Published online Mar 25, 2026. doi: 10.5501/wjv.v15.i1.118988
Revised: February 10, 2026
Accepted: March 9, 2026
Published online: March 25, 2026
Processing time: 53 Days and 22.6 Hours
The coronavirus disease 2019 (COVID-19) pandemic remains a major global public health threat, and ongoing viral mutations continue to complicate control efforts. To inform local prevention strategies, this study investigated the epidemiological and molecular characteristics of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in Lu’an city from 2020 to 2022, and analyzed their association with clinical outcomes.
To analyze the molecular epidemiology and risk factors for severe COVID-19, and provide a scientific basis for guiding local epidemic prevention and control strategies.
Biological samples were collected from confirmed COVID-19 patients in Lu’an city between 2020 and 2022. Complete SARS-CoV-2 genomic sequences were obtained through sequencing. Epidemiological and clinical data were collected concurrently for each patient. Statistical analyses were conducted using IBM SPSS 29.0 software to assess risk factors associated with severe COVID-19. Viral genomic sequences were analyzed using MEGA software to characterize the molecular features of circulating SARS-CoV-2 strains.
Sequencing identified the original SARS-CoV-2 strain in samples from 2020-2021, while the Omicron variant was detected in samples from 2022. The predominant clinical manifestations among patients were cough (67.03%) and fever (65.56%). Laboratory and imaging examinations revealed that 78.85% of infected patients exhibited abnor
The development of severe COVID-19 was predominantly associated with pre-existing comorbidities rather than with SARS-CoV-2 variant type. These findings provide evidence to inform targeted clinical management and public health planning for vulnerable populations.
Core Tip: To conduct whole-genome sequencing of the severe acute respiratory syndrome coronavirus 2 strains circulating in Lu’an city, analyze the association between different genotypes and major clinical symptoms, and investigate the relationship between severe coronavirus disease 2019 and both viral genotypes and patients’ underlying diseases.
- Citation: Chang HW, Yang W, Yu L, Gao DW, Zhang F, Chen ZC, Chen BL, Zhang LM, Zhu R, Zhang Q, Li ZY, Rao JG. Study on the correlation between molecular characteristics of SARS-CoV-2 and its epidemiology and clinical manifestations in Lu’an city. World J Virol 2026; 15(1): 118988
- URL: https://www.wjgnet.com/2220-3249/full/v15/i1/118988.htm
- DOI: https://dx.doi.org/10.5501/wjv.v15.i1.118988
In December 2019, cases of pneumonia of unknown etiology were reported in several hospitals in Wuhan city, Hubei Province, China[1]. Pharyngeal and nasal swab samples were collected from affected patients, and the causative viruses were identified and the full genomic sequences were determined. The novel coronavirus, which has never been found in humans before, was identified. The International Committee on Taxonomy of Viruses then named it severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)[2]. On February 8, 2020, the National Health Commission temporarily named the associated disease “Novel Coronavirus Pneumonia”, and on February 11, 2020, the World Health Organization officially named it coronavirus disease 2019 (COVID-19)[3].
The pneumonia epidemic caused by the original strain of SARS-CoV-2 reached its peak in China from late January to early February 2020, with the majority of cases occurring in Hubei Province and later spreading to other regions[4]. Due to international travel and population mobility, COVID-19 subsequently spread worldwide, placing a substantial burden on global health systems and economies. The annual nucleotide mutation rate of the full SARS-CoV-2 genome is 6.677 × 10-4[5]. As the COVID-19 pandemic progressed, multiple viral variant strains emerged, and various types of vaccines have been successively introduced, ushering the world into an unprecedented phase of COVID-19 prevention and control[6].
On January 20, 2020, the laboratory of the Lu’an Municipal Center for Disease Control and Prevention (CDC) detected a positive result for SARS-CoV-2 nucleic acid in a sputum sample from a suspected COVID-19 patient at the Lu’an People’s Hospital, representing the first confirmed COVID-19 case in the city[7]. By the end of 2022, four major epidemic waves had occurred in the city, two waves were caused by the original SARS-CoV-2 strain and the other two by the Omicron variant.
Whole-genome nucleotide sequencing was performed on SARS-CoV-2 nucleic acid-positive samples and the obtained sequences were analyzed to determine the different clinical manifestations of various SARS-CoV-2 genotypes in this study. Furthermore, whole-genome sequencing and nucleotide polymorphism analysis of the SARS-CoV-2 virus from clustered cases were performed to clarify the transmission chain relationship among clustered cases, thereby providing a scientific method for epidemic traceability.
A total of ninety-one complete SARS-CoV-2 genome sequences were obtained from laboratory-confirmed patients admitted to designated hospitals within the jurisdiction of Lu’an city. All samples were collected between 2020 and 2022, and sequencing was performed by the Lu’an Municipal Center for Disease CDC using either second- or third-generation sequencing platforms. Case diagnoses adhered to the criteria specified in the Diagnosis and Treatment Protocol for Novel Coronavirus Pneumonia issued by the National Health Commission of China. Sputum and throat swab specimens were collected by relevant medical institutions and district- or county-level CDCs, stored under appropriate conditions, and transported to the Lu’an municipal CDC laboratory for testing in accordance with the Novel Coronavirus Pneumonia Prevention and Control Protocol.
Data collection: Epidemiological data were extracted from standardized case investigation forms for all confirmed COVID-19 cases within Lu’an city. Clinical data were primarily retrieved from the Infectious Diseases Department of Lu’an Municipal People’s Hospital and other designated county-level COVID-19 treatment hospitals. For the 91 included cases, demographic information (sex, age, date of symptom onset, case source, and exposure history) and clinical characteristics were compiled and analyzed.
Nucleic acid detection: Viral RNA was extracted using the BTK-96 Nucleic Acid Extractor and corresponding extraction kits (Wuxi Biotech Co., Ltd.) following the national prevention and control protocol[8]. Reverse transcription polymerase chain reaction amplification was performed on an ABI 7500 real-time polymerase chain reaction system[9] using the SARS-CoV-2 Nucleic Acid Detection Kit (Beijing Zhuocheng Huisheng Biotechnology Co., Ltd.). All experimental procedures and result interpretations strictly followed the manufacturers’ instructions. Equipment was calibrated regularly, reagents were used within their validity periods, and each batch included appropriate quality controls.
Library preparation and whole-genome sequencing: For a subset of SARS-CoV-2-positive samples with adequate viral load, extracted RNA was used as the template for whole-genome amplification using the SARS-CoV-2 Whole Genome Capture Kit (Hangzhou Baiyi Company). Amplified products were quantified with a Qubit 4.0 Fluorometer. Sequencing libraries were prepared with the Ligation Sequencing Kit (SQK-LSK109; Oxford Nanopore Technologies). A portion of the samples was sequenced on an Oxford Nanopore Mk1C system using R9.4.1 flow cells. The remaining samples were sequenced on an Illumina MiSeq platform using Infinium Automation Kit-8 Tip Tecan LIMS Ready flow cells[10].
Whole-genome sequence assembly and analysis: For Nanopore sequencing, runs exceeding 12 hours were monitored using MinKNOW software (v19.12.5)[11]. Raw FAST5 data were base-called to FASTQ format using Guppy (v4.2.2). Quality assessment was performed with NanoPlot (v1.30.0), and reads shorter than 200 bp or of low quality were filtered using Filtlong (v0.2.0). Cleaned reads were aligned to the SARS-CoV-2 reference genome Wuhan-Hu-1 (GenBank: MN908947) and assembled with Medaka (v1.1.2), which was also used for variant calling[12].
For Illumina data, sequencing-by-synthesis was conducted for over 10 hours[13]. Subsequent bioinformatic processing followed a similar alignment and assembly workflow. Final genome sequences were genotyped using MEGA software (v4.0)[14].
Data were compiled in Microsoft Excel 2016 and analyzed using IBM SPSS Statistics (v29.0). Categorical variables were summarized as counts and percentages. Group comparisons were performed using the χ2 test or Fisher’s exact test, as appropriate. A two-sided P < 0.05 was considered statistically significant.
The demographic and clinical features of 91 laboratory-confirmed COVID-19 patients treated in designated hospitals in Lu’an city between 2020 and 2022 are summarized in Table 1. The three most common clinical manifestations were abnormal findings on chest computed tomography (78.41%, 69/89), cough (67.03%, 61/91), and fever (65.56%, 59/90). The prevalence of each of these manifestations varied significantly across the three study years (all P < 0.01).
| Item | 2020 | 2021 | 2022 | Total | Statistics of different years | |||||
| n | % | n | % | n | % | n | % | χ2 | P value | |
| Cough | 47 | 90.38 (47/52) | 5 | 31.25 (5/16) | 9 | 39.13 (9/23) | 61 | 67.03 (61/91) | 30.205 | < 0.001 |
| Fever | 47 | 90.38 (47/52) | 4 | 25.00 (4/16) | 8 | 36.36 (8/22) | 59 | 65.56 (59/90) | 34.154 | < 0.001 |
| Chest tightness/chest discomfort | 22 | 47.83 (22/46) | 0 | 0 (0/8) | 0 | 0 (0/15) | 22 | 31.88 (22/69) | 16.149 | < 0.001 |
| Chills | 1 | 8.33 (1/12) | 0 | 0 (0/1) | 1 | 20.00 (1/5) | 2 | 11.11 (2/18) | 0.619 | 0.734 |
| Feeling cold/rigors | 4 | 26.67 (4/15) | 4 | 25.00 (4/16) | 0 | 0 (0/4) | 8 | 22.86 (8/35) | 1.350 | 0.509 |
| Mild fatigue | 6 | 31.58 (6/19) | 0 | 0 (0/1) | 2 | 40.00 (2/5) | 8 | 32.00 (8/25) | 0.619 | 0.734 |
| Headache | 3 | 17.65 (3/17) | 3 | 33.33 (3/9) | 0 | 0 (0/12) | 6 | 15.79 (6/38) | 4.377 | 0.112 |
| Chest pain | 2 | 9.52 (2/21) | 0 | 0 (0/14) | 0 | 0 (0/23) | 2 | 3.45 (2/58) | 3.650 | 0.161 |
| Vomiting | 4 | 12.50 (4/32) | 1 | 6.25 (1/16) | 0 | 0 (0/23) | 5 | 7.04 (5/71) | 2.923 | 0.232 |
| Nausea | 8 | 22.22 (8/36) | 1 | 6.25 (1/16) | 1 | 4.35 (1/23) | 10 | 13.33 (10/75) | 3.981 | 0.137 |
| Abdominal pain | 4 | 14.29 (4/28) | 0 | 0 (0/16) | 1 | 4.35 (1/23) | 5 | 7.46 (5/67) | 3.501 | 0.174 |
| Diarrhea | 12 | 37.50 (12/32) | 5 | 31.25 (5/16) | 0 | 0 (0/23) | 17 | 23.94 (17/71) | 10.939 | 0.004 |
| Convulsion/impaired consciousness | 1 | 2.00 (1/50) | 0 | 0 (0/16) | 0 | 0 (0/23) | 1 | 11.24 (10/89) | 0.789 | 0.674 |
| Poor mental state, appetite, or sleep | 24 | 46.15 (24/52) | 8 | 50.00 (8/16) | 3 | 13.04 (3/23) | 35 | 38.46 (35/91) | 8.478 | 0.014 |
| Lymphadenopathy | 0 | 0 (0/52) | 0 | 0 (0/16) | 0 | 0 (0/23) | 0 | 0 (0/91) | - | - |
| CT findings/CT abnormalities | 47 | 90.38 (47/52) | 12 | 80.00 (12/15) | 10 | 47.62 (10/21) | 69 | 78.41 (69/88) | 16.187 | < 0.001 |
| Pharyngeal redness | 27 | 51.92 (27/52) | 10 | 62.50 (10/16) | 12 | 57.14 (12/21) | 49 | 55.06 (49/89) | 0.602 | 0.74 |
| Tonsillar enlargement | 1 | 1.92 (1/52) | 1 | 6.25 (1/16) | 10 | 43.48 (10/23) | 12 | 13.19 (12/91) | 24.870 | < 0.001 |
| Blood oxygen saturation | 30 | 81.08 (30/37) | 2 | 12.50 (2/16) | 0 | 0 (0/23) | 32 | 42.11 (32/76) | 45.538 | < 0.001 |
| Sore throat | 4 | 22.22 (4/18) | 3 | 50.00 (3/6) | 7 | 63.64 (7/11) | 14 | 40.00 (14/35) | 5.181 | 0.075 |
| Brudzinski’s sign | 0 | 0 (0/52) | 0 | 0 (0/16) | 0 | 0 (0/22) | 0 | 0 (0/90) | - | - |
| Kernig’s sign | 0 | 0 (0/52) | 0 | 0 (0/16) | 0 | 0 (0/22) | 0 | 0 (0/90) | - | - |
| Babinski’s sign | 0 | 0 (0/52) | 0 | 0 (0/16) | 0 | 0 (0/22) | 0 | 0(0/90) | - | - |
| CRP | 28 | 60.87 (28/46) | 7 | 43.75 (7/16) | 4 | 23.53 (4/17) | 39 | 49.37 (39/79) | 7.177 | 0.028 |
| Myalgia | 1 | 3.03 (1/33) | 0 | 0 (0/14) | 0 | 0 (0/16) | 1 | 15.87 (10/63) | 0.924 | 0.63 |
| Elevated WBC count | 2 | 3.85 (2/52) | 3 | 18.75 (3/16) | 0 | 0 (0/18) | 5 | 5.81 (5/86) | 6.368 | 0.014 |
| Decreased WBC count | 6 | 11.54 (6/52) | 1 | 6.25 (1/16) | 3 | 16.67 (3/18) | 10 | 11.63 (10/86) | 0.895 | 0.639 |
| Clinical classification | 15.100 | 0.001 | ||||||||
| Asymptomatic | 0 | 0 (0/52) | 3 | 18.75 (3/16) | 5 | 21.74 (5/23) | 8 | 8.79 (8/91) | 11.800 | 0.003 |
| Mild | 5 | 9.62 (5/52) | 2 | 12.5 (2/16) | 8 | 34.78 (8/23) | 15 | 16.48 (15/91) | 7.561 | 0.023 |
| Moderate | 41 | 78.85 (41/52) | 11 | 68.75 (11/16) | 10 | 43.48 (10/23) | 62 | 68.13 (62/91) | 11.069 | 0.004 |
| Severe | 6 | 11.54 (6/52) | 0 | 0 (0/16) | 0 | 0 (0/23) | 6 | 6.59 (6/91) | 4.818 | 0.09 |
Excluding asymptomatic cases, patients were categorized as mild, moderate, severe, or critical according to the national “Diagnosis and Treatment Protocol for COVID-19 (Trial Version 10)”. Logistic regression analysis of the 91 patients with whole-genome sequencing data indicated that age and the presence of comorbidities were independent risk factors for severe COVID-19. The odds of severe disease increased with age (odds ratio = 1.05, 95% confidence interval: 1.04-1.06).
Of the 91 patients, 48 were male and 43 were female. Severe disease occurred in 6 patients (five males and one female), with no statistically significant difference in severity between sexes (χ2 = 1.276, P = 0.259).
Eight patients had underlying medical conditions, including hypertension, diabetes, or liver impairment. Among these patients, four developed severe disease, corresponding to a severe disease incidence of 50%. Patients with such comorbidities exhibited a significantly elevated risk of severe COVID-19 (rate ratio = 9.625, 95% confidence interval: 2.960-31.300), and the difference in severe illness rates between patients with and without comorbidities was statistically significant (χ2 = 12.213, P < 0.001).
In 2020, the severe illness rate among patients infected with the ancestral SARS-CoV-2 strain was 11.54% (6/52). By contrast, none of the 16 patients infected in 2021 (n = 16) or 2022 (n = 23) progressed to severe disease, suggesting that viral variant type was not a primary determinant of disease severity in this cohort.
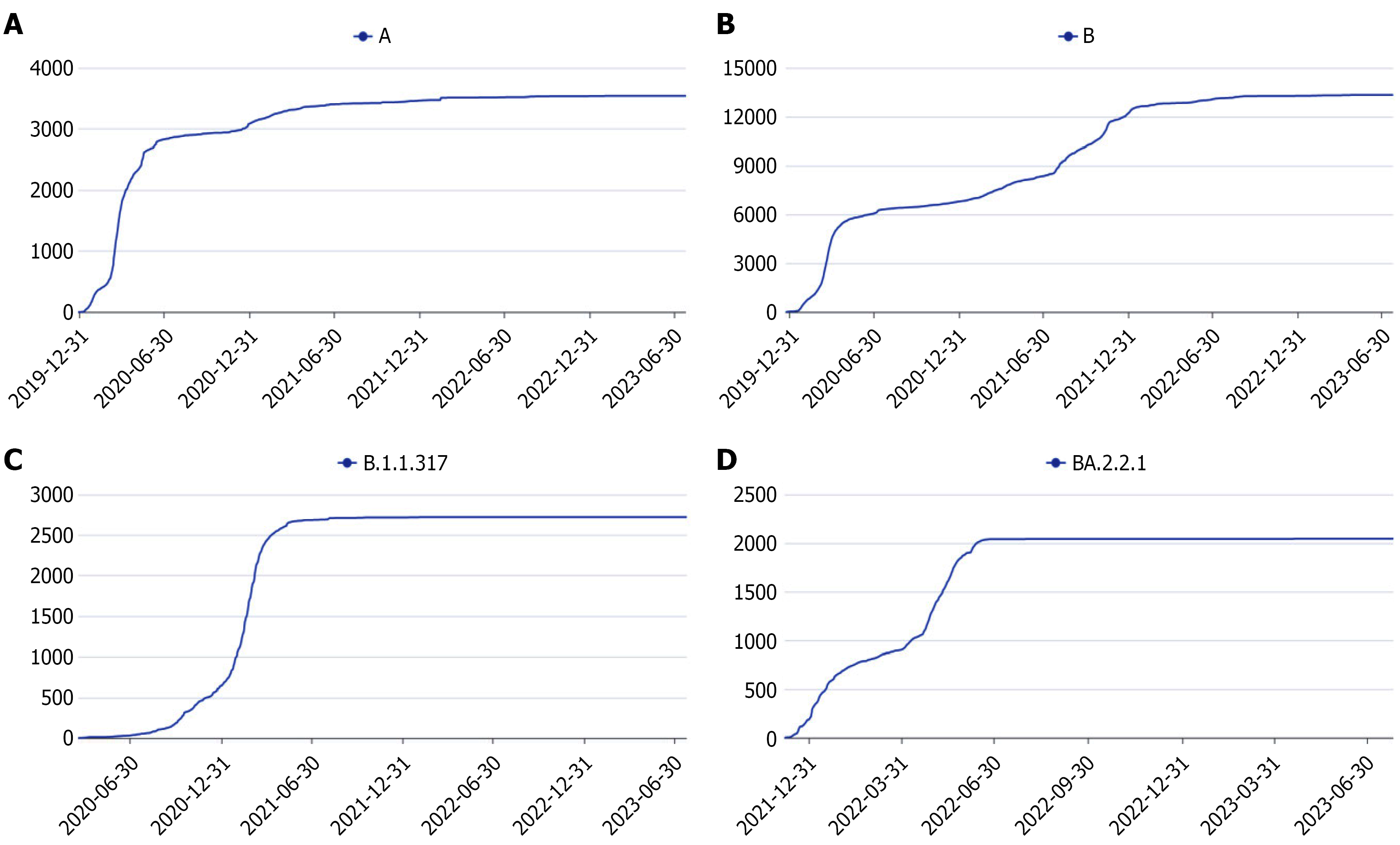
In 2020, 52 complete SARS-CoV-2 genomes were obtained (GenBank: OM065338-OM065389). Pangolin lineage assi
In May 2021, 16 complete genomes were sequenced (GenBank: OM098414-OM098426, OQ586439-OQ586441). All were assigned to lineage B.1.1.317. During this period, global submissions of B.1.1.317 were limited; according to public da
In April 2022, 23 complete genomes were obtained (GenBank: OQ586645-OQ586667). Pangolin analysis identified all as BA.2.2.1. Globally, submissions of BA.2.2.1 increased exponentially during this interval, establishing it as the predo
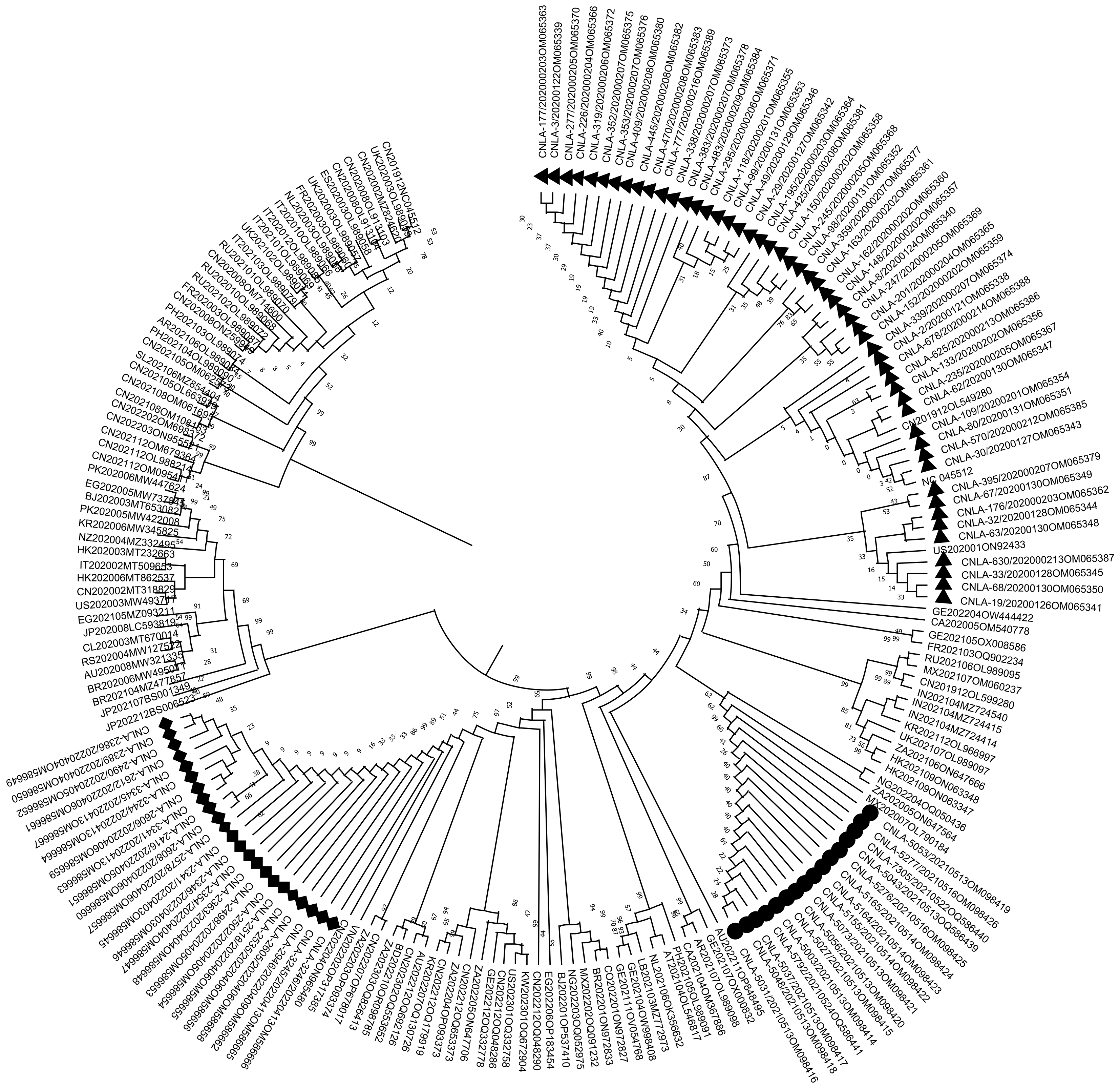
Whole-genome sequences of SARS-CoV-2 from different time periods, geographic regions, and lineages were retrieved from public databases, together with the Wuhan-Hu-1 reference strain (National Center for Biotechnology Information accession NC_045512) and the 91 Local sequences generated in this study[18]. A maximum-likelihood phylogenetic tree was constructed using MEGA software (Figure 2)[19].
The phylogenetic tree revealed clear temporal clustering, with sequences from each of the three years forming distinct monophyletic groups. The 2020 isolates were divided into lineage A and lineage B clusters. Viruses from 2021 and 2022 were consistently classified as B.1.1.317 and BA.2.2.1, respectively, supporting the lineage assignments obtained by Pangolin.
The clinical data from COVID-19 patients in this study indicate that cough, fever, and computed tomography imaging abnormalities were the primary clinical manifestations, consistent with relevant reports[20], each occurring in more than 90% of the identified cases. According to existing literature, fever has been reported in 88.7%-98.6% of patients, while cough occurs in 59.4%-79.0% of cases[21-23]. Notably, the positivity rates of these clinical indicators varied substantially across different years. In 2020, the incidence of fever and cough symptoms in COVID-19 patients exceeded 90%, whereas in 2021 and 2022, the prevalence of both symptoms decreased to below 40%. Reports from early 2020 in Wuhan documented fever and cough rates of 78.2% and 15.6%, respectively[24]. These discrepancies may be attributed to differences in the circulating SARS-CoV-2 variants across years, as well as variations in case detection and the timing of treatment initiation, all of which may influence clinical presentation. In 2020, patients were typically hospitalized only after the onset of symptoms. In contrast, during 2021 and 2022, widespread nucleic acid testing of close contacts was implemented. Consequently, many infected individuals, including asymptomatic cases, received antiviral treatment immediately after testing positive, which likely contributed to improved clinical outcomes and reduced positivity rates of these indicators.
To support genomic epidemiology, 91 complete genome sequences of SARS-CoV-2 were obtained from Lu’an city between 2020 and 2022 using next-generation sequencing technology. These sequences have been submitted to the GenBank database [accession numbers OM065338-OM065389(2020), OM098414-OM098426(2021),
The temporal pattern of circulating strains in 2020 and 2022 was consistent with national SARS-CoV-2 variant trends[16]. However, in May 2021, the identified genotype B.1.1.317 differed from the globally dominant variant at that time. Epidemiological investigation revealed that this outbreak originated from an employee in another province who handled imported frozen products; this individual was likely infected through contact with contaminated products and subsequently transmitted the virus, resulting in a cluster of 20 cases in Lu’an. As SARS-CoV-2 can persist on frozen products, this genotype, while not dominant nationally, circulated locally and in several provinces on a limited scale. Timely public health interventions led to the rapid containment of this outbreak. According to spatiotemporal distribution data, Russia and Germany reported the highest numbers of B.1.1.317 sequences in May 2021, suggesting that the infected worker may have handled products imported from these regions.
This study delineates the evolving clinical presentation and genomic landscape of SARS-CoV-2 in Lu’an city from 2020 to 2022. The marked decrease in the prevalence of classic symptoms, such as fever and cough in later years underscores the impact of widespread testing, early treatment, and potentially viral evolution on clinical manifestations. Genomic surveillance effectively tracked the transition from ancestral variants to the Omicron BA.2.2.1 subvariant and identified a discrete, imported outbreak linked to contaminated frozen products in 2021. The accumulation of mutations over time, particularly in the Omicron variant, highlights the rapid evolutionary dynamics of SARS-CoV-2.
Following the implementation of class B management for COVID-19 infection in early 2023[25], a significant increase in infections was observed nationwide. This surge in case numbers is likely to promote the emergence of new viral variants. Therefore, strengthening ongoing genomic surveillance and molecular epidemiology is paramount. Data generated through such monitoring are essential for understanding transmission dynamics, detecting emerging variants promptly, and informing evidence-based public health strategies for the effective prevention and control of the pandemic.
Comparative genomic analysis against the Wuhan reference strain (NC_045512)[26] revealed a progressive increase in mutational burden over time. The 2020 strains had 0-6 mutations across 59 sites, the 2021 strains had 15-17 mutations across 23 sites (15 of which were shared), and the 2022 strains exhibited 76-80 mutations at 82 sites (76 of which were shared)[27]. Given the rapid mutation rate of SARS-CoV-2, approximately one mutation every two weeks[28], with some reports suggesting an even faster rate[29] -genomic analysis serves as a crucial tool for tracing the origins of clustered cases[30].
We extend our sincere gratitude to all frontline healthcare workers and public health staff in Lu’an city for their dedicated efforts in patient care, sample collection, and data recording during the COVID-19 pandemic. We also thank the Lu’an Municipal Center for Disease Control and Prevention, as well as the participating hospitals for their collaboration and support in facilitating this study.
| 1. | Zhu N, Zhang D, Wang W, Li X, Yang B, Song J, Zhao X, Huang B, Shi W, Lu R, Niu P, Zhan F, Ma X, Wang D, Xu W, Wu G, Gao GF, Tan W; China Novel Coronavirus Investigating and Research Team. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N Engl J Med. 2020;382:727-733. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21952] [Cited by in RCA: 17523] [Article Influence: 2920.5] [Reference Citation Analysis (15)] |
| 2. | Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat Microbiol. 2020;5:536-544. [PubMed] [DOI] [Full Text] |
| 3. | World Health Organization. Naming the coronavirus disease (COVID-19) and the virus that causes it. [cited 18 January 2026]. Available from: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/technical-guidance/naming-the-coronavirus-disease-(covid-2019)-and-the-virus-that-causes-it. |
| 4. | Fisher D, Heymann D. Q&A: The novel coronavirus outbreak causing COVID-19. BMC Med. 2020;18:57. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 172] [Cited by in RCA: 128] [Article Influence: 21.3] [Reference Citation Analysis (1)] |
| 5. | Wang S, Xu X, Wei C, Li S, Zhao J, Zheng Y, Liu X, Zeng X, Yuan W, Peng S. Molecular evolutionary characteristics of SARS-CoV-2 emerging in the United States. J Med Virol. 2022;94:310-317. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 85] [Cited by in RCA: 64] [Article Influence: 16.0] [Reference Citation Analysis (1)] |
| 6. | Surpris ACA, Jungerman MR, Preston LE, Gertz AM, Duong KK, Roy S, Morales M, Olmstead J, Delea K, Alvarado-Ramy F, Brown C, Chen TH. Public Health Travel Restrictions Implemented for Persons at Risk of Transmitting SARS-CoV-2 Infection-United States, January 1, 2020-April 6, 2022. J Public Health Manag Pract. 2025;31:472-476. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 7. | Yang W, Xie SY, Gao DW, Chen ZC, Fan YZ, Lv Y, Gong TQ, Chang HW. [Epidemiological and Etiological Study of COVID-19 Cases in Lu’an City]. Guoji Bingduxue Zazhi. 2021;28:325-328. [DOI] [Full Text] |
| 8. | Cui LL, Xia AH, Wang W, Shao KK. [Analysis of novel coronavirus nucleic acid detection results by different detection]. Linchuang Jianyan Zazhi. 2020;11:823-826. [DOI] [Full Text] |
| 9. | Zhang WH, Ma W, Li FG, Xi J, Jin ZY, Cui XQ, He RF, Zhang JH, Piao WH. [Comparative analysis of the performance of different nucleic acid detection reagents for 2019 novel coronavirus]. Ningxia Yixue Zazhi. 2020;42:913-916. [DOI] [Full Text] |
| 10. | Freed NE, Vlková M, Faisal MB, Silander OK. Rapid and inexpensive whole-genome sequencing of SARS-CoV-2 using 1200 bp tiled amplicons and Oxford Nanopore Rapid Barcoding. Biol Methods Protoc. 2020;5:bpaa014. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 251] [Cited by in RCA: 193] [Article Influence: 32.2] [Reference Citation Analysis (1)] |
| 11. | Oeck S, Tüns AI, Schramm A. Nanopore Sequencing Techniques: A Comparison of the MinKNOW and the Alignator Sequencers. Methods Mol Biol. 2023;2649:209-221. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 12. | Baker DJ, Aydin A, Le-Viet T, Kay GL, Rudder S, de Oliveira Martins L, Tedim AP, Kolyva A, Diaz M, Alikhan NF, Meadows L, Bell A, Gutierrez AV, Trotter AJ, Thomson NM, Gilroy R, Griffith L, Adriaenssens EM, Stanley R, Charles IG, Elumogo N, Wain J, Prakash R, Meader E, Mather AE, Webber MA, Dervisevic S, Page AJ, O'Grady J. CoronaHiT: high-throughput sequencing of SARS-CoV-2 genomes. Genome Med. 2021;13:21. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 109] [Cited by in RCA: 77] [Article Influence: 15.4] [Reference Citation Analysis (1)] |
| 13. | Gunadi, Wibawa H, Hakim MS, Marcellus, Trisnawati I, Khair RE, Triasih R, Irene, Afiahayati, Iskandar K, Siswanto, Anggorowati N, Daniwijaya EW, Supriyati E, Nugrahaningsih DAA, Budiono E, Retnowulan H, Puspadewi Y, Puspitawati I, Sianipar O, Afandy D, Simanjaya S, Widitjiarso W, Puspitarani DA, Fahri F, Riawan U, Fauzi AR, Kalim AS, Ananda NR, Setyati A, Setyowireni D, Laksanawati IS, Arguni E, Nuryastuti T, Wibawa T; the Yogyakarta-Central Java COVID-19 study group. Molecular epidemiology of SARS-CoV-2 isolated from COVID-19 family clusters. BMC Med Genomics. 2021;14:144. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 14] [Article Influence: 2.8] [Reference Citation Analysis (1)] |
| 14. | Molecular evolutionary Genetics Analysis. [cited 18 January 2026]. Available from: https://megasoftware.net/. |
| 15. | Rambaut A, Holmes EC, O'Toole Á, Hill V, McCrone JT, Ruis C, du Plessis L, Pybus OG. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat Microbiol. 2020;5:1403-1407. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2480] [Cited by in RCA: 1861] [Article Influence: 310.2] [Reference Citation Analysis (0)] |
| 16. | China National Center for Bioinformation. [National Genomics Data Center Resource for Coronavirus 2019]. [cited 18 January 2026]. Available from: https://ngdc.cncb.ac.cn/ncov/. |
| 17. | National Center for Biotechnology Information. sars-cov-2 Lu'an. [cited 18 January 2026]. Available from: https://www.ncbi.nlm.nih.gov/nuccore/?term=sars-cov-2+Lu%27an. |
| 18. | National Center for Biotechnology Information. Nucleotide. [cited 18 January 2026]. Available from: https://www.ncbi.nlm.nih.gov/nuccore. |
| 19. | Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol Biol Evol. 2018;35:1547-1549. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32778] [Cited by in RCA: 23114] [Article Influence: 2889.3] [Reference Citation Analysis (4)] |
| 20. | Struyf T, Deeks JJ, Dinnes J, Takwoingi Y, Davenport C, Leeflang MM, Spijker R, Hooft L, Emperador D, Domen J, Horn SRA, Van den Bruel A; Cochrane COVID-19 Diagnostic Test Accuracy Group. Signs and symptoms to determine if a patient presenting in primary care or hospital outpatient settings has COVID-19. Cochrane Database Syst Rev. 2021;2:CD013665. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 75] [Cited by in RCA: 76] [Article Influence: 15.2] [Reference Citation Analysis (1)] |
| 21. | Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J, Wang B, Xiang H, Cheng Z, Xiong Y, Zhao Y, Li Y, Wang X, Peng Z. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus-Infected Pneumonia in Wuhan, China. JAMA. 2020;323:1061-1069. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17217] [Cited by in RCA: 14472] [Article Influence: 2412.0] [Reference Citation Analysis (7)] |
| 22. | Guan WJ, Ni ZY, Hu Y, Liang WH, Ou CQ, He JX, Liu L, Shan H, Lei CL, Hui DSC, Du B, Li LJ, Zeng G, Yuen KY, Chen RC, Tang CL, Wang T, Chen PY, Xiang J, Li SY, Wang JL, Liang ZJ, Peng YX, Wei L, Liu Y, Hu YH, Peng P, Wang JM, Liu JY, Chen Z, Li G, Zheng ZJ, Qiu SQ, Luo J, Ye CJ, Zhu SY, Zhong NS; China Medical Treatment Expert Group for Covid-19. Clinical Characteristics of Coronavirus Disease 2019 in China. N Engl J Med. 2020;382:1708-1720. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21520] [Cited by in RCA: 18438] [Article Influence: 3073.0] [Reference Citation Analysis (24)] |
| 23. | Zhou F, Yu T, Du R, Fan G, Liu Y, Liu Z, Xiang J, Wang Y, Song B, Gu X, Guan L, Wei Y, Li H, Wu X, Xu J, Tu S, Zhang Y, Chen H, Cao B. Clinical course and risk factors for mortality of adult inpatients with COVID-19 in Wuhan, China: a retrospective cohort study. Lancet. 2020;395:1054-1062. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21249] [Cited by in RCA: 18041] [Article Influence: 3006.8] [Reference Citation Analysis (10)] |
| 24. | Wei Y, Lu Y, Xia L, Yuan X, Li G, Li X, Liu L, Liu W, Zhou P, Wang CY, Zhang H. Analysis of 2019 novel coronavirus infection and clinical characteristics of outpatients: An epidemiological study from a fever clinic in Wuhan, China. J Med Virol. 2020;92:2758-2767. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 27] [Article Influence: 4.5] [Reference Citation Analysis (1)] |
| 25. | National Health Commission of the People's Republic of China. [Notice on Issuing the Overall Plan for Implementing the "Class B, Level 2" Management of Novel Coronavirus Infection]. Feb 26, 2022 [cited 18 January 2026]. Available from: https://www.nhc.gov.cn/xcs/zhengcwj/202212/8eaeef83bb7a4fdb851a3ae204295df1.shtml. |
| 26. | National Center for Biotechnology Information. Severe acute respiratory syndrome coronavirus 2 isolate Wuhan-Hu-1, complete genome. Jul 18, 2020 [cited 18 January 2026]. Available from: https://www.ncbi.nlm.nih.gov/nuccore/NC_045512. |
| 27. | Wu F, Zhao S, Yu B, Chen YM, Wang W, Song ZG, Hu Y, Tao ZW, Tian JH, Pei YY, Yuan ML, Zhang YL, Dai FH, Liu Y, Wang QM, Zheng JJ, Xu L, Holmes EC, Zhang YZ. A new coronavirus associated with human respiratory disease in China. Nature. 2020;579:265-269. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9823] [Cited by in RCA: 7448] [Article Influence: 1241.3] [Reference Citation Analysis (5)] |
| 28. | Gupta S, Gupta D, Bhatnagar S. Analysis of SARS-CoV-2 genome evolutionary patterns. Microbiol Spectr. 2024;12:e0265423. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 17] [Article Influence: 8.5] [Reference Citation Analysis (1)] |
| 29. | Morales AC, Rice AM, Ho AT, Mordstein C, Mühlhausen S, Watson S, Cano L, Young B, Kudla G, Hurst LD. Causes and Consequences of Purifying Selection on SARS-CoV-2. Genome Biol Evol. 2021;13:evab196. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 42] [Cited by in RCA: 30] [Article Influence: 6.0] [Reference Citation Analysis (1)] |
| 30. | Houwaart T, Belhaj S, Tawalbeh E, Nagels D, Fröhlich Y, Finzer P, Ciruela P, Sabrià A, Herrero M, Andrés C, Antón A, Benmoumene A, Asskali D, Haidar H, von Dahlen J, Nicolai J, Stiller M, Blum J, Lange C, Adelmann C, Schroer B, Osmers U, Grice C, Kirfel PP, Jomaa H, Strelow D, Hülse L, Pigulla M, Kreuzer P, Tyshaieva A, Weber J, Wienemann T, Kohns Vasconcelos M, Hoffmann K, Lübke N, Hauka S, Andree M, Scholz CJ, Jazmati N, Göbels K, Zotz R, Pfeffer K, Timm J, Ehlkes L, Walker A, Dilthey AT; German COVID-19 OMICS Initiative (DeCOI); German COVID-19 OMICs Initiative (DeCOI). Integrated genomic surveillance enables tracing of person-to-person SARS-CoV-2 transmission chains during community transmission and reveals extensive onward transmission of travel-imported infections, Germany, June to July 2021. Euro Surveill. 2022;27:2101089. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 12] [Article Influence: 3.0] [Reference Citation Analysis (1)] |