Copyright: ©Author(s) 2026.
World J Stem Cells. Apr 26, 2026; 18(4): 118621
Published online Apr 26, 2026. doi: 10.4252/wjsc.v18.i4.118621
Published online Apr 26, 2026. doi: 10.4252/wjsc.v18.i4.118621
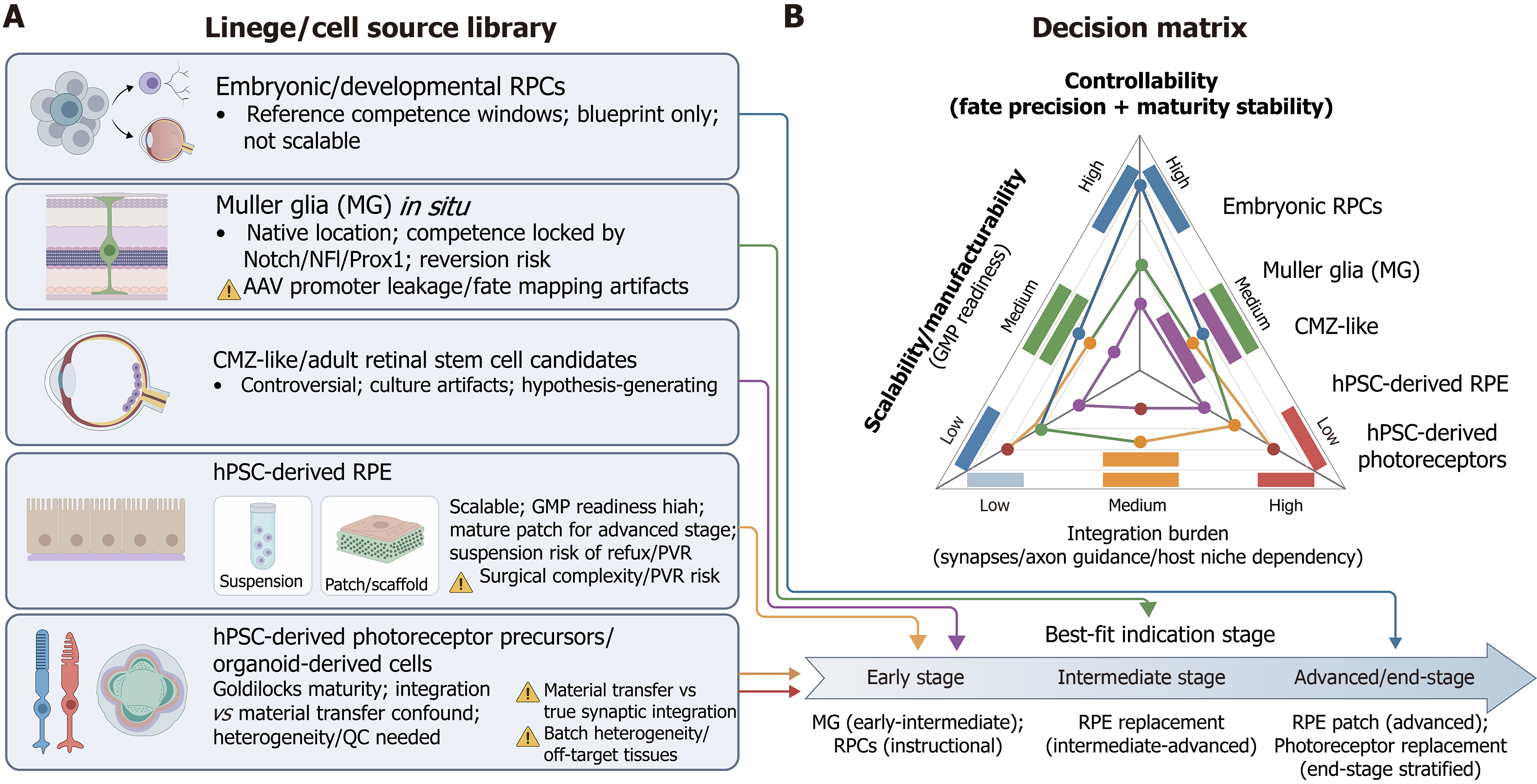
Figure 1 Retinal regenerative lineage library: Sources vs controllability/scalability/translational fit.
A: Lineage/cell source library: A categorization of the major regenerative substrates discussed in this review. Embryonic retinal progenitor cells (RPCs) serve primarily as developmental blueprints (reference only). Among adult-associated substrates, Müller glia (MG)-based approaches and human pluripotent stem cell-derived products represent the most tractable therapeutic routes, whereas ciliary marginal zone-like/adult-candidate populations are presented here as hypothesis-generating developmental-edge programs with currently limited evidence as a near-term clinical cell source; B: Decision matrix & indication stage: A multi-axis evaluation of these sources based on three translational criteria: Controllability: The precision of fate specification and stability of the maturation state [high in RPCs/retinal pigment epithelium (RPE); lower in MG/ciliary marginal zone]. Scalability: Good Manufacturing Practice manufacturing readiness (highest in human pluripotent stem cell-derived RPE). Integration burden: The complexity of host connectivity required for function (highest for photoreceptors/RPCs; lowest for RPE). The bottom timeline maps these sources to their “best-fit” disease stages, proposing MG reprogramming for early-stage intervention and cell replacement (RPE/photoreceptors) for intermediate-to-end-stage degeneration. Critical cautions: Key risks are highlighted, including adeno-associated virus promoter leakage artifacts in MG studies, the confounding of material transfer vs synaptic integration in photoreceptor grafts, and surgical complexity for RPE patches. RPCs: Retinal progenitor cells; Nuclear factor I; Prox1: Prospero homeobox 1; AAV: Adeno-associated virus; CMZ: Ciliary marginal zone; hPSC: Human pluripotent stem cell; RPE: Retinal pigment epithelium; GMP: Good Manufacturing Practice; PVR: Proliferative vitreoretinopathy; QC: Quality control.
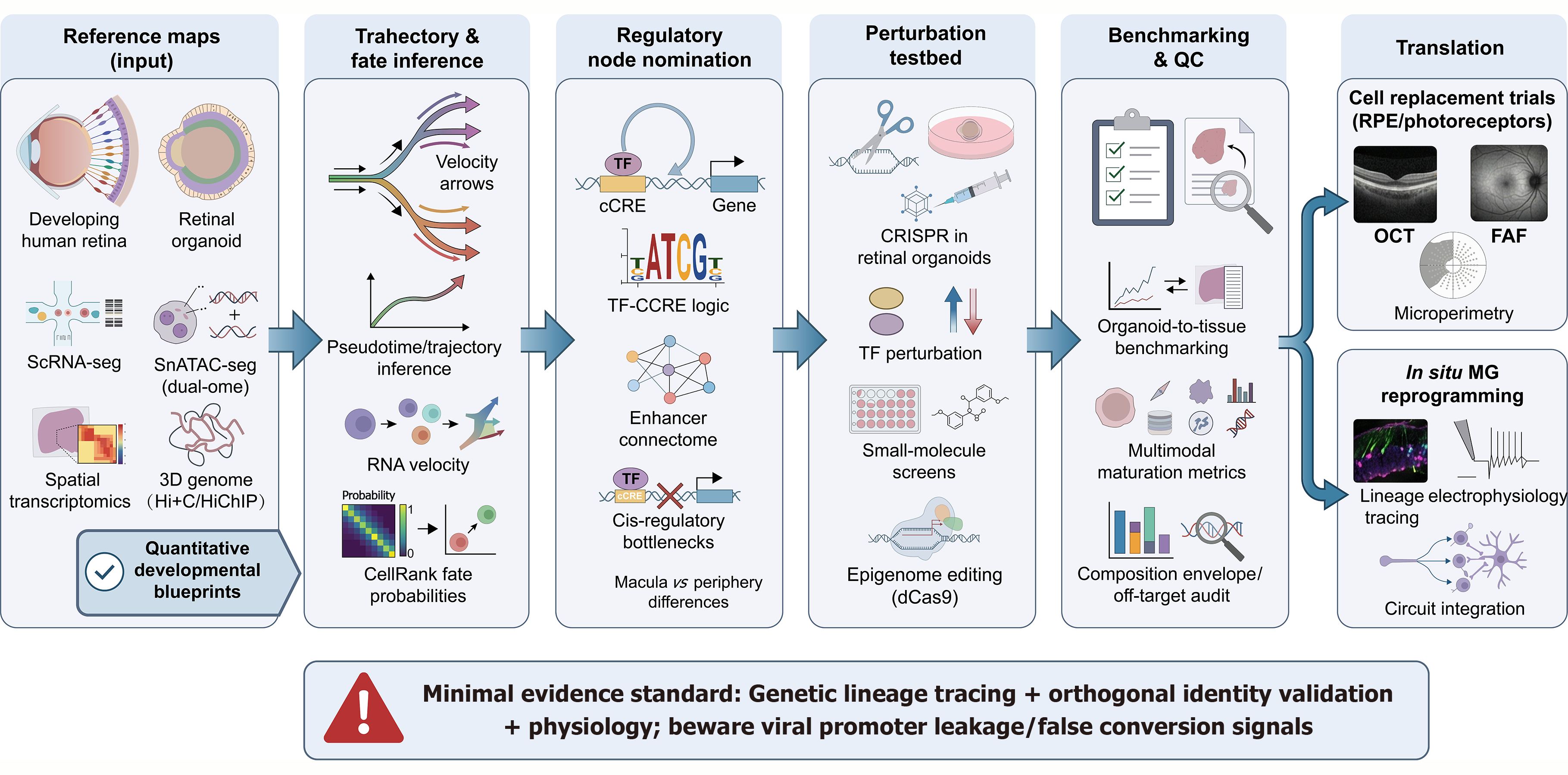
Figure 2 The “atlas-to-engineering” workflow: From quantitative blueprints to validated regenerative therapies.
This pipeline illustrates how multi-omic data is operationalized for fate engineering: (1) Reference maps (input): The integration of single-cell transcriptomics (single-cell RNA sequencing), dual-omic profiling (single nucleus assay for transposase-accessible chromatin sequencing), and spatial transcriptomics creates quantitative developmental blueprints; (2) Trajectory & fate inference: Computational frameworks such as RNA velocity and CellRank use these blueprints to define lineage directionality and probabilistic fate decision regions; (3) Regulatory node nomination: Inference of gene regulatory networks identifies the specific transcription factor and cis-regulatory element logic driving lineage restrictions; (4) Perturbation testbed: Candidate control nodes are causally validated using CRISPR-Cas9 or small-molecule screens within human retinal organoids to test their ability to shift fate; (5) Benchmarking & quality control: Engineered cells are evaluated against primary tissue references using multimodal metrics to ensure they meet composition and maturation envelopes; and (6) Translation: Validated protocols are deployed for cell replacement (with structural/functional endpoints) or in situ reprogramming. Critical caution (bottom): A minimal evidence standard is required to distinguish true regeneration from artifacts, necessitating genetic lineage tracing, orthogonal identity validation, and physiological proof-of-function to rule out viral promoter leakage. QC: Quality control; scRNA: Single-cell RNA; snATAC: Single nucleus assay for transposase-accessible chromatin; 3D: Three-dimensional; TF: Transcription factor; cCRE: Cis-regulatory element; dCas9: Catalytically dead Cas9; RPE: Retinal pigment epithelium; OCT: Optical coherence tomography; FAF: Fundus autofluorescence; MG: Müller glia.
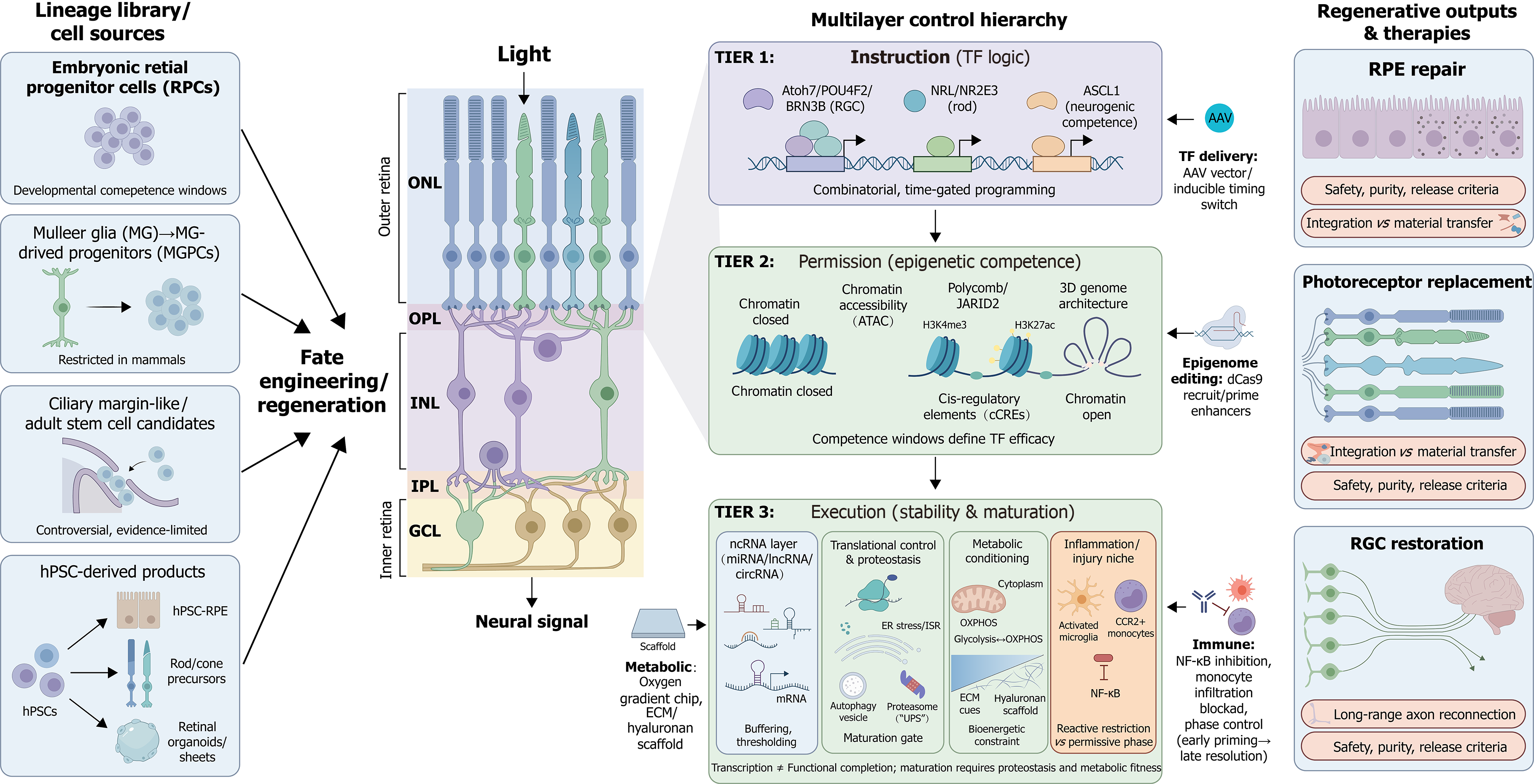
Figure 3 The multilayered fate-control paradigm for retinal regeneration: From lineage sources to therapeutic outputs.
This schematic illustrates the hierarchical framework governing retinal cell fate engineering. The lineage library: Distinct cell sources available for regeneration, ranging from developmental references (embryonic retinal progenitor cells) and endogenous candidates (Müller glia, ciliary marginal zone-like cells) to scalable manufacturing sources (human pluripotent stem cell-derived retinal pigment epithelium, organoids, and photoreceptors) (left). The multilayer control hierarchy: Successful fate engineering requires the convergence of three regulatory tiers: (1) Instruction (tier 1): Lineage-determining transcription factors (e.g., Atoh7, neural retina leucine-zipper, ASCL1) that provide combinatorial and time-gated programming logic; (2) Permission (tier 2): The epigenetic landscape (chromatin accessibility, histone modifications, 3D architecture) that defines competence windows and determines whether transcription factor instructions can access cis-regulatory elements; and (3) Execution (tier 3): The stabilization and maturation layer, comprising non-coding RNAs, translational/proteostatic control, metabolic conditioning (e.g., glycolysis/OXPHOS balance), and the immune microenvironment (e.g., inflammation phase control), which collectively ensure the functional completion of the transcriptional program (middle). Regenerative outputs & therapies: The application of this logic to specific clinical targets - retinal pigment epithelium repair, photoreceptor replacement, and retinal ganglion cell restoration - highlighting key translational bottlenecks such as synaptic integration, long-range axon guidance, and safety/purity release criteria (right). hPSC: Human pluripotent stem cell; RPE: Retinal pigment epithelium; RGC: Retinal ganglion cell; NRL: Neural retina leucine-zipper; ATAC: Assay for transposase-accessible chromatin; TFs: Transcription factors; miRNA: MicroRNA; lncRNA: Long non-coding RNA; circRNA: Circular RNA; ECM: Extracellular matrix; ER: Endoplasmic reticulum; CCR2+: C-C chemokine receptor type 2-positive; NF-κB: Nuclear factor kappa B; AAV: Adeno-associated virus; dCas9: Catalytically dead Cas9.
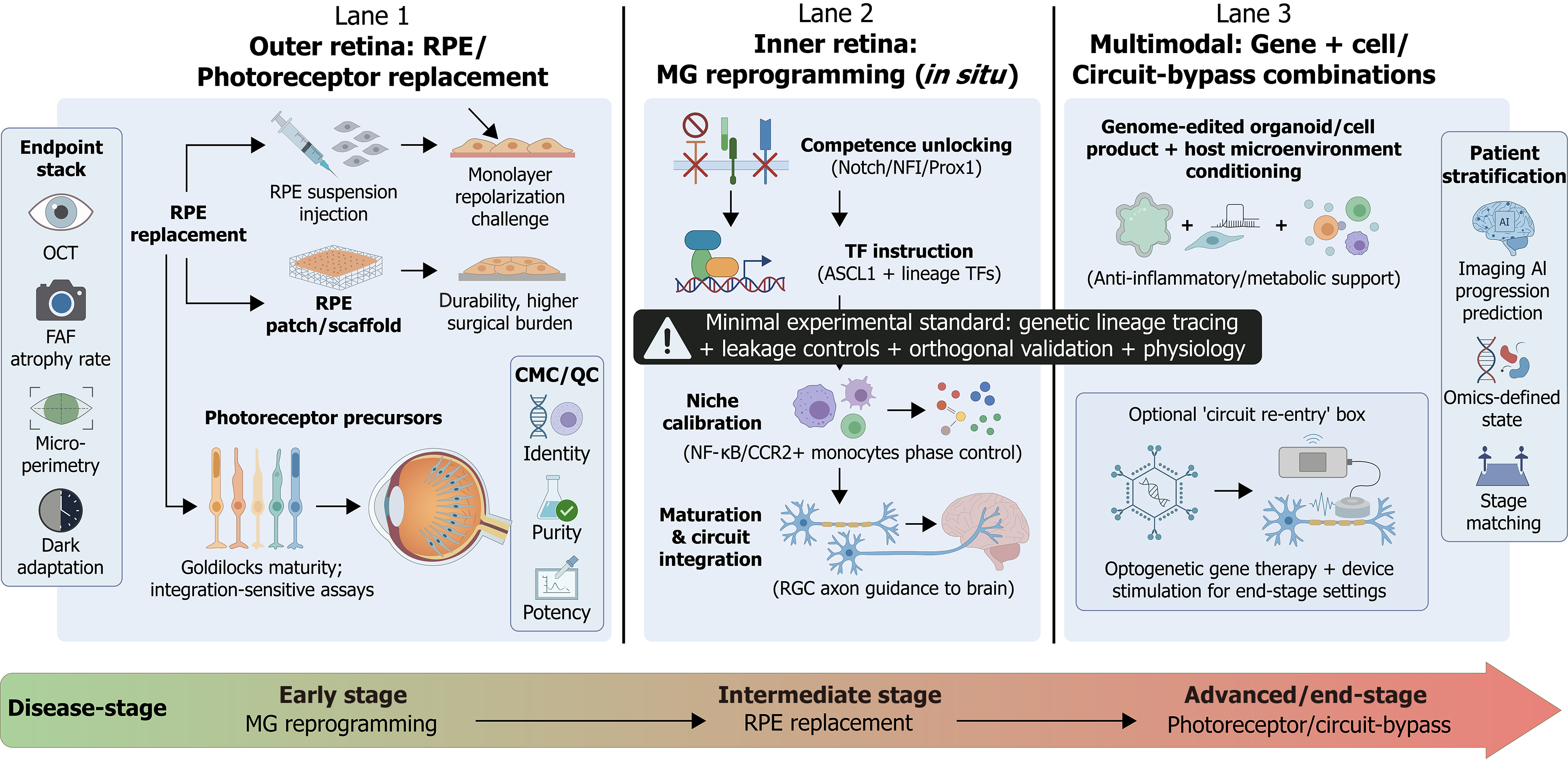
Figure 4 Regenerative ophthalmology strategies for retinal degeneration: Stage-matched, multimodal fate engineering.
This schematic synthesizes the clinical pathways for retinal fate engineering across three distinct therapeutic lanes, aligned with disease progression. Outer retina replacement: Strategies for RPE and photoreceptor replacement (discussed in “Regenerative strategy II: Clinical pathway and evidence hierarchy for cell-replacement therapy”), detailing delivery modalities (suspension vs scaffold) and the requirement for a multimodal endpoint stack (optical coherence tomography, fundus autofluorescence, microperimetry) alongside rigorous chemistry, manufacturing, and controls release criteria (identity, purity, potency) to ensure graft function (lane 1). Inner retina reprogramming: In situ Müller glia reprogramming (discussed in the section of Regenerative strategy III: MG reprogramming - from “feasible” to controllable, reproducible, and translatable) is depicted as a staged control sequence: Unlocking competence (e.g., via Notch/nuclear factor I inhibition), identifying lineage instruction (ASCL1), and calibrating the immune niche. A critical warning highlights the “minimal experimental standard” required to rule out viral leakage artifacts (lane 2). Multimodal combinations: “Gene + cell” and circuit-bypass strategies (discussed in “Gene + cell and multimodal combination strategies: Turning fate-control networks into therapies”) that couple biological replacement with host microenvironment conditioning or optogenetic re-entry (lane 3). Patient stratification: The timeline illustrates the necessity of precision stage-matching - deploying reprogramming for early-stage rescue, cell replacement for intermediate degeneration, and circuit-bypass for end-stage atrophy - guided by artificial intelligence-driven progression prediction (bottom). RPE: Retinal pigment epithelium; OCT: Optical coherence tomography; FAF: Fundus autofluorescence; CMC: Chemistry, manufacturing, and controls; QC: Quality control; MG: Müller glia; TF: Transcription factor; NFI: Nuclear factor I; Prox1: Prospero homeobox 1; NF-κB: Nuclear factor kappa B; CCR2+: C-C chemokine receptor type 2-positive; RGC: Retinal ganglion cell; AI: Artificial intelligence.
- Citation: Xie QQ, Zeng MQ, Mao LN, Han SJ, Sun D, Zheng ZG. Multilayered control of retinal stem/progenitor cell fate in the single-cell and organoid era: Developmental blueprints and regenerative opportunities. World J Stem Cells 2026; 18(4): 118621
- URL: https://www.wjgnet.com/1948-0210/full/v18/i4/118621.htm
- DOI: https://dx.doi.org/10.4252/wjsc.v18.i4.118621