Published online Jun 20, 2026. doi: 10.5662/wjm.v16.i2.115059
Revised: October 31, 2025
Accepted: January 5, 2026
Published online: June 20, 2026
Processing time: 199 Days and 1.2 Hours
The diagnosis of endogenous Cushing’s syndrome (CS) can be complicated and often delayed, given its low incidence (estimated globally at 1.8 cases to 4.5 cases per million people per year) and its clinical features that mimic far more prevalent metabolic disorders, such as central obesity, hypertension, and glucose intole
Core Tip: The diagnosis of Cushing’s syndrome remains challenging due to its rarity and its resemblance to common metabolic disorders. Large language models and other artificial intelligence-capable systems are potential diagnostic tools for early detection and differential diagnosis; nevertheless, they are likely to strengthen both human cognitive and training data biases. Large language models are susceptible to biases in the textual data they are trained on, reflecting human cognitive biases, while traditional machine learning models are susceptible to biases in structured data, leading to spectrum and measurement bias. Spectrum bias, exclusion of demographic variables, and heterogeneity of the data undermine diagnostic validity and justice. These heuristics are similar to clinicians’ mental shortcuts - anchoring, availability, and framing - and share the same diagnostic bias. Transparency, data diversity, and clinically relevant predictors are required to build unbiased, interpretable artificial intelligence solutions for endocrine diagnosis.
- Citation: Savvidis C, Liakopoulos C, Ilias I. Biases of large language models in diagnosing Cushing’s syndrome. World J Methodol 2026; 16(2): 115059
- URL: https://www.wjgnet.com/2222-0682/full/v16/i2/115059.htm
- DOI: https://dx.doi.org/10.5662/wjm.v16.i2.115059
Cushing’s syndrome (CS) is defined by persistent pathological hypercortisolism[1,2]. CS is difficult to diagnose due to its rarity[3] and the non-specific nature of its clinical signs[2]. The estimated global incidence of CS ranges from 1.8 cases to 4.5 cases per million people per year[4]. The clinical presentation of CS (with central obesity, hypertension, and glucose intolerance) may overlap to a large extent with many prevalent conditions such as metabolic syndrome, obesity, and polycystic ovary syndrome and can lead to misclassification as pseudo-Cushing states[2,5]. Adrenal CS, in particular, has remained associated with a high complication rate in the 2000, despite becoming less florid[6]. Diagnosis is frequently delayed, with a mean time of 34 months from symptom onset to diagnosis[7].
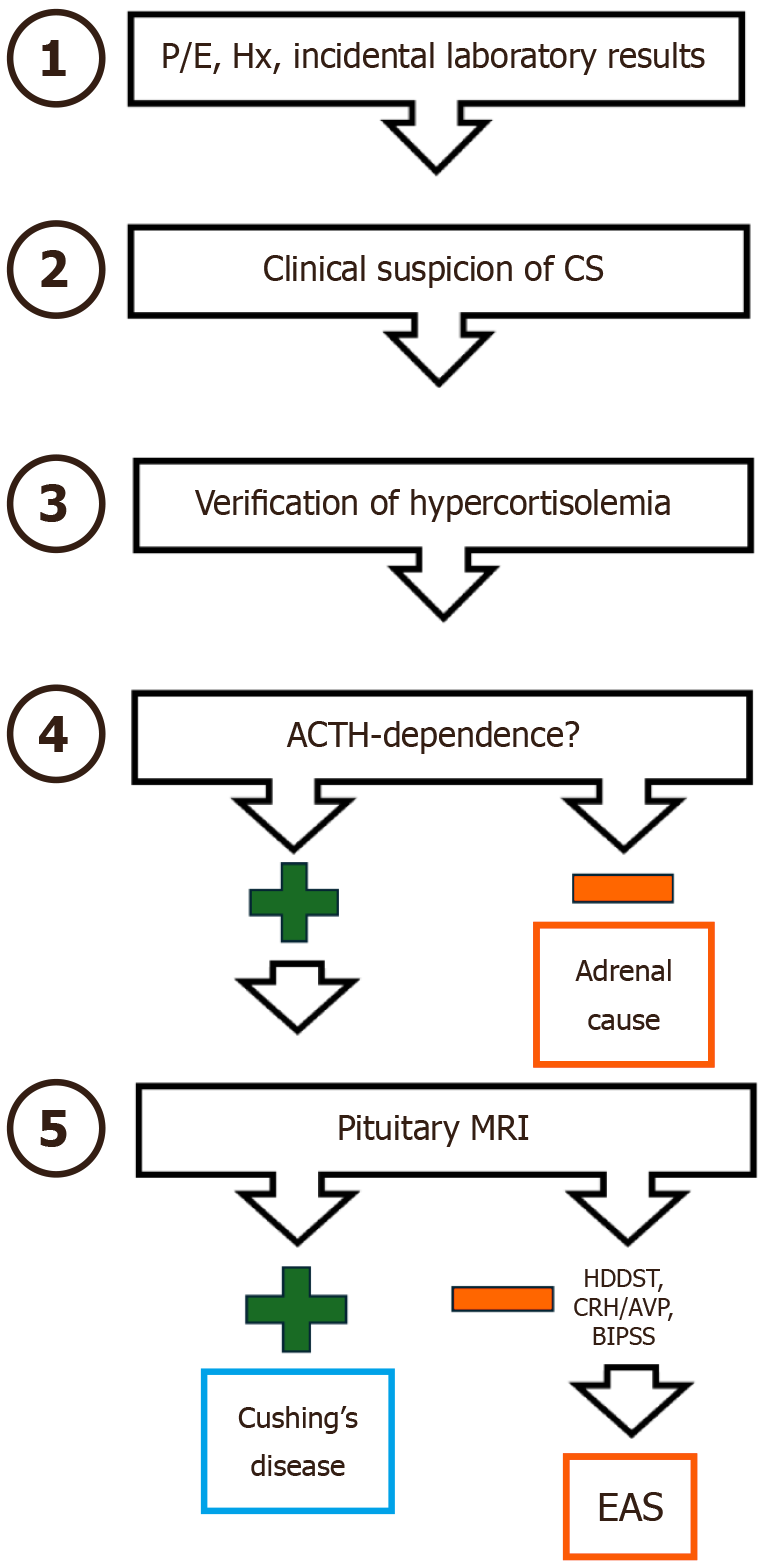
The diagnostic protocol is sequential and cost-intensive and resource-intensive, comprising preliminary screening through biochemical tests - late-night salivary cortisol, 24-hour urinary free cortisol, or 1-mg dexamethasone suppression tests - followed by advanced localization tests in the presence of adrenocorticotropic hormone (ACTH)-dependent etiologies (Figure 1)[1,2].
To separate Cushing’s disease (CD) from ectopic ACTH secretion (EAS) can necessitate corticotropin-releasing hor
Diagnostic challenges have led to growing interest in artificial intelligence (AI) as a means to provide non-invasive, rapid, and cost-effective decision support[10]. However, as machine learning tools become more integrated into clinical workflows, it is essential to reduce the risk of algorithmic bias that has the potential to undermine the goals of improved diagnostic quality and fairness[11].
In clinical medicine, the majority of diagnostic errors are attributed to cognitive rather than procedural or technical factors[12,13]. Cognitive biases - systematic departures from rational thinking - are most evident in high-stakes, time-limited, or vague clinical contexts, where doctors often use heuristics (mental shortcuts) that can lead to diagnostic failure[12]. Furthermore, cognitive biases are implicated in as many as 40%-80% of preventable deaths resulting from diagnostic or medical errors annually in the United States[14].
Among biases in human diagnosis, anchoring bias is one of the most common[12]. Clinicians place excessive value on the first information given in their way, even after later contradictory data[15]. For example, anchoring was observed when “CHF” (congestive heart failure) was spelled out in triage notes for patients with shortness of breath, leading to reduced pulmonary embolism testing and diagnostic delays[15]. Comparable limitations exist for endocrinology. Cortisol physiology and diagnostic test outcomes are significantly age, sex, ethnicity, and comorbidity-dependent, indicating assay-related bias and further reducing diagnostic accuracy across populations[1].
Beyond technical shortcomings, human cognitive biases remain a critical but underappreciated driver of misdiagnosis in CS (Table 1)[13].
| Type of bias | Mechanism of action in CS context | Clinical impact/example | Ref. | Potential mitigation strategies |
| Anchoring/intrinsic cognitive bias | Overreliance on early, salient information (heuristics) that suggest a common diagnosis (e.g., obesity or metabolic syndrome). LLMs are misled by case-intrinsic biasing information (SDFs) | Delayed diagnosis of true CS because symptoms are anchored to common benign conditions (e.g., “just obesity” or “pseudo-Cushing”). LLM accuracy declines when distracting features are present | [14,15,24] | Utilize LLM self-reflection or sequential prompting frameworks to challenge initial impressions and improve accuracy[14,24] |
| Spectrum bias/effect | Training data derived from highly specialized referral centers, skew the spectrum toward severe or advanced cases. Performance is overestimated compared to general practice populations | Diagnostic algorithms report inflated accuracy metrics when applied in diverse community settings where presentation overlaps heavily with pseudo-Cushing states | [4,5,23] | Require inclusion of representative cohorts across the full clinical spectrum and report results via subgroup analysis based on disease severity[23] |
| Exclusion/demographic bias | Exclusion of demographic factors (e.g., gender), which may be statistically irrelevant in model optimization, ignores their clinical relevance and association with diagnostic delays in real-world practice | An ML model for CS diagnosis excluded sex due to low statistical association in the training dataset[14], potentially failing to perform optimally for female subgroups who already face provider bias/stigma[30] | [4,11,30] | Employ mathematical modeling or stratification to control for demographic confounders[37]. Use adversarial debiasing or reweighting techniques to ensure equitable treatment across demographic groups[18,41] |
| Measurement bias (methodological) | The variability in laboratory methods (e.g., immunoassays vs LC-MS/MS for cortisol) used across different training centers, compromises eventual model transferability and predictive stability | A model developed using non-standardized immunoassay data from a single center[4] may perform poorly when used in a clinic relying on mass spectrometry, as results are not standardized | [4,23] | Demand transparency regarding data acquisition protocols and device/software versions used (STARD-AI items 13 and 14)[41]. Ensure dataset diversity from multiple centers with stringent protocols |
| Small sample size/class imbalance | Relying on limited samples for rare subtypes (e.g., EAS) affects model robustness and reproducibility. Reliance on simple oversampling (SMOTE) may bias accuracy | The differential diagnosis model for ACTH-dependent CS included only 26 EAS patients, limiting robustness and generalizability[8] | [4,8] | Use collaborative learning techniques across multiple centers to pool data while maintaining privacy and security[8]. Conduct multi-center, collaborative trials to achieve larger, more diverse sample sizes[4,37] |
Anchoring bias: Early attributions (e.g., weight gain explained solely by obesity) may obscure evidence of CS.
Availability bias: Common conditions such as depression or diabetes can overshadow rarer diagnoses like CS[16].
Framing bias: The way in which clinical data are presented may disproportionately shape interpretation.
Systemic bias may also persist through limitations in clinical data sources, including incomplete documentation, inaccurate coding, and inadequate representation of minority populations[11,17]. Low diversity also worsens inequities: Underrepresentation of minority groups reduces diagnostic accuracy and worsens health disparities[11,18]. For instance, Black patients with CD experience a median diagnostic delay 1.1 years longer than White patients with the same disease severity[19]. Similarly, racial and ethnic bias influences disease severity and outcomes: Hispanic/Latino and African American children with CD have greater severity and higher recurrence risk[20]. Biases can emerge through indirect indicators such as geographic localization of subjects/patients (e.g., via ZIP codes), which may also serve as surrogates for race and socioeconomic status[21]. Race has been employed as a surrogate for genetic risk or prevalence in the past, without control for structural or socioeconomic determinants of health[21]. Annotation bias introduced during clinical coding can also distort data quality. Overreliance on textbook descriptions or coding conventions can fail to capture mild or atypical CS presentations, leading to delays in diagnosis. Guidelines emphasize wide variability in presentation and the danger of false positives[22]. Adrenal CS cases in the 2000 are often less florid, making features like moon face and central obesity critical for early detection[6].
Spectrum bias is another pitfall: Methods/assays developed in “extreme spectrum” cases do not generalize well to the real-world, heterogeneous populations[5,23]. In traditional CS diagnostics, test accuracy varies between pediatric and adult populations or between mild and severe disease; population-specific interpretation is therefore recommended[22].
A prospective multicenter endocrine study confirmed knowledge deficit and cognitive biases as the main drivers of diagnostic errors[13]. Given its highly variable and often misleading presentation, CS remains particularly susceptible to diagnostic errors driven by cognitive bias, clinical misattribution, and population-level disparities.
Large language models LLMs, trained on human-authored clinical texts and published medical literature, may internalize cognitive patterns and biases in them[11,14]. LLMs are susceptible to biases in the textual data they are trained on, reflecting human cognitive biases, while traditional machine learning (ML) models for CS diagnosis are susceptible to biases in structured data (such as lab values or imaging features)[4,14]. The overall diagnostic pipeline may integrate both, and biases from either source can compromise the final output[14].
Although AI seems less susceptible to biases related to clinician fatigue or affective state on judgment compared to humans[11], investigations have discovered LLMs to be surprisingly vulnerable to errors influenced by the format or framing of the clinical presentation in the clinical narrative itself[24].
Comparative studies of Chat Generative Pre-trained Transformer (version 3.5 and version 4.0) with internal medicine residents established that the LLM’s performance was compromised when vignettes included embedded clinical cues that biased diagnostic reasoning, such as prominent but misleading clinical features (e.g., anchoring cues)[24]. For instance, if a new diagnosis was masked by salient clinical features pointing towards a common, look-alike condition, the LLM was often misled[24]. This phenomenon is attributed to LLMs’ pattern-based inference mechanisms, which can approximate the anchoring bias observed in human physicians[14]. For example, when Generative Pre-Trained Transformer-4 generated incorrect initial diagnoses for clinical vignettes, those conclusions consistently influenced its later reasoning[14].
Conversely, LLMs appeared to be less susceptible to external influences such as recency bias or patient behavior (e.g., availability of a recent similar case or patient uncooperativeness) than did human residents[24]. However, the LLM vulnerability to bias stemming from the structure and framing of clinical information - a common context in the differential diagnosis of CS where pseudo-Cushing signs are misleading[5] - is an important consideration for use in diagnostic workflows[14]. Furthermore, LLMs may exhibit suggestibility bias, prioritizing user agreement over independent reasoning, potentially leading them to adopt incorrect answers when confronted with persuasive but inaccurate prompts[14].
Due to the rarity of CS, the development of diagnostic models is constrained by limited data availability, most of which originates from tertiary referral centers[5]. The performance of a diagnostic test (sensitivity and specificity) may be spectrum-dependent, a state in which the spectrum of patients used during assessment changes[5,23]. Methods and assays that have been devised in tertiary centers have often faced severe or complicated cases[5]; when they are applied to populations in the community - where the cases are milder, heterogeneous, and usually blend with non-CS mimics - the performance measures are overstated[5,23].
In ML applications, spectrum bias is a recognized limitation in model generalizability[23]. Models developed for screening facial features of hypercortisolism or to distinguish CS from pseudo-Cushing states may suffer from limited generalizability in the absence of a clinically representative patient cohort[4,25,26]. Demographics, comorbidities, and distribution of disease severity in development and validation cohorts must be reported fully. The high prevalence of obesity in the United States population (over 40%) complicates the application of European CSSs for CS[9,27,28].
CS exhibits a demographic pattern in which 55%-60% of the cases occur in females[29]; this calls for accounting for sex-based or gender-based diagnostic differences[18]. When crafting ML models, variables that are statistically insignificant can be omitted to enhance model performance within the training dataset. In one study that built a generalized linear model to diagnose CS with 97.03% accuracy, gender was excluded after it was found to have a near-zero correlation with CS in that specific dataset[4]. Although this exclusion was statistically justified within that dataset[4], this omitted variable bias risks reproducing disparities and erasing generalizability[11,18], particularly for women whose symptoms are minimized by providers[30]. Such bias is problematic because race and sex influence CS disease severity, outcomes, and diagnostic delays[11,18].
Language-based diagnostic models may replicate cognitive biases observed in clinical reasoning.
Tendency to conform to leading prompts even when they contradict clinical evidence, as seen when clinical models adopt incorrect outputs from persuasive prompts[14].
Overemphasis on frequent clinical scenarios in the training data, leading to neglect of rare conditions such as CS[14,16].
Preference for evidence confirming initial diagnostic impressions, which can be encoded when training labels reinforce prevailing clinical assumptions, leading to premature diagnostic closure in conditions like diabetes or obesity while missing CS[14].
Variations in diagnostic interpretation based on minor differences in case framing or terminology, potentially steering clinicians away from rare diagnoses such as CS[24].
Disproportionate influence of initial clinical cues or working diagnoses, such as attributing hypertension to primary causes despite discordant endocrine findings[13,14].
Evidence suggests female patients with CS are more frequently identified by language-based diagnostic models, reflecting both true prevalence and under-recognition of atypical groups. Episodic or cyclic CS is also frequently overlooked[31]. Disease presentation and diagnostic accuracy and presentation of CS vary across sex, age, and racial groups: Black patients with CD are more often diagnosed with macroadenomas and hypopituitarism[21], and pediatric vs adult populations require age-specific diagnostic thresholds in diagnostic testing[22].
The “black-box” nature of AI systems further complicates identification of diagnostic bias, as retrospective interpretability methods rarely capture underlying computational reasoning or decision pathways[8,14]. This resembles the challenge of interpreting laboratory tests in endocrine diagnostics, where the results of cortisol and ACTH immunoassays may vary due to lack of standardization, calibration differences, and cross-reactivity[32-34]. Such variability diminishes confidence and reproducibility and highlights the need for assay-specific reference thresholds, methodological precision and careful interpretation[33].
Beyond generalized biases, algorithm-based diagnostic models for CS face methodological limitations stemming from the rarity of disease subtypes. For instance, ectopic CS accounts for only about 10% to 20% of ACTH-dependent cases[4,8]. A model developed to discriminate ACTH-dependent CS was based on only a relatively small number of EAS patients (n = 26), reflecting its clinical rarity[8]. The authors of that study used oversampling techniques (synthetic minority oversampling technique and adaptive synthetic sampling) to address class imbalance[8]. However, the authors acknowledge that small sample size limits the robustness and reproducibility of the model, stating that the reliability of algorithmic predictions is highly dependent on the size and representativeness of the training dataset[4,8]. While the oversampling used (synthetic minority oversampling technique) alleviates class imbalance, it may likely bias the accuracy since new data samples are created from old samples and cannot introduce significant changes to the dataset[8,35].
Furthermore, heterogeneity in methods of biochemical testing poses a measurement bias[33]. Cortisol measurements obtained using older methods, such as chemiluminescence immunoassays, may differ significantly from those obtained with more specific techniques like liquid chromatography-tandem mass spectrometry[4,32,36]. If a model is developed using data from a single institution using one method (e.g., immunoassays), its diagnostic accuracy may be reduced when deployed in a setting using a different methodology, limiting generalizability across clinical settings[4,23]. The generalized linear model developed by Aydemir et al[4] was developed using data from a single medical center, limiting its generalizability, as differences in testing methods across centers could affect predictive performance.
For algorithm-based tools to function as reliable adjuncts in clinical decision-making in endocrinology, researchers must seek to address explicitly the complex dynamics of biases arising from cognitive heuristics and dataset limitations. This implies rigorous adherence to traditional research methodology principles, adapted for application to algorithm-based diagnostic models (Table 1)[37].
One of the simplest methods of addressing bias in algorithmic diagnostic models is in demanding transparency, including comprehensive documentation of model creation, strengths and weaknesses, and use intention[11]. Structured model documentation tools such as model cards or datasheets for datasets provide information on model limitations[38,39], composition of development datasets, ethical concerns, and performance metrics, allowing clinicians and scientists to understand potential mistakes[38]. Specific frameworks for reporting AI studies should be adopted to ensure transparency.
DECIDE-AI: A reporting guideline focused on the early and live clinical evaluation of AI decision-support systems[40], focusing on clinical utility, safety, human factors, and preparation for subsequent larger trials[40].
Standards for Reporting of Diagnostic Accuracy studies-AI: A guideline for reporting diagnostic accuracy studies using AI[41], which includes items specifically requiring details on dataset practices, data acquisition protocols, performance error analysis, and assessments of algorithmic bias and fairness (item 23 and item 35, respectively)[41].
Interpretable modeling techniques, including SHapley Additive exPlanations (SHAP)[8], may be implemented to show the diagnostic contribution of individual model features[8]. For instance, SHAP results were found to be vital in predicting ACTH-dependent CS etiology, confirming the contribution of variables like suppression in high-dose dexa
New diagnostic algorithms should be designed in accordance with strict epidemiological research standards[37].
Preventing spectrum bias: To minimize spectrum bias, algorithmic model development requires inclusion of a representative clinical spectrum, especially non-CS control groups that include a representative spectrum of those seen in clinical practice[5,23]. Subgroup analysis by patient characteristics and disease severity should be included as part of studies, and these results should become part of the performance interpretation[23].
Controlling for confounding and demographics: Methods of controlling confounding variables must be employed[37]. While methods like restriction limit generalization[37], mathematical modeling (for example, logistic regression) and stratification are preferred to control for possible confounders, especially for demographic characteristics like age, sex, and socioeconomic status[18,37]. Researchers must avoid the temptation to exclude demographic variables due to lack of significance within a restricted sample[4]. Furthermore, advanced strategies such as adversarial debiasing or reweighting can be utilized to address issues of fairness by preventing the AI system from systematically underperforming or misclassifying specific subgroups[18,41].
Data diversity and sample size maintenance: As CS subtypes such as EAS are rare, explicit restriction by small datasets directly affects the reproducibility and stability of ML-based diagnostic models[8]. Follow-up studies should then be directed toward multi-center, collaborative trials using stringent protocols to achieve larger, more diverse sample sizes and increase the generalizability of the findings[37]. The ethical issue of enrolling representative populations must be resolved methodically to avoid developing biased model outputs[11,18]. When developing models for rare diseases like CS, specific strategies for multicenter collaboration, such as federated or collaborative learning, should be explored to enable the pooling of data while maintaining data confidentiality and fragmentation[8].
The integration of LLMs and ML-based diagnostic algorithms into the diagnostic workflow for CS has the potential to overcome the inherent complexity and the human fallibility associated with this rare endocrine disease. Algorithmic tools have already demonstrated their potential in non-invasive risk stratification and differentiation of ACTH-dependent etiologies[8,25,26]. The development of specialized CSSs tailored to specific populations, such as the United States-based RAPID CD CSS, further demonstrates the utility of ML/AI in accelerating diagnosis[9].
However, these advanced diagnostic systems are prone to bias, reflecting the cognitive limitations observed in human clinical reasoning[14,24] and introducing new system-level errors due to dataset bias and methodological limitations[4,5]. Despite less florid clinical presentations in adrenal CS in recent decades, the high prevalence of CS-related complications persists, necessitating new models for early detection[6].
For the endocrine community, the future entails not just the implementation of algorithm-based diagnostics, but also the critical scrutiny of their development and application. Diagnostic models should be grounded in high-quality, heterogeneous datasets that resolve the spectrum effect[5] and do not quietly exclude, but actively incorporate, clinically relevant demographic factors such as sex and race[4,11]. The large-scale deployment of structured transparency tools (e.g., Developmental and Exploratory Clinical Investigations of DEcision support systems driven-AI, Standards for Reporting of Diagnostic Accuracy studies-AI)[40,41] and interpretable modeling techniques (e.g., SHAP)[8] will support trust and ensure that language-based models and structured ML models serve as fair and accountable assistants, accelerating accurate diagnosis of CS and ultimately benefiting all patient populations.
| 1. | Fleseriu M, Auchus R, Bancos I, Ben-Shlomo A, Bertherat J, Biermasz NR, Boguszewski CL, Bronstein MD, Buchfelder M, Carmichael JD, Casanueva FF, Castinetti F, Chanson P, Findling J, Gadelha M, Geer EB, Giustina A, Grossman A, Gurnell M, Ho K, Ioachimescu AG, Kaiser UB, Karavitaki N, Katznelson L, Kelly DF, Lacroix A, McCormack A, Melmed S, Molitch M, Mortini P, Newell-Price J, Nieman L, Pereira AM, Petersenn S, Pivonello R, Raff H, Reincke M, Salvatori R, Scaroni C, Shimon I, Stratakis CA, Swearingen B, Tabarin A, Takahashi Y, Theodoropoulou M, Tsagarakis S, Valassi E, Varlamov EV, Vila G, Wass J, Webb SM, Zatelli MC, Biller BMK. Consensus on diagnosis and management of Cushing's disease: a guideline update. Lancet Diabetes Endocrinol. 2021;9:847-875. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 767] [Cited by in RCA: 647] [Article Influence: 129.4] [Reference Citation Analysis (3)] |
| 2. | Nieman LK. Cushing's syndrome: update on signs, symptoms and biochemical screening. Eur J Endocrinol. 2015;173:M33-M38. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 237] [Cited by in RCA: 180] [Article Influence: 16.4] [Reference Citation Analysis (0)] |
| 3. | Braun LT, Riester A, Oßwald-Kopp A, Fazel J, Rubinstein G, Bidlingmaier M, Beuschlein F, Reincke M. Toward a Diagnostic Score in Cushing's Syndrome. Front Endocrinol (Lausanne). 2019;10:766. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 73] [Cited by in RCA: 55] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 4. | Aydemir M, Çakir M, Oral O, Yilmaz M. Diagnosis of Cushing's syndrome with generalized linear model and development of mobile application. Medicine (Baltimore). 2025;104:e42910. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 1] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 5. | Ayala AR, Ilias I, Nieman LK. The spectrum effect in the evaluation of Cushing syndrome. Curr Opin Endocrinol Diabetes Obes. 2003;10:272-276. [RCA] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 7] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 6. | Katabami T, Asai S, Matsuba R, Sone M, Izawa S, Ichijo T, Tsuiki M, Okamura S, Yoshimoto T, Otsuki M, Takeda Y, Naruse M, Tanabe A; ACPA-J Study Group. Changes in clinical features of adrenal Cushing syndrome: a national registry study. Endocr Connect. 2025;14:e240684. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 3] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 7. | Rubinstein G, Osswald A, Hoster E, Losa M, Elenkova A, Zacharieva S, Machado MC, Hanzu FA, Zopp S, Ritzel K, Riester A, Braun LT, Kreitschmann-Andermahr I, Storr HL, Bansal P, Barahona MJ, Cosaro E, Dogansen SC, Johnston PC, Santos de Oliveira R, Raftopoulos C, Scaroni C, Valassi E, van der Werff SJA, Schopohl J, Beuschlein F, Reincke M. Time to Diagnosis in Cushing's Syndrome: A Meta-Analysis Based on 5367 Patients. J Clin Endocrinol Metab. 2020;105:dgz136. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 130] [Cited by in RCA: 105] [Article Influence: 17.5] [Reference Citation Analysis (0)] |
| 8. | Demir AN, Ayata D, Oz A, Sulu C, Kara Z, Sahin S, Ozaydin D, Korkmazer B, Arslan S, Kizilkilic O, Ciftci S, Celik O, Ozkaya HM, Tanriover N, Gazioglu N, Kadioglu P. Machine Learning May Be an Alternative to BIPSS in the Differential Diagnosis of ACTH-dependent Cushing Syndrome. J Clin Endocrinol Metab. 2025;110:e412-e422. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 4] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 9. | Salcedo-Sifuentes JE, Shih R, Heaney AP, Bergsneider M, Wang MB, Donangelo I, Lee J, Delery W, Karsy M, Kshettry VR, Yuen KCJ, Evans JJ, Barkhoudarian G, Pacione DR, Gardner PA, Fernandez-Miranda JC, Benjamin C, Zada G, Rennert RC, Silverstein JM, Chicoine MR, Kim J, Li G, Little AS, Kim W. Cushing Disease Clinical Phenotype and Tumor Behavior Vary With Age: Diagnostic and Perioperative Implications. J Clin Endocrinol Metab. 2025;110:2595-2604. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 10. | Laws ER, Catalino MP. Editorial. Machine learning and artificial intelligence applied to the diagnosis and management of Cushing disease. Neurosurg Focus. 2020;48:E6. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 10] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 11. | Williams NH. Artificial Intelligence and Healthcare. The International Library of Bioethics. Cham: Springer, 2023. [DOI] [Full Text] |
| 12. | Cunningham N, Cook H, Leach D, Klein J, Harrison J. Demystifying cognitive bias in the diagnostic process for frontline clinicians and educators; new words for old ideas. Diagnosis (Berl). 2025;12:322-332. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 5] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 13. | Frey J, Braun LT, Handgriff L, Kendziora B, Fischer MR, Reincke M, Zwaan L, Schmidmaier R. Insights into diagnostic errors in endocrinology: a prospective, case-based, international study. BMC Med Educ. 2023;23:934. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 2] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 14. | Mahajan A, Obermeyer Z, Daneshjou R, Lester J, Powell D. Cognitive bias in clinical large language models. NPJ Digit Med. 2025;8:428. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 15] [Article Influence: 15.0] [Reference Citation Analysis (0)] |
| 15. | Ly DP, Shekelle PG, Song Z. Evidence for Anchoring Bias During Physician Decision-Making. JAMA Intern Med. 2023;183:818-823. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 68] [Cited by in RCA: 55] [Article Influence: 18.3] [Reference Citation Analysis (0)] |
| 16. | Gómez-Gutiérrez MA, Huertas-Cañas JM, Bedoya-Ossa A. From Knee Pain Consultation to Pituitary Surgery: The Challenge of Cushing Disease Diagnosis. JCEM Case Rep. 2024;2:luae048. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 17. | Mohammed S, Budach L, Feuerpfeil M, Ihde N, Nathansen A, Noack N, Patzlaff H, Naumann F, Harmouch H. The effects of data quality on machine learning performance on tabular data. Inf Syst. 2025;132:102549. [DOI] [Full Text] |
| 18. | Cirillo D, Catuara-Solarz S, Morey C, Guney E, Subirats L, Mellino S, Gigante A, Valencia A, Rementeria MJ, Chadha AS, Mavridis N. Sex and gender differences and biases in artificial intelligence for biomedicine and healthcare. NPJ Digit Med. 2020;3:81. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 423] [Cited by in RCA: 244] [Article Influence: 40.7] [Reference Citation Analysis (0)] |
| 19. | Omotosho YB, Mcglotten R, Chittiboina P, Nieman L. MON-113 Racial Disparity in the Time to Diagnosis of Cushing’s Disease. J Endocr Soc. 2024;8:bvae163.2408. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 1] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 20. | Gkourogianni A, Sinaii N, Jackson SH, Karageorgiadis AS, Lyssikatos C, Belyavskaya E, Keil MF, Zilbermint M, Chittiboina P, Stratakis CA, Lodish MB. Pediatric Cushing disease: disparities in disease severity and outcomes in the Hispanic and African-American populations. Pediatr Res. 2017;82:272-277. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 22] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 21. | Ioachimescu AG, Goswami N, Handa T, Pappy A, Veledar E, Oyesiku NM. Racial Disparities in Acromegaly and Cushing's Disease: A Referral Center Study in 241 Patients. J Endocr Soc. 2022;6:bvab176. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 9] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 22. | Nieman LK, Biller BM, Findling JW, Newell-Price J, Savage MO, Stewart PM, Montori VM. The diagnosis of Cushing's syndrome: an Endocrine Society Clinical Practice Guideline. J Clin Endocrinol Metab. 2008;93:1526-1540. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2133] [Cited by in RCA: 1739] [Article Influence: 96.6] [Reference Citation Analysis (0)] |
| 23. | Tseng AS, Shelly-Cohen M, Attia IZ, Noseworthy PA, Friedman PA, Oh JK, Lopez-Jimenez F. Spectrum bias in algorithms derived by artificial intelligence: a case study in detecting aortic stenosis using electrocardiograms. Eur Heart J Digit Health. 2021;2:561-567. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 15] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 24. | Schmidt HG, Rotgans JI, Mamede S. Bias Sensitivity in Diagnostic Decision-Making: Comparing ChatGPT with Residents. J Gen Intern Med. 2025;40:790-795. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 9] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 25. | Wei R, Jiang C, Gao J, Xu P, Zhang D, Sun Z, Liu X, Deng K, Bao X, Sun G, Yao Y, Lu L, Zhu H, Wang R, Feng M. Deep-Learning Approach to Automatic Identification of Facial Anomalies in Endocrine Disorders. Neuroendocrinology. 2020;110:328-337. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 39] [Cited by in RCA: 30] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 26. | Qiang J, Liu H, Guo X, Song C, Lu L, Long X, Pan H, Zhao Q, Huang J, Li J, Chen S. Deep Learning-based Multiview Facial Identification as a Screening Tool for Cushing Syndrome. Endocr Pract. 2025;31:1601-1607. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 27. | Stierman B, Afful J, Carroll MD, Chen TC, Davy O, Fink S, Fryar CD, Gu Q, Hales CM, Hughes JP, Ostchega Y, Storandt RJ, Akinbami LJ. National Health and Nutrition Examination Survey 2017-March 2020 Prepandemic Data Files-Development of Files and Prevalence Estimates for Selected Health Outcomes. Natl Health Stat Report. 2021. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 284] [Cited by in RCA: 285] [Article Influence: 57.0] [Reference Citation Analysis (0)] |
| 28. | NCD Risk Factor Collaboration (NCD-RisC). Worldwide trends in underweight and obesity from 1990 to 2022: a pooled analysis of 3663 population-representative studies with 222 million children, adolescents, and adults. Lancet. 2024;403:1027-1050. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1583] [Cited by in RCA: 1359] [Article Influence: 679.5] [Reference Citation Analysis (15)] |
| 29. | Reincke M, Fleseriu M. Cushing Syndrome: A Review. JAMA. 2023;330:170-181. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 203] [Cited by in RCA: 156] [Article Influence: 52.0] [Reference Citation Analysis (0)] |
| 30. | Jones SC, Nutter S, Saunders JF. "The healthcare system did fail me repeatedly": a qualitative study on experiences of healthcare among Canadian women with Cushing's syndrome. BMC Prim Care. 2024;25:329. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 31. | Świątkowska-Stodulska R, Berlińska A, Stefańska K, Kłosowski P, Sworczak K. Cyclic Cushing's Syndrome - A Diagnostic Challenge. Front Endocrinol (Lausanne). 2021;12:658429. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 16] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 32. | Casals G, Hanzu FA. Cortisol Measurements in Cushing's Syndrome: Immunoassay or Mass Spectrometry? Ann Lab Med. 2020;40:285-296. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 74] [Cited by in RCA: 62] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 33. | Flowers KC, Shipman KE. Pitfalls in the Diagnosis and Management of Hypercortisolism (Cushing Syndrome) in Humans; A Review of the Laboratory Medicine Perspective. Diagnostics (Basel). 2023;13:1415. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 10] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 34. | Brixey-Mccann R, Tennant S, Geen J, Armston A, Barth JH, Keevil BG, Rees A, Evans C. Effect of cortisol assay bias on the overnight dexamethasone suppression test: implications for the investigation of Cushing's syndrome. Endocr Abstr. 2015. [DOI] [Full Text] |
| 35. | Chawla NV, Bowyer KW, Hall LO, Kegelmeyer WP. SMOTE: Synthetic Minority Over-sampling Technique. J Artif Intell Res. 2002;16:321-357. [RCA] [DOI] [Full Text] [Cited by in Crossref: 10955] [Cited by in RCA: 7310] [Article Influence: 304.6] [Reference Citation Analysis (0)] |
| 36. | Taylor DR, Ghataore L, Couchman L, Vincent RP, Whitelaw B, Lewis D, Diaz-Cano S, Galata G, Schulte KM, Aylwin S, Taylor NF. A 13-Steroid Serum Panel Based on LC-MS/MS: Use in Detection of Adrenocortical Carcinoma. Clin Chem. 2017;63:1836-1846. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 101] [Cited by in RCA: 83] [Article Influence: 9.2] [Reference Citation Analysis (0)] |
| 37. | Collins GS, Moons KGM, Dhiman P, Riley RD, Beam AL, Van Calster B, Ghassemi M, Liu X, Reitsma JB, van Smeden M, Boulesteix AL, Camaradou JC, Celi LA, Denaxas S, Denniston AK, Glocker B, Golub RM, Harvey H, Heinze G, Hoffman MM, Kengne AP, Lam E, Lee N, Loder EW, Maier-Hein L, Mateen BA, McCradden MD, Oakden-Rayner L, Ordish J, Parnell R, Rose S, Singh K, Wynants L, Logullo P. TRIPOD+AI statement: updated guidance for reporting clinical prediction models that use regression or machine learning methods. BMJ. 2024;385:e078378. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1587] [Cited by in RCA: 1487] [Article Influence: 743.5] [Reference Citation Analysis (7)] |
| 38. | Mitchell M, Wu S, Zaldivar A, Barnes P, Vasserman L, Hutchinson B, Spitzer E, Raji ID, Gebru T. Model Cards for Model Reporting. FAT* '19: Proceedings of the Conference on Fairness, Accountability, and Transparency; 2019 Jan 29-31; NY, United States. Association for Computing Machinery, 2019: 220-229. [DOI] [Full Text] |
| 39. | Gebru T, Morgenstern J, Vecchione B, Vaughan JW, Wallach H, Iii HD, Crawford K. Datasheets for datasets. Commun ACM. 2021;64:86-92. [DOI] [Full Text] |
| 40. | Vasey B, Nagendran M, Campbell B, Clifton DA, Collins GS, Denaxas S, Denniston AK, Faes L, Geerts B, Ibrahim M, Liu X, Mateen BA, Mathur P, McCradden MD, Morgan L, Ordish J, Rogers C, Saria S, Ting DSW, Watkinson P, Weber W, Wheatstone P, McCulloch P; DECIDE-AI expert group. Reporting guideline for the early stage clinical evaluation of decision support systems driven by artificial intelligence: DECIDE-AI. BMJ. 2022;377:e070904. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 281] [Cited by in RCA: 210] [Article Influence: 52.5] [Reference Citation Analysis (0)] |
| 41. | Sounderajah V, Guni A, Liu X, Collins GS, Karthikesalingam A, Markar SR, Golub RM, Denniston AK, Shetty S, Moher D, Bossuyt PM, Darzi A, Ashrafian H; STARD-AI Steering Committee. The STARD-AI reporting guideline for diagnostic accuracy studies using artificial intelligence. Nat Med. 2025;31:3283-3289. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 87] [Cited by in RCA: 76] [Article Influence: 76.0] [Reference Citation Analysis (0)] |