Published online Mar 27, 2026. doi: 10.4240/wjgs.v18.i3.115320
Revised: November 7, 2025
Accepted: January 9, 2026
Published online: March 27, 2026
Processing time: 164 Days and 8.7 Hours
The coexistence of pancreatic ductal adenocarcinoma (PDAC) and serous cysta
A 62-year-old male patient presented to our hospital with a chief complaint of right upper abdominal discomfort. Extensive preoperative imaging examinations consistently indicated a mass located at the pancreatic tail with accompanying pancreatic duct dilation. Initially, an intraductal papillary mucinous neoplasm was suspected. However, intraoperatively, we discovered a mass at the pancreatic head and a mass at the pancreatic tail, and the entire pancreas demonstrated evid
MUC16 is associated with poor prognosis in coexistent PDAC and SCN. Moreover, enhancing the precision and accuracy of preoperative imaging diagnoses is of paramount importance, due to the difficulty in differential dia
Core Tip: The coexistent pancreatic ductal adenocarcinoma and serous cystadenoma occur rarely, accompanied by challenges in diagnosis. MUC16 is associated with poor prognosis in coexistent pancreatic ductal adenocarcinoma and serous cystadenoma. Enhancing the precision and accuracy of preoperative imaging diagnoses is of paramount importance in the differential diagnosis of coexistent pancreatic lesions.
- Citation: Li YT, Zhang YP, Fang QD, Wu HW, Xiao Y, Wu WM, Xue HD, Jin ZY, Wang ZW. MUC16 indicates poor prognosis in coexistent pancreatic ductal adenocarcinoma and serous cystadenoma: A case report and review of literature. World J Gastrointest Surg 2026; 18(3): 115320
- URL: https://www.wjgnet.com/1948-9366/full/v18/i3/115320.htm
- DOI: https://dx.doi.org/10.4240/wjgs.v18.i3.115320
Among pancreatic neoplasms, serous cystadenoma (SCN) accounts for approximately 1%-2% of all pancreatic tumors, while pancreatic ductal adenocarcinoma (PDAC) is even less frequent[1-4]. The coexistent tumors in the pancreas are even rarer. To our knowledge, there have been only seven case reports[5-11]. The incidence rate of pancreatic cancer has been increasing annually, accompanied by exceedingly high mortality rates and poor prognostic outcomes[12]. With advancements in next-generation sequencing (NGS) technology, sequencing has increasingly been applied to patients with pancreatic tumors, in hopes of discovering key driving genes. The widely recognized driver genes associated with pancreatic cancer include KRAS, TP53, CDKN2A, and SMAD4[13-15]. In recent years, MUC16 and several other genes have also been identified as risk genes for malignancy and poor prognosis in pancreatic cancers. However, there remains a lack of research focusing on the coexistence of pancreatic cancer and other pancreatic tumors. Therefore, it is necessary to discuss the prognosis and intrinsic genomics in this subtype of pancreatic cancer, which could provide valuable insights for future clinical diagnosis and treatment.
With the continuous advancements in medical imaging technology, abdominal imaging has witnessed a growing application in patients with pancreatic tumors, especially when presenting with nonspecific symptoms[16], while the complexity of tumor coexistence may pose potential diagnostic challenges. Among the fundamental imaging modalities, computed tomography (CT) and ultrasound (US) are widely employed. For the evaluation of dilated common bile ducts or pancreatic duct (PD), magnetic resonance cholangiopancreatography (MRCP) has been established as the gold stan
This study presented a rare case and aimed to identify critical gene mutations in the context of pancreatic dual tumors with spatial distance, through the technique of whole-genome sequencing. Meanwhile, we highlighted the challenges encountered in the differential diagnosis of intraductal papillary mucinous neoplasm (IPMN), PDAC, SCN, and chronic pancreatitis. Although these entities are known to exhibit distinct manifestations in various imaging modalities, it is important to recognize that the actual clinical scenario can be more complex and intriguing than anticipated.
The coexistent pancreatic tumors were discovered during routine medical care, with continuous follow-up extending from March 2015 to May 2016 until death. Detailed information regarding clinical symptoms, physical examinations, and medical history was collected. Auxiliary examinations included laboratory, imaging, and pathological profiles, all of which were obtained from the medical record data system of Peking Union Medical College Hospital. Laboratory indicators included blood routine, liver and kidney function, and tumor markers. The serum normal range reference values applied in our study were reported as follows: (1) Carbohydrate antigen (CA) 19-9: 0-34 U/mL; (2) Carcinoembryonic antigen: 0-5 ng/mL; and (3) CA24-2: 0-20 U/mL. The imaging characteristics of this disease often resembled those of other conditions, complicating the differential diagnosis. Preoperative imaging studies included CT, positron emission tomography-CT (PET-CT), MRCP, and EUS. Image analysis was conducted and verified by two experienced radiologists through a retrospective consistent review. Pathological profiles included macroscopic specimens and pathological staining sections embedded in paraffin, examined and reported by two experienced pathologists. This study was approved by the institutional ethics review board, and informed consent was obtained.
In March 2015, a 62-year-old male presented with right upper abdominal discomfort for one year.
He denied any concomitant symptoms or prior treatment.
Heavy tobacco and alcohol consumption, and cerebral infarction history were reported.
Nothing special was found in his physical examination.
The genomic DNAs (gDNA) of the tumor tissue were extracted using the standard protocols with the QIAamp DNA FFPE Tissue Kit (Qiagen, 56404). DNA concentrations were determined by Qubit dsDNA HS Assay Kit (Invitrogen, Q32854). The gDNAs were fragmented to sizes ranging from 150 bp to 250 bp using the Covaris S220. A next-generation sequencing gDNA library was prepared for whole-genome sequencing (WGS) using 150-300 ng of fragmented gDNA. In brief, the library was prepared by KAPA Hyper Prep Kit (KAPA, KK8504) according to the manual. The concentrations of the library were determined using Qubit, and the size distributions of the library were analyzed using the Bioanalyzer 2100 (Agilent Technologies).
The pooled DNAs of the library were mixed with 1 μL of DNA blocker (Integrated DNA Technologies) and 5 μL of SureSelect QXT Fast Blocker Mix (Agilent) before being dried by a vacuum concentrator. The dried mixture was then dissolved in a 13 μL hybridization buffer, supplied by the hybridization of SureSelect QXT Reagent kit (Agilent). Sequencing of the whole-genome libraries was performed using 150bp paired-end runs on the Illumina HiSeq × 10 (Illumina). The 150 bp paired-end sequencing data were aligned to the human reference genome hg19 (GRCh37) by bwa v0.7.17[20], and variant calling analysis was performed using VarScan v2.3.8[21]. Calls were annotated using ANNOVAR v2015.06[22] and COSMIC v70[23]. To identify relevant gene targets for pancreatic cancer, a Hotspot Panel was selected based on common variants from pancreatic cancer samples through WGS. A Literature review was also conducted to help narrow down the gene list. And laboratory examinations, including CA19-9 (14.2 U/mL), carcinoembryonic antigen (4.78 ng/mL), and CA242 (6.4 U/mL) revealed no significant abnormalities.
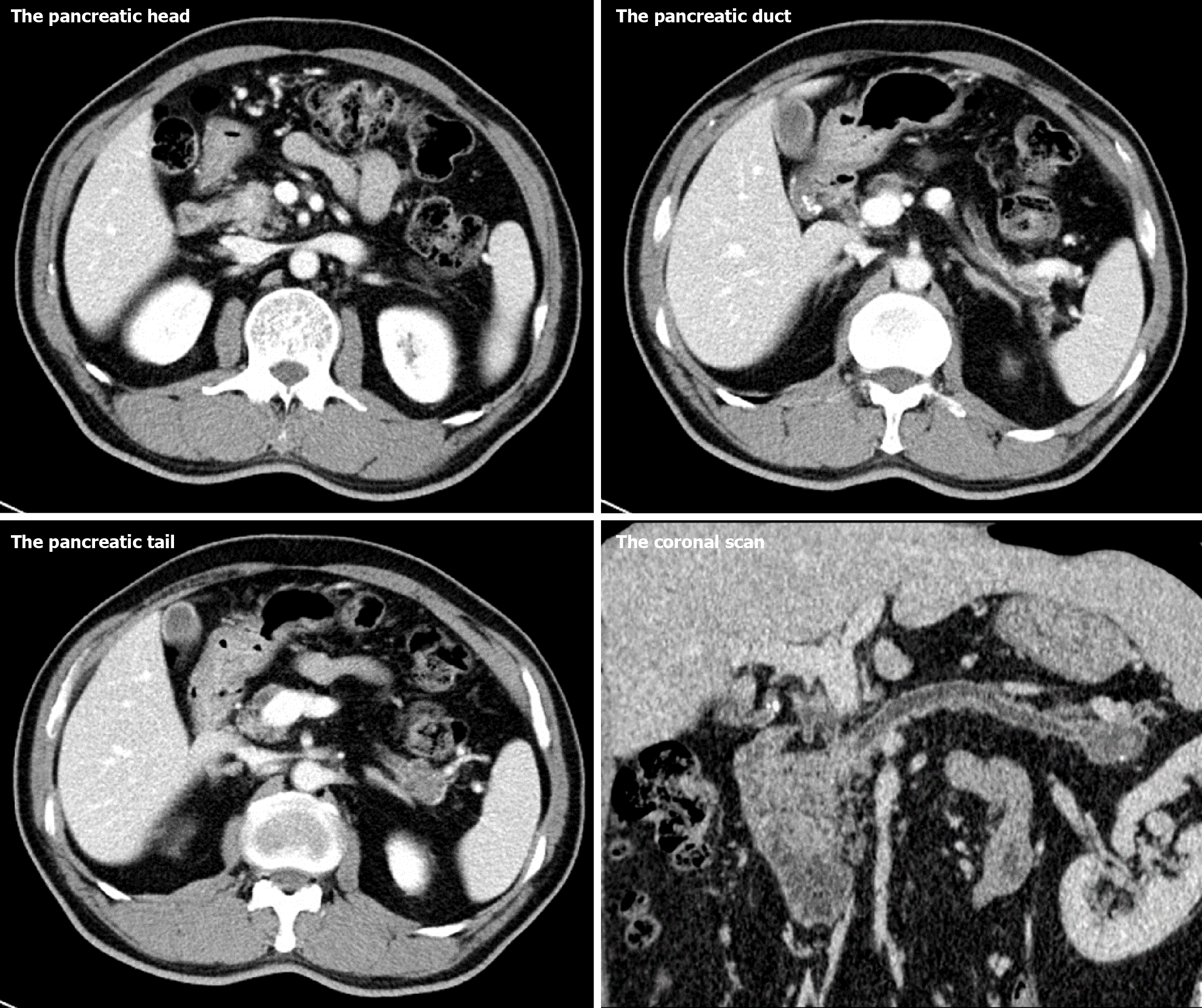
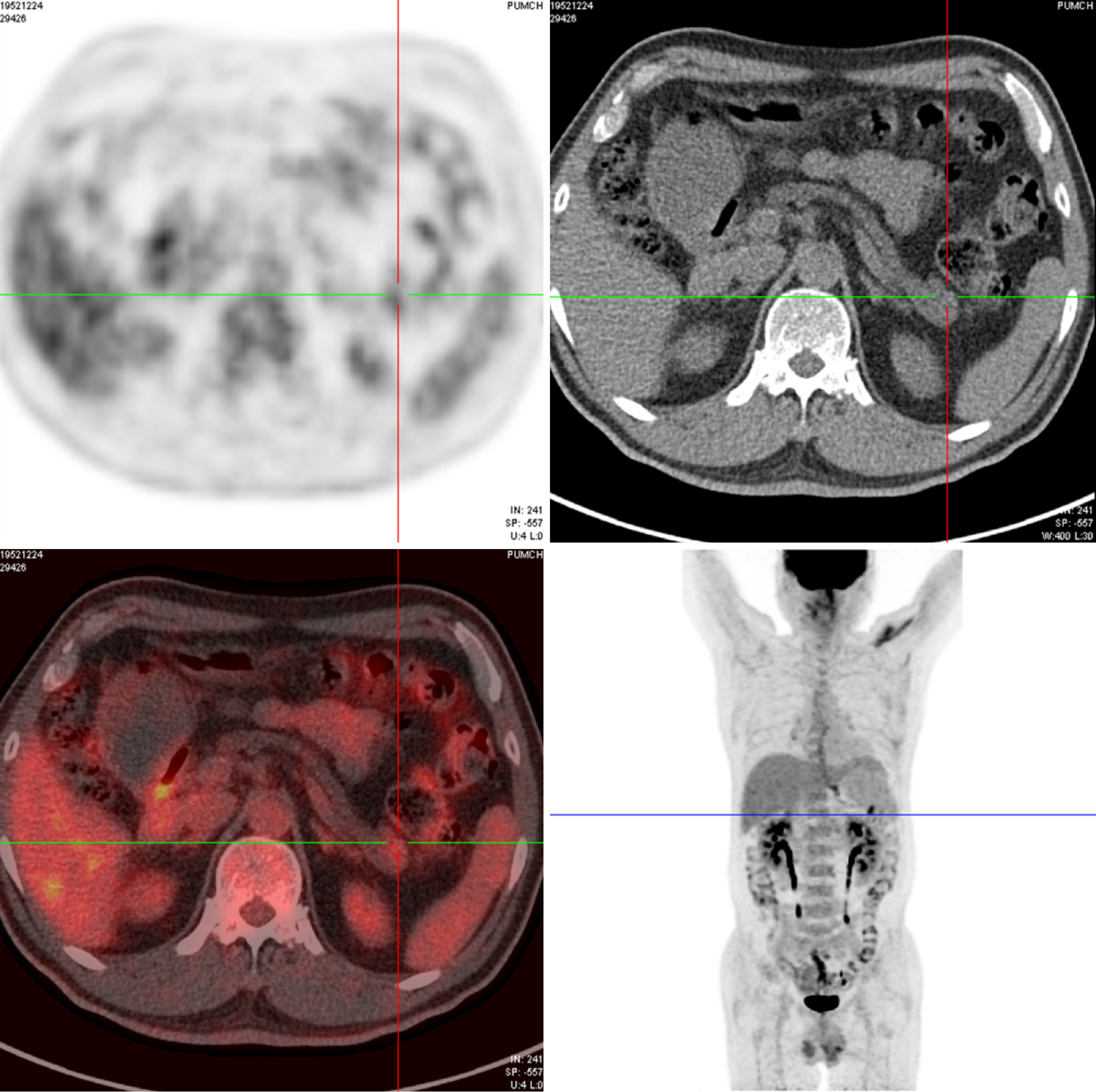
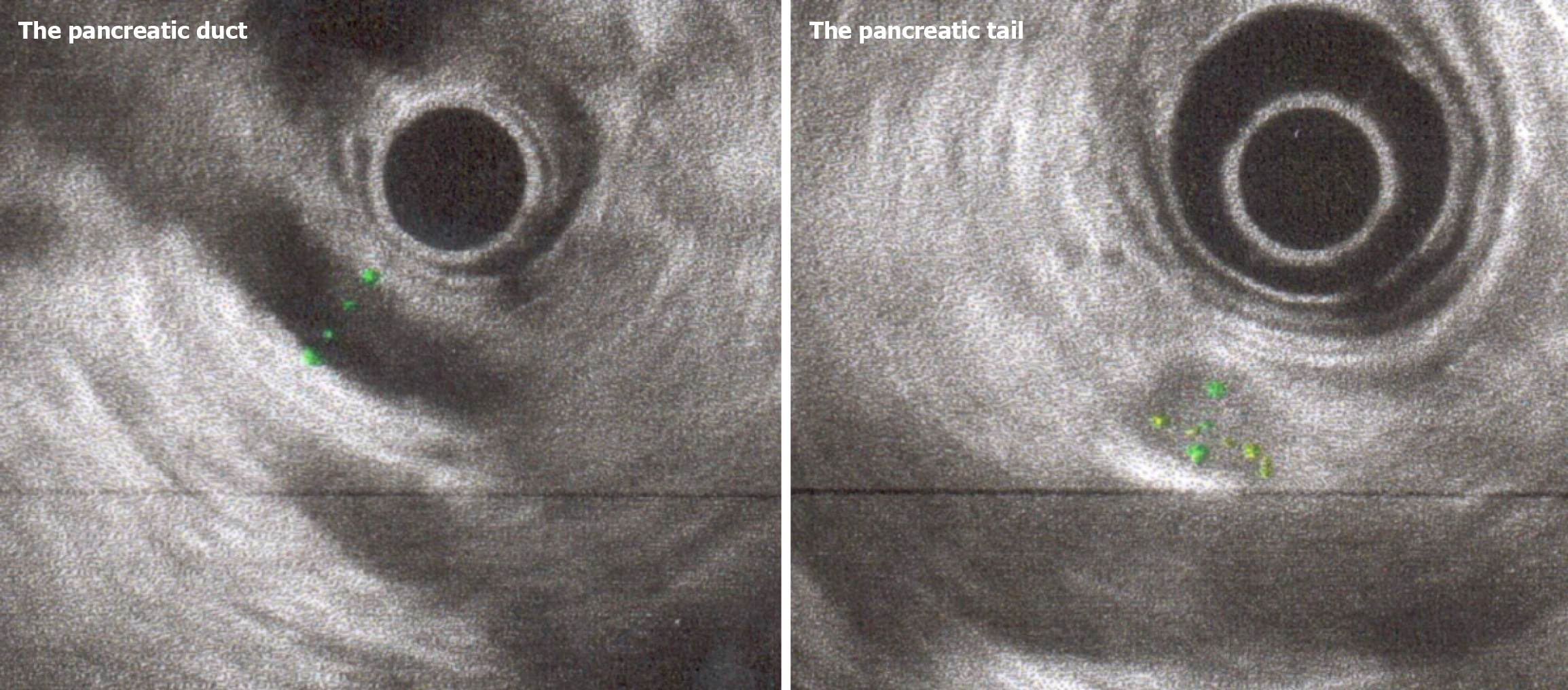
MRCP revealed dilation of PD and common bile ducts, along with a cystic lesion at the tail. The high-resolution CT scan with three-dimensional reconstruction revealed atrophy in the pancreatic neck, body, and tail. It also documented a dilated PD and a low-density lesion at the tail, measuring 3.0 cm × 2.0 cm, with well-defined borders and multiple small cystic-like foci, some connected to the main PD, further supporting the diagnosis of IPMN (Figure 1). PET-CT confirmed pancreatic atrophy and identified a mass (1.7 cm × 2.0 cm) at the tail with increased radioactive uptake (Figure 2). In the pancreatic tail, EUS displayed a cystic PD dilation and an irregularly enhanced echo (2.5 cm × 1.8 cm) containing non-echoic dots. At the head, the PD was measured 0.6 cm in width, and a hyperechoic nodule was identified in the neck region of the PD (Figure 3). These findings collectively pointed towards a strong suspicion of IPMN.
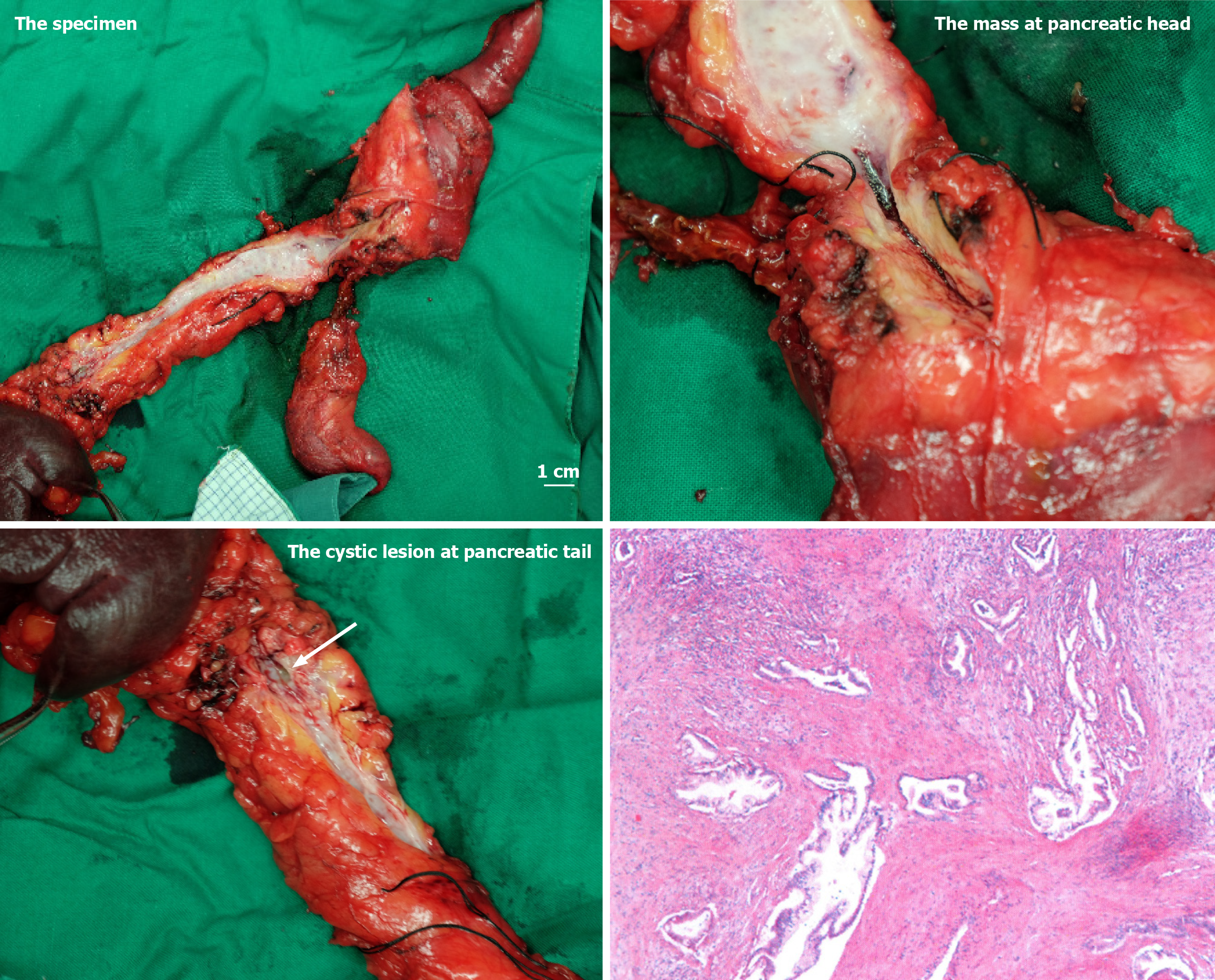
Surprisingly, postoperative pathology revealed well-differentiated PDAC at the head, infiltrating surrounding adipose and neural tissues, classified as stage T3N0M0 (Figure 4); while the tail mass was identified as SCN, with minimal cytological atypia and no invasive features, in concomitant context of chronic pancreatitis.
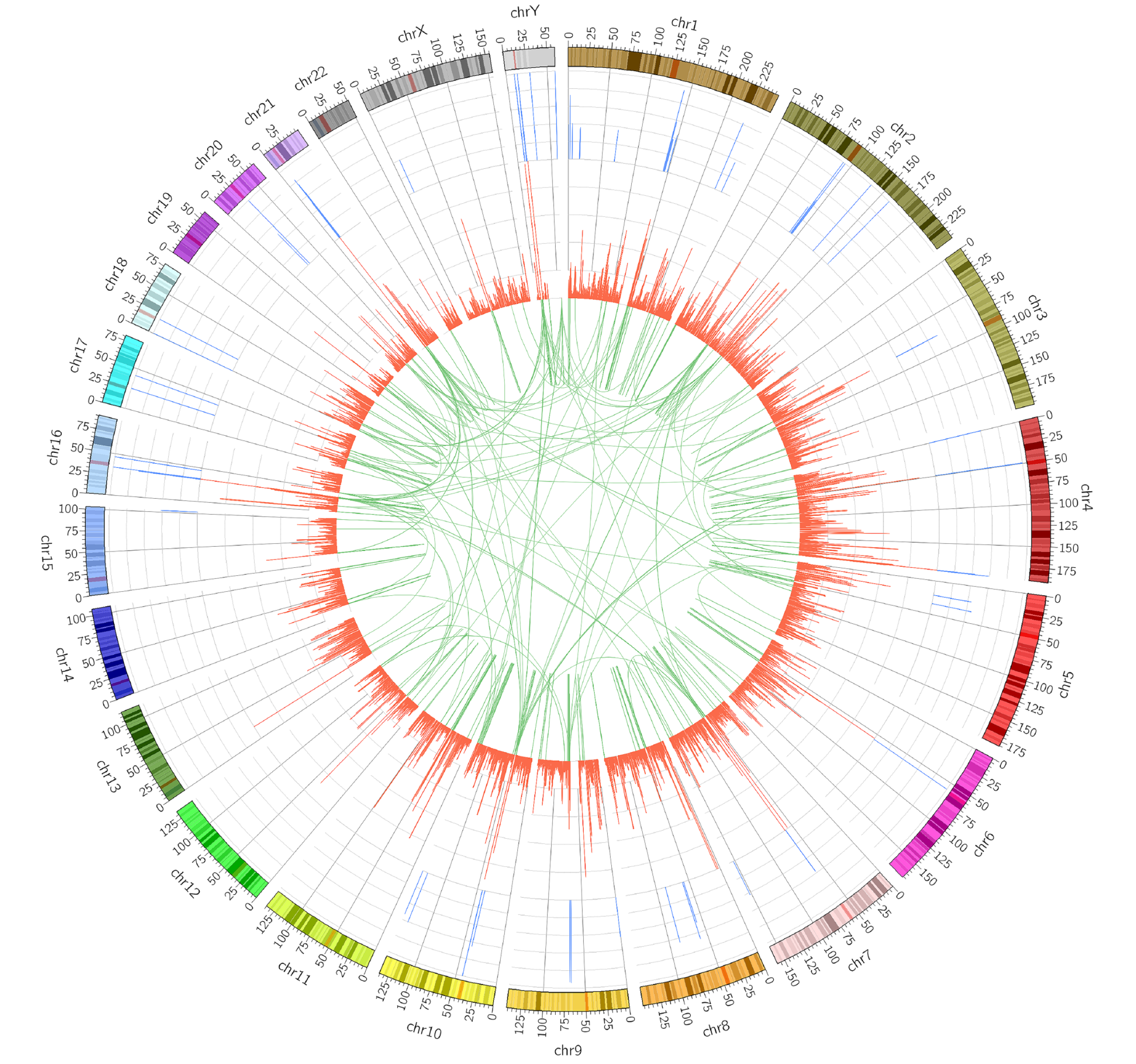
The gDNA from the coexistent tumor tissues was extracted and sequenced. The sequencing depth of WGS was 40 ×. Structure variations, spot mutation variations, and copy number variations were analyzed and integrated (Figure 5). Over 24-thousand genes with effective single nucleotide polymorphism were detected. Among them, important mutations were reported for MUC16, TP53, KRAS, CDKN2A, and SMAD4, compared to sequencing result of normal pancreatic tissue from the same patient.
Compared to the other mutations, MUC16 was regarded as an important mutant gene associated with the coexistence of PDAC and SCN, as well as the poor prognosis. Through whole genome sequencing, we identified two categories of mutation sites in MUC16, including deleterious sites and tolerated sites (Figure 6).
The deleterious mutant sites were: (1) MUC16_p.A12850P; (2) MUC16_p.D12871A; (3) MUC16_p.D13589G; (4) MUC16_p.L13050F; (5) MUC16_p.L13171F; (6) MUC16_p.S12849F; (7) MUC16_p.T13579I; (8) MUC16_p.T13585P; and (9) MUC16_p.W13569C. Refer to Figure 6 for the tolerated mutation sites and additional information. The abundant deleterious mutations hinder the normal protein expression of MUC16 in the patient, consequently leading to the development of pancreatic tumors and unfavorable prognostic outcomes.
The patient underwent surgery in April 2015. Intraoperatively, a hard and irregular mass was identified at the pancreatic head and a well-defined, pliable yet firm mass at the tail. The entire pancreas exhibited evident atrophy and marked PD dilation (Figure 4). Consequently, a total pancreatectomy with splenectomy was performed. Postoperatively, adjuvant therapy was not administered due to the patient’s weakened physical condition.
Unfortunately, lymph node metastasis, abdominal cavity metastasis, and suspected lung metastasis were observed 10 months after the surgery, and the patient succumbed 13 months after the surgery.
This article focused on the intrinsic genomic profiles, prognostic characteristics, and imaging diagnosis of coexistent pancreatic tumors. Our study presented a mass in the pancreatic tail with PD dilation in imaging examinations, while ultimately confirmed pathologically as PDAC in the head and SCN in the tail. Coexistent PDAC and SCN, spatially distant from each other, are exceedingly rare. Whole genome sequencing suggested that MUC16 is associated with dual tumor coexistence and poor prognosis.
SCN is a relatively uncommon and benign neoplasm, representing approximately 25% of all pancreatic cystic tumors[7]. Its coexistence with other pancreatic neoplasms, such as ductal adenocarcinoma and endocrine tumors, is infrequent[5]. Recently, the World Health Organization updated the classification of SCN, introducing the category of mixed serous neuroendocrine neoplasms[24]. Some reported cases have documented the presence of SCN concomitant with endocrine tumors. However, our literature search revealed that only 6 cases of coexistent PDAC and SCN had been reported before our study[5,6,8-10], in which the dual tumors all had spatial distance between them. Therefore, the coexistence of PDAC and SCN is exceptionally rare, rendering prognosis analysis challenging.
PDAC is a highly aggressive carcinoma, and epidemiological projections indicate a rise in the incidence of pancreatic cancer in the coming decade[1,2]. The simultaneous occurrence of PDAC and SCN may confer a more unfavorable prognosis compared to either PDAC or SCN in isolation, given their presumed shared origins. As proposed by Nitta and Montag, aberrations in pancreatic exocrine cells may contribute to the development of both lesions[7,9]. Consequently, SCN coexisting with PDAC may possess a greater malignant potential, with an increased likelihood of evolving into serous cystadenocarcinoma. Furthermore, patients with dual neoplasms, especially two distinct lesions located in separate regions of the pancreas, may experience more extensive organ damage, resulting in a greater loss of normal pancreatic tissue and an enlarged surgical resection field. Consequently, the prognosis for the coexistence of PDAC and SCN is anticipated to be worse.
Our study raised two questions: What is the reason for the coexistence of PDAC and SCN, and what is the relationship between the two lesions? Precursor lesions of pancreatic cancer include pancreatic intraepithelial neoplasia (PanIN), IPMN, and mucinous cystic neoplasm. PanINs develop within the exocrine ducts of pancreatic lobules, which are categorized into PanIN-1, PanIN-2, and PanIN-3 based on the degree of dysplasia. Approximately 5%-10% of non-invasive cystic lesions may advance to invasive adenocarcinoma, referred to as intraductal papillary mucinous carcinoma, accounting for 10% of all the resected PDACs[25]. Differentiate from IPMN, mucinous cystic neoplasm lacks communication with the PD, carrying an estimated 10% risk of malignant transformation[26]. Additionally, a small subset of SCNs may progress into serous cystadenocarcinomas, posing a risk of invasive malignancy[27]. However, regardless of precancerous types, coexistent precursor and cancer lesions typically share an adjacent spatial location, a microscopically benign-malignant transition zone, and similar immunohistochemical characteristics[11]. Since our two lesions were located at opposite ends of the pancreas, we believed that they were geographically independent, with a much lower likelihood of sharing the same origin. One potential explanation could be that the SCN induced PD obstruction and chronic pancreatitis, thereby increasing the risk of PDAC occurrence. Andea et al[28] indicated that PanIN or PDAC occurred more frequently in the context of chronic pancreatitis, compared to normal pancreatic tissue. Another possibility is that this patient may harbor unidentified genetic predispositions to coexisting tumors. Therefore, we hoped that genome-wide sequencing could provide some clues.
The updated report from the American Cancer Society ranks pancreatic cancer as the third leading cause of cancer-related mortality, trailing behind lung cancer, prostate or breast cancer. In the United States, the mortality rate of pancreatic cancer has been increasing annually by 0.3%. Pancreatic cancer accounts for around 3% of newly diagnosed cancers, ranking 10th among males and 7th among females[12].
From an evolutionary perspective, the development of pancreatic cancer can be divided into three stages: (1) Initiation; (2) Clonal expansion; and (3) Invasive dissemination[29]. In the initiation stage, induced by external exposures or DNA repair deficiencies, driver gene mutations occur in normal pancreatic cells. Subsequently, during the clonal expansion phase, mutation-carrying cells proliferate into tumor clones, which could be described in a stepwise progression model[30] or a punctuated evolution model[31]. In the invasive dissemination stage, it is crucial to emphasize interactions between tumor cells and their surrounding microenvironment, including factors like desmoplastic stroma, oxygen concentrations, and immunological landscape[29].
Traditionally, the widely recognized driver genes for pancreatic cancer include KRAS, CDKN2A, TP53, and SMAD4. Under the influence of these genes, normal pancreatic cells develop into PanIN, eventually progressing to invasive cancer[13,14,32]. Recent genomic analyses[30,33] have revealed that KRAS mutations are present in over 99% of PanIN-1 Lesions, CDKN2A inactivation can be detected as early as PanIN-2, while TP53 and SMAD4 mutations occur in PanIN-3. In the human genome of PDACs, KRAS mutation is the most common (90%-95%), while CDKN2A, TP53, and SMAD4 are inactivated in 50%-80% of cases[15,34]. These four key driver genes act synergistically, forming a complex network that promotes tumorigenesis[29].
With the continuous iteration of genomic sequencing technology, the research scope has expanded to include point mutations, gene copy number variations, and chromosome structure variations. Consequently, several genes or pathways beyond the core genes have been newly discovered, such as GATA6[31], KMT2C[15], SF3B1[15,35], and SLIT/ROBO axon signal pathway[35]. Moreover, germline mutations associated with DNA stability, including BRCA1/2 and PALB2, are also involved[36].
Our whole genome sequencing also exhibited these four gene mutations, which played a driving role in carcinoma formation. Notably, a special gene mutation, MUC16, was also detected in our patient. In 2017, Balachandran et al[37] reported the unique expression of MUC16 in PDAC with heterogeneous immunology, sparking widespread discussion and attention. MUC16, situated on chromosome 19p13.2, spans 374k base pairs and contains 92 exons. It encodes a high-molecular-weight mucin-like glycoprotein, commonly found in the epithelial cells of adult body cavities, also known as CA125, comprising a large number of O-glycosylated tandem repeat domains and extension structures.
It was demonstrated that MUC16 was the third most common mutated gene, following KRAS and TP53, in two independent PDAC cohorts[37]. High levels of CA125 were observed in 50%-70% of PDAC patients[38]. Numerous studies have shown that elevated expression and abnormal O-glycosylation of MUC16 were independent risk factors for tumor progression, increased metastasis, and poor survival in PDAC patients[38-40]. However, to date, there have been no reports or articles focused on the correlation between genetics and the coexistence of pancreatic tumors. In our study, MUC16 was pronouncedly highlighted through genome-wide sequencing. In consistency with the previous studies, we believe that MUC16 played an important role in the poor prognosis of pancreatic cancer. Since the coexistence of pancreatic tumors is exceedingly rare, only one set of sequencing data was available in this study. Moreover, PDAC inherently carries an extremely poor prognosis. Therefore, we cannot exclude the possibility that the short survival observed in this case may have been influenced by other factors, such as resistance to radiotherapy or chemotherapy. Nevertheless, based on our findings and a review of the existing literature, we propose that MUC16 may be associated with both the coexistence and poor prognosis of dual or multiple pancreatic tumors. Further studies with larger sample sizes are warranted to validate this potential association.
In addition to the common features of pancreatic cancer, inter-individual heterogeneity has become a hot topic in recent research. Heterogeneity implies differences in etiology, clinical manifestations, treatment responses, and prognosis. Stratification or grouping based on this heterogeneity aids in personalized diagnosis and treatment of pancreatic cancer. Collisson et al[41] were the first to report genomic variations among PDAC patients, categorizing them into classical, quasi-mesenchymal, and exocrine-like subtypes. Subsequently, Moffitt et al[42] identified two subtypes, classical and basal-like; while Waddell et al[15] determined four subtypes: Stable, locally rearranged, scattered, and unstable. Lately, through genomic sequencing, Bailey et al[43] classified PDACs into four characteristic subgroups: Squamous, pancreatic progenitor, immunogenic, and aberrantly differentiated endocrine-exocrine. There exists correspondence or overlap among all these reported classifications.
Our study focused on an important clinical subtype of pancreatic cancer: Coexistent PDAC and IPMN. There presents important reference value in sharing clinical features, imaging findings, pathological profiles, and prognosis of this specific subtype. Furthermore, exploring the correlation between these features and the intrinsic genomic information holds indicative significance.
Imaging examinations are crucial for diagnosing pancreatic masses, as pancreatic diseases often remain asymptomatic until tissue damage and malignancies have advanced or metastasized. Contrast-enhanced multidetector CT and PET-CT play central roles in diagnosing, staging, and treatment planning for PDAC patients[44]. Besides, it is reported that MRI could better detect cystic lesions, while EUS could be more accurate for small solid lesions and convenient for fine needle aspiration[3,10].
The uniqueness of our study lies in its distinctive imaging presentation, which posed challenges in differential diagnosis: (1) The radiological findings of the pancreatic tail mass and PD dilation strongly suggested IPMN, contradicting the final pathology of SCN; and (2) The mass in the pancreatic head displayed iso-density and iso-enhancement with surrounding normal tissue, complicating the identification of PDAC and differential diagnosis of PD dilation.
IPMN originates from the epithelium of the main PD or its branches and often presents variable duct dilation. Neoplastic cells in IPMN typically exhibit a papillary morphology, although flat epithelium can also be observed[45]. Diagnostic modalities such as MRCP, CT, and EUS are commonly employed to evaluate IPMN. Key indicators of IPMN include a dilated PD with a connected lesion, a patulous or “fish-eye” papilla, and mucin presented at the papillae[45]. Furthermore, analyzing protein expressions in various cyst fluids may also contribute to diagnosis[46].
In this study, determining the etiology of PD dilation was a crucial aspect of differential diagnosis. According to our imaging findings, IPMN appeared to be the most plausible etiology. However, the possibility of chronic pancreatitis or a mass at the pancreatic head should also be considered, as both conditions could lead to PD dilation. Simultaneously, it is important to emphasize that the ultimate diagnosis confirmation relies on pathological examination. Additionally, in patients with IPMN, the coexistence of distinct pancreatic cancer is possible, particularly when early-stage cancer may not be readily detectable by CT, MRI, or EUS[47]. In certain instances, concurrent chronic pancreatitis and IPMN may also occur[16]. Consequently, it is essential to consider all possibilities, including IPMN, chronic pancreatitis, and pancreatic mass at the head, when arriving at a comprehensive diagnosis. Given the challenging differential diagnosis of coexistent pancreatic tumors, it is imperative to improve the accuracy of preoperative imaging examinations. Nevertheless, there is limited evidence regarding the accuracy of imaging screening, highlighting a research gap that needs to be filled.
Our article indicated the association between MUC16 mutation and poor prognosis, along with coexistent pancreatic tumors, paving a new avenue for future studies. There have been several studies exploring the application of this correlation in clinical settings. MUC16 fluorescent probes were created to aid in lesion marking during surgery[48], while immune imaging probes were expected to improve early recognition of pancreatic cancer in imaging examinations[49]. Additionally, anti-MUC16 nanoantibodies might serve as a novel immunotherapeutic agent in PDAC patients[38].
There are some limitations in this study. The sample size of coexistent PDAC and SCN was limited, and the evidence strength was not sufficient enough. However, we presented the seventh case worldwide of coexistent pancreatic tumors with spatially distant locations. The clinical and genomic profiles are extremely rare and valuable, especially in this subtype of pancreatic cancer. Consistent with the results from Balachandran et al[37], our findings hold considerable credibility and suggestive significance. Future studies with larger samples are required for further validation. Ad
In conclusion, our study focused on coexistent PDAC and SCN. Whole-genome sequencing indicated an association between MUC16 with poor prognosis and tumor coexistence. Imaging examinations with higher accuracy and more studies are needed.
| 1. | Raimondi S, Maisonneuve P, Lowenfels AB. Epidemiology of pancreatic cancer: an overview. Nat Rev Gastroenterol Hepatol. 2009;6:699-708. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 498] [Cited by in RCA: 451] [Article Influence: 26.5] [Reference Citation Analysis (0)] |
| 2. | Stolzenberg-Solomon RZ, Amundadottir LT. Epidemiology and Inherited Predisposition for Sporadic Pancreatic Adenocarcinoma. Hematol Oncol Clin North Am. 2015;29:619-640. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 26] [Article Influence: 2.6] [Reference Citation Analysis (1)] |
| 3. | Capurso G, Signoretti M, Valente R, Arnelo U, Lohr M, Poley JW, Delle Fave G, Del Chiaro M. Methods and outcomes of screening for pancreatic adenocarcinoma in high-risk individuals. World J Gastrointest Endosc. 2015;7:833-842. [PubMed] [DOI] [Full Text] |
| 4. | van Huijgevoort NCM, Del Chiaro M, Wolfgang CL, van Hooft JE, Besselink MG. Diagnosis and management of pancreatic cystic neoplasms: current evidence and guidelines. Nat Rev Gastroenterol Hepatol. 2019;16:676-689. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 215] [Cited by in RCA: 174] [Article Influence: 24.9] [Reference Citation Analysis (1)] |
| 5. | Montag AG, Fossati N, Michelassi F. Pancreatic microcystic adenoma coexistent with pancreatic ductal carcinoma. A report of two cases. Am J Surg Pathol. 1990;14:352-355. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 25] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 6. | Posniak HV, Olson MC, Demos TC. Coexistent adenocarcinoma and microcystic adenoma of the pancreas. Clin Imaging. 1991;15:220-222. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 14] [Article Influence: 0.4] [Reference Citation Analysis (1)] |
| 7. | Nitta H, Hirota M, Ohkado A, Motomura Y, Chikamoto A, Shibata M, Takamori H, Kanemitsu K, Imamura T, Yamamoto T, Baba H. Coexistence of serous cystadenoma and ductal adenocarcinoma in the pancreas: a case report. Pancreas. 2008;36:218-219. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 8] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 8. | Izumi S, Onoda Y, Shiota K. A Case of Serous Cystic Adenoma Coexistent with Pancreas Head Carcinoma. Jpn J Gastroenterol Surg. 2008;41:1821-1826. [DOI] [Full Text] |
| 9. | Ando N, Kato J, Mabuchi M, Obara K, Iwasa J, Ohshima Y, Iwata K, Imose M, Ohnishi T, Shimizu S, Sugihara J, Iwata H, Yasuda I, Moriwaki H. [A case report of pancreatic serous cystadenoma coexistent with adenosquamous carcinoma]. Nihon Shokakibyo Gakkai Zasshi. 2012;109:442-450. [PubMed] |
| 10. | Rowan D, Pant M, Hagen C, Giorgadze T. Coexisting pancreatic serous cystadenoma and pancreatic ductal adenocarcinoma: A cytological-pathologic correlation with literature review. Ann Diagn Pathol. 2019;42:87-91. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 11. | Yoshida M, Naitoh I, Hayashi K, Jinno N, Natsume M, Hori Y, Kato A, Kachi K, Asano G, Matsuo Y, Takahashi S, Kataoka H. Metachronous Pancreatic Ductal Adenocarcinoma with Adjacent Serous Cystadenoma that Was Preoperatively Diagnosed by EUS-FNA: A Case Report and Review of the Literature. Intern Med. 2020;59:649-656. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 4] [Article Influence: 0.7] [Reference Citation Analysis (1)] |
| 12. | Siegel RL, Giaquinto AN, Jemal A. Cancer statistics, 2024. CA Cancer J Clin. 2024;74:12-49. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7368] [Cited by in RCA: 6584] [Article Influence: 3292.0] [Reference Citation Analysis (4)] |
| 13. | Maitra A, Hruban RH. Pancreatic cancer. Annu Rev Pathol. 2008;3:157-188. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 613] [Cited by in RCA: 585] [Article Influence: 32.5] [Reference Citation Analysis (5)] |
| 14. | Connor AA, Gallinger S. Pancreatic cancer evolution and heterogeneity: integrating omics and clinical data. Nat Rev Cancer. 2022;22:131-142. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 302] [Cited by in RCA: 255] [Article Influence: 63.8] [Reference Citation Analysis (1)] |
| 15. | Waddell N, Pajic M, Patch AM, Chang DK, Kassahn KS, Bailey P, Johns AL, Miller D, Nones K, Quek K, Quinn MC, Robertson AJ, Fadlullah MZ, Bruxner TJ, Christ AN, Harliwong I, Idrisoglu S, Manning S, Nourse C, Nourbakhsh E, Wani S, Wilson PJ, Markham E, Cloonan N, Anderson MJ, Fink JL, Holmes O, Kazakoff SH, Leonard C, Newell F, Poudel B, Song S, Taylor D, Waddell N, Wood S, Xu Q, Wu J, Pinese M, Cowley MJ, Lee HC, Jones MD, Nagrial AM, Humphris J, Chantrill LA, Chin V, Steinmann AM, Mawson A, Humphrey ES, Colvin EK, Chou A, Scarlett CJ, Pinho AV, Giry-Laterriere M, Rooman I, Samra JS, Kench JG, Pettitt JA, Merrett ND, Toon C, Epari K, Nguyen NQ, Barbour A, Zeps N, Jamieson NB, Graham JS, Niclou SP, Bjerkvig R, Grützmann R, Aust D, Hruban RH, Maitra A, Iacobuzio-Donahue CA, Wolfgang CL, Morgan RA, Lawlor RT, Corbo V, Bassi C, Falconi M, Zamboni G, Tortora G, Tempero MA; Australian Pancreatic Cancer Genome Initiative, Gill AJ, Eshleman JR, Pilarsky C, Scarpa A, Musgrove EA, Pearson JV, Biankin AV, Grimmond SM. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature. 2015;518:495-501. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2296] [Cited by in RCA: 2077] [Article Influence: 188.8] [Reference Citation Analysis (5)] |
| 16. | Carriere V, Conway J, Evans J, Shokoohi S, Mishra G. Which patients with dilated common bile and/or pancreatic ducts have positive findings on EUS? J Interv Gastroenterol. 2012;2:168-171. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 17] [Article Influence: 1.2] [Reference Citation Analysis (1)] |
| 17. | Levin DC, Rao VM, Parker L, Frangos AJ, Sunshine JH. Recent trends in utilization rates of abdominal imaging: the relative roles of radiologists and nonradiologist physicians. J Am Coll Radiol. 2008;5:744-747. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 36] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 18. | Ledro-Cano D. Suspected choledocholithiasis: endoscopic ultrasound or magnetic resonance cholangio-pancreatography? A systematic review. Eur J Gastroenterol Hepatol. 2007;19:1007-1011. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 40] [Article Influence: 2.1] [Reference Citation Analysis (5)] |
| 19. | Nickl NJ, Bhutani MS, Catalano M, Hoffman B, Hawes R, Chak A, Roubein LD, Kimmey M, Johnson M, Affronti J, Canto M, Sivak M, Boyce HW, Lightdale CJ, Stevens P, Schmitt C. Clinical implications of endoscopic ultrasound: the American Endosonography Club Study. Gastrointest Endosc. 1996;44:371-377. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 83] [Cited by in RCA: 66] [Article Influence: 2.2] [Reference Citation Analysis (1)] |
| 20. | Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754-1760. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46154] [Cited by in RCA: 36919] [Article Influence: 2171.7] [Reference Citation Analysis (5)] |
| 21. | Koboldt DC, Zhang Q, Larson DE, Shen D, McLellan MD, Lin L, Miller CA, Mardis ER, Ding L, Wilson RK. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 2012;22:568-576. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4444] [Cited by in RCA: 3765] [Article Influence: 268.9] [Reference Citation Analysis (5)] |
| 22. | Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12850] [Cited by in RCA: 11256] [Article Influence: 703.5] [Reference Citation Analysis (5)] |
| 23. | Bamford S, Dawson E, Forbes S, Clements J, Pettett R, Dogan A, Flanagan A, Teague J, Futreal PA, Stratton MR, Wooster R. The COSMIC (Catalogue of Somatic Mutations in Cancer) database and website. Br J Cancer. 2004;91:355-358. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 862] [Cited by in RCA: 986] [Article Influence: 44.8] [Reference Citation Analysis (1)] |
| 24. | Assarzadegan N, Montgomery E. What is New in the 2019 World Health Organization (WHO) Classification of Tumors of the Digestive System: Review of Selected Updates on Neuroendocrine Neoplasms, Appendiceal Tumors, and Molecular Testing. Arch Pathol Lab Med. 2021;145:664-677. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 150] [Cited by in RCA: 135] [Article Influence: 27.0] [Reference Citation Analysis (1)] |
| 25. | Mas L, Lupinacci RM, Cros J, Bachet JB, Coulet F, Svrcek M. Intraductal Papillary Mucinous Carcinoma Versus Conventional Pancreatic Ductal Adenocarcinoma: A Comprehensive Review of Clinical-Pathological Features, Outcomes, and Molecular Insights. Int J Mol Sci. 2021;22:6756. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 27] [Article Influence: 5.4] [Reference Citation Analysis (1)] |
| 26. | Chandwani R, Allen PJ. Cystic Neoplasms of the Pancreas. Annu Rev Med. 2016;67:45-57. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 35] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 27. | King JC, Ng TT, White SC, Cortina G, Reber HA, Hines OJ. Pancreatic serous cystadenocarcinoma: a case report and review of the literature. J Gastrointest Surg. 2009;13:1864-1868. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 68] [Cited by in RCA: 55] [Article Influence: 3.2] [Reference Citation Analysis (1)] |
| 28. | Andea A, Sarkar F, Adsay VN. Clinicopathological correlates of pancreatic intraepithelial neoplasia: a comparative analysis of 82 cases with and 152 cases without pancreatic ductal adenocarcinoma. Mod Pathol. 2003;16:996-1006. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 238] [Cited by in RCA: 203] [Article Influence: 8.8] [Reference Citation Analysis (1)] |
| 29. | Makohon-Moore A, Iacobuzio-Donahue CA. Pancreatic cancer biology and genetics from an evolutionary perspective. Nat Rev Cancer. 2016;16:553-565. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 281] [Cited by in RCA: 326] [Article Influence: 32.6] [Reference Citation Analysis (1)] |
| 30. | Hruban RH, Goggins M, Parsons J, Kern SE. Progression model for pancreatic cancer. Clin Cancer Res. 2000;6:2969-2972. [PubMed] |
| 31. | Notta F, Chan-Seng-Yue M, Lemire M, Li Y, Wilson GW, Connor AA, Denroche RE, Liang SB, Brown AM, Kim JC, Wang T, Simpson JT, Beck T, Borgida A, Buchner N, Chadwick D, Hafezi-Bakhtiari S, Dick JE, Heisler L, Hollingsworth MA, Ibrahimov E, Jang GH, Johns J, Jorgensen LG, Law C, Ludkovski O, Lungu I, Ng K, Pasternack D, Petersen GM, Shlush LI, Timms L, Tsao MS, Wilson JM, Yung CK, Zogopoulos G, Bartlett JM, Alexandrov LB, Real FX, Cleary SP, Roehrl MH, McPherson JD, Stein LD, Hudson TJ, Campbell PJ, Gallinger S. A renewed model of pancreatic cancer evolution based on genomic rearrangement patterns. Nature. 2016;538:378-382. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 443] [Cited by in RCA: 423] [Article Influence: 42.3] [Reference Citation Analysis (5)] |
| 32. | Yachida S, Jones S, Bozic I, Antal T, Leary R, Fu B, Kamiyama M, Hruban RH, Eshleman JR, Nowak MA, Velculescu VE, Kinzler KW, Vogelstein B, Iacobuzio-Donahue CA. Distant metastasis occurs late during the genetic evolution of pancreatic cancer. Nature. 2010;467:1114-1117. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2120] [Cited by in RCA: 1962] [Article Influence: 122.6] [Reference Citation Analysis (5)] |
| 33. | Kanda M, Matthaei H, Wu J, Hong SM, Yu J, Borges M, Hruban RH, Maitra A, Kinzler K, Vogelstein B, Goggins M. Presence of somatic mutations in most early-stage pancreatic intraepithelial neoplasia. Gastroenterology. 2012;142:730-733.e9. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 622] [Cited by in RCA: 590] [Article Influence: 42.1] [Reference Citation Analysis (1)] |
| 34. | Hayashi A, Hong J, Iacobuzio-Donahue CA. The pancreatic cancer genome revisited. Nat Rev Gastroenterol Hepatol. 2021;18:469-481. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 179] [Cited by in RCA: 154] [Article Influence: 30.8] [Reference Citation Analysis (1)] |
| 35. | Biankin AV, Waddell N, Kassahn KS, Gingras MC, Muthuswamy LB, Johns AL, Miller DK, Wilson PJ, Patch AM, Wu J, Chang DK, Cowley MJ, Gardiner BB, Song S, Harliwong I, Idrisoglu S, Nourse C, Nourbakhsh E, Manning S, Wani S, Gongora M, Pajic M, Scarlett CJ, Gill AJ, Pinho AV, Rooman I, Anderson M, Holmes O, Leonard C, Taylor D, Wood S, Xu Q, Nones K, Fink JL, Christ A, Bruxner T, Cloonan N, Kolle G, Newell F, Pinese M, Mead RS, Humphris JL, Kaplan W, Jones MD, Colvin EK, Nagrial AM, Humphrey ES, Chou A, Chin VT, Chantrill LA, Mawson A, Samra JS, Kench JG, Lovell JA, Daly RJ, Merrett ND, Toon C, Epari K, Nguyen NQ, Barbour A, Zeps N; Australian Pancreatic Cancer Genome Initiative, Kakkar N, Zhao F, Wu YQ, Wang M, Muzny DM, Fisher WE, Brunicardi FC, Hodges SE, Reid JG, Drummond J, Chang K, Han Y, Lewis LR, Dinh H, Buhay CJ, Beck T, Timms L, Sam M, Begley K, Brown A, Pai D, Panchal A, Buchner N, De Borja R, Denroche RE, Yung CK, Serra S, Onetto N, Mukhopadhyay D, Tsao MS, Shaw PA, Petersen GM, Gallinger S, Hruban RH, Maitra A, Iacobuzio-Donahue CA, Schulick RD, Wolfgang CL, Morgan RA, Lawlor RT, Capelli P, Corbo V, Scardoni M, Tortora G, Tempero MA, Mann KM, Jenkins NA, Perez-Mancera PA, Adams DJ, Largaespada DA, Wessels LF, Rust AG, Stein LD, Tuveson DA, Copeland NG, Musgrove EA, Scarpa A, Eshleman JR, Hudson TJ, Sutherland RL, Wheeler DA, Pearson JV, McPherson JD, Gibbs RA, Grimmond SM. Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature. 2012;491:399-405. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1782] [Cited by in RCA: 1689] [Article Influence: 120.6] [Reference Citation Analysis (5)] |
| 36. | Roberts NJ, Norris AL, Petersen GM, Bondy ML, Brand R, Gallinger S, Kurtz RC, Olson SH, Rustgi AK, Schwartz AG, Stoffel E, Syngal S, Zogopoulos G, Ali SZ, Axilbund J, Chaffee KG, Chen YC, Cote ML, Childs EJ, Douville C, Goes FS, Herman JM, Iacobuzio-Donahue C, Kramer M, Makohon-Moore A, McCombie RW, McMahon KW, Niknafs N, Parla J, Pirooznia M, Potash JB, Rhim AD, Smith AL, Wang Y, Wolfgang CL, Wood LD, Zandi PP, Goggins M, Karchin R, Eshleman JR, Papadopoulos N, Kinzler KW, Vogelstein B, Hruban RH, Klein AP. Whole Genome Sequencing Defines the Genetic Heterogeneity of Familial Pancreatic Cancer. Cancer Discov. 2016;6:166-175. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 317] [Cited by in RCA: 291] [Article Influence: 29.1] [Reference Citation Analysis (5)] |
| 37. | Balachandran VP, Łuksza M, Zhao JN, Makarov V, Moral JA, Remark R, Herbst B, Askan G, Bhanot U, Senbabaoglu Y, Wells DK, Cary CIO, Grbovic-Huezo O, Attiyeh M, Medina B, Zhang J, Loo J, Saglimbeni J, Abu-Akeel M, Zappasodi R, Riaz N, Smoragiewicz M, Kelley ZL, Basturk O; Australian Pancreatic Cancer Genome Initiative; Garvan Institute of Medical Research; Prince of Wales Hospital; Royal North Shore Hospital; University of Glasgow; St Vincent’s Hospital; QIMR Berghofer Medical Research Institute; University of Melbourne, Centre for Cancer Research; University of Queensland, Institute for Molecular Bioscience; Bankstown Hospital; Liverpool Hospital; Royal Prince Alfred Hospital, Chris O’Brien Lifehouse; Westmead Hospital; Fremantle Hospital; St John of God Healthcare; Royal Adelaide Hospital; Flinders Medical Centre; Envoi Pathology; Princess Alexandria Hospital; Austin Hospital; Johns Hopkins Medical Institutes; ARC-Net Centre for Applied Research on Cancer, Gönen M, Levine AJ, Allen PJ, Fearon DT, Merad M, Gnjatic S, Iacobuzio-Donahue CA, Wolchok JD, DeMatteo RP, Chan TA, Greenbaum BD, Merghoub T, Leach SD. Identification of unique neoantigen qualities in long-term survivors of pancreatic cancer. Nature. 2017;551:512-516. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1030] [Cited by in RCA: 935] [Article Influence: 103.9] [Reference Citation Analysis (5)] |
| 38. | Thomas D, Sagar S, Liu X, Lee HR, Grunkemeyer JA, Grandgenett PM, Caffrey T, O'Connell KA, Swanson B, Marcos-Silva L, Steentoft C, Wandall HH, Maurer HC, Peng XL, Yeh JJ, Qiu F, Yu F, Madiyalakan R, Olive KP, Mandel U, Clausen H, Hollingsworth MA, Radhakrishnan P. Isoforms of MUC16 activate oncogenic signaling through EGF receptors to enhance the progression of pancreatic cancer. Mol Ther. 2021;29:1557-1571. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 56] [Cited by in RCA: 55] [Article Influence: 11.0] [Reference Citation Analysis (1)] |
| 39. | Shimizu A, Hirono S, Tani M, Kawai M, Okada K, Miyazawa M, Kitahata Y, Nakamura Y, Noda T, Yokoyama S, Yamaue H. Coexpression of MUC16 and mesothelin is related to the invasion process in pancreatic ductal adenocarcinoma. Cancer Sci. 2012;103:739-746. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 93] [Cited by in RCA: 102] [Article Influence: 7.3] [Reference Citation Analysis (1)] |
| 40. | Liang C, Shi S, Meng Q, Liang D, Hua J, Qin Y, Zhang B, Xu J, Ni Q, Yu X. MiR-29a, targeting caveolin 2 expression, is responsible for limitation of pancreatic cancer metastasis in patients with normal level of serum CA125. Int J Cancer. 2018;143:2919-2931. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 24] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 41. | Collisson EA, Sadanandam A, Olson P, Gibb WJ, Truitt M, Gu S, Cooc J, Weinkle J, Kim GE, Jakkula L, Feiler HS, Ko AH, Olshen AB, Danenberg KL, Tempero MA, Spellman PT, Hanahan D, Gray JW. Subtypes of pancreatic ductal adenocarcinoma and their differing responses to therapy. Nat Med. 2011;17:500-503. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1688] [Cited by in RCA: 1481] [Article Influence: 98.7] [Reference Citation Analysis (4)] |
| 42. | Moffitt RA, Marayati R, Flate EL, Volmar KE, Loeza SG, Hoadley KA, Rashid NU, Williams LA, Eaton SC, Chung AH, Smyla JK, Anderson JM, Kim HJ, Bentrem DJ, Talamonti MS, Iacobuzio-Donahue CA, Hollingsworth MA, Yeh JJ. Virtual microdissection identifies distinct tumor- and stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nat Genet. 2015;47:1168-1178. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1898] [Cited by in RCA: 1686] [Article Influence: 153.3] [Reference Citation Analysis (5)] |
| 43. | Bailey P, Chang DK, Nones K, Johns AL, Patch AM, Gingras MC, Miller DK, Christ AN, Bruxner TJ, Quinn MC, Nourse C, Murtaugh LC, Harliwong I, Idrisoglu S, Manning S, Nourbakhsh E, Wani S, Fink L, Holmes O, Chin V, Anderson MJ, Kazakoff S, Leonard C, Newell F, Waddell N, Wood S, Xu Q, Wilson PJ, Cloonan N, Kassahn KS, Taylor D, Quek K, Robertson A, Pantano L, Mincarelli L, Sanchez LN, Evers L, Wu J, Pinese M, Cowley MJ, Jones MD, Colvin EK, Nagrial AM, Humphrey ES, Chantrill LA, Mawson A, Humphris J, Chou A, Pajic M, Scarlett CJ, Pinho AV, Giry-Laterriere M, Rooman I, Samra JS, Kench JG, Lovell JA, Merrett ND, Toon CW, Epari K, Nguyen NQ, Barbour A, Zeps N, Moran-Jones K, Jamieson NB, Graham JS, Duthie F, Oien K, Hair J, Grützmann R, Maitra A, Iacobuzio-Donahue CA, Wolfgang CL, Morgan RA, Lawlor RT, Corbo V, Bassi C, Rusev B, Capelli P, Salvia R, Tortora G, Mukhopadhyay D, Petersen GM; Australian Pancreatic Cancer Genome Initiative, Munzy DM, Fisher WE, Karim SA, Eshleman JR, Hruban RH, Pilarsky C, Morton JP, Sansom OJ, Scarpa A, Musgrove EA, Bailey UM, Hofmann O, Sutherland RL, Wheeler DA, Gill AJ, Gibbs RA, Pearson JV, Waddell N, Biankin AV, Grimmond SM. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature. 2016;531:47-52. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3189] [Cited by in RCA: 2792] [Article Influence: 279.2] [Reference Citation Analysis (4)] |
| 44. | Furlow B. Computed Tomography of Pancreatitis and Pancreatic Cancer. Radiol Technol. 2015;86:645CT-664CT; quiz 665CT. [PubMed] |
| 45. | Freeman HJ. Intraductal papillary mucinous neoplasms and other pancreatic cystic lesions. World J Gastroenterol. 2008;14:2977-2979. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 7] [Cited by in RCA: 6] [Article Influence: 0.3] [Reference Citation Analysis (1)] |
| 46. | Allen PJ, Qin LX, Tang L, Klimstra D, Brennan MF, Lokshin A. Pancreatic cyst fluid protein expression profiling for discriminating between serous cystadenoma and intraductal papillary mucinous neoplasm. Ann Surg. 2009;250:754-760. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 42] [Article Influence: 2.5] [Reference Citation Analysis (1)] |
| 47. | Mori Y, Ohtsuka T, Tamura K, Ideno N, Aso T, Kono H, Nagayoshi Y, Ueda J, Takahata S, Aishima S, Ookubo F, Oda Y, Tanaka M. Intraoperative irrigation cytology of the remnant pancreas to detect remnant distinct pancreatic ductal adenocarcinoma in patients with intraductal papillary mucinous neoplasm undergoing partial pancreatectomy. Surgery. 2014;155:67-73. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 12] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 48. | Olson MT, Aguilar EN, Brooks CL, Isder CC, Muilenburg KM, Talmon GA, Ly QP, Carlson MA, Hollingsworth MA, Mohs AM. Preclinical Evaluation of a Humanized, Near-Infrared Fluorescent Antibody for Fluorescence-Guided Surgery of MUC16-Expressing Pancreatic Cancer. Mol Pharm. 2022;19:3586-3599. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 11] [Article Influence: 2.8] [Reference Citation Analysis (1)] |
| 49. | Babeker H, Ketchemen JP, Annan Sudarsan A, Andrahennadi S, Tikum AF, Nambisan AK, Fonge H, Uppalapati M. Engineering of a Fully Human Anti-MUC-16 Antibody and Evaluation as a PET Imaging Agent. Pharmaceutics. 2022;14:2824. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 10] [Reference Citation Analysis (1)] |