Published online Apr 15, 2026. doi: 10.4251/wjgo.v18.i4.115635
Revised: December 4, 2025
Accepted: February 4, 2026
Published online: April 15, 2026
Processing time: 169 Days and 4.1 Hours
Lymph node metastasis (LNM) is closely linked to poor prognosis in patients with colorectal cancer (CRC). An accurate preoperative evaluation of lymph node status is crucial for tailoring individualized treatment plans. However, depen
To develop a nomogram combining clinical features, radiomics and deep learning (DL) features for predicting LNM and prognosis in patients with resectable CRC.
Two hundred and seventy-eight patients with pathologically confirmed CRC from two hospitals were retrospectively enrolled. Radiomics and DL features were extracted from preoperative three-phase contrast-enhanced computed tomography images within the intratumoral and peritumoral-3 mm regions. Using logistic regression combined with three feature selection methods, a clinical-DL-radiomics nomogram (DLRN) was developed. SHapley Additive exPlanations were employed to interpret the model. Additionally, Cox regression analysis was performed to identify risk factors for 3-year recurrence-free survival and to construct a prognostic model.
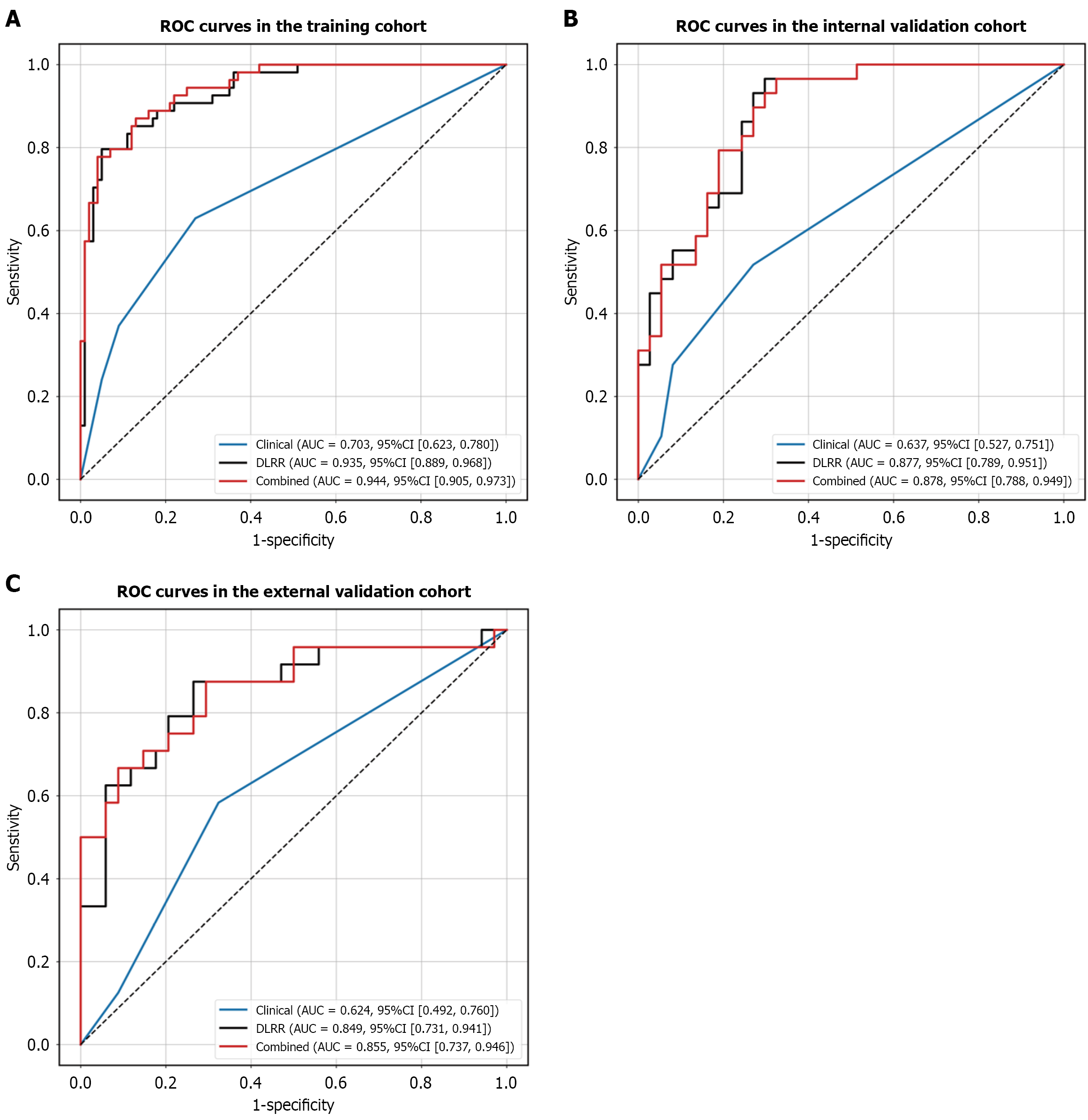
The DLRN demonstrated significantly superior performance in differentiating LNM compared to both the clinical model and the radiomics model. The area under the receiver operating characteristic curve for the DLRN was 0.944, 0.878, and 0.855 in the training, internal validation, and external validation cohorts, respectively. Calibration curves and decision curve analysis confirmed that the DLRN exhibited excellent calibration and strong clinical utility. Furthermore, the prognostic model based on the DLRN score showed robust performance in predicting 3-year recurrence-free survival, achieving an area under the receiver operating characteristic curves of 0.826, 0.788, and 0.755 across the three cohorts, respectively.
The DLRN demonstrated excellent performance in predicting LNM in CRC. Additionally, the prognostic model derived from the DLRN score effectively stratified patients according to their risk of recurrence.
Core Tip: Accurate preoperative prediction of lymph node metastasis is crucial for optimizing treatment strategies in colorectal cancer. In this study, we developed an interpretable clinical-deep learning-radiomics nomogram (DLRN) by integrating clinical features with multi-regional radiomics and deep learning features. Moreover, the DLRN-based prognostic model effectively predicted 3-year recurrence-free survival. As a noninvasive preoperative tool, the DLRN demonstrated strong predictive accuracy for lymph node metastasis in colorectal cancer and offers a practical means for individualized risk stratification and informed treatment decision-making.
- Citation: Lei XD, Qian GX, Sun ZG, Tang ZQ, Liu YC, Du R, Li YH. Deep learning radiomics nomogram based on multi-regional features for predicting lymph node metastasis and prognosis in colorectal cancer. World J Gastrointest Oncol 2026; 18(4): 115635
- URL: https://www.wjgnet.com/1948-5204/full/v18/i4/115635.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v18.i4.115635
Colorectal cancer (CRC) is one of the most prevalent malignant tumors encountered in clinical practice. According to the 2022 Global Cancer Statistics Report, CRC ranks as the third most common malignancy worldwide and is the second leading cause of cancer-related mortality[1]. Lymph node metastasis (LNM) represents the most frequent metastatic route in CRC and is a critical factor contributing to postoperative recurrence and patient mortality[2]. Importantly, the presence or absence of LNM not only directly influences the extent of surgical intervention and the formulation of adjuvant therapy regimens but also significantly affects long-term patient survival[3,4]. Therefore, accurate preoperative assessment of lymph node status holds considerable clinical significance for optimizing personalized treatment strategies and prognostic evaluation in CRC patients.
Currently, contrast-enhanced computed tomography (CECT) is a commonly used method for the preoperative assessment of LNM in CRC. However, its accuracy in detecting LNM is limited by several factors, including equipment performance, the standardization of imaging protocols, the expertise of radiologists, and individual patient variability. Radiomics, a rapidly evolving field, shows promising potential for LNM prediction by extracting high-throughput quantitative features from medical images and analyzing them[5,6]. Concurrently, deep learning (DL), a key branch of machine learning, can automatically extract and characterize deep-level tumor image features in an end-to-end manner[7], achieving remarkable results in tumor diagnosis and prognostic prediction[8-10]. Several studies have demonstrated that although radiomics and DL individually possess considerable classification capabilities, they each excel at capturing different aspects of image information, radiomics in texture and morphology, and DL in high-level semantic features, and their combination often yields superior performance[11,12]. Nevertheless, current research on LNM prediction in CRC primarily focuses on the independent use of either radiomics or DL features, with limited exploration of their effective integration.
CRC exhibits substantial spatial heterogeneity, encompassing both the tumor itself (intratumoral region) and the surrounding peritumoral area. Although peritumoral tissue may appear similar to normal tissue macroscopically, studies have revealed significant microstructural heterogeneity within this region[13]. Moreover, tumor cells have a biological propensity to infiltrate and migrate into the peritumoral area, suggesting that this region may harbor critical information related to LNM[14,15]. With the advancement of radiomics, analysis of peritumoral features has become increasingly prevalent in tumor prediction research and has demonstrated considerable value in LNM prediction[16,17]. For instance, Yuan et al[18] reported the potential of computed tomography-based intratumoral and peritumoral radiomics models for preoperative LNM prediction in rectal cancer; however, their study lacked a comprehensive external validation cohort, limiting its clinical applicability.
Therefore, this study aims to develop and validate a comprehensive nomogram [DL-radiomics nomogram (DLRN)] that integrates clinical features with multi-regional radiomics and DL features for the preoperative prediction of LNM in CRC patients. Furthermore, building upon this predictive model, a prognostic model for 3-year recurrence-free survival (RFS) is constructed to provide a reliable tool for individualized clinical decision-making and risk stratification.
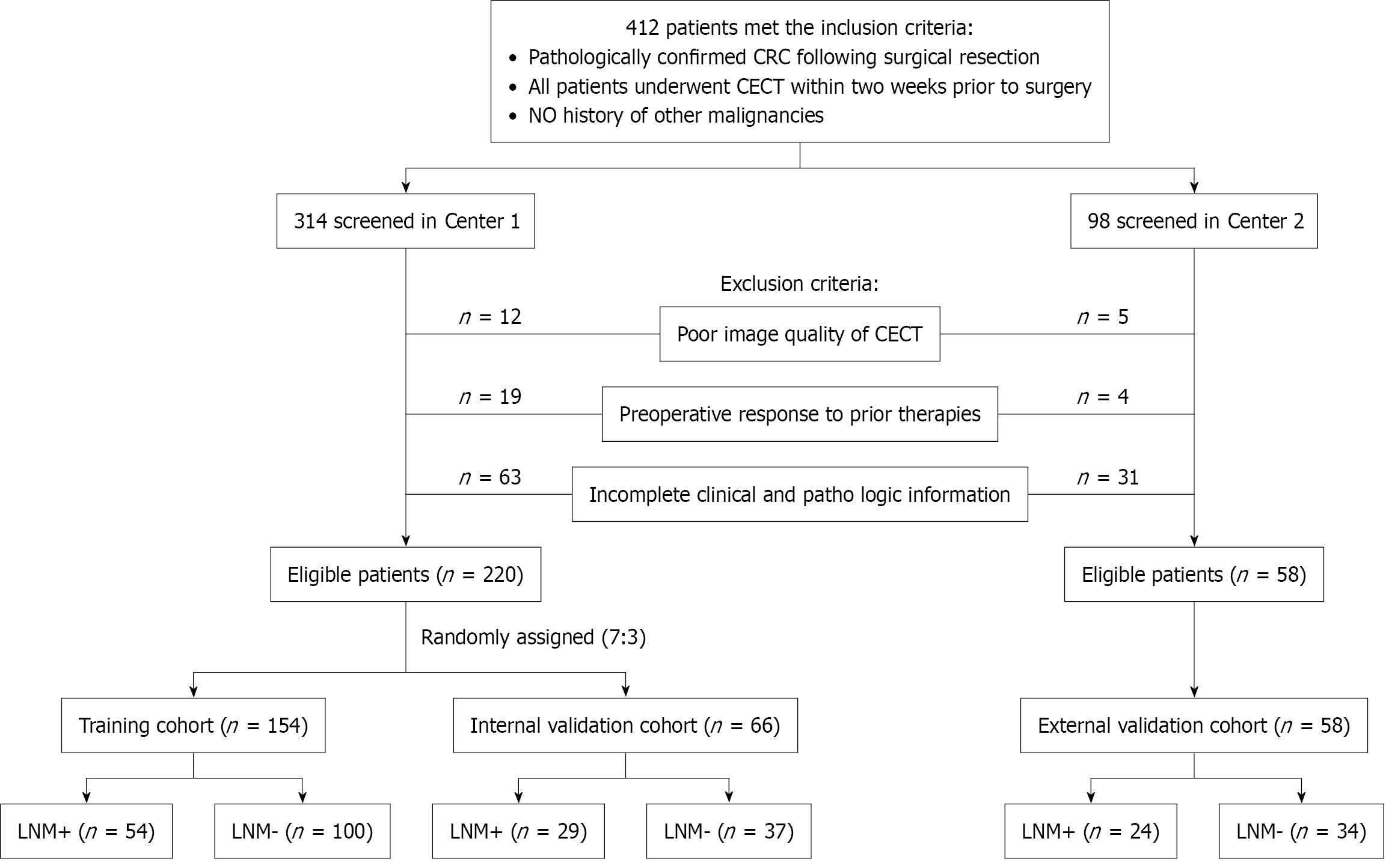
This retrospective study was approved by our institutional review board, with a waiver of informed consent, approval No. 2025-158-01. Between January 1, 2018, and December 31, 2021, a total of 412 patients with resectable CRC were consecutively enrolled from the First People’s Hospital of Hefei (Center 1) and Binhu Hospital of Hefei (Center 2). Inclusion criteria were: (1) Pathologically confirmed CRC following surgery; (2) CECT performed within two weeks prior to surgery; and (3) No history of other malignant tumors. Exclusion criteria included: (1) Poor CECT image quality due to bias, motion artifacts, or low resolution; (2) Receipt of neoadjuvant chemoradiotherapy or other treatments before surgery; and (3) Incomplete clinical or pathological data. Ultimately, 220 patients from Center 1 and 58 patients from Center 2 met the criteria and were included. The 220 patients from Center 1 were randomly divided into a training cohort (n = 154) and an internal validation cohort (n = 66) at a 7:3 ratio, while the 58 patients from Center 2 formed the external validation cohort. The study population flowchart is presented in Figure 1.
Arterial, venous, and delayed phase CECT images were retrieved from the picture archiving and communication system for further analysis. Detailed CT acquisition parameters for each center are provided in Table 1. A non-ionic iodinated contrast agent (300-400 mgI/mL) was administered intravenously at a dose of 1-1.5 mL/kg, with an injection rate of 3 mL/second. After contrast administration, contrast images were obtained at the arterial phase (25-35 seconds), portal venous phase (65-80 seconds), and delayed phase (3-5 minutes). To minimize batch effects arising from different scanners, ComBat harmonization was applied. Additionally, to standardize spatial resolution and intensity across datasets, all original CECT images were resampled to an isotropic voxel size of 1 mm × 1 mm × 0.625 mm (slice thickness 0.625 mm), and z-score normalization was performed on image intensities to reduce scanner-related variability. Two experienced radiologists independently evaluated the CECT images: Radiologist A, with 5 years of experience in CRC imaging, and radiologist B, with 12 years of experience. Both were blinded to clinical and pathological information. In cases of disagreement, a senior radiologist (radiologist C), with over 20 years of expertise in CRC CECT diagnosis, reviewed the conflicting cases and facilitated a consensus decision.
| Center | Scanner | Field of view (cm) | Image matrix (mm) | Tube voltage (kv) | Tube current (mA) | Pixel size (mm) | Slice thickness (mm) |
| 1 | Revolution CT (256-slice) | 50 | 512 × 512 | 70-140 | Auto | 0.977 | 0.625-5 |
| Discovery CT750 HD (Gemstone) | 50 | 512 × 512 | 80/100/120/140 | Auto | 0.977 | 0.625-5 | |
| 2 | SOMATOM definition flash (dual-source CT) | 50 | 512 × 512 | 70-140 | Auto | 0.977 | 0.625-5 |
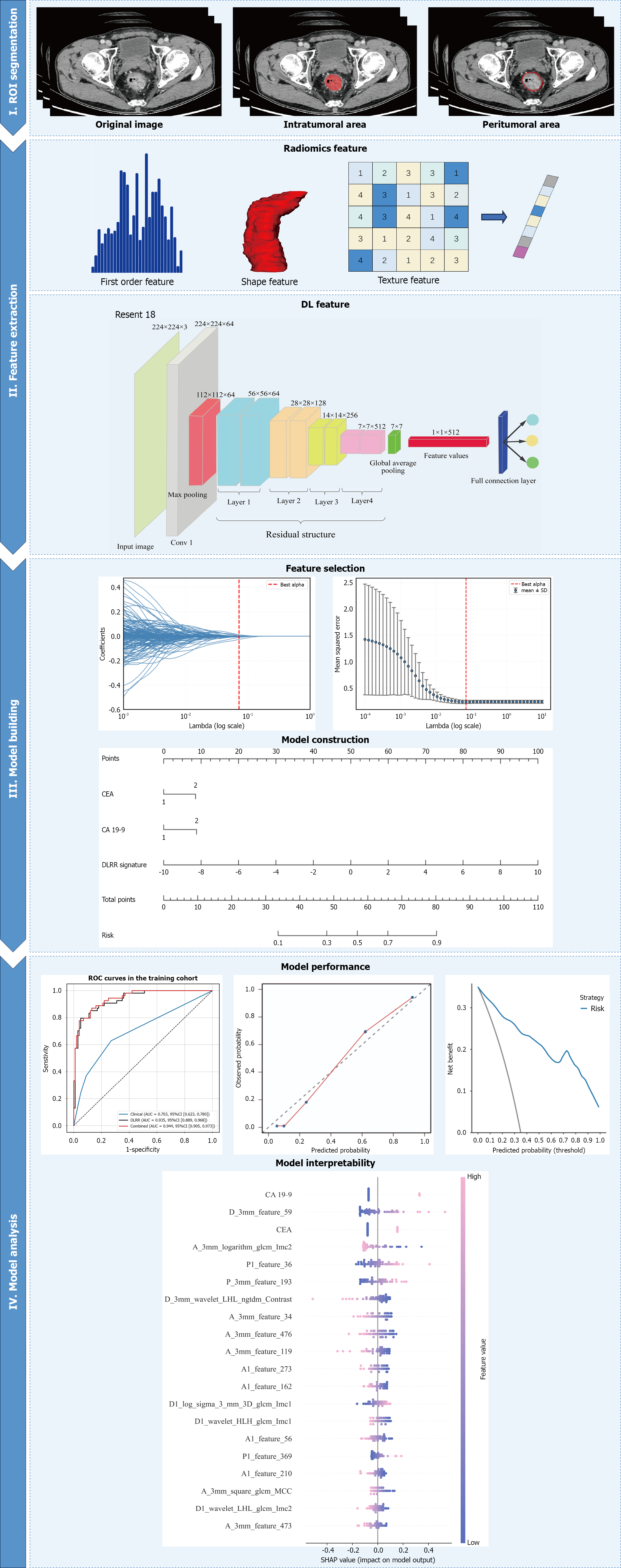
The radiomics workflow is illustrated in Figure 2. Radiologist A manually segmented the three-dimensional (3D) region of interest of the tumor using ITK-SNAP software (version 3.8.0; www.itksnap.org). To capture information from the invasive margin, the peritumoral region was automatically generated by expanding the tumor boundary outward by 3 mm using Python-based morphological erosion and dilation algorithms, resulting in a 3 mm-thick ring around the tumor. When this ring overlapped with adjacent structures such as the bladder, kidney, or surrounding normal intestine, these areas were manually excluded. To assess the reproducibility of radiomics and DL feature extraction, radiologist A and radiologist B randomly selected 50 patients for repeated region of interest segmentation two weeks later, including both the intratumoral region and the automatically defined 3-mm peritumoral region. Intra-class and inter-class correlation coefficients (ICCs) were then calculated for the features derived from both regions to evaluate intra-observer and inter-observer reliability.
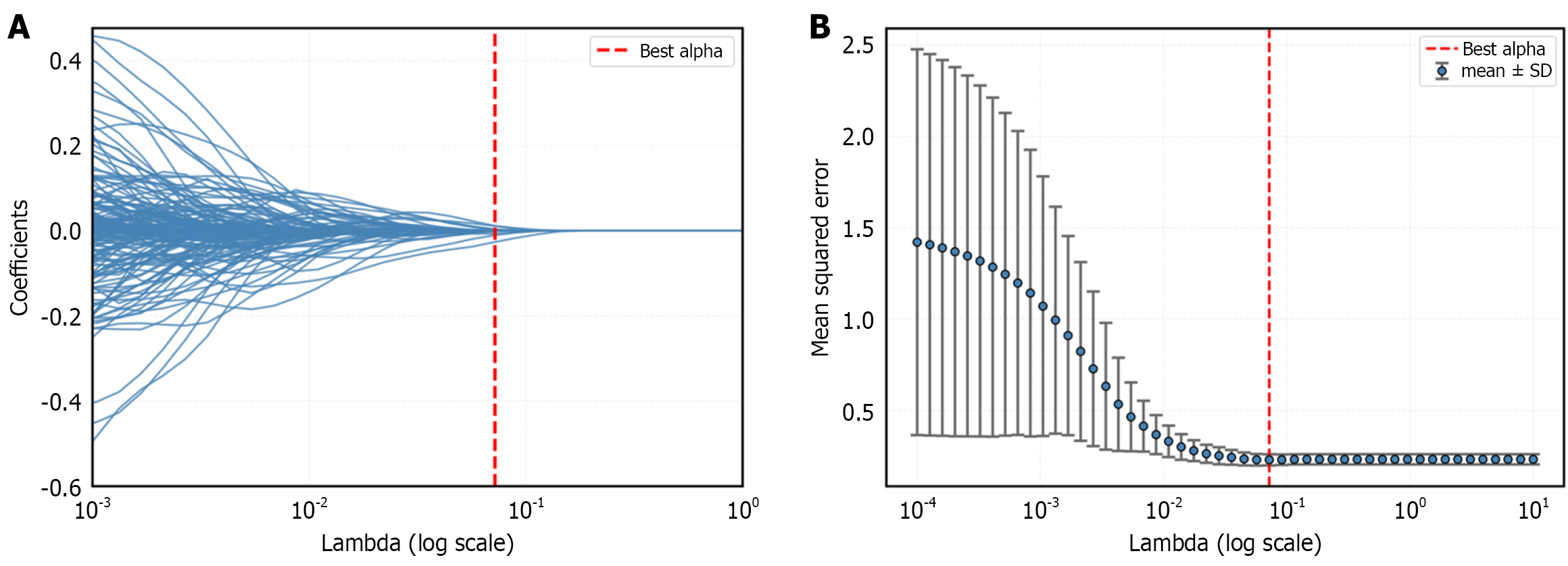
Radiomics feature extraction was conducted using the Pyradiomics package (version 2.2.0) in Python (version 3.7.1) on the CECT images. Radiomics features were extracted from each volume of interest and included first-order statistics, gray-level co-occurrence matrix, gray-level run length matrix, gray-level size zone matrix, gray-level dependence matrix, and shape features. Based on the training cohort data, the radiomics features were filtered and reduced: (1) All radiomics features were standardized using z-score normalization to eliminate scale differences; (2) Reproducibility was assessed using the ICC, and features with ICC < 0.85 were excluded; (3) The minimum redundancy-maximum relevance (mRMR) algorithm was applied to remove redundant and irrelevant features; and (4) The least absolute shrinkage and selection operator (LASSO) regression was used to identify the optimal feature subset within the training dataset. The selected features were then linearly combined and weighted by their respective LASSO coefficients to calculate a radiomics score (rad-score) for each patient, which was used to construct the radiomics model (R).
To address the limitations of a small sample size, DL features were extracted using a 3D ResNet-18 convolutional neural network model pre-trained on ImageNet. Based on the training cohort data, all DL features were standardized through z-score normalization. Feature selection involved ICC analysis, mRMR, and LASSO regression to identify the most relevant features. The DL-score was then calculated by linearly combining these selected features weighted by their LASSO coefficients. This process resulted in the construction of a DL radiomics (DLR) model.
First, univariate analysis was conducted on all clinical risk factors. Factors with a significance level of P < 0.100 were then included in a multivariate analysis to identify independent risk factors (P < 0.05). A clinical model was subsequently developed based on these independent predictors. By integrating radiomics and DL features, a DL-radiomics integrated model (DLRR) was constructed using logistic regression. Finally, a comprehensive clinical-DL-radiomics combined model was established by combining all three components.
Receiver operating characteristic curves and the corresponding area under the curve (AUC) values were used to assess the discriminative performance of the developed models across the training, internal validation, and external validation cohorts. Accuracy, sensitivity, specificity, positive predictive value, and negative predictive value were calculated based on the cutoff value determined by the Youden index in the training cohort. Following a comprehensive comparison of model performances, a nomogram was constructed using the optimal predictive model to facilitate clinical application. Calibration of the nomogram was evaluated using calibration curves and the Hosmer-Lemeshow goodness-of-fit test. Additionally, decision curve analysis was performed to assess the clinical utility of the nomogram.
Patients underwent regular follow-up after surgery. Those with postoperative pathology of T1-2N0M0 did not receive adjuvant therapy. Patients classified as T3-4N0M0 or T1-4N1-2M0 were administered adjuvant chemotherapy based on the XELOX regimen (capecitabine plus oxaliplatin). Recurrence was evaluated every 3 months to 6 months postoperatively through clinical assessments and imaging studies. The primary endpoint of the study was 3-year RFS. Follow-up concluded on December 31, 2024.
Statistical analyses were performed using IBM SPSS Statistics (version 29.0; IBM Corp., New York, NY, United States) and R software (version 4.3.2; http://www.r-project.org). Continuous variables are expressed as either means ± SD or medians with interquartile ranges, while categorical variables are shown as n (%). Independent sample t-tests were used for normally distributed variables, while the Mann-Whitney U test was applied to variables with skewed distributions. Categorical variables were analyzed using the χ² test or Fisher’s exact test, as appropriate. A two-tailed P value < 0.05 was considered statistically significant.
A total of 278 CRC patients who underwent surgical treatment and were pathologically confirmed postoperatively were included in this study. The clinical characteristics of all patients are summarized in Table 2. Among them, 107 patients developed LNM. There were no statistically significant differences among the training, internal validation, and external validation cohorts in terms of sex, age, height, weight, body mass index, location, low-density lipoprotein, C-reactive protein, carcinoembryonic antigen (CEA), and carbohydrate antigen 19-9 (CA 19-9). In the training cohort, CEA and CA 19-9 levels differed significantly between patients with and without LNM, suggesting that elevated CEA and CA 19-9 levels may be associated with LNM. However, other factors, including age, sex, and body mass index, showed no significant correlation with LNM in any cohort.
| Characteristic | Training cohort (n = 154) | Internal validation cohort (n = 66) | External validation cohort (n = 58) | P value | ||||||
| LNM (n = 54) | non-LNM (n = 100) | P value | LNM (n = 29) | non-LNM | P value | LNM (n = 24) | non-LNM | P value | ||
| Sex | 0.493 | - | 0.530 | - | 0.286 | 0.112 | ||||
| Male | 37 (68.5) | 63 (63.0) | 15 (51.7) | 22 (59.5) | 14 (58.3) | 15 (44.1) | ||||
| Female | 17 (31.5) | 37 (37.0) | 14 (48.3) | 15 (40.5) | 10 (41.7) | 19 (55.9) | ||||
| Age (years), median (IQR) | 64.50 (57.00, 74.00) | 69.50 (61.00,77.00) | 0.054 | 70.00 (58.50, 72.50) | 72.00 (62.50, 78.50) | 0.047 | 66.50 (58.25, 71.00) | 67.00 (62.75, 72.00) | 0.448 | 0.203 |
| Height (cm), median (IQR) | 165.00 (160.00, 170.25) | 165.00 (160.00, 170.00) | 0.968 | 165.00 (159.00,171.00) | 165.00 (158.00, 172.00) | 0.801 | 161.50 (156.75, 170.00) | 164.00 (155.00, 168.00) | 0.776 | 0.245 |
| Weight (kg), median (IQR) | 62.00 (54.75, 68.25) | 62.00 (55.00, 68.00) | 0.847 | 65.00 (61.00, 70.00) | 60.00 (53.00, 67.50) | 0.108 | 60.50 (52.50, 69.50) | 60.50 (54.50, 66.50) | 0.937 | 0.424 |
| BMI, median (IQR) | 22.98 (20.42, 24.37) | 22.63 (20.55, 24.65) | 0.913 | 23.44 (22.22, 24.67) | 22.21 (20.13, 25.37) | 0.272 | 23.14 (20.70, 24.93) | 22.99 (20.68, 24.93) | 0.819 | 0.616 |
| Location | 0.112 | - | 0.375 | - | 0.314 | 0.254 | ||||
| RCC | 21 (38.9) | 23 (23.0) | 6 (20.7) | 8 (21.6) | 6 (25.0) | 6 (17.6) | ||||
| LCC | 14 (25.9) | 31 (31.0) | 9 (31.0) | 17 (45.9) | 8 (33.3) | 7 (20.6) | ||||
| RC | 19 (35.2) | 46 (46.0) | 14 (48.3) | 12 (3.24) | 10 (41.7) | 21 (61.8) | ||||
| CRP (mg/L), median (IQR) | 2.06 (0.53, 7.02) | 3.19 (0.50, 11.82) | 0.740 | 2.39 (0.63, 8.17) | 5.00 (0.67, 8.43) | 0.725 | 5.00 (0.66, 10.99) | 0.87 (0.50, 8.39) | 0.068 | 0.595 |
| LDL (mmol/L), median (IQR) | 2.13 (1.61, 2.85) | 2.21 (1.85, 2.90) | 0.153 | 2.40 (1.91, 2.98) | 1.97 (1.55, 2.41) | 0.041 | 2.07 (1.72, 2.70) | 2.85 (2.24, 3.62) | 0.011 | 0.058 |
| CEA (ng/mL) | < 0.001 | - | 0.366 | - | 0.467 | 0.678 | ||||
| ≤ 5 | 27 (50.0) | 77 (77.0) | 19 (65.5) | 28 (75.7) | 14 (58.3) | 23 (67.6) | ||||
| > 5 | 27 (50.0) | 23 (23.0) | 10 (34.5) | 9 (24.3) | 10 (41.7) | 11 (32.4) | ||||
| CA 19-9 (U/mL) | < 0.001 | - | 0.035 | - | 0.651 | 0.333 | ||||
| ≤ 37 | 34 (63.0) | 91 (91.0) | 21 (72.4) | 34 (91.9) | 21 (87.5) | 31 (91.2) | ||||
| > 37 | 20 (37.0) | 9 (9.0) | 8 (27.6) | 3 (8.1) | 3 (12.5) | 3 (8.8) | ||||
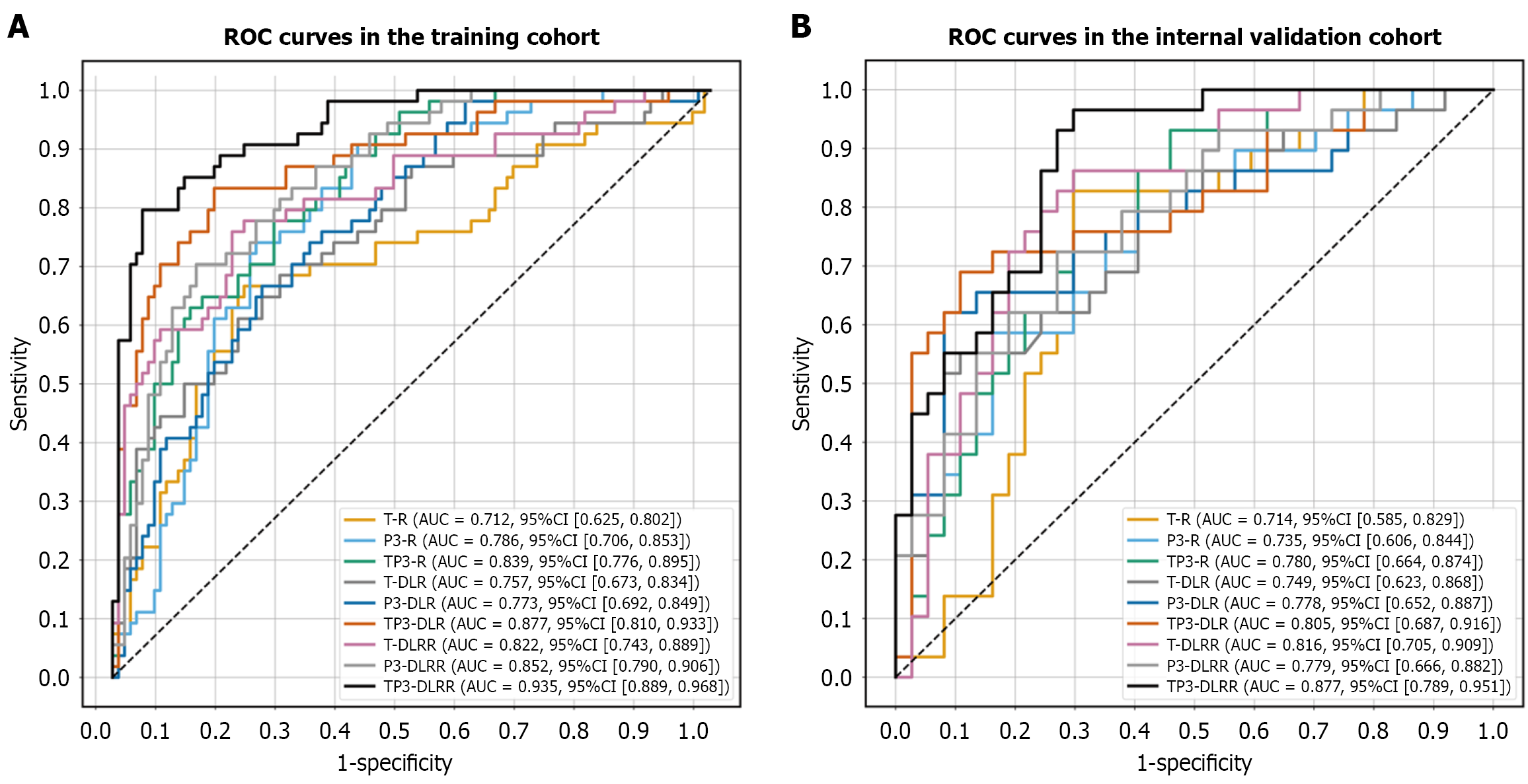
For feature extraction, a total of 12388 intratumoral and peritumoral-3 mm radiomics features were extracted from each 3D segmentation using Pyradiomics. Following a three-step feature selection process, comprising ICC, mRMR, and LASSO regression, 7 intratumoral and 7 peritumoral radiomics features were retained (Figure 3). Additionally, 3066 intratumoral and peritumoral-3 mm DL features were extracted using ResNet-18. After applying the same three-step selection process, 12 intratumoral and 7 peritumoral-3 mm DL features were selected. Based on these features, three models were constructed: A multi-regional radiomics model, a multi-regional DLR model, and a multi-regional DLRR model. The performance of each model is presented in Figure 4. The results demonstrated that the DLRR model, which integrates both intratumoral and peritumoral-3 mm features, achieved the best performance in the training and internal validation cohorts, with AUCs of 0.935 [95% confidence interval (CI): 0.889-0.968] and 0.877 (95%CI: 0.789-0.951), respectively.
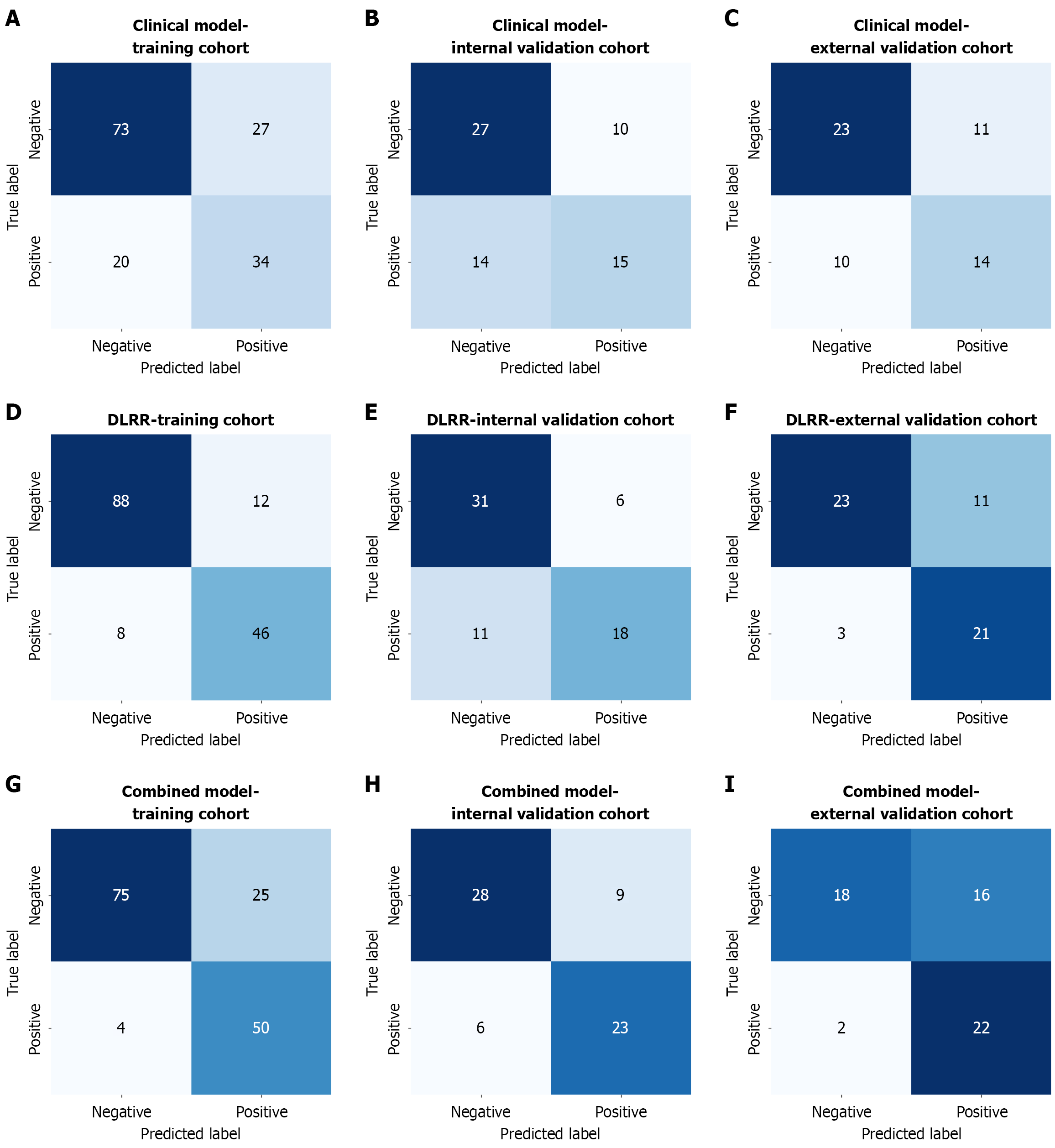
Univariate and multivariate regression analyses identified CEA and CA 19-9 as independent risk factors for LNM. Consequently, a clinical model was constructed based on these two factors (Table 3). Subsequently, the clinical factors were combined with radiomics and DL features to develop a combined predictive model. The performance of the clinical, DLRR, and combined models is presented in Figure 5 and Table 4. The combined model demonstrated superior predictive performance, with AUC values of 0.944 (95%CI: 0.905-0.973), 0.878 (95%CI: 0.788-0.949), and 0.855 (95%CI: 0.737-0.946) in the training, internal validation, and external validation cohorts, respectively. According to the DeLong test, the differences in AUCs between the combined model and the clinical model, as well as between the DLRR and clinical models, were statistically significant (P < 0.01 for all cohorts). However, no significant difference was observed between the combined model and the DLRR model in the training, internal validation, and external validation cohorts (P = 0.238, 0.940, and 0.630, respectively). Detailed classification results for each model are visualized using confusion matrices (Figure 6).
| Variable | Univariate analysis | Multivariate analysis | ||
| OR (95%CI) | P value | OR (95%CI) | P value | |
| Sex | ||||
| Male | 1 (Ref.) | - | - | - |
| Female | 0.782 (0.387-1.581) | 0.494 | - | - |
| Age | 0.975 (0.949-1.002) | 0.070 | 0.982 (0.953-1.012) | 0.235 |
| Height | 1.009 (0.962-1.060) | 0.703 | - | - |
| Weight | 0.997 (0.961-1.034) | 0.865 | - | - |
| BMI | 0.972 (0.869-1.008) | 0.624 | - | - |
| Location | ||||
| RCC | 1 (Ref.) | - | 1 (Ref) | - |
| LCC | 0.495 (0.208-1.175) | 0.111 | 0.552 (0.227-1.342) | 0.190 |
| RC | 0.452 (0.204-1.004) | 0.051 | 0.583 (0.228-1.492) | 0.261 |
| CRP | 0.997 (0.988-1.006) | 0.520 | - | - |
| LDL | 0.697 (0.445-1.094) | 0.117 | - | - |
| CEA | ||||
| ≤ 5 | 1 (Ref.) | - | 1 (Ref) | - |
| > 5 | 3.348 (1.649-6.797) | < 0.001 | 2.597 (1.199-5.623) | 0.015 |
| CA 19-9 | ||||
| ≤ 37 | 1 (Ref.) | - | 1 (Ref) | - |
| > 37 | 5.948 (2.467-14.337) | < 0.001 | 3.704 (1.435-9.563) | 0.007 |
| Cohort | Models | AUC | ACC | SEN | SPE | PPV | NPV |
| Training cohort | Clinical | 0.703 (0.623-0.780) | 0.695 | 0.630 | 0.730 | 0.557 | 0.785 |
| DLRR | 0.935 (0.889-0.968) | 0.870 | 0.852 | 0.880 | 0.793 | 0.917 | |
| Combined | 0.944 (0.905-0.973) | 0.812 | 0.926 | 0.750 | 0.667 | 0.949 | |
| Internal validation cohort | Clinical | 0.637 (0.527-0.751) | 0.636 | 0.517 | 0.730 | 0.600 | 0.659 |
| DLRR | 0.877 (0.789-0.951) | 0.742 | 0.621 | 0.838 | 0.750 | 0.738 | |
| Combined | 0.878 (0.788-0.949) | 0.773 | 0.793 | 0.757 | 0.719 | 0.824 | |
| External validation cohort | Clinical | 0.624 (0.492-0.760) | 0.638 | 0.583 | 0.676 | 0.560 | 0.697 |
| DLRR | 0.849 (0.731-0.941) | 0.759 | 0.875 | 0.676 | 0.656 | 0.885 | |
| Combined | 0.855 (0.737-0.946) | 0.690 | 0.917 | 0.529 | 0.579 | 0.900 |
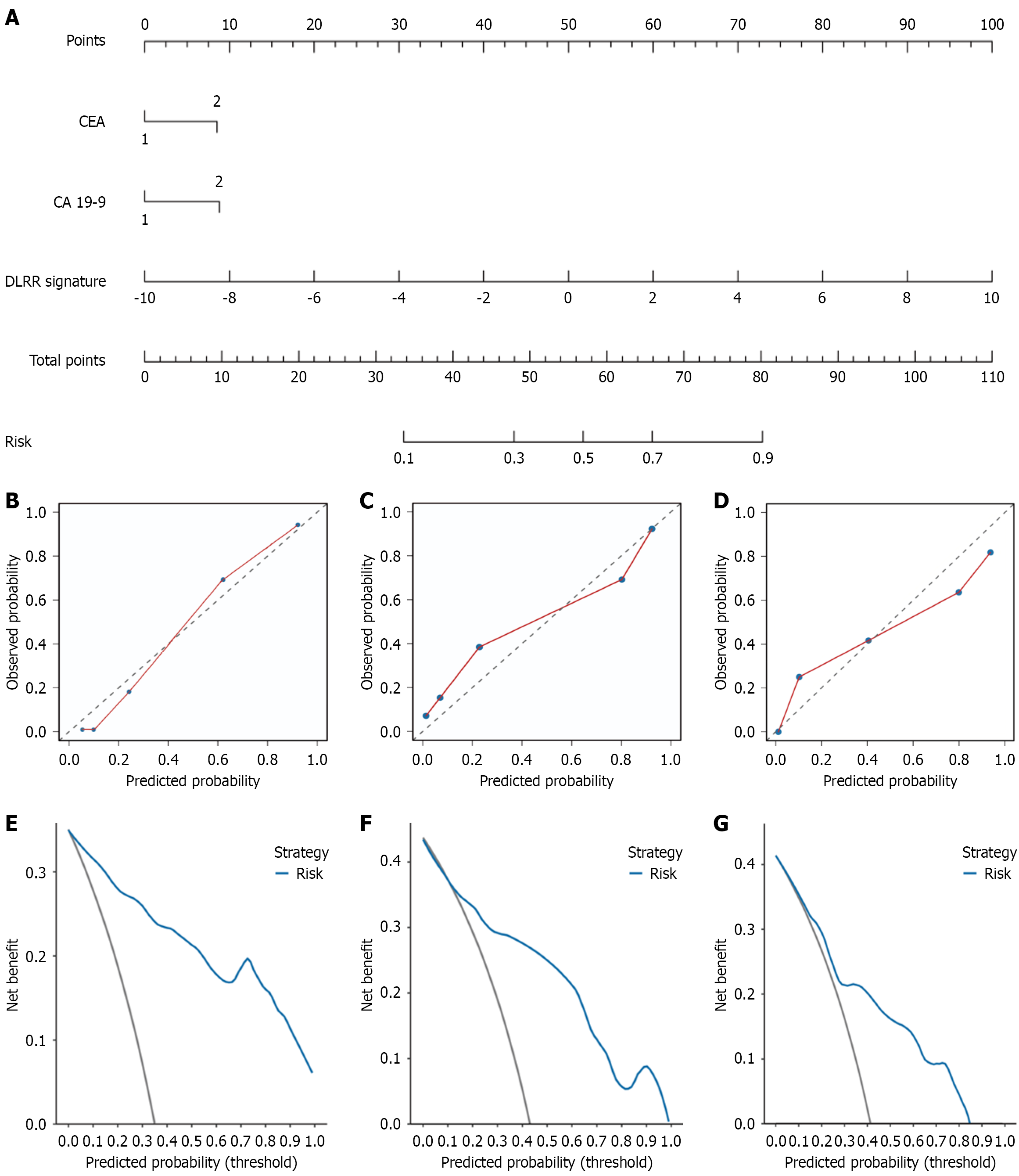
To visualize the combined model, a DLRN was constructed. Calibration curves were plotted to assess the agreement between predicted and observed outcomes (Figure 7A). The Hosmer-Lemeshow goodness-of-fit test demonstrated that the DLRN exhibited good calibration in the training, internal validation, and external validation cohorts, with P values of 0.775, 0.242, and 0.116, respectively (Figure 7B-D). Decision curve analysis further indicated that, across relevant threshold probabilities, the DLRN provided a greater net clinical benefit compared to other models (Figure 7E-G).
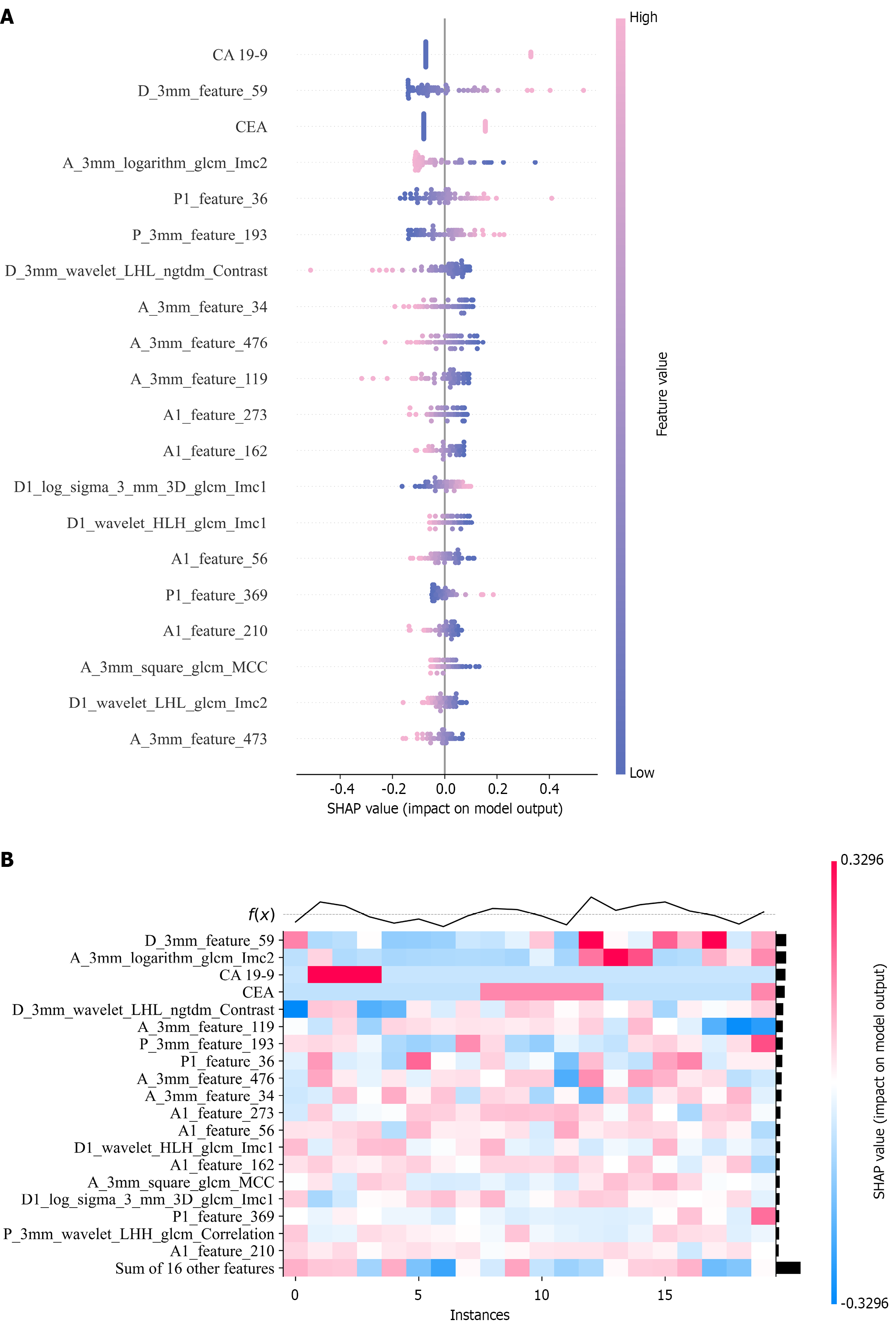
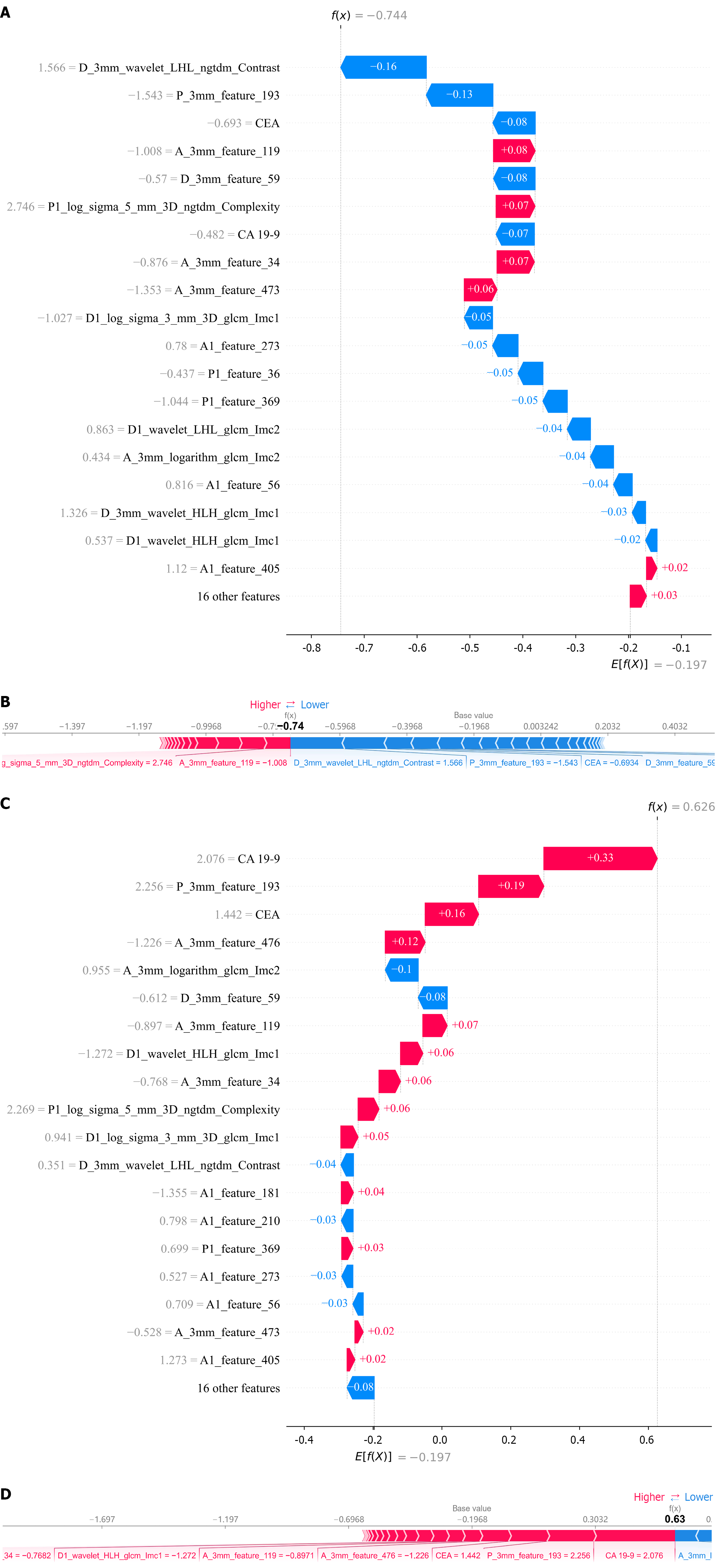
To evaluate the contribution of each feature to the model, SHapley Additive exPlanations (SHAP) were used to quantify the impact of individual features on the model’s predictions. In the overall visualization, the SHAP beeswarm plot (Figure 8A) highlighted the top 20 features contributing most to the model, with the four strongest predictors being CA 19-9, D_3mm_feature_59, CEA, and A_3mm_logarithm_glcm_Imc2. The SHAP heatmap (Figure 8B) illustrated the direction and magnitude of each feature’s influence across all instances in the dataset. For individual case interpretation, waterfall and force plots (Figure 9) demonstrated two representative examples of accurate predictions for LNM and non-LNM cases. Figure 9A and C depict a patient confirmed as non-LNM by postoperative pathology. The model assigned this patient a 74.4% probability of non-LNM, significantly higher than the baseline probability of 19.7%. Conversely, Figure 9B and D show a patient with pathologically confirmed LNM, for whom the model predicted a 62.6% probability of LNM, also substantially exceeding the baseline probability. The waterfall plots (Figure 9A) detail the individual contribution of each feature to the final prediction, while the force plots (Figure 9C and D) visually represent the direction and magnitude of each feature’s influence on the model’s output for these cases.
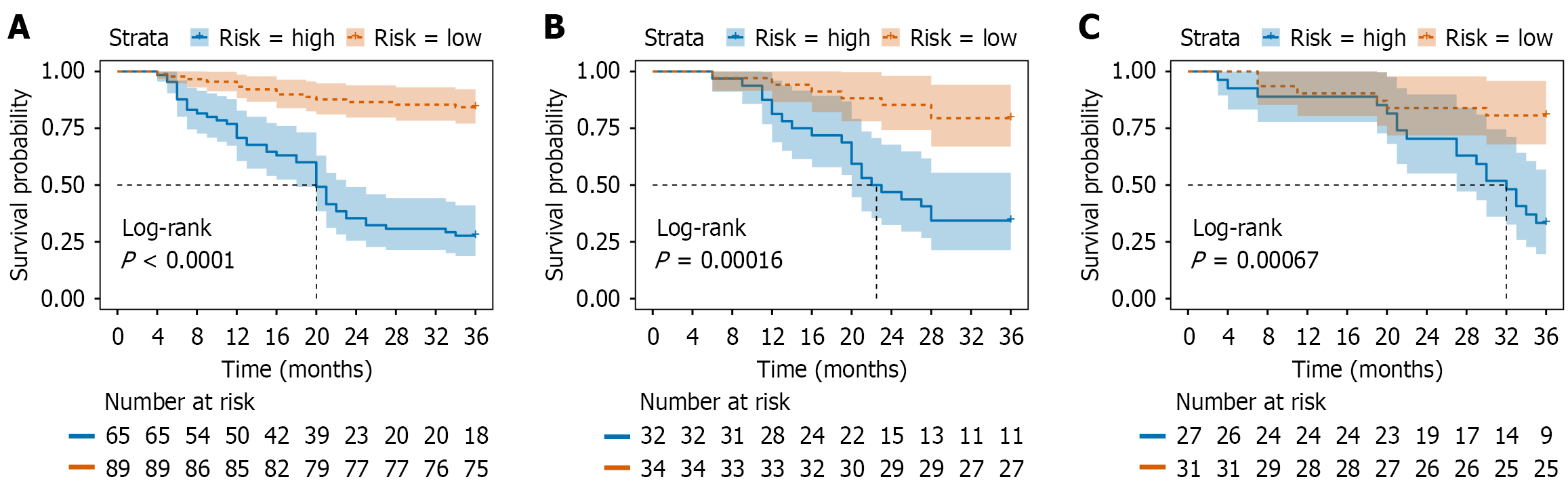
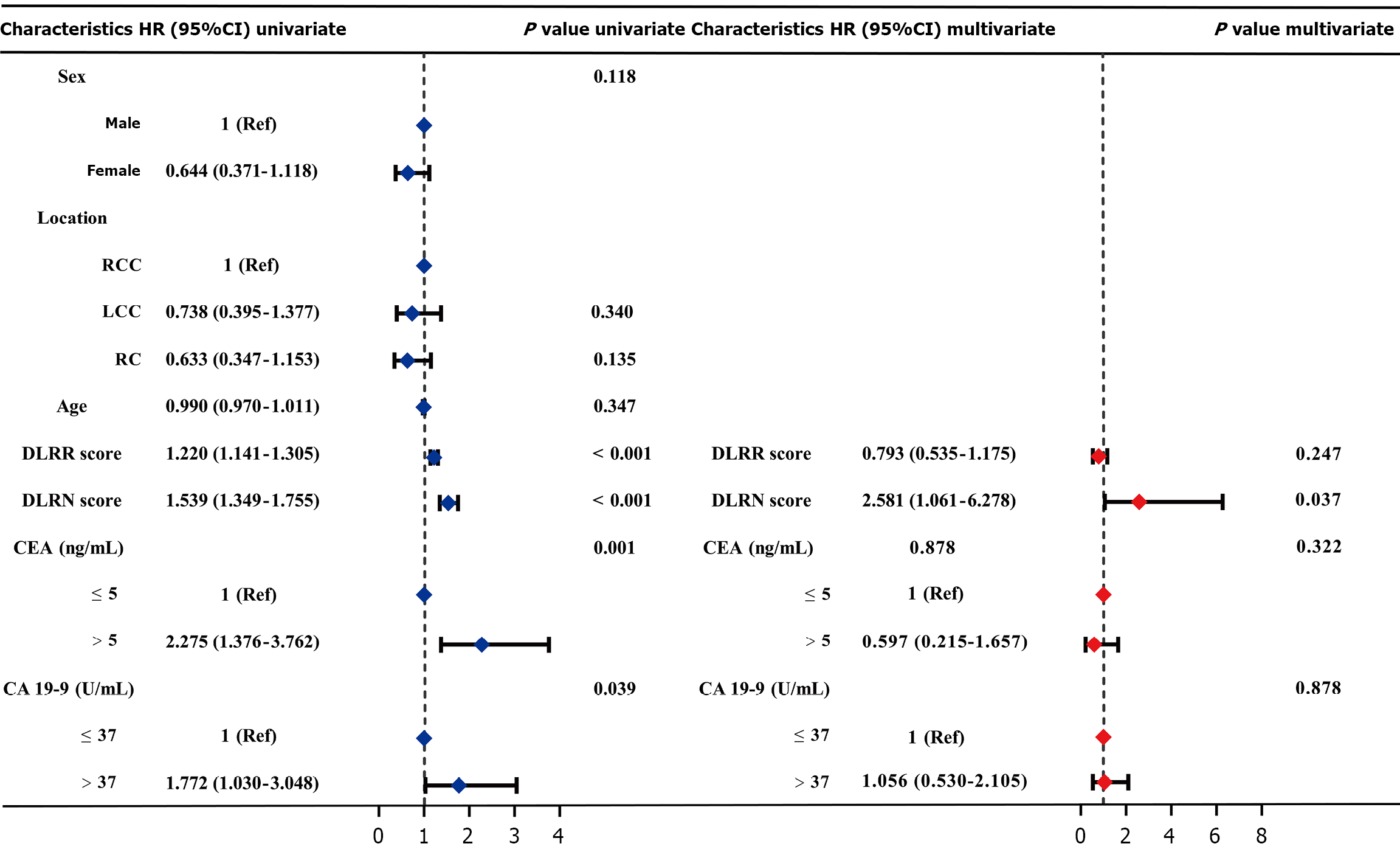
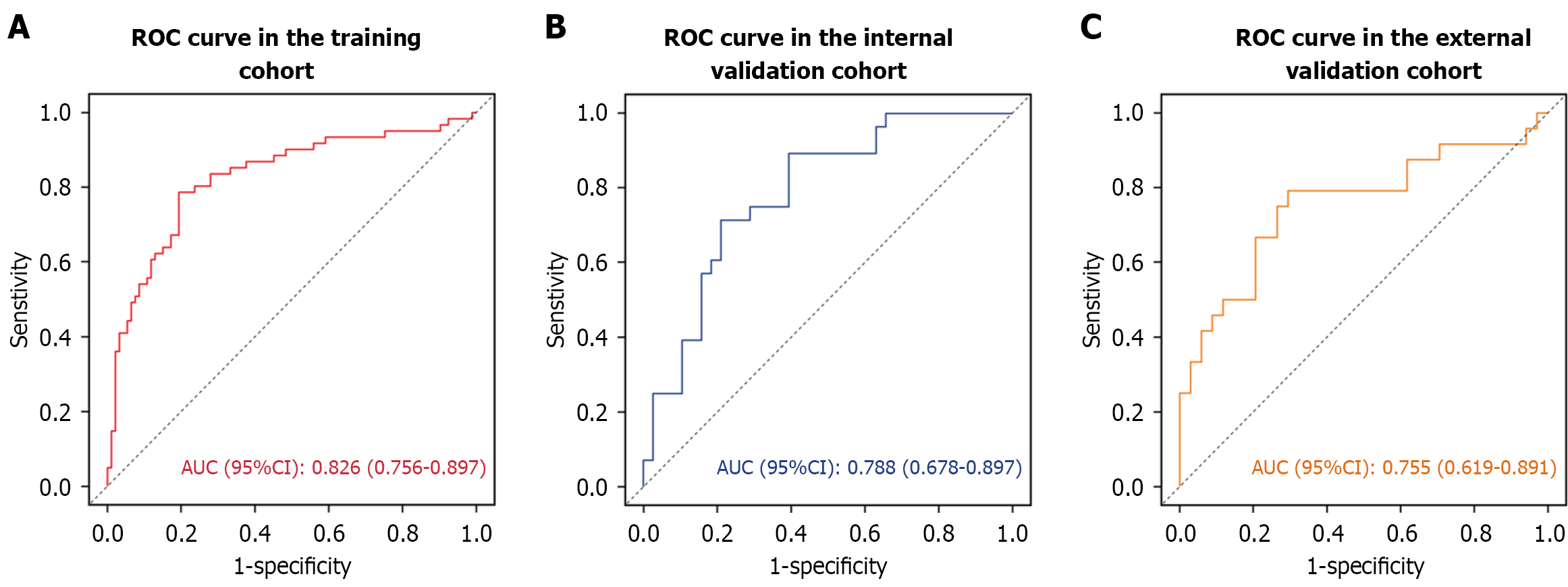
Kaplan-Meier survival curve analysis was conducted using the receiver operating characteristic cutoff value corresponding to the maximum Youden index (0.406) as the optimal threshold to stratify patients into low-risk and high-risk groups for 3-year RFS. The survival curves demonstrated that patients with low DLRN scores (≤ -0.380) had significantly better 3-year RFS compared to those with high DLRN scores (> -0.380) (Figure 10). In the training cohort, relevant factors were initially identified through univariate Cox analysis, and those exhibiting a P value < 0.100 were subsequently included in the multivariate Cox regression model, which showed that the DLRN score (hazard ratio = 2.581, 95%CI: 1.061-6.278, P = 0.037) was an independent risk factor for 3-year RFS (Figure 11). Subsequently, a prognostic model for 3-year RFS was developed based on the DLRN score, demonstrating robust performance with AUCs of 0.826, 0.788, and 0.755 in the training, internal validation, and external validation cohorts, respectively (Figure 12).
In this study, we developed and validated an interpretable DLRN based on CECT for preoperative, individualized prediction of LNM in CRC patients. Using SHAP analysis, we intuitively clarified the contribution of each variable to the model’s predictions. Our results demonstrated that the combined model, which integrates CEA, CA 19-9, intratumoral and peritumoral-3 mm radiomics features, and DL features, achieved excellent performance in predicting LNM in CRC. Furthermore, the preoperative prognostic model constructed using the DLRN score also showed strong predictive ability for 3-year RFS, underscoring its potential utility in preoperative risk stratification and personalized treatment planning.
Several clinical factors have been shown to be useful for preoperative prediction of LNM in CRC patients[19,20]. Li et al[21] reported that age, sex, preoperative CA 19-9 level, and preoperative CEA level were independent risk factors for LNM. Consistent with these findings, our multivariate analysis revealed that elevated preoperative CEA levels (odds ratio = 2.597, 95%CI: 1.199-5.623, P = 0.015) and CA 19-9 levels (odds ratio = 3.704, 95%CI: 1.435-9.563, P = 0.007) were significantly associated with LNM. However, other clinical characteristics such as age and sex did not significantly contribute to LNM prediction, which may be related to the characteristics of the study population itself. Nonetheless, relying solely on clinical factors for LNM prediction remains suboptimal, with AUC values of only 0.703, 0.637, and 0.624 in the training, internal validation, and external validation cohorts, respectively. This highlights the need for more accurate predictive approaches. Radiomics offers a non-invasive means to identify tumor biological behavior by extracting and analyzing high-throughput quantitative data from medical images. Recently, an increasing number of radiomics models have been developed to predict LNM in CRC patients[22]. For example, Niu et al[23] constructed a CECT-based radiomics model for LNM prediction in rectal cancer, achieving AUCs of 0.886 and 0.711 in the training and validation cohorts, respectively, significantly outperforming their clinical model, which had AUCs of 0.699 and 0.657. Although their study lacked external validation, it corroborates the value of radiomics in LNM prediction, consistent with the findings of our study.
Unlike the low-level image features extracted by traditional radiomics, DL, as an advanced machine learning techni
Recent studies have demonstrated that multi-regional radiomics can more comprehensively capture tumor heterogeneity[29-31]. In this study, we selected a 3 mm peritumoral region surrounding the tumor based on prior research[32]. Our results showed that the combined model integrating both intratumoral and peritumoral radiomics features signi
Previous research has demonstrated that integrating multiple relevant factors can lead to more robust predictive models[35,36]. Accordingly, we developed a multi-feature combined model by integrating clinical factors, multi-regional radiomics features, and multi-regional DL features. This combined model achieved AUCs of 0.944, 0.878, and 0.855 in the training, internal validation, and external validation cohorts, respectively, significantly outperforming both the clinical model (AUCs of 0.703, 0.637, and 0.624) and the DLRR (AUCs of 0.935, 0.877, and 0.849). These results indicate that combining features from diverse data sources enables a more comprehensive characterization of tumor biology, thereby enhancing the accuracy and robustness of LNM prediction. This approach offers stronger support for clinical individualized treatment decision-making.
Machine learning techniques are often regarded as “black boxes” due to their lack of interpretability. To address this, we employed the SHAP method combined with logistic regression to quantify and visualize the contribution of each feature to the model’s predictions. This enhances model transparency and allows clinicians to intuitively understand the relative importance of each variable in predicting LNM, facilitating focused clinical attention on key predictors. Furthermore, individual case analyses illustrated the specific impact of each variable in individual cases, enabling clinicians to more precisely assess LNM risk and tailor individualized treatment strategies.
Another noteworthy finding is that the DLRN not only demonstrated excellent performance in preoperative LNM prediction but was also significantly associated with 3-year RFS. Kaplan-Meier analysis revealed that patients with low DLRN scores had significantly better 3-year RFS compared to those with high scores. Multivariate Cox regression confirmed the DLRN score as an independent prognostic factor for 3-year RFS (hazard ratio = 1.748, 95%CI: 1.137-2.689, P = 0.011), with AUCs of 0.826, 0.788, and 0.755 in the training, internal validation, and external validation cohorts, respectively. Thus, the DLRN model not only accurately predicts preoperative LNM status but also reflects patients’ long-term prognosis through a quantified risk score, providing valuable guidance for preoperative risk stratification and personalized treatment planning.
This study has several limitations. First, its retrospective design may introduce selection bias, necessitating validation through prospective studies. Second, data were collected from only two medical centers, and the external validation cohort was relatively small. Therefore, in future research, we will incorporate data from additional centers to reduce the impact of insufficient sample size on model accuracy and enhance the model’s generalizability. Third, we included only common and easily accessible clinical variables; future studies should expand the variable range and include more clinical factors to further improve predictive performance and clinical applicability.
In conclusion, this study demonstrated that a nomogram integrating clinical features, intratumoral and peritumoral-3 mm radiomics features, and DL features effectively predicts LNM in CRC patients. The use of SHAP analysis enhanced the model’s interpretability and clinical utility. Furthermore, the prognostic model based on the DLRN score showed strong performance in predicting 3-year RFS. These findings hold promise for preoperative identification of high-risk patients and support personalized treatment decision-making.
Dr. Duan Bo, Dr. Zhang Zhenhua, and Dr. Wang Linlin (Department of Radiology, The First People’s Hospital of Hefei) are acknowledged for their expert guidance in tumor region of interest delineation. We gratefully recognize the patients who participated in this study.
| 1. | Bray F, Laversanne M, Sung H, Ferlay J, Siegel RL, Soerjomataram I, Jemal A. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2024;74:229-263. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16785] [Cited by in RCA: 14765] [Article Influence: 7382.5] [Reference Citation Analysis (21)] |
| 2. | Backes Y, Elias SG, Bhoelan BS, Groen JN, van Bergeijk J, Seerden TCJ, Pullens HJM, Spanier BWM, Geesing JMJ, Kessels K, Kerkhof M, Siersema PD, de Vos Tot Nederveen Cappel WH, van Lelyveld N, Wolfhagen FHJ, Ter Borg F, Offerhaus GJA, Lacle MM, Moons LMG; Dutch T1 CRC Working Group. The prognostic value of lymph node yield in the earliest stage of colorectal cancer: a multicenter cohort study. BMC Med. 2017;15:129. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 28] [Cited by in RCA: 28] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 3. | Bosch SL, Teerenstra S, de Wilt JH, Cunningham C, Nagtegaal ID. Predicting lymph node metastasis in pT1 colorectal cancer: a systematic review of risk factors providing rationale for therapy decisions. Endoscopy. 2013;45:827-834. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 356] [Cited by in RCA: 314] [Article Influence: 24.2] [Reference Citation Analysis (0)] |
| 4. | Oka S, Tanaka S, Kajiwara Y, Saito S, Fukunaga Y, Takamatsu M, Kawachi H, Hotta K, Ikematsu H, Kojima M, Saito Y, Yamada M, Kanemitsu Y, Sekine S, Nagata S, Yamada K, Kobayashi N, Ishihara S, Saitoh Y, Matsuda K, Togashi K, Komori K, Ishiguro M, Kuwai T, Okuyama T, Ohuchi A, Ohnuma S, Sakamoto K, Sugai T, Katsumata K, Matsushita HO, Yamano HO, Eda H, Uraoka T, Akimoto N, Kobayashi H, Sugihara K, Ueno H. Treatment Decision for Locally Resected T1 Colorectal Carcinoma-Verification of the Japanese Guideline Criteria for Additional Surgery Based on Long-Term Clinical Outcomes. Am J Gastroenterol. 2024;119:2019-2027. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 15] [Article Influence: 7.5] [Reference Citation Analysis (1)] |
| 5. | Tang Y, Su YX, Zheng JM, Zhuo ML, Qian QF, Shen QL, Lin P, Chen ZK. Radiogenomic analysis for predicting lymph node metastasis and molecular annotation of radiomic features in pancreatic cancer. J Transl Med. 2024;22:690. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 10] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 6. | Gu W, Chen Y, Zhu H, Chen H, Yang Z, Mo S, Zhao H, Chen L, Nakajima T, Yu X, Ji S, Gu Y, Chen J, Tang W. Development and validation of CT-based radiomics deep learning signatures to predict lymph node metastasis in non-functional pancreatic neuroendocrine tumors: a multicohort study. EClinicalMedicine. 2023;65:102269. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 39] [Article Influence: 13.0] [Reference Citation Analysis (1)] |
| 7. | Tran KA, Kondrashova O, Bradley A, Williams ED, Pearson JV, Waddell N. Deep learning in cancer diagnosis, prognosis and treatment selection. Genome Med. 2021;13:152. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 750] [Cited by in RCA: 507] [Article Influence: 101.4] [Reference Citation Analysis (4)] |
| 8. | Kim H, Goo JM, Lee KH, Kim YT, Park CM. Preoperative CT-based Deep Learning Model for Predicting Disease-Free Survival in Patients with Lung Adenocarcinomas. Radiology. 2020;296:216-224. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 115] [Cited by in RCA: 99] [Article Influence: 16.5] [Reference Citation Analysis (0)] |
| 9. | Skrede OJ, De Raedt S, Kleppe A, Hveem TS, Liestøl K, Maddison J, Askautrud HA, Pradhan M, Nesheim JA, Albregtsen F, Farstad IN, Domingo E, Church DN, Nesbakken A, Shepherd NA, Tomlinson I, Kerr R, Novelli M, Kerr DJ, Danielsen HE. Deep learning for prediction of colorectal cancer outcome: a discovery and validation study. Lancet. 2020;395:350-360. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 553] [Cited by in RCA: 416] [Article Influence: 69.3] [Reference Citation Analysis (7)] |
| 10. | Chen PT, Wu T, Wang P, Chang D, Liu KL, Wu MS, Roth HR, Lee PC, Liao WC, Wang W. Pancreatic Cancer Detection on CT Scans with Deep Learning: A Nationwide Population-based Study. Radiology. 2023;306:172-182. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 146] [Cited by in RCA: 83] [Article Influence: 27.7] [Reference Citation Analysis (0)] |
| 11. | Wei Z, Xv Y, Liu H, Li Y, Yin S, Xie Y, Chen Y, Lv F, Jiang Q, Li F, Xiao M. A CT-based deep learning model predicts overall survival in patients with muscle invasive bladder cancer after radical cystectomy: a multicenter retrospective cohort study. Int J Surg. 2024;110:2922-2932. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 22] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 12. | Lin CY, Guo SM, Lien JJ, Lin WT, Liu YS, Lai CH, Hsu IL, Chang CC, Tseng YL. Combined model integrating deep learning, radiomics, and clinical data to classify lung nodules at chest CT. Radiol Med. 2024;129:56-69. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 55] [Cited by in RCA: 47] [Article Influence: 23.5] [Reference Citation Analysis (0)] |
| 13. | Long H, Zhang P, Bi Y, Yang C, Wu M, He D, Huang S, Yang K, Qi S, Wang J. MRI radiomic features of peritumoral edema may predict the recurrence sites of glioblastoma multiforme. Front Oncol. 2022;12:1042498. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 24] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 14. | Li XR, Jin JJ, Yu Y, Wang XH, Guo Y, Sun HZ. PET-CT radiomics by integrating primary tumor and peritumoral areas predicts E-cadherin expression and correlates with pelvic lymph node metastasis in early-stage cervical cancer. Eur Radiol. 2021;31:5967-5979. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 29] [Article Influence: 5.8] [Reference Citation Analysis (3)] |
| 15. | Ma Q, Wang J, Tu Z, She J, Zhu J, Jiang F, Zhang C. Prediction model of axillary lymph node status using an automated breast volume ultrasound radiomics nomogram in early breast cancer with negative axillary ultrasound. Front Immunol. 2025;16:1460673. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 6] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 16. | Mao N, Shi Y, Lian C, Wang Z, Zhang K, Xie H, Zhang H, Chen Q, Cheng G, Xu C, Dai Y. Intratumoral and peritumoral radiomics for preoperative prediction of neoadjuvant chemotherapy effect in breast cancer based on contrast-enhanced spectral mammography. Eur Radiol. 2022;32:3207-3219. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 88] [Cited by in RCA: 72] [Article Influence: 18.0] [Reference Citation Analysis (0)] |
| 17. | Pan YJ, Wu SJ, Zeng Y, Cao ZR, Shan Y, Lin J, Xu PJ. Intra- and Peri-tumoral Radiomics Based on Dynamic Contrast Enhanced-MRI to Identify Lymph Node Metastasis and Prognosis in Intrahepatic Cholangiocarcinoma. J Magn Reson Imaging. 2024;60:2669-2680. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 14] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 18. | Yuan H, Xu X, Tu S, Chen B, Wei Y, Ma Y. The CT-based intratumoral and peritumoral machine learning radiomics analysis in predicting lymph node metastasis in rectal carcinoma. BMC Gastroenterol. 2022;22:463. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 15] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 19. | Wang K, He H, Lin Y, Zhang Y, Chen J, Hu J, He X. A new clinical model for predicting lymph node metastasis in T1 colorectal cancer. Int J Colorectal Dis. 2024;39:46. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 4] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 20. | Guo K, Feng Y, Yuan L, Wasan HS, Sun L, Shen M, Ruan S. Risk factors and predictors of lymph nodes metastasis and distant metastasis in newly diagnosed T1 colorectal cancer. Cancer Med. 2020;9:5095-5113. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 49] [Cited by in RCA: 44] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 21. | Li M, Zhang J, Dan Y, Yao Y, Dai W, Cai G, Yang G, Tong T. A clinical-radiomics nomogram for the preoperative prediction of lymph node metastasis in colorectal cancer. J Transl Med. 2020;18:46. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 110] [Cited by in RCA: 105] [Article Influence: 17.5] [Reference Citation Analysis (1)] |
| 22. | Zhang YC, Li M, Jin YM, Xu JX, Huang CC, Song B. Radiomics for differentiating tumor deposits from lymph node metastasis in rectal cancer. World J Gastroenterol. 2022;28:3960-3970. [PubMed] [DOI] [Full Text] |
| 23. | Niu Y, Yu X, Wen L, Bi F, Jian L, Liu S, Yang Y, Zhang Y, Lu Q. Comparison of preoperative CT- and MRI-based multiparametric radiomics in the prediction of lymph node metastasis in rectal cancer. Front Oncol. 2023;13:1230698. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 16] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 24. | Tharmaseelan H, Vellala AK, Hertel A, Tollens F, Rotkopf LT, Rink J, Woźnicki P, Ayx I, Bartling S, Nörenberg D, Schoenberg SO, Froelich MF. Tumor classification of gastrointestinal liver metastases using CT-based radiomics and deep learning. Cancer Imaging. 2023;23:95. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 17] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 25. | Zhu Z, Wu K, Lu J, Dai S, Xu D, Fang W, Yu Y, Gu W. Gd-EOB-DTPA-enhanced MRI radiomics and deep learning models to predict microvascular invasion in hepatocellular carcinoma: a multicenter study. BMC Med Imaging. 2025;25:105. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 6] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 26. | Liao H, Yuan J, Liu C, Zhang J, Yang Y, Liang H, Jiang S, Chen S, Li Y, Liu Y. Feasibility and effectiveness of automatic deep learning network and radiomics models for differentiating tumor stroma ratio in pancreatic ductal adenocarcinoma. Insights Imaging. 2023;14:223. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 8] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 27. | Wu L, Li S, Wu C, Wu S, Lin Y, Wei D. Ultrasound-based deep learning radiomics nomogram for differentiating mass mastitis from invasive breast cancer. BMC Med Imaging. 2024;24:189. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 8] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 28. | Gao Y, Wang W, Yang Y, Xu Z, Lin Y, Lang T, Lei S, Xiao Y, Yang W, Huang W, Li Y. An integrated model incorporating deep learning, hand-crafted radiomics and clinical and US features to diagnose central lymph node metastasis in patients with papillary thyroid cancer. BMC Cancer. 2024;24:69. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 27] [Article Influence: 13.5] [Reference Citation Analysis (0)] |
| 29. | Xu R, Wang K, Peng B, Zhou X, Wang C, Lu T, Shi J, Zhao J, Zhang L. Evaluating peritumoral and intratumoral radiomics signatures for predicting lymph node metastasis in surgically resectable non-small cell lung cancer. Front Oncol. 2024;14:1427743. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 30. | Lin P, Xie W, Li Y, Zhang C, Wu H, Wan H, Gao M, Liang F, Han P, Chen R, Cheng G, Liu X, Fan S, Huang X. Intratumoral and peritumoral radiomics of MRIs predicts pathologic complete response to neoadjuvant chemoimmunotherapy in patients with head and neck squamous cell carcinoma. J Immunother Cancer. 2024;12:e009616. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 40] [Cited by in RCA: 30] [Article Influence: 15.0] [Reference Citation Analysis (0)] |
| 31. | Xia TY, Zhou ZH, Meng XP, Zha JH, Yu Q, Wang WL, Song Y, Wang YC, Tang TY, Xu J, Zhang T, Long XY, Liang Y, Xiao WB, Ju SH. Predicting Microvascular Invasion in Hepatocellular Carcinoma Using CT-based Radiomics Model. Radiology. 2023;307:e222729. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 205] [Cited by in RCA: 174] [Article Influence: 58.0] [Reference Citation Analysis (6)] |
| 32. | Li H, Chen XL, Liu H, Liu YS, Li ZL, Pang MH, Pu H. MRI-based multiregional radiomics for preoperative prediction of tumor deposit and prognosis in resectable rectal cancer: a bicenter study. Eur Radiol. 2023;33:7561-7572. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 34] [Article Influence: 11.3] [Reference Citation Analysis (1)] |
| 33. | Liang ZY, Yu ML, Yang H, Li HJ, Xie H, Cui CY, Zhang WJ, Luo C, Cai PQ, Lin XF, Liu KF, Xiong L, Liu LZ, Chen BY. Beyond the tumor region: Peritumoral radiomics enhances prognostic accuracy in locally advanced rectal cancer. World J Gastroenterol. 2025;31:99036. [PubMed] [DOI] [Full Text] |
| 34. | Wang Y, Zhao H, Fu P, Tian L, Su Y, Lyu Z, Gu W, Wang Y, Liu S, Wang X, Zheng H, Du J, Zhang R. Preoperative prediction of lymph node metastasis in colorectal cancer using (18)F-FDG PET/CT peritumoral radiomics analysis. Med Phys. 2024;51:5214-5225. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 2.5] [Reference Citation Analysis (1)] |
| 35. | Liu Z, Luo C, Chen X, Feng Y, Feng J, Zhang R, Ouyang F, Li X, Tan Z, Deng L, Chen Y, Cai Z, Zhang X, Liu J, Liu W, Guo B, Hu Q. Noninvasive prediction of perineural invasion in intrahepatic cholangiocarcinoma by clinicoradiological features and computed tomography radiomics based on interpretable machine learning: a multicenter cohort study. Int J Surg. 2024;110:1039-1051. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 54] [Cited by in RCA: 57] [Article Influence: 28.5] [Reference Citation Analysis (0)] |
| 36. | Fu N, Fu W, Chen H, Chai W, Qian X, Wang W, Jiang Y, Shen B. A deep-learning radiomics-based lymph node metastasis predictive model for pancreatic cancer: a diagnostic study. Int J Surg. 2023;109:2196-2203. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 41] [Article Influence: 13.7] [Reference Citation Analysis (0)] |