Published online Mar 27, 2026. doi: 10.4254/wjh.v18.i3.117134
Revised: December 23, 2025
Accepted: February 4, 2026
Published online: March 27, 2026
Processing time: 117 Days and 2.8 Hours
Hepatocellular carcinoma (HCC) is a life-threatening malignant tumor with complex pathogenesis and limited treatment. The core clinical protocols include surgical resection, liver transplantation, local ablation, transcatheter arterial che
To systematically elucidate the expression pattern, clinical significance, and mo
CUL2 protein expression in 125 HCC tissue specimens was assessed via immunohistochemistry. We integrated global HCC microarray and high-throughput sequencing datasets from a multicenter cohort (9283 specimens: 5204 HCC and 4079 non-tumor tissues) for CUL2 mRNA expression analysis. Single-cell RNA sequencing and spatial transcriptomics were employed to characterize CUL2 expression patterns. The correlation between CUL2 expression and clinicopathological parameters was analyzed in the Cancer Genome Atlas-LIHC cohort (n = 371). Gene set enrichment analysis (GSEA) was performed using Hallmark, Kyoto Encyclopedia of Genes and Genomes, and Gene Ontology gene sets. CRISPR functional screening and gene co-expression network analysis were applied to elucidate molecular mechanisms.
CUL2 was significantly overexpressed in HCC tissues at both mRNA and protein levels. Integrated multicenter analysis (n = 5204) confirmed marked CUL2 mRNA up-regulation (standardized mean difference = 0.70, P < 0.05) with moderate diagnostic value (area under the curve = 0.78). Patients with high CUL2 expression showed sig
CUL2 is significantly overexpressed in HCC tissues, which highlights its potential as a molecular biomarker and a promising therapeutic target for HCC management.
Core Tip: Cullin-2 (CUL2) is highly expressed in mRNA and protein levels in hepatocellular carcinoma (HCC). Multicenter analysis (n = 5204) confirmed its high expression (standardized mean difference = 0.70) and diagnostic value (area under the curve = 0.78). The overall survival of patients with high expression was shortened by about 4.1 years. Functional enrichment showed that it was associated with E2F, G2/M and MYC pathways and promoted cell cycle progression. Knockout of CUL2 significantly inhibited the growth of HCC cells, indicating that CUL2 drives HCC progression through proliferation and migration.
- Citation: Tang P, Nong QH, Xiong KH, Tang HH, Huang BB, Huang GL, Li Q, He SP, Chen G, Feng ZB, Li JD. Overexpression and clinicopathological significance of cullin-2 in hepatocellular carcinoma progression. World J Hepatol 2026; 18(3): 117134
- URL: https://www.wjgnet.com/1948-5182/full/v18/i3/117134.htm
- DOI: https://dx.doi.org/10.4254/wjh.v18.i3.117134
Liver cancer is the sixth most common cancer in the world and the third leading cause of cancer-related mortality. Hepatocellular carcinoma (HCC) is the most common histological subtype of liver cancer, accounting for about 80% of all primary liver cancers[1,2]. While significant progress has been made in relation to treatment strategies for HCC in recent years, the five-year survival rate of patients with HCC remains suboptimal due to the heterogeneity and invasiveness of HCC[3]. This situation urgently requires us to further elucidate the molecular pathogenesis of HCC and explore new biomarkers and therapeutic targets to improve the prognosis of patients. This situation indicates an urgent need to further elucidate the molecular pathogenesis of HCC and explore new biomarkers and therapeutic targets to improve HCC patients’ prognosis.
The Cullin family proteins are the core components of Cullin-RING E3 ubiquitin ligases (CRLs) and an important part of the ubiquitin-proteasome system. By mediating the ubiquitination and degradation of various substrate proteins such as cell cycle regulators and transcription factors, CRLs control key cellular processes, including cell cycle progression, DNA damage repair, transcriptional regulation, and signal transduction[4]. As a member of the Cullin family, Cullin-2 (CUL2) acts as a central scaffold protein in the CUL2–RBX1-EloC-EloB complex. It binds to substrate recognition subunits, such as the von Hippel-Lindau tumor suppressor, to form an E3 ubiquitin ligase complex similar to the SKP1-CUL1-F-box protein complex[5,6]. This multi-protein combination plays a key role in a number of basic physiological and pathological pathways, including oxygen sensing, cell proliferation, apoptosis, and angiogenesis[7]. There is a growing body of evidence that CUL2 is abnormally expressed in various malignant tumors, suggesting it promotes tumorigenesis and progression by regulating the stability of oncoproteins or tumor suppressors[8]. For example, dysregulation of CUL2 has been documented in gastric cancer[9], cervical cancer[10], and breast cancer[11], while its expression is associated with an aggressive phenotype and a poor patient prognosis. Despite this, the expression patterns, clinical relevance, and potential molecular mechanisms of CUL2 in HCC have not yet been fully characterized. It is worth noting that other Cullin family members have also been shown to be associated with the pathogenesis of HCC, forming a complex regulatory network. Studies have shown that CUL4A is highly expressed in HCC tissues, and it can promote the occurrence of HCC by promoting cell cycle progression and epithelial-mesenchymal transition[12]. In addition, the loss of CUL3 expression leads to abnormal activation of Nrf2, which in turn promotes the proliferation of biliary epithelial cells and promotes the development of cholangiocarcinoma[13]. The above research shows that Cullin family proteins constitute a wide and complex regulatory network in HCC, which further highlights the necessity of further exploring the role of CUL2 in this malignant tumor.
To address this gap in the literature, the present study sought to systematically elucidate the significant overexpression and prognostic value of CUL2 in HCC tissues at both the protein and mRNA levels by integrating HCC clinical tissue samples, immunohistochemistry (IHC) results, and large-scale global HCC microarray and high-throughput sequencing data. The results of this study will help clarify the key role of CUL2 in HCC and indicate new molecular markers and potential targets for prognostic evaluation and targeted therapy in HCC.
A total of 125 paired HCC and corresponding adjacent non-tumorous liver tissues were obtained from patients who underwent radical resection at Wuming Hospital of Guangxi Medical University between January 2019 and June 2023. All specimens were histopathologically confirmed as HCC. The study protocol was approved by the Institutional Ethics Committee of the hospital.
IHC staining was performed using a rabbit anti-human CUL2 polyclonal antibody (ABclonal Technology Co., Ltd., 1:500 dilution). Two independent pathologists, blinded to the clinical data, evaluated the staining intensity and the percentage of CUL2-positive HCC cells. The scoring criteria were as follows: (1) The proportion of positive cells was scored as zero (0%-5%), one (6%-25%), two (26%-50%), three (51%-75%), or four (> 75%); (2) Staining intensity was graded as: Zero (negative), one (weak/Light yellow), two (moderate/brownish-yellow), or three (strong/dark brown); and (3) A semi-quantitative H-score was calculated using the formula: H-score = staining intensity score × percentage score.
To investigate the mRNA expression level of CUL2, this study acquired relevant data from The Cancer Genome Atlas (TCGA, https://portal.gdc.cancer.gov/), the Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo/), the Sequence Read Archive (https://www.ncbi.nlm.nih.gov/sra), and the Array Express database (https://www.ebi.ac.uk/arrayexpress/). Relevant high-throughput sequencing datasets and datasets from the same platforms were merged separately. Appropriate data processing tools were utilized to remove batch effects between different datasets, resulting in multiple HCC expression matrices. To analyze the differential expression of the CUL2 protein between HCC tissues and non-cancerous liver tissues, violin plots of its expression level and receiver operating characteristic curves were plotted. If the area under the curve (AUC) was not less than 0.70 or 0.90, it indicated that the protein possessed moderate or strong discriminatory capability, respectively.
The association between CUL2 mRNA expression levels and patient prognosis in HCC was analyzed using the UALCAN online portal. The analytical procedure was as follows: The TCGA module was selected on the UALCAN homepage, followed by entry of the target gene CUL2 and selection of HCC. After clicking “Explore”, the “Survival” option under the “link for analysis” tab was utilized to perform the survival analysis and export the resultant plots.
Based on the HCC single-cell RNA sequencing dataset (GSE112271), t-SNE nonlinear dimensionality reduction and supervised learning algorithms were employed to visualize CUL2 mRNA expression across distinct HCC cellular subpopulations. Differential expression of CUL2 mRNA between normal hepatocytes and HCC cells was assessed using non-parametric tests.
Utilizing the HCC spatial transcriptomics dataset (GSE245908), stringent quality control was implemented by applying the thresholds of nFeature_Spatial > 500 and percent_mito < 25. Spatial localization and expression patterns of CUL2 mRNA, along with established HCC markers ARG1 and GPC3, were visualized on the HCC tissue sections.
Based on gene expression values, genes positively correlated with high CUL2 expression in HCC were identified through batch calculation using the following criteria: (1) Standardized mean difference (SMD) > 0 with P < 0.05 (or a confidence interval not spanning zero); and (2) Spearman correlation coefficient ≥ 0.30 with P < 0.05. Genes meeting these thresholds were subsequently subjected to Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses.
Publicly available clustered regularly interspaced short palindromic repeats (CRISPR) screening data were utilized to analyze the impact of CUL2 knockout on cellular growth across a panel of 21 HCC cell lines. A CRISPR gene effect score of < 0 was interpreted as indicative of a growth-promoting role for the gene.
In order to clarify the mechanism of CUL2 in liver cancer, the samples were divided into high and low expression groups based on the median expression of CUL2 in the TCGA-LIHC cohort, and the differences were analyzed by limma package. Using the R language clusterProfiler package, the genes were sorted by t value, and gene set enrichment analysis (GSEA) analysis was performed on the Hallmark, KEGG, and GO Biological Processes gene sets in the MSigDB database. The size of the included gene set was 15 to 500, and after 1000 permutation tests, P < 0.05 was considered significant after correction.
Based on the TCGA-LIHC cohort (n = 371), the correlation between CUL2 mRNA expression and key clinicopathological parameters was analyzed, including age, gender, tumor stage, histological grade, and T stage. The median expression was divided into high and low expression groups. The categorical variables were tested by χ2 test or Fisher’s exact test, and the continuous variables were tested by t-test or Wilcoxon rank sum test. All analyses were two-sided tests, and P < 0.05 was considered statistically significant.
Data analysis was performed using SPSS software (version 26.0) and R (version 4.2.1). Differences between the two groups were compared using student’s t-tests or Wilcoxon rank-sum tests, as appropriate. Survival analysis was conducted using the Kaplan-Meier method with log-rank tests for comparison. A two-sided P value of < 0.05 was considered statistically significant.
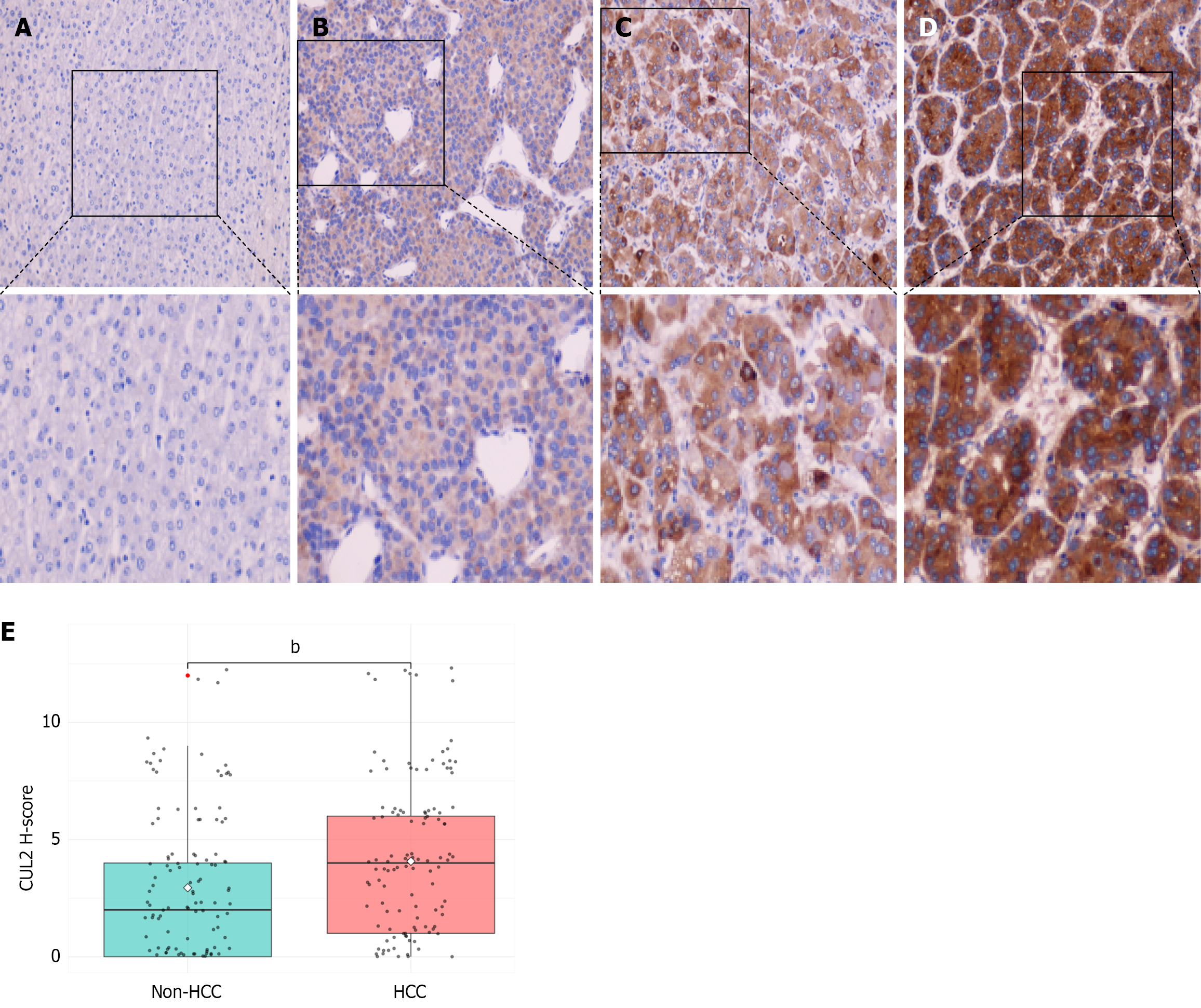
Immunohistochemical staining was performed to assess CUL2 protein expression in 125 paired HCC and corresponding adjacent non-tumorous liver tissues. Quantitative evaluation of the staining revealed predominant subcellular localization of CUL2 within the cytoplasmic compartment of HCC cells (Figure 1A-D). Comparative analysis using an independent samples t-test demonstrated significantly higher CUL2 protein levels in HCC tissues compared to matched peritumoral controls (Figure 1E, P < 0.01).
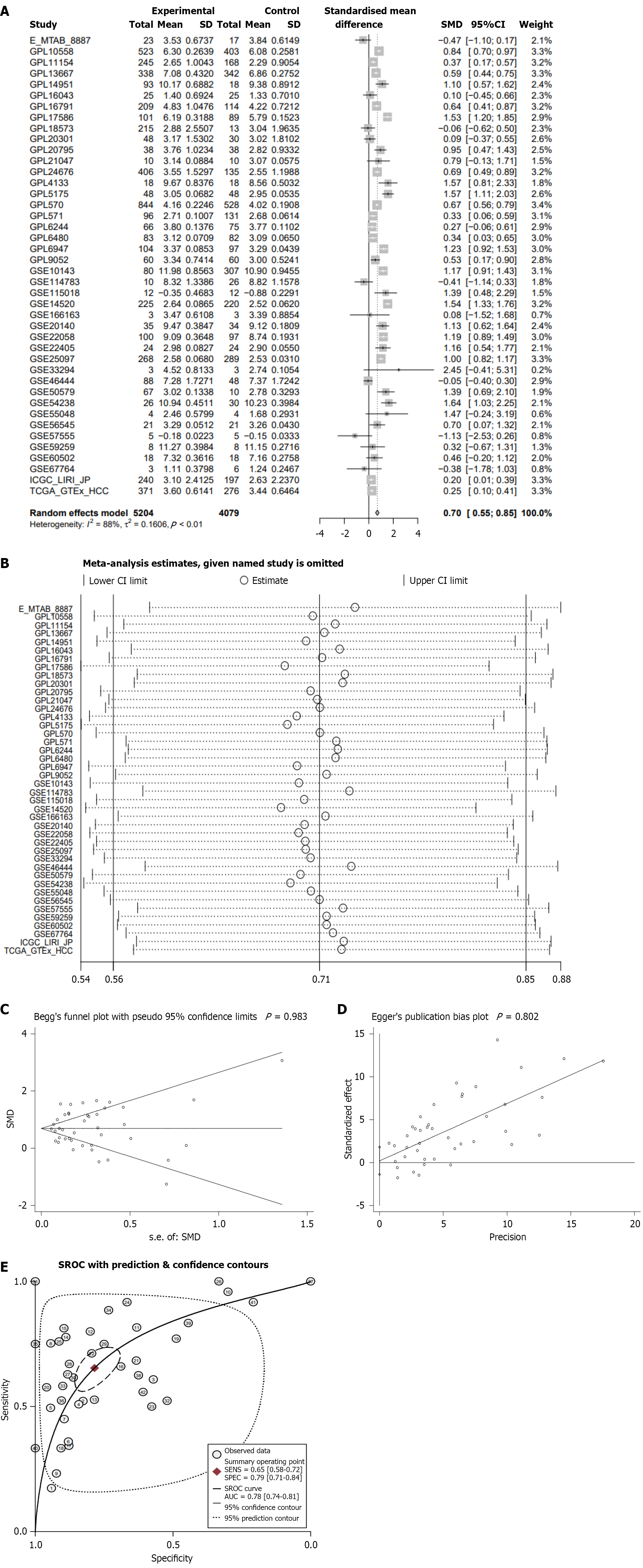
Integrated analysis of global HCC microarray and high-throughput sequencing datasets demonstrated significant upregulation of CUL2 mRNA in 5204 HCC tissues compared to 4079 non-neoplastic controls (Figure 2A) (SMD = 0.70, P < 0.05). Sensitivity analysis confirmed the stability of the pooled SMD estimate, as no individual dataset significantly altered the overall effect size upon sequential exclusion (Figure 2B). Both Begg’s and Egger’s tests indicated no substantial publication bias (P > 0.05) (Figure 2C and D). The summary receiver operating characteristic curve further evaluated the diagnostic performance of CUL2 mRNA in distinguishing HCC tissues from non-neoplastic controls, showing an AUC of 0.78 (95% confidence interval: 0.74-0.81), which supported the potential diagnostic value of CUL2 mRNA (Figure 2E).
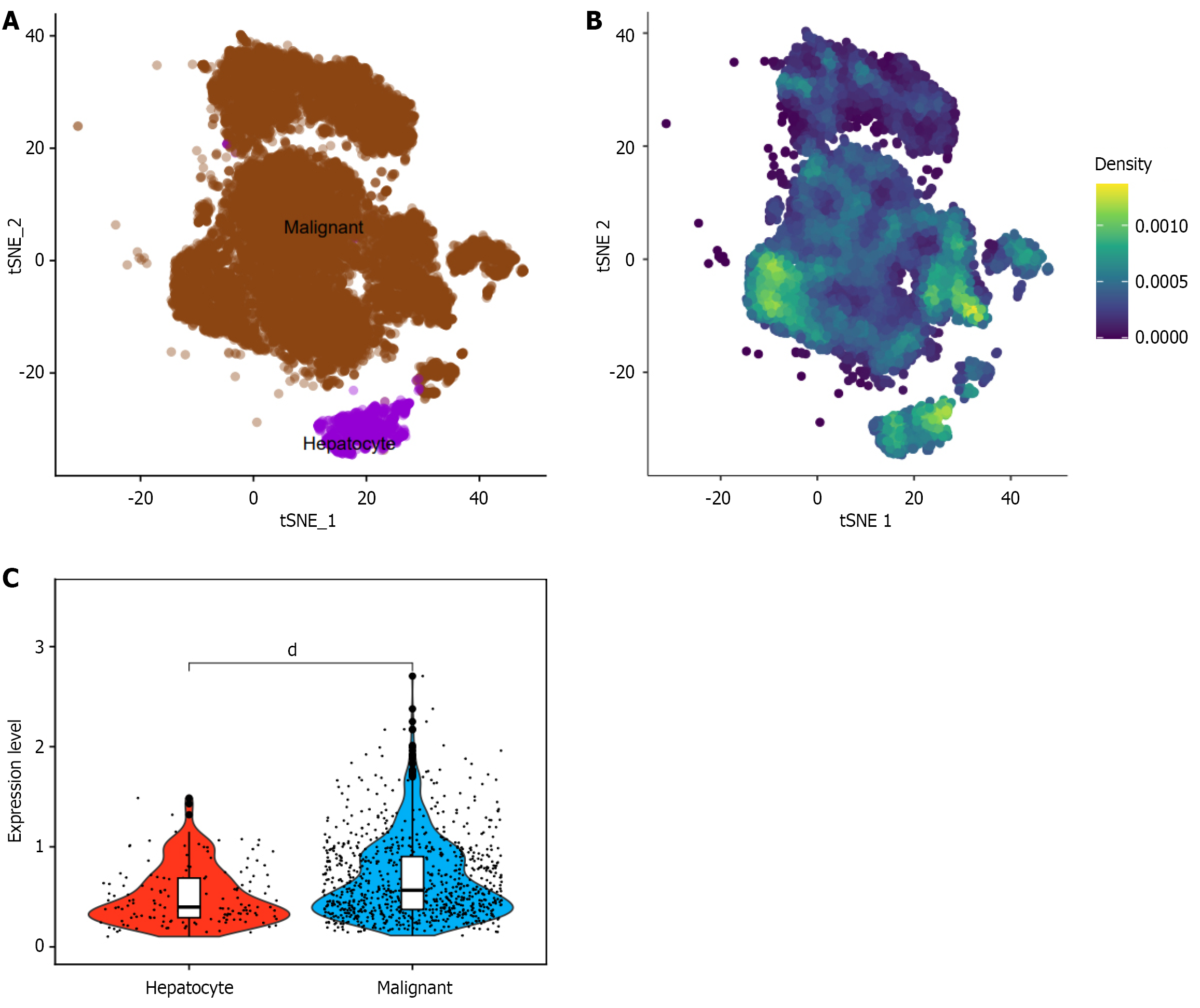
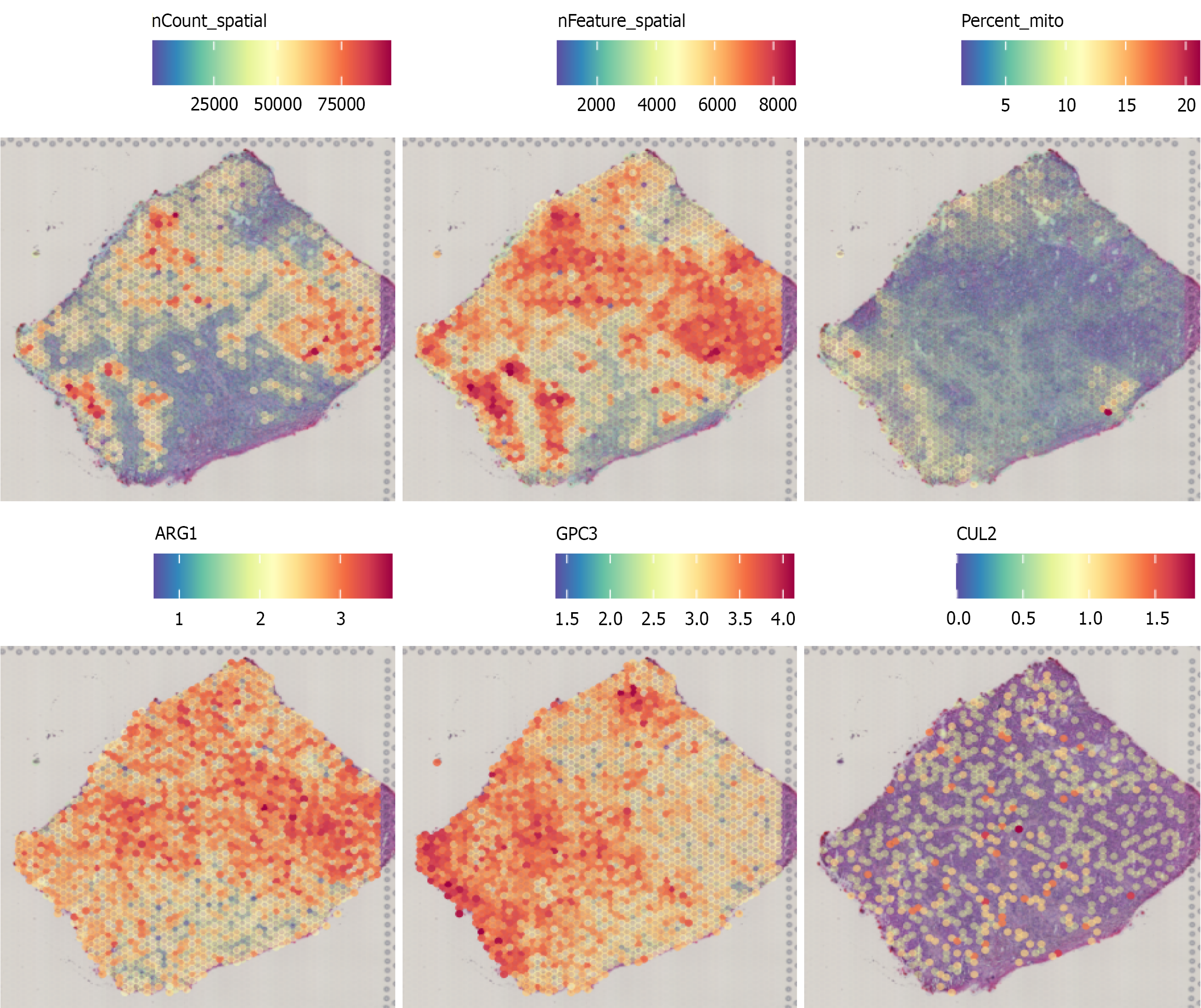
Density plots illustrated the characteristic expression pattern of CUL2 mRNA across cellular subpopulations, demonstrating significant overexpression in HCC single-cell clusters compared to normal hepatocytes (Figure 3) (P < 0.05). Spatial transcriptomic analysis further revealed a scattered, heterogeneous distribution pattern of CUL2 mRNA in HCC tissues, contrasting with the diffuse expression characteristics of established markers ARG1 and GPC3 (Figure 4).
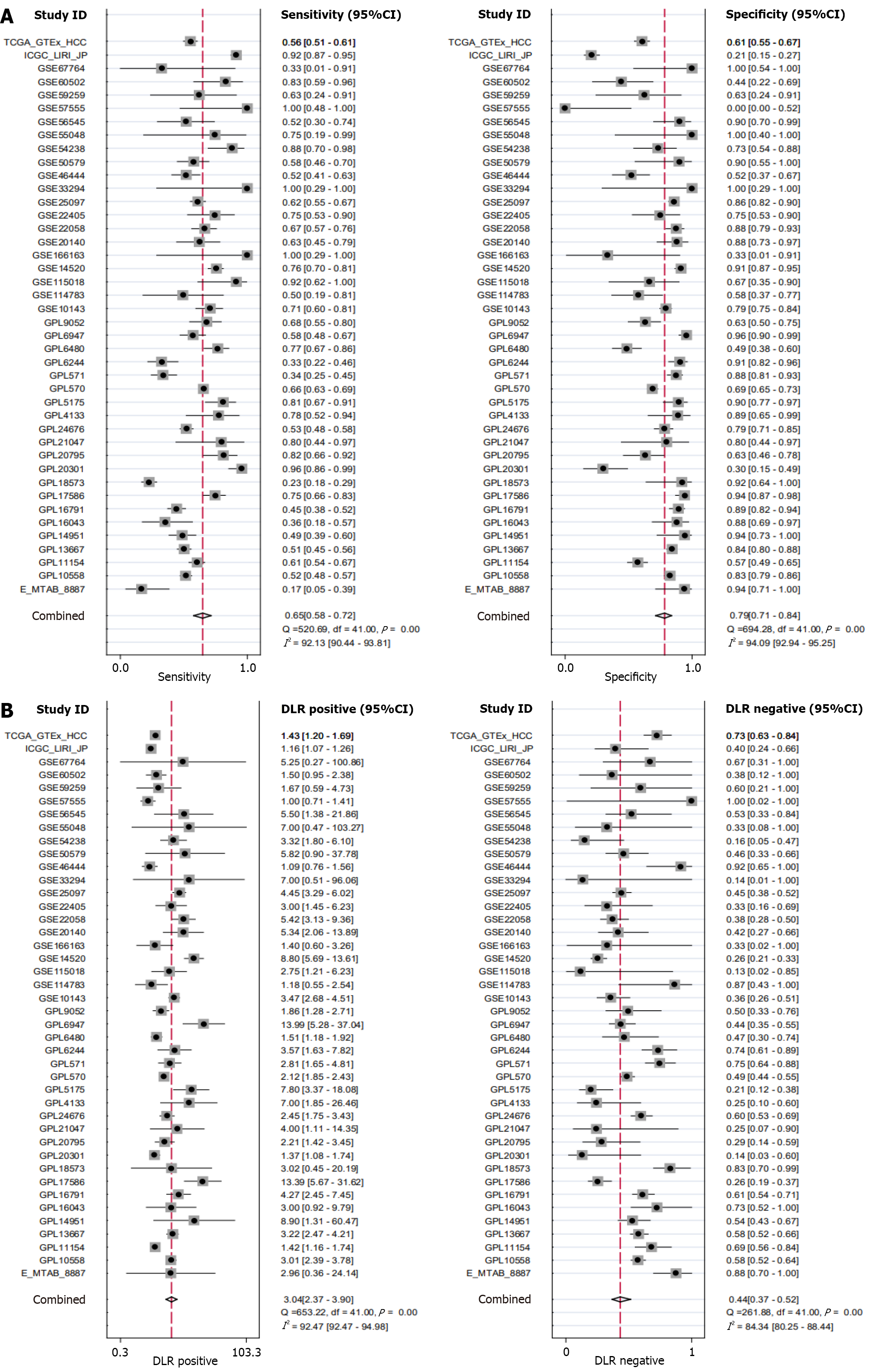
CUL2 mRNA exhibited moderate discriminatory power for distinguishing HCC tissues (Figure 5) (AUC = 0.78, P < 0.05), with a sensitivity of 0.65, specificity of 0.79, and positive and negative likelihood ratios of 3.04 and 0.44, respectively.
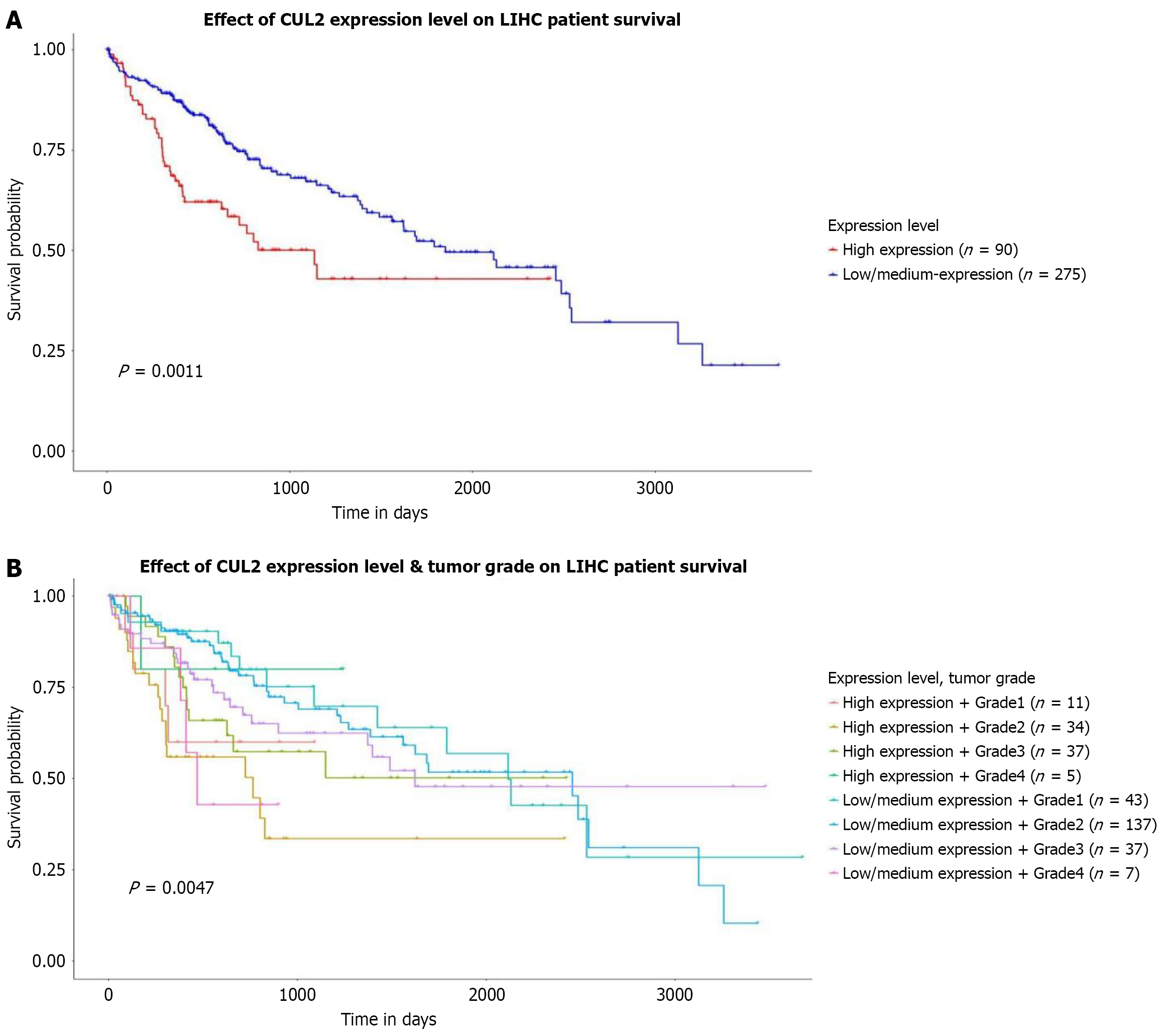
Analysis via the UALCAN database further elucidated the prognostic impact of CUL2 mRNA expression in HCC tissues. The results demonstrated that patients with high CUL2 mRNA expression exhibited significantly shorter overall survival (OS) compared to those with low expression (P = 0.0011) (Figure 6A). Furthermore, even within the same tumor grade, high CUL2 mRNA expression remained significantly associated with reduced OS (P = 0.0047) (Figure 6B).
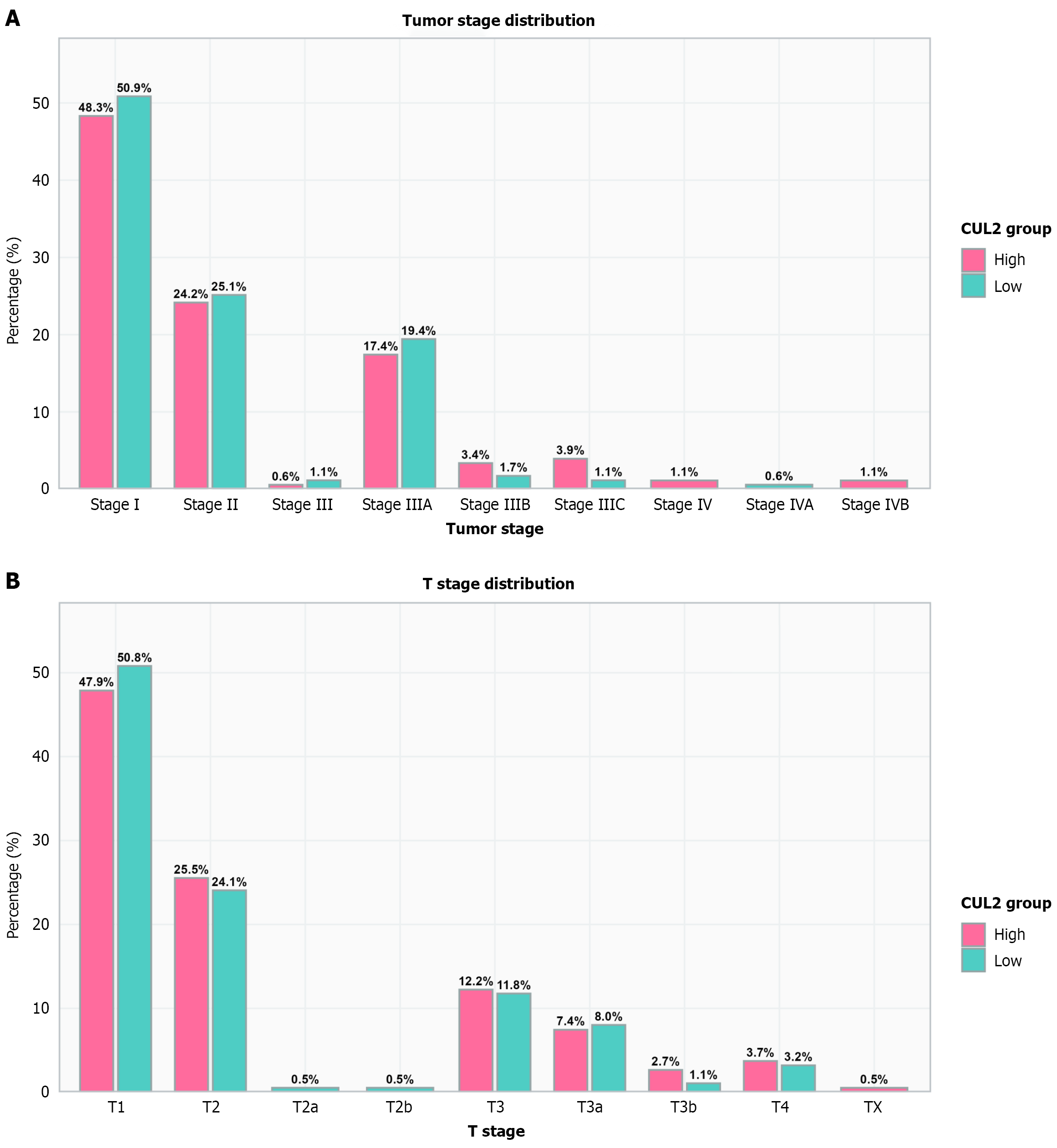
To evaluate the clinical significance of CUL2 expression, we analyzed its correlation with major clinicopathological parameters in the TCGA-LIHC cohort (n = 371). The distribution of primary tumor T stage between CUL2 expression groups indicates that both groups are predominantly comprised of patients with early-stage lesions (T1-T2) (Figure 7A). However, there is no significant difference in the overall AJCC stage distribution between CUL2 expression groups (Figure 7B), suggesting that the up-regulation of CUL2 may be an early event in the occurrence of HCC and persist throughout the disease progression.
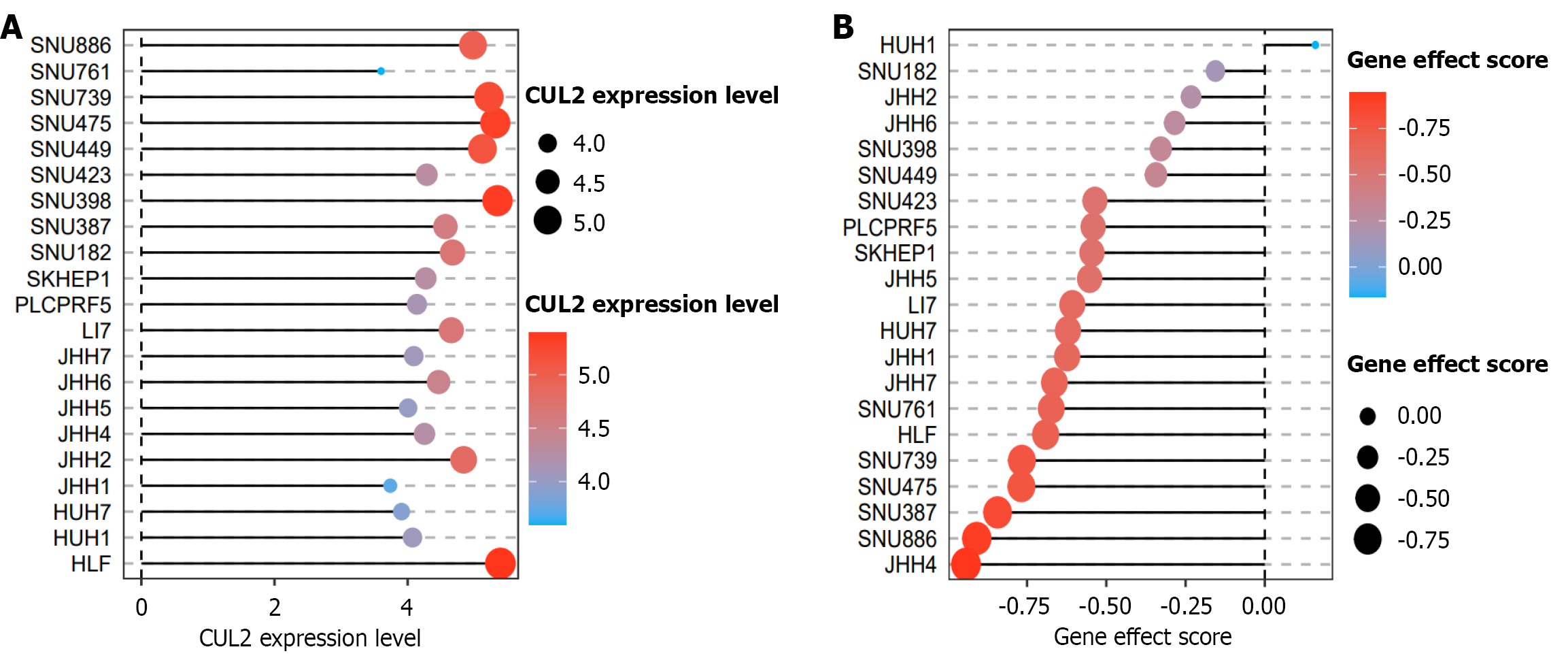
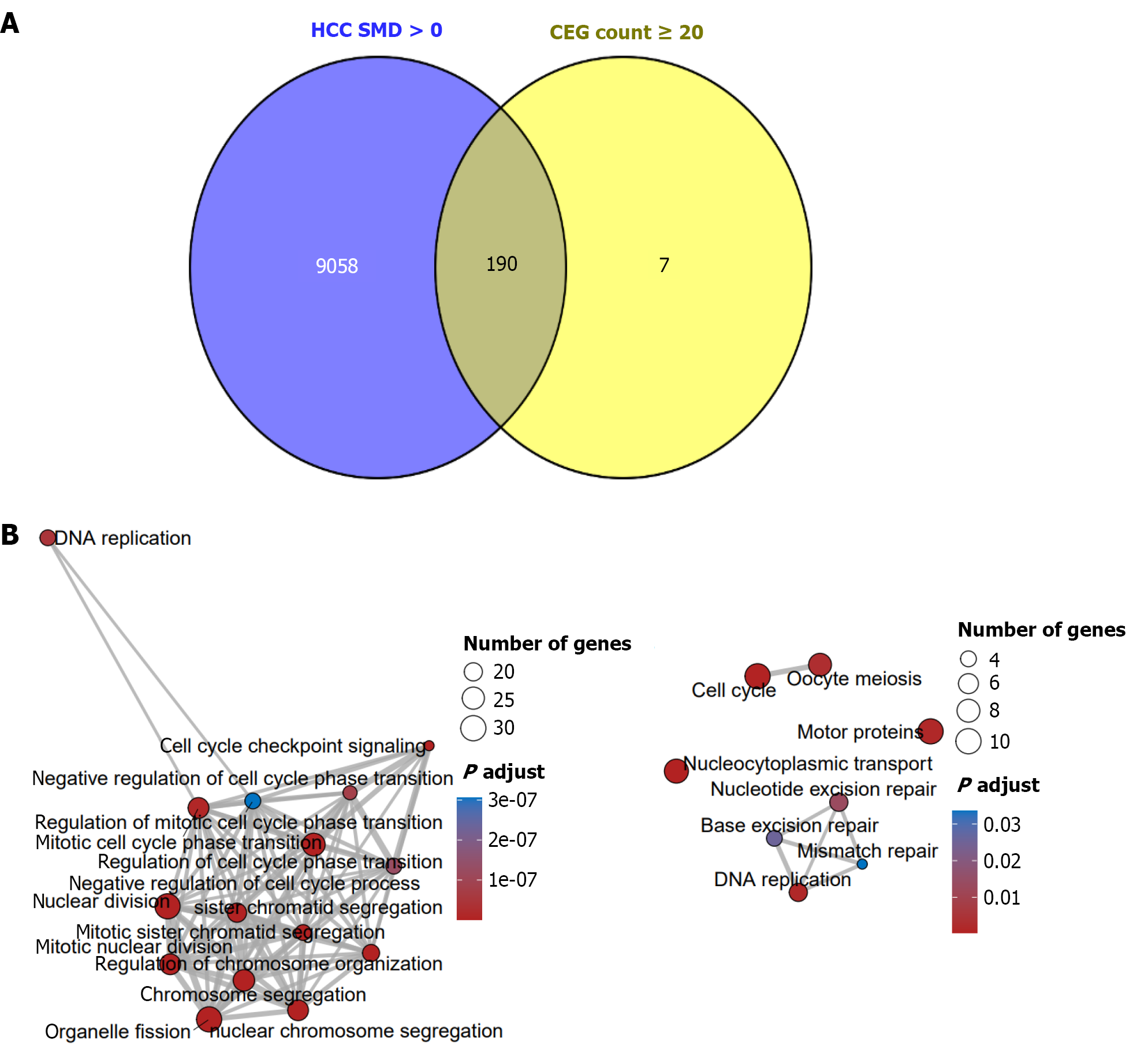
To further investigate the potential function of CUL2 in HCC, this study first confirmed ubiquitous expression of CUL2 across all 21 examined HCC cell lines. CRISPR-mediated knockout screens revealed that genetic ablation of CUL2 significantly suppressed cellular growth in 20 out of the 21 HCC cell lines (Figure 8) (CRISPR gene effect score < 0), strongly suggesting an essential pro-proliferative role of CUL2 in HCC. Building upon these findings, we subsequently dissected the underlying molecular mechanisms of CUL2 in HCC. Integrated analysis identified 190 genes positively correlated with high CUL2 expression in HCC (Figure 9A). Subsequent enrichment analysis demonstrated that these CUL2-coexpressed genes were predominantly associated with GO terms including nuclear division, chromosome segregation, and mitotic cell cycle phase transition, and were significantly enriched in key KEGG pathways such as nucleocytoplasmic transport and cell cycle regulation (Figure 9B).
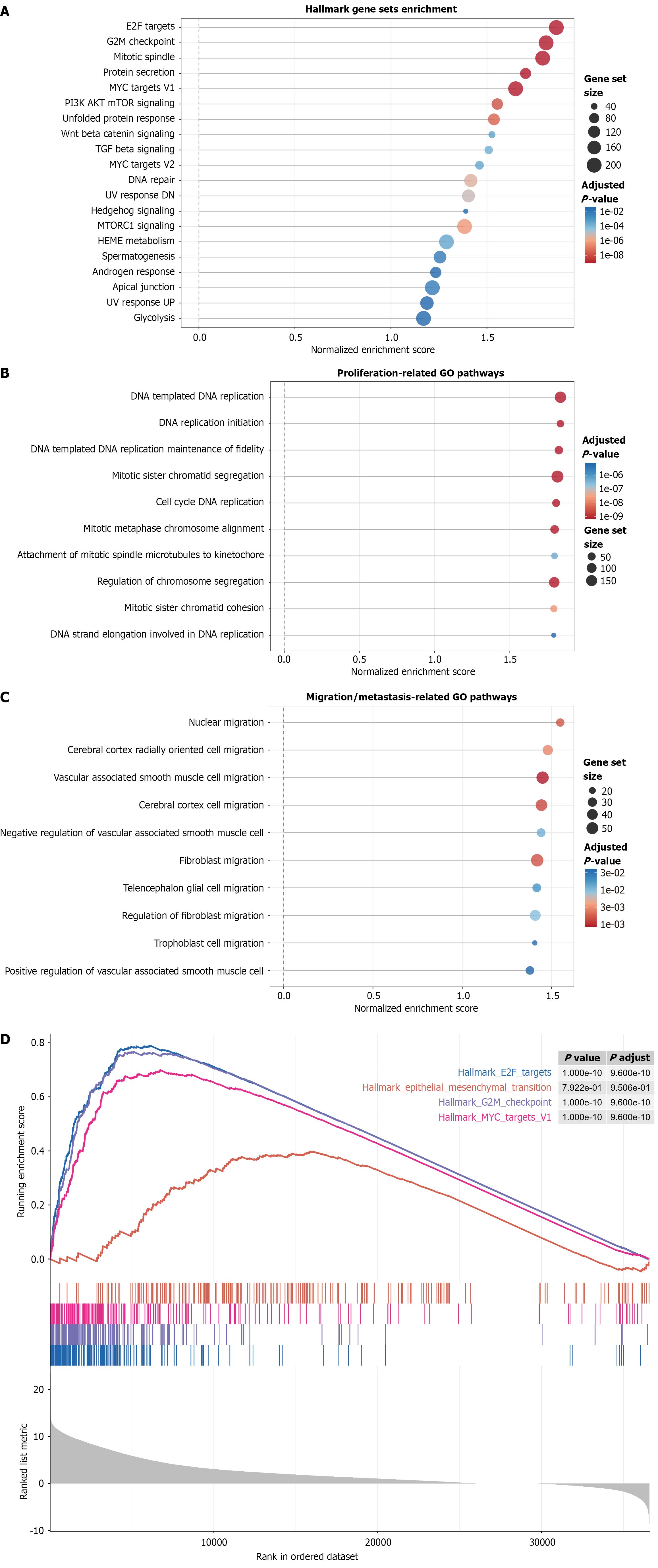
In order to systematically elucidate the molecular mechanism of CUL2 function in HCC, we performed a comprehensive GSEA. Hallmark pathway analysis showed that the characteristics of proliferation-related genes were significantly enriched in CUL2-overexpressing tumors (Figure 10A). The most significantly enriched pathways include E2F target [Normalized Enrichment Score (NES) = 2.18, P.adj < 0.001], G2/M checkpoint (NES = 2.05, P.adj < 0.001) and the v-myc avian MYC target V1 (NES = 1.85, P.adj < 0.001), which are classical driving pathways for cell cycle progression and cell proliferation. Other enrichment pathways cover mitotic spindle assembly, protein secretion, and mTOR signaling, all of which point to the central role of CUL2 in coordinating cell growth and division procedures. In order to further analyze the specific biological processes regulated by CUL2, we used the GO biological process gene set, focusing on proliferation and migration mechanisms for targeted GSEA analysis. Proliferation-related GO analysis showed that pathways regulating DNA replication, mitotic sister chromatid separation, and cell cycle phase transition were significantly enriched (Figure 10B), and the normalized enrichment scores of the top-ranked pathways exceeded 1.5 (all P.adj < 0.05). Among them, DNA-templated DNA replication-related pathways are most strongly associated with high CUL2 expression. GO pathway analysis related to migration and metastasis showed that nuclear migration, cerebral cortex cell migration, and vascular associated smooth muscle cell migration were significantly enriched in CUL2-overexpressing tumors (Figure 10C). The visualization of Hallmark pathway GSEA enrichment results further validated the significant enrichment trend of the three key proliferation-related pathways (E2F target, G2/M checkpoint, and v-myc avian MYC target V1) in CUL2-overexpressing tumors (Figure 10D).
This multi-center study comprehensively reveals that CUL2 is significantly overexpressed in HCC tissues, and CUL2 is closely related to the poor prognosis of patients. Functional experiments and bioinformatics studies further confirmed the important role of CUL2 in maintaining the growth of HCC cell lines, which may be achieved by regulating key signaling pathways involved in cell cycle control and nuclear transport. In summary, our study for the first time comprehensively elucidated the key role of CUL2 in HCC progression, highlighting its dual potential as a prognostic biomarker and a new therapeutic target for HCC.
Our results clearly confirmed the significant overexpression of CUL2 in HCC tissues. Immunohistochemical analysis showed that CUL2 protein was mainly localized in the cytoplasm, and the expression level of CUL2 protein in HCC specimens was significantly higher than that in matched adjacent non-tumor tissues. This observation is consistent with the previously recorded up-regulation trend of CUL2 in a variety of malignant tumors[14-17]. Specifically, our clinical specimen validation and database analysis confirmed the oncogenic characteristics of CUL2 in HCC. In addition, our findings are further supported by large-scale multi-queue data integration. By systematically integrating the global HCC microarray and high-throughput sequencing data sets, we verified that CUL2 mRNA was significantly up-regulated in 5204 HCC tissues, demonstrating a stable SMD estimate and no significant publication bias. These consistent findings across experimental platforms indicate that CUL2 overexpression is a common phenomenon and may be related to the pathogenesis and progression of HCC. In addition, CUL2 mRNA showed moderate discrimination in HCC detection (AUC = 0.78), indicating its potential as a biomarker for HCC treatment.
In addition, this study used single-cell and spatial transcriptome sequencing to achieve a detailed characterization of CUL2 mRNA expression patterns in HCC. Single-cell analysis showed that CUL2 mRNA was significantly overexpressed in HCC cell subsets compared with normal hepatocytes, confirming that CUL2 was up-regulated in single cell resolution and suggesting its potential functional significance in specific tumor subsets. Spatial transcriptomics further characterized the heterogeneous distribution of CUL2 mRNA in HCC tissue sections, showing a dispersed and non-diffuse expression pattern, which was different from the diffuse expression characteristics of the established HCC markers ARG1[18] and GPC3[19]. This spatial heterogeneity may reflect the dynamic expression of CUL2 in the tumor microenvironment or the functional diversity of different tumor regions[20,21], which provides a new perspective for understanding the mul
Finally, our study provides preliminary insights into the molecular mechanism of CUL2 in HCC through functional verification and bioinformatics exploration. CRISPR-mediated gene knockout screening showed that CUL2 gene ablation significantly inhibited cell growth in 20 of 21 HCC cell lines. This finding is consistent with previous reports that down-regulation of CUL2 inhibits the proliferation and migration of HCC cells[23], thereby consolidating its tumor-promoting function. These results provide an experimental basis for CUL2 as a potential therapeutic target for HCC. Subsequent bioinformatics analysis showed that 190 genes positively correlated with CUL2 overexpression were significantly en
Our GSEA revealed the mechanism by which CUL2 drives HCC progression. The results showed that E2F targets, G2/M checkpoints and MYC signaling pathways were significantly enriched in tumors with high CUL2 expression, suggesting a direct link between CUL2 overexpression and cell cycle out of control. CUL2 is the core component of the CRL2 ubiquitin ligase complex, which is known to regulate the stability of key cell cycle proteins (such as E2F). Therefore, the high expression of CUL2 may accelerate the entry of cells into the S phase by destroying the normal regulation of E2F[29], thereby promoting proliferation. At the same time, the enrichment of G2/M checkpoint genes indicates that abnormal expression of CUL2 may interfere with the degradation process of checkpoint proteins, weaken the ability of cells to monitor DNA damage, and enable damaged cells to enter mitosis[30]. In addition, the enrichment of MYC target genes suggests that CUL2 may affect the activity of MYC signaling pathway through an undefined mechanism[31]. The analysis also found that the nuclear transport pathway was significantly enriched in CUL2-overexpressing tumors. Nuclear-cytoplasmic transport disorder is one of the characteristics of cancer, which may affect the localization and function of many key proteins[32]. It is worth noting that pathways related to cell migration and metastasis are also enriched, such as nuclear migration and smooth muscle cell migration. This is consistent with previous findings that CUL2 circular RNA can promote epithelial-mesenchymal transition in liver cancer, suggesting that CUL2 may also be involved in regulating tumor invasiveness at the linear mRNA level.
Despite these meaningful findings, several limitations should be acknowledged. Although we have gained preliminary insights into CUL2’s function and potential mechanisms in HCC through comprehensive analysis of clinical specimens, public databases, and in vitro CRISPR screening, the specific molecular mechanisms require further validation through cell line models and in vivo animal studies. Future investigations should focus on: (1) Identifying specific CUL2 substrate proteins in HCC cells through co-immunoprecipitation and mass spectrometry; (2) Validating the functional role of CUL2 in HCC progression using genetic manipulation in cell lines and xenograft models; and (3) Conducting prospective clinical studies to validate CUL2 as a prognostic biomarker and predictive marker for targeted therapies.
This study confirmed that CUL2 is generally overexpressed in HCC and is an independent prognostic indicator. Based on the analysis of 9283 multi-omics samples, CUL2 overexpression had moderate diagnostic value (AUC = 0.78) and strong prognostic ability for HCC. CRISPR screening verified that it played a key role in promoting proliferation in 21 HCC cell lines. Mechanistically, CUL2 drives cell cycle out of control and proliferation by regulating signaling pathways such as E2F, G2/M checkpoint, and MYC. At the same time, it also affects nuclear transport and cell migration-related pathways, which may promote tumor invasion and metastasis. These functions are consistent with its molecular role as a core component of E3 ubiquitin ligase. In view of the key role of CUL2 in HCC, it is expected to become a new therapeutic target. In the future, it is necessary to further clarify the specific substrates of CUL2 and explore individualized treatment strategies based on its expression level.
We would like to express our heartfelt gratitude to all participants involved in this study. At the same time, we also appreciate the technical support provided by the Guangxi Zhuang Autonomous Region Clinical Medicine Research Center for Molecular Pathology and Intelligent Pathology Precision Diagnosis. We also appreciate the valuable data resources shared by public scientific databases and the professional guidance of experts in related fields in the research process.
| 1. | Mohamed NK, Hamad MA, Hafez MZ, Wooley KL, Elsabahy M. Nanomedicine in management of hepatocellular carcinoma: Challenges and opportunities. Int J Cancer. 2017;140:1475-1484. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 58] [Cited by in RCA: 52] [Article Influence: 5.8] [Reference Citation Analysis (1)] |
| 2. | Chan SL, Sun HC, Xu Y, Zeng H, El-Serag HB, Lee JM, Schwartz ME, Finn RS, Seong J, Wang XW, Paradis V, Abou-Alfa GK, Rimassa L, Kao JH, Zhang BH, Llovet JM, Bruix J, Yip TC, Wong VW, Wong GL, Chan LL, Liu MQ, Gao Q, Shen F, Kelley RK, Cheng AL, Kurosaki M, Toyoda H, Chen WX, Murakami T, Liang P, Zucman-Rossi J, Minami Y, Miyayama S, Wang K, Man K, Hasegawa K, Li Q, Tsuchiya K, Xu L, Chew V, Chow P, Hoshida Y, Lujambio A, Ng IO, Sakamoto M, Park YN, Yau T, Kudo M, Fan J, Zhou J. The Lancet Commission on addressing the global hepatocellular carcinoma burden: comprehensive strategies from prevention to treatment. Lancet. 2025;406:731-778. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 82] [Article Influence: 82.0] [Reference Citation Analysis (1)] |
| 3. | Ziogas IA, Tsoulfas G. Advances and challenges in laparoscopic surgery in the management of hepatocellular carcinoma. World J Gastrointest Surg. 2017;9:233-245. [PubMed] [DOI] [Full Text] |
| 4. | Petroski MD, Deshaies RJ. Function and regulation of cullin-RING ubiquitin ligases. Nat Rev Mol Cell Biol. 2005;6:9-20. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1859] [Cited by in RCA: 1755] [Article Influence: 83.6] [Reference Citation Analysis (1)] |
| 5. | Berndsen CE, Wolberger C. New insights into ubiquitin E3 ligase mechanism. Nat Struct Mol Biol. 2014;21:301-307. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 621] [Cited by in RCA: 567] [Article Influence: 47.3] [Reference Citation Analysis (1)] |
| 6. | Toma-Fukai S, Shimizu T. Structural Diversity of Ubiquitin E3 Ligase. Molecules. 2021;26:6682. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 140] [Cited by in RCA: 126] [Article Influence: 25.2] [Reference Citation Analysis (1)] |
| 7. | Megumi Y, Miyauchi Y, Sakurai H, Nobeyama H, Lorick K, Nakamura E, Chiba T, Tanaka K, Weissman AM, Kirisako T, Ogawa O, Iwai K. Multiple roles of Rbx1 in the VBC-Cul2 ubiquitin ligase complex. Genes Cells. 2005;10:679-691. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 18] [Article Influence: 0.9] [Reference Citation Analysis (1)] |
| 8. | Liu X, Zurlo G, Zhang Q. The Roles of Cullin-2 E3 Ubiquitin Ligase Complex in Cancer. Adv Exp Med Biol. 2020;1217:173-186. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 20] [Article Influence: 3.3] [Reference Citation Analysis (1)] |
| 9. | Chen P, Yao GD. The role of cullin proteins in gastric cancer. Tumour Biol. 2016;37:29-37. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 17] [Article Influence: 1.7] [Reference Citation Analysis (1)] |
| 10. | Xu J, Fang Y, Wang X, Wang F, Tian Q, Li Y, Xie X, Cheng X, Lu W. CUL2 overexpression driven by CUL2/E2F1/miR-424 regulatory loop promotes HPV16 E7 induced cervical carcinogenesis. Oncotarget. 2016;7:31520-31533. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 22] [Article Influence: 2.2] [Reference Citation Analysis (5)] |
| 11. | Liu A, Zhang S, Shen Y, Lei R, Wang Y. Association of mRNA expression levels of Cullin family members with prognosis in breast cancer: An online database analysis. Medicine (Baltimore). 2019;98:e16625. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 9] [Article Influence: 1.3] [Reference Citation Analysis (1)] |
| 12. | Liu X, Peng D, Cao Y, Zhu Y, Yin J, Zhang G, Peng X, Meng Y. Upregulated lncRNA DLX6-AS1 underpins hepatocellular carcinoma progression via the miR-513c/Cul4A/ANXA10 axis. Cancer Gene Ther. 2021;28:486-501. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 21] [Article Influence: 4.2] [Reference Citation Analysis (1)] |
| 13. | Zhao M, Quan Y, Zeng J, Lyu X, Wang H, Lei JH, Feng Y, Xu J, Chen Q, Sun H, Xu X, Lu L, Deng CX. Cullin3 deficiency shapes tumor microenvironment and promotes cholangiocarcinoma in liver-specific Smad4/Pten mutant mice. Int J Biol Sci. 2021;17:4176-4191. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 17] [Article Influence: 3.4] [Reference Citation Analysis (1)] |
| 14. | Costessi A, Mahrour N, Sharma V, Stunnenberg R, Stoel MA, Tijchon E, Conaway JW, Conaway RC, Stunnenberg HG. The human EKC/KEOPS complex is recruited to Cullin2 ubiquitin ligases by the human tumour antigen PRAME. PLoS One. 2012;7:e42822. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 40] [Cited by in RCA: 40] [Article Influence: 2.9] [Reference Citation Analysis (1)] |
| 15. | Sui XY, Ma XY, Hou Y, Cao SW, Wang ZQ, Jia LJ, Fan L, Shao ZM, Zhang WJ. Elongin B promotes breast cancer progression by ubiquitinating tumor suppressor p14/ARF. Cell Biol Toxicol. 2024;40:24. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 6] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 16. | Zhao W, Wen S, Wang X, Wang J, Zhang L, Wang T. Targeted regulation of miR-154-5p/Cullin2 pathway by hsa_circ_TRIM22 in promoting human papillomavirus 16 positive cervical cancer progression. J Cancer. 2024;15:2137-2146. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 4] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 17. | Zheng S, Wu Y, Li Z. Integrating cullin2-RING E3 ligase as a potential biomarker for glioblastoma multiforme prognosis and radiosensitivity profiling. Radiother Oncol. 2021;154:36-44. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 13] [Article Influence: 2.6] [Reference Citation Analysis (1)] |
| 18. | Radwan NA, Ahmed NS. The diagnostic value of arginase-1 immunostaining in differentiating hepatocellular carcinoma from metastatic carcinoma and cholangiocarcinoma as compared to HepPar-1. Diagn Pathol. 2012;7:149. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 96] [Cited by in RCA: 72] [Article Influence: 5.1] [Reference Citation Analysis (1)] |
| 19. | Chen M, Li G, Yan J, Lu X, Cui J, Ni Z, Cheng W, Qian G, Zhang J, Tu H. Reevaluation of glypican-3 as a serological marker for hepatocellular carcinoma. Clin Chim Acta. 2013;423:105-111. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 44] [Cited by in RCA: 44] [Article Influence: 3.4] [Reference Citation Analysis (1)] |
| 20. | Liu X, Zhang K, Kaya NA, Jia Z, Wu D, Chen T, Liu Z, Zhu S, Hillmer AM, Wuestefeld T, Liu J, Chan YS, Hu Z, Ma L, Jiang L, Zhai W. Tumor phylogeography reveals block-shaped spatial heterogeneity and the mode of evolution in Hepatocellular Carcinoma. Nat Commun. 2024;15:3169. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 15] [Article Influence: 7.5] [Reference Citation Analysis (1)] |
| 21. | Zheng S, Hu C, Lin H, Li G, Xia R, Zhang X, Su D, Li Z, Zhou Q, Chen R. circCUL2 induces an inflammatory CAF phenotype in pancreatic ductal adenocarcinoma via the activation of the MyD88-dependent NF-κB signaling pathway. J Exp Clin Cancer Res. 2022;41:71. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 51] [Cited by in RCA: 50] [Article Influence: 12.5] [Reference Citation Analysis (1)] |
| 22. | Jia L, Zhang X, Zhou T, Xie J, Jin J, Zhang D, Zhu C, Wan R. Comprehensive prognostic and immunological analysis of Cullin2 in pan-cancer and its identification in hepatocellular carcinoma. Aging (Albany NY). 2024;16:8898-8921. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 23. | Meng J, Chen S, Han JX, Qian B, Wang XR, Zhong WL, Qin Y, Zhang H, Gao WF, Lei YY, Yang W, Yang L, Zhang C, Liu HJ, Liu YR, Zhou HG, Sun T, Yang C. Twist1 Regulates Vimentin through Cul2 Circular RNA to Promote EMT in Hepatocellular Carcinoma. Cancer Res. 2018;78:4150-4162. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 260] [Cited by in RCA: 262] [Article Influence: 32.8] [Reference Citation Analysis (4)] |
| 24. | Jang SM, Redon CE, Thakur BL, Bahta MK, Aladjem MI. Regulation of cell cycle drivers by Cullin-RING ubiquitin ligases. Exp Mol Med. 2020;52:1637-1651. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 69] [Cited by in RCA: 62] [Article Influence: 10.3] [Reference Citation Analysis (1)] |
| 25. | Zhou L, Zhang W, Sun Y, Jia L. Protein neddylation and its alterations in human cancers for targeted therapy. Cell Signal. 2018;44:92-102. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 208] [Cited by in RCA: 202] [Article Influence: 25.3] [Reference Citation Analysis (1)] |
| 26. | Fouad S, Wells OS, Hill MA, D'Angiolella V. Cullin Ring Ubiquitin Ligases (CRLs) in Cancer: Responses to Ionizing Radiation (IR) Treatment. Front Physiol. 2019;10:1144. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 59] [Cited by in RCA: 46] [Article Influence: 6.6] [Reference Citation Analysis (1)] |
| 27. | Zhao Y, Sun Y. Cullin-RING Ligases as attractive anti-cancer targets. Curr Pharm Des. 2013;19:3215-3225. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 244] [Cited by in RCA: 240] [Article Influence: 18.5] [Reference Citation Analysis (1)] |
| 28. | Hill R, Cautain B, de Pedro N, Link W. Targeting nucleocytoplasmic transport in cancer therapy. Oncotarget. 2014;5:11-28. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 96] [Cited by in RCA: 92] [Article Influence: 7.7] [Reference Citation Analysis (1)] |
| 29. | Clijsters L, Hoencamp C, Calis JJA, Marzio A, Handgraaf SM, Cuitino MC, Rosenberg BR, Leone G, Pagano M. Cyclin F Controls Cell-Cycle Transcriptional Outputs by Directing the Degradation of the Three Activator E2Fs. Mol Cell. 2019;74:1264-1277.e7. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 107] [Cited by in RCA: 95] [Article Influence: 13.6] [Reference Citation Analysis (1)] |
| 30. | Bassermann F, Frescas D, Guardavaccaro D, Busino L, Peschiaroli A, Pagano M. The Cdc14B-Cdh1-Plk1 axis controls the G2 DNA-damage-response checkpoint. Cell. 2008;134:256-267. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 332] [Cited by in RCA: 319] [Article Influence: 17.7] [Reference Citation Analysis (1)] |
| 31. | Stine ZE, Walton ZE, Altman BJ, Hsieh AL, Dang CV. MYC, Metabolism, and Cancer. Cancer Discov. 2015;5:1024-1039. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1124] [Cited by in RCA: 1054] [Article Influence: 95.8] [Reference Citation Analysis (5)] |
| 32. | Zheng Y, Yu K, Lin JF, Liang Z, Zhang Q, Li J, Wu QN, He CY, Lin M, Zhao Q, Zuo ZX, Ju HQ, Xu RH, Liu ZX. Deep learning prioritizes cancer mutations that alter protein nucleocytoplasmic shuttling to drive tumorigenesis. Nat Commun. 2025;16:2511. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 3.0] [Reference Citation Analysis (1)] |