Copyright: ©Author(s) 2026.
World J Hepatol. Mar 27, 2026; 18(3): 117134
Published online Mar 27, 2026. doi: 10.4254/wjh.v18.i3.117134
Published online Mar 27, 2026. doi: 10.4254/wjh.v18.i3.117134
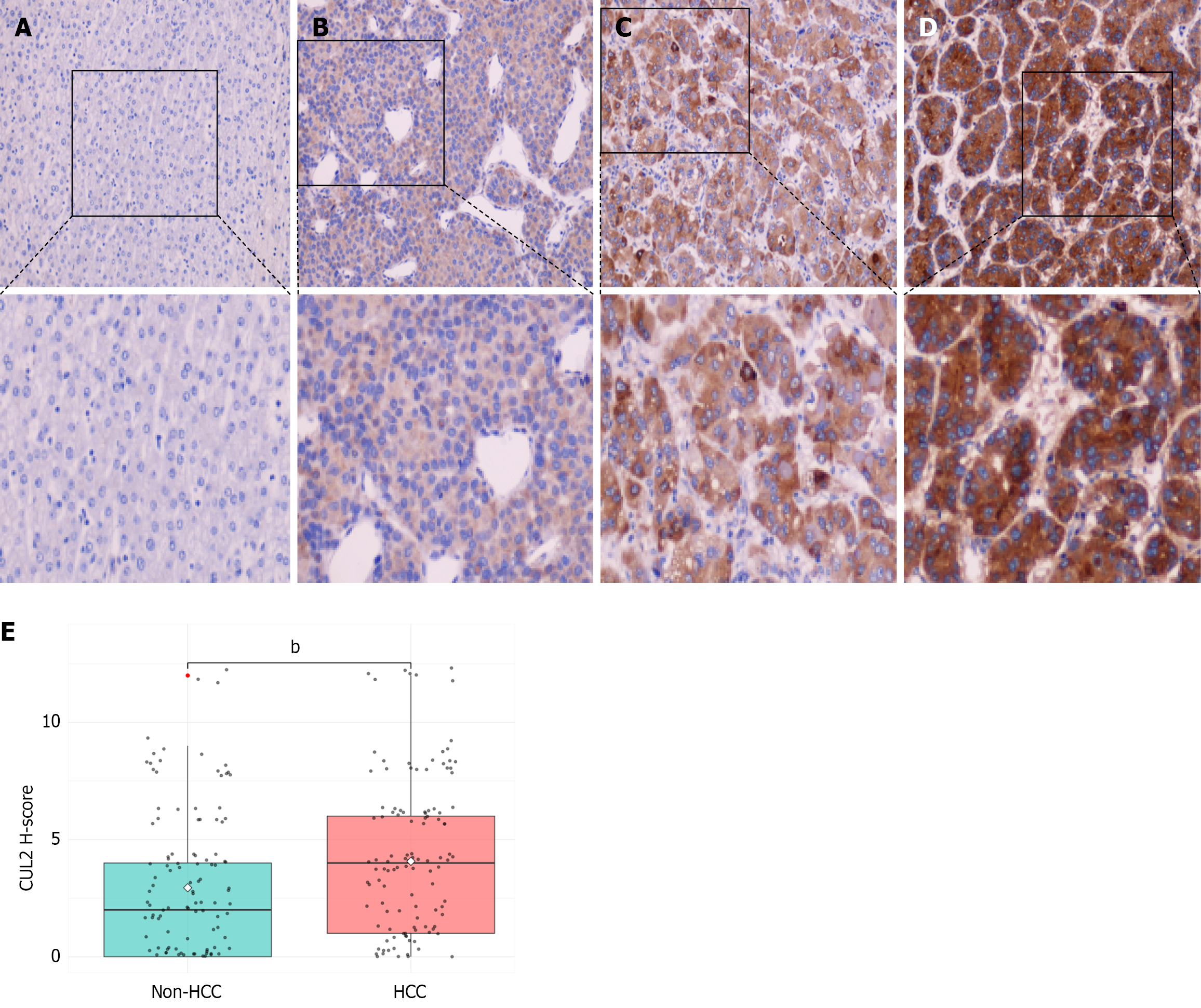
Figure 1 Immunohistochemical analysis of Cullin-2 protein in hepatocellular carcinoma tissues.
A: Adjacent liver tissue (200 ×); B-D: Hepatocellular carcinoma tissues (200 ×); E: Box plot of H-score distribution. bP < 0.01. HCC: Hepatocellular carcinoma; CUL2: Cullin-2.
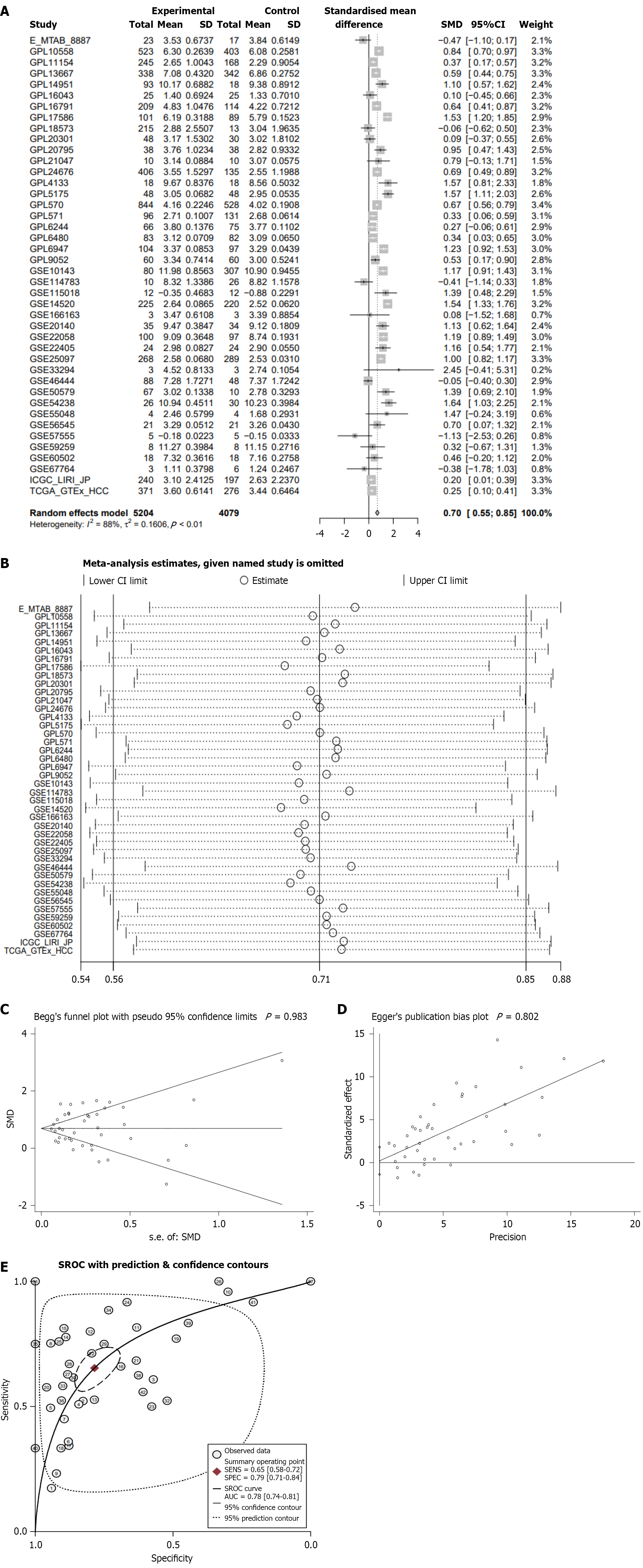
Figure 2 Integrated analysis of global hepatocellular carcinoma gene microarray and high-throughput sequencing data.
A: Cullin-2 mRNA is significantly overexpressed in 5204 hepatocellular carcinoma tissues compared to 4079 non-cancerous tissues; B: Sensitivity analysis results indicate stability of the standardized mean difference, as assessed by publication bias test; C: Begg’s test; D: Egger’s test; E: Pooled receiver operating characteristic curve analysis. SROC: Summary receiver operating characteristic; SMD: Standardized mean difference.
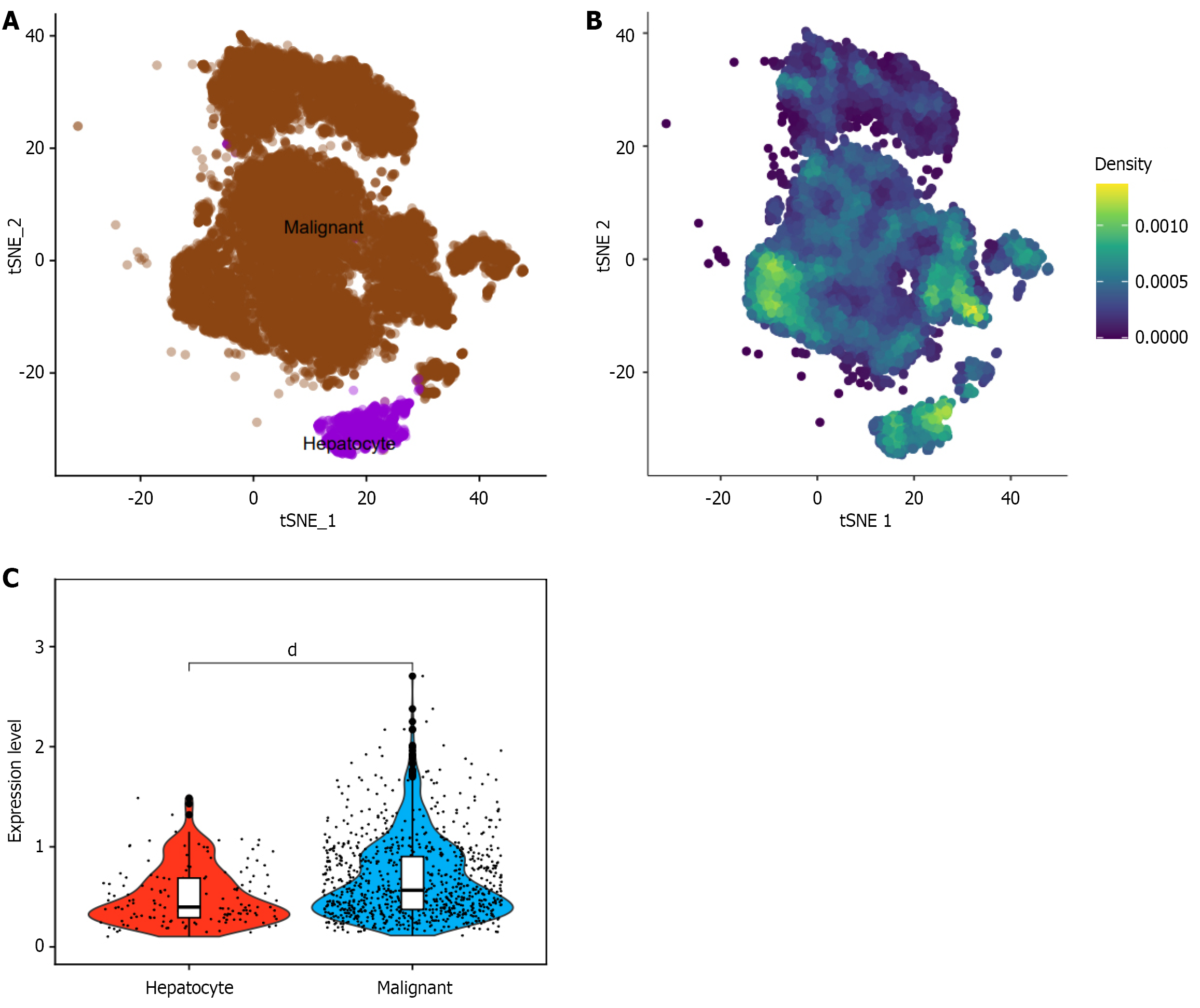
Figure 3 Single-cell sequencing analysis of CUL2 mRNA.
A: Annotation of hepatocellular carcinoma (HCC) cells; B: Expression of Cullin-2 (CUL2) mRNA in HCC subpopulations; C: Compared with normal hepatocytes, CUL2 mRNA is significantly overexpressed in HCC single-cell subpopulations. dP < 0.0001.
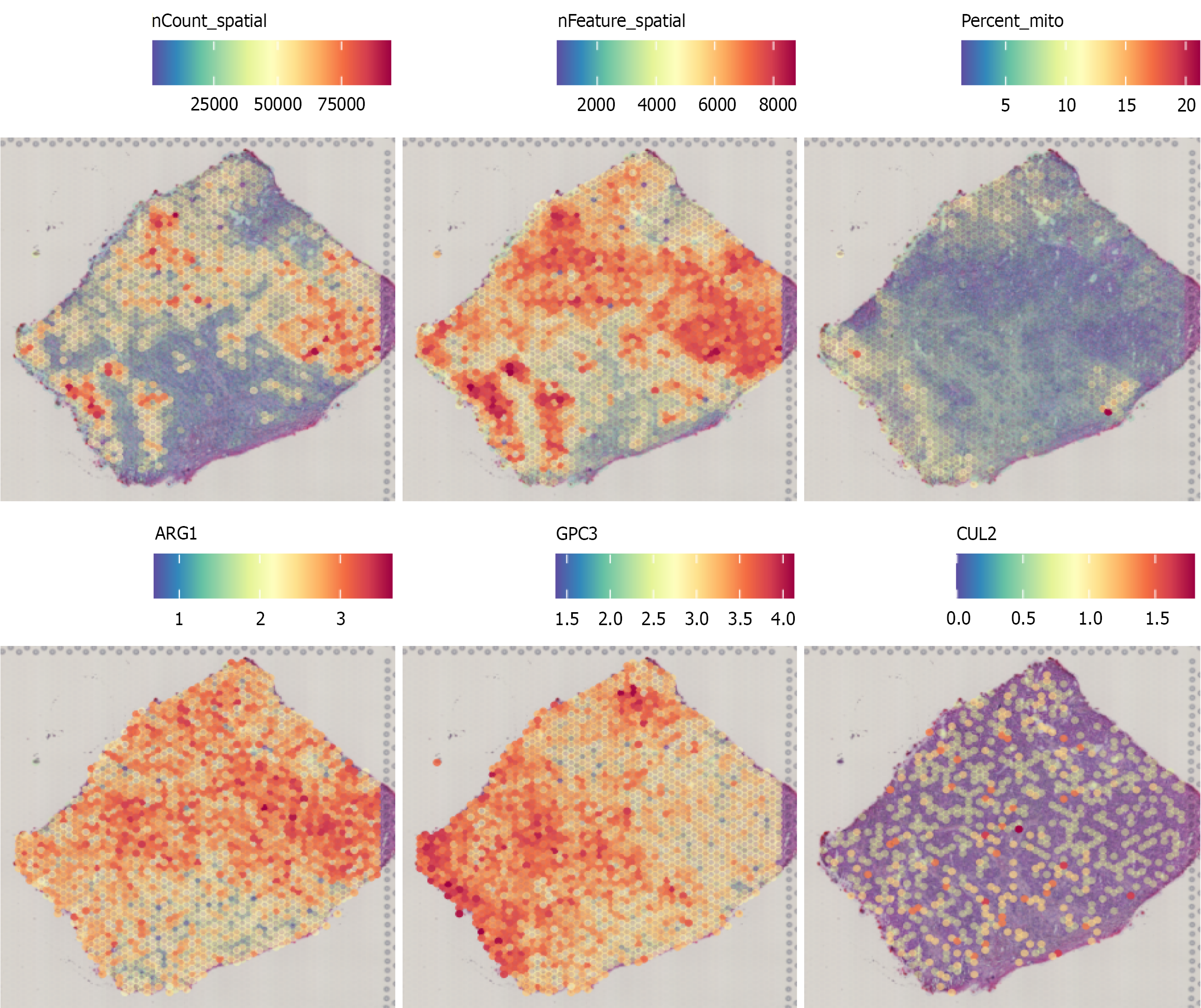
Figure 4
Spatial transcriptomics analysis of Cullin-2 mRNA.
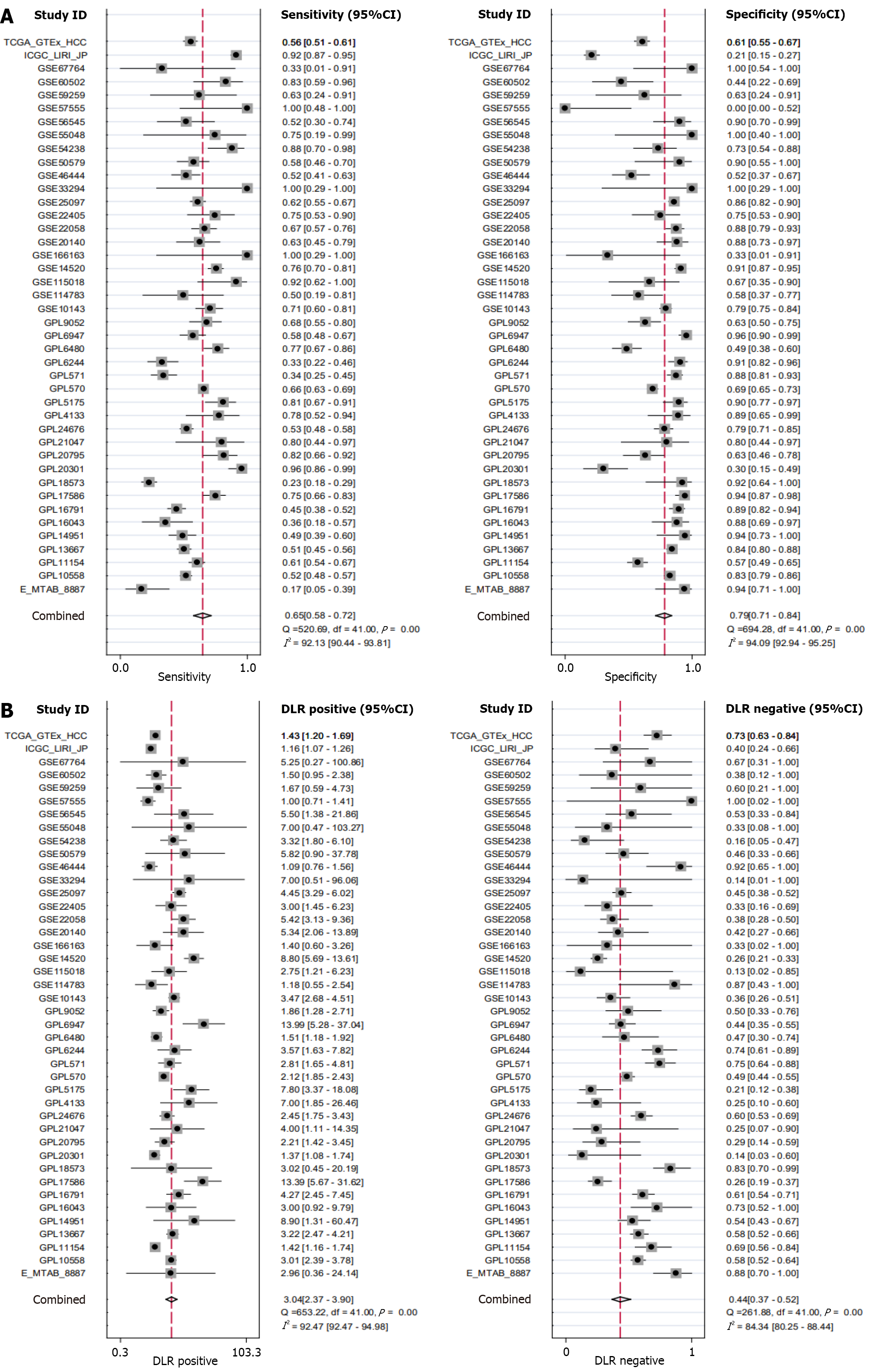
Figure 5 Forest plot of Cullin-2 mRNA sensitivity, specificity, and likelihood ratios.
A: Sensitivity and specificity of Cullin-2 (CUL2) mRNA for discriminating hepatocellular carcinoma (HCC) tissues; B: Positive likelihood ratio and negative likelihood ratio of CUL2 mRNA for discriminating HCC tissues.
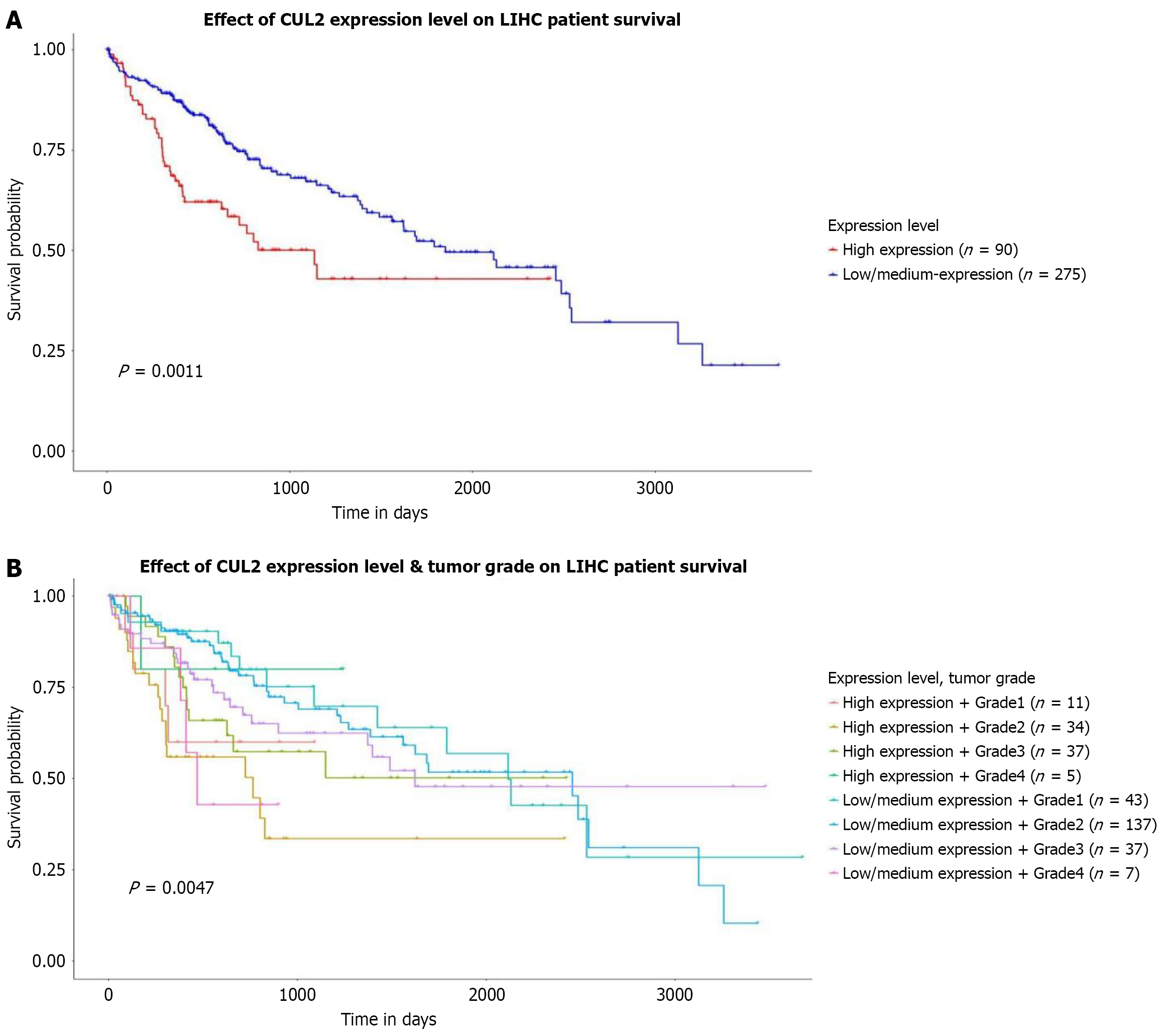
Figure 6 Analysis of the relationship between Cullin-2 mRNA expression levels and prognosis in hepatocellular carcinoma using the UALCAN database.
A: Impact of Cullin-2 (CUL2) mRNA expression levels on the survival rate of hepatocellular carcinoma (HCC) patients; B: Combined effects of CUL2 mRNA expression levels and tumor grade on the survival rate of HCC patients. CUL2: Cullin-2.
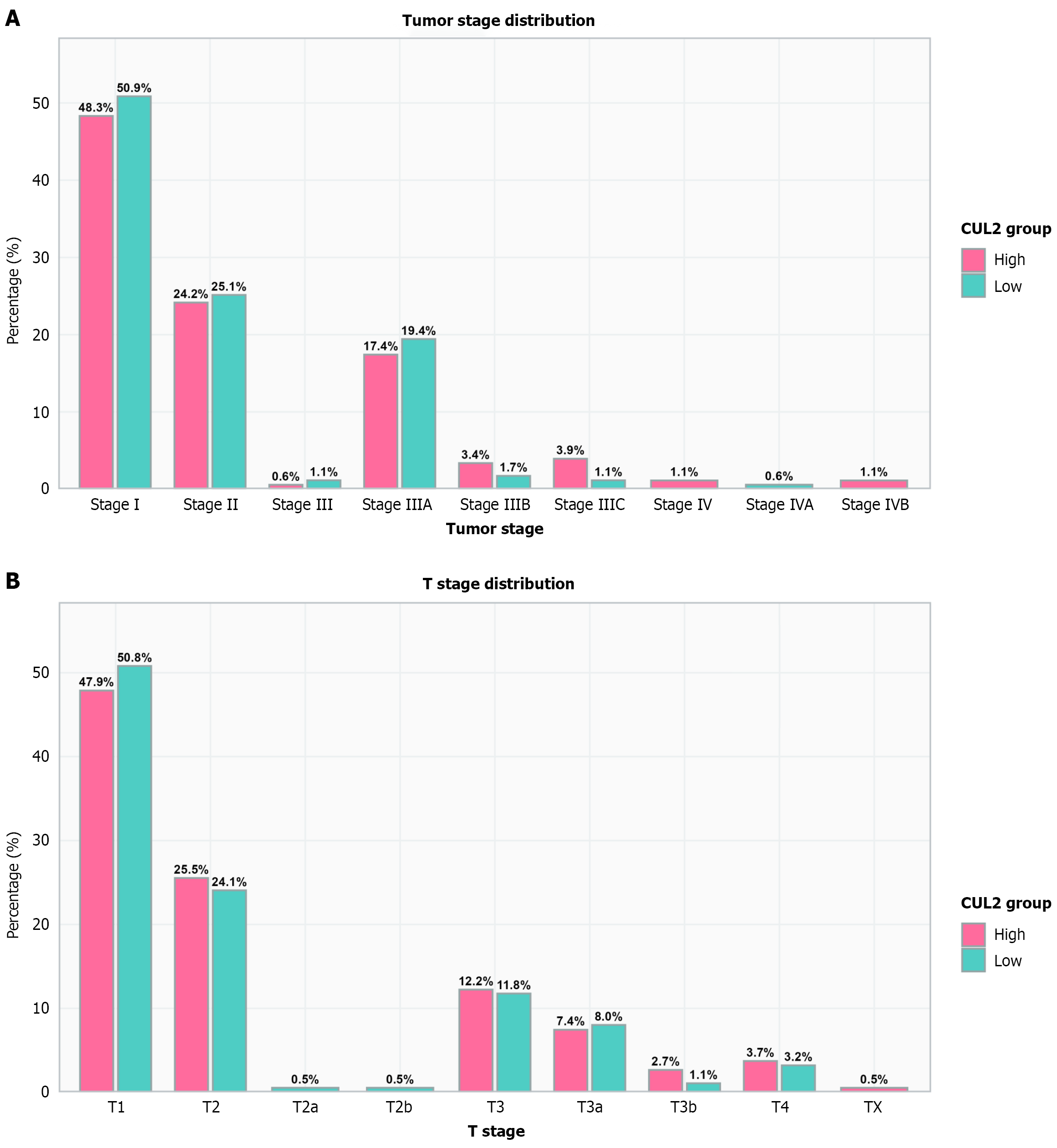
Figure 7 Correlation between Cullin-2 expression and clinical-pathological features in The Cancer Genome Atlas-LIHC cohort.
A: The distribution of primary tumor T stage between Cullin-2 (CUL2) expression groups indicates that both groups are predominantly comprised of patients with early-stage lesions (T1-T2); B: There is no significant difference in the overall AJCC stage distribution between CUL2 expression groups. CUL2: Cullin-2.
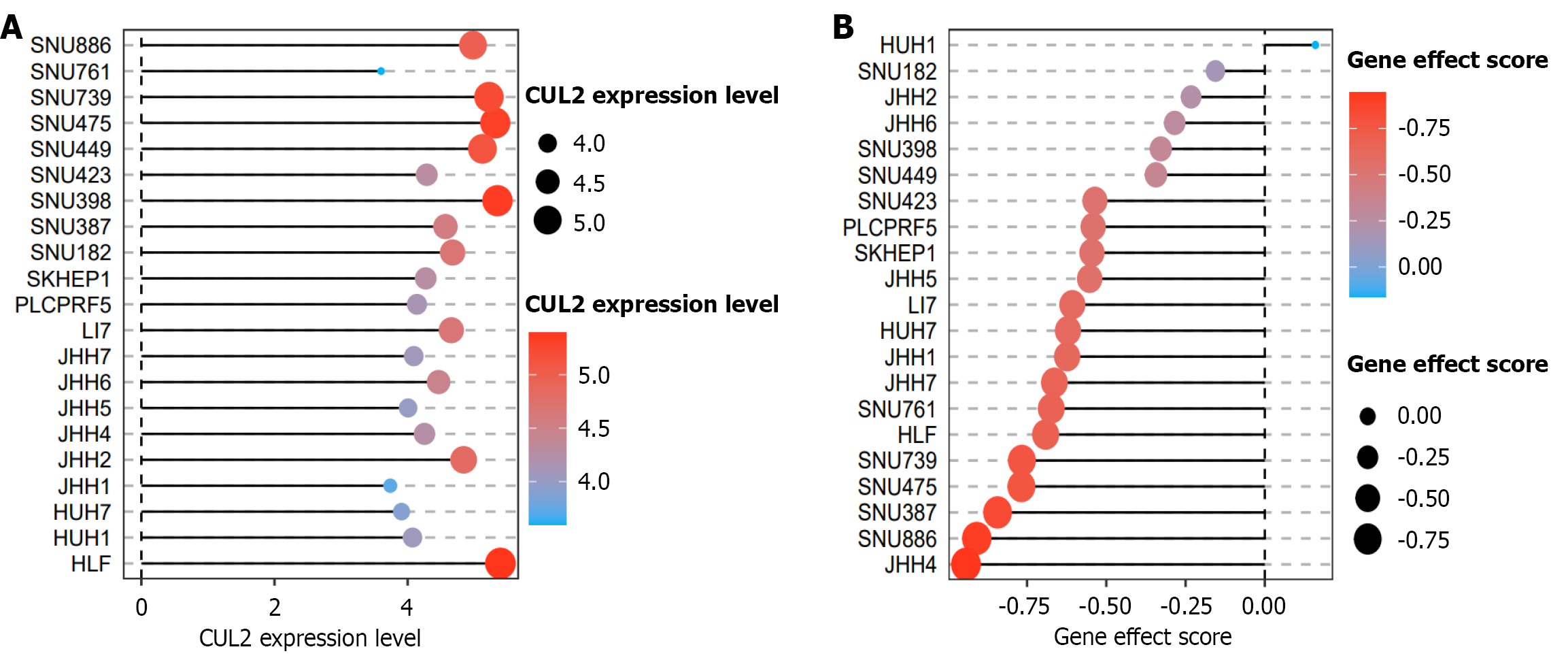
Figure 8 Hepatocellular carcinoma Cullin-2 mRNA CRISPR analysis.
A: Cullin-2 (CUL2) expression in hepatocellular carcinoma (HCC) cell lines; B: HCC CUL2 CRISPR score. CUL2: Cullin-2.
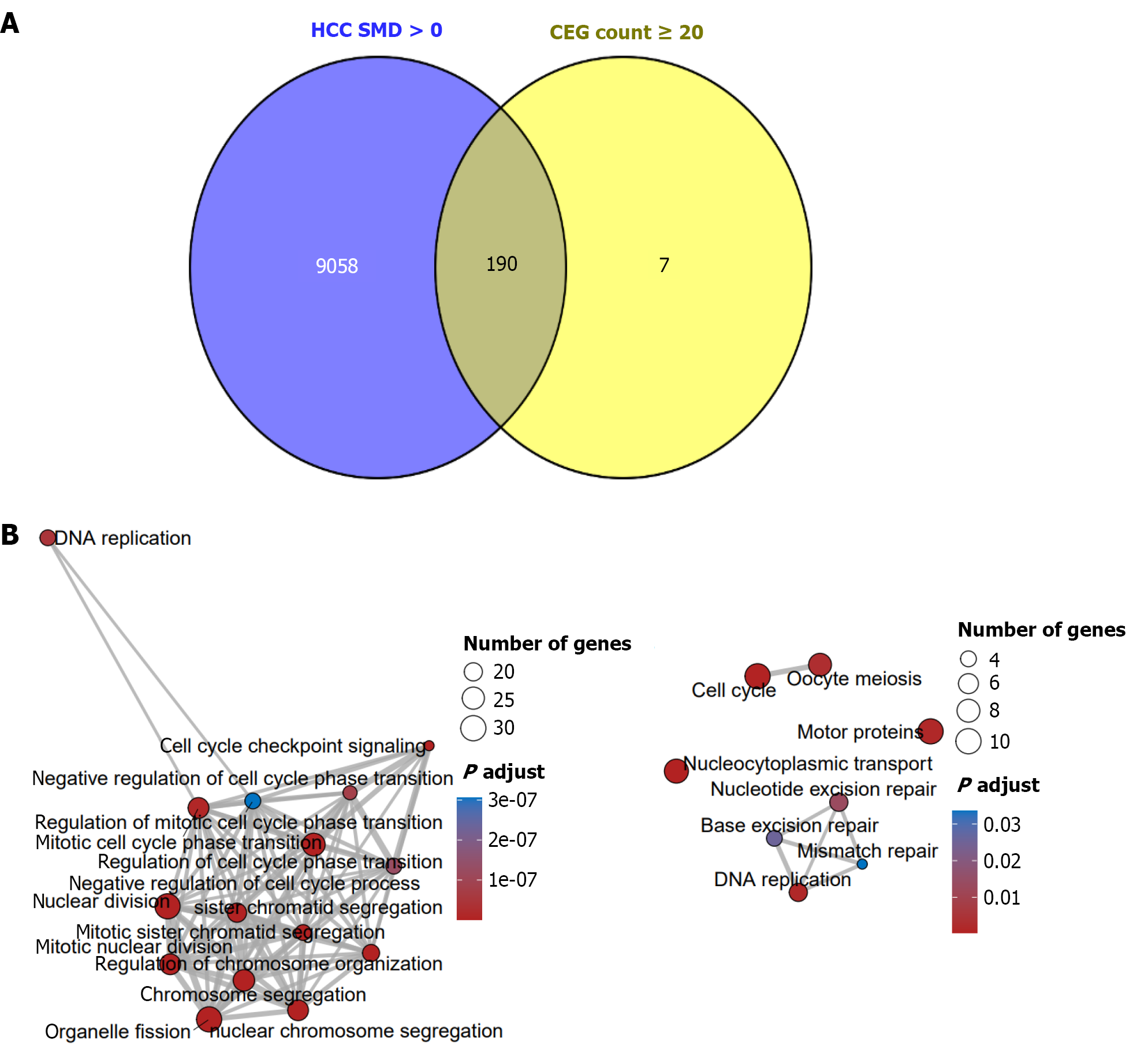
Figure 9 Analysis of potential molecular mechanisms of Cullin-2 in hepatocellular carcinoma.
A: This study identified 190 hepatocellular carcinoma highly expressed Cullin-2 (CUL2)-positively correlated genes; B: The CUL2 co-expression gene regulatory network is closely related to Gene Ontology terms such as nuclear division, chromosome segregation, and mitotic cell cycle transition, and significantly enriched in signaling pathways including nucleocytoplasmic transport and cell cycle. HCC: Hepatocellular carcinoma; SMD: Standardized mean difference.
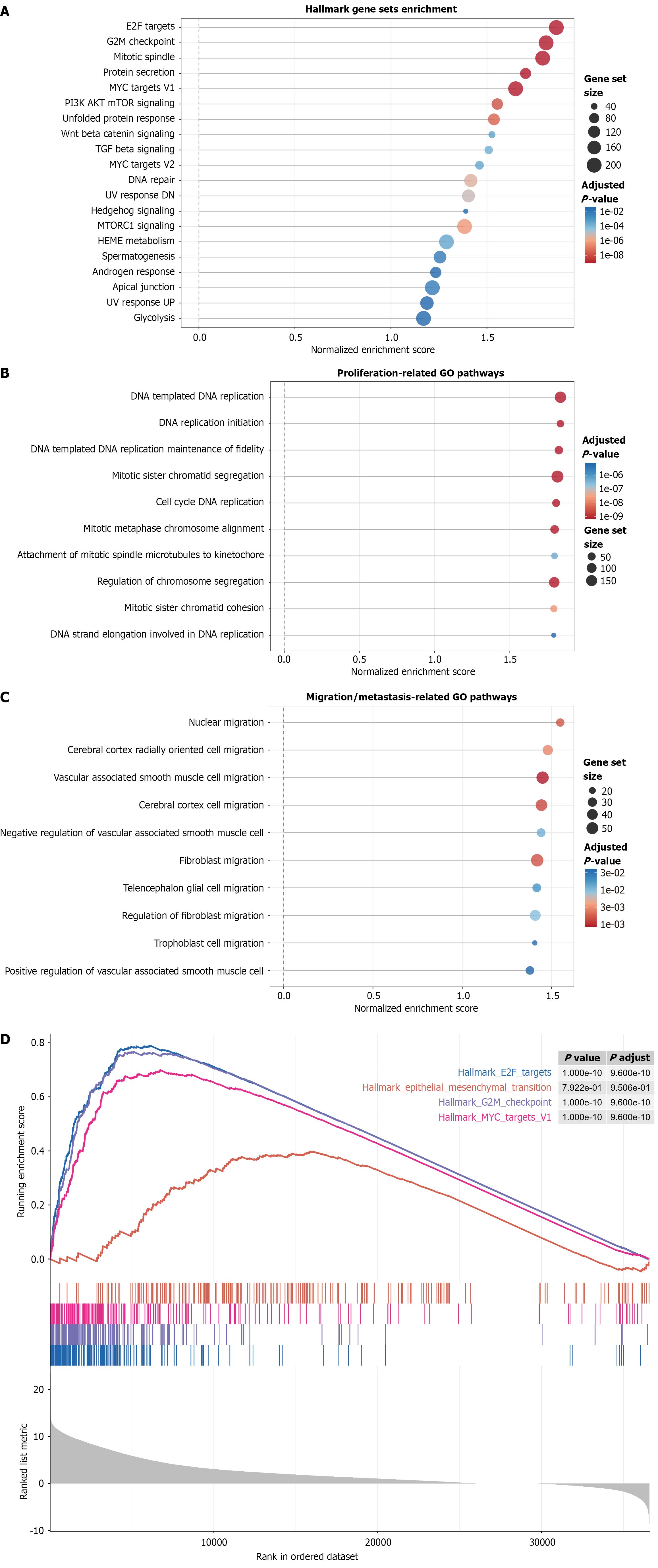
Figure 10 Functional enrichment analysis of Cullin-2 in hepatocellular carcinoma.
A: Hallmark pathway analysis reveals significant enrichment of E2F targets, G2M checkpoint, and MYC signaling in Cullin-2 (CUL2)-high tumors; B: Gene Ontology analysis shows association with DNA replication, sister chromatid segregation, and cell cycle progression; C: CUL2 is also enriched in pathways related to nuclear migration and cellular motility; D: Gene set enrichment analysis enrichment plots confirm coordinated upregulation of key proliferation and EMT pathways in CUL2-high samples.
- Citation: Tang P, Nong QH, Xiong KH, Tang HH, Huang BB, Huang GL, Li Q, He SP, Chen G, Feng ZB, Li JD. Overexpression and clinicopathological significance of cullin-2 in hepatocellular carcinoma progression. World J Hepatol 2026; 18(3): 117134
- URL: https://www.wjgnet.com/1948-5182/full/v18/i3/117134.htm
- DOI: https://dx.doi.org/10.4254/wjh.v18.i3.117134