Copyright: ©Author(s) 2026.
World J Gastrointest Oncol. Apr 15, 2026; 18(4): 116504
Published online Apr 15, 2026. doi: 10.4251/wjgo.v18.i4.116504
Published online Apr 15, 2026. doi: 10.4251/wjgo.v18.i4.116504
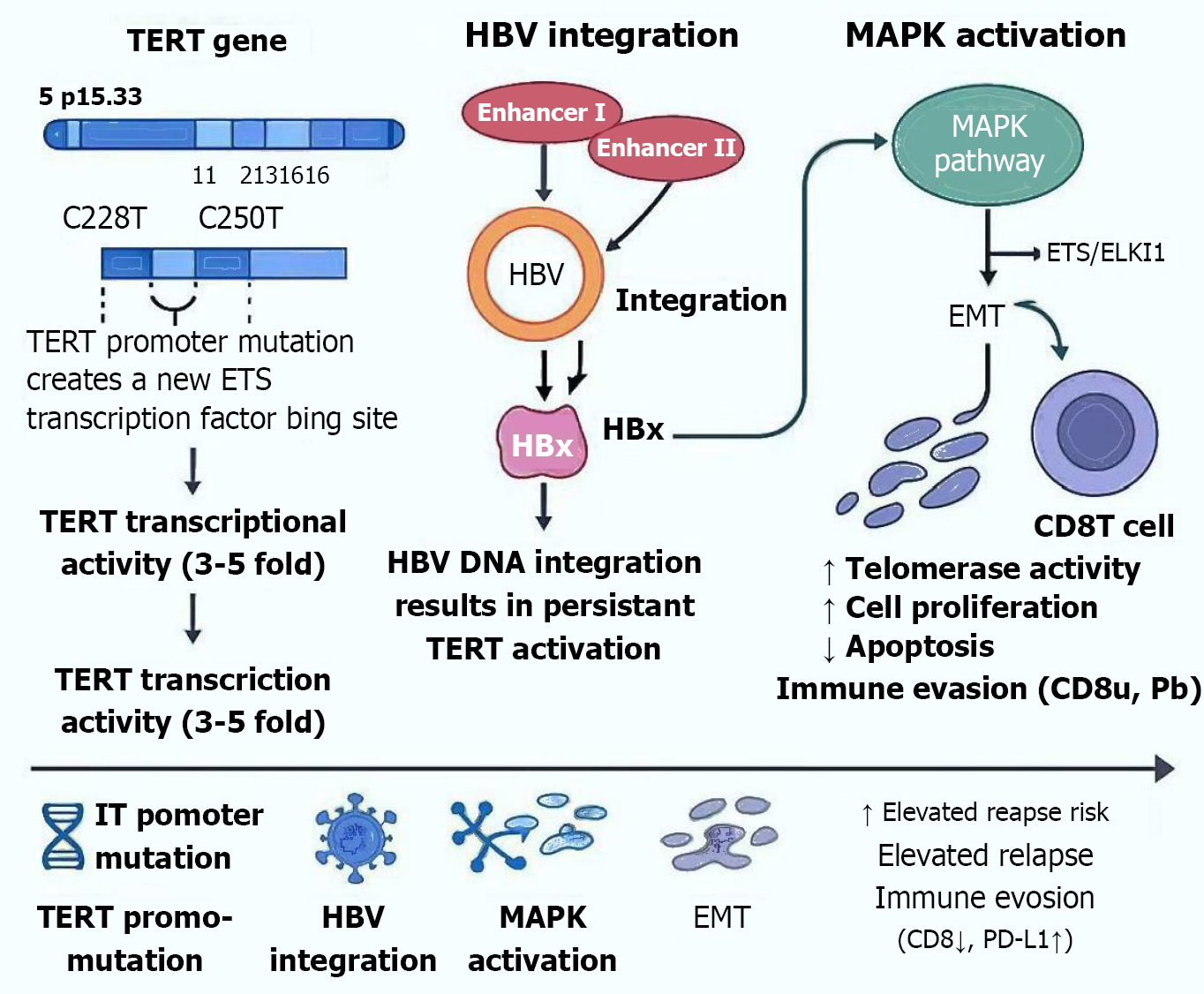
Figure 1 Integrated molecular model.
Illustrating how hepatitis B virus DNA integration and telomerase reverse transcriptase promoter mutations synergistically activate telomerase, enhance oncogenic signaling (mitogen-activated protein kinase/E-twenty-six, nuclear factor kappaB, Wnt/β-catenin), promote epithelial-mesenchymal transition (epithelial-mesenchymal transition), and drive immune evasion, contributing to recurrence in hepatitis B virus-related hepatocellular carcinoma. TERT: Telomerase reverse transcriptase; ETS: E-twenty-six; IT: Integrated transcription; HBV: Hepatitis B virus; HBx: Hepatitis B virus X protein; MAPK: Mitogen-activated protein kinase; EMT: Epithelial-mesenchymal transition; ELKI1: Ets-like protein 1; PD-L1: Programmed death ligand-1.
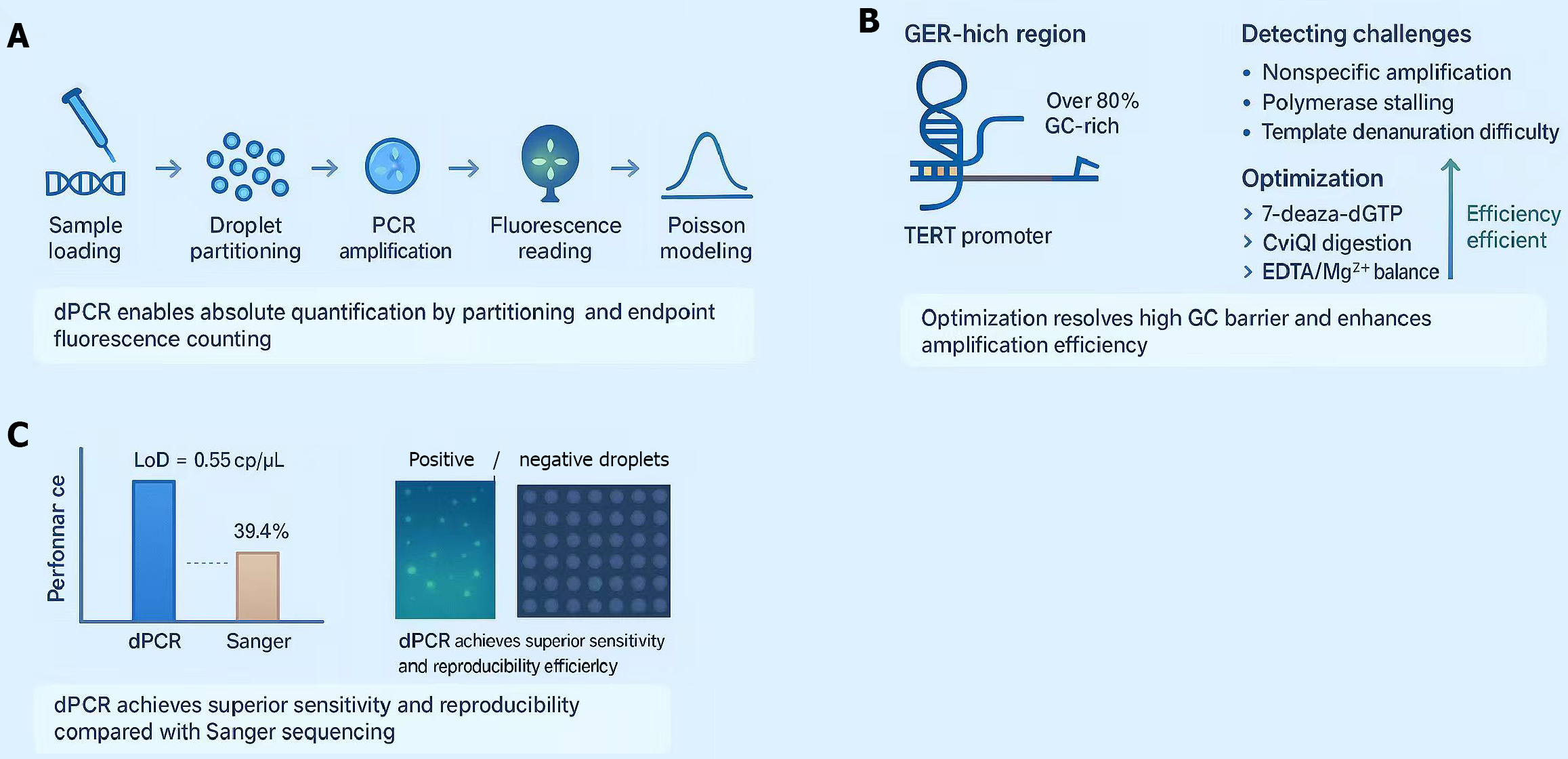
Figure 2 Optimized digital polymerase chain reaction system for ultra-sensitive detection of the guanine-cytosine-rich telomerase reverse transcriptase promoter C228T mutation.
A: Workflow schematic: Illustration of the digital polymerase chain reaction (dPCR) process, including sample loading, droplet partitioning, thermal cycling, fluorescence signal detection, and Poisson-based absolute quantification. This system enables precise molecular counting through endpoint fluorescence analysis; B: Optimization strategies: The guanine-cytosine-rich telomerase reverse transcriptase promoter region [> 80% guanine-cytosine (GC) content] forms stable secondary structures that hinder amplification. Optimization with 7-deaza-dPCR (reducing GC stacking), CviQ1 restriction enzyme digestion (simplifying templates), and Mg2+/ethylenediaminetetraacetic acid balancing (enhancing specificity and signal strength) effectively overcomes GC-barriers and improves amplification efficiency; C: Performance validation: Comparative analysis between dPCR and Sanger sequencing demonstrates superior performance of dPCR, achieving 100% sensitivity, 90% specificity, and a detection limit of 0.55 cp/μL. Representative fluorescence droplet images illustrate clear separation of positive (green) and negative (gray) droplets at the endpoint. dPCR: Digital polymerase chain reaction; GER: G-quadruplex-forming element-rich; GC: Guanine-cytosine; LOD: Limit of detection; PCR: Polymerase chain reaction; TERT: Telomerase reverse transcriptase; dGTP: Deoxyguanosine triphosphate; EDTA: Ethylenediaminetetraacetic acid.
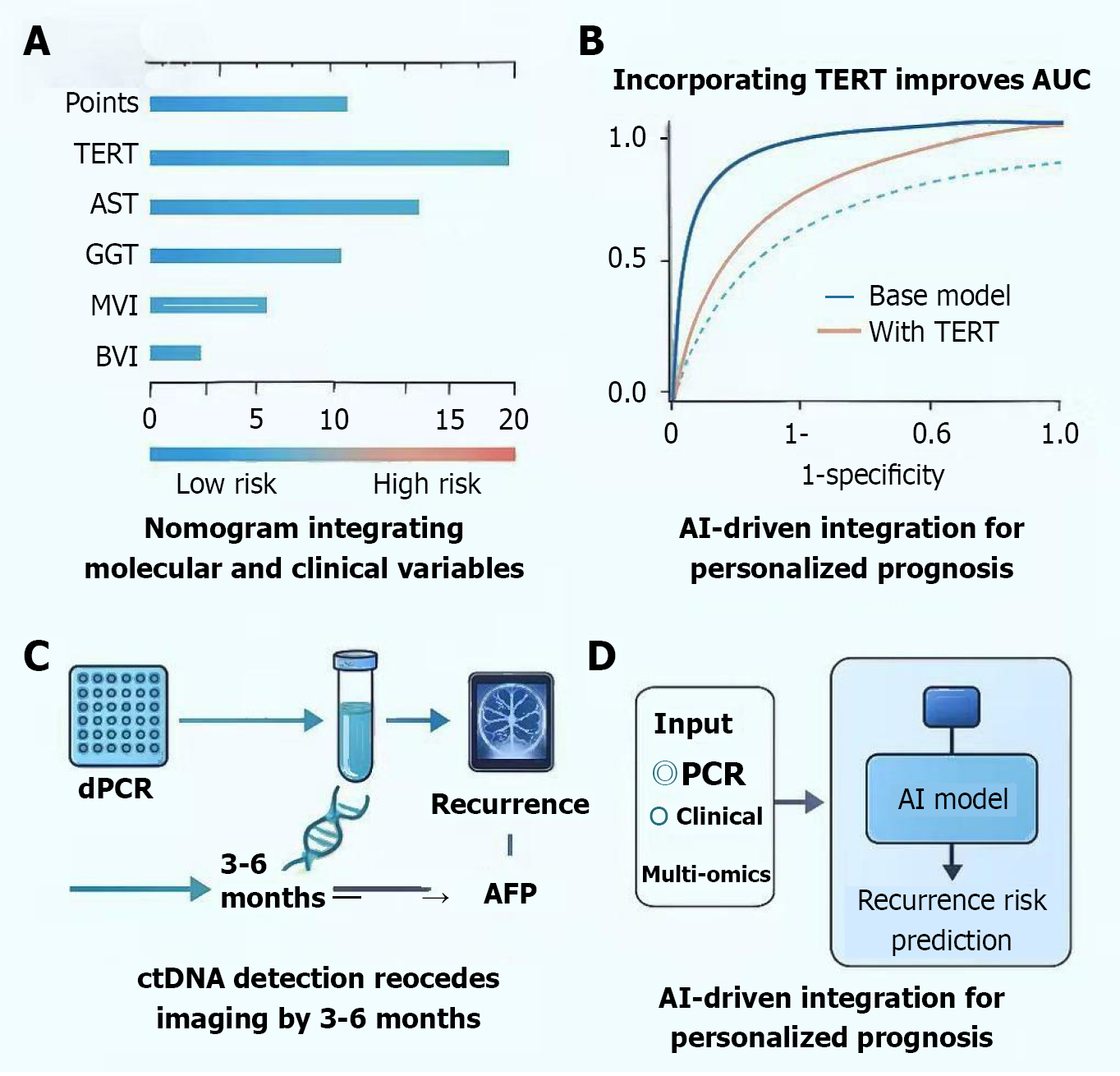
Figure 3 Clinical translation of telomerase reverse transcriptase mutation detection integrating molecular diagnostics, prognostic modeling, and artificial intelligence prediction.
A: Nomogram model combining telomerase reverse transcriptase (TERT) mutation status with clinical variables (aspartate aminotransferase, γ-glutamyl transpeptidase, microvascular invasion, blood vessel invasion) predicts 1-year, 2-year, and 5-year overall survival/disease-free survival, distinguishing high-risk (red) and low-risk (blue) groups; B: Model performance comparison: Incorporation of TERT mutation improves predictive accuracy (area under the curve 0.83 vs 0.75) and decision curve benefit; C: Dynamic monitoring: Digital polymerase chain reaction-based ctDNA detection identifies TERT mutations 3-6 months before radiologic recurrence and alpha-fetoprotein elevation, enabling early relapse prediction; D: Artificial intelligence -integrated framework: Machine learning model (random forest/extreme gradient boosting) fuses molecular, clinical, and multi-omics data for individualized recurrence risk assessment. TERT: Telomerase reverse transcriptase; AST: Aspartate aminotransferase; GGT: γ-glutamyl transpeptidase; MVI: Microvascular invasion; BVI: Blood vessel invasion; dPCR: Digital polymerase chain reaction; AFP: Alpha-fetoprotein; AUC: Area under the curve; AI: Artificial intelligence; PCR: Polymerase chain reaction.
- Citation: Shao S, Xiong Y, Kong MW, Yu Y, Zhang CX. Digital polymerase chain reaction detection of telomerase reverse transcriptase promoter mutations in hepatitis B virus related hepatocellular carcinoma. World J Gastrointest Oncol 2026; 18(4): 116504
- URL: https://www.wjgnet.com/1948-5204/full/v18/i4/116504.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v18.i4.116504