Copyright: ©Author(s) 2026.
World J Gastrointest Oncol. Apr 15, 2026; 18(4): 115635
Published online Apr 15, 2026. doi: 10.4251/wjgo.v18.i4.115635
Published online Apr 15, 2026. doi: 10.4251/wjgo.v18.i4.115635
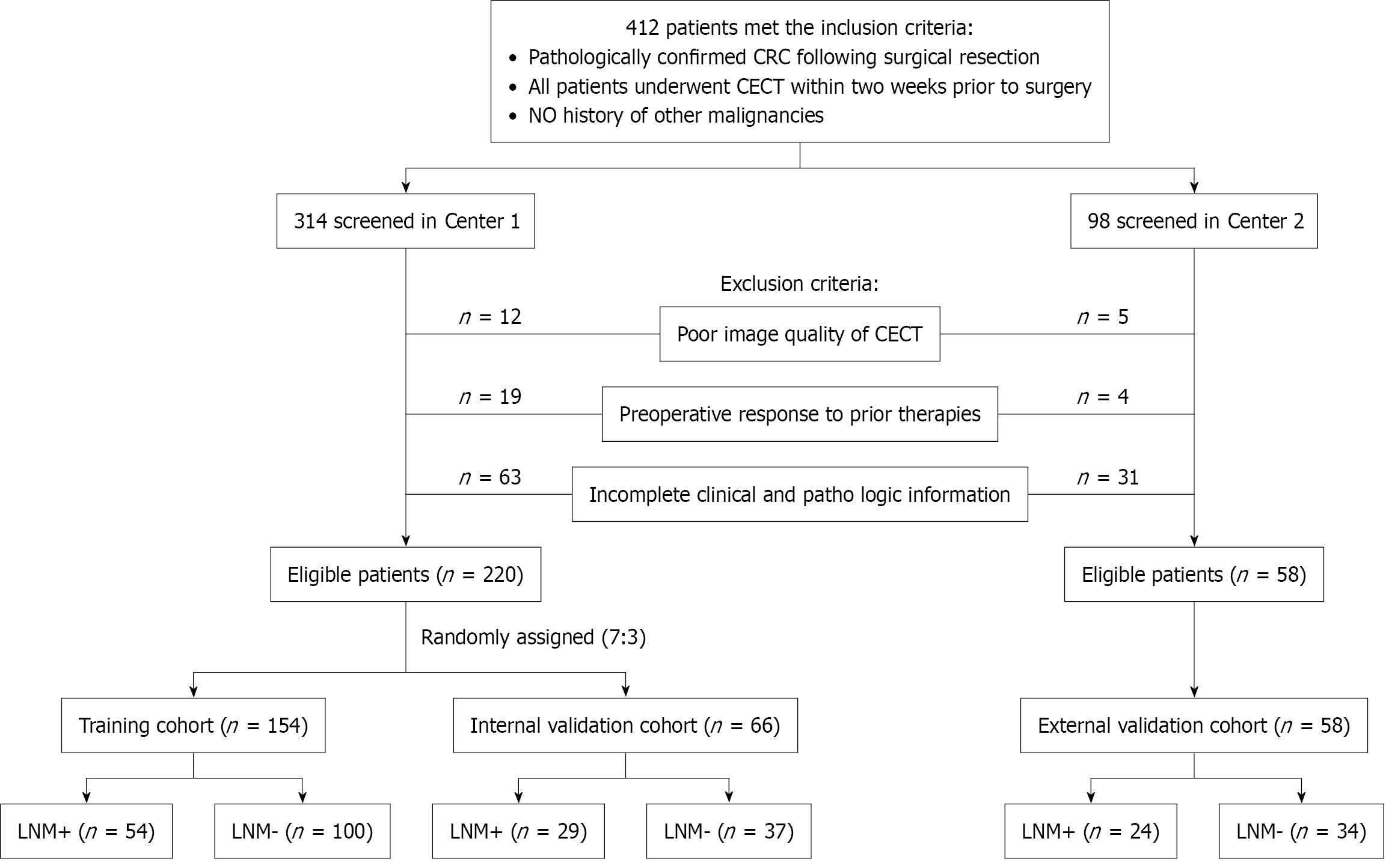
Figure 1 Flowchart of patient enrollment.
CRC: Colorectal Cancer; CECT: Contrast-enhanced computed tomography; LNM: Lymph node metastasis.
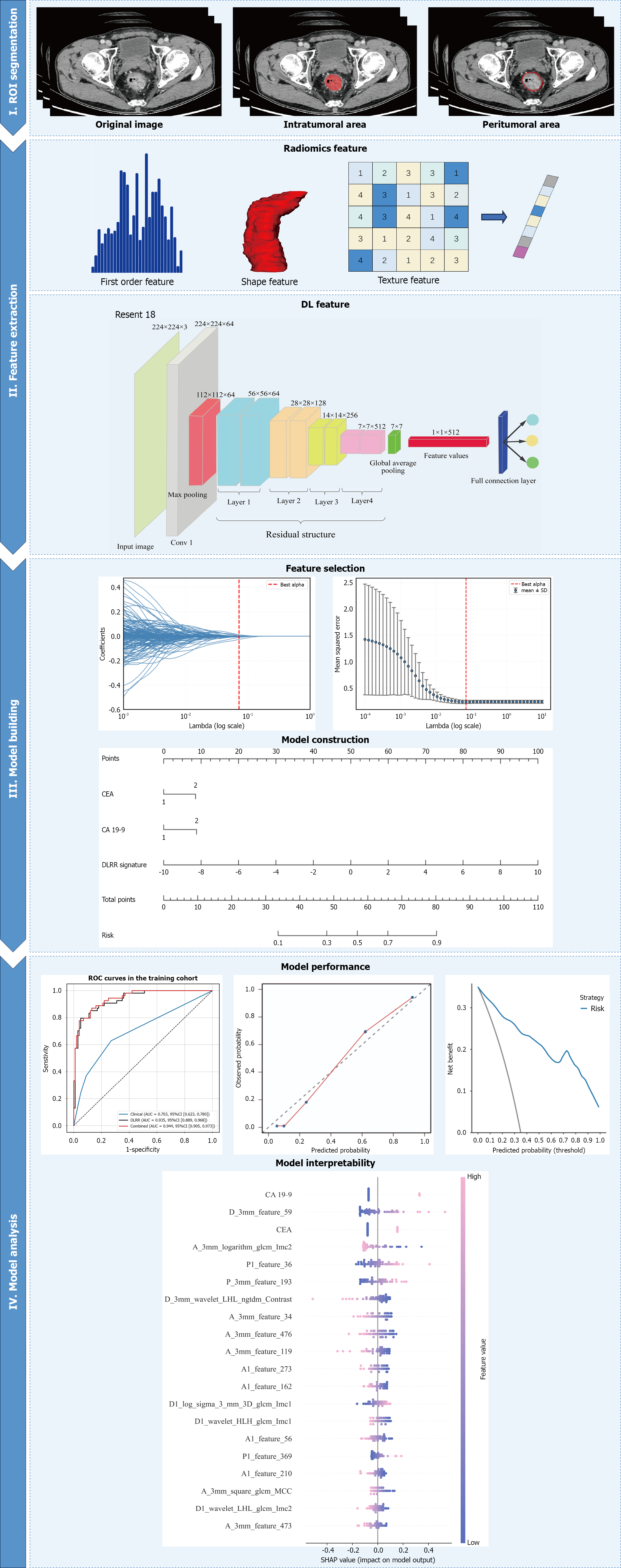
Figure 2 Workflow of the study.
ROI: Region of interest; DL: Deep learning; ROC: Receiver operating characteristic; CA 19-9: Carbohydrate antigen 19-9; CEA: Carcinoembryonic antigen; DLRR: Deep learning-radiomics integrated model; SHAP: SHapley Additive exPlanation.
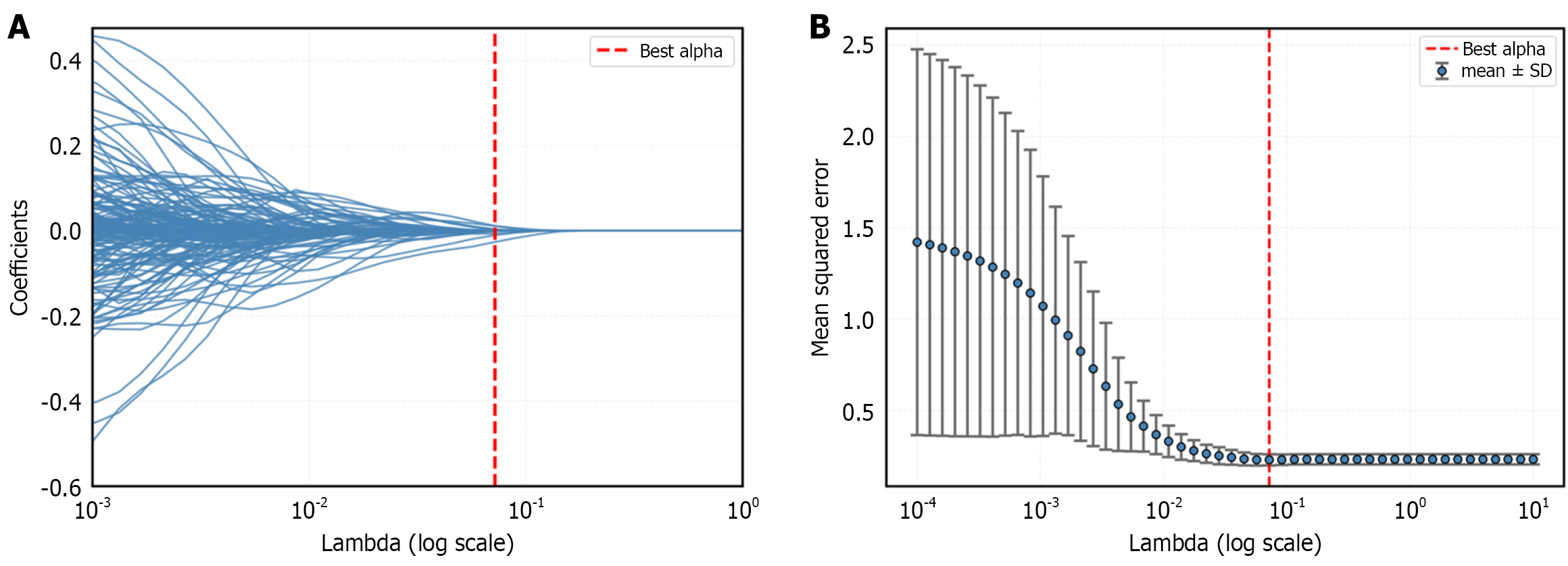
Figure 3 Radiomics and deep learning feature selection.
Radiomics feature selection using least absolute shrinkage and selection operator logistic regression: A: Selection of the tuning parameter; B: Least absolute shrinkage and selection operator coefficient profiles of the radiomics features.
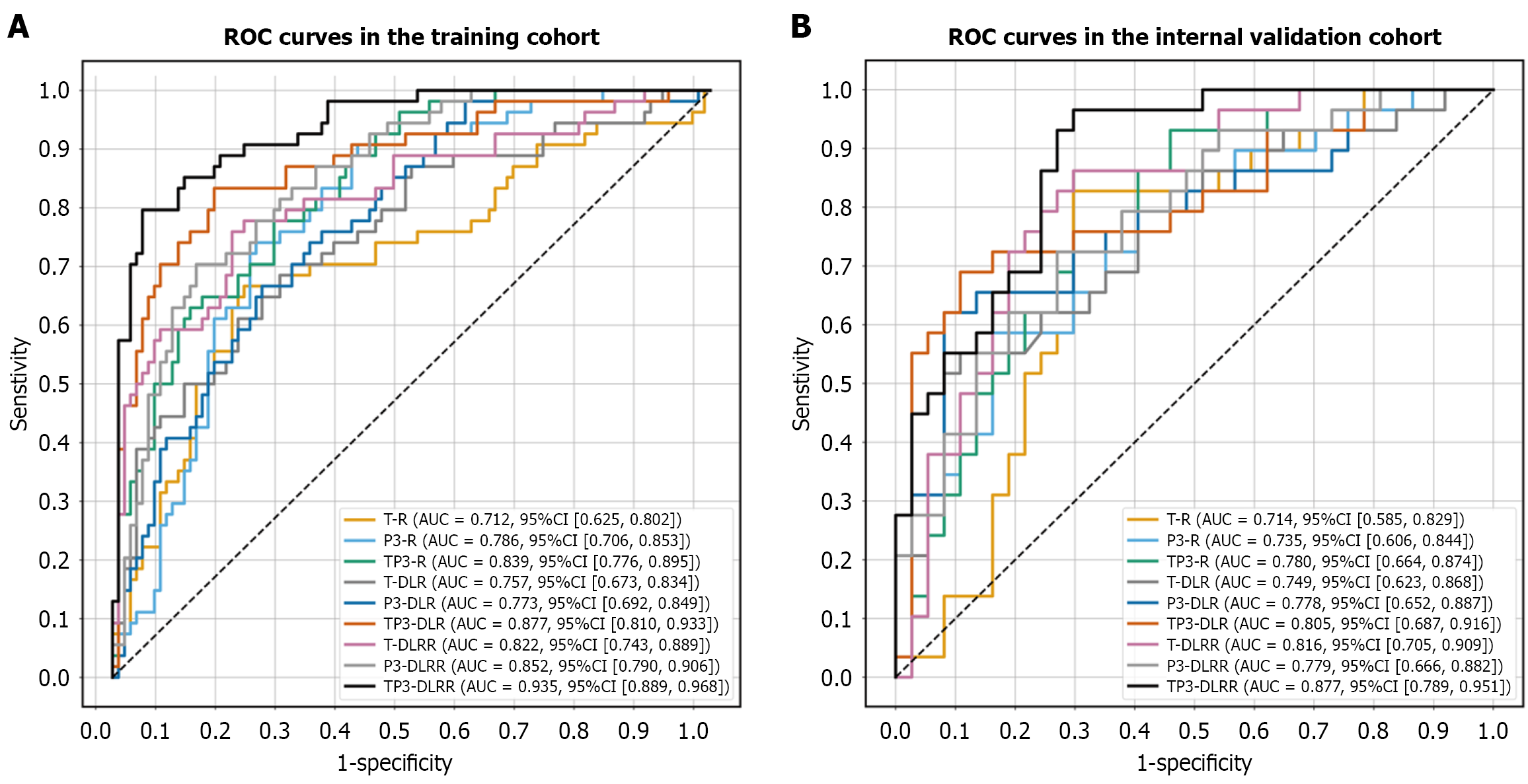
Figure 4 Comparative analysis of area under the receiver operating characteristic curve for the nine models.
A: Comparison of the area under the receiver operating characteristic curve of the nine models in the training cohort; B: Comparison of the area under the receiver operating characteristic curve of the nine models in the internal validation cohort. ROC: Receiver operating characteristic; AUC: Area under the receiver operating characteristic curve; T: The intratumoral model; P3: The peritumoral-3mm model; TP3: The intra-peritumoral-3mm model; R: The radiomics model; DLR: The deep learning radiomics model; DLRR: Deep learning-radiomics integrated model.
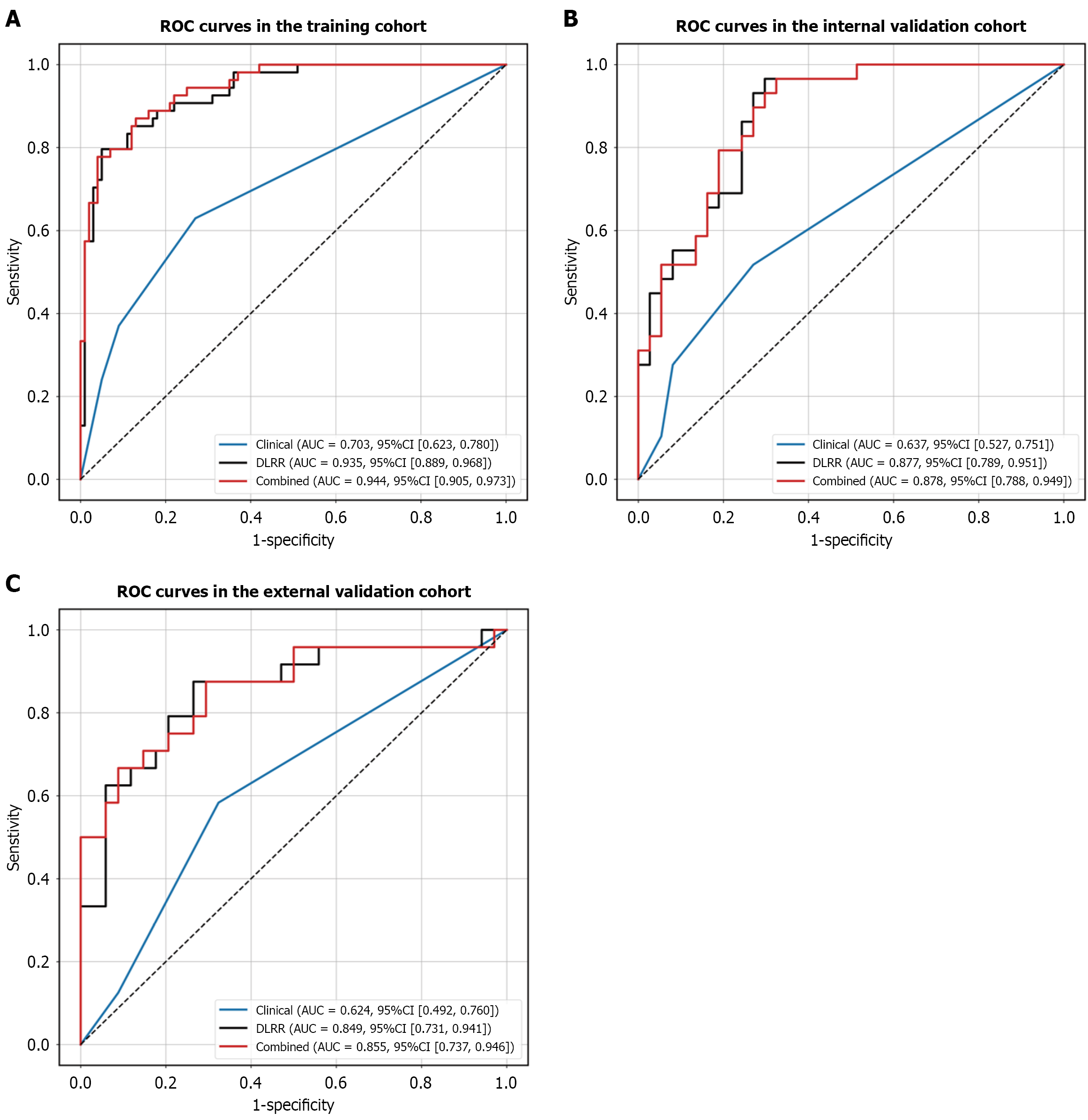
Figure 5 Receiver operating characteristic curves of the three models.
A-C: Receiver operating characteristic curves of the clinical, deep learning-radiomics integrated model and combined model for predicting lymph node metastasis in the training cohort (A), internal validation cohort (B), and external validation cohort (C), respectively. ROC: Receiver operating characteristic; AUC: Area under the receiver operating characteristic curve; DLRR: Deep learning-radiomics integrated model; Combined: The clinical-deep learning-radiomics model; CI: Confidence interval.
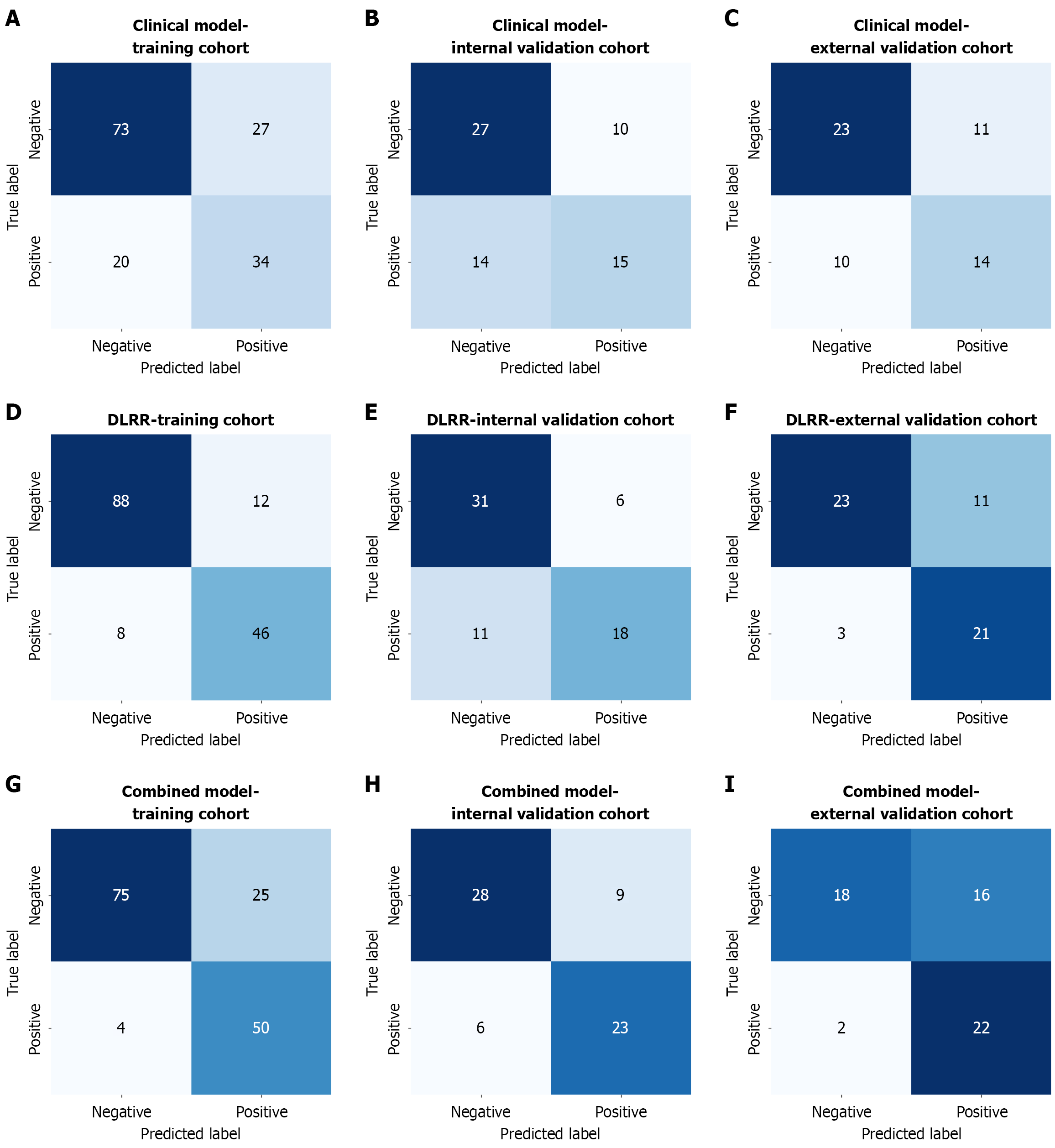
Figure 6 Confusion matrix of patient-level models in the training, internal validation and external validation cohorts, respectively.
A-C: Clinical model; D-F: Deep learning-radiomics integrated model; G-I: Combined model. Positive represents lymph node metastasis status; negative represents non-lymph node metastasis status. DLRR: Deep learning-radiomics integrated model; Combined: The clinical-deep learning-radiomics model.
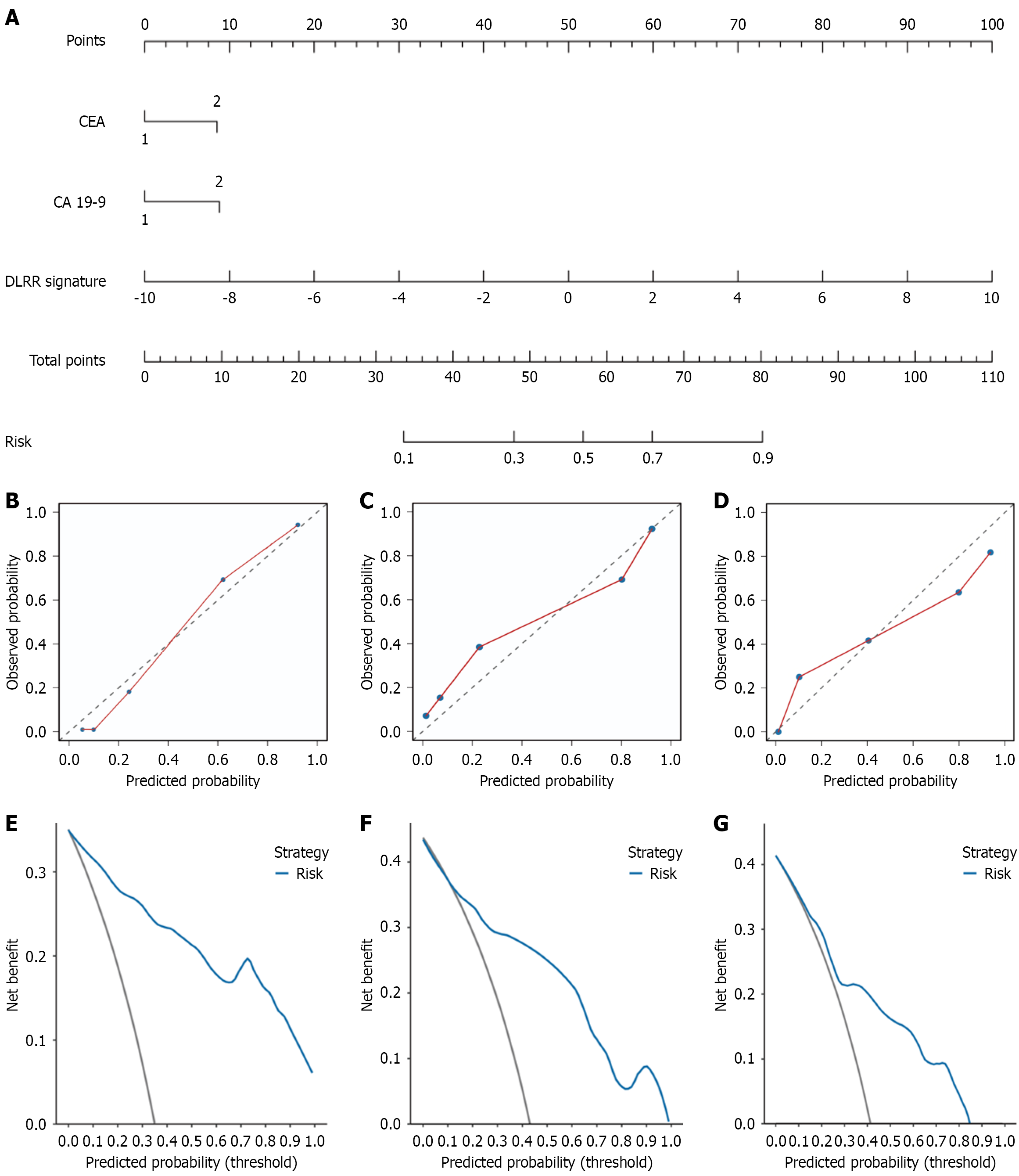
Figure 7 The clinical-deep learning-radiomics nomogram construction and performance evaluation.
A: Is a nomogram for individual prediction of lymph node metastasis risk combined with the deep learning-radiomics integrated model and independent clinical features; B-D: Are the calibration curves of the clinical-deep learning-radiomics nomogram in the training, internal validation and external validation cohorts, respectively; E-G: Are the decision curves of the clinical-deep learning-radiomics nomogram in the training, internal validation and external validation cohorts, respectively. CA 19-9: Carbohydrate antigen 19-9; CEA: Carcinoembryonic antigen; DLRR: Deep learning-radiomics integrated model.
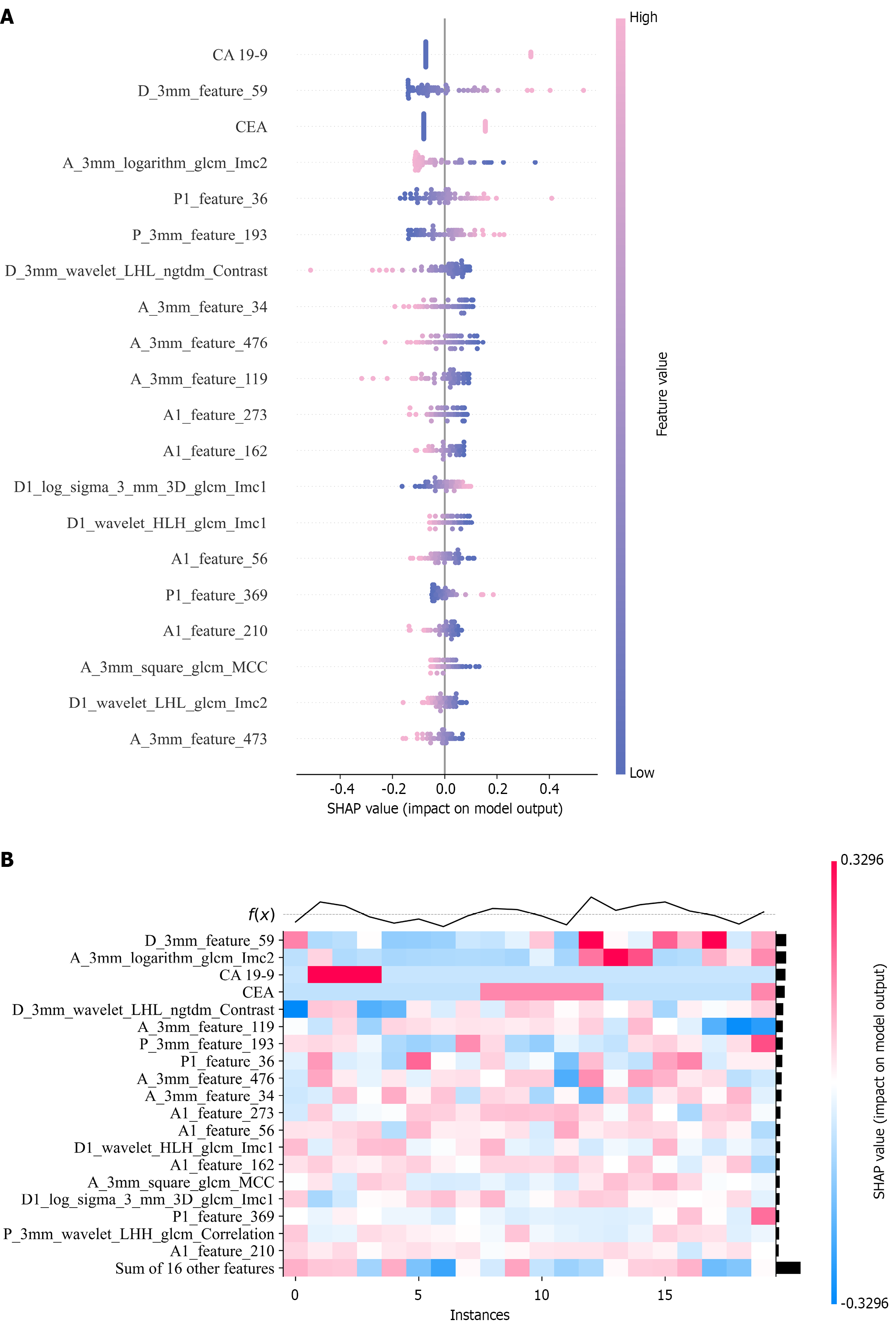
Figure 8 Global model explanation by the SHapley Additive exPlanation method.
A: The SHapley Additive exPlanation beeswarm plot shows the positive or negative effects of each feature on the prediction probability through red and blue colors; B: The SHapley Additive exPlanation heatmap plot shows the direction and intensity of influence for each feature of all cases in the model. SHAP: SHapley Additive exPlanation; CA 19-9: Carbohydrate antigen 19-9; CEA: Carcinoembryonic antigen.
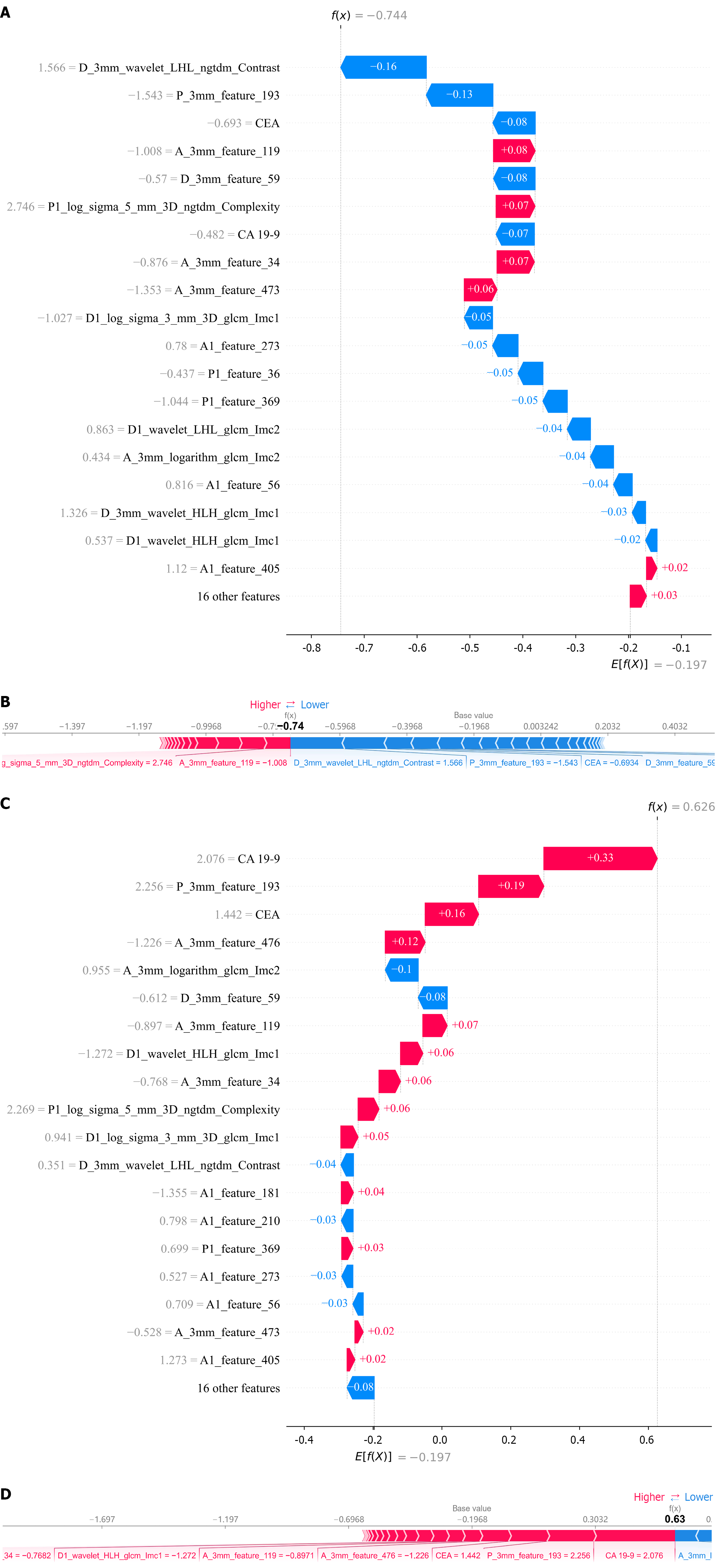
Figure 9 Local model explanation by the SHapley Additive exPlanation method.
A and B: Shows examples of correctly predicted lymph node metastasis-negative cases; C and D: Shows examples of correctly predicted lymph node metastasis-positive cases. CA 19-9: Carbohydrate antigen 19-9; CEA: Carcinoembryonic antigen.
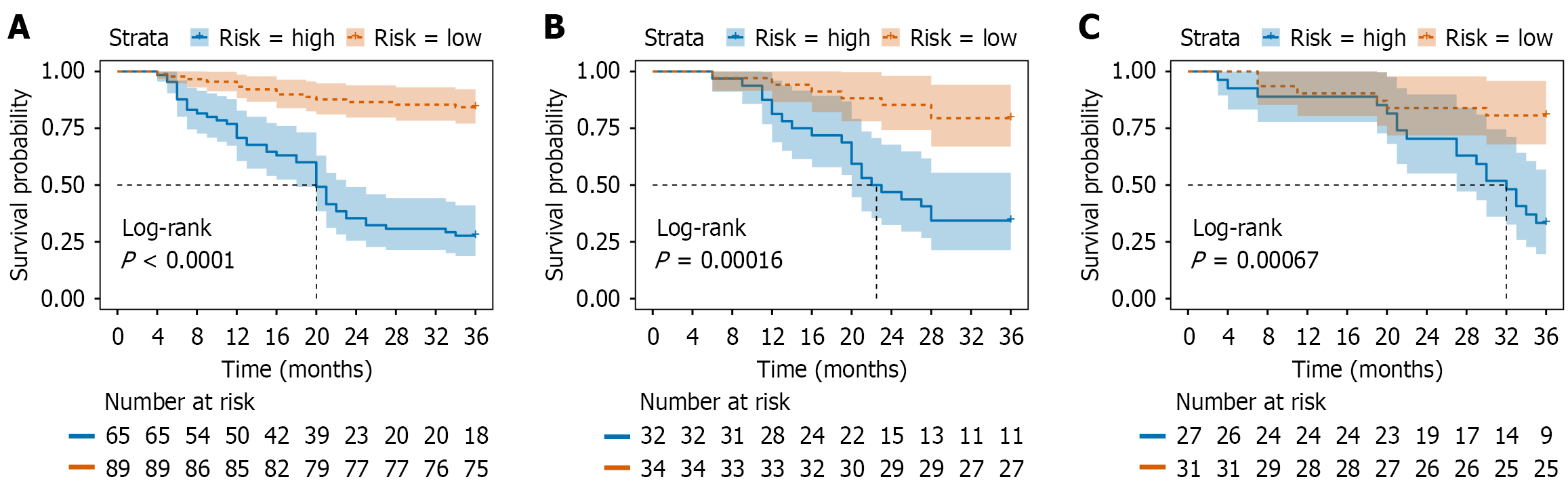
Figure 10 Kaplan-Meier curves of 3-year recurrence-free survival.
A: Training cohort; B: Internal validation cohort; C: External validation cohort, respectively.
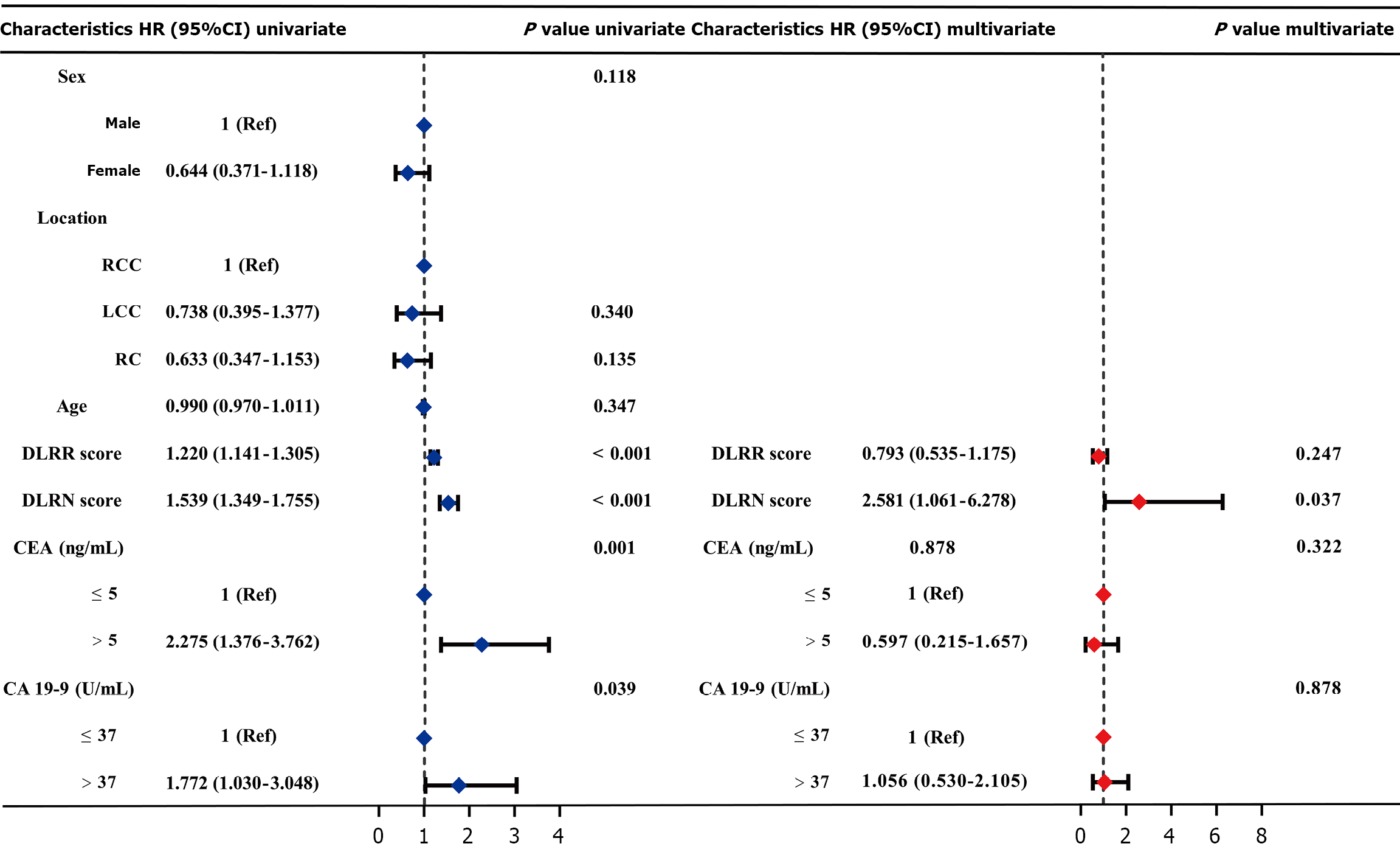
Figure 11 Forest plot of univariate Cox and multivariate Cox analysis based on the training cohort.
HR: Hazard ratio; CI: Confidence interval; RCC: Right-sided colon cancer; LCC: Left-sided colon cancer; RC: Rectal cancer; DLRR: Deep learning-radiomics integrated model; DLRN: The clinical-deep learning-radiomics nomogram; CEA: Carcinoembryonic antigen; CA 19-9: Carbohydrate antigen 19-9.
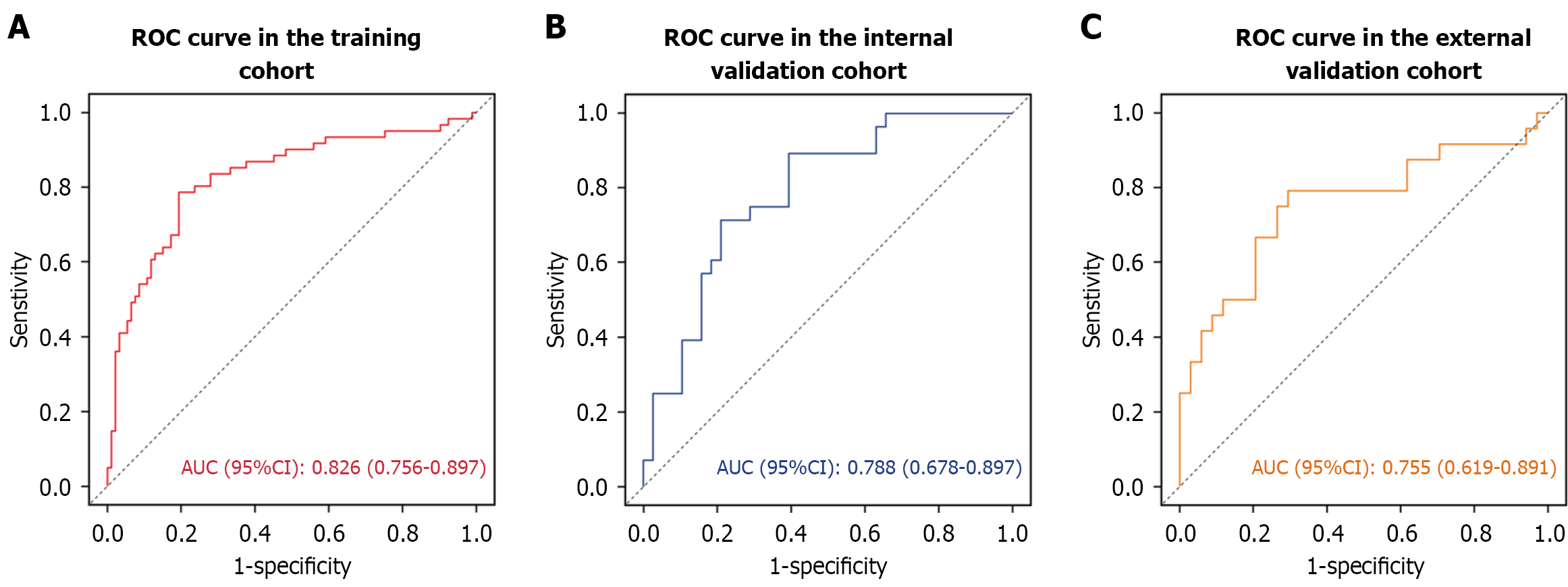
Figure 12 The receiver operating characteristic curves of a prognostic model.
A: Performance of a prognostic model in the training cohort; B: Performance of a prognostic model in the internal validation cohort; C: Performance of a prognostic model in the external validation cohort. ROC: Receiver operating characteristic. AUC: Area under the receiver operating characteristic curve; CI: Confidence interval.
- Citation: Lei XD, Qian GX, Sun ZG, Tang ZQ, Liu YC, Du R, Li YH. Deep learning radiomics nomogram based on multi-regional features for predicting lymph node metastasis and prognosis in colorectal cancer. World J Gastrointest Oncol 2026; 18(4): 115635
- URL: https://www.wjgnet.com/1948-5204/full/v18/i4/115635.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v18.i4.115635