Copyright: ©Author(s) 2026.
World J Gastroenterol. May 14, 2026; 32(18): 114713
Published online May 14, 2026. doi: 10.3748/wjg.v32.i18.114713
Published online May 14, 2026. doi: 10.3748/wjg.v32.i18.114713
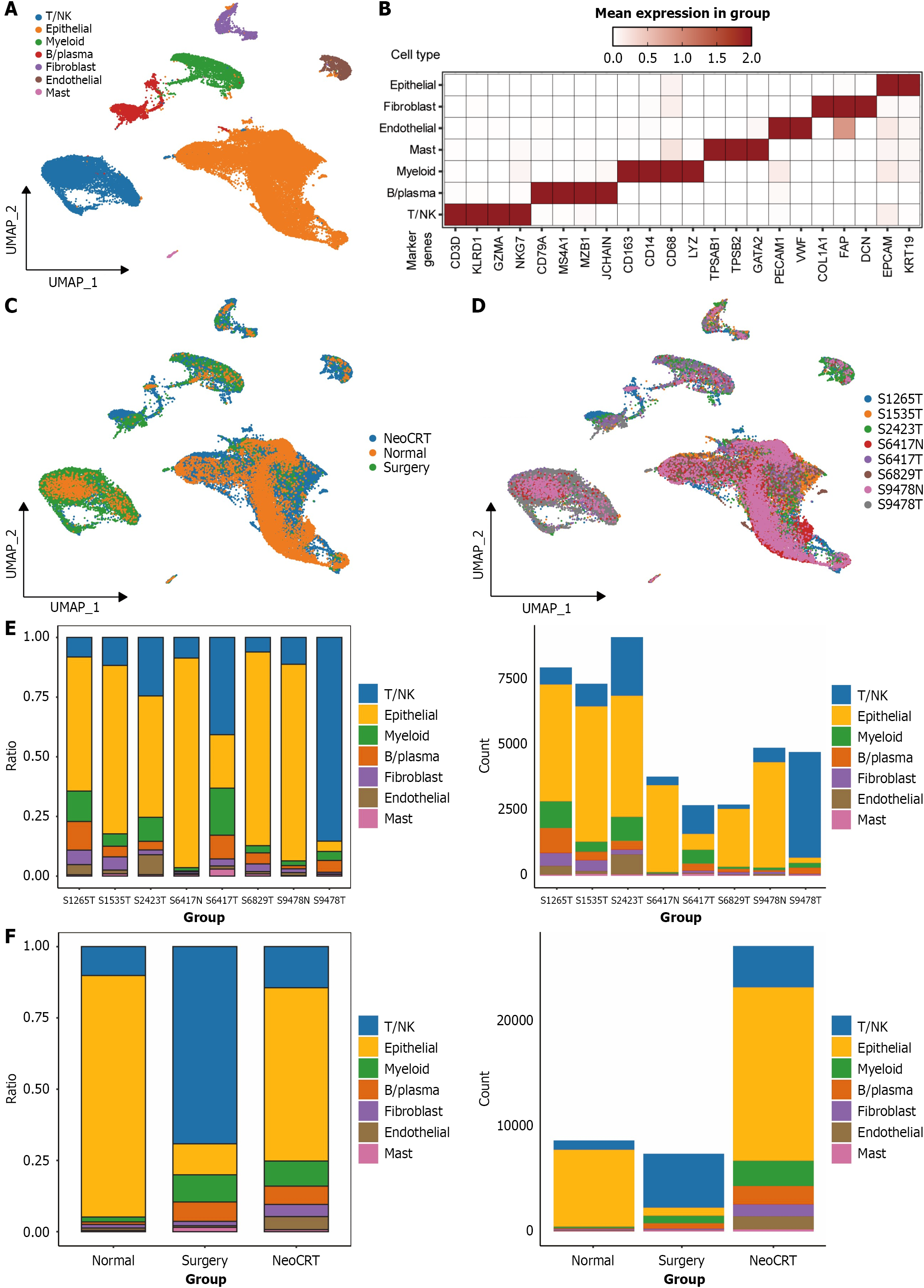
Figure 1 Comprehensive analysis of single-cell subpopulations in esophageal squamous cell carcinoma patients receiving neoadjuvant chemoradiotherapy and surgical treatment.
A: UMAP visualization of 43407 high-quality cells across all 7 cellular types; B: Marker genes for distinct cell types, with coloring based on Z-score normalized expression levels; C: UMAP visualization of all cells derived from normal tissue, surgical and neoadjuvant chemoradiotherapy groups; D: UMAP visualization of all cells obtained from 8 samples; E: Distribution and quantities of distinct cell types across 8 specimens; F: Distribution and quantities of distinct cell types within the 3 groups. NeoCRT: Neoadjuvant chemoradiotherapy; NK: Natural killer.
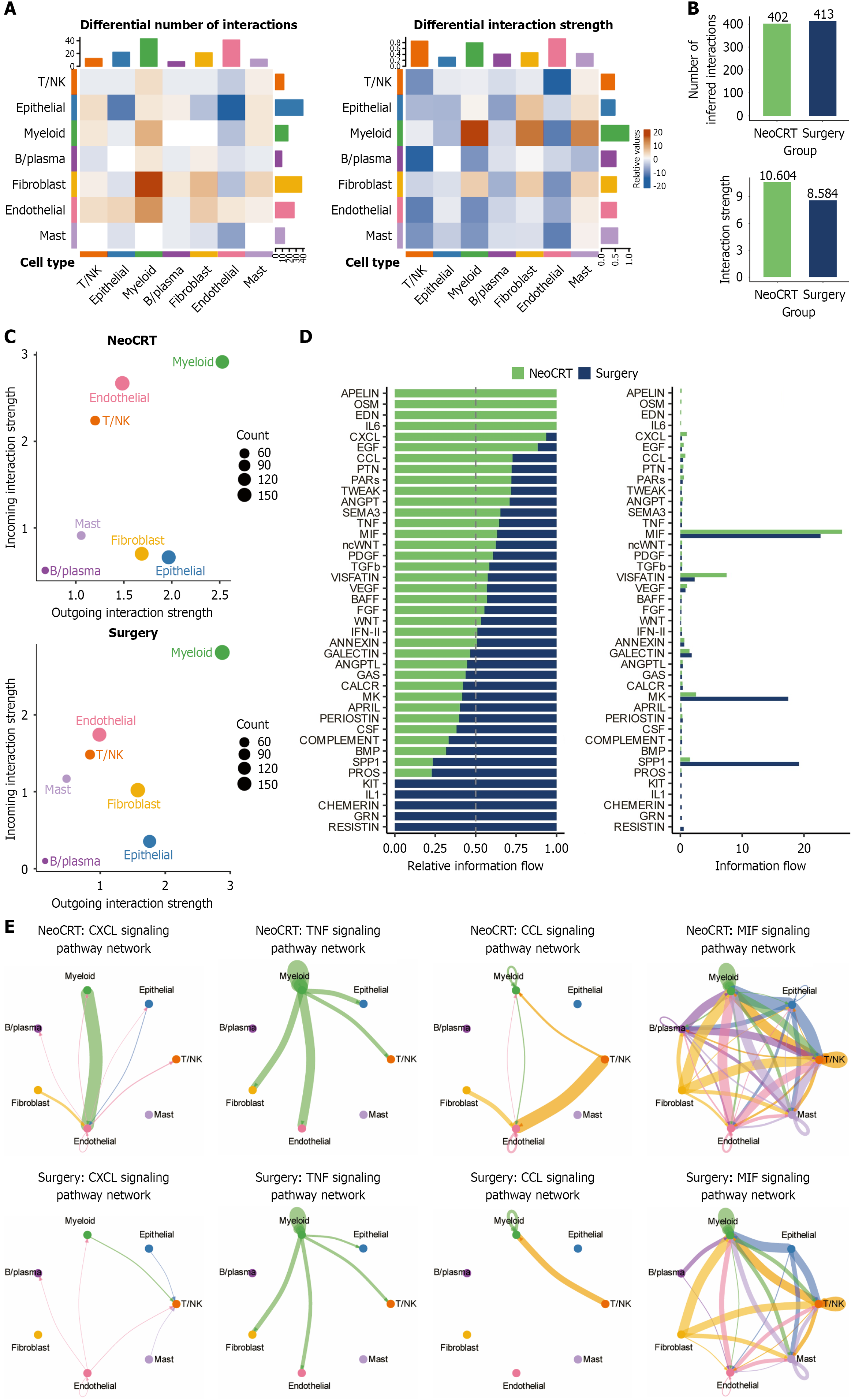
Figure 2 Cellchat analysis demonstrates neoadjuvant chemoradiotherapy-mediated development of a pro-inflammatory microen
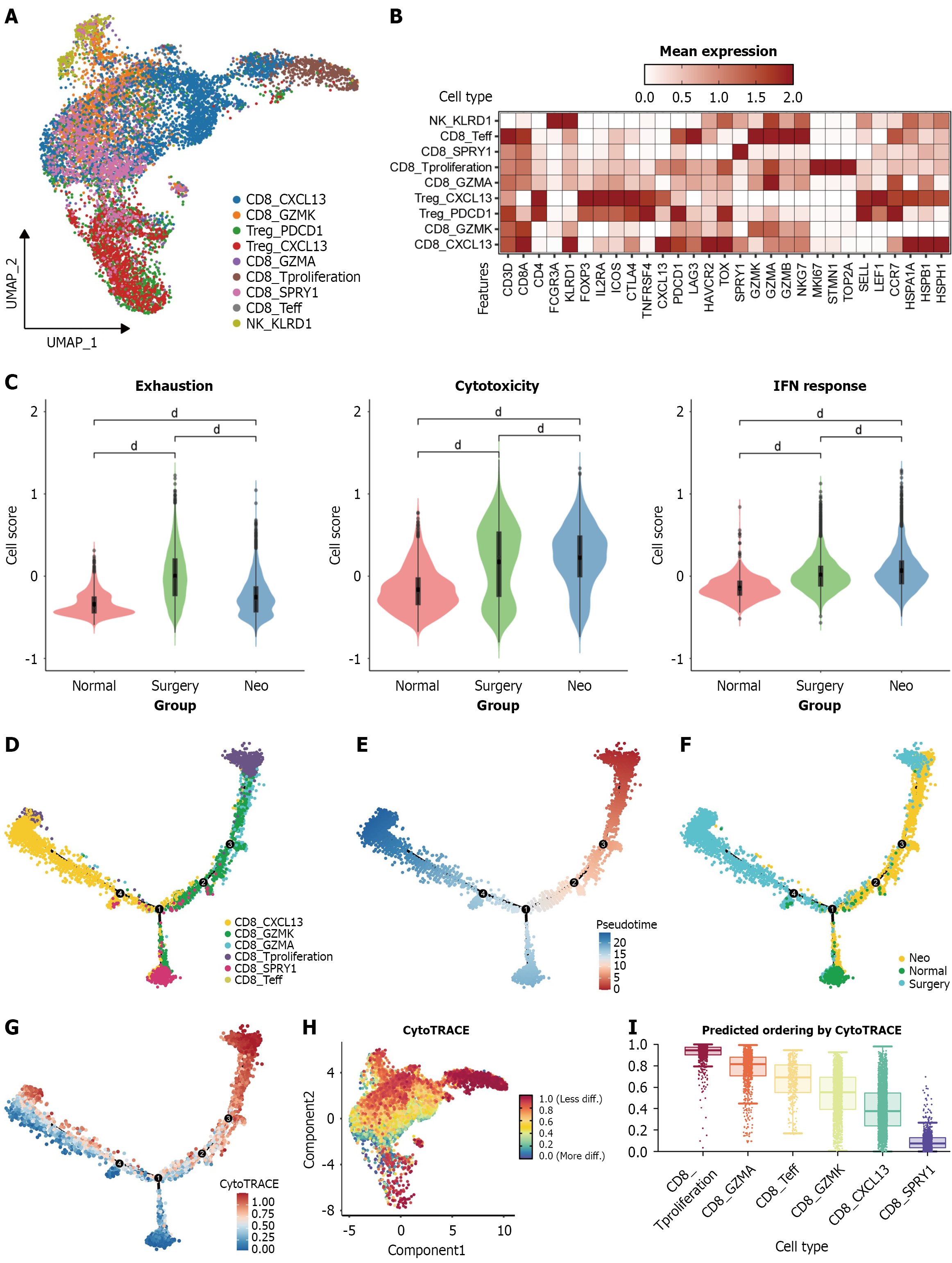
Figure 3 Neoadjuvant chemoradiotherapy facilitated CD8+ T cell activation.
A: UMAP visualization displaying 9 distinct clusters comprising 9968 T/natural killer cells; B: Heatmap illustrating marker gene expression across T cell subsets; C: Comparative analysis of T-cell functional scores; D: Temporal trajectory analysis of the entire CD8+ T cell population; E: Positioning of CD8+ T cell subtypes along the cellular developmental trajectory; F: Distinct groups along the cellular developmental trajectory; G: CytoTRACE score distribution throughout cellular developmental pathways; H: UMAP representation of CytoTRACE score distribution; I: Comparative differentiation levels among various CD8+ T cell subsets. dP < 0.0001. IFN: Interferon; Neo: Neoadjuvant.
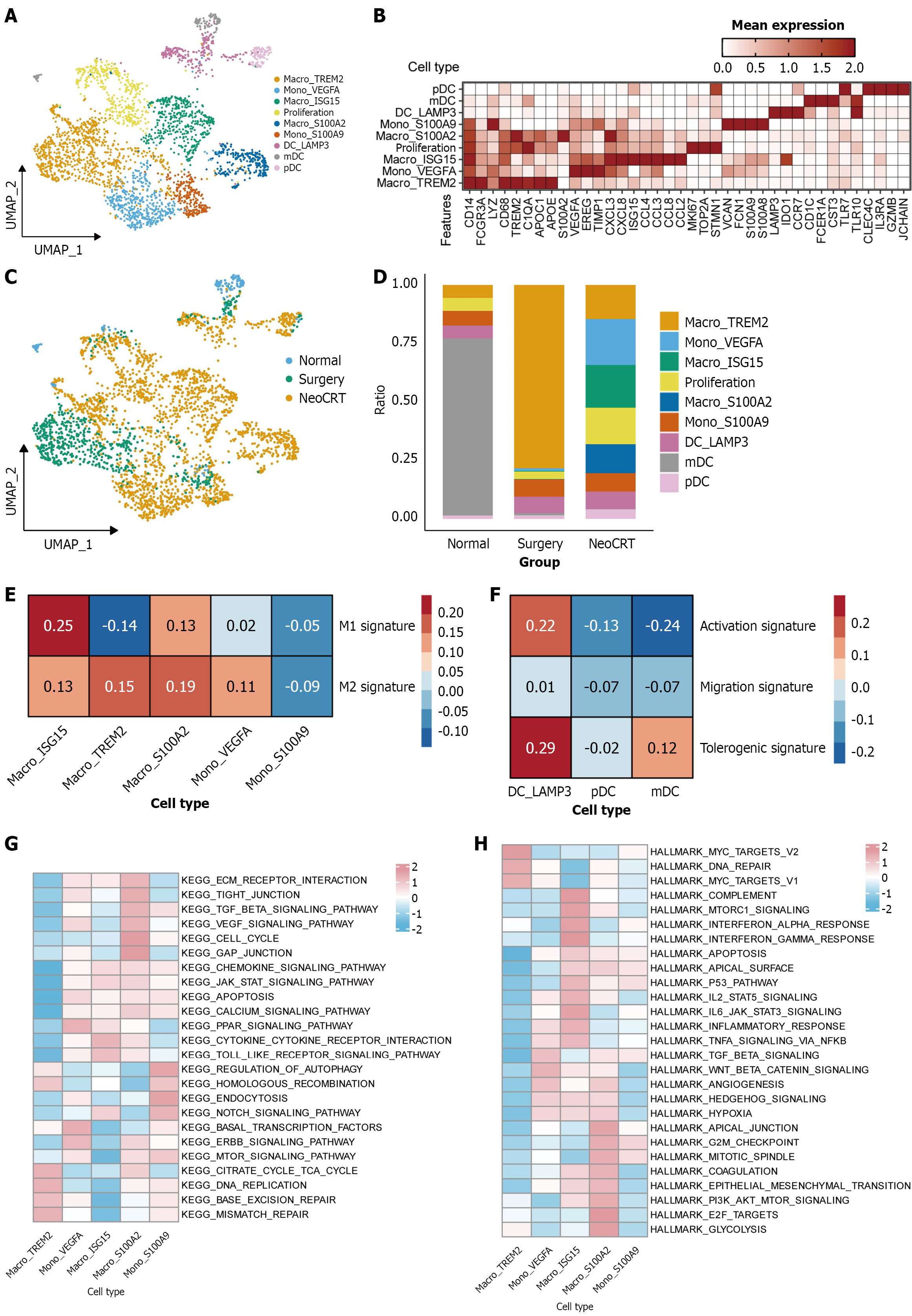
Figure 4 Characteristic alterations in myeloid cells following neoadjuvant chemoradiotherapy.
A: UMAP visualization of 9 distinct clusters comprising monocytes, macrophages, and dendritic cells; B: Marker genes identifying the 9 clusters; C: UMAP visualization depicting the distribution pattern of myeloid cells across diverse groups; D: Compositional analysis of 9 myeloid cell subsets across diverse groups; E: M1 and M2 signature scoring of monocytes and macrophages; F: Activation, migration, and tolerance signature scoring of dendritic cells; G: Kyoto Encyclopedia of Genes and Genomes pathway enrichment in distinct monocyte and macrophage populations; H: HALLMARK pathway enrichment in distinct monocyte and macrophage populations. NeoCRT: Neoadjuvant chemoradiotherapy.
Figure 5 Detection of malignant epithelial cells.
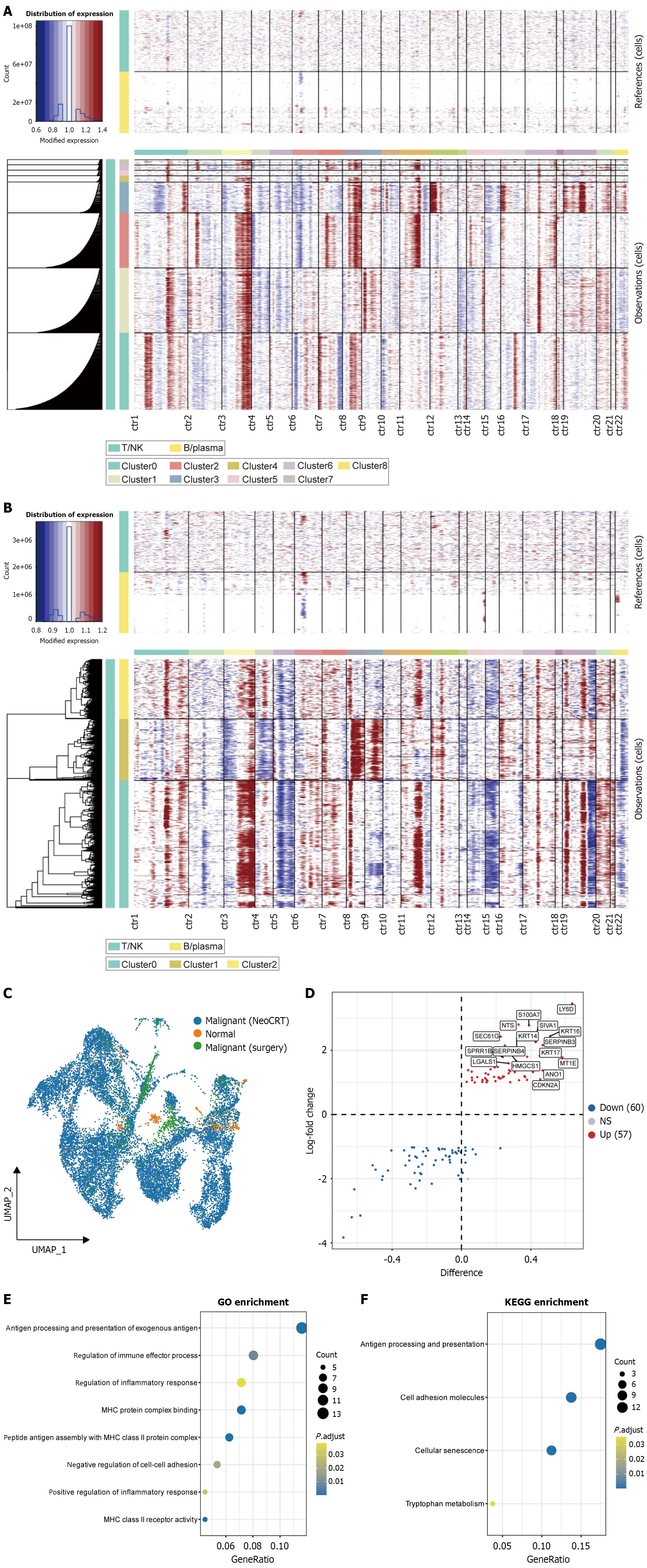
A and B: The inferCNV algorithm was employed to infer malignant epithelial cells in the neoadjuvant chemoradiotherapy and surgery groups; C: UMAP visualization depicting the spatial distribution of malignant and normal epithelial cells; D: Differentially expressed genes in malignant epithelial cells between neoadjuvant chemoradiotherapy and surgery groups; E and F: Gene Ontology and Kyoto Encyclopedia of Genes and Genomes pathway enrichment analyses of the differentially expressed genes. NeoCRT: Neoadjuvant chemoradiotherapy; NK: Natural killer.
Figure 6 Detection of essential drug resistance-associated genes.
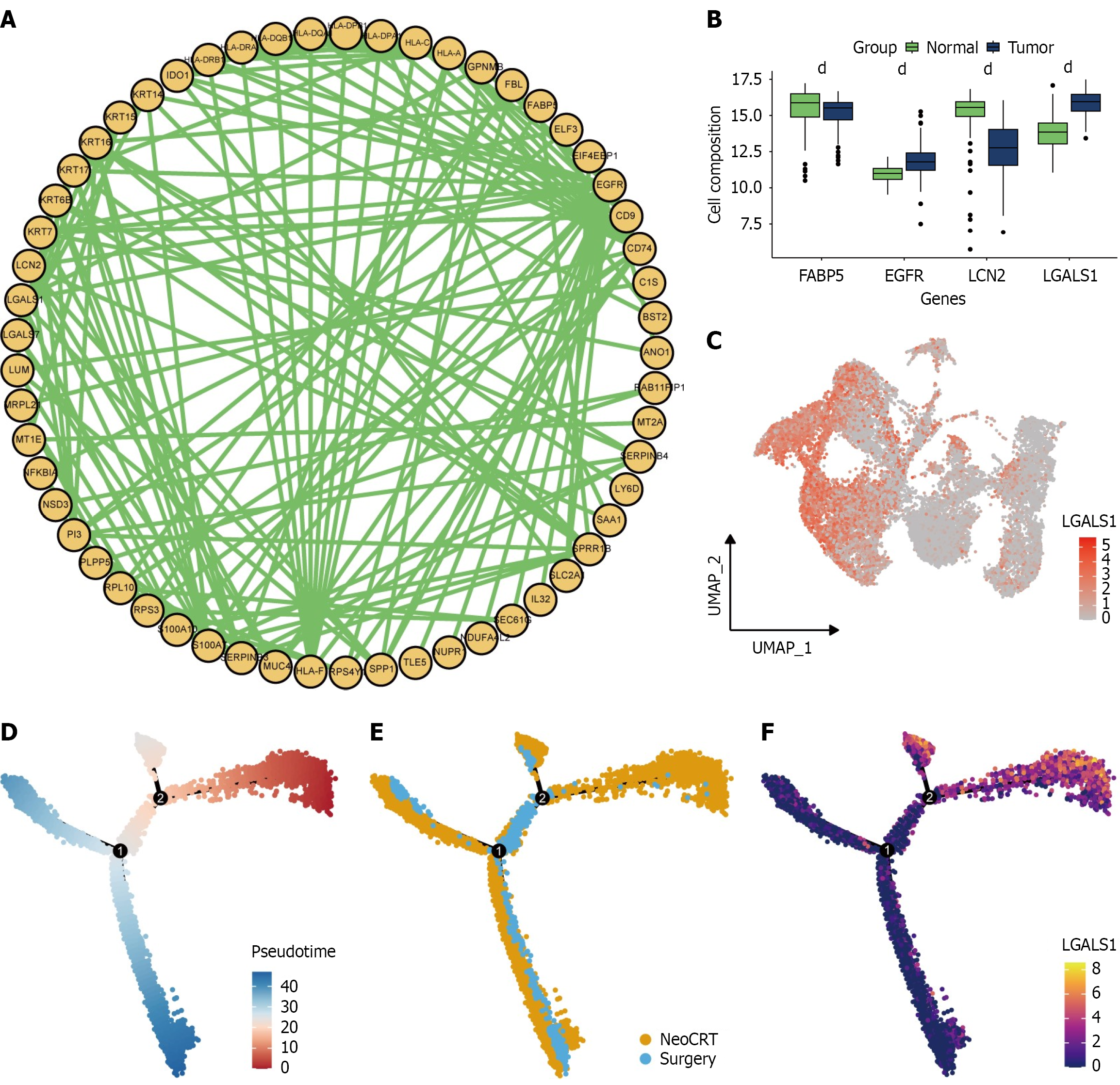
A: Protein-protein interaction networks for differentially expressed genes; B: Expression of key genes in tumor and normal tissues from GSE53625; C: LGALS1 expression across all epithelial cells; D: Pseudotime analysis of all malignant epithelial cells, with red indicating the starting point of cell development; E: Distribution of neoadjuvant chemoradiotherapy and surgery groups along cell developmental trajectories; F: Distribution of LGALS1. dP < 0.0001. NeoCRT: Neoadjuvant chemoradiotherapy.
Figure 7 LGALS1 diminishes esophageal squamous cell carcinoma cell responsiveness to cisplatin treatment.
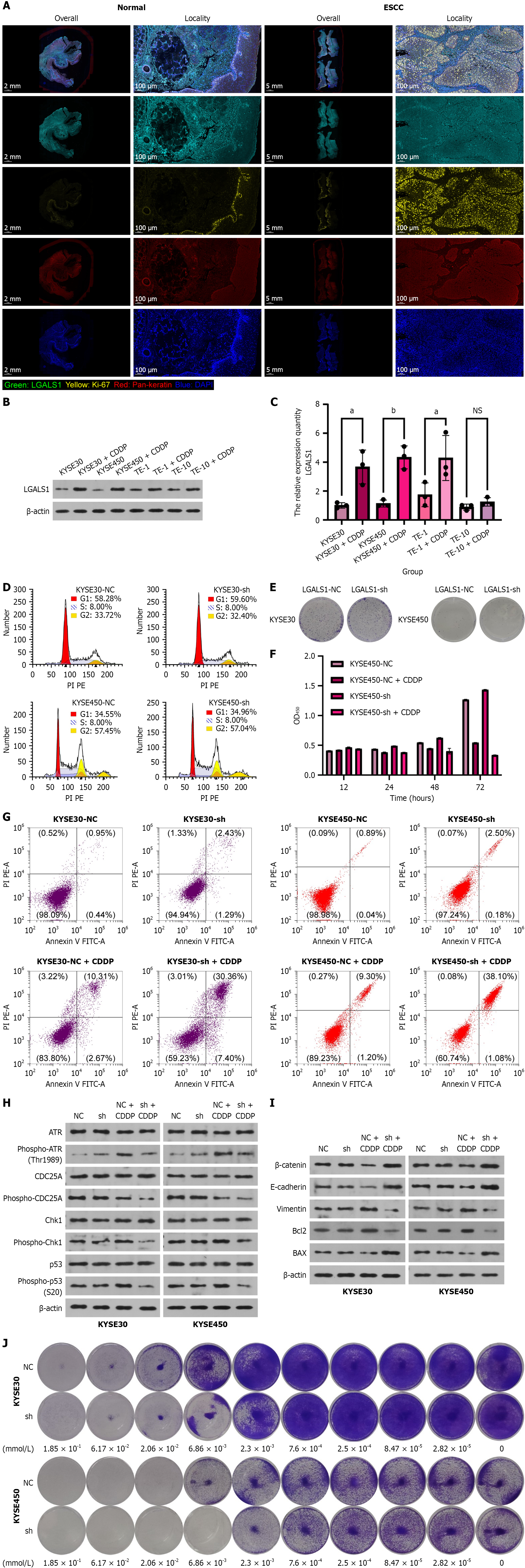
A: LGALS1, Ki67, and Pan-keratin expression levels in esophageal squamous cell carcinoma specimens compared with adjacent normal esophageal tissue samples; B: Assessment of LGALS1 expression changes under cisplatin administration utilizing quantitative real-time pathological complete remission methodology; C: Evaluation of LGALS1 protein levels following cisplatin exposure as determined by western blot analysis; D: The influence of LGALS1 expression on the cell cycle was analyzed by flow cytometry; E: Clone formation assay examined the influence of LGALS1 expression on cell proliferation; F: Cholecystokinin-8 examined the influence of LGALS1 expression on tumor cell viability under cisplatin treatment; G: The influence of LGALS1 expression on tumor cell apoptosis was ascertained by flow cytometry with cisplatin treatment; H: The phosphorylation levels of the DNA repair-related proteins ATR, CDC25A, Chk1, and p53 were measured by western blot; I: The expression of epithelial-mesenchymal transition and apoptosis-related proteins beta-catenin, E-cadherin, Vimentin, and Bcl2 and BAX was measured by western blot; J: Crystal violet staining was utilized to detect LGALS1 expression on tumor cell proliferation by cisplatin treatment with a concentration gradient. aP < 0.05, bP < 0.01, NS: Not significant. ESCC: Esophageal squamous cell carcinoma.
Figure 8 LGALS1 diminishes the cisplatin sensitivity of esophageal squamous cell carcinoma CDX model.
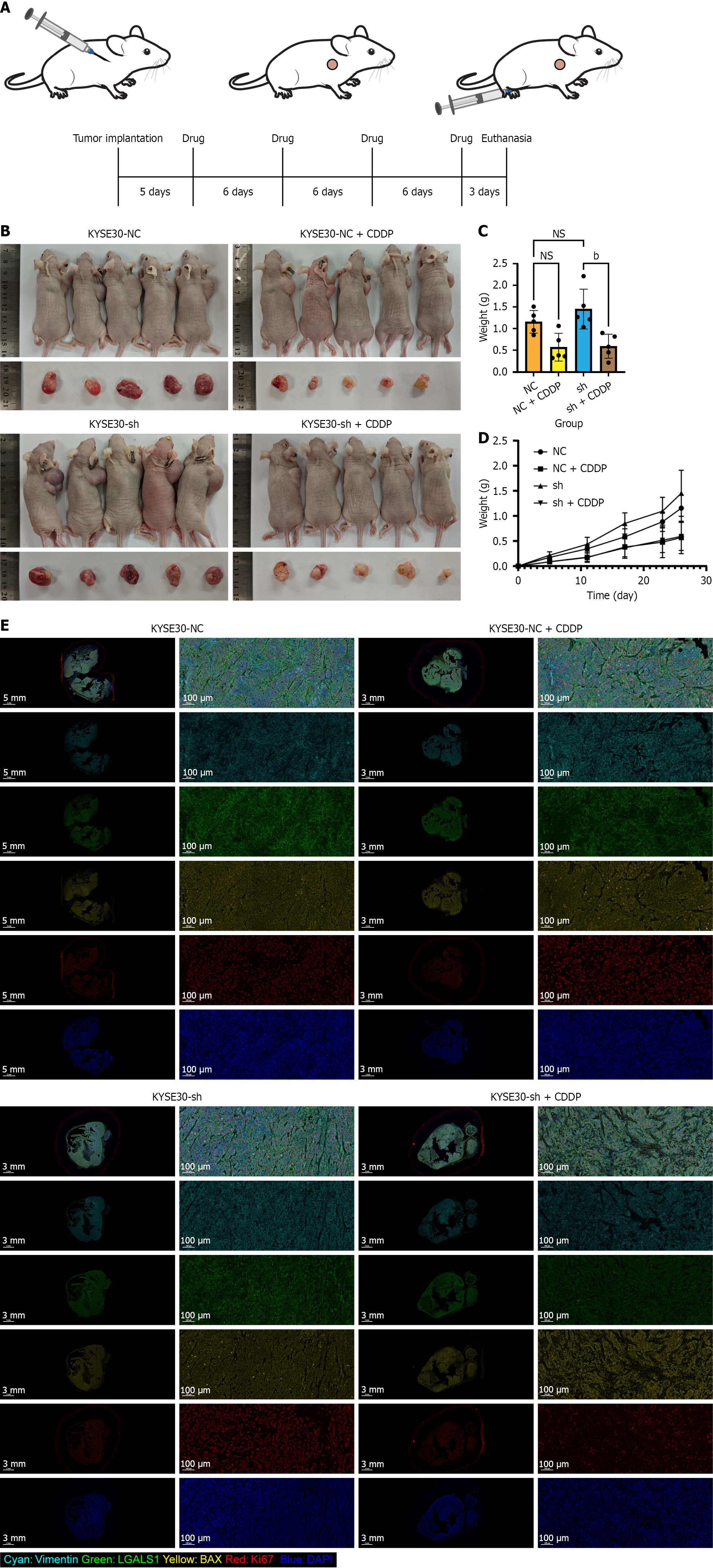
A: Experimental protocol for subcutaneous tumor formation and dosing experiments in mice; B: Mice and individual tumors after different treatment approaches; C: Statistical plots of tumor weights in mice following diverse treatments; D: Growth curves of mice following various treatments; E: The LGALS1, Ki67, and BAX and vimentin protein were detected in tumor sections by multiplex immunohistochemistry staining. bP < 0.01, NS: Not significant.
- Citation: Yan QH, Xu CD, Li ZG, Zheng YZ, Wong WS, Liang DC, Yang J, Chang WG, Wang JY. LGALS1 drives chemoresistance in esophageal squamous cell carcinoma by modulating epithelial-mesenchymal transition and tumor immunity. World J Gastroenterol 2026; 32(18): 114713
- URL: https://www.wjgnet.com/1007-9327/full/v32/i18/114713.htm
- DOI: https://dx.doi.org/10.3748/wjg.v32.i18.114713