Published online Mar 27, 2026. doi: 10.4254/wjh.v18.i3.116233
Revised: November 23, 2025
Accepted: January 7, 2026
Published online: March 27, 2026
Processing time: 140 Days and 23.7 Hours
This minireview summarizes recent advances in artificial intelligence (AI) for imaging-based diagnosis of hepatocellular carcinoma (HCC), emphasizing how technical progress translates to clinical practice. Developments from convolutional neural networks to transformer-based, data-efficient, and spatiotemporal models have improved lesion detection, segmentation, classification, and treatment response assessment. These innovations underpin emerging clinical systems such as intelligent quantitative liver imaging analysis system and smart liver imaging analysis system, demonstrating growing translational impact. Yet persistent challenges-including cross-site generalizability, interpretable decision support, seamless workflow integration, and regulatory oversight-continue to shape re
Core Tip: Artificial intelligence (AI) for image-guided hepatocellular carcinoma diagnosis improves sensitivity to small lesions, enables standardized quantitative readouts, and reduces reporting time, thereby complementing conventional radiology. Built on diverse imaging datasets and evolving model designs, AI has advanced across detection, segmentation, characterization, and response assessment, with early clinical uptake. Outstanding obstacles-cross-institution generalization and clinically meaningful explainability-must be resolved to achieve scalable, trustworthy deployment.
- Citation: Zhou ZX, Xiao JJ, Ning ZX. Advances in artificial intelligence for imaging-based diagnosis of hepatocellular carcinoma. World J Hepatol 2026; 18(3): 116233
- URL: https://www.wjgnet.com/1948-5182/full/v18/i3/116233.htm
- DOI: https://dx.doi.org/10.4254/wjh.v18.i3.116233
Hepatocellular carcinoma (HCC) is one of the most common and deadliest primary liver malignancies worldwide. Its incidence and mortality rates remain alarmingly high, posing a significant challenge to global public health[1]. For example, in 2020, HCC ranked as the sixth most common cancer worldwide and the third leading cause of cancer-related deaths. Furthermore, with the global spread of metabolic dysfunction-associated steatotic liver disease, the disease burden of HCC is expected to worsen[2]. Given this, the development of more efficient and precise diagnostic and therapeutic strategies has become crucial[3].
Traditional HCC diagnosis mainly relies on imaging techniques, including ultrasound (US), computed tomography (CT), magnetic resonance imaging (MRI), and positron emission tomography (PET). These imaging methods play a foundational role in HCC screening, diagnosis, treatment planning, and long-term monitoring. However, existing imaging technologies face many limitations in clinical practice. For example, small or atypical lesions are prone to misdiagnosis; there are discrepancies in image interpretation between different readers and medical institutions, leading to inconsistent diagnoses; and the growing volume of imaging exams imposes significant pressure on clinical workflows, potentially affecting diagnostic efficiency and quality[4]. In particular, in the routine interpretation of liver CT scans, an 8-mm early HCC lesion may pass through a radiologist's field of view in just 0.3 seconds, creating a physiological blind spot that greatly increases the risk of missing early intervention opportunities.
To overcome these inherent challenges of traditional imaging diagnosis, artificial intelligence (AI) technologies, es
In the imaging diagnosis of HCC, AI intervention is considered to effectively address many challenges faced by tra
Imaging technologies are crucial in the diagnosis of HCC, and different types of imaging data have unique characteristics and advantages. According to different imaging devices, common types of imaging data include CT, MRI, and US. The performance of each modality depends on its imaging principle, resolution, contrast, and acquisition speed (Table 1).
| Modality | Advantages | Limitations |
| CT | (1) Rapid, whole-liver coverage; (2) High spatial resolution for vascular and calcified structures; and (3) Widely available with standardized interpretation | (1) Ionizing radiation and contrast-related risks; (2) Limited soft-tissue contrast; (3) Lower sensitivity for sub-centimeter lesions; and (4) Susceptible to respiratory artifacts |
| MRI | (1) Excellent soft-tissue contrast and multi-parametric capability; (2) Specific contrast agents improve HCC detection accuracy; (3) Functional imaging (e.g., DWI, elastography); and (4) No ionizing radiation | (1) Long acquisition time and high cost; (2) Contraindicated for patients with metallic implants; (3) Limited availability; and (4) Less effective in showing calcifications |
| US | (1) Real-time and dynamic imaging of hepatic blood flow; (2) Useful for biopsy guidance and follow-up; (3) Contrast-enhanced US improves detection of small lesions; and (4) Low cost and radiation-free | (1) Highly operator-dependent; (2) Reduced accuracy in obese patients or with bowel gas; (3) Low sensitivity for sub-centimeter HCC; and (4) Limited for definitive qualitative diagnosis |
Overall, the three imaging techniques of CT, MRI, and US each have their own advantages and limitations. CT can quickly complete a full liver scan, and it is superior to MRI in displaying calcifications and fatty infiltration, with clear vascular imaging and standardized interpretation. However, it has risks of ionizing radiation and contrast agents, limited soft tissue resolution, a lower detection rate for tiny lesions smaller than 1 cm compared to MRI, and may be affected by respiratory artifacts[15,16].
MRI has significant advantages in multi-parameter imaging. Combined with specific contrast agents, it can improve the accuracy of HCC diagnosis. Diffusion-weighted imaging can evaluate small lesions, elastography can quantify liver fibrosis, and it has no radiation. But it has a long examination time, is contraindicated for patients with metal implants, has high costs and low popularity, and is not as good as CT in displaying calcifications[15,17].
US can perform real-time imaging and dynamically evaluate intrahepatic blood flow, which is irreplaceable in guiding liver biopsy. Contrast-enhanced US can improve the detection rate of small lesions, with low cost and no radiation, making it suitable for primary care and follow-up. However, it is highly dependent on the operator's experience, obesity or intestinal gas can easily lead to missed diagnoses, and its sensitivity for sub-centimeter HCC remains relatively low, requiring complementary imaging for definitive diagnosis[15,16].
In practice, no single technique is sufficient for comprehensive HCC evaluation; instead, complementary use of CT, MRI, and US is often required to balance sensitivity, specificity, and practical constraints. AI technologies further enhance this multimodal strategy by integrating heterogeneous imaging data and mitigating the limitations inherent to individual modalities, supporting more accurate and efficient liver disease diagnosis.
With the wide application of AI technology in imaging-assisted diagnosis of HCC, the deep learning algorithm system continues to evolve, especially showing trends of diversification and integration in terms of model architecture, which has significantly promoted the improvement of performance in HCC image recognition and analysis. Initially, traditional CNNs served as the basic algorithm and played an important role in liver lesion recognition, tumor classification, and localization. Through local receptive fields, they achieve translation-invariant feature extraction, suitable for automatic diagnosis of two-dimensional (2D) images. Kopp et al[18] evaluated the performance of a CNN-based anthropomorphic model observer in liver lesion detection tasks through phantom studies. The results showed that the CNN model observer could accurately predict the performance of human observers under all lesion sizes and dose levels, supporting the important role of CNN in liver lesion recognition. Vanmore and Chougule[19] proposed a liver lesion classification system for CT images, using a CNN deep learning model to improve classification accuracy. Through a sequential CNN architecture model including input convolutional layers, hidden convolutional layers, and output convolutional layers, they classified liver lesions in CT images of different sizes and achieved high accuracy, demonstrating the application value of CNN in liver lesion classification. Said et al[20] evaluated the performance and capability of a CNN-based pipeline for semi-automatic segmentation of HCC tumors on MRI. The experimental results showed that the CNN model had moderate to good performance in semi-automatic HCC segmentation, providing technical support for more rapid and accurate clinical evaluation of HCC tumors, reflecting the role of CNN in tumor localization and segmentation.
However, CNNs have difficulty modeling complex dependencies in both spatial and temporal dimensions when processing dynamic multi-phase data[21]. 2D convolution cannot fully utilize 3D spatial information, while 3D convolution has problems of high computational cost and large GPU memory consumption. Li et al[22] proposed a novel hybrid densely connected UNet, consisting of a 2D DenseUNet and a 3D DenseUNet. The former can extract intra-slice features, while the latter can hierarchically aggregate volumetric context. Through a hybrid feature fusion layer, it can jointly optimize intra-slice representation and inter-slice features. The research results showed that this method outperformed other technologies in tumor segmentation on the medical image computing and computer-assisted intervention 2017 liver tumor segmentation challenge dataset and the 3D liver and tumor dataset for computer-aided diagnosis dataset, and also had excellent performance in liver segmentation. Therefore, researchers further proposed 2D/3D hybrid network architectures, using 2D convolution to extract local spatial features and 3D convolution to model cross-slice structural and temporal consistency, significantly improving the ability to model complex 3D liver structures and lesions.
On this basis, to address CNNs’ limited ability to capture global semantic relationships-n important challenge in detecting small or low-contrast HCC lesions-ransformer models were introduced into HCC image analysis. Rather than emphasizing their technical mechanisms, it is more relevant clinically that Transformers help the model “see the bigger picture”, allowing better differentiation between tumors and surrounding tissues in cases where contrast is subtle. When combined with CNNs, Transformers provide global contextual awareness while CNNs preserve fine structural details, leading to more accurate boundary delineation and improved detection of tiny lesions.
In addition, multi-phase spatiotemporal modeling was developed to better utilize the diagnostic value of arterial, portal venous, and delayed phases-each capturing different enhancement characteristics of HCC. Instead of focusing on the underlying mathematical operations, these models help align multi-phase images and extract temporal changes that radiologists rely on for staging and characterization. This approach improves the recognition of the dynamic evolution of lesions, which is critical for distinguishing early HCC from benign nodules and for refining treatment planning.
Meanwhile, few-shot learning has emerged to address the common clinical challenge of limited annotated medical images. Instead of describing each algorithmic variant, the key idea is that few-shot techniques allow models to learn effectively from very small datasets, making them valuable for rare HCC subtypes or centers with limited imaging resources. By leveraging data augmentation, pre-trained models, and cross-modal feature alignment, few-shot learning improves model robustness and enhances diagnostic performance even under low-resource conditions.
In summary, from traditional CNNs to 2D/3D hybrid networks to transformer fusion, multi-phase spatiotemporal modeling, and few-shot learning, the core algorithm architectures for HCC image analysis are gradually becoming more intelligent, refined, and efficient, providing solid technical support for early diagnosis and individualized treatment (Table 2).
| Model architecture type | Core principles/technical features | Core advantages | Main limitations | Clinical application scenarios (HCC imaging analysis) |
| Traditional CNNs | Extract features through local receptive fields to achieve translation invariance; available in 2D/3D forms, relying on stacked sequential convolutional layers | (1) 2D CNNs offer high computational efficiency and adapt to conventional 2D images; (2) 3D CNNs can capture partial spatial structures; and (3) Mature technology with strong generalization | (1) 2D CNNs fail to fully utilize 3D spatial information; (2) 3D CNNs have high computational cost and large GPU memory consumption; and (3) Difficulty in modeling complex spatiotemporal dependencies and global semantic associations | Liver lesion recognition, tumor classification/localization, semi-automatic segmentation of CT/MRI images |
| Transformer models | Capture global semantic associations based on self-attention mechanism, emphasizing “global contextual awareness”, often used in fusion with CNNs | (1) Break through the limitation of local features to accurately capture global associations; (2) Clearly distinguish tumors from surrounding tissues under weak contrast; and (3) Complement CNNs to balance global context and detailed features | (1) Prone to losing fine structural details when used alone; and (2) High demand for annotated data volume | Detection of small/low-contrast HCC lesions, accurate delineation of tumor boundaries, global imaging feature association analysis |
| 2D/3D hybrid networks (e.g., H-DenseUNet) | Integrate 2D CNNs (extract intra-slice features) and 3D CNNs (aggregate volumetric context), optimizing performance through hybrid feature fusion layers | (1) Balance spatial information utilization and computational efficiency; (2) Enhance the ability to model cross-slice structural consistency and temporal consistency; and (3) Superior segmentation performance | Relatively complex technical architecture, requiring targeted optimization of feature fusion logic | Accurate segmentation of liver tumors, cross-slice feature association analysis of multi-phase images |
| Multi-phase spatiotemporal modeling | Align arterial phase/portal venous phase/delayed phase images and extract temporal change features of dynamic lesion enhancement | (1) Fully utilize the diagnostic value of multi-phase images; (2) Facilitate tumor staging and characterization; and (3) Distinguish early HCC from benign nodules | Rely on complete multi-phase imaging data with high requirements for data quality | Recognition of dynamic lesion evolution, differentiation between early HCC and benign nodules, treatment plan optimization |
| Few-shot learning | Achieve efficient learning from extremely small datasets based on data augmentation, pre-trained models, and cross-modal feature alignment | (1) Adapt to scenarios with limited annotated data; (2) Improve model robustness under low-resource conditions; and (3) Suitable for analyzing rare HCC subtypes | Performance still needs improvement in complex scenarios (e.g., multi-center heterogeneous data) | Diagnosis of rare HCC subtypes, lesion detection/classification in centers with limited imaging resources |
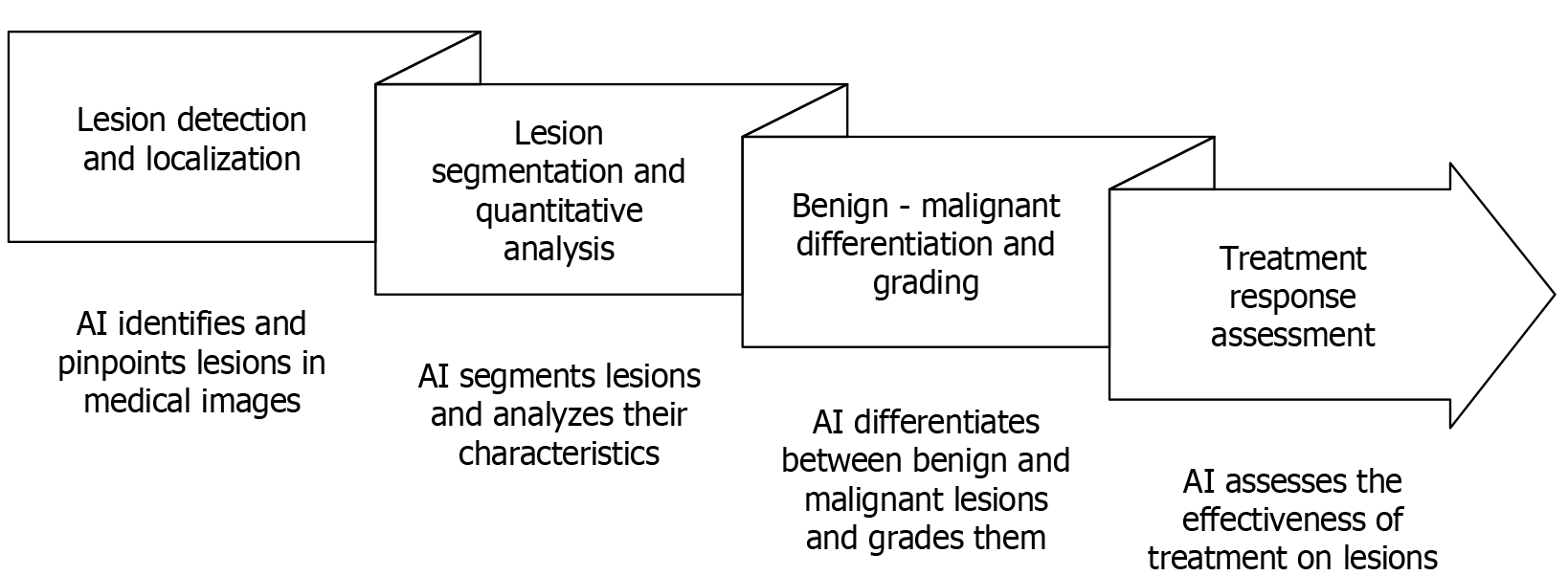
AI technology, through deep learning, enables the automation and intelligent analysis of imaging data, thereby reshaping the entire process of liver cancer diagnosis and treatment. This chapter will focus on four core application scenarios: Efficient and precise detection and localization of lesions, breakthroughs in lesion segmentation and quantitative analysis, clinical practical value in differentiating and grading benign and malignant conditions, and the intelligent advancements in treatment response assessment. It will systematically analyze the transformative role of AI in optimizing diagnostic and treatment processes, enhancing decision-making accuracy, and reducing human errors (Figure 1).
Early diagnosis and precise treatment of HCC are inseparable from effective detection and accurate localization of liver lesions. Traditional medical imaging methods, such as CT and MRI, can provide some help in HCC diagnosis but have limitations such as dependence on doctor experience, diagnostic inconsistency, and insufficient sensitivity to early tiny lesions[23]. Roberts et al[24] conducted a systematic review and meta-analysis of relevant literature, showing that in cirrhotic patients, the sensitivity of MRI in detecting HCC was 82%, and that of CT was 66%. For lesions smaller than 1 cm, the detection effect of both modalities was poor. Traditional methods are difficult to distinguish HCC from cirrhotic nodules, easily leading to missed diagnosis of early tumors and affecting early clinical diagnoses. However, with the rapid development of AI technology, especially deep learning models, the accuracy of HCC image analysis has been significantly improved. By training on a large amount of medical image data, AI models can automatically extract complex features related to HCC and perform efficient lesion detection and localization. Li et al[25] developed a deep learning model, DISMIR, based on circulating cell-free DNA methylation and sequence information for non-invasive detection of HCC. In the test cohort, the subject-level sensitivity of DISMIR was 93.94% and specificity was 100%, with the AI model showing significantly better sensitivity than traditional methods in this field. Furthermore, with technological progress, single-modal image analysis can no longer meet the needs of complex HCC diagnosis, so multi-modal image fusion has become an emerging research direction. Parsai et al[26] retrospectively fused fluorodeoxyglucose PET/CT and MRI data to analyze 150 liver lesions, finding that the fused PET/MRI significantly outperformed single modalities (PET/CT or MRI) in detection sensitivity (91.9%), specificity (97.4%), and accuracy (94.7%). For tiny lesions with a diameter of ≤ 0.8 cm, the structural resolution of MRI compensated for the deficiency of PET metabolic signals, while the metabolic information of PET helped exclude benign hyperplastic nodules that are easily misdiagnosed by MRI. Although the study did not directly use AI, it provided a clinical basis for subsequent AI models to integrate multi-modal data. By combining PET and MRI image data, AI can make up for the limitations of a single modality and provide more comprehensive tumor detection information. PET provides metabolic information of tumors, especially with high sensitivity in the early stage, while MRI can provide high-resolution structural images, facilitating precise localization of lesions.
Clinical impact and decision-making: AI-assisted detection can increase the likelihood that subcentimeter lesions are flagged for confirmatory MRI or short-interval follow-up, supporting earlier eligibility for curative therapies (e.g., ablation or resection). In multidisciplinary settings, automated heatmaps and triage lists help prioritize reading order, potentially shortening time to diagnosis and reducing interreader variability, which is critical in surveillance programs for cirrhosis.
In imaging-assisted diagnosis of HCC, lesion segmentation and quantitative analysis are key links in achieving early detection, treatment planning, and efficacy evaluation. Firstly, traditional image processing methods such as edge detection and threshold segmentation can complete basic segmentation tasks in simple cases, but their accuracy and robustness are seriously insufficient in complex situations such as blurred lesion boundaries, irregular shapes, and low contrast with surrounding tissues, making it difficult to meet clinical needs for precise lesion localization and quantitative evaluation. Especially in patients with high tumor heterogeneity, traditional methods cannot provide stable and consistent segmentation results. Sun et al[27] pointed out that traditional threshold segmentation and region-growing methods have significant defects in HCC segmentation: The dice coefficient for tumors with blurred boundaries (such as infiltratively growing HCC) is only 0.65-0.72, and the missed diagnosis rate for low-contrast lesions (such as isodense metastases) exceeds 30%. The deep learning model RHEU-Net improved the Dice coefficient of tumor segmentation to 0.7019 through multi-scale feature fusion and hybrid attention mechanisms, and its robustness was significantly better than traditional methods when dealing with heterogeneous tumors. The study emphasized that traditional methods lack the ability to model contextual information and cannot effectively distinguish tumors from surrounding fibrotic or necrotic tissues. To solve these problems, in recent years, AI technology, especially deep learning models, has shown breakthrough advantages in lesion segmentation. Huang et al[28] proposed AdwU-Net based on differentiable neural architecture search, which realizes adaptive optimization for different medical imaging tasks by automatically adjusting the depth (number of convolutional layers) and width (number of channels) of U-Net. In multi-organ segmentation datasets (such as liver and pancreas), the Dice coefficient of AdwU-Net is 3%-5% higher than that of traditional U-Net, especially in processing tumors with blurred boundaries, it significantly reduces over-segmentation and under-segmentation problems by dynamically adjusting the receptive field. By sharing network weights of different depth/width configurations, the model reduces computational costs while ensuring accuracy, making it suitable for clinical real-time segmentation needs. Gao et al[29] proposed a nested U-Net structure for liver tumor segmentation, realizing dynamic aggregation of multi-scale features through dilated dense short skip connections and adaptive atrous spatial pyramid pooling (Adaptive ASPP). On CT and US datasets, ASU-Net++ achieved a Dice coefficient of 0.9153-0.9413 for segmentation of tumors with complex edges, and the segmentation error for tiny lesions with a diameter of ≤ 2 cm was reduced by 40%. The Adaptive ASPP module can automatically adjust the receptive field according to tumor mor
Clinical impact and decision-making: Volumetric and peritumoral quantitative metrics derived from AI segmentation can inform transplant listing criteria (e.g., tumor burden), radiotherapy planning margins, and selection between ablation vs transarterial therapies. Automated portal vein tumor thrombus mapping supports timely anticoagulation or modification of locoregional strategies, while reproducible measurements improve response adjudication in tumor boards and can reduce time spent on manual contouring.
Early diagnosis of HCC is crucial for patient prognosis, but traditional imaging methods such as CT, MRI, and US, although important in HCC diagnosis, are often affected by the heterogeneity of imaging manifestations and differences in doctor experience, leading to great challenges in distinguishing benign and malignant lesions. With the development of AI technology, automated benign-malignant differentiation and grading based on imaging data have become important research directions in imaging-assisted diagnosis of HCC. By improving the processing and analysis accuracy of imaging data, AI has significantly improved the recognition rate of HCC and its metastatic lesions and provided strong support for precise grading. Chen et al[32] used the Inception V3 model for fully automatic grading of H&E-stained sections, achieving an accuracy of 96.0% in benign-malignant classification and 89.6% in high-medium-low differentiation prediction. Meanwhile, the model can predict gene mutations such as CTNNB1 and TP53 through histomorphological features [area under the curve (AUC) 0.71-0.89], linking imaging phenotypes with molecular characteristics, providing a new paradigm for precise pathological diagnosis. In the differentiation of benign and malignant HCC, traditional imaging (such as CT and MRI) relies on enhancement patterns to distinguish liver lesions, but the manifestations of HCC and metastatic liver cancer in arterial phase hyperenhancement and portal phase washout overlap. For example, the washout of HCC may be delayed to more than 2 minutes, while metastatic cancer may show rapid and complete was
Clinical impact and decision-making: Benign-malignant differentiation tools can reduce unnecessary biopsies and help determine candidacy for curative approaches (e.g., resection vs ablation) when findings are indeterminate. Risk scores for MVI and grade support surgical margin planning, transplant prioritization, and adjuvant therapy selection by indicating recurrence risk. Linking imaging phenotypes to likely molecular alterations can also guide trial enrollment for targeted and immunotherapies.
Assessment of treatment response in HCC is of great significance for treatment decisions and patient prognosis, especially in the context of the increasing application of immunotherapy and targeted therapy. Traditional imaging methods have gradually exposed their limitations, especially in evaluating dynamic changes in tumor necrotic areas, active tumor areas, and immunotherapy-related responses. The introduction of AI technology has provided new solutions for evaluating treatment response in HCC. Xia et al[35] proposed a deep learning model based on multi-phase CT, integrating imaging features of the arterial phase, portal phase, and delayed phase to predict the response to lenvatinib combined with immunotherapy. By quantifying tumor vascular density (such as iodine uptake rate) and microenvironmental fibrosis degree, the model achieved an AUC of 0.802 in 120 patients, with a prediction accuracy of 78.3% for treatment response. The study found that portal phase images contributed the most to model performance (AUC 0.760), and AI-generated heatmaps could accurately locate active tumor regions to assist clinical the decision-making. Through AI, automated analysis of tumor images is realized, which can more accurately extract imaging features of tumors, thereby quantifying tumor necrotic and active areas and accurately predicting treatment response based on changes between baseline and early images, providing an objective and personalized decision-making basis for clinical treatment. Zheng et al[36] proposed a 4D deep learning model based on 3D convolution and convolutional long short-term memory, realizing precise segmentation of HCC lesions by integrating temporal information of dynamically enhanced MRI (arterial phase, portal phase, delayed phase). The dice coefficient of the model on multi-center datasets reached 0.825, and the accuracy of identifying active tumor regions (91.3%) was significantly higher than that of traditional manual segmentation (78.6%). The study confirmed that AI models can effectively distinguish necrotic areas from active tumors by capturing dynamic changes in tumor enhancement patterns (such as "fast-in fast-out" vs "persistent enhancement"), providing a quantitative basis for treatment response assessment. Traditionally, the evaluation of tumor regions relies on manual delineation by doctors, which is easily affected by image quality and doctor experience. However, AI technology realizes automatic identification, segmentation, and quantitative analysis of HCC tumor regions through deep learning algorithms, greatly improving the objectivity and accuracy of evaluation. In addition, AI can dynamically monitor based on baseline and early imaging data, timely capture possible pseudoprogression of tumors during immunotherapy, avoid misjudgment of treatment effects, and provide real-time feedback for adjusting treatment plans. Li et al[37] developed a CT-based radiomics model, which accurately predicted pseudoprogression (PP) and hyperprogression in 105 non-small cell lung cancer patients by extracting texture features of intra-tumor and peritumoral regions (such as gray-level co-occurrence matrix entropy and fractal dimension). The AUC of the model in the training set and validation set was 0.95 and 0.88, respectively, significantly better than the traditional response evaluation criteria in solid tumors criteria. The study found that tumor heterogeneity in baseline CT (such as gray-level co-occurrence matrix GLCM contrast) was a key feature for predicting PP, and AI-quantified immune-related adverse events (such as hepatitis) were significantly correlated with PP risk (HR = 2.13, P < 0.01). This enables AI not only to accurately predict immunotherapy response but also to effectively distinguish early tumor response from pseudoprogression, ensuring the continuity and effectiveness of treatment. By combining multi-dimensional features and dynamic changes of tumor images, AI provides more refined solutions for predicting immunotherapy response, improving the accuracy of clinical treatment response. In summary, the application of AI technology effectively makes up for the shortcomings of tra
Clinical impact and decision-making: Response models that quantify viable tumor burden can trigger earlier treatment adaptation (e.g., switch of systemic agent, add-on transarterial chemoembolization, or repeat ablation) when conventional size metrics lag behind biological change. Early identification of pseudoprogression helps avoid premature discontinuation of immunotherapy, and standardized AI-derived biomarkers improve consistency across centers in clinical trials and real-world registries.
With the continuous development of AI technology, more and more AI tools have obtained clinical certification and begun to be widely used in imaging-assisted diagnosis of HCC. For example, the intelligent quantitative liver imaging analysis system-liver (IQQA-Liver) system is the first HCC imaging AI tool to obtain Conformité Européenne (CE) certification, realizing 3D reconstruction of the liver, tumors, and blood vessels based on 4D image processing technology. In 200 HCC patients, the correlation between AI-segmented tumor volume and pathological measurement values was extremely strong (intraclass correlation coefficient = 0.96), and the accuracy of identifying vascular invasion (87.6%) was significantly higher than that of traditional 2D evaluation (68.2%). The system has been used for preoperative planning in more than 30 hospitals in Europe, shortening the surgical plan design time from 2 hours to 15 minutes and reducing intraoperative blood loss by 23%[38]. On the other hand, the smart liver segmentation algorithm (SALSA) tool developed by Vall d'Hebron Institute of Oncology realizes fully automatic detection and segmentation of HCC in CT images through deep learning. In 607 patients from 8 centers in 5 countries, the detection accuracy of AI for HCC and liver metastases was 99.2% and 97.8%, respectively, with a dice coefficient of 0.812, significantly better than the average level of radiologists (0.724). The tool has passed the European Union Medical Device Regulation certification and is used in hospitals in Spain, Germany, etc., to assist doctors in quickly evaluating tumor burden (such as total tumor volume and vascular density), providing a quantitative basis for targeted therapy and liver transplantation decisions[39].
However, although AI tools show high accuracy in single-center trials, in multi-center clinical applications, the sensitivity of AI may be affected by different equipment and scanning protocols used in different medical institutions. Gresser et al[40] found that in 142 prostate cancer patients from 14 institutions using 7 types of MRI equipment (1.5T/3T), the AUC of radiomics-based machine learning models for classifying clinically significant prostate cancer was 0.78-0.83, which was higher than that of the traditional prostate imaging reporting and data system score (AUC 0.78), but there were significant inter-institutional differences in model performance. The study found that differences in gradient echo sequence parameters (such as echo time) and reconstruction algorithms (such as iterative reconstruction vs filtered back projection) of different scanners led to quantitative deviations in texture features (such as GLCM contrast), thereby affecting model generalization ability. Despite the use of feature selection (minimum redundancy maximum relevance) and class weight adjustment (synthetic minority over-sampling technique), the heterogeneity of multi-center data still caused a 12% fluctuation in model sensitivity (from 82% to 70%). Therefore, to improve the adaptability and stability of AI tools in multi-center settings, it is necessary to standardize different scanning protocols or optimize algorithms.
Beyond technical variability, multi-center generalization remains a key barrier to clinical deployment. Even within certified systems, model performance may degrade when applied to underrepresented populations, differing disease spectra, or scanners not included in the original training dataset. Site-specific biases-such as differences in patient demographics, contrast protocols, and liver disease prevalence-can lead to systematic over- or underestimation of HCC risk, raising concerns about fairness and reproducibility. Prospective, multi-institutional validation studies with harmonized imaging pipelines and shared reference datasets are urgently needed to ensure consistent performance across clinical environments.
In addition to performance variability, regulatory and ethical challenges significantly influence real-world adoption. Current AI approval pathways (e.g., CE, Food and Drug Administration, Medical Device Regulation) primarily assess safety and efficacy at one point in time, but AI models continue to evolve with retraining, creating a “moving target” for regulation. This raises questions about post-deployment monitoring, algorithm update approval, and responsibility in case of diagnostic errors. Moreover, algorithmic bias-arising from imbalanced training data-can disproportionately affect minority groups or patients with rare HCC subtypes. Data privacy is another concern, as multi-center collaboration often requires cross-border data sharing, which must comply with General Data Protection Regulation (GDPR) and Health Insurance Portability and Accountability Act (HIPAA) regulations. De-identification alone may not guarantee anonymity when image metadata or genetic signatures are available. Transparency and interpretability also remain limited in many commercial AI systems; clinicians often receive a predicted probability or color map without understanding how the model reached its conclusion. This “black box” nature can hinder physician trust and accountability. Addressing these issues requires the integration of explainable AI techniques, continuous auditing of performance drift, and establishing clear governance frameworks that define data ownership, consent management, and ethical oversight.
In summary, while clinically certified AI tools such as IQQA-Liver and SALSA demonstrate strong potential in preoperative planning and tumor quantification, their broader translation depends on addressing multi-center heterogeneity, establishing transparent regulatory pathways, and building ethical safeguards for trustworthy AI use in HCC imaging. Future progress will rely on standardized imaging protocols, federated learning frameworks that preserve privacy, and regulatory mechanisms capable of adapting to continuously learning systems.
Although AI shows great potential in imaging-assisted diagnosis of HCC, its clinical application and large-scale promotion still face multiple challenges. Firstly, technical bottlenecks: AI tools perform well in a single center, but in a multi-center environment, due to differences in image acquisition equipment, scanning protocols, and patient groups, their generalization problem is particularly prominent, and performance is prone to decline. To this end, domain adaptation, which adjusts feature extraction to adapt to different data domains, and federated learning, which uses multi-center data for collaborative training under the premise of protecting privacy, have become key strategies to improve cross-center generalization ability.
In practice, federated learning can be implemented through hospital-level “federation nodes”, where each center trains models locally on its data and only shares encrypted parameters with a central server. This allows continuous improvement of AI algorithms while complying with privacy laws such as GDPR and HIPAA. Similarly, domain adaptation models can be embedded into the deployment workflow so that algorithms automatically adjust to variations in scanners or contrast protocols without requiring manual retraining.
In addition, medical applications not only require high accuracy but also extremely rely on the interpretability of models to establish clinical trust. The “black box” nature of AI hinders doctors’ understanding of its decision-making process. Fortunately, technologies such as heatmap guidance and interpretability frameworks (such as gradient-weighted class activation mapping and SHapley Additive exPlanations) are promoting the establishment of a "doctor-AI collaboration" diagnostic model by visualizing decision-making basis and providing evidence support, enhancing transparency and acceptance. Furthermore, facing the complex and changing clinical environment, AI algorithms need to improve adaptability. This can be achieved through reinforcement learning, which simulates doctor decisions for continuous optimization, and multi-modal learning, which integrates multi-source data such as images, serum markers, and clinical information to reduce misdiagnosis risks and improve comprehensive diagnostic capabilities. In real-world deployment, reinforcement learning can be applied to treatment recommendation systems that learn from feedback on therapeutic outcomes-such as imaging-based tumor response-to continuously refine decision strategies. Multi-modal AI can be realized through data integration platforms that merge radiology, pathology, and laboratory information, enabling more holistic decision-making in multidisciplinary tumor boards.
Secondly, clinical integration obstacles are also significant. Integrating AI into existing workflows faces the challenge of process reconstruction. Traditionally, the diagnostic process is “technician scanning → doctor reading images → writing reports”, with limited efficiency. The AI-driven process “AI pre-analysis during scanning → doctor reviewing AI markers → structured reporting” can improve efficiency and free up doctors’ energy, but due to difficulties in docking hospital information systems with AI tools, inconsistent interfaces and data standards, etc., the complexity of integration is increased. Therefore, breakthroughs depend not only on the model itself but also on the collaborative upgrading and process redesign of medical information systems.
At the same time, the positioning of the human-machine collaboration model needs to be clarified: AI as a “second reader” can improve diagnostic sensitivity and consistency, and as a "screening tool" can improve screening efficiency. However, differences in doctors’ trust in AI, the low false positive rate required for screening tools, and collaboration with doctors to avoid cognitive conflicts rather than replacement are urgent problems in practical deployment.
Finally, regulatory and ethical issues cannot be ignored, which are directly related to the safety and fairness of AI applications. The primary challenge is the risk of algorithmic bias, which stems from the lack of representativeness of training data (such as the lack of data on patients with chronic liver diseases such as cirrhosis), leading to reduced diagnostic accuracy and lack of fairness of the model for specific groups (especially cirrhotic patients). The key to breakthrough is to build diverse and representative datasets and introduce fairness and bias detection mechanisms for continuous monitoring.
Beyond bias, legal responsibility for AI-related misdiagnosis remains ambiguous. In real-world use, shared accountability models are being explored: Clinicians retain ultimate authority for diagnostic interpretation, while developers are responsible for algorithm integrity and updates. Future legal frameworks may include certification renewal mechanisms tied to model performance and traceability logs that record every AI decision to facilitate responsibility attribution.
Ethical and legal considerations also call for transparent “AI governance frameworks”. These should include institutional ethics committees overseeing algorithm validation, periodic audits for bias and drift, and clear patient consent processes for AI-assisted diagnosis. Importantly, patients should be informed when AI contributes to their diagnosis, and audit trails should ensure explainability and accountability.
In addition, a sound regulatory framework and ethical compliance are prerequisites for clinical promotion. Currently, a special regulatory system for AI medical devices is not yet sound. There is an urgent need to establish strict approval processes (drawing on medical device approval) to ensure safety and effectiveness, promote the standardization of AI ethics, and mandatorily require the protection of data privacy (such as compliance with GDPR), enhance model transparency and interpretability, eliminate the uncertainty caused by the “black box”, and improve clinical trust.
Looking ahead, the sustainable advancement of AI in HCC imaging will depend on a coordinated technological and governance roadmap. The gradual harmonization of international imaging standards-including upcoming digital imaging and communications in medicine extensions tailored for AI metadata-is expected to mitigate inter-institutional variability arising from heterogeneous scanners, acquisition parameters, and data formats, thereby strengthening cross-center reproducibility at the source. Concurrently, the establishment of collaborative, multi-institutional validation frameworks could support global HCC AI consortia that continuously benchmark algorithmic performance and monitor generalizability across diverse patient populations and clinical settings. For “continuously learning” AI systems, dynamic regulatory mechanisms will be essential to ensure traceable management of model updates and to align regulatory oversight with technological evolution, maintaining safety, efficacy, and controllability throughout the model lifecycle. From an ethical and governance perspective, incorporating algorithm auditability, fairness metrics, and explainability requirements into certification pathways will be critical for building a transparent and trustworthy AI ecosystem, fostering clinical confidence and facilitating deeper integration of AI into diagnostic decision-making. Collectively, the synergistic progress in technical standardization, collaborative validation, adaptive regulation, and ethical stewardship will provide the structural foundation for scaling HCC imaging AI from localized deployment to standardized, large-scale clinical adoption, ultimately positioning it as a core component of precision oncology.
AI has shown significant value and broad prospects in the field of imaging-assisted diagnosis of HCC. It effectively makes up for the limitations of traditional diagnostic methods by improving the sensitivity of identifying tiny lesions, quantifying imaging features, and shortening reporting time. Imaging data represented by CT, MRI, and US, together with evolving algorithm architectures from CNN to Transformer fusion and multi-phase spatiotemporal modeling, form a solid technical foundation, supporting application progress in lesion detection and localization, segmentation and quantitative analysis, benign-malignant differentiation and grading, and treatment response assessment. Although tools such as IQQA-Liver and SALSA have achieved clinical translation, they still face challenges such as cross-center generalization, model interpretability, clinical integration, and regulatory ethics. To truly bridge the gap between research innovation and widespread clinical adoption, a coordinated effort across technical development, clinical workflow redesign, and regulatory governance is urgently needed. Future progress will depend not only on advancing algorithms but also on building interoperable data infrastructures, embedding AI seamlessly into real-world radiology workflows, and establishing transparent and adaptive oversight mechanisms. Addressing the challenges discussed throughout this review-particularly multi-center variability, ethical safeguards, and the practical realities of human-AI collaboration-will be essential for ensuring trustworthy deployment. At this pivotal stage, the field must transition from demonstrating feasibility to implementing scalable, clinician-centered solutions that can reliably enhance decision-making, reduce diagnostic disparities, and ultimately improve patient outcomes. With sustained interdisciplinary collaboration, AI has the potential to become an integral, safe, and equitable component of HCC diagnosis and precision care.
| 1. | Calderaro J, Žigutytė L, Truhn D, Jaffe A, Kather JN. Artificial intelligence in liver cancer - new tools for research and patient management. Nat Rev Gastroenterol Hepatol. 2024;21:585-599. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 36] [Article Influence: 18.0] [Reference Citation Analysis (0)] |
| 2. | Yin C, Zhang H, Du J, Zhu Y, Zhu H, Yue H. Artificial intelligence in imaging for liver disease diagnosis. Front Med (Lausanne). 2025;12:1591523. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 16] [Article Influence: 16.0] [Reference Citation Analysis (1)] |
| 3. | Calderaro J, Seraphin TP, Luedde T, Simon TG. Artificial intelligence for the prevention and clinical management of hepatocellular carcinoma. J Hepatol. 2022;76:1348-1361. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 242] [Cited by in RCA: 201] [Article Influence: 50.3] [Reference Citation Analysis (1)] |
| 4. | Feng B, Ma XH, Wang S, Cai W, Liu XB, Zhao XM. Application of artificial intelligence in preoperative imaging of hepatocellular carcinoma: Current status and future perspectives. World J Gastroenterol. 2021;27:5341-5350. [PubMed] [DOI] [Full Text] |
| 5. | Koh B, Danpanichkul P, Wang M, Tan DJH, Ng CH. Application of artificial intelligence in the diagnosis of hepatocellular carcinoma. eGastroenterology. 2023;1:e100002. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 13] [Article Influence: 4.3] [Reference Citation Analysis (2)] |
| 6. | Zhang X, Yang L, Liu C, Yuan X, Zhang Y. An Artificial Intelligence Pipeline for Hepatocellular Carcinoma: From Data to Treatment Recommendations. Int J Gen Med. 2025;18:3581-3595. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 5.0] [Reference Citation Analysis (1)] |
| 7. | Kucukkaya AS, Zeevi T, Chai NX, Raju R, Haider SP, Elbanan M, Petukhova-Greenstein A, Lin M, Onofrey J, Nowak M, Cooper K, Thomas E, Santana J, Gebauer B, Mulligan D, Staib L, Batra R, Chapiro J. Predicting tumor recurrence on baseline MR imaging in patients with early-stage hepatocellular carcinoma using deep machine learning. Sci Rep. 2023;13:7579. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 14] [Article Influence: 4.7] [Reference Citation Analysis (1)] |
| 8. | Ma L, Li C, Li H, Zhang C, Deng K, Zhang W, Xie C. Deep learning model based on contrast-enhanced MRI for predicting post-surgical survival in patients with hepatocellular carcinoma. Heliyon. 2024;10:e31451. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 1.5] [Reference Citation Analysis (1)] |
| 9. | Zhang T, Yang F, Zhang P. Progress and clinical translation in hepatocellular carcinoma of deep learning in hepatic vascular segmentation. Digit Health. 2024;10:20552076241293498. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 1] [Article Influence: 0.5] [Reference Citation Analysis (1)] |
| 10. | Xu FWX, Tang SS, Soh HN, Pang NQ, Bonney GK. Augmenting care in hepatocellular carcinoma with artificial intelligence. Art Int Surg. 2023;3:48-63. [DOI] [Full Text] |
| 11. | Ayuso C, Bruix J. The challenges of novel contrast agents for the imaging diagnosis of hepatocellular carcinoma. Hepatol Int. 2014;8:4-6. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 0.3] [Reference Citation Analysis (1)] |
| 12. | Hamm CA, Wang CJ, Savic LJ, Ferrante M, Schobert I, Schlachter T, Lin M, Duncan JS, Weinreb JC, Chapiro J, Letzen B. Deep learning for liver tumor diagnosis part I: development of a convolutional neural network classifier for multi-phasic MRI. Eur Radiol. 2019;29:3338-3347. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 306] [Cited by in RCA: 236] [Article Influence: 33.7] [Reference Citation Analysis (8)] |
| 13. | Feng Z, Li H, Liu Q, Duan J, Zhou W, Yu X, Chen Q, Liu Z, Wang W, Rong P. CT Radiomics to Predict Macrotrabecular-Massive Subtype and Immune Status in Hepatocellular Carcinoma. Radiology. 2023;307:e221291. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 142] [Cited by in RCA: 124] [Article Influence: 41.3] [Reference Citation Analysis (3)] |
| 14. | Huang J, Wittbrodt MT, Teague CN, Karl E, Galal G, Thompson M, Chapa A, Chiu ML, Herynk B, Linchangco R, Serhal A, Heller JA, Abboud SF, Etemadi M. Efficiency and Quality of Generative AI-Assisted Radiograph Reporting. JAMA Netw Open. 2025;8:e2513921. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 21] [Article Influence: 21.0] [Reference Citation Analysis (1)] |
| 15. | Hanna RF, Miloushev VZ, Tang A, Finklestone LA, Brejt SZ, Sandhu RS, Santillan CS, Wolfson T, Gamst A, Sirlin CB. Comparative 13-year meta-analysis of the sensitivity and positive predictive value of ultrasound, CT, and MRI for detecting hepatocellular carcinoma. Abdom Radiol (NY). 2016;41:71-90. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 187] [Cited by in RCA: 171] [Article Influence: 17.1] [Reference Citation Analysis (4)] |
| 16. | Vilgrain V, Lagadec M, Ronot M. Pitfalls in Liver Imaging. Radiology. 2016;278:34-51. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 34] [Article Influence: 3.4] [Reference Citation Analysis (1)] |
| 17. | Welle CL, Guglielmo FF, Venkatesh SK. MRI of the liver: choosing the right contrast agent. Abdom Radiol (NY). 2020;45:384-392. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 26] [Article Influence: 4.3] [Reference Citation Analysis (1)] |
| 18. | Kopp FK, Catalano M, Pfeiffer D, Fingerle AA, Rummeny EJ, Noël PB. CNN as model observer in a liver lesion detection task for x-ray computed tomography: A phantom study. Med Phys. 2018;45:4439-4447. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 17] [Article Influence: 2.1] [Reference Citation Analysis (1)] |
| 19. | Vanmore SV, Chougule SR. Liver Lesions Classification System using CNN with Improved Accuracy. J Adv Appl Sci Res. 2022;4. [DOI] [Full Text] |
| 20. | Said D, Carbonell G, Stocker D, Hectors S, Vietti-Violi N, Bane O, Chin X, Schwartz M, Tabrizian P, Lewis S, Greenspan H, Jégou S, Schiratti JB, Jehanno P, Taouli B. Semiautomated segmentation of hepatocellular carcinoma tumors with MRI using convolutional neural networks. Eur Radiol. 2023;33:6020-6032. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 16] [Article Influence: 5.3] [Reference Citation Analysis (1)] |
| 21. | Kord S, Taghikhany T, Akbari M. A novel spatiotemporal 3D CNN framework with multi-task learning for efficient structural damage detection. Struct Health Monit. 2024;23:2270-2287. [DOI] [Full Text] |
| 22. | Li X, Chen H, Qi X, Dou Q, Fu CW, Heng PA. H-DenseUNet: Hybrid Densely Connected UNet for Liver and Tumor Segmentation From CT Volumes. IEEE Trans Med Imaging. 2018;37:2663-2674. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2111] [Cited by in RCA: 922] [Article Influence: 115.3] [Reference Citation Analysis (6)] |
| 23. | Hong SB, Choi SH, Kim SY, Shim JH, Lee SS, Byun JH, Park SH, Kim KW, Kim S, Lee NK. MRI Features for Predicting Microvascular Invasion of Hepatocellular Carcinoma: A Systematic Review and Meta-Analysis. Liver Cancer. 2021;10:94-106. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 144] [Cited by in RCA: 138] [Article Influence: 27.6] [Reference Citation Analysis (6)] |
| 24. | Roberts LR, Sirlin CB, Zaiem F, Almasri J, Prokop LJ, Heimbach JK, Murad MH, Mohammed K. Imaging for the diagnosis of hepatocellular carcinoma: A systematic review and meta-analysis. Hepatology. 2018;67:401-421. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 398] [Cited by in RCA: 372] [Article Influence: 46.5] [Reference Citation Analysis (4)] |
| 25. | Li J, Wei L, Zhang X, Zhang W, Wang H, Zhong B, Xie Z, Lv H, Wang X. DISMIR: Deep learning-based noninvasive cancer detection by integrating DNA sequence and methylation information of individual cell-free DNA reads. Brief Bioinform. 2021;22:bbab250. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 55] [Cited by in RCA: 46] [Article Influence: 9.2] [Reference Citation Analysis (1)] |
| 26. | Parsai A, Miquel ME, Jan H, Kastler A, Szyszko T, Zerizer I. Improving liver lesion characterisation using retrospective fusion of FDG PET/CT and MRI. Clin Imaging. 2019;55:23-28. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 18] [Article Influence: 2.6] [Reference Citation Analysis (1)] |
| 27. | Sun L, Jiang L, Wang M, Wang Z, Xin Y. A Multi-Scale Liver Tumor Segmentation Method Based on Residual and Hybrid Attention Enhanced Network with Contextual Integration. Sensors (Basel). 2024;24:5845. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 2] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 28. | Huang Z, Ye J, Wang H, Deng Z, Yang Z, Su Y, Liu J, Li T, Gu Y, Zhang S, Qiao Y, Gu L, He J. Revisiting model scaling with a U-net benchmark for 3D medical image segmentation. Sci Rep. 2025;15:29795. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 7] [Article Influence: 7.0] [Reference Citation Analysis (1)] |
| 29. | Gao Q, Almekkawy M. ASU-Net++: A nested U-Net with adaptive feature extractions for liver tumor segmentation. Comput Biol Med. 2021;136:104688. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 30] [Article Influence: 6.0] [Reference Citation Analysis (1)] |
| 30. | Yang Z, Yang X, Cao Y, Shao Q, Tang D, Peng Z, Di S, Zhao Y, Li S. Deep learning based automatic internal gross target volume delineation from 4D-CT of hepatocellular carcinoma patients. J Appl Clin Med Phys. 2024;25:e14211. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 7] [Article Influence: 3.5] [Reference Citation Analysis (1)] |
| 31. | He K, Liu X, Shahzad R, Reimer R, Thiele F, Niehoff J, Wybranski C, Bunck AC, Zhang H, Perkuhn M. Advanced Deep Learning Approach to Automatically Segment Malignant Tumors and Ablation Zone in the Liver With Contrast-Enhanced CT. Front Oncol. 2021;11:669437. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 15] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 32. | Chen M, Zhang B, Topatana W, Cao J, Zhu H, Juengpanich S, Mao Q, Yu H, Cai X. Classification and mutation prediction based on histopathology H&E images in liver cancer using deep learning. NPJ Precis Oncol. 2020;4:14. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 207] [Cited by in RCA: 151] [Article Influence: 25.2] [Reference Citation Analysis (2)] |
| 33. | Jang HJ, Kim TK, Wilson SR. Imaging of malignant liver masses: characterization and detection. Ultrasound Q. 2006;22:19-29. [PubMed] |
| 34. | Javed S, Mahmood A, Dias J, Werghi N, Rajpoot N. Spatially Constrained Context-Aware Hierarchical Deep Correlation Filters for Nucleus Detection in Histology Images. Med Image Anal. 2021;72:102104. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 11] [Article Influence: 2.2] [Reference Citation Analysis (1)] |
| 35. | Xia TY, Zhou ZH, Meng XP, Zha JH, Yu Q, Wang WL, Song Y, Wang YC, Tang TY, Xu J, Zhang T, Long XY, Liang Y, Xiao WB, Ju SH. Predicting Microvascular Invasion in Hepatocellular Carcinoma Using CT-based Radiomics Model. Radiology. 2023;307:e222729. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 205] [Cited by in RCA: 176] [Article Influence: 58.7] [Reference Citation Analysis (6)] |
| 36. | Zheng R, Wang Q, Lv S, Li C, Wang C, Chen W, Wang H. Automatic Liver Tumor Segmentation on Dynamic Contrast Enhanced MRI Using 4D Information: Deep Learning Model Based on 3D Convolution and Convolutional LSTM. IEEE Trans Med Imaging. 2022;41:2965-2976. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 97] [Cited by in RCA: 38] [Article Influence: 9.5] [Reference Citation Analysis (5)] |
| 37. | Li Y, Wang P, Xu J, Shi X, Yin T, Teng F. Noninvasive radiomic biomarkers for predicting pseudoprogression and hyperprogression in patients with non-small cell lung cancer treated with immune checkpoint inhibition. Oncoimmunology. 2024;13:2312628. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 20] [Article Influence: 10.0] [Reference Citation Analysis (1)] |
| 38. | Smagulova KK, Kaidarova DR, Chingissova ZK, Nasrytdinov TS, Khozhayev AA. Effects of metastatic CRC predictors on treatment outcomes. Surg Oncol. 2022;44:101835. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 0.8] [Reference Citation Analysis (1)] |
| 39. | Balaguer-Montero M, Marcos Morales A, Ligero M, Zatse C, Leiva D, Atlagich LM, Staikoglou N, Viaplana C, Monreal C, Mateo J, Hernando J, García-Álvarez A, Salvà F, Capdevila J, Elez E, Dienstmann R, Garralda E, Perez-Lopez R. A CT-based deep learning-driven tool for automatic liver tumor detection and delineation in patients with cancer. Cell Rep Med. 2025;6:102032. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 8] [Article Influence: 8.0] [Reference Citation Analysis (1)] |
| 40. | Gresser E, Schachtner B, Stüber AT, Solyanik O, Schreier A, Huber T, Froelich MF, Magistro G, Kretschmer A, Stief C, Ricke J, Ingrisch M, Nörenberg D. Performance variability of radiomics machine learning models for the detection of clinically significant prostate cancer in heterogeneous MRI datasets. Quant Imaging Med Surg. 2022;12:4990-5003. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 12] [Article Influence: 3.0] [Reference Citation Analysis (1)] |