Copyright: ©Author(s) 2026.
World J Gastrointest Oncol. May 15, 2026; 18(5): 116568
Published online May 15, 2026. doi: 10.4251/wjgo.v18.i5.116568
Published online May 15, 2026. doi: 10.4251/wjgo.v18.i5.116568
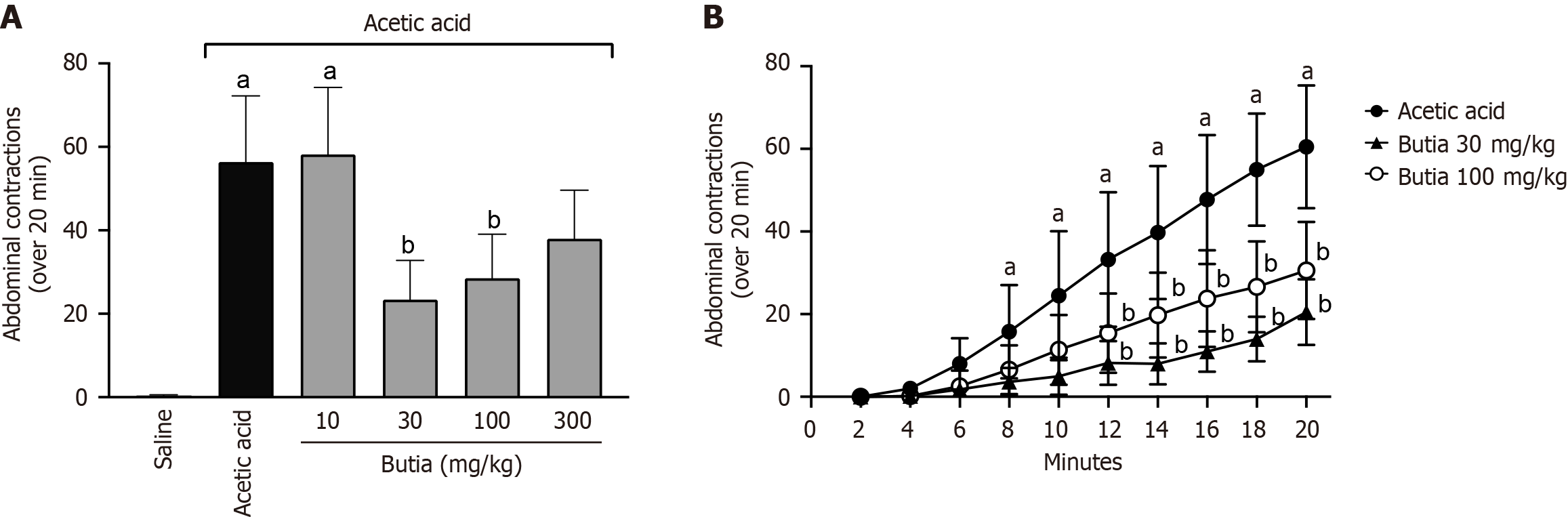
Figure 1 Antinociceptive activity of the hydroalcoholic extract of Butia capitata fruit pulp.
A: Abdominal contractions (over 20 minutes); B: Accumulated abdominal contortions every two minutes (over 20 minutes). Results presented ± representative error of 8 animals per group. aP < 0.05 vs saline group; bP < 0.05 vs acetic acid. One-way analysis of variance followed by Tukey’s t-test. Error bar: Standard error of the mean.
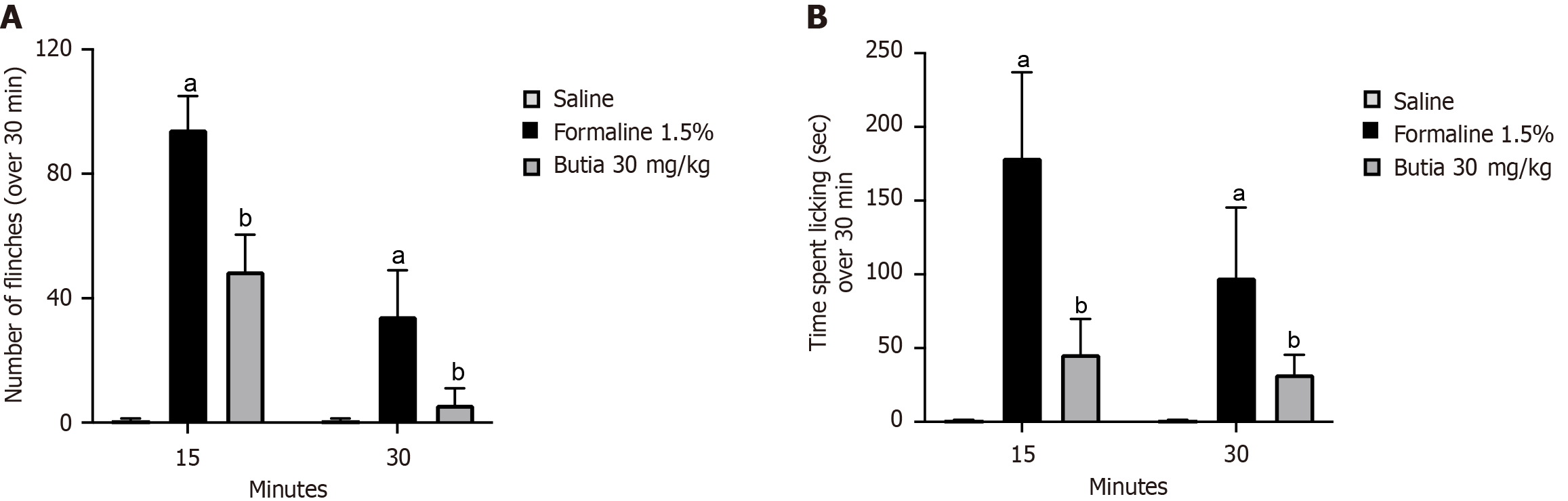
Figure 2 Antinociceptive and anti-inflammatory activities of the hydroalcoholic extract of Butia capitata fruit pulp.
A: Number of paw flinching over 30 minutes; B: Time spent licking the paw over 30 minutes. Results presented ± representative error of 8 animals per group. aP < 0.05 vs saline group; bP < 0.05 vs acetic acid. One-way analysis of variance followed by Tukey’s t-test. Error bar: Standard error of the mean.
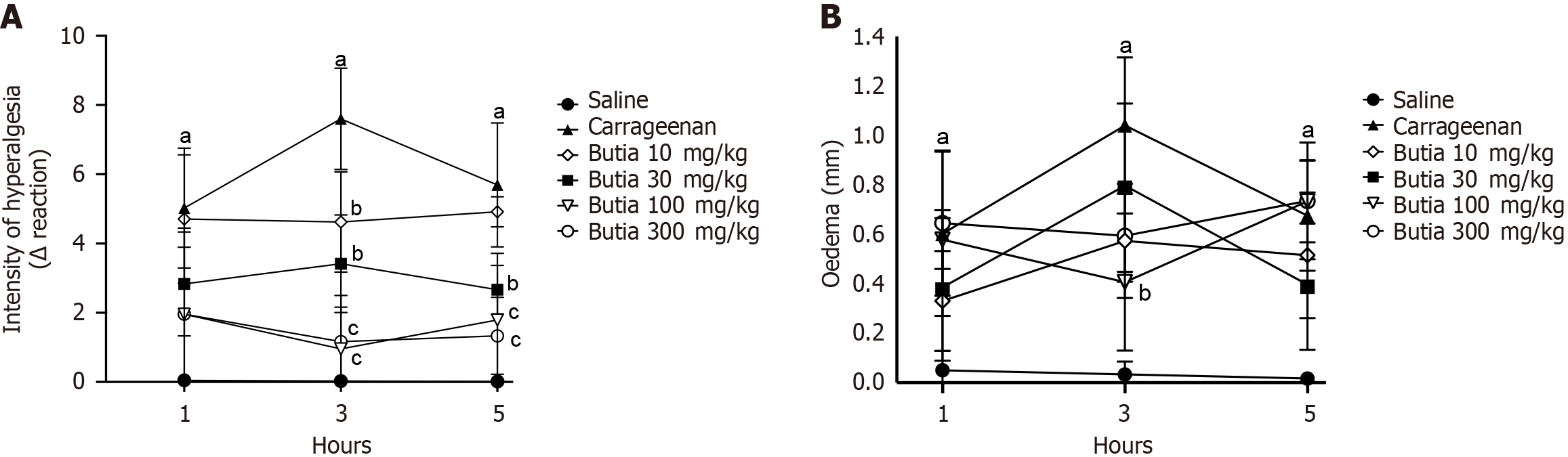
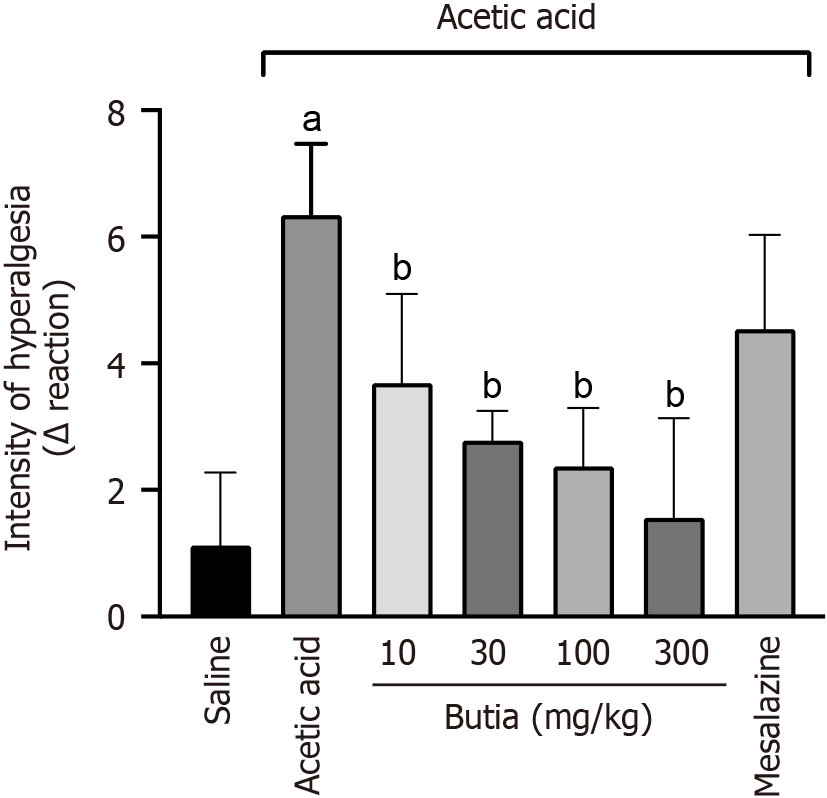
Figure 3 Anti-hyperalgesia and anti-edematous activities of the hydroalcoholic extract of Butia capitata fruit pulp.
A: Intensity of hyperalgesia; B: Intensity of edema. Results presented ± representative error of 6 animals per group. aP < 0.05 vs saline group; bP < 0.05 vs carrageenan; cP < 0.05 vs acetic acid, extract at 10 mg/kg. One-way analysis of variance followed by Tukey’s t-test. Error bar: Standard error of the mean.
Figure 4 Macroscopic indices of colonic segments from mice with induced colitis post-treatment.
Scores: 0 = no lesion; 1 = focal hyperemia without ulceration; 2 = ulceration without hypertrophy of the intestinal wall; 3 = ulceration with inflammation spot according to Wallace et al[18]. Results presented ± representative error of 8 animals per group. aP < 0.05 vs saline group; bP < 0.05 vs acetic acid. One-way analysis of variance followed by Tukey’s t-test. Error bar: Standard error of the mean.
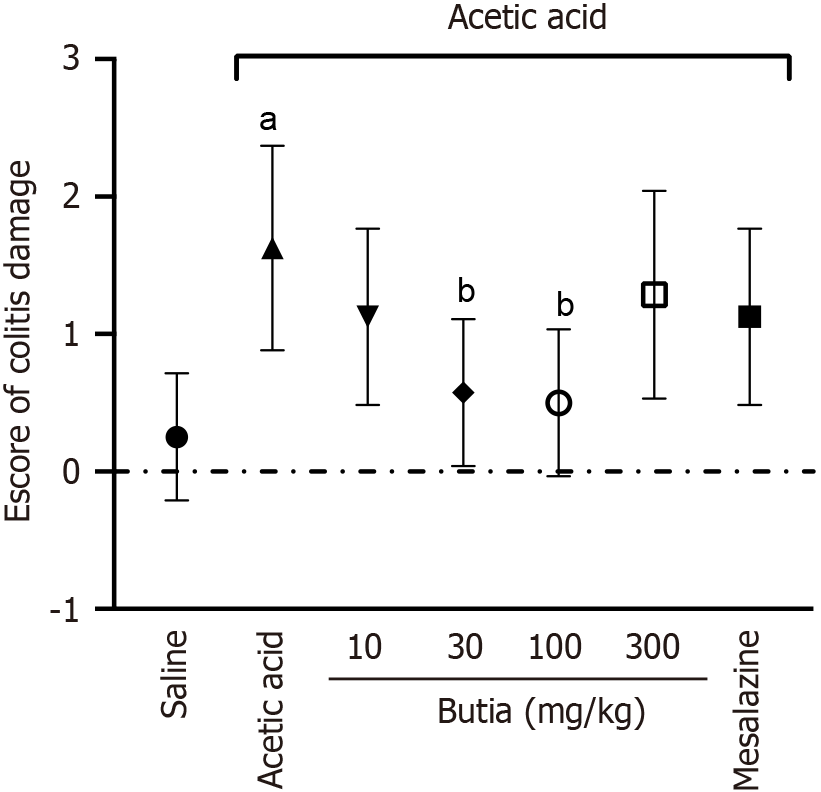
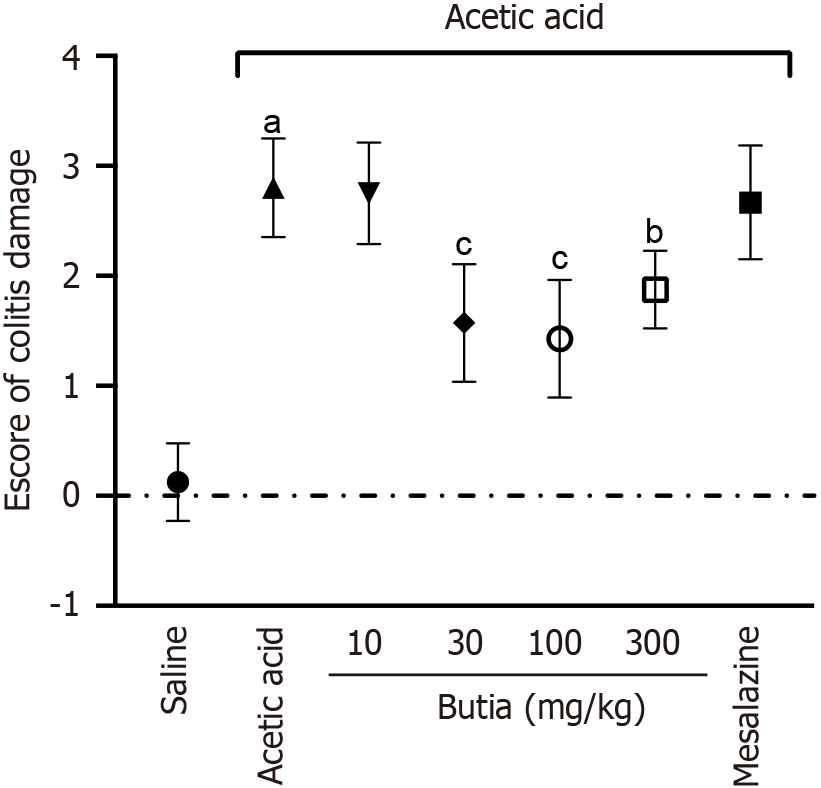
Figure 5 Colitis damage score from histological analysis of colonic segments from mice with induced colitis post-treatment.
Scores: 0 = no signs of inflammation; 1 = mild inflammation; 2 = moderate inflammation; 3 = intense inflammation. Results presented ± representative error of 8 animals per group. aP < 0.05 vs saline group; bP < 0.05 vs extract at 10 mg/Kg, acetic acid; cP < 0.05 vs acetic acid. One-way analysis of variance followed by Tukey’s t-test. Error bar: Standard error of the mean.
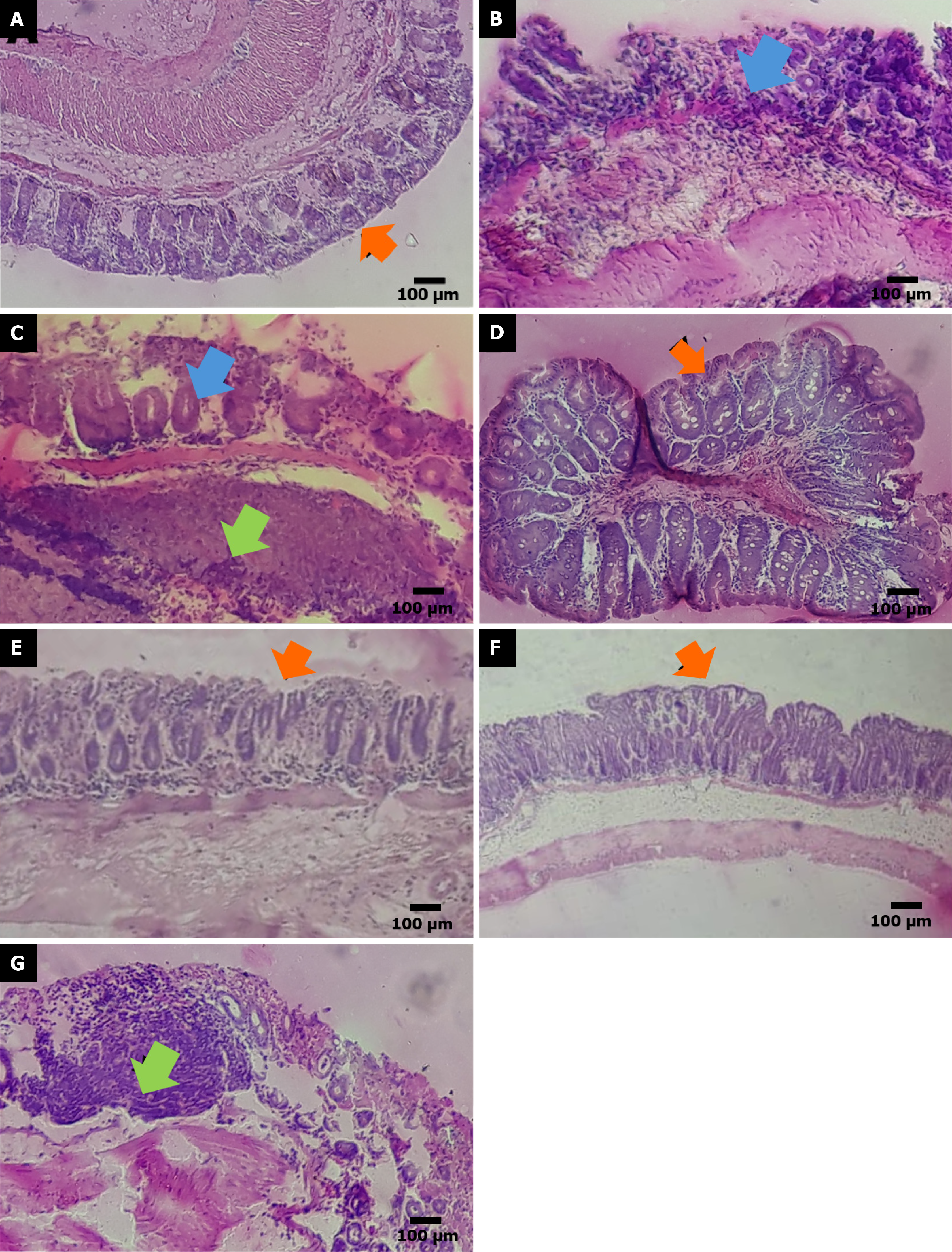
Figure 6 Photomicrographs representing histological sections of colon tissue stained with hematoxylin-eosin.
A: Negative control group (saline); B: Acetic acid-treated group; C: Keto-alcoholic extract of Butia capitata at 10 mg/kg; D: Keto-alcoholic extract of Butia capitata at 30 mg/kg; E: Keto-alcoholic extract of Butia capitata at 100 mg/kg; F: Keto-alcoholic extract of Butia capitata at 300 mg/kg; G: Mesalazine treated group. The green arrows indicate inflammatory infiltrate, the blue arrows show loss of crypt architecture, and the orange arrows demonstrates a spot with preserved crypts and epithelial barrier.
Figure 7 Antinociceptive activity of the hydroalcoholic extract of Butia capitata fruit pulp in the acute colitis model.
Results presented ± representative error of 8 animals per group. aP < 0.05 vs saline group; bP < 0.05 vs acetic acid. One-way analysis of variance followed by Tukey's t-test. Error bar: Standard error of the mean.
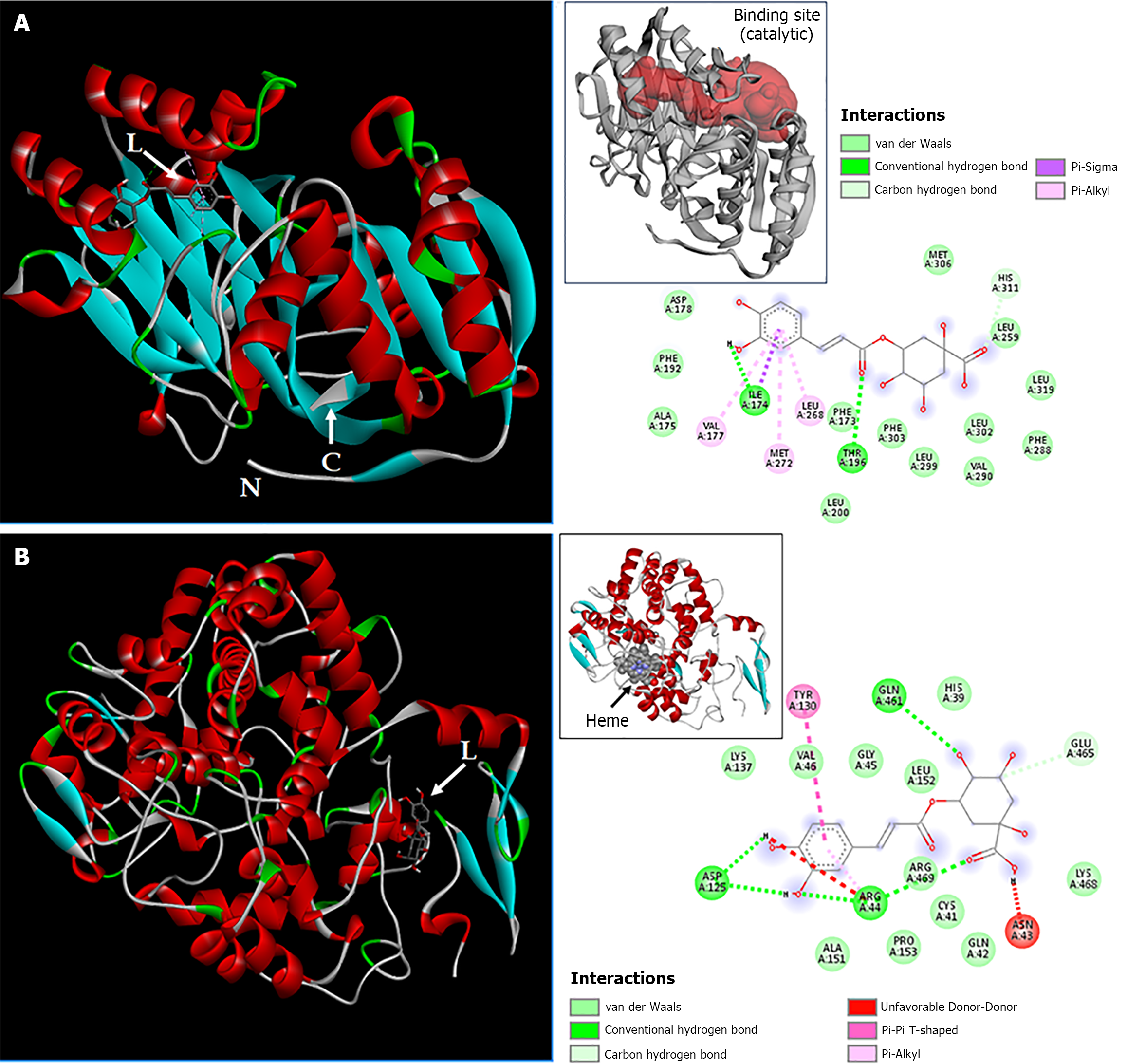
Figure 8 Interactions of the sphingosine kinase 1/chlorogenic acid and cyclooxygenase-2/chlorogenic acid complexes.
A: Chain A of sphingosine kinase 1 bound to chlorogenic acid. Inside the square, the red region indicates the binding site of chain A; B: Chain A of cyclooxygenase-2 bound to chlorogenic acid. Inside the square, the arrow indicates the heme group. The two-dimensional diagrams show the chemical bonds between receptor amino acid residues and chlorogenic acid atoms. The internal legend indicates the type of chemical bond according to the color code. Amino acid residues are represented within spheres by the three-letter code followed by the residue position in the primary sequence. L: Ligand; N: Amino-terminal; C: Carboxy-terminal.
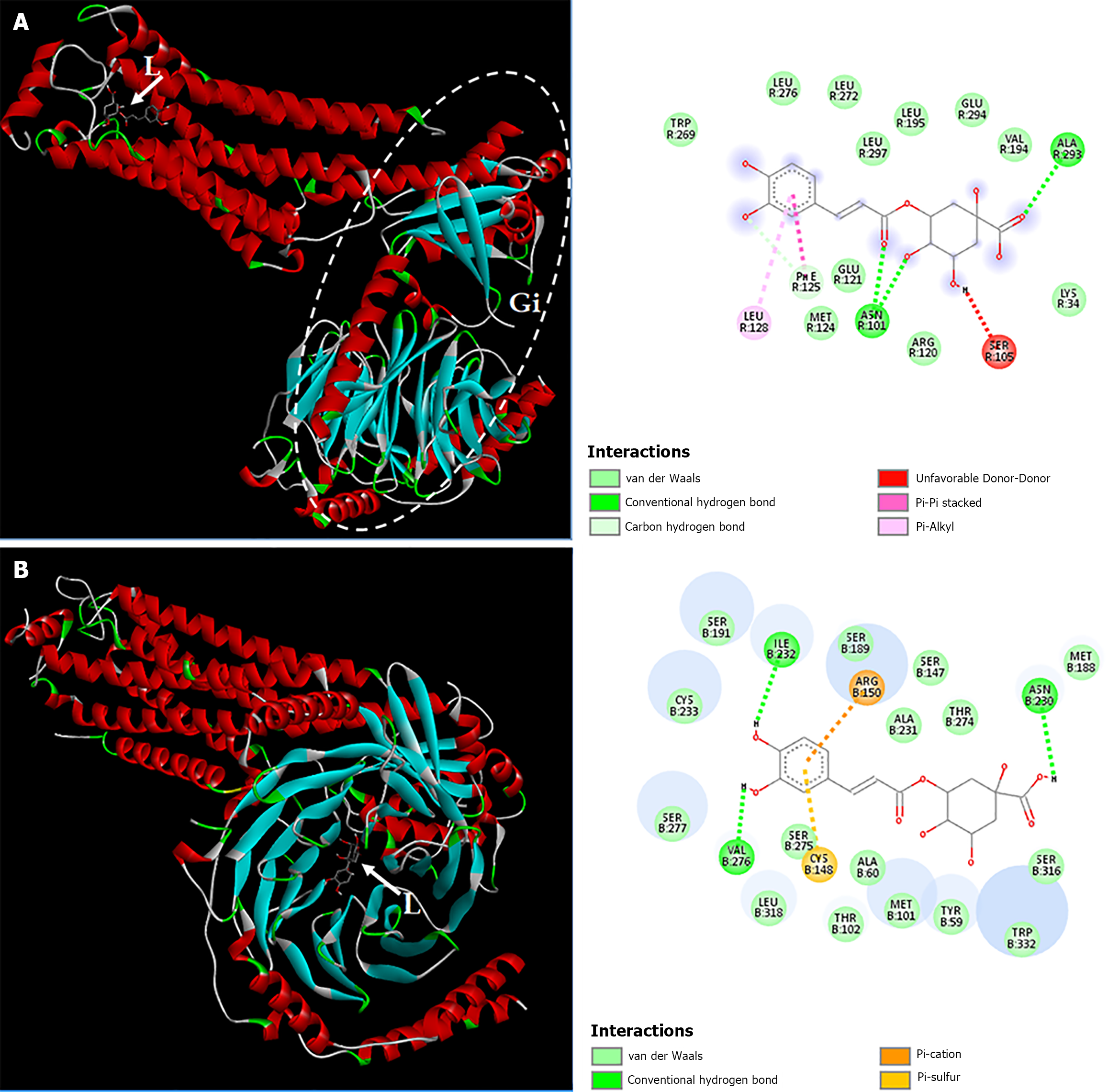
Figure 9 Interactions of the sphingosine 1-phosphate receptor 1/chlorogenic acid complex.
A: Complex formed by the 7-transmembrane receptor and chlorogenic acid. The region delimited by the dashed line corresponds to the Gi protein; B: Complex formed by the β subunit of the Gi protein and chlorogenic acid. The two-dimensional diagrams show the chemical bonds between receptor amino acid residues and chlorogenic acid atoms. The internal legend indicates the type of chemical bond according to the color code. Amino acid residues are represented within spheres by the three-letter code followed by the residue position in the primary sequence. L: Ligand; Gi: Gi protein.
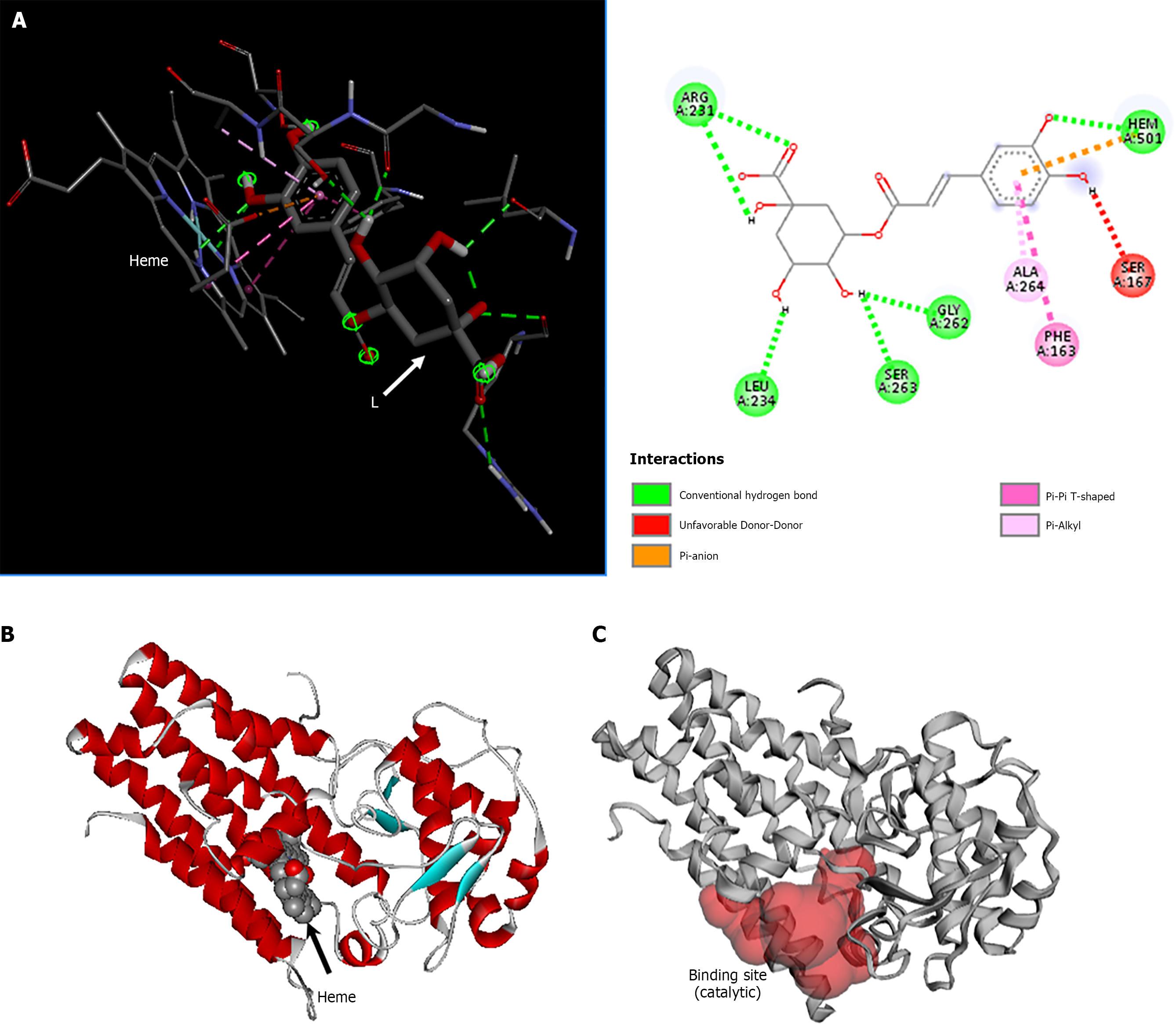
Figure 10 Interactions of the indoleamine 2,3-dioxygenase 1/chlorogenic acid complex.
A: Chemical bonds between chlorogenic acid and the amino acid residues/heme of indoleamine 2,3-dioxygenase 1 (IDO1); B: Chain A of IDO1 indicating the heme group; C: The red region indicates the binding site of chain A of IDO1. The two-dimensional diagrams show the chemical bonds between receptor amino acid residues and chlorogenic acid atoms. The internal legend indicates the type of chemical bond according to the color code. Amino acid residues are represented within spheres by the three-letter code followed by the residue position in the primary sequence. L: Ligand.
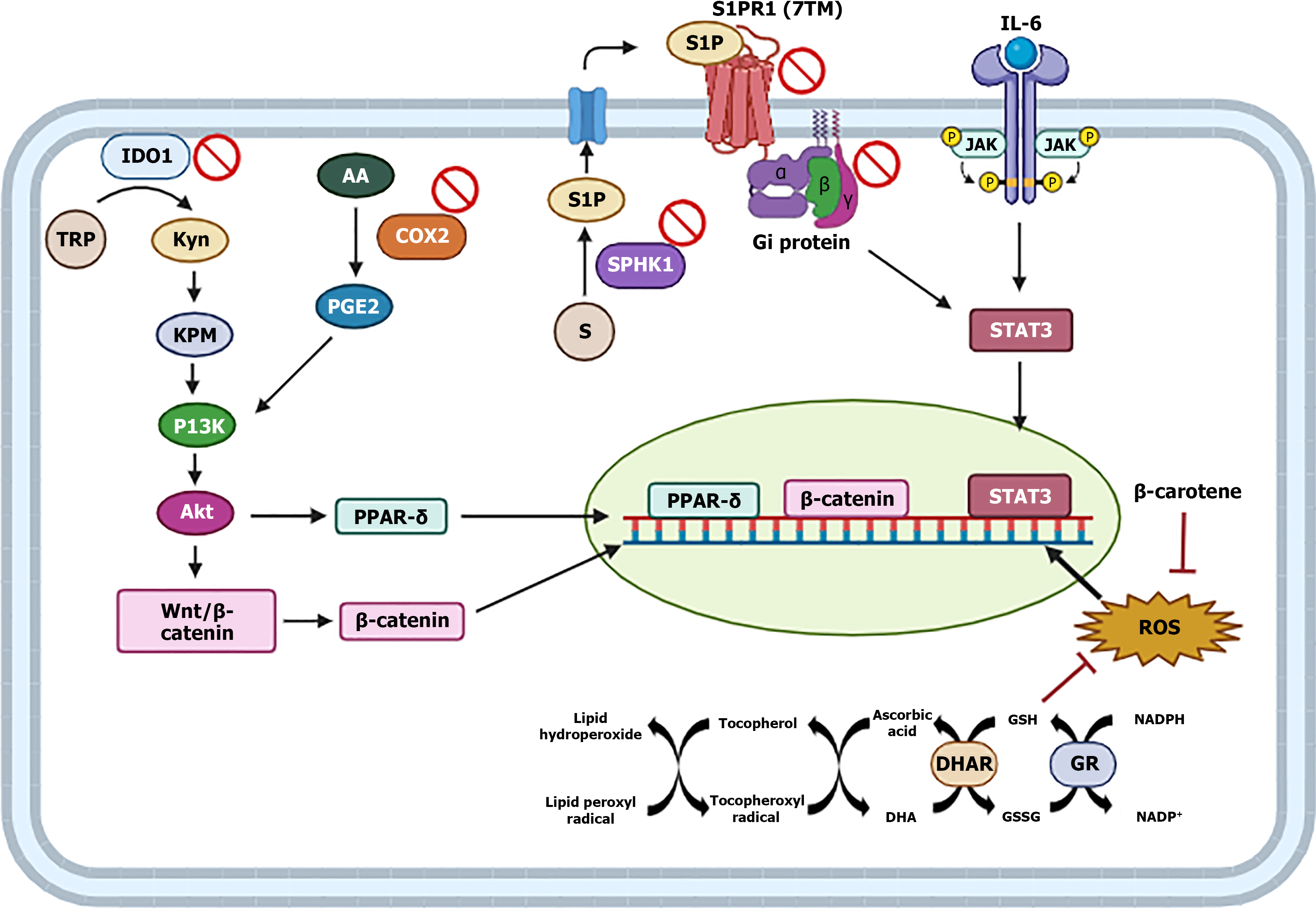
Figure 11 Schematic of the supposed joint activity of antioxidants and chlorogenic acid from the fruit of Butia capitata on cellular targets of signaling pathways associated with colitis inflammation and DNA damage.
TRP: Tryptophan; Kyn: Kynurenine; KPM: Kynurenine metabolites; AA: Arachidonic acid; S: Sphingosine; S1P: Sphingosine 1-phosphate; DHA: Dehydroascorbate; DHAR: Semidehydroascorbate reductase; GR: Glutathione reductase. ROS: Reactive oxygen species; STAT3: Signal transducer and activator of transcription 3; IDO1: Indoleamine 2,3-dioxygenase 1; IL: Interleukin; 7TM: Seven-transmembrane.
- Citation: Da Silva PB, Lopes-Bute L, Fischer J, Parcianello AP, Pires-Duarte R, Luiz RM, Brandalize APC, Melo FF, Wendel CF, Teixeira KN, Zarpelon-Schutz AC. Protective effects of Butia capitata extract against colitis-driven colorectal tumorigenesis: Insights into its anti-inflammatory and antinociceptive activities. World J Gastrointest Oncol 2026; 18(5): 116568
- URL: https://www.wjgnet.com/1948-5204/full/v18/i5/116568.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v18.i5.116568