Published online Mar 21, 2026. doi: 10.3748/wjg.v32.i11.115393
Revised: November 22, 2025
Accepted: December 30, 2025
Published online: March 21, 2026
Processing time: 150 Days and 8.4 Hours
Tumor metastasis is the main cause of death in patients with colorectal cancer (CRC). Despite the rapid development of new therapeutic methods and drugs in recent years, the survival rate of patients with metastatic CRC has not improved significantly. Proteins play pivotal roles in CRC development and progression. Thus, studying their expression profiles and regulatory mechanisms in CRC may facilitate the development of novel diagnostic and therapeutic strategies.
To explore the role and regulatory mechanism of LIM domain only protein 7 (LMO7) in CRC metastasis.
We screened differentially expressed proteins associated with CRC through pro
Data-independent acquisition proteomic analysis of 78 paired CRC tissues and adjacent normal tissues identified 1144 differentially expressed proteins, among which LMO7 was significantly upregulated in CRC. We found that the expression of LMO7 in CRC tissues was increased and correlated with the clinical stage of CRC. Additionally, compared with that in primary colon cancer sites or negative lymph nodes, the protein expression of LMO7 in metastatic lymph nodes was greater. LMO7 promoted CRC cell proliferation, migration, and invasion, as well as the growth and metastasis of CRC tumors. Transcriptome sequencing revealed that LMO7 is associated with cell migration and EMT. We also confirmed that LMO7 promoted the EMT ability of CRC cells by regulating the expression of E-cadherin, N-cadherin, and vimentin. Furthermore, we found that LMO7 could bind to zinc finger E-box binding homeobox 1 (ZEB1) and regulate the expression of LMO7 through ZEB1. Moreover, induction with transforming growth factor (TGF)-β led to increased expression of ZEB1 and LMO7 proteins in CRC cells, and knockdown of ZEB1 attenuated this induction effect of TGF-β on LMO7 protein expression.
This study demonstrated that LMO7 facilitates EMT, as well as cell migration and invasion, ultimately driving the progression and metastasis of CRC cells. Furthermore, the TGF-β/ZEB1 signaling pathway positively regulates LMO7 expression in CRC.
Core Tip: LIM domain only protein 7 (LMO7) is upregulated in colorectal cancer (CRC), and is correlated with clinical stage and metastasis. It promotes CRC cell proliferation, migration, invasion, epithelial-mesenchymal transition and tumor metastasis. transforming growth factor-β induces LMO7 expression by upregulating zinc finger E-box binding homeobox 1, which binds to the LMO7 promoter, forming a transforming growth factor-β/zinc finger E-box binding homeobox 1/LMO7 axis that drives CRC progression, highlighting LMO7 as a potential therapeutic target for CRC.
- Citation: Bai X, Zheng B, Li XM, Wang JD, Huang XH, Fu SM, Chen X. LIM domain only protein 7 promotes metastasis in colorectal cancer by TGF-β/ZEB1 pathway. World J Gastroenterol 2026; 32(11): 115393
- URL: https://www.wjgnet.com/1007-9327/full/v32/i11/115393.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i11.115393
Colorectal cancer (CRC) is a common gastrointestinal malignant tumor that accounts for approximately 9.4% (the second most common cause) of cancer-related deaths worldwide. The global incidence of CRC is expected to increase to 2.5 million new cases by 2035[1]. CRC incidence and mortality rates are still rising rapidly in many low- and middle-income countries, whereas trends are stable or declining in highly developed countries[2]. Although in the United States, the incidence of CRC has shown a decreasing trend of 3.3% annually among patients aged 65 years and above and an increasing trend of 2.2% for early-onset or young CRC among those under 50 years, this age-related trend has not been observed in China[3].
The prognosis of CRC is related mainly to tumor stage and metastasis. Approximately 20% of CRC patients present with distant metastasis (stage IV) at the time of diagnosis. Furthermore, 35%-45% of patients with local lesions (stage II and stage III) experience recurrence within 5 years after surgery, and the majority of these recurrences occur in the form of metastasis[4]. CRC and its metastasis are correlated with early molecular alterations during the tumor progression process, which also determine its metastatic ability. Existing studies have revealed that cellular signaling pathways, such as the Wnt/β-catenin, p53, transforming growth factor (TGF)-β/SMAD, nuclear factor kappa B, Notch, vascular en
Metastasis is driven predominantly by the dissemination of cancer cells that have shed from the primary tumor and are circulating in the bloodstream, namely, the transformation of epithelial tumor cells into circulating tumor cells. This involves a series of biochemical and cellular alterations to facilitate migration, invasion, resistance to apoptosis, and regulation of extracellular matrix production. The epithelial-mesenchymal transition (EMT) plays a critical role in promoting this process. A variety of oncogenic factors can induce EMT, including the TGF-β, Wnt, and Notch signaling pathways, and the activation of proteins such as zinc finger E-box binding homeobox 1 (ZEB1), TWIST, and Snail. These genetic and phenotypic changes ultimately promote the migration of cancer cells[5]. Although favorable clinical res
Deep proteome profiling has become a reliable and essential tool for providing highly valuable and clinically relevant real-time information at all stages of cancer progression[6]. In this study, we used in-depth proteomics to identify differentially expressed proteins in 78 pairs of CRC and adjacent normal tissue samples. We screened functional differential proteins, confirmed the clinical significance and biological role of differential protein LIM domain only protein 7 (LMO7) in the development of CRC, and further explored its expression regulatory mechanisms.
The tissue samples for proteomics and validation were obtained from CRC tissues and paired adjacent tissues procured from patients during endoscopic examinations. A total of 80 cases for proteomic analysis were collected, among which two cases were excluded because of substandard proteomic detection quality. A total of 78 cases for validation were included, with 3 pieces of tissue taken from each case. All cancer tissue samples were pathologically diagnosed as adenocarcinoma. All the enrolled patients had no history of other tumors and had not received any antitumor treatment before enrollment. All patients signed informed consent forms. Sample collection and research were approved by the Ethics Committee of the First Affiliated Hospital of Chengdu Medical College (No. 2023CYFYIRB-BA-Feb001). The tissue microarrays were purchased from Shanghai Xinchao Biotechnology Co., Ltd.
The proteomic analysis was adapted from Dong et al[7]. All fraction samples were analyzed by mass spectrometry in data-independent acquisition (DIA) mode. The spectral library was generated using Spectronaut 14.2 (Biognosys) on the basis of the UNIPROT human database (reviewed entries only). Virtual spectra were generated by matching peptide ions and fragment ions using DIA-Umpire. If three peptides or a unique peptide of a certain protein were identified, that protein was considered to be identified. After the fold difference and P value were calculated through the MSstats software package for the t-test, a fold change ≥ 1.5 and P < 0.05 were used as the screening criteria for significantly differentially expressed proteins.
All the BALB/c-nu mice were male, 4-6 weeks old, weighed 15-18 g, and were purchased from Chengdu Yaoke Biotechnology Co., Ltd. The mice were housed under standard conditions at 24 ± 2 °C with food and water provided. After a 1-week acclimation period, the groups were assigned using simple random sampling. A random sequence was generated via a random number table, and the mice were sequentially allocated to the following groups: (1) Negative control [empty vector-transfected cells, cells transfected with the black vector (CTL)] SW480 cells; (2) LMO7 knockdown (sh-LMO7) SW480 cells; (3) Negative control (empty vector-transfected cells, CTL) LoVo cells; and (4) LMO7-knockdown (sh-LMO7) LoVo cells. All animal experimental protocols were approved by the animal experimental teaching and research commi
The cells used in this study were normal human intestinal epithelial cell lines (HIECs) and the colon adenocarcinoma cell lines HT29, Caco2, HCT116, SW480, and LoVo (metastatic tumor nodules from the left clavicle region of a male colon adenocarcinoma patient). The cell lines to be transfected were seeded in six-well plates and cultured until the cell density reached 30%-50%. Afterward, 2 mL of virus and 1 mL of culture medium were added, and the mixture was incubated overnight. After 48 hours, LMO7-overexpressing and LMO7-knockdown plasmid (sh-LMO7-1, sh-LMO7-2)-transfected cells were selected with 2 µg/mL puromycin, ZEB1-overexpressing cells were selected with 1000 µg/mL neomycin, and ZEB1-knockdown plasmid (sh-ZEB1-1, sh-ZEB1-2)-transfected cells were selected with 500 µg/mL hygromycin. The selection process lasted for 7-10 days.
A total of 1000-2000 cells were inoculated into each well of a 96-well plate, with 5 plates for each cell type and 200 μL of culture medium in each well. The cells were incubated at 37 °C and 5% CO2. After the cells adhered to the wall, cholecystokinin-8 (CCK-8) solution was added, and the plates were incubated for 1 hour. The OD value at a wavelength of 450 nm was subsequently measured. The above steps were repeated for 3-4 days to generate a proliferation curve.
The cells were seeded into 24-well culture plates at a density of approximately 1 × 105 per cells well. After the cells reached confluence, a 20 μL pipette tip was used to create a scratch. Twenty-four and 36 hours later, the width of the scratch was observed under a microscope and photographed.
The diluted Matrigel in serum-free medium was evenly coated on the upper side of the polycarbonate membrane of the Transwell chamber and placed in a 37 °C incubator for 1 hour. A total of 100 μL (approximately 1 × 105 cells per well) of the cell suspension was seeded in the upper chamber of the Transwell system, and 600 μL of medium containing 10% fetal bovine serum was added to the lower chamber. After 24-48 hours, the chamber was removed, the cells and matrix gel on the upper chamber were wiped off, and the chamber was fixed with formaldehyde for 30 minutes and then stained with crystal violet for 10 minutes. The cells were observed under a microscope, and 5 random fields were photographed. The average number of cells that passed through the membrane was calculated to evaluate the invasion ability of the cells.
The cells (6 × 106) were subcutaneously injected into the backs of the nude mice. Seven nude mice were included in each group. The tumor size was measured every 4 days, and the tumor volume was calculated as follows: Tumor volume = (length × width2)/2. After 42 days of subcutaneous injection of SW480 cells and 21 days of injection of LoVo cells, the mice were euthanized, and the tumors were removed and photographed. The data were plotted using GraphPad. The endpoint of the experiment was when the tumor volume reached 1500-2000 mm³, or the nude mice showed a 15% decrease in body weight.
For the lung metastasis experiment, 2 × 106 cells were injected into the tail vein of nude mice to evaluate their metastatic ability. Seven nude mice were included in each group. After 58 days of SW480 cell injection and 60 days of LoVo cell injection, the mice were euthanized, and the lung tissues were removed. The number of lung tumor metastases was calculated. The endpoint of the experiment was when the nude mice showed obvious anxiety, restlessness, and a 15% decrease in body weight.
As previously described[7], RNA sequencing was performed using SW480 cells and LMO7-knockdown SW480 cells. Functional enrichment analyses, including clustering, Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Ge
Immunohistochemistry was performed as previously described[8]. The obtained tissue was fixed in 4% paraformaldehyde, followed by dehydration, clearing, and paraffin embedding at 60 °C. The sections were then dewaxed and rehydrated, followed by antigen retrieval in citrate buffer (pH 6) and blocking with 3% hydrogen peroxide followed by bovine serum albumin. The sections were incubated with primary antibody (1:500-1:1000 dilution) overnight at 4 °C. Secondary antibody (HRP-labeled) was incubated at room temperature, followed by DAB staining and hematoxylin counterstaining. The sections were subjected to graded dehydration, and mounted with neutral resin for sealing.
Proteins were extracted using nuclear and cytoplasmic extraction reagents according to the manufacturer’s instructions. The protein concentration was determined by the BCA method. The proteins were separated by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis and transferred to a polyvinylidene difluoride membrane. The membrane was incubated in bovine serum albumin blocking solution (1 hour, room temperature) and then incubated overnight with antibodies against LMO7 (Sigma), ZEB1 (CST), E-cadherin (CST), N-cadherin (CST), and vimentin (CST) at 4 °C. After the membrane was washed three times with Tris-buffered saline containing Tween (10 minutes each), the secondary antibody was added, and the mixture was incubated at room temperature for 1 hour. A freshly prepared ECL solution was added to the protein side of the membrane, which was exposed in a dark room and then scanned.
Total RNA was extracted by a total RNA extraction kit (Beyotime Biotechnology) according to the manufacturer’s protocol. Genomic cDNA eraser and cDNA synthesis (Beyotime Biotechnology) were also performed following the manufacturers’ protocols. The mRNA levels of LMO7 were determined by a real-time PCR system (Life Technologies). The primers used for the amplification of the LMO7 gene were as follows: (1) Forward, 5′-TGTTGCCTGTGAGTGTGAC-3′; and (2) Reverse, 5′-ACAGTGCTTTCGTATGGAGG-3′. The results were repeated in at least three independent RNA preparations. The expression levels were normalized to those of glyceraldehyde-3-phosphate dehydrogenase, which was used as an internal control gene.
Formaldehyde was used to cross-link DNA with DNA-binding proteins in 293T cells. The cells were resuspended in 500 μL of cell lysis buffer and disrupted by ultrasonic treatment. The cell lysate was subjected to ultrasonic treatment to break down double-stranded DNA under the following conditions: (1) 15% ultrasonic power, 4 °C; and (2) On 2 seconds/off 4 seconds, for a total of 5 minutes. Then, 250 μL of the supernatant was incubated with 5 μL of ZEB1 antibody or 5 μL of IgG antibody at 4 °C overnight. Activated magnetic beads were added to the samples and incubated at 4 °C for 1 hour. After eluting and decrosslinking the protein-DNA complexes, the DNA was purified using a DNA purification kit. The precipitated samples were subsequently subjected to qPCR detection.
A luciferase reporter assay was performed as previously described[9]. The pGL3-LMO7-promoter vector was constructed by ELK Biotechnology Co., Ltd. The internal reference gene plasmid carried Renilla luciferase (pRL-TK). A t-test was used for statistical analysis to compare the differences in luciferase activity.
TGF-β pathway intervention experiments were performed and the expression levels of ZEB1 and LMO7 were assessed. SW480/LoVo cells were treated with 0 ng/mL, 5 ng/mL, 10 ng/mL, or 20 ng/mL TGF-β; real-time reverse-transcriptase PCR was used to detect the LMO7 mRNA level to determine the optimal intervention concentration of TGF-β. Wild-type and ZEB1-knockdown (Sh-ZEB1) SW480/LoVo cells were then treated with this concentration of TGF-β, and western blotting was performed to assess ZEB1/LMO7 protein levels.
Experimental data in this study were analyzed using Statistical Package for the Social Sciences 19.0 and GraphPad Prism 8.0. Some figures were generated using GraphPad Prism 8.0 tools. Normally distributed quantitative data are presented as mean ± SD, while median and interquartile range represent non-normally distributed quantitative data. The t-tests were used to analyze two normally distributed groups of quantitative data. Analysis of variance was used for three groups. The Wilcoxon signed-rank test was applied for two groups with non-normal distribution, and the Kruskal-Wallis test for three groups. Count data were analyzed using χ² tests. Spearman correlation analysis was used to assess relationships. Results were considered statistically significant at P < 0.05. All mRNA and protein expression levels were quantified using the 2-ΔΔCt method to calculate relative expression, expressed as dimensionless relative ratios (normal colonic epithelial cells/tissues were set as the reference group, defined as 1.00). Quantitative data are presented as mean ± SD, with error bars represented as “± SD”. Intergroup differences were assessed using t-tests or one-way analysis of variance. Significance thresholds were uniformly defined as: (1) P < 0.05 (significant, denoted by “aP”); and (2) P < 0.01 (highly significant, denoted by “bP”). All cell line experiments were performed in triplicate by default, and transcriptomic sequencing included three replicate samples per group. Real-time reverse-transcriptase PCR, Western blot, CCK-8 proliferation assays, scratch assays, Transwell invasion assays, chromatin immunoprecipitation analysis, dual-luciferase reporter assays, and TGF-β pathway intervention experiments were each performed in triplicate.
A total of 78 samples from CRC patients (31 females and 47 males) were included in this proteomics analysis. Only 59 patients underwent surgery and had complete clinical data available for follow-up. Among the 59 patients, 4 were in stage I, 23 were in stage II, 19 were in stage III, and 13 were in stage IV. CRC proteomics identified 1144 differentially expressed proteins, of which 577 proteins were upregulated and 567 proteins were downregulated. Further analysis revealed that the difference in LMO7 protein expression was 2.16 (cancer tissue vs paracarcinoma tissue). LMO7 was stably expressed in all the CRC tissue samples, and the KEGG pathway was enriched in adherens junctions.
We analyzed the differential expression of LMO7 in two public databases containing clinical characteristics and gene expression data. The results revealed that LMO7 protein levels were higher in CRC than in other cancer types in the Human Protein Atlas (www.proteinatlas.org) HPA020923 dataset. In addition, the R package was used to analyze the expression of LMO7 in the TCGA database in CRC and its correlation with the clinical features of CRC. TCGA data revealed that LMO7 expression was higher in CRC tissues than in paired paracarcinoma tissues (Supplementary Figure 1A). Clinical correlation analysis revealed that high LMO7 expression was positively correlated with CRC stage, N stage, and M stage (Supplementary Figure 1B-D). We then analyzed the correlation between the relative abundance of LMO7 expression and different clinical stages in 59 CRC patients whose complete clinical data were available. LMO7 expression was positively correlated with CRC TNM stage, and the LMO7 expression level was significantly greater in stage IV CRC than in other stages. Moreover, the expression level was positively correlated with the N and M stages (P < 0.05; Supple
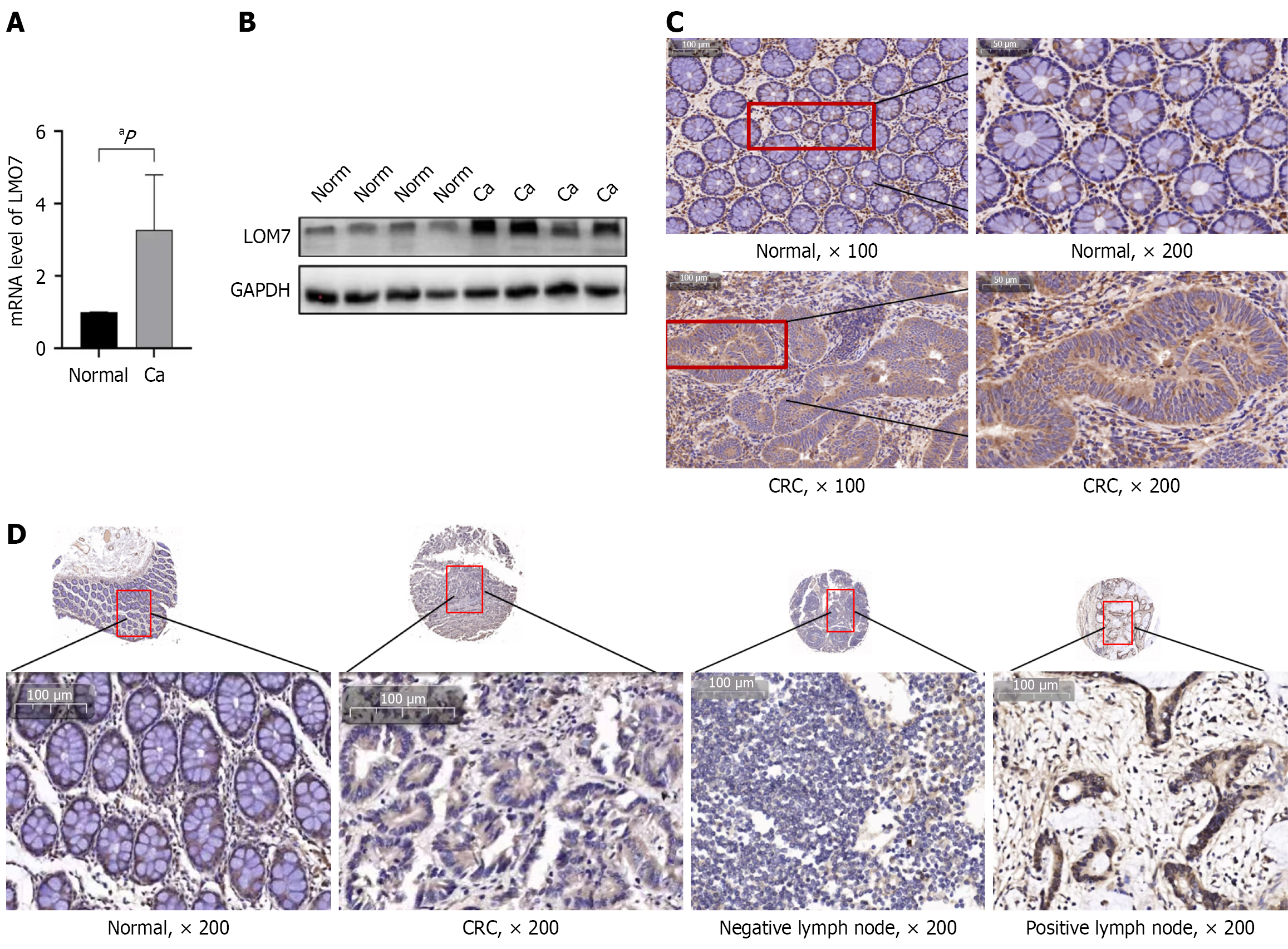
Next, the expression of LMO7 mRNA and protein in CRC and paracarcinoma normal tissues was assessed. Compared with those in paired normal tissues, LMO7 mRNA and protein expression in cancer tissues was significantly increased (Figure 1A and B). Immunohistochemical results further verified that the expression of LMO7 in CRC tissues was significantly increased. LMO7 is expressed mainly in the cytoplasm but is also positively expressed in the membrane (Figure 1C). To further investigate the expression of LMO7 in metastatic tissues, we also assessed the expression of LMO7 protein in CRC tissue chips with metastatic lymph nodes, and the results revealed that the expression of LMO7 protein in colon cancer tissues was increased compared with that in normal colon tissues. LMO7 expression in metastatic positive lymph nodes was greater than that in primary colon cancer and significantly greater than that in negative lymph nodes (Figure 1D). These results indicate that LMO7 is highly expressed in CRC tissues and may be associated with CRC metastasis. However, LMO7 is primarily localized in the cytoplasm of both normal intestinal tissues and primary CRC lesions, whereas it is significantly enriched in the nuclei of metastasis-positive lymph nodes (Figure 1C and D).
To investigate the potential role of the LMO7 gene in CRC, we first assessed the expression of LMO7 mRNA and protein in a normal colon epithelial cell line (HIEC) and five colon cancer epithelial cell lines: CaCo2, HCT116, SW480, LoVo, and HT29. Compared with those in HIECs, LMO7 mRNA and protein expression in CaCo2, SW480, LoVo, and HT29 cells was increased, and LMO7 protein expression in SW480 and LoVo cells was significantly increased. The expression of LMO7 mRNA in the HCT116 cell line was not significantly different from that in HIECs (Supplementary Figure 3).
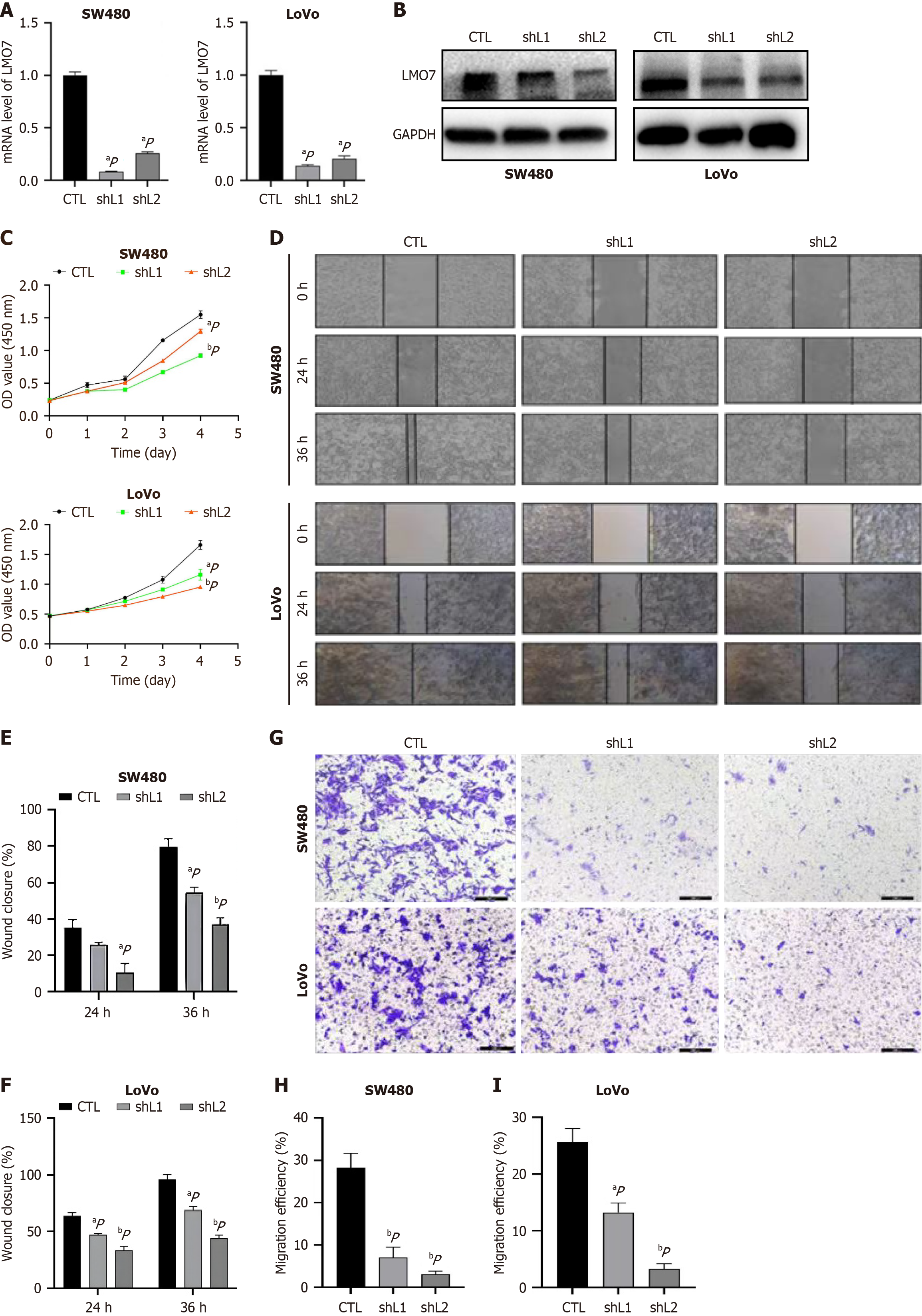
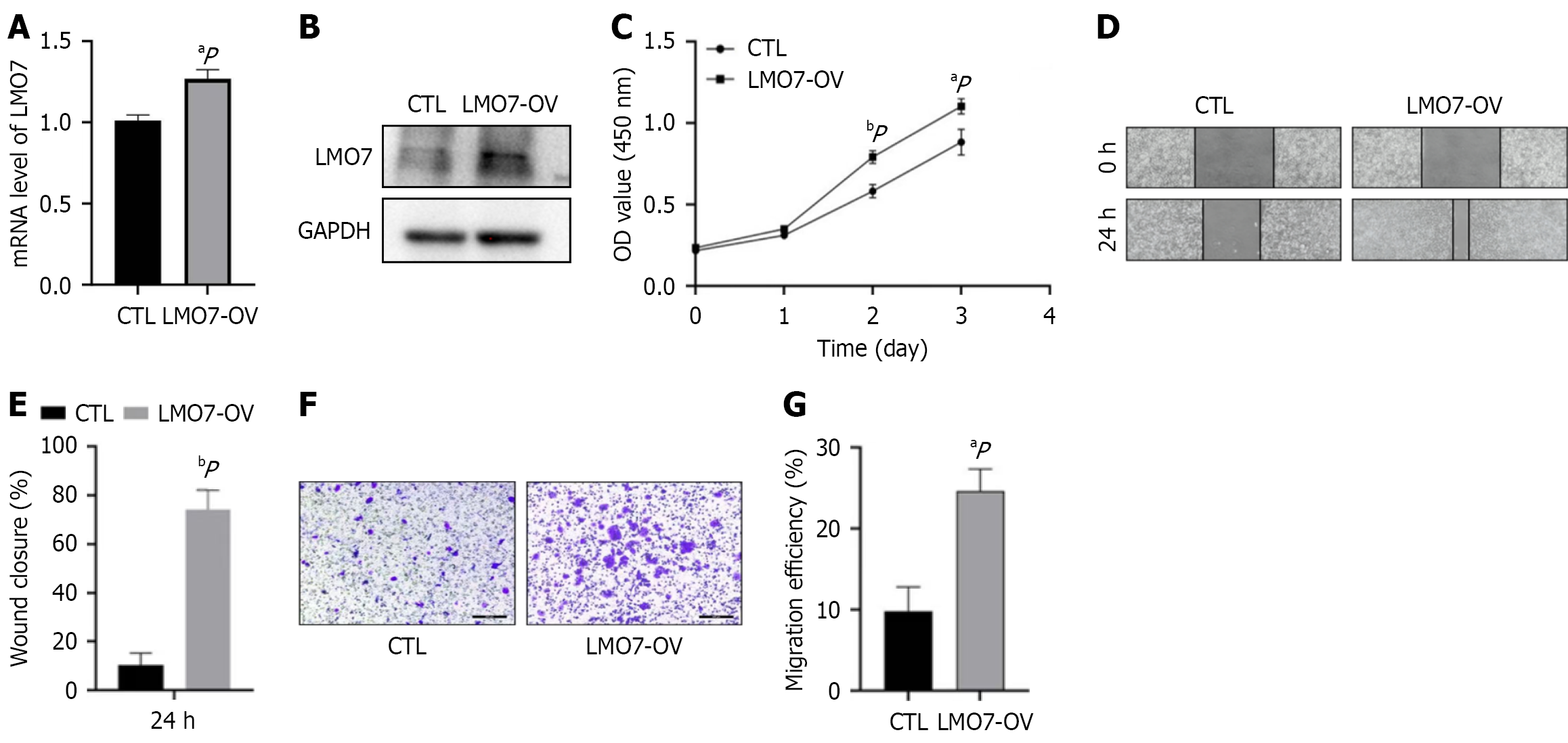
To explore the potential role of the LMO7 gene in CRC, LMO7-knockdown (cells transfected with sh-ZEB1-1 and cells transfected with sh-ZEB1-2) SW480, LoVo (Figure 2A and B) and LMO7-overexpressing HCT116 cell lines were con
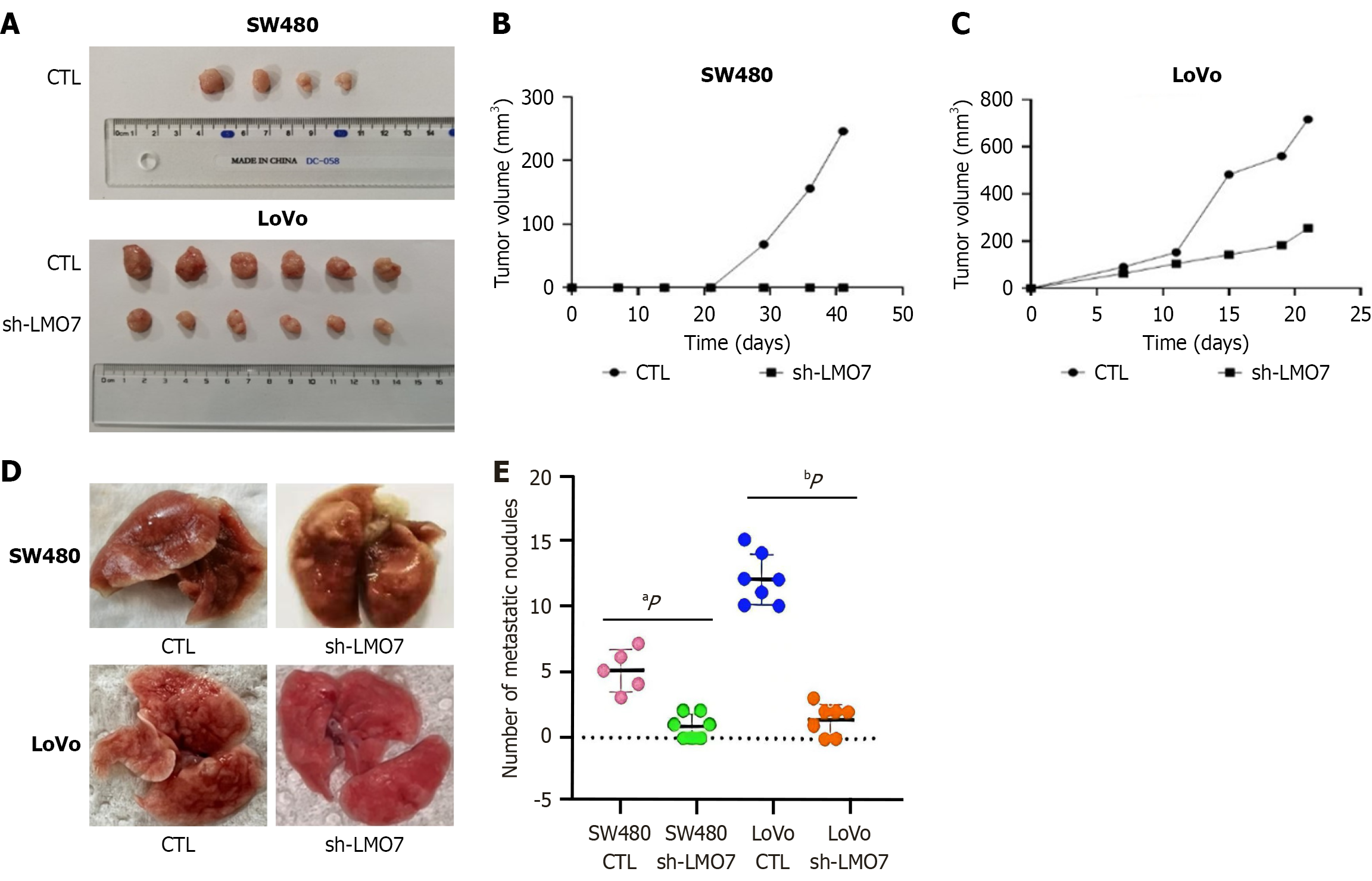
Following in vitro validation that LMO7 promotes the proliferation, migration, and invasion of CRC cells, we next investigated whether LMO7 affects the proliferation and metastasis of CRC cells in vivo. As shown in Figure 4, 42 days after the subcutaneous injection of SW480 cells, 4 out of the 7 nude mice injected with the blank vector exhibited tumor formation, and 1 died, whereas no subcutaneous tumor formation was observed in the mice with LMO7 knockdown. After 21 days of subcutaneous injection of LoVo cells, the subcutaneous tumor volume of some control mice (CTL, injected with CTL) reached the experimental end point. All mice with LMO7 knockdown (sh-LMO7, injected with cells transfected with sh-LMO7-1) had subcutaneous tumor formation, but the tumor volume was significantly reduced. These results demonstrate that inhibiting the expression of LMO7 can suppress the development of CRC. Additionally, the tumorigenicity of LoVo cells was greater than that of SW480 cells; thus, LoVo cells are more suitable for subcutaneous tumorigenesis experiments.
Next, we evaluated the role of LMO7 in lung metastasis in nude mice. White nodules were observed in both lungs of the mice injected with blank vector-transfected SW480 cells (CTLs), but white nodules were not obvious in the lungs of the LMO7-knockdown mice, and two mice presented no nodular masses in either lung (Figure 4D and E). Sixty days after the tail vein injection of LoVo cells into the mice, multiple white nodules were visible in the lungs of the mice (CTL), with many more nodules than in the lungs of the mice injected with SW480 cells. In contrast, the lungs of the LMO7-knock
Transcriptome sequencing was performed to further investigate the biological functions of LMO7 in CRC and provide robust evidence for the subsequent exploration of its specific mechanism of action. There were 393 DEGs, including 122 upregulated genes (31%) and 271 downregulated genes (67%). DEGs with the same or similar expression patterns were clustered, and the clustering results of the DEGs are shown in Supplementary Figure 4A and B. The top 50 DEGs were all downregulated, and LMO7 ranked third. GO analysis revealed that cell junctions can be enriched in cell components, and genes related to molecular function can be enriched in binding, indicating that LMO7 can affect cell structure, junctions, and adhesion (Supplementary Figure 4C). KEGG pathway analysis revealed changes in metabolic pathways related to cell migration and EMT, including focal adhesion, ECM-receptor interaction, the Wnt signaling pathway, and the PI3K/Akt signaling pathway (Supplementary Figure 4D). Taken together, these findings suggest that LMO7 may affect cell junctions and adhesion. Cell-to-cell interactions and cell adhesion are key regulatory factors in cancer progression and contribute to immune evasion and tumor metastasis[10].
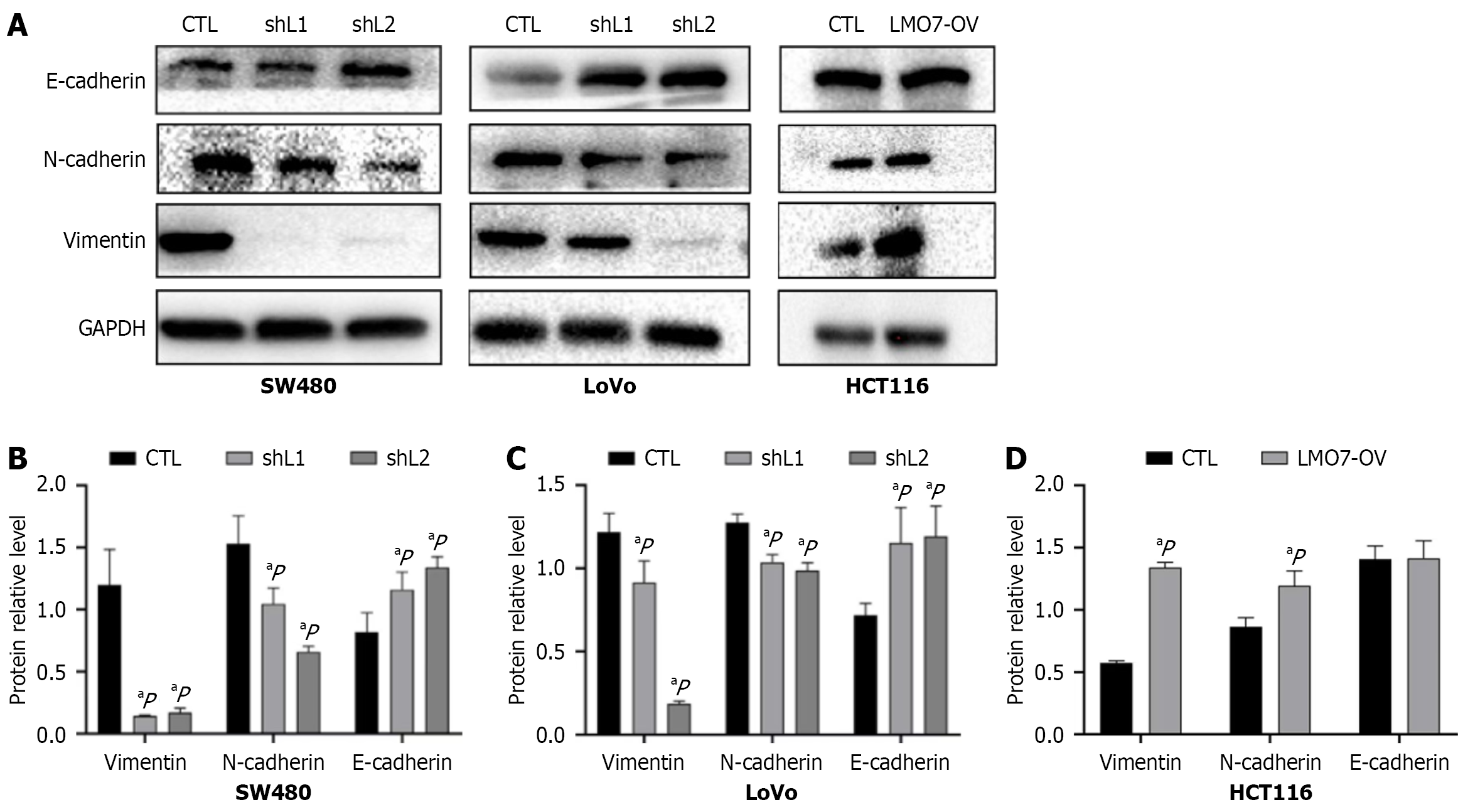
EMT is a developmental process that enables quiescent epithelial cells to acquire the ability to migrate and invade. During the process of carcinogenesis, EMT is related to tumor occurrence, invasion, metastasis, and even resistance to treatment[5,11]. We next detected the expression of the EMT-related markers E-cadherin, N-cadherin and vimentin (Figure 5). Compared with that in the cells transfected with the empty vector (CTL), the expression of the epithelial marker E-cadherin increased, whereas the expression of the mesenchymal markers N-cadherin and vimentin decreased in the LMO7-knockdown cells. Epithelial-to-mesenchymal cell transformation promotes cell metastasis and invasion; thus, these results indicate that LMO7 deletion reduces the EMT and metastasis ability of CRC cells. We further detected the expression of E-cadherin, N-cadherin and vimentin in HCT116 cells overexpressing LMO7. The protein expression of N-cadherin and vimentin increased, but the protein level of E-cadherin did not significantly change, which was inconsistent with our expectations.
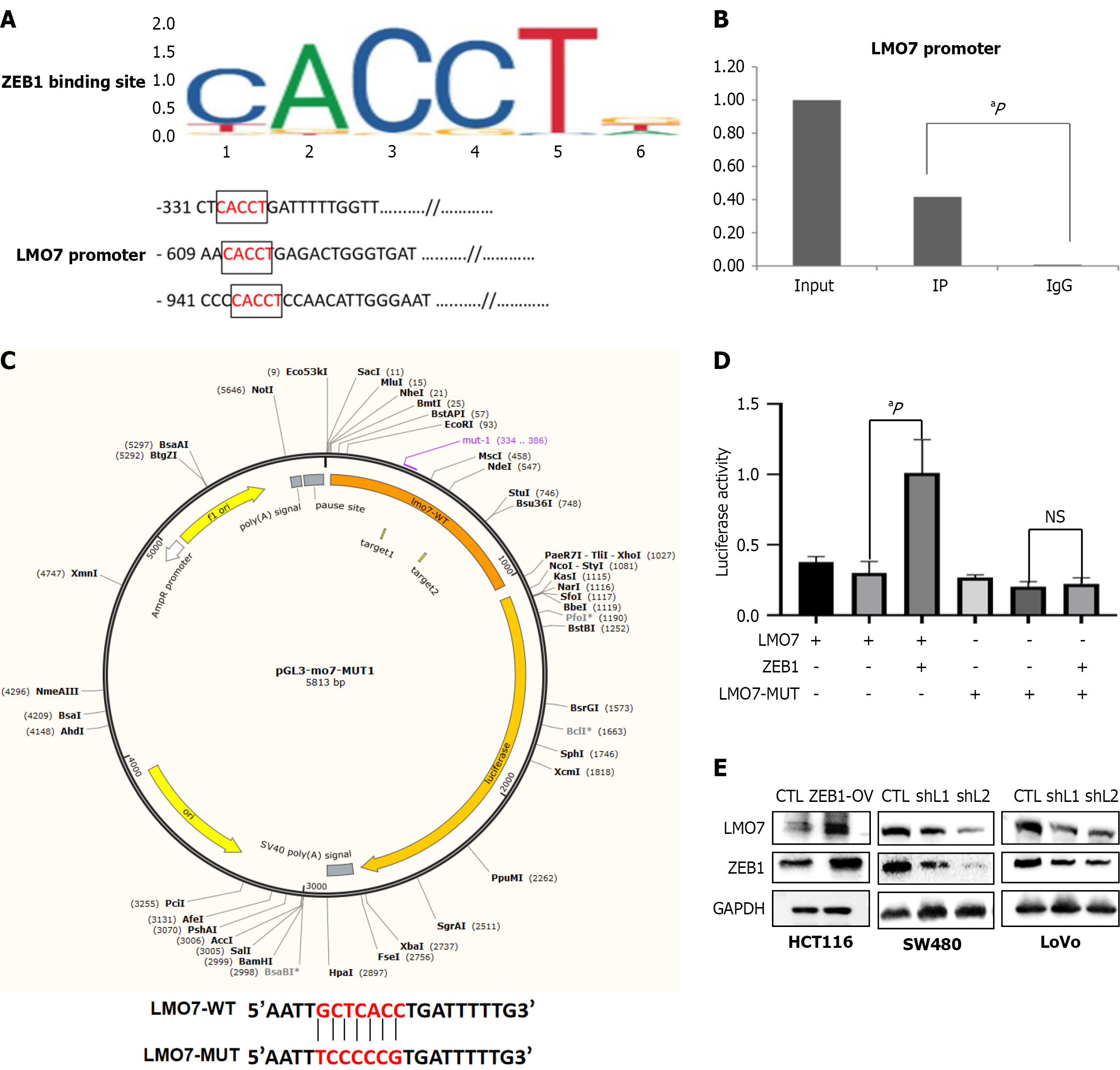
Our results confirmed that LMO7 plays an important role in CRC cell EMT, but the underlying regulatory mechanism remains unclear. We obtained the promoter sequence in the upstream 2 kb region of the human LMO7 gene through NCBI (https://www.ncbi.nlm.nih.gov). The binding sites of EMT-related transcription factors, such as SMAD2, Smad3, Snail, Slug, ZEB1, and TWIST, were obtained from jaspar.genereg.net. A prediction analysis revealed that the LMO7 promoter region contains three transcription binding sites for ZEB1 (Figure 6A). Chromatin immunoprecipitation ana
We next constructed ZEB1-overexpressing and ZEB1-knockdown (cells transfected with sh-ZEB1-1, cells transfected with sh-ZEB1-2) cells to examine the effect of ZEB1 on LMO7 expression. The results revealed that LMO7 protein expression was increased in ZEB1-overexpressing HCT116 cells compared with that in cells transfected with the blank vector (CTL), whereas it was decreased in ZEB1-knockdown SW480 and LoVo cells (Figure 6E). These results confirm that ZEB1 regulates the expression of the LMO7 gene in CRC.
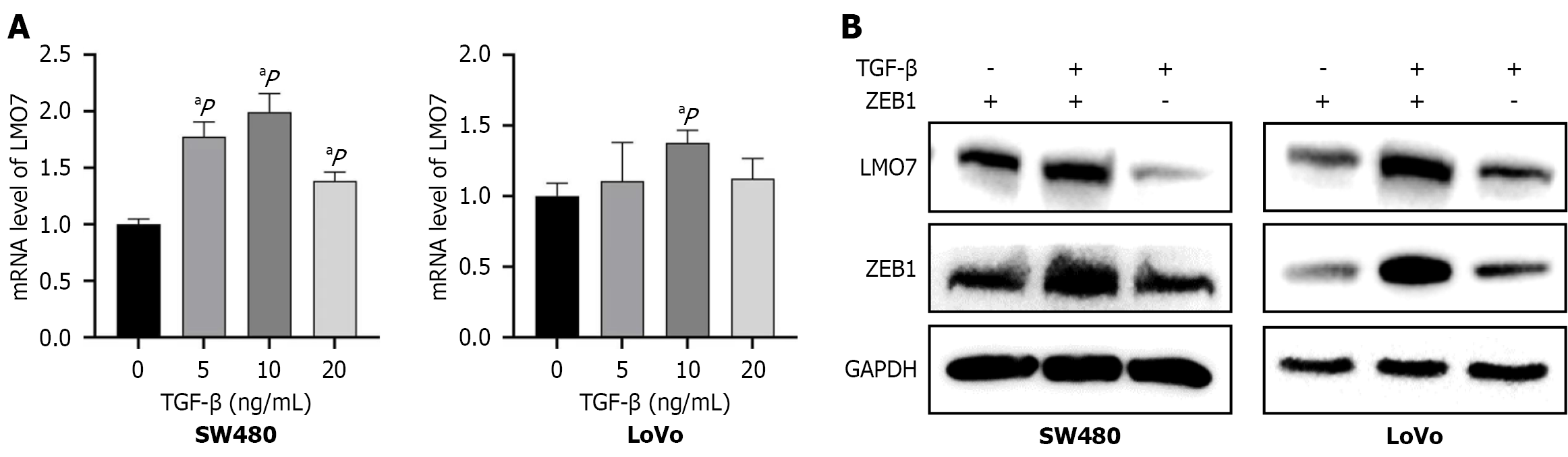
Previous studies have reported that TGF-β can induce the expression of ZEB1 and LMO7[12,13]. TGF-β is a cytokine that can regulate cell proliferation, migration and the differentiation of various cell types and plays an important role in tumor metastasis, partly because it can stimulate EMT in tumor cells. To gain further insight into the regulatory effect of TGF-β on LMO7 expression in CRC, we investigated the effect of TGF-β on the expression of ZEB1 and LMO7 in CRC cells. First, SW480 and LoVo cells were treated with 0 ng/mL, 5 ng/mL, 10 ng/mL and 20 ng/mL TGF-β1, and LMO7 mRNA levels were assessed. The results revealed that LMO7 mRNA levels increased after the addition of TGF-β, with the most significant increase occurring at 10 ng/mL (Figure 7A). Next, SW480 cells, ZEB1-knockdown SW480 cells, LoVo cells, and ZEB1-knockdown LoVo cells were treated with 10 ng/mL TGF-β1, and changes in ZEB1 and LMO7 protein expression were detected. The results revealed that ZEB1 and LMO7 expression increased in SW480 and LoVo cells but decreased in ZEB1-knockdown cells (Figure 7B). These results strongly suggest that TGF-β can induce the expression of LMO7 in CRC. Moreover, ZEB1 is likely involved in the regulation of LMO7 expression by TGF-β in CRC.
We identified a differentially expressed protein, LMO7, which has been rarely studied in CRC, through DIA proteomics of clinical CRC samples. LMO7 gene expression is elevated in CRC and is correlated with clinical stage and metastasis, promoting the proliferation, migration and invasion of CRC tumor cells. We also provide evidence that LMO7 promotes CRC cell proliferation, migration and invasion in vitro and that LMO7 knockdown inhibits subcutaneous tumor formation and lung metastasis in vivo. These findings indicate that LMO7 may have a significant protumorigenic effect on CRC progression and is associated with the distant metastasis of CRC.
Previous studies have shown that LMO7 is upregulated in multiple tumors, including CRC, breast cancer, lung cancer, thyroid cancer and liver cancer[14,15]. LMO7 can significantly promote the occurrence of pancreatic tumors by affecting the cell cycle and programmed cell death. Gene knockdown or knockout of LMO7 not only significantly inhibits the proliferation, colony formation and motility of pancreatic cancer cells in vitro but also significantly slows tumor growth and metastasis in vivo[16]. LMO7 immunoreactivity has been shown to be an adverse prognostic factor and indicates poor patient survival in non-small cell lung cancer[17], but another report proposed that LMO7 is negatively correlated with the development and prognosis of human lung adenocarcinoma[18] and papillomavirus-positive oropharyngeal squamous cell carcinoma[19]. In CRC, the difference in LMO7 expression was statistically significant in survival analysis at both the RNA and protein levels and was identified as a potential clinical biomarker. Although LMO7 may be involved in CRC metastasis, it may not be a prognosis-related risk factor at the population level[20]. Until recently, the biological role of LMO7 has not been widely discussed, and LMO7 has not generally been regarded as a mature oncogene. Notably, more research is needed to fully understand the molecular mechanisms through which LMO7 functions in cancer.
As a cytoskeletal protein, LMO7 plays a role in promoting cell migration[21,22]. Cell migration is a cytoskeleton-driven cellular process involving many physiological and pathological events, such as embryonic development and cancer metastasis. Recent research has shown that the absence of LMO7 inhibits the directed migration of normal fibroblasts on fibronectin-rich surfaces, indicating that LMO7 is important for directed migration[23]. The invasion and migration of tumor cells are often achieved through existing cellular mechanisms such as adherens junction signal transduction pathways. Adherens junctions have multiple functions, such as initiating and stabilizing intercellular adhesion, and regulating the actin cytoskeleton[24]. LMO7 is an organizer of epithelial cell polarity and cell-cell adhesion, and can regulate the actin cytoskeleton and the assembly of adherens junctions in epithelial cells[25]. The loss of LMO7 in renal tubular epithelial cells can lead to the loss of integrity of the epithelial layer connection[26]. Helicobacter pylori can upregulate the expression of the LMO7 gene in gastric mucosal epithelial cells to reduce E-cadherin expression, which regulates cell adhesion and contact inhibition in gastric mucosal epithelial cells[27]. The plasticity of cancer invasion and metastasis depends on the ability of cancer cells to switch between collective migration and single-cell dissemination, a process controlled by cell-cell adhesion[28]. Previous studies have shown that LMO7 is significantly upregulated in breast cancer cells and can regulate cell migration and adhesion by mediating the activation of the Rho-MRTF-SRF signaling pathway[29], which is involved participating in the progression and lymph node metastasis of breast cancer[30]. On the basis of the existing evidence of our results, the effect of LMO7 on cell adhesion in CRC may be an important mechanism promoting tumor cell migration and invasion and may be related to its role in promoting CRC metastasis. However, the potential regulatory mechanism through which LMO7 affects cell adhesion is still poorly understood.
EMT plays a crucial role in promoting tumor metastasis and invasion, and is the process through which tumor cells lose intercellular adhesion and thereby acquire invasive and metastatic activity. The hallmark of EMT is the loss of epithelial surface markers, most notably a decrease in E-cadherin levels, while cells acquire a mesenchymal phenotype, such as elevated expression of vimentin and N-cadherin, which reduces cell adhesion and thereby confers the ability to metastasize and invade[31]. Notably, non-coding RNAs also regulate CRC EMT. Luo et al[32] reported that circWBSCR22 promotes metastasis by increasing CHD4 stability, indicating that EMT is controlled by multiple mechanisms (including LMO7 and circRNAs). We found that the loss of LMO7 increased the expression of E-cadherin and decreased the expression of N-cadherin and vimentin, suggesting that LMO7 may be involved in the EMT of CRC cells by regulating the expression of E-cadherin, N-cadherin and vimentin, thereby reducing the adhesion ability of CRC cells, increasing the migration of tumor cells and promoting the metastasis of tumor cells. However, unexpectedly, overexpression of LMO7 in CRC cells did not lead to a decrease in the expression of E-cadherin. Researchers have proposed a relationship between E-cadherin downregulation and cancer invasion. Interestingly, a previous study revealed that breast cancer tumor cells lacking E-cadherin in mice rarely metastasize, and both primary and metastatic tumors, as well as tumor cells in the bloodstream, express E-cadherin[33]. Compared with that in primary CRC tumors, E-cadherin expression is increased in liver and lymph node metastases[34]. However, another study revealed that E-cadherin-negative breast cancer cells can also induce lung metastasis of primary tumors in mice, and these metastases re-express E-cadherin[35]. The expression of E-cadherin in metastatic cancer cells may contribute to selective growth advantages and play a key role in the phenotypic changes of primary and secondary cancer lesions, supporting tumor cell plasticity.
The recently-proposed cancer eco-evolutionary theory indicates that cancer is a pathological ecosystem involving dynamic interactions between tumor cells and the microenvironment, where phenotypes such as EMT are adaptive strategies in response to microenvironmental stress rather than being driven by a single gene[36,37]. Traditionally regarded as a cytoplasmic protein, LMO7 participates in cell morphogenesis by regulating cell adhesion complexes (e.g., the E-cadherin-catenin complex), and facilitates the disruption of epithelial layer constraints[25]; thus, LMO7 may become pivotal for tumor niche establishment. Therefore, we hypothesize that LMO7 may play a pivotal role in enabling CRC cells to adapt to the metastatic microenvironment and that this role requires the participation of E-cadherin.
EMT transcription factors regulate epithelial/mesenchymal markers directly or indirectly in the oncogenesis and development of cancer. Among these transcription factors, the promoter region of LMO7 contains only the transcriptional binding site of ZEB1. We further confirmed that ZEB1 specifically binds to the LMO7 promoter and regulates the expression of the LMO7 protein. ZEB1 is thought to be associated with tumorigenesis and poor prognosis in various tumors[13]. ZEB1 offers cellular plasticity, which serves as a prerequisite for the ability of cells to evade the primary tumor mass and adapt to the new environment during the process of metastatic colony formation. In the absence of ZEB1, the colonization ability of the isolated and cultured tumor cell lines specifically decreased. Similarly, during the process of malignant transformation, ZEB1-mediated EMT is the main and most important driving force for tumor metastasis[38]. ZEB1 also plays a significant role in the invasion and metastasis of CRC[39,40]. Cai et al[41] reported that the TRP
In epithelial tumor cells, TGF-β signaling promotes EMT by inducing the expression of transcription factors such as Snail1, Slug, and ZEB1; inhibits the expression of epithelial cell adhesion proteins; promotes the loss of cell polarity and intercellular contact; and promotes the acquisition of a migratory and invasive phenotype, thus facilitating the spread of tumor cells[42]. TGF-β can also regulate the expression of ZEB1 and thereby be involved in cell proliferation, migration and angiogenesis[43]. In the present study, we demonstrated that TGF-β intervention can induce the expression of ZEB1 and LMO7 proteins in CRC cells, confirming that TGF-β can regulate the expression of ZEB1 and LMO7, which is consistent with the findings of previous studies. Additionally, we found that the ability of TGF-β to induce LMO7 expre
Tumor metastasis is a complex biological process regulated by multiple molecular signaling pathways. This study is the first to explore the role and partial regulatory mechanism of LMO7 in CRC. LMO7 expression is upregulated in CRC and is positively associated with clinical stage, lymph node metastasis, and distant metastasis. LMO7 can promote the proliferation of CRC cells and enhance their migration and invasion capabilities by influencing the adhesion function of CRC cells, ultimately facilitating the progression and metastasis of CRC. TGF-β may induce LMO7 expression by upregulating the expression of the transcription factor ZEB1, accordingly promoting EMT and enabling epithelial tumor cells to acquire the ability to metastasize. Nonetheless, whether LMO7 is correlated with overall survival and disease-free survival in CRC still needs to be further analyzed in subsequent follow-up analyses. Further experimental exploration is needed to determine whether TGF-β directly regulates the expression of ZEB1/LMO7. However, since the progression and meta
| 1. | Dekker E, Tanis PJ, Vleugels JLA, Kasi PM, Wallace MB. Colorectal cancer. Lancet. 2019;394:1467-1480. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4063] [Cited by in RCA: 3536] [Article Influence: 505.1] [Reference Citation Analysis (10)] |
| 2. | Kasi PM, Shahjehan F, Cochuyt JJ, Li Z, Colibaseanu DT, Merchea A. Rising Proportion of Young Individuals With Rectal and Colon Cancer. Clin Colorectal Cancer. 2019;18:e87-e95. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 114] [Cited by in RCA: 110] [Article Influence: 15.7] [Reference Citation Analysis (1)] |
| 3. | Wang J, Liu L, Cai Y, Gao Y, Guo Z, Yu F, Ping Z. Trends in the age-related incidence of colon and rectal cancers in China, 2005-2015. Dig Liver Dis. 2021;53:908-914. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 11] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 4. | Tauriello DV, Calon A, Lonardo E, Batlle E. Determinants of metastatic competency in colorectal cancer. Mol Oncol. 2017;11:97-119. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 207] [Cited by in RCA: 202] [Article Influence: 22.4] [Reference Citation Analysis (7)] |
| 5. | Noubissi Nzeteu GA, Geismann C, Arlt A, Hoogwater FJH, Nijkamp MW, Meyer NH, Bockhorn M. Role of Epithelial-to-Mesenchymal Transition for the Generation of Circulating Tumors Cells and Cancer Cell Dissemination. Cancers (Basel). 2022;14:5483. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 20] [Article Influence: 5.0] [Reference Citation Analysis (3)] |
| 6. | Ding Z, Wang N, Ji N, Chen ZS. Proteomics technologies for cancer liquid biopsies. Mol Cancer. 2022;21:53. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 215] [Cited by in RCA: 168] [Article Influence: 42.0] [Reference Citation Analysis (0)] |
| 7. | Dong L, Lu D, Chen R, Lin Y, Zhu H, Zhang Z, Cai S, Cui P, Song G, Rao D, Yi X, Wu Y, Song N, Liu F, Zou Y, Zhang S, Zhang X, Wang X, Qiu S, Zhou J, Wang S, Zhang X, Shi Y, Figeys D, Ding L, Wang P, Zhang B, Rodriguez H, Gao Q, Gao D, Zhou H, Fan J. Proteogenomic characterization identifies clinically relevant subgroups of intrahepatic cholangiocarcinoma. Cancer Cell. 2022;40:70-87.e15. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 307] [Cited by in RCA: 271] [Article Influence: 67.8] [Reference Citation Analysis (3)] |
| 8. | Chen X, Fu XS, Li CP, Zhao HX. ER stress and ER stress-induced apoptosis are activated in gastric SMCs in diabetic rats. World J Gastroenterol. 2014;20:8260-8267. [PubMed] [DOI] [Full Text] |
| 9. | Yun C, DasGupta R. Luciferase reporter assay in Drosophila and mammalian tissue culture cells. Curr Protoc Chem Biol. 2014;6:7-23. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 17] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 10. | Läubli H, Borsig L. Altered Cell Adhesion and Glycosylation Promote Cancer Immune Suppression and Metastasis. Front Immunol. 2019;10:2120. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 227] [Cited by in RCA: 205] [Article Influence: 29.3] [Reference Citation Analysis (0)] |
| 11. | Pastushenko I, Blanpain C. EMT Transition States during Tumor Progression and Metastasis. Trends Cell Biol. 2019;29:212-226. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2351] [Cited by in RCA: 2088] [Article Influence: 298.3] [Reference Citation Analysis (3)] |
| 12. | Nakamura H, Mukai M, Komatsu K, Tanaka-Okamoto M, Itoh Y, Ishizaki H, Tatsuta M, Inoue M, Miyoshi J. Transforming growth factor-beta1 induces LMO7 while enhancing the invasiveness of rat ascites hepatoma cells. Cancer Lett. 2005;220:95-99. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 21] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 13. | Wu HT, Zhong HT, Li GW, Shen JX, Ye QQ, Zhang ML, Liu J. Oncogenic functions of the EMT-related transcription factor ZEB1 in breast cancer. J Transl Med. 2020;18:51. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 136] [Cited by in RCA: 122] [Article Influence: 20.3] [Reference Citation Analysis (0)] |
| 14. | te Velthuis AJ, Bagowski CP. PDZ and LIM domain-encoding genes: molecular interactions and their role in development. ScientificWorldJournal. 2007;7:1470-1492. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 92] [Cited by in RCA: 82] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 15. | Zeng Q, Jiang T, Wang J. Role of LMO7 in cancer (Review). Oncol Rep. 2024;52:117. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 45] [Cited by in RCA: 32] [Article Influence: 16.0] [Reference Citation Analysis (0)] |
| 16. | Liu X, Yuan H, Zhou J, Wang Q, Qi X, Bernal C, Avella D, Kaifi JT, Kimchi ET, Timothy P, Cheng K, Miao Y, Jiang K, Li G. LMO7 as an Unrecognized Factor Promoting Pancreatic Cancer Progression and Metastasis. Front Cell Dev Biol. 2021;9:647387. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 21] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 17. | Karlsson T, Kvarnbrink S, Holmlund C, Botling J, Micke P, Henriksson R, Johansson M, Hedman H. LMO7 and LIMCH1 interact with LRIG proteins in lung cancer, with prognostic implications for early-stage disease. Lung Cancer. 2018;125:174-184. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 19] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 18. | Nakamura H, Hori K, Tanaka-Okamoto M, Higashiyama M, Itoh Y, Inoue M, Morinaka S, Miyoshi J. Decreased expression of LMO7 and its clinicopathological significance in human lung adenocarcinoma. Exp Ther Med. 2011;2:1053-1057. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 28] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 19. | Israelsson P, Oda H, Öfverman C, Stefansson K, Lindquist D. Immunoreactivity of LMO7 and other molecular markers as potential prognostic factors in oropharyngeal squamous cell carcinoma. BMC Oral Health. 2024;24:729. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 1] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 20. | Zhou B, Guo R. Genomic and regulatory characteristics of significant transcription factors in colorectal cancer metastasis. Sci Rep. 2018;8:17836. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 22] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 21. | Zhang Y, Wang W, Duan C, Li M, Gao L. Novel Long Non-Coding RNA (lncRNA) Transcript AL137782.1 Promotes the Migration of Normal Lung Epithelial Cells through Positively Regulating LMO7. Int J Mol Sci. 2023;24:13904. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 2] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 22. | Wu H, Zhou J, Mei S, Wu D, Mu Z, Chen B, Xie Y, Ye Y, Liu J. Circulating exosomal microRNA-96 promotes cell proliferation, migration and drug resistance by targeting LMO7. J Cell Mol Med. 2017;21:1228-1236. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 142] [Cited by in RCA: 142] [Article Influence: 15.8] [Reference Citation Analysis (0)] |
| 23. | Bun T, Sato Y, Futami H, Tagawa Y, Murakami Y, Takahashi M. Cytoskeletal fractionation identifies LMO7 as a positive regulator of fibroblast polarization and directed migration. Biochem Biophys Res Commun. 2023;638:58-65. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 1] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 24. | Campàs O, Noordstra I, Yap AS. Adherens junctions as molecular regulators of emergent tissue mechanics. Nat Rev Mol Cell Biol. 2024;25:252-269. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 143] [Cited by in RCA: 95] [Article Influence: 47.5] [Reference Citation Analysis (0)] |
| 25. | Ooshio T, Irie K, Morimoto K, Fukuhara A, Imai T, Takai Y. Involvement of LMO7 in the association of two cell-cell adhesion molecules, nectin and E-cadherin, through afadin and alpha-actinin in epithelial cells. J Biol Chem. 2004;279:31365-31373. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 143] [Cited by in RCA: 126] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 26. | Zhen YY, Wu CH, Chen HC, Chang EE, Lee JJ, Chen WY, Chang JM, Tseng PY, Wang YF, Hung CC. Coordination of LMO7 with FAK Signaling Sustains Epithelial Integrity in Renal Epithelia Exposed to Osmotic Pressure. Cells. 2022;11:3805. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 6] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 27. | Lim JW, Kim H, Kim KH. Cell adhesion-related gene expression by Helicobacter pylori in gastric epithelial AGS cells. Int J Biochem Cell Biol. 2003;35:1284-1296. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 64] [Cited by in RCA: 60] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 28. | Ilina O, Gritsenko PG, Syga S, Lippoldt J, La Porta CAM, Chepizhko O, Grosser S, Vullings M, Bakker GJ, Starruß J, Bult P, Zapperi S, Käs JA, Deutsch A, Friedl P. Cell-cell adhesion and 3D matrix confinement determine jamming transitions in breast cancer invasion. Nat Cell Biol. 2020;22:1103-1115. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 355] [Cited by in RCA: 255] [Article Influence: 42.5] [Reference Citation Analysis (0)] |
| 29. | Hu Q, Guo C, Li Y, Aronow BJ, Zhang J. LMO7 mediates cell-specific activation of the Rho-myocardin-related transcription factor-serum response factor pathway and plays an important role in breast cancer cell migration. Mol Cell Biol. 2011;31:3223-3240. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 62] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 30. | Sasaki M, Tsuji N, Furuya M, Kondoh K, Kamagata C, Kobayashi D, Yagihashi A, Watanabe N. PCD1, a novel gene containing PDZ and LIM domains, is overexpressed in human breast cancer and linked to lymph node metastasis. Anticancer Res. 2003;23:2717-2721. [PubMed] |
| 31. | Serrano-Gomez SJ, Maziveyi M, Alahari SK. Regulation of epithelial-mesenchymal transition through epigenetic and post-translational modifications. Mol Cancer. 2016;15:18. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 670] [Cited by in RCA: 654] [Article Influence: 65.4] [Reference Citation Analysis (0)] |
| 32. | Luo J, Xu S, Wang J, He L, Li Z. Circular RNA circWBSCR22 facilitates colorectal cancer metastasis by enhancing CHD4's protein stability. Int J Biol Macromol. 2024;282:137135. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 4] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 33. | Padmanaban V, Krol I, Suhail Y, Szczerba BM, Aceto N, Bader JS, Ewald AJ. E-cadherin is required for metastasis in multiple models of breast cancer. Nature. 2019;573:439-444. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 727] [Cited by in RCA: 600] [Article Influence: 85.7] [Reference Citation Analysis (5)] |
| 34. | Buda A, Pignatelli M. E-cadherin and the cytoskeletal network in colorectal cancer development and metastasis. Cell Commun Adhes. 2011;18:133-143. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 57] [Cited by in RCA: 54] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 35. | Chao YL, Shepard CR, Wells A. Breast carcinoma cells re-express E-cadherin during mesenchymal to epithelial reverting transition. Mol Cancer. 2010;9:179. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 331] [Cited by in RCA: 318] [Article Influence: 19.9] [Reference Citation Analysis (0)] |
| 36. | Luo W. Nasopharyngeal carcinoma ecology theory: cancer as multidimensional spatiotemporal "unity of ecology and evolution" pathological ecosystem. Theranostics. 2023;13:1607-1631. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 184] [Cited by in RCA: 160] [Article Influence: 53.3] [Reference Citation Analysis (1)] |
| 37. | Luo WR. [Rethinking cancer]. Zhonghua Zhong Liu Za Zhi. 2025;47:463-467. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 9] [Reference Citation Analysis (1)] |
| 38. | Brabletz S, Brabletz T, Stemmler MP. Road to perdition: Zeb1-dependent and -independent ways to metastasis. Cell Cycle. 2017;16:1729-1730. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 5] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 39. | Luo D, Ge W. MeCP2 Promotes Colorectal Cancer Metastasis by Modulating ZEB1 Transcription. Cancers (Basel). 2020;12:758. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 28] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 40. | Vu T, Datta PK. Regulation of EMT in Colorectal Cancer: A Culprit in Metastasis. Cancers (Basel). 2017;9:171. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 407] [Cited by in RCA: 404] [Article Influence: 44.9] [Reference Citation Analysis (5)] |
| 41. | Cai J, Zhang W, Lu Y, Liu W, Zhou H, Liu M, Bi X, Liu J, Chen J, Yin Y, Deng Y, Luo Z, Yang Y, Chen Q, Chen X, Xu Z, Zhang Y, Wu C, Long Q, Huang C, Yan C, Liu Y, Guo L, Li W, Yuan P, Jiao Y, Song W, Wang X, Huang Z, Ying J, Zhao H. Single-cell exome sequencing reveals polyclonal seeding and TRPS1 mutations in colon cancer metastasis. Signal Transduct Target Ther. 2024;9:247. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 17] [Article Influence: 8.5] [Reference Citation Analysis (0)] |
| 42. | Hao Y, Baker D, Ten Dijke P. TGF-β-Mediated Epithelial-Mesenchymal Transition and Cancer Metastasis. Int J Mol Sci. 2019;20:2767. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 977] [Cited by in RCA: 895] [Article Influence: 127.9] [Reference Citation Analysis (5)] |
| 43. | Zhang Y, Fan K, Xu X, Wang A. The TGF-β1 Induces the Endothelial-to-Mesenchymal Transition via the UCA1/miR-455/ZEB1 Regulatory Axis in Human Umbilical Vein Endothelial Cells. DNA Cell Biol. 2020;39:1264-1273. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 13] [Article Influence: 2.2] [Reference Citation Analysis (0)] |