TO THE EDITOR
Hepatocellular carcinoma (HCC) remains a formidable foe in the global cancer landscape, ranking as the third leading cause of cancer-related mortality worldwide[1]. Its aggressiveness is notoriously fueled not just by the cancer cells themselves, but by the environment they create – the tumor immune microenvironment[2,3]. Within this corrupted landscape, a specific group of immune cells, known as tumor-associated macrophages (TAM), are coerced into becoming traitors[4]. Typically exhibiting an “M2-like” polarization, these macrophages abandon their tumor-fighting duties and instead secrete a cocktail of factors that promote cancer growth[5], relapse[6], and metastasis[7]. This hijacking of the body's innate defenses is a key reason why current therapies, including some immunotherapies, often show limited effectiveness[8].
A new study published in the World Journal of Gastroenterology by Escobedo-Calvario et al[9] now points to a promising new strategy that targets this very weakness. The research focuses on growth differentiation factor 11 (GDF11), a member of the transforming growth factor-beta superfamily[10,11]. The superfamily’s notorious relative transforming growth factor-beta, which primarily signals through the TβRII/ALK5 receptor complex and leads to the phosphorylation of Smad2/3, sustained ALK5 signaling often drives pro-fibrotic and pro-metastatic gene expression in advanced cancers[12]. In contrast, GDF11, which signals through ActRIIB and likely utilizes different anaplastic lymphoma kinase (ALK) isoforms (e.g., ALK4, ALK5, ALK7), can lead to a distinct phospho-Smad complex that regulates a different set of target genes, which possess anti-tumor effects on HCC cells[13]. This new work, however, reveals a more profound and exciting dimension: GDF11 doesn't just attack the cancer cells, it effectively “re-educates” the pro-tumoral M2 macrophages, disarming them and turning their secretions from a cancer fuel into a cancer-suppressing agent. This dual-pronged mechanism could represent a paradigm shift in how we approach liver cancer therapy.
APPROACH: MODELING THE TUMOR MICROENVIRONMENT IN A DISH
To investigate the mechanism behind GDF11’s anti-tumoral effects, the research team led by Escobedo-Calvario et al[9] proposed a compelling “two-axis” hypothesis. They posited that GDF11’s power extends beyond directly suppressing HCC cells to also neutralizing key pro-tumoral forces within the tumor immune microenvironment, specifically the M2-polarized TAMs.
This focus is critical due to the opposing roles of macrophage subsets[14]. Classically activated M1 macrophages typically secrete immunostimulatory cytokines (e.g., interferon-gamma) and combat tumors. In contrast, alternatively activated M2 macrophages, polarized by factors like interleukin (IL)-4 and IL-13, adopt an immunosuppressive and pro-tumoral profile, secreting factors such as IL-10 that promote cancer growth and aggression by fostering an immune suppressive microenvironment[15].
To test this dual-targeting hypothesis, the researchers established a well-established in vitro model. They used THP-1 human monocytes, differentiating them first into neutral macrophages (M0) and then polarizing them into M2-like macrophages using IL-4 and IL-13, effectively recreating a key component of the immunosuppressive tumor niche. The human Huh7 cell line was used to model HCC. The experimental strategy was two-pronged: (1) To characterize the direct effects of GDF11 (50 ng/mL, three doses every 24 hours) on the biology of M2-like macrophages; and (2) To determine how these GDF11-induced changes subsequently impact the proliferation and migratory capacity of the liver cancer cells.
MECHANISM: A STORY OF METABOLIC REWIRING AND RE-EDUCATION
The results of the study provide a compelling narrative of how GDF11 reprograms pro-tumoral macrophages. The first crucial finding was that GDF11 successfully activates the canonical Smad2/3 signaling pathway in macrophages, confirming the cells are receptive to its signal. Importantly, this activation occurred without inducing cell death or affecting proliferation, suggesting GDF11’s action is modulatory rather than cytotoxic – a desirable trait for a potential therapeutic that aims to retrain, not kill, immune cells.
A key marker of pro-tumoral M2 macrophages is a surface protein called CD206, a member of the group 6, C-type lectin receptor family[16]. While the precise mechanism by which CD206 promotes immunosuppression is not fully elucidated, its known role in pathogen recognition and endocytosis suggests a hypothesis: Within the tumor microenvironment, CD206 may facilitate the uptake of cellular debris (e.g., apoptotic cells)[17,18]. This process can activate tissue remodeling and immunoregulatory pathways that suppress anti-tumor immunity[19]. Although this function is essential for normal tissue homeostasis, in cancer it is co-opted to support tumor growth. Supporting this view, inhibitors of CD206 have been shown to reprogram macrophages from a pro-tumorigenic M2 state toward a more inflammatory, tumor-attacking M1 phenotype.
The researchers demonstrated that GDF11 treatment significantly reduced the percentage of CD206+ cells. In an extended treatment protocol, the effect was even more dramatic, reducing CD206+ subsets from 90% to 30%. This indicates that GDF11 actively impairs the M2 polarization by stripping away the M2 macrophages’ badge of aggressiveness, essentially forcing these macrophages to shed their pro-tumoral identity.
M2 macrophages are known for their distinct metabolic profile, which often involves reduced mitochondrial function[20]. This study found that GDF11 treatment “rewired” this metabolism, increasing the oxygen consumption rate and restoring mitochondrial activity to levels resembling neutral or even anti-tumoral macrophages. Furthermore, GDF11 triggered a significant increase in the production of reactive oxygen species. While high reactive oxygen species can be damaging, in the context of immune cells, it is a critical weapon for attacking pathogens and, potentially, cancer cells. This GDF11-induced reactive oxygen specie boost suggests it may be re-arming the macrophages for anti-tumoral activity.
The most compelling evidence came from experiments using conditioned media (CM), which directly connected GDF11’s action on macrophages to its subsequent effect on cancer cells. As anticipated, CM from untreated M2 macrophages dramatically enhanced the migration and proliferation of Huh7 cells. In stark contrast, CM harvested from GDF11-treated M2 macrophages completely neutralized this pro-tumoral effect. Remarkably, the cancer cells lost their enhanced capacity to migrate and close wounds in the lab assays. This proved that the changes GDF11 induced in the macrophages functionally translated into a less aggressive environment for the cancer cells. Crucially, the team meticulously ruled out the possibility that traces of the GDF11 protein itself in the CM were causing this effect, confirming that the shift was due to the altered secretome of the re-educated macrophages.
To understand what caused this functional shift, the researchers analyzed the cytokine profile in the CM. They found that GDF11 treatment downregulated key pro-tumoral factors like IL-6[21], ENA-78 (CXC motif chemokine ligand 5)[22], and angiogenin[23], all of which are known to promote cancer growth, invasion, and blood vessel formation. Simultaneously, there was an increase in a mix of other chemokines and growth factors, indicating a complex reprogramming of the macrophage secretome away from a pro-tumoral and toward a more neutral or even pro-inflammatory state.
TRANSLATIONAL INSIGHTS: A NEW AVENUE FOR COMBINATION THERAPY
This study positions GDF11 as a master regulator with a unique dual function. It directly targets cancer cells, as shown in their previous work[13], and simultaneously neutralizes one of the tumor's most powerful allies – the M2 macrophage. By rewiring macrophage metabolism, reducing their pro-tumoral markers, increasing their anti-tumor reactive oxygen species production, and altering their secretome, Therefore, GDF11 effectively dismantles a key support pillar of the tumor microenvironment.
The implications for liver cancer treatment are significant. The limited success of some immunotherapies in HCC, particularly in cases driven by metabolic dysfunction, is often due to the immunosuppressive nature of the tumor microenvironment, largely orchestrated by TAMs[24]. A therapeutic agent like GDF11, which can counteract this immunosuppression and “re-educate” TAMs, could be powerfully synergistic. It could be used to precondition the tumor microenvironment, making it more susceptible to subsequent immunotherapy, potentially breaking the cycle of treatment resistance.
THE LIMITATIONS OF IN VITRO MODELS IN RECAPITULATING MACROPHAGE HETEROGENEITY AND PLASTICITY
While the study by Escobedo-Calvario et al[9] presents the exciting potential of using GDF11 to simultaneously reprogram macrophages toward an immunostimulatory state and hinder cancer cell invasion, the in vitro system used in this study inherently simplifies the profound heterogeneity and plasticity of macrophages in vivo. The research relies on models like the THP-1 human monocyte cell line, which cannot fully capture the complexity of the liver macrophage niche. This niche is predominantly composed of Kupffer cells, tissue-resident macrophages of embryonic origin[25]. In HCC, while most TAM are bone marrow-derived, lineage-tracing studies have confirmed that embryonic-derived Kupffer cells can also infiltrate tumors and exhibit significant phenotypic plasticity, particularly when monocyte recruitment is impaired[26].
Furthermore, the tumor microenvironment comprises a complex ecosystem of immune cells, including T cells, natural killer cells, and myeloid-derived suppressor cells. The effect of GDF11 on these populations remains unknown and warrants investigation in preclinical models and human samples. It would be particularly valuable to correlate GDF11 expression with TAM phenotypes and patient prognosis. Consequently, while informative, the in vitro macrophage model used in this study cannot capture the full complexity of cellular interactions and the dynamic adaptability of macrophage subsets within the actual tumor microenvironment.
UNRESOLVED MECHANISMS OF GDF11 ACTION ON MACROPHAGE REPROGRAMMING
This study by Escobedo-Calvario et al[9] identifies a significant phenomenon: GDF11 reprograms macrophages by reducing the expression of CD206, a defining M2 marker with direct immunosuppressive functions. While this finding is compelling, it opens a critical gap in our knowledge. The mechanism by which an extracellular ligand like GDF11 leads to the specific downregulation of a cell surface receptor of target cell remains elusive. Indeed, the specific transcriptional regulators, signaling intermediates, and potential epigenetic modifications involved are currently a “black box”. Key questions about the involved intracellular signaling circuits, potential crosstalk with other pathways, and resultant changes in gene expression are unanswered. Therefore, future work leveraging multi-omics approaches, such as whole transcriptomic (e.g., single cell or bulk RNA-seq) and epigenetic profiling (e.g., ATAC-seq and CUT and Tag-seq) of GDF11-treated macrophages would provide indispensable insights into the gene regulatory networks that underpin this phenotypic reprogramming[27].
CONCLUSION
In conclusion, the study conducted by Escobedo-Calvario et al[9] elucidates the dual function of GDF11. Compared with traditional macrophage targeting therapy, GDF11 can be utilized to re-educate pro-tumoral M2 macrophages. This finding signifies a paradigm shift from merely inhibiting macrophages to actively reprogramming them.
Another intriguing perspective emerging from this study is the potential of GDF11 – a factor known for its rejuvenating effects in aged tissues[28-30] – to reprogram, rather than merely inhibit, TAMs in HCC. This connects the findings of this study to the burgeoning field of cellular senescence within the tumor microenvironment[31-34]. Accumulating evidence indicates that senescent immune cells, including T cells[35,36] and natural killer cells[37,38], accumulate in tumors and exhibit functional decline, characterized by the upregulation of stress-related genes and a concomitant loss of cytotoxic activity[39].
The role of macrophage senescence is more complex and less defined. While macrophages can clear senescent cells[40], this interaction can sometimes induce an immunosuppressive state in the macrophages themselves[41]. Furthermore, the senescence-associated secretory phenotype from other cells can recruit and polarize macrophages toward a pro-tumoral M2 phenotype, creating a vicious cycle[42]. The critical questions of whether TAMs themselves undergo senescence and whether targeting this state could be a viable therapeutic strategy remain largely unanswered. This study provides a compelling angle by demonstrating that a known rejuvenation factor can redirect macrophages from a pro-tumor to an anti-tumor state. This intriguing concept of “rejuvenation therapy” for the immune microenvironment warrants extensive future investigation (Figure 1).
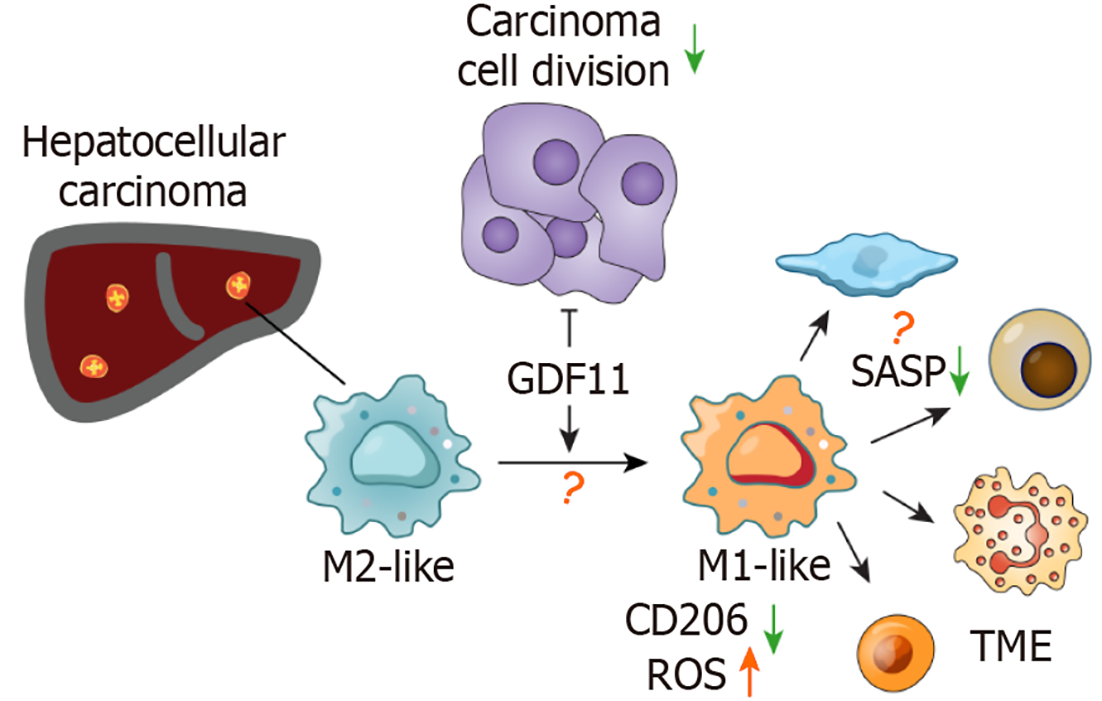
Figure 1 This intriguing concept of “rejuvenation therapy” for the immune microenvironment warrants extensive future investigation.
In the immunosuppressive tumor microenvironment, pro-tumoral M2-like macrophages promote cancer cell proliferation. growth differentiation factor 11 counteracts this by directly suppressing tumor growth and simultaneously reprogramming M2-like macrophages toward an anti-tumoral M1-like state. This reprogramming is characterized by the downregulation of the M2 marker CD206 and an increase in reactive oxygen species. Future investigations will determine the roles of underlying metabolic rewiring, epigenetic reprogramming, and the potential for growth differentiation factor 11 to suppress a pro-tumoral senescence-associated secretory phenotype. GDF11: Growth differentiation factor 11; ROS: Reactive oxygen species; SASP: Senescence-associated secretory phenotype; TME: Tumor microenvironment.
Provenance and peer review: Invited article; Externally peer reviewed.
Peer-review model: Single blind
Specialty type: Gastroenterology and hepatology
Country of origin: China
Peer-review report’s classification
Scientific Quality: Grade A, Grade B, Grade B
Novelty: Grade A, Grade B, Grade C
Creativity or Innovation: Grade A, Grade B, Grade C
Scientific Significance: Grade A, Grade A, Grade B
Open Access: This article is an open-access article that was selected by an in-house editor and fully peer-reviewed by external reviewers. It is distributed in accordance with the Creative Commons Attribution NonCommercial (CC BY-NC 4.0) license, which permits others to distribute, remix, adapt, build upon this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See: https://creativecommons.org/Licenses/by-nc/4.0/
P-Reviewer: Ivanga M, PhD, Gabon; You R, Associate Chief Physician, China S-Editor: Luo ML L-Editor: A P-Editor: Zhang L