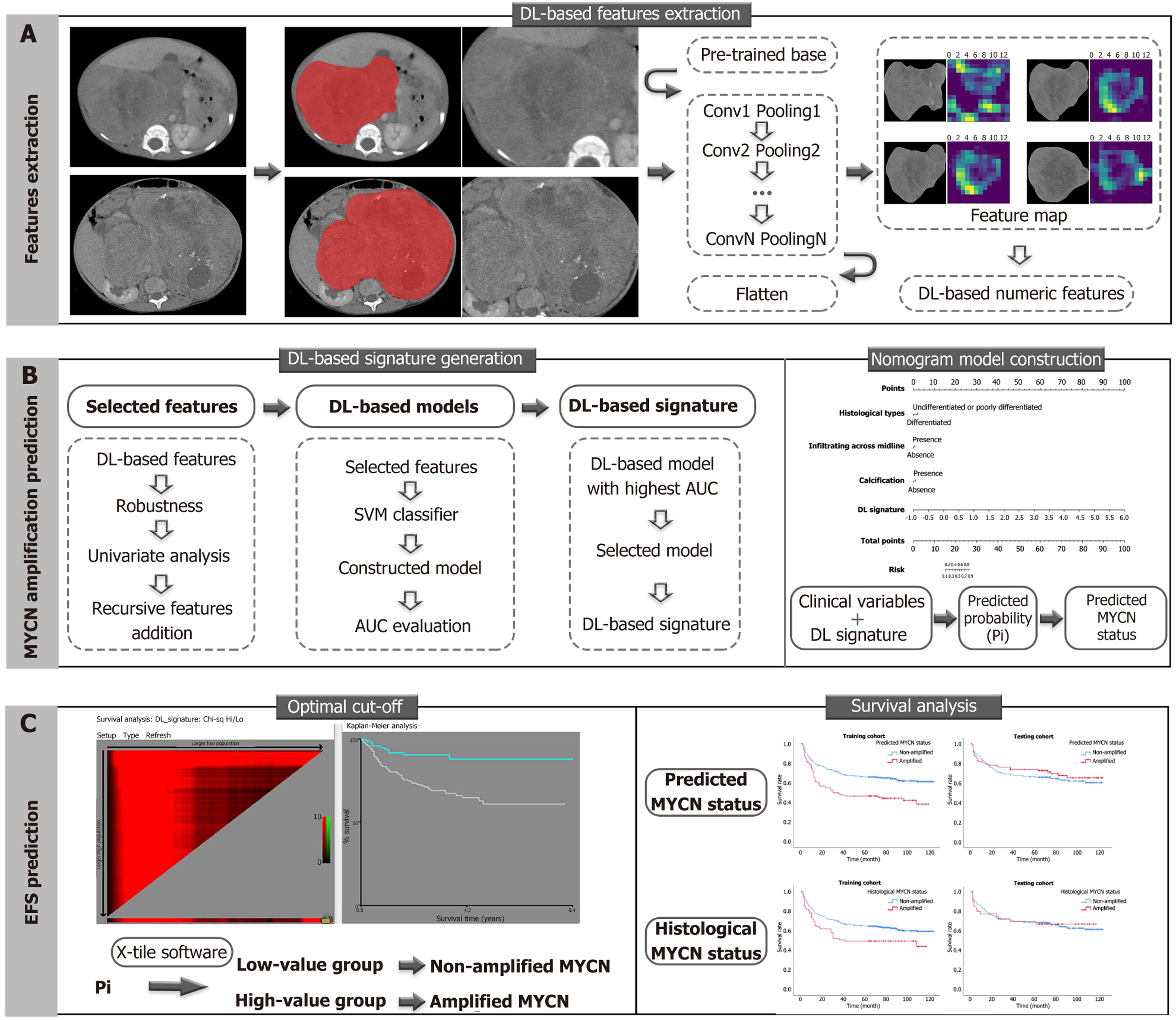
Figure 1 A general flowchart of data analysis.
A: Deep learning (DL)-based feature extraction was performed on contrast-enhanced computed tomography images of neuroblastoma by pre-trained convolutional neural network algorithms, which transformed the original images into numeric DL-based features; B: The MYCN amplification prediction was approached by DL-based features via DL-based signature generation and integrated nomogram model construction. The DL-based signature was generated on the DL-based model with the largest area under the receiver operating characteristic curve value in the testing cohort. The integrated nomogram model combined significant clinical variables and the DL-based signature; C: The survival analysis in the prediction of event-free survival was evaluated on MYCN amplification identified by nomogram-predicted and histopathological results. Predicted probabilities (Pi) are the total points that derive from independent variables in the nomogram model for differentiating non-amplified and amplified MYCN status. The optimal cut-off point of Pi was -1.996, defined by X-tile software, which indicated Pi ≤ -1.996 is considered as non-amplified MYCN status, and Pi > -1.996 is considered as amplified MYCN status. DL: Deep learning; SVM: Support vector machine; AUC: Area under the receiver operating characteristic curve; EFS: Event-free survival; Pi: Predicted probabilities.
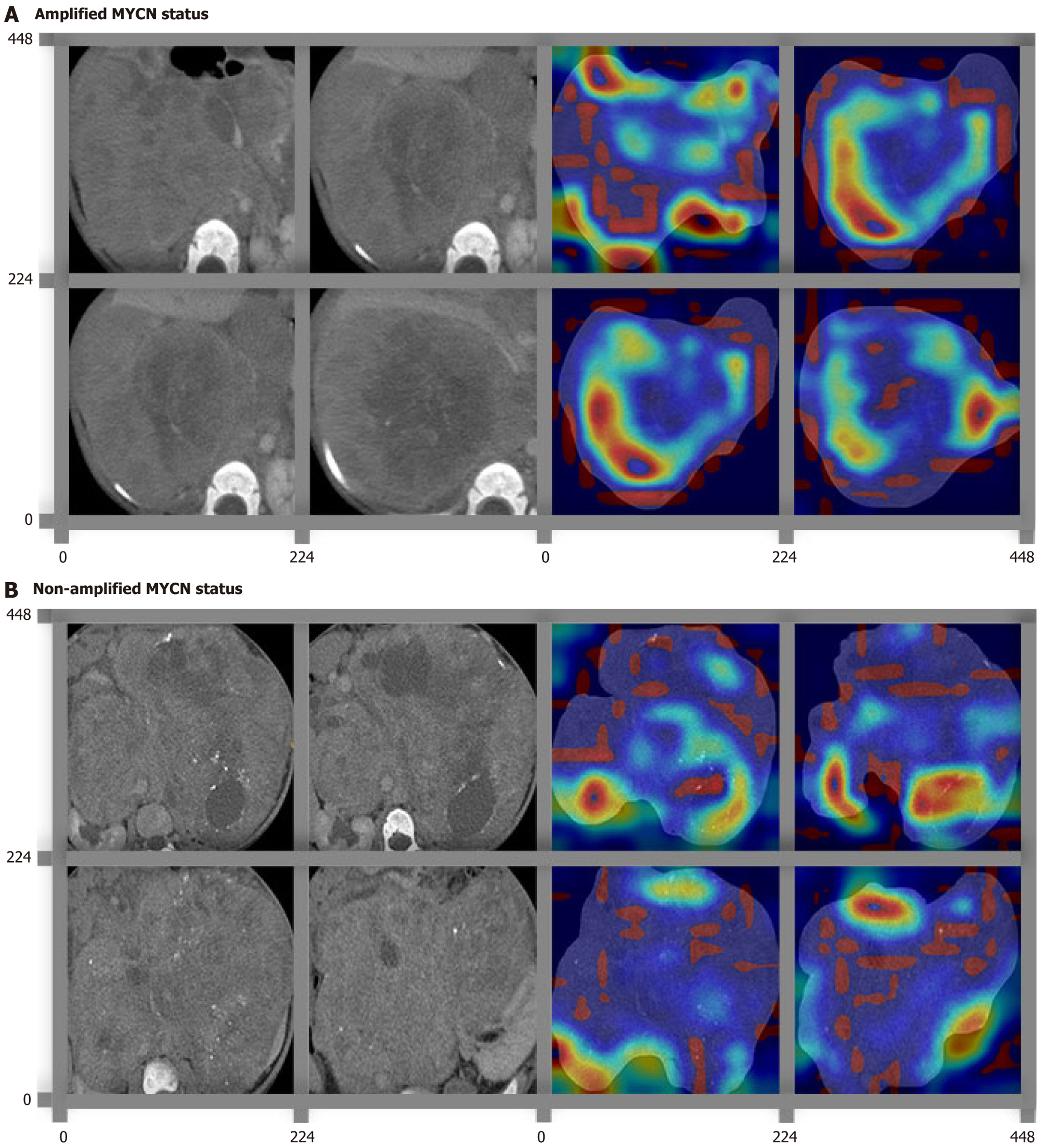
Figure 2 Feature heatmaps of representative histological patterns in bladder cancer on the deep learning ResNet50 algorithm via the Guided Grad-Class activation mapping.
A and B: The original computed tomography images and their corresponding feature heatmaps were shown in left and right sides, respectively. The red color highlighted the region of interest to classify MYCN amplification. Different subregions were recognized and highlighted in different MYCN statuses, including amplified MYCN status (A) and non-amplified MYCN status (B).
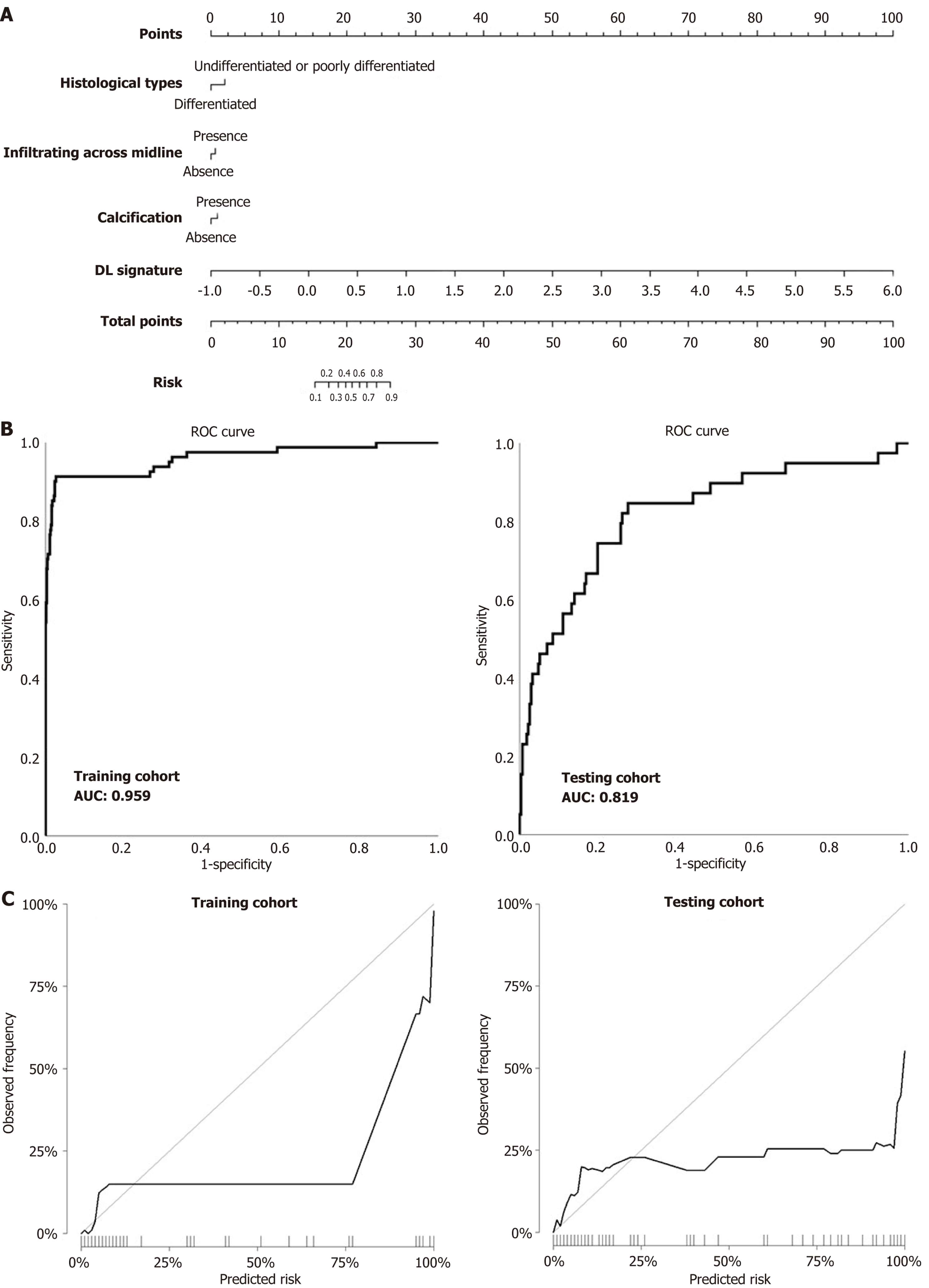
Figure 3 Evaluation of predictive performances for the integrated nomogram model for the prediction of MYCN amplification in neu
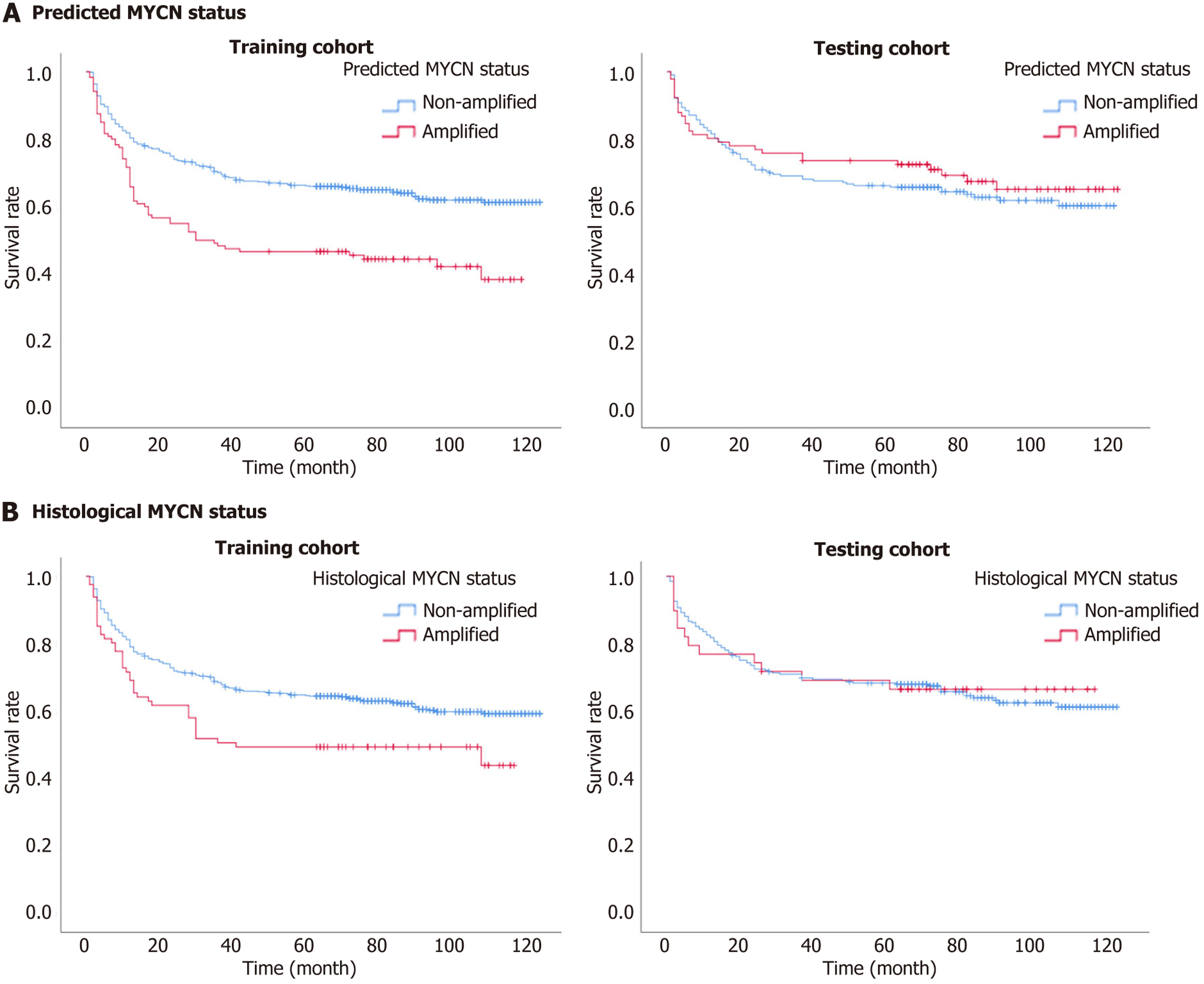
Figure 4 Kaplan-Meier survival curves on event-free survival of MYCN amplification status for patients with neuroblastoma in the training and testing cohorts.
A: Kaplan-Meier survival curves of nomogram model-predicted results, non-amplified vs amplified MYCN status, in the training and testing cohorts, respectively. Predicted probabilities (Pi) are the total points that derive from independent variables in the integrated nomogram model for differentiating non-amplified and amplified MYCN status. The optimal cut-off point of Pi was -1.996, defined by X-tile software, which indicated Pi ≤ -1.996 is considered as non-amplified MYCN status, and Pi > -1.996 is considered as amplified MYCN status; B: Kaplan-Meier survival curves of histopathological results, non-amplified vs amplified MYCN status, in the training and testing cohorts, respectively.
- Citation: Yang YH, Li Y. Deep learning radiomic analysis in the prediction of MYCN status and survival outcome in children with neuroblastoma. World J Clin Oncol 2026; 17(3): 114744
- URL: https://www.wjgnet.com/2218-4333/full/v17/i3/114744.htm
- DOI: https://dx.doi.org/10.5306/wjco.v17.i3.114744