Published online Mar 25, 2026. doi: 10.5501/wjv.v15.i1.118362
Revised: January 27, 2026
Accepted: February 13, 2026
Published online: March 25, 2026
Processing time: 73 Days and 12.4 Hours
Paediatric gut microbiome research has long been bacteriocentric, overlooking the extensive viral component known as the gut virome. Composed of bacteriophages, eukaryotic viruses, and endogenous viral elements, the paediatric gut virome is the most abundant and genetically diverse biological entity in the intestine. Emerging evidence indicates that the virome is a key regulator of microbial ecology, immune maturation, and systemic physiological programming during early life. This narrative review synthesizes current knowledge on the establishment, development, and functional roles of the paediatric gut virome, with emphasis on its interactions with the bacterial microbiome and host immune system. We highlight how early-life viral exposures influence mucosal immune imprinting, epithelial barrier integrity, and immune tolerance, particularly during the first 1000 days of life. Virome dysbiosis is increasingly associated with paediatric gastrointestinal disorders, including inflammatory bowel disease, necrotizing enterocolitis, celiac disease, and functional gastrointestinal disorders. Beyond the gut, the virome also contributes to metabolic regulation, type 1 diabetes risk, and gut-brain axis signaling, influencing neurodevelopment. Mechanistic pathways involving phage-mediated bacterial modulation, innate immune sensing, cytokine signaling, and metabolic intermediates are discussed, positioning the paediatric gut virome as a central regulator of gas
Core Tip: The pediatric gut virome is an active biological rheostat that shapes the developmental trajectory of the infant immune system and metabolic health. Beyond serving as a reservoir for pathogens, this complex viral ecosystem-dominated by bacteriophages-governs bacterial population dynamics through lytic pressure and horizontal gene transfer. During the critical first 1000 days of life, virome disturbances triggered by antibiotics, formula feeding, or neonatal intensive care unit hospitalization can drive the pathogenesis of necrotizing enterocolitis, inflammatory bowel disease, and systemic autoimmunity. Moving forward, “illuminating” viral dark matter and leveraging precision tools such as phage therapy will be essential to transitioning toward virome-inclusive, proactive pediatric medicine.
- Citation: Saeed NK, Elbeltagi YM, Al-Beltagi M. Unveiling the viral dimension: The paediatric gut virome as a key modulator of gastrointestinal metabolic, and neurodevelopmental health. World J Virol 2026; 15(1): 118362
- URL: https://www.wjgnet.com/2220-3249/full/v15/i1/118362.htm
- DOI: https://dx.doi.org/10.5501/wjv.v15.i1.118362
The human gastrointestinal tract hosts a dense and dynamic microbial consortium-the gut microbiome-that serves as a fundamental determinant of host physiology, metabolic homeostasis, and immune maturation. For decades, microbiome research has been characterized by a “bacteriocentric” focus, successfully unravelling the profound influence of bacterial communities on human health[1]. However, this perspective overlooks the most abundant and genetically diverse biological entities within the enteric ecosystem: Viruses. The gut virome, the collective community of viruses inhabiting the intestine, represents the “dark matter” of the microbiome. It is a multi-kingdom assembly comprising bacteriophages (viruses that infect bacteria), eukaryotic viruses (infecting host cells), archaeal viruses, and endogenous viral elements (EVEs)-remnants of ancient viral integrations within the host genome that continue to influence health and immunity[2].
A significant paradigm shift is currently underway in our understanding of these entities. Once viewed exclusively through the lens of pathogenesis, the virome is now recognized as a critical suite of commensals and regulators. Rather than acting solely as transient invaders, these viruses are permanent architects of the microbial landscape[3]. Bacteriophages, through predatory “kill-the-winner” dynamics or symbiotic “piggyback-the-winner” lysogeny, orchestrate the structure and function of the bacterial microbiome. Simultaneously, eukaryotic viruses and EVEs engage in a sophisticated cross-talk with the host, modulating inflammation, promoting epithelial barrier integrity, and “priming” the immune system for future challenges[4].
Nowhere is the influence of this viral ecosystem more pivotal than during the “First 1000 Days” of life. This formative period represents a unique developmental window where the virome is first established and undergoes rapid, non-random succession. Unlike the relatively stable adult virome, the pediatric virome is highly plastic and exquisitely sensitive to perinatal variables, including mode of delivery, infant feeding (breast milk vs formula), and antibiotic exposure[5]. This early-life maturation coincides with a critical period of immune imprinting, where the virome assists in training the nascent immune system to distinguish between self, commensal, and pathogen.
Despite its importance, the virome remains underrepresented in the broader microbiome literature. Emerging evidence suggests that disruptions in this delicate viral equilibrium-viral dysbiosis-are not merely confined to the gut. While alterations are linked to gastrointestinal conditions such as necrotizing enterocolitis (NEC), inflammatory bowel disease (IBD), and celiac disease, the virome’s reach is truly systemic[6]. Distortions in viral colonization have been implicated in the pathogenesis of type 1 diabetes (T1D), obesity, asthma, and even neurodevelopmental disorders, suggesting a “gut-virome-axis” that extends to the lungs, pancreas, and brain[7].
Given the disparate nature of current research and the lack of standardized methodologies, there is an urgent need for a comprehensive synthesis of this burgeoning field. This review aims to unveil the viral dimension of pediatric health by characterizing the composition and maturation of the virome in healthy early life, evaluating its association with both gastrointestinal and systemic pathologies, and discussing how perinatal factors shape its trajectory. By integrating viral-microbial-host interactions, we seek to highlight the translational potential of the virome in early-life diagnostics and therapeutic innovation.
This narrative review was informed by a comprehensive literature search conducted using PubMed, Scopus, and Web of Science databases up to December 2025. Search terms included combinations of paediatric, infant, gut virome, bacteriophages, early-life microbiome, immune development, metabolic programming, and neurodevelopment. Priority was given to original human studies, birth cohort analyses, mechanistic animal models, and high-quality systematic reviews focusing on early-life viral dynamics. Given the rapidly evolving nature of virome research and the absence of standardized methodologies, we adopted an integrative narrative approach to synthesize heterogeneous evidence across microbiology, immunology, gastroenterology, and developmental biology.
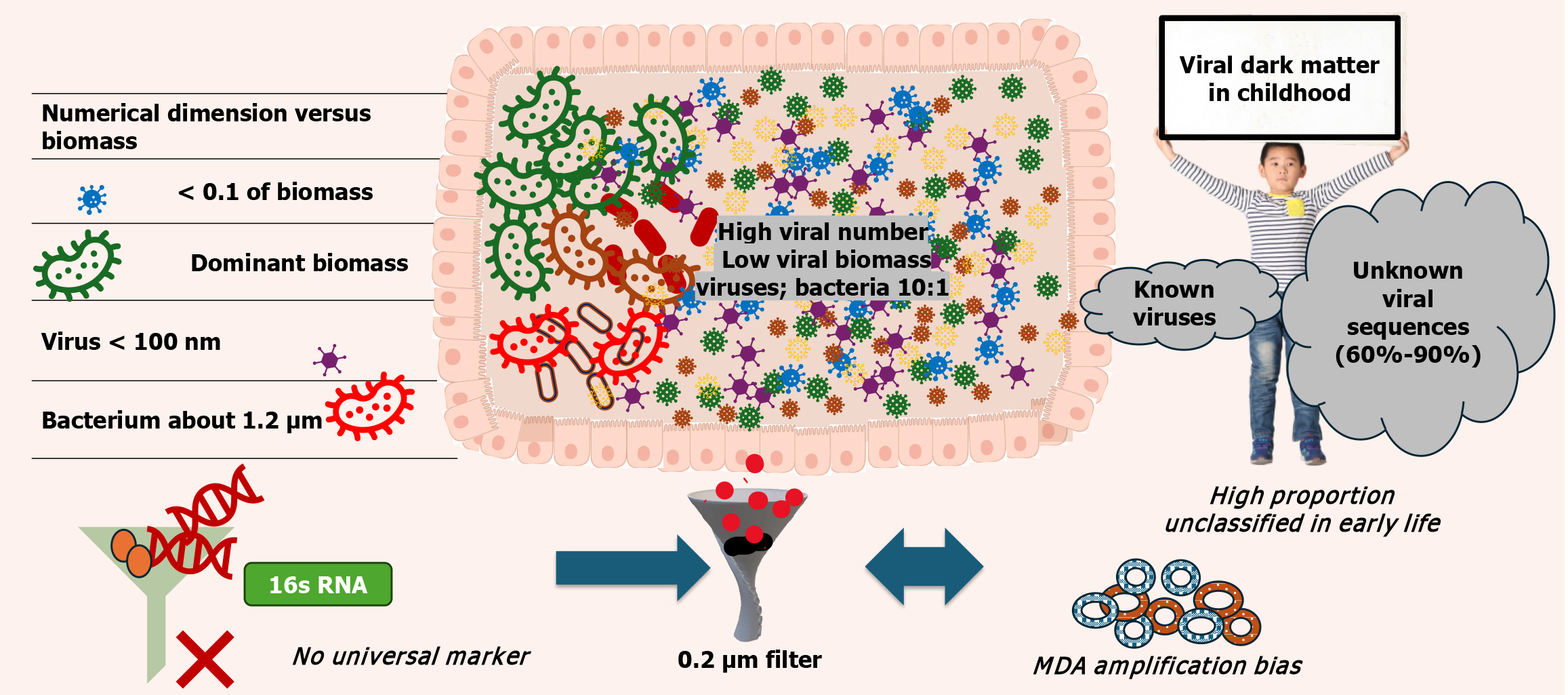
Although bacteria dominate the biomass of the gut microbiome, viruses overwhelmingly dominate its numerical landscape, creating what is often referred to as the viral-bacterial paradox. In the human gastrointestinal tract, virus-like particles-the vast majority of which are bacteriophages-typically outnumber bacterial cells by approximately 10:1, with reported ratios ranging from 1:1 to over 20:1, depending on the intestinal compartment, mucosal proximity, age, and health status[8,9]. This striking numerical superiority stands in sharp contrast to their negligible contribution to total microbial mass: Viruses collectively account for less than 0.1% of the gut microbial biomass (Figure 1 and Table 1).
| Metric | Gut bacteriome | Gut virome |
| Numerical abundance | 1011-1012 cells per gram of feces | 109-1012 particles per gram of feces (VLPs)1 |
| Biomass contribution | Major (99.9% microbial biomass) | Minimal (< 0.1% of microbial biomass) |
| Universal genetic marker | Present (16S rRNA gene) | None (Requires shotgun metagenomics) |
| Database maturity | High (most sequences are identifiable) | Low (40%-90% “viral dark matter”)2 |
| Identification logic | Taxonomy-based (16S) or functional | Homology-based or de novo assembly |
| Major constituents | Bacteria, archaea | Bacteriophages, eukaryotic viruses, archaeal viruses, and EVEs |
| Replication strategy | Predominantly binary fission | Lytic, lysogenic (prophages), and chronic |
| Pediatric trajectory | Diversity increases linearly with age | Richness often peaks in infancy and fluctuates |
| Study cost/complexity | Lower (standardized and high-throughput) | Higher (requires enrichment and heavy compute) |
This disproportion reflects the fundamental biological differences between viruses and bacteria. Viruses are orders of magnitude smaller and lack the complex cellular machinery that characterizes bacterial cells. While bacteria possess ribosomes, cell walls, and metabolic networks, viruses consist primarily of nucleic acid enclosed within a protein capsid, which may be surrounded by a lipid envelope. As a result, viruses contribute minimally to bulk biomass measurements yet exert a disproportionately large ecological and functional influence through their interactions with bacterial hosts and the human immune system[10,11].
From a methodological perspective, this size and simplicity present substantial challenges. The extraction of sufficient quantities of high-quality viral DNA or RNA from gut samples is inherently difficult, particularly in paediatric populations where stool volumes are limited and viral loads fluctuate rapidly with age and environmental exposure[12]. Early microbiome studies relied heavily on bulk metagenomic sequencing, in which total microbial DNA was extracted without prior viral enrichment. In such datasets, the abundant bacterial DNA signal effectively “drowned out” viral sequences, resulting in a profoundly bacteriocentric view of the gut ecosystem that persisted for more than a decade[13].
The technical barriers to virome characterization extend beyond biomass considerations. Bacterial community profiling was revolutionized by the discovery of the 16S ribosomal RNA gene, a conserved genetic marker present in all bacteria and archaea that enables rapid, cost-effective taxonomic identification. Viruses, in contrast, lack any equivalent universal marker[14]. Each viral family-and often each genus-possesses a distinct genetic architecture, rendering targeted amplification approaches impractical. Consequently, virome research relies almost exclusively on shotgun metagenomic sequencing, which is significantly more expensive, computationally demanding, and analytically complex[15].
Even with advanced sequencing technologies, viral identification remains limited by incomplete reference databases. Currently, 60%-90% of viral sequences recovered from gut virome studies fail to match any known viral genome-a phenomenon commonly referred to as “viral dark matter”. This knowledge gap is particularly pronounced in paediatrics, where early-life viromes are highly dynamic and enriched with age-specific and transient viruses that have yet to be catalogued[16,17]. In essence, paediatric virome researchers are navigating a biological landscape in which most of the genomic territory remains uncharted.
Additional sources of bias arise during sample processing. Standard microbiological filtration methods, such as 0.2 µm filters, were originally designed to separate bacteria from smaller particles. However, these filters may inadvertently retain large viruses (e.g., giant phages or enveloped eukaryotic viruses) while allowing smaller virions to pass through, thereby skewing viral community profiles. Such biases are rarely uniform across studies, contributing to poor inter-study comparability[18,19].
Moreover, because viral nucleic acid yields are often insufficient for direct sequencing, many studies employ multiple displacement amplification (MDA) to amplify viral DNA prior to library preparation[20]. While MDA improves detectability, it introduces its own distortions by preferentially amplifying small, circular, single-stranded DNA (ssDNA) viruses, most notably members of the Microviridae family, while underrepresenting linear or RNA viruses. These amplification artifacts can give the false impression of viral dominance or stability where none exists[21].
Taken together, these biological and technical factors explain why the gut virome, despite its numerical dominance and functional importance, has historically remained understudied, underappreciated, and poorly characterized, especially in children. Recognizing and addressing this viral-bacterial paradox is essential for accurately interpreting microbiome-host interactions and for advancing a more integrated, ecosystem-level understanding of paediatric gastrointestinal and systemic health.
The pediatric gut virome develops through a coordinated “multi-hit” process in which maternal transmission establishes the initial viral community, and subsequent influences such as diet, environment, and early medical exposures shape its maturation[22]. This early assembly follows a distinct, rapidly evolving trajectory driven by highly dynamic pioneer viral species. Disruptions during this sensitive window-such as cesarean delivery or early antibiotic use-can alter normal virome progression and potentially lead to long-term consequences for gut and immune development[23].
The traditional “sterile womb” hypothesis has been challenged by recent studies detecting viral genetic signatures in meconium (the first stool) and amniotic fluid. While some researchers have identified fragments of bacteriophages and even eukaryotic viruses, such as torque teno virus (TTV), in newborns, the field remains divided[24]. Critics argue these findings may result from low-biomass contamination or the translocation of viral nucleic acids rather than active viral replication. Regardless of whether a “prenatal virome” exists, it is widely accepted that the birth event serves as the primary “big bang” for viral colonization[25].
The delivery room is the first major determinant of viral composition. Infants born vaginally are seeded with a virome that mirrors the maternal vaginal and fecal microbiota. These infants typically display an early dominance of bacteriophages-specifically those belonging to the caudoviricetes class-which target pioneer bacterial colonizers like Bacteroides and Bifidobacterium[26]. Infants delivered via cesarean section miss this vertical seeding, instead showing a virome initially dominated by skin-associated viruses and environmental contaminants. This “founder effect” can delay the maturation of the phage community, potentially altering the pace at which the bacterial microbiome stabilizes[27].
Feeding serves as a continuous “viral infusion”. Human milk is not sterile; it contains a diverse virome, including bacteriophages and eukaryotic viruses (e.g., cytomegalovirus, human immunodeficiency virus, and TTV). Beyond direct transmission, human milk oligosaccharides (HMOs) in breast milk act as “decoy receptors”, preventing pathogenic viruses from binding to the infant’s intestinal epithelium while simultaneously supporting the growth of bacterial hosts for beneficial phages[28]. Formula-fed infants exhibit a distinct virome signature, often characterized by greater richness of certain eukaryotic viruses and a lack of maternal immune-modulating factors found in breast milk. This difference may explain the “protective effect” breastfeeding has against early-life viral gastroenteritis[29].
The virome is exquisitely sensitive to the child’s physical surroundings. The presence of older siblings and furry pets acts as a catalyst for virome diversification. Siblings often act as vectors for “socially transmitted” eukaryotic viruses, which, while potentially pathogenic, also serve to “train” the developing immune system[30]. Urban vs. rural living conditions significantly shift the phage-to-eukaryote ratio, with rural environments generally promoting a more diverse and resilient viral ecosystem[31].
Antibiotics are perhaps the most disruptive force in early-life virome development. Because bacteriophages are obligate parasites of bacteria, the depletion of bacterial hosts via antibiotics leads to a secondary viral collapse[32]. Sub-lethal doses of antibiotics can trigger the “SOS response” in bacteria, causing dormant prophages to wake up, replicate, and lyse their hosts. Early-life antibiotic exposure reduces phage richness and can lead to a “bloom” of antibiotic-resistance genes (ARGs) carried by the surviving viral population, effectively turning the virome into a reservoir for future bacterial resistance[33].
Key mechanisms by which the paediatric gut virome modulates host metabolic and endocrine pathways include: (1) Phage-mediated regulation of bacterial metabolic networks through selective lysis and lysogenic integration, thereby shaping short-chain fatty acid production, bile acid metabolism, and nutrient harvest; (2) Horizontal transfer of auxiliary metabolic genes (AMGs) that enhance or reprogram microbial enzymatic capacity; (3) Immune-mediated metabolic signaling, whereby viral sensing via innate immune receptors [e.g., toll-like receptors (TLR)] induces cytokine and interferon responses that influence insulin sensitivity and systemic inflammation; and (4) Barrier-dependent viral translocation, permitting viral nucleic acids to access systemic compartments and modulate endocrine-relevant immune pathways. Collectively, these mechanisms position the early-life virome as a dynamic regulator of metabolic pro
The transition from the neonatal period through childhood is marked by a sophisticated successional choreography and a series of age-specific steps, likely driven by the selection pressures of the maturing bacterial microbiome. Table 2 shows the different determinants of the developing paediatric gut virome[34]. While the bacterial microbiome typically increases in diversity and richness linearly with age, the virome follows a more complex, “biphasic” trajectory defined by early-life expansion, a eukaryotic “spike”, and subsequent stabilization. The virome assembles immediately at birth, with extreme volatility[35]. Seminal metagenomic studies, such as the work by Breitbart and colleagues, have demonstrated that the infant virome undergoes rapid compositional changes even between the first and second weeks of life[36].
| Factor | Primary impact on virome composition | Age-dependent outcome |
| Mode of delivery | Vaginal: Vertical seeding of maternal phages (e.g., Caudoviricetes). C-section: Initial colonization by skin-associated and environmental viruses | VD: Earlier stabilization of the bacteriome-phage axis. CS: Delayed maturation and reduced alpha-diversity up to 24 months |
| Infant diet | Breast milk: Direct transfer of TTV/CMV; HMOs act as “decoy receptors” for pathogens. Formula: Distinct eukaryotic signature; lacks maternal immune factors | Breastfed: Sustained “healthy” phage-host dynamics; lower risk of early-life viral gastroenteritis. Formula-fed: More rapid diversification but may lack protective “pioneer” phages |
| Antibiotic use | “Viral collapse”: Loss of bacterial hosts leads to secondary phage depletion; induction of prophages (lytic cycle) | Reduced richness and a “bloom” of antibiotic-resistance genes within the viral reservoir; long-term instability |
| Social environment | Presence of older siblings and pets acts as a vector for “socially transmitted” eukaryotic viruses | Increased eukaryotic viral richness at 1-2 years; potentially beneficial “immune training” through non-pathogenic exposure |
| Geography | Urban vs. Rural: Rural living is associated with higher viral diversity and distinct metabolic gene profiles (vAMGs) | Rural viromes tend to converge toward a stable “adult-like” state more resiliently than urban counterparts |
| Host maturity (age) | Shift from a phage-dominated (predatory) state to a eukaryotic-inclusive (commensal) state | Conversion from chaotic/volatile infancy to a stable, “adult-like” personal virome by age 2 to 5 |
Despite this rapid change, the neonatal virome is initially characterized by low initial diversity, the “dark matter” predominance, and prophage induction. Contrary to earlier assumptions, viral richness is often lower in the earliest-in-life specimens compared to later timepoints in the first year. Moreover, a vast majority of neonatal sequences (often > 60%) represent “viral dark matter” that does not match any known genomic references[37]. In addition, the primary reservoir for these initial viruses is not necessarily the environment, but the bacterial hosts themselves. Most pioneer bacteriophages are “seeded” as prophages integrated into the genomes of colonizing bacteria, which then enter the lytic cycle to establish the free-living virome[26].
As the infant moves through the first year of life, a distinct ecological pattern emerges: The inverse correlation between phage and bacterial richness. Initial months are characterized by high phage richness and low bacterial diversity (bacteriophage contraction)[38]. As the child ages, bacterial richness expands significantly (bacterial expansion). This growth is paralleled by a corresponding reduction in bacteriophage richness and diversity. This suggests a “top-down” prey-predator model, in which a vast array of pioneer phages “sculpt” the burgeoning bacterial populations, eventually reaching a state of equilibrium in which bacterial diversity dominates the ecosystem[39,40].
A defining feature of maturation is the shift from a purely prokaryotic virome to one that includes host-infecting viruses (The Eukaryotic Expansion and Immune “Taming”). By approximately age two, the eukaryotic family Anelloviridae often becomes the major component of the gut virome[39]. This transient surge in eukaryotic viruses is largely attributed to the child’s immature adaptive immune system (immune immaturity window). During this “window of opportunity”, viruses that the host would otherwise suppress are allowed to proliferate, potentially serving as a training ground for immune imprinting[41]. Once a fully competent immune response is achieved, the eukaryotic viral load is “tamed” and significantly reduced, settling into the minor fraction seen in healthy adults (post-maturation decline)[42].
Between ages 2 and 5, the pediatric virome loses its chaotic infant signature and converges toward an “adult-like” configuration (convergence toward the persistent personal virome). There is a notable transition from lytic phages (predatory “kill-the-winner” dynamics) in early infancy to temperate phages (symbiotic prophages) as the child grows (lytic to temperate shift)[43]. While the infant virome is in constant flux, the adult-like virome becomes remarkably stable over time, forming a “persistent personal virome” that remains unique to the individual[35].
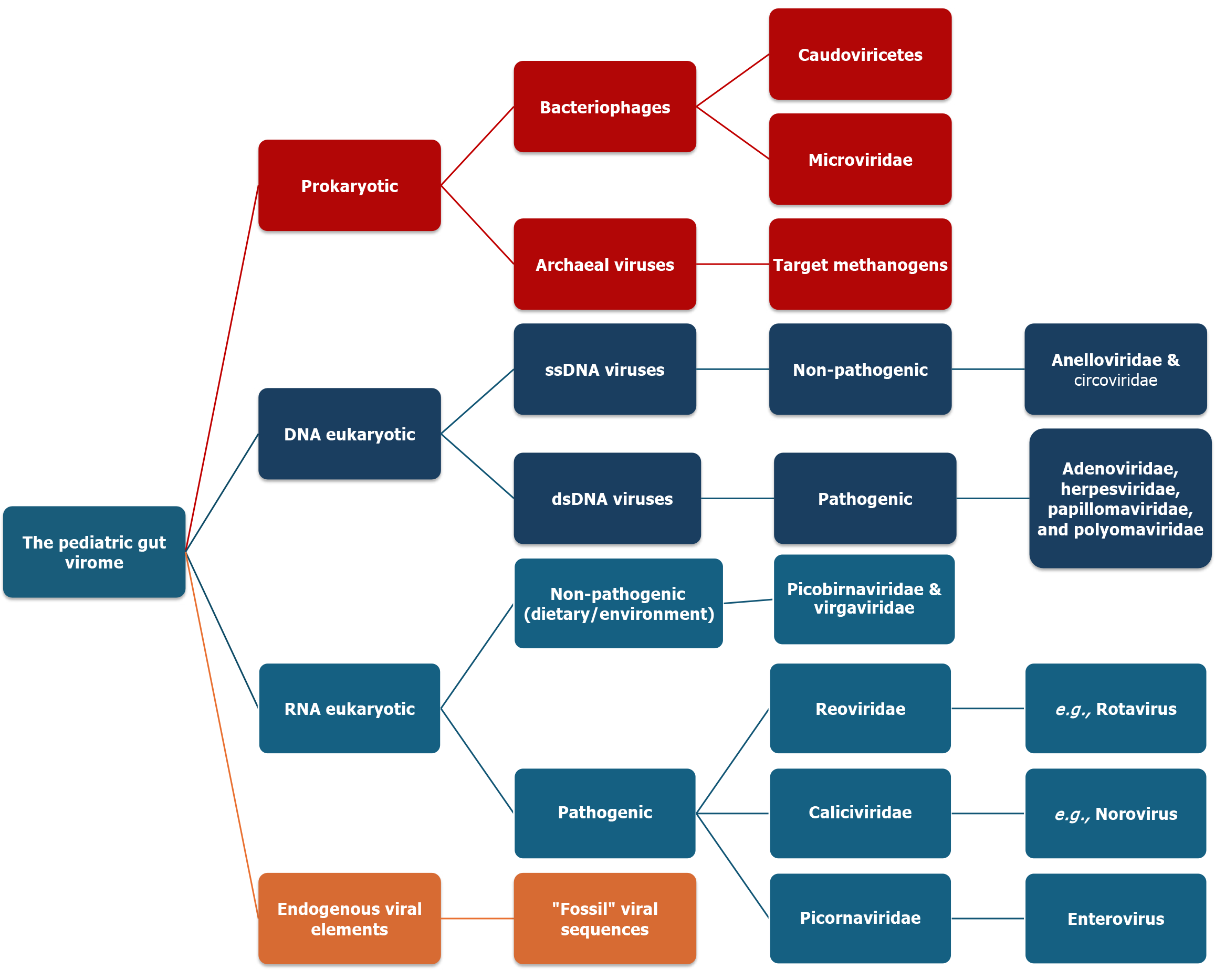
To understand the functional impact of the virome, we must categorize its members not only by name but also by their ecological roles within the pediatric gut (Figure 2).
The phage superfamilies (prokaryotic): Bacteriophages are the most abundant and diverse members of the gut virome. They employ two primary life-cycle strategies that shape the bacterial landscape: Lysogenic and lytic cycles[44]. In the lysogenic cycle, Phages integrate their genome into the bacterial host chromosome (as prophages) and replicate synchronously with the host. This represents the primary reservoir of phages in the infant gut, suggesting that early viral colonization is largely “encoded” within the pioneer bacterial species[26]. In the lytic cycle, Viruses hijack the host’s molecular machinery to produce new virions, which are ultimately released via host cell lysis or chronic shedding. Two main taxa predominate in the pediatric virome: Caudovirales and Microviridae[45].
Caudoviricetes (Tailed Phages) are the most prevalent phages in the pediatric gut. They include the Myoviridae, Podoviridae, and Siphoviridae families, as well as the recently discovered crAss-like phages. These are central to modulating bacterial diversity[46]. They are critical for modulating bacterial diversity and are often the first to respond to dietary changes or antibiotic stressors. Microviridae are small, ssDNA phages that are disproportionately abundant in early infancy. Their role is less understood, but they are known to have high mutation rates, potentially allowing them to adapt rapidly to a changing infant gut environment[47].
DNA eukaryotic viruses: From commensals to pathobionts: These viruses infect human cells and are categorized by their strandedness: Either single- or double-stranded DNA viruses. ssDNA Viruses are predominantly represented by Anelloviridae and Circoviridae. Anelloviridae (e.g., TTV) are often considered the “commensal” non-pathogenic eukaryotic viruses of humans[48]. By age two, they often become a major component of the virome, potentially because the child’s adaptive immune system is immature. Their abundance in the pediatric gut is highly correlated with the host’s immune status; high levels are often seen in immunosuppressed states or during periods of rapid immune “training”. dsDNA Viruses include families such as Adenoviridae, Herpesviridae, Papillomaviridae, and Polyomaviridae. Unlike ssDNA viruses, many members of these families are associated with clinically relevant pediatric infections[49].
RNA eukaryotic viruses: Pathogenic and dietary signals: The RNA virome provides a snapshot of both the child’s health and their environment. They are either pathogenic or non-pathogenic RNA eukaryotic viruses. The non-pathogenic/dietary viruses, the most abundant families, are often Picobirnaviridae and Virgaviridae (dietary and environmental viruses)[50]. The latter is a clade of plant viruses, indicating that a significant portion of the intestinal RNA virome is derived from the diet. They may reside transiently in the gut without causing overt disease, contributing to the “background noise” of the virome. The pathogenic viruses are smaller but clinically significant populations of Reoviridae (e.g., Rotavirus), Caliciviridae (e.g., Norovirus), and Picornaviridae (e.g., Enterovirus). They are the primary drivers of pediatric viral gastroenteritis[51].
Archaeal viruses and EVEs: The archaeal frontier, while less studied, is beginning to reveal that viruses infecting Methanobrevibacter (archaeal viruses) are increasingly common as the gut becomes more anaerobic with age. EVEs are “fossil” viral sequences integrated into the human genome. In the pediatric context, EVEs can be transcriptionally active, playing a role in innate immune signaling and protecting infants against related exogenous viral pathogens[52].
Bacteriophages are the primary architects of the gut microbial landscape, exerting a “top-down” regulatory pressure that shapes the structure, function, and evolutionary trajectory of the bacterial community. These interactions are not merely predatory but represent a complex continuum ranging from lethal parasitism to symbiotic integration[53].
The balance of the gut ecosystem is maintained through two primary ecological strategies that dictate how phages respond to shifts in bacterial populations. In the “Kill the Winner” (lytic predation) model, phages act as density-dependent regulators. When a specific bacterial species becomes dominant (the “winner”), its corresponding phages proliferate rapidly, lysing the bacterial cells and preventing any single taxon from monopolizing the niche[54]. This predatory pressure is essential for maintaining high bacterial diversity by creating space for less-dominant species. Conversely, in “Piggyback the Winner” (lysogenic coexistence) model, many phages in the gut, particularly as the microbiome matures, adopt a lysogenic strategy[55]. Instead of killing a successful bacterial host, the phage integrates its genome into the host chromosome as a prophage. This “piggybacking” allows the virus to replicate alongside a successful bacterial lineage, often providing the host with new functional traits that enhance its fitness in the gut environment[56].
Phages serve as critical mediators of bacterial evolution by transferring genetic material. During the lysogenic cycle, phages can facilitate gene transfer between different bacterial species, a process that significantly affects the functional capacity of the microbiome[57]. Phages often carry AMGs that enable their bacterial hosts to utilize new nutrient sources, which is particularly vital during dietary transitions in early life. The viral reservoir can act as a vehicle for ARGs. When bacterial hosts are under stress, phages may mobilize these genes across the community, potentially complicating clinical treatments[58].
Through these multifaceted interactions, the virome acts as a master rheostat for the gut’s bacterial “chassis”. The inverse relationship observed in early life-where high phage richness precedes the expansion of bacterial diversity-highlights the role of phages in “sculpting” the initial microbial assembly[59]. By controlling bacterial “blooms” and facilitating the exchange of beneficial genes, phages ensure the microbiome remains resilient to environmental perturbations, such as dietary changes or antibiotic exposure[60]. In the pediatric gut, the transition from an early lytic-dominated environment to a more stable, lysogenic (prophage-rich) state is a hallmark of healthy maturation. This shift reflects a move from active predatory regulation to a sophisticated symbiotic partnership that supports long-term microbial balance[61].
The paediatric virome is not merely a passenger but an active immunomodulator. Beyond their role in shaping bacterial communities, viruses in the paediatric gut engage in a sophisticated “dialogue” with the host immune system. This interaction is foundational for immune maturation, helping the infant’s nascent defenses learn to distinguish between harmless commensals and dangerous pathogens[62].
The gut virome acts as a continuous source of nucleic acid ligands that “prime” the innate immune system. Both eukaryotic viruses and bacteriophages are recognized by pattern recognition receptors (PRRs), primarily TLR located on intestinal epithelial cells and dendritic cells[63]. Bacteriophages are a significant source of DNA within the gut lumen; their DNA can be sensed by TLR9, which modulates interferon production and helps maintain intestinal homeostasis. In addition, eukaryotic viruses are recognized by TLR3 (double-stranded RNA), TLR7/8 (single-stranded RNA), and TLR9 (DNA)[64]. These interactions provide the “tonic” signaling necessary to keep the immune system in a state of readiness without triggering overt inflammation. This constant low-level stimulation is crucial for immune imprinting during the “First 1000 Days” ensuring the adaptive immune response matures correctly[65].
The virome plays a direct role in maintaining the physical and biochemical integrity of the gut barrier. Many phages possess capsid proteins that allow them to bind to mucin glycoproteins (bacteriophage adherence to mucus). This creates a “non-host-derived” immunity layer within the mucus, where phages can intercept and lyse invading bacteria before they reach the epithelial surface[66]. In addition, commensal viral signaling has been shown to support the expression of tight junction proteins (e.g., zonulin, occludin) and strengthen epithelial Junctions. By strengthening these junctions, the virome helps prevent “leaky gut” and the systemic translocation of inflammatory microbial products[67].
A hallmark of a healthy paediatric virome is its ability to promote immune tolerance while retaining the capacity to activate the immune system. Non-pathogenic “commensal” viruses, such as Anelloviridae, are often found at high levels in infants with immature immune systems. Their presence may help induce regulatory T-cells (Tregs), which suppress unnecessary inflammatory responses to food antigens and beneficial bacteria[68]. Conversely, an overgrowth of certain eukaryotic viruses or a sudden lytic “bloom” of phages (viral dysbiosis) can tip the scale toward immune activation, triggering the release of pro-inflammatory cytokines that contribute to tissue damage in conditions like NEC or IBD[69].
The development of a competent immune system is intrinsically linked to the viral load in the gut. As the child reaches approximately age two, the peak in eukaryotic viruses like Anelloviridae is eventually “tamed” by the developing adaptive immune response (Adaptive Transition). This transition suggests that the virome serves as an essential “training manual”; once the immune system has learned to recognize and control these viral populations, it achieves a state of adult-like maturity and stability (long-term programming). Disruptions in this early-life viral-host cross-talk may lead to “faulty programming”, potentially explaining the rise in autoimmune and allergic diseases later in childhood[70,71].
Beyond their localized effects within the intestinal lumen, components of the paediatric gut virome can access extra-intestinal compartments and contribute to systemic immune signaling. Under physiological conditions, an intact mucosal barrier tightly regulates the passage of viral particles and viral nucleic acids, ensuring that immune sensing remains largely compartmentalized within the gut. However, during early life-when epithelial junctions are still maturing-or in pathological states characterized by increased intestinal permeability, limited viral translocation may occur[72].
Low-level translocation of viral particles, phage-derived DNA, or viral nucleic acids into the lamina propria and systemic circulation can serve as an additional layer of immune education[73]. Viral nucleic acids detected beyond the gut are sensed by innate immune pathways, including endosomal and cytosolic PRRs, which lead to controlled interferon responses and shape systemic immune tone. This phenomenon may contribute to the calibration of antiviral immunity and peripheral immune tolerance during critical windows of immune development[74].
Importantly, bacteriophages-long considered confined to the gut lumen-have been detected in mesenteric lymph nodes, liver, and circulation, suggesting that phages can cross epithelial and endothelial barriers under specific conditions. Phage-derived nucleic acids may indirectly modulate inflammatory thresholds and immune responsiveness by influencing systemic immune cells, including monocytes and dendritic cells. In this context, the virome functions not only as a local ecological regulator but also as a long-range signaling entity within the host[75].
When tightly regulated, viral translocation appears to support immune maturation and homeostasis. In contrast, excessive or dysregulated translocation-often associated with viral dysbiosis, epithelial barrier disruption, or inflammatory disease-can amplify systemic inflammation[76]. This mechanism has been implicated in severe paediatric conditions such as NEC, in which barrier breakdown permits widespread exposure to viral and microbial products and may contribute to extra-intestinal manifestations of IBD[77].
Collectively, these observations position the paediatric gut virome as a critical interface between the intestinal environment and systemic immunity. Viral translocation and downstream signaling represent a key pathway through which early-life gut events may exert lasting effects on immune function beyond the gastrointestinal tract, reinforcing the concept of the virome as a central modulator of whole-body immune programming. Table 3 summarizes the various mechanisms by which the virome modulates the child’s immune system[78].
| Virome function | Key viral components | Host sensors/pathways | Primary immune effects | Clinical relevance |
| Innate immune priming (“tonic signaling”) | Bacteriophages; eukaryotic viruses | TLR9 (DNA); TLR3 (dsRNA); TLR7/8 (ssRNA) on IECs and dendritic cells | Basal interferon signaling; immune readiness without inflammation | Proper immune imprinting during the first 1000 days |
| Immune sensing and pattern recognition | Phage DNA; viral RNA/DNA | PRRs (TLRs, downstream interferon pathways) | Distinction between commensals and pathogens | Failure may predispose to immune dysregulation |
| Mucosal barrier reinforcement | Mucus-adherent bacteriophages | Bacteriophage adherence to mucus; epithelial junction signaling | Interception of invading bacteria; enhanced tight junction integrity | Protection against “leaky gut” and microbial translocation |
| Maintenance of immune tolerance | Commensal viruses (e.g., Anelloviridae) | Induction of regulatory immune pathways | Promotion of Treg differentiation; suppression of unnecessary inflammation | Reduced risk of food allergy and inflammatory disorders |
| Immune activation (pathological) | Viral overgrowth; phage lytic blooms | Excess PRR stimulation; cytokine release | Pro-inflammatory cytokine production; tissue injury | NEC, IBD, post-infectious inflammation |
| Immune maturation and adaptive transition | Early-life eukaryotic viruses | Adaptive immune control over viral load | Development of immune competence and stability | Long-term immune programming |
| Long-term immune programming | Early virome composition | Host-virome co-adaptation | Adult-like immune homeostasis | Dysregulation linked to allergy and autoimmunity |
| Viral translocation and systemic signaling | Limited passage of viral particles, bacteriophages, or viral nucleic acids across the intestinal barrier into the lamina propria and systemic circulation, particularly during early life or barrier disruption | Pattern recognition receptors (TLR3, TLR7/8, cGAS-STING), type I and III interferon signaling, dendritic cell and monocyte activation | Immune education, and calibration of antiviral immunity and peripheral immune tolerance during critical windows of immune development | Immune maturation and calibration of systemic antiviral responses; excessive translocation linked to systemic inflammation, necrotizing enterocolitis, inflammatory bowel disease, and extra-intestinal immune dysregulation |
While the bacterial microbiome is often termed the body’s “metabolic factory”, the virome acts as its primary supervisor. The virome influences host metabolic health through a combination of indirect ecological shifts and the direct introduction of functional metabolic genes[79].
Indirect modulation via bacterial intermediaries: Phages dictate the “metabolic potential” of the gut by controlling the abundance of key fermentative bacteria. By regulating the population dynamics of bacteria such as Bifidobacterium and Bacteroides, phages indirectly modulate the production of short-chain fatty acids (SCFAs)-metabolites essential for energy homeostasis and intestinal barrier signaling[80]. The significant expansion of bacterial diversity in the first two years is paralleled by a reduction in phage richness, suggesting that phages “thin the ranks” of dominant bacteria, allowing a more metabolically diverse community to emerge[81]. Vertical transmission of Bifidobacterium (specifically via maternal milk) supports the early establishment of Bifidobacterium populations. These bacteria are critical for the metabolism of HMOs, a foundational process for neonatal energy balance[82].
Viral AMG: Bacteriophages are not merely genetic parasites; they are biological “upgraders”. Through lysogenic integration, phages introduce viral AMG into the bacterial host’s genome[83]. These genes can be involved in carbon, nitrogen, or sulfur metabolism pathways. In the pediatric gut, these genes may allow bacteria to more efficiently utilize available nutrients during the transition to solid foods (weaning), essentially boosting the microbiome’s functional repertoire during times of environmental stress[84].
Systemic metabolic programming: Early-life virome perturbations may have long-lasting effects on host adiposity and glucose regulation. Shifts in the viral-bacterial balance can alter the metabolic “set-point” during the first 1000 days. A lack of temperate phages or a “bloom” of lytic phages can cause a loss of metabolic diversity, which has been linked to conditions such as pediatric obesity and the early stages of T1D[85].
The assembly of the paediatric virome is not purely biological inevitability; it is an ecological process heavily influenced by external pressures. From medical interventions to daily nutrition, these modifiers act as “environmental sculptors” that can promote a resilient, diverse viral ecosystem or drive it toward persistent dysbiosis[86]. Understanding these factors is critical for developing preventive public health strategies (Table 4).
| Modifier | Key impact on virome | Long-term consequence |
| Antibiotics | SOS response and prophage induction | Bacterial instability; expansion of ARGs |
| Breast milk | Transfer of sIgA and maternal phages | Enhanced barrier protection; regulated assembly |
| Oral vaccines | Competition with resident viruses | Potential for altered vaccine “take” and immune priming |
| NICU Stay | Reduced diversity; hospital signatures | Increased risk of NEC and delayed immune maturation |
Antibiotics are among the most frequently prescribed medications in paediatrics, and their impact on the virome is profound and often overlooked. While antibiotics target bacteria, they indirectly trigger a “viral explosion”. Many antibiotics induce DNA damage in bacteria, triggering the SOS response. This causes dormant prophages to “wake up”, enter the lytic cycle, and burst from their bacterial hosts. This leads to massive, sudden viral blooms and the depletion of beneficial bacterial populations[87]. By killing the bacterial hosts that phages depend on for replication, broad-spectrum antibiotics can cause a secondary “virome collapse”, leading to a significant loss of viral diversity that may take months to recover. While less common in infancy, the use of antivirals (e.g., for respiratory syncytial virus or flu) can inadvertently suppress commensal eukaryotic viruses (like Anelloviridae), potentially removing the “tonic” immune stimulation necessary for healthy maturation[88].
Diet and feeding patterns: The foundational input: Diet is the primary architect of the early-life microbiome. The transition from milk to solid foods represents a major “regime shift” for the virome. Breast Milk acts as a continuous delivery system for maternal secretory IgA, which coats and neutralizes potentially pathogenic viruses. It also harbors its own unique virome and HMOs that support specific bacteria (e.g., Bifidobacterium), which, in turn, support a healthy, stable phage community[89]. Infants fed exclusively on formula often exhibit a different successional pattern, sometimes characterized by an earlier expansion of certain bacteriophages and a lack of the protective maternal antibodies found in milk. The “beech-nut” transition (introduction of complex plant polysaccharides) leads to a surge in viral diversity as new bacterial niches are created, inviting a more complex array of temperate and lytic phages into the ecosystem[90].
Vaccination: Oral vaccines and virome shaping: Vaccines do more than protect against specific pathogens; they interact dynamically with the resident virome. Oral Vaccines (Rotavirus and Polio) introduce live-attenuated viruses directly into the gut. They must compete with the resident virome for “space” and immune attention[91]. Emerging evidence suggests that a child’s baseline virome composition can influence the efficacy of oral vaccines. A “noisy” or highly inflamed virome may interfere with the vaccine’s ability to replicate and induce immunity. Conversely, the introduction of vaccine strains can lead to transient shifts in the resident phage communities, effectively “re-shuffling” the microbial deck[92,93].
Hospitalization and neonatal intensive care unit exposure: The environment of a neonatal intensive care unit (NICU) is fundamentally different from a home environment, leading to an “artificial” virome assembly. Infants in the NICU are often colonized by hospital-associated viral signatures. These viromes tend to be less diverse and more unstable, dominated by phages that target common NICU pathobionts such as Klebsiella and Enterococcus[94]. High levels of sanitation and the use of medical equipment can limit the exposure to “old friend” commensal viruses, while simultaneously creating niches for resistant viral-bacterial communities to persist. Paradoxically, the hospital’s high-hygiene environment can prevent the healthy “priming” of the immune system by reducing the overall diversity of viral encounters during the critical first weeks of life[95].
The paediatric gut virome influences health and disease through both local gastrointestinal mechanisms and systemic immune, metabolic, and neurodevelopmental pathways. Alterations in virome composition, diversity, and functional capacity-collectively termed virome dysbiosis-have been increasingly implicated in a wide range of paediatric disorders. Importantly, the causal direction of these associations remains an area of active investigation, as researchers work to determine whether viral shifts are drivers or markers of pathology[37].
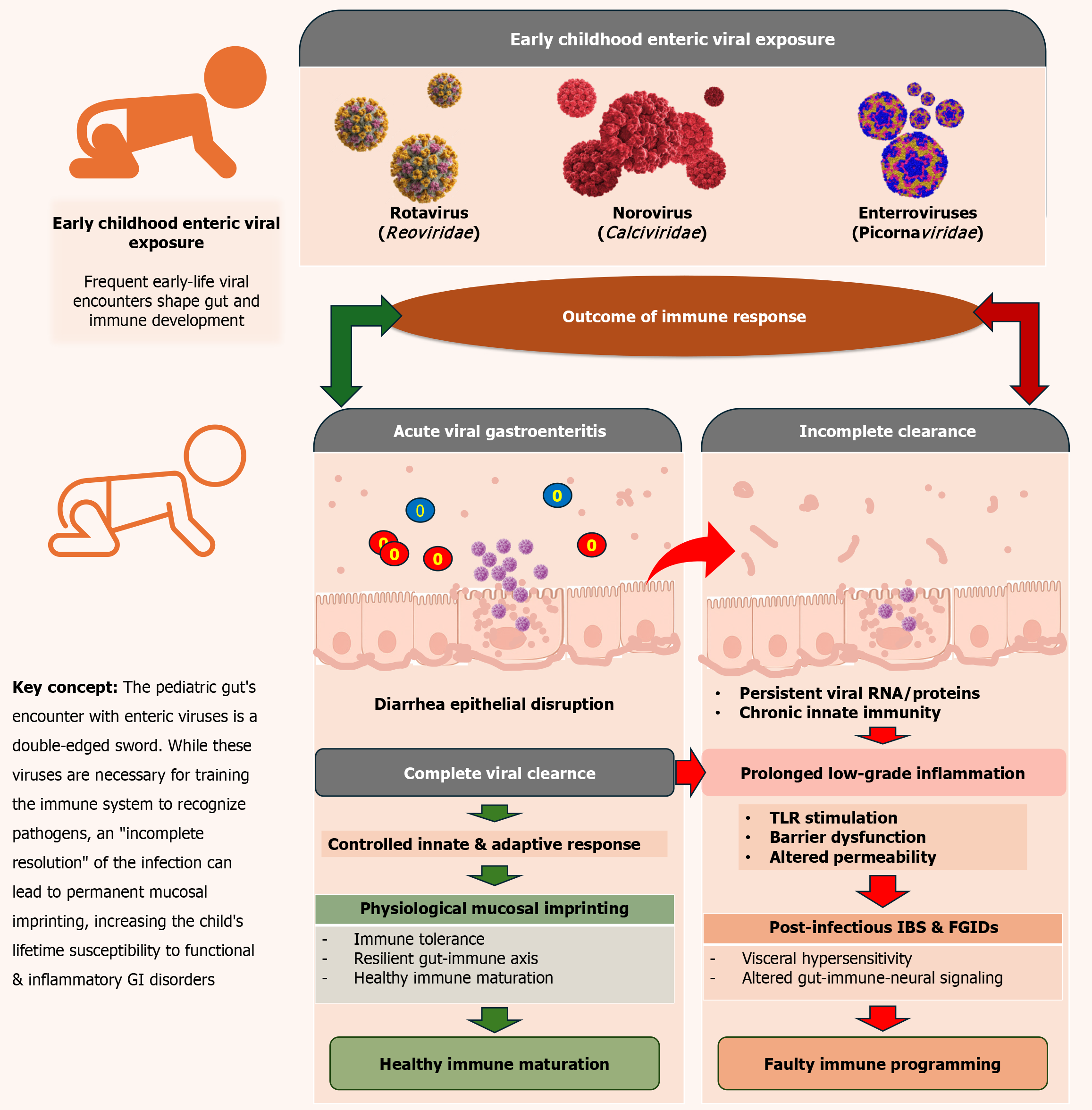
Infectious and post-infectious gastrointestinal conditions: Early childhood is characterized by frequent encounters with enteric viruses, which act as primary drivers of pediatric morbidity and essential catalysts for immune development. Acute viral gastroenteritis is the most common clinical manifestation of these encounters, typically involving highly contagious RNA viruses such as Rotavirus (family Reoviridae), Norovirus (family Caliciviridae), and Enteroviruses (family Picornaviridae)[96]. While these infections are often viewed as self-limiting, emerging evidence suggests that the viral “shadow” they leave behind can have lasting consequences (Table 5).
| Condition | Primary virome alterations (effects) | Putative biological mechanism |
| Acute gastroenteritis | High load of Reoviridae (Rotavirus), Caliciviridae (Norovirus) | Mucosal Imprinting: Incomplete clearance leads to prolonged immune activation and “sculpting” of future immune tone |
| Inflammatory bowel disease | Expansion of Caudoviricetes; Contraction of overall viral diversity | Lytic pressure and HGT: Phage blooms deplete beneficial bacteria (e.g., Faecalibacterium prausnitzii) and transfer virulence genes via horizontal gene transfer |
| Necrotizing enterocolitis | Sudden “phage blooms” preceding symptoms; Low virome stability | The triple hit: Antibiotic-induced SOS response triggers phage lysis; viral translocation across leaky barrier activates TLR-mediated necrosis |
| Celiac disease | Presence of candidate viruses (Reovirus, Enterovirus) | Loss of Tolerance: Viruses act as “danger signals” that disrupt oral tolerance to gluten, triggering Th1-mediated immune priming in HLA-susceptible children |
| Functional gastrointestinal disorders | Specific early-life signatures; post-infectious viral “shadows” | Neuro-immune sensitization: Low-grade inflammation and viral-neural crosstalk sensitize nociceptors, leading to visceral hypersensitivity |
In many cases, viral clearance is incomplete, leading to the persistence of viral genomic fragments or proteins within the intestinal mucosa. This persistence maintains prolonged immune activation, in which the innate immune system remains “primed” long after clinical resolution of symptoms[97]. The transition from an acute infection to a chronic functional disorder is a significant concern in pediatrics. Post-infectious irritable bowel syndrome (IBS) is thought to arise when an acute viral insult disrupts the delicate balance of the gut-immune-neural axis. This disruption can lead to persistent low-grade inflammation, altered intestinal permeability, and visceral hypersensitivity, which are hallmarks of functional abdominal pain disorders[98].
A central theme in pediatric viromics is that early-life infections do not occur in a vacuum; they provide the “first draft” of the gut’s immunological memory. This mucosal imprinting means that the specific viral signatures encountered in infancy determine the “tone” of future immune responses[43]. Incomplete immune resolution during these early stages may lead to “faulty” programming. For example, while a robust adaptive immune response is required to “tame” the eukaryotic viral load as a child matures, an aberrant response may instead lead to chronic inflammatory sensitivity[99]. The pediatric gut’s encounter with enteric viruses is a double-edged sword. While these viruses are necessary for training the immune system to recognize pathogens, an “incomplete resolution” of the infection can lead to permanent mucosal imprinting, increasing the child’s lifetime susceptibility to functional and inflammatory gastrointestinal disorders. Figure 3 shows the viral pathways and gut immune development[100].
IBD: IBD, encompassing Crohn’s disease (CD) and ulcerative colitis (UC), represents a profound failure of immune-microbial homeostasis. In the pediatric context, IBD is often more extensive and aggressive than in adults, suggesting that early-life disruptions in the microbial ecosystem are central to its pathogenesis[101]. While the bacterial “bacteriome” has been extensively mapped, the virome is now emerging as a critical anchor in the inflammatory cascade[102].
Pediatric IBD patients exhibit a distinct “viral signature” that differs significantly from healthy peers. This dysbiosis is characterized by two primary shifts. The first shift is a contraction of diversity, which is a marked reduction in the overall alpha-diversity (richness and evenness) of the gut virome[103]. This “shrunken” viral landscape mirrors the loss of bacterial diversity typically seen in IBD[104]. The second shift is the expansion of Caudoviricetes. Paradoxically, while overall diversity falls, there is often a massive expansion of bacteriophages belonging to the Caudoviricetes class (tailed phages). This bloom is not uniform; it is frequently characterized by a “stochastic” or chaotic increase in specific phage taxa that correlates with high levels of intestinal inflammation[105].
The expansion of Caudoviricetes is not merely a bystander effect; it actively destabilizes the bacterial community. A “bloom” of lytic phages (lytic pressure) can target and deplete beneficial, anti-inflammatory bacteria (such as Faecalibacterium prausnitzii), essentially removing the “brakes” on intestinal inflammation[106]. Phages in the IBD gut serve as hyperactive vehicles for horizontal gene transfer (HGT). They can transfer virulence factors or metabolic genes that allow pathobionts to thrive in an oxidative, inflamed environment, further entrenching the disease state (HGT)[107].
In addition, the intestinal epithelium and underlying immune cells are equipped with receptors designed to “see” viruses. In IBD, this sensing goes into overdrive. Viral DNA and RNA fragments, released during massive phage-driven bacterial lysis, act as damage-associated molecular patterns (DAMPs)[108]. These are sensed by TLR9 (DNA) and TLR3/7 (RNA), triggering the production of Type I interferons and pro-inflammatory cytokines, such as tumor necrosis factor-alpha (TNF-α). Moreover, in the “leaky gut” of a pediatric IBD patient, these viral particles translocate across the epithelial barrier, directly activating mucosal dendritic cells and T-cells, thereby sustaining chronic inflammation (Barrier Failure)[109].
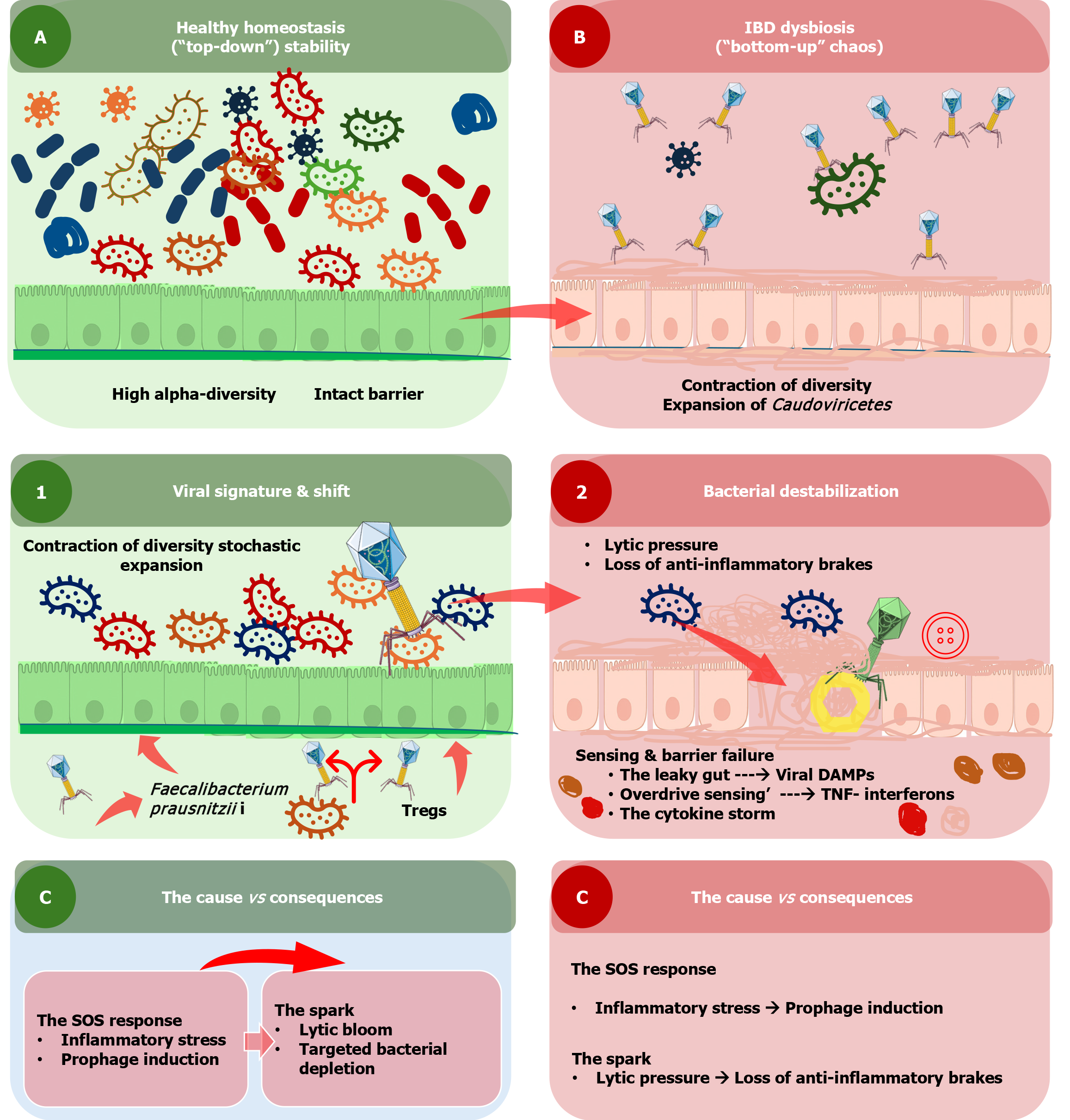
However, one of the most debated topics in pediatric viromics is whether virome dysbiosis triggers IBD or is a result of the inflamed microenvironment (cause vs consequence). In the “consequence” argument, inflammation, oxidative stress, and rapid transit times in the IBD gut may trigger the “SOS response” in lysogenic bacteria, leading to prophages entering the lytic cycle and “bloom[110]”. In the “cause” Argument, longitudinal studies in at-risk children suggest that viral shifts can sometimes be detected before clinical symptoms emerge, hinting that the virome may act as an environmental “spark” that ignites the inflammatory process in genetically susceptible individuals (Figure 4)[111]. Therefore, in pediatric IBD, the virome shifts from a state of “top-down” stability to one of “bottom-up” chaos. The expansion of tailed phages and the resulting loss of bacterial diversity create a pro-inflammatory feedback loop that is sensed by the host’s innate immune system, making the virome a promising target for future precision diagnostics and “phage-based” therapeutics[37].
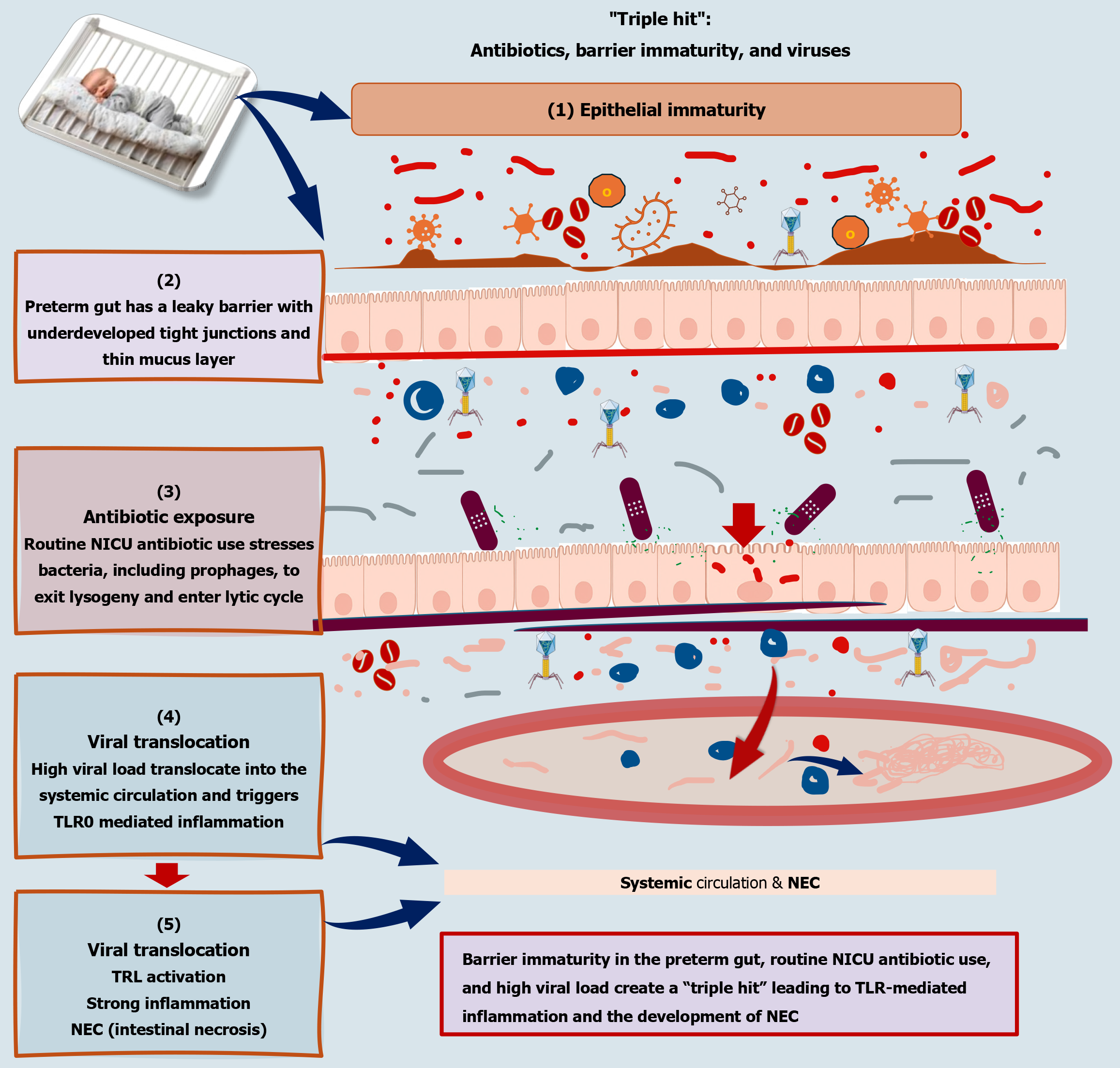
Panel A: Healthy homeostasis (“top-down” stability): In health, the gut virome exhibits high alpha-diversity, composed of a balanced mixture of bacteriophages and eukaryotic viruses. This diverse virome coexists with a rich, stable bacterial community, including anti-inflammatory commensals such as Faecalibacterium prausnitzii. An intact epithelial barrier with well-formed tight junctions and a thick mucus layer prevents microbial translocation. Immune tolerance predominates, supported by Tregs within the lamina propria and low-level, controlled immune signaling.
Panel B: IBD dysbiosis (“bottom-up” chaos): IBD is characterized by contraction of virome diversity and a disproportionate expansion of Caudoviricetes bacteriophages, as evidenced by stochastic phage blooms. Increased lytic phage activity leads to targeted depletion of beneficial bacteria, removing key anti-inflammatory “brakes”. Concurrently, phage-mediated HGT may enhance bacterial virulence in pathobionts. Barrier integrity deteriorates, resulting in a “leaky gut” that allows viral nucleic acids and viral DAMPs to interact with epithelial pattern-recognition receptors, including TLR3, TLR7, and TLR9. This overactivation triggers exaggerated cytokine responses, notably TNF-α and type I interferons, which promote mucosal inflammation. Activated dendritic cells and effector T cells further amplify tissue injury and immune dysregulation.
Panel C: Cause-consequence feedback loop: A self-perpetuating pro-inflammatory loop links host inflammation and virome instability. Inflammatory stress induces bacterial SOS responses, leading to prophage activation and lytic induction. Subsequent phage blooms cause further bacterial depletion and dysbiosis, reinforcing mucosal inflammation and sustaining disease activity in paediatric IBD.
NEC: NEC remains one of the most devastating and unpredictable gastrointestinal emergencies in the NICU. While the role of bacterial “dysbiosis” (specifically the Proteobacteria bloom) is well documented, the virome is now recognized as a critical, albeit understudied, layer in the pathogenesis of this condition[112].
Infants born prematurely do not just have immature lungs and brains; they have a “stunted” virome assembly (im
Moreover, recent longitudinal studies suggest that the virome may provide a “window into the future” regarding NEC risk. A significant expansion or “bloom” of specific bacteriophages (phage blooms) has been observed to occur days before the clinical onset of NEC symptoms[115]. These blooms are often directed at pioneer bacterial colonizers. By lysing these bacteria, phages release endotoxins and inflammatory triggers (DAMPs) directly into the gut lumen, potentially acting as the “spark” that initiates the necrotic process (destabilization)[116].
Therefore, we can view NEC pathogenesis in the context of the virome as a “triple hit” model. The first hit is marked by epithelial immaturity. Preterm infants have a “leaky” intestinal barrier with poorly developed tight junctions and a thin mucus layer. The second hit could be triggered by antibiotic exposure. Routine antibiotic use in the NICU acts as a massive stressor. It induces the SOS response in bacteria, causing prophages to “wake up” and enter the lytic cycle, leading to mass bacterial lysis and viral blooms. These changes are followed by the third hit: Viral translocation. The combination of barrier immaturity and high viral load allows phages and viral nucleic acids to translocate into the systemic circulation, where they trigger TLR-mediated inflammatory cascades that lead to intestinal necrosis[117-119] (Figure 5).
Because viral changes often precede bacterial changes and clinical symptoms, the virome holds immense potential for precision neonatology. Identifying specific “NEC-associated viral signatures” in stool samples could enable early intervention, such as adjusting feeding protocols or preemptively using probiotics, before the disease becomes life-threatening[115]. It is important to note that while the associations are compelling, the field is still in its infancy. Current evidence is primarily based on small, longitudinal cohort studies. We are still moving from correlation (observing viral blooms in NEC cases) to causation (proving that these blooms actually drive necrosis).
Celiac disease: Celiac disease is a complex autoimmune-like disorder triggered by the ingestion of gluten in genetically susceptible individuals (carrying HLA-DQ2 or HLA-DQ8). While gluten is the necessary antigen and HLA genes provide the genetic “fertile ground”, only a fraction of at-risk individuals develop the disease[120]. The gut virome is increasingly recognized as a critical environmental trigger that can disrupt the delicate balance between oral tolerance and inflammatory immunity[58]. In the “hit and run” model of autoimmunity, a transient viral infection in early life may per
The primary mechanism by which viruses contribute to Celiac disease is the disruption of oral tolerance. Research has shown that certain viruses, specifically Reoviruses (e.g., type 1 lang strain), can interfere with the way the gut processes food antigens[122]. These viruses trigger a pro-inflammatory Th1 immune response in the presence of gluten, overriding the natural Treg response that usually promotes tolerance. Viral sensing leads to the production of Interferon-regulatory factor 1, which blocks the protective signaling of retinoic acid, a key molecule for maintaining oral tolerance[123].
Moreover, Celiac disease pathogenesis is often described as a “perfect storm” involving three converging factors: HLA susceptibility, gut permeability, and immune activation. Genetic predisposition (HLA-DQ2/8) enables the efficient pre
As noted, reoviruses are the most studied mechanistic candidates for initiating Celiac-related immune skewing. In addition, prospective studies (such as the TEDDY study) have linked prolonged Enterovirus infections in early life with a higher risk of developing Celiac disease-associated antibodies. In addition, early, severe Rotavirus infections have also been correlated with increased risk, possibly due to the massive epithelial damage and subsequent immune priming[126]. However, it is vital to distinguish the virome’s role in Celiac disease from classical infectious diseases. The virus does not “cause” celiac disease in the traditional sense; rather, it modulates the immune system’s response to an external antigen (gluten)[127]. In addition, the viral infection may be long gone by the time clinical symptoms of Celiac disease appear (temporal gap), making it a “molecular scar” on the immune system rather than an ongoing infection[128]. In Celiac disease, the virome acts as a biological “switch”. By activating innate immune pathways during critical windows of early development, viruses can turn a benign dietary exposure into a lifelong autoimmune condition, particularly in genetically vulnerable children[129].
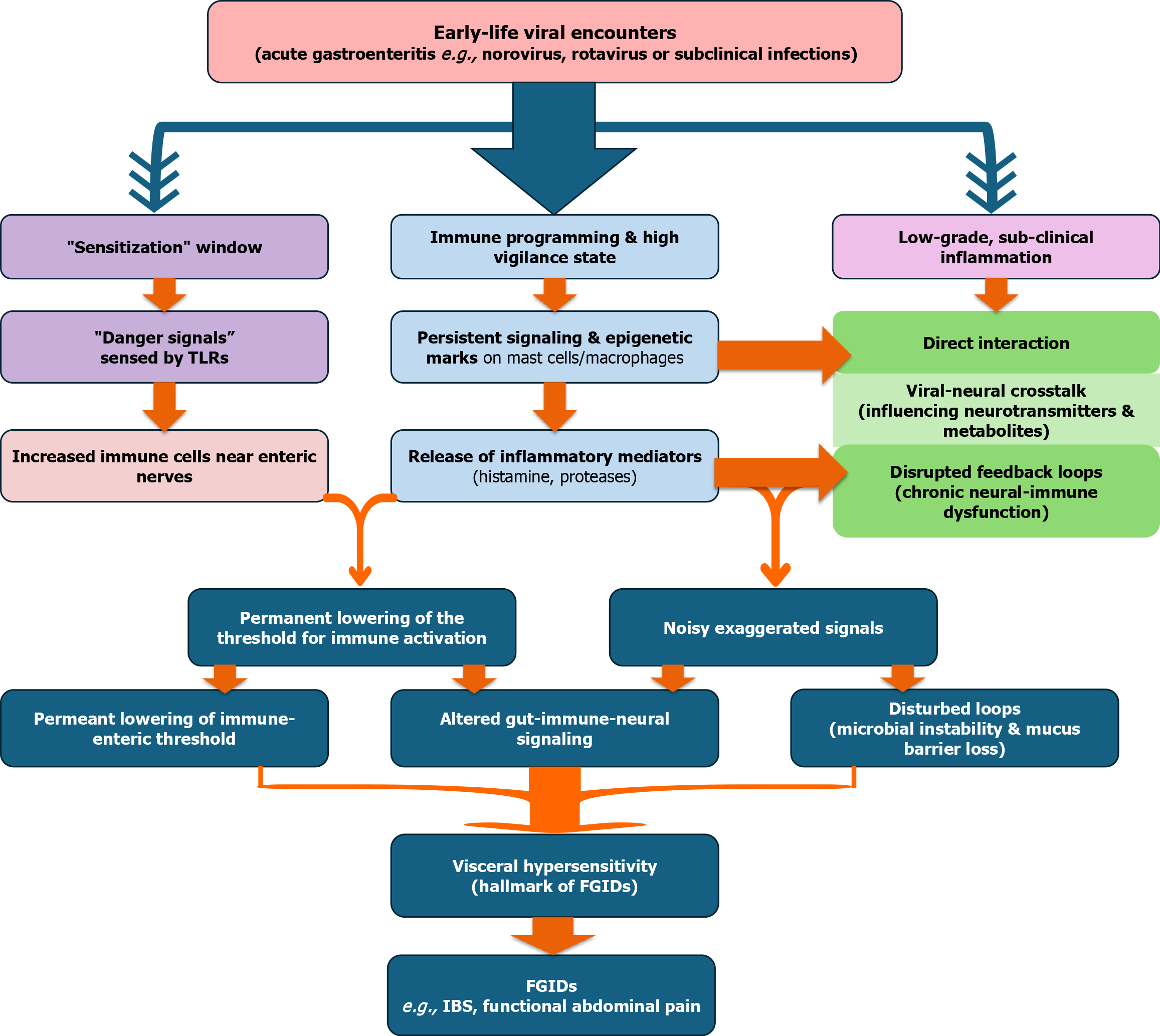
Functional gastrointestinal disorders: Functional gastrointestinal disorders (FGIDs), such as IBS and functional ab
A hallmark of FGIDs is visceral hypersensitivity-a state where normal intestinal processes (like gas or peristalsis) are perceived as painful. Acute viral insults can leave behind a “molecular hangover” of low-grade, sub-clinical inflammation. While the gut looks “normal” on a standard biopsy, there is an increased density and activation of immune cells near enteric nerve endings. Inflammatory mediators released in response to viral triggers (such as histamine or proteases) directly sensitize nociceptors (pain-sensing neurons). Over time, this results in “peripheral sensitization”, in which the gut becomes hyperreactive to even minor distension[134,135].
The virome sits at the intersection of a complex three-way conversation between the microbiome, the immune system, and the ENS. Some bacteriophages and eukaryotic viruses can interact directly with the ENS (viral-neural crosstalk)[136]. By modulating the local microbial environment, viruses influence the production of neurotransmitters (like serotonin) and microbial metabolites that regulate gut motility and pain perception. In children with FGIDs, this signaling loop is often “noisy” or exaggerated[137]. The virome’s role in maintaining the mucus barrier and regulating bacterial “blooms” is essential; when this balance is lost, the resulting microbial instability keeps the neural-immune axis in a state of chronic dysfunction (disrupted feedback loops) (Figure 6)[138].
The strongest clinical evidence for the virome’s role in FGIDs comes from post-infectious syndromes. A significant portion of pediatric IBS cases begin immediately following an episode of acute viral gastroenteritis (e.g., norovirus or rotavirus)[139]. While most children return to baseline, a subset-potentially due to genetic predisposition or specific early-life virome signatures-fails to “reset” their gut-immune-neural axis. This leads to a transition from an acute infectious state to a chronic functional pain disorder. In FGIDs, the virome is a “silent programmer”. It doesn’t cause visible damage like IBD, but it shapes the neuro-immune architecture of the developing gut. Early viral “hits” can imprint a state of visceral hypersensitivity, turning a transient childhood infection into a long-term functional challenge[110].
While the gut is the primary site of viral activity, the virome’s influence extends far beyond the intestinal wall. By modulating systemic inflammation, metabolic precursors, and neuro-immune signaling, the paediatric virome acts as a systemic “biostat” that can influence the risk of endocrine, metabolic, and neurodevelopmental disorders[140] (Table 6).
| Condition | Primary virome alterations (effects) | Putative biological mechanism |
| Obesity and insulin resistance | Reduced viral diversity; altered phage-to-bacteria ratios | Metabolic supervision: Phages modulate the abundance of SCFA-producing bacteria, influencing energy harvest and systemic lipid metabolism |
| Type 1 diabetes | Enteroviral persistence; altered phage communities | Molecular mimicry and priming: Enteroviruses may directly target beta cells, while phages prime the innate immune system toward pancreatic autoimmunity |
| Neurodevelopment (ASD/ADHD) | Distinct viral “fingerprints”; reduced phage diversity | Microglial activation: Gut-derived viral signals influence neuroinflammation and aberrant synaptic pruning during critical developmental windows |
| Allergy and asthma | Low infantile viral diversity; absence of “Old Friend” commensal viruses | Th2 skewing: Lack of viral-driven Th1/Treg induction prevents the correction of neonatal Th2 bias; disrupted Gut-Lung Axis signaling |
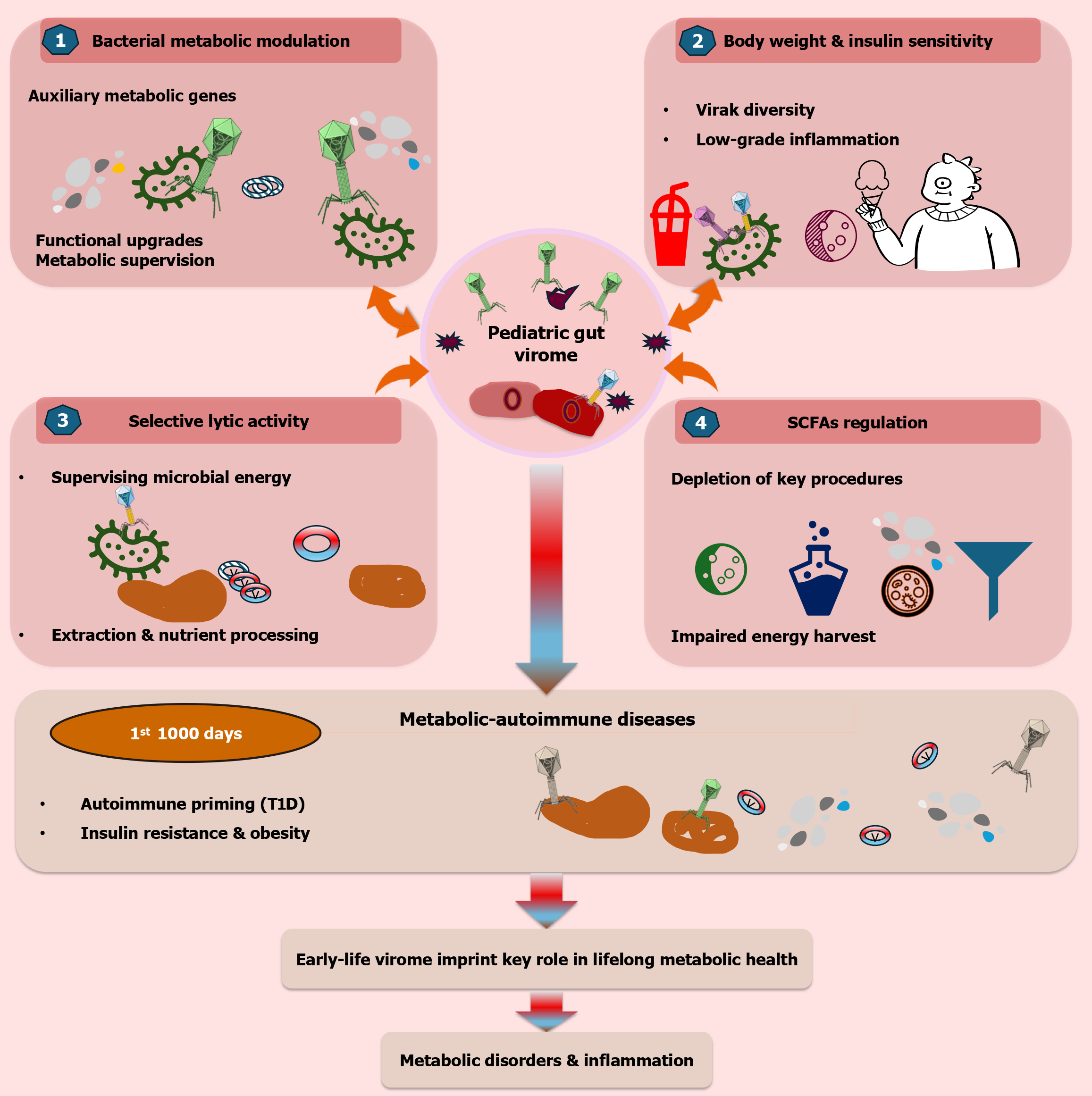
The intersection between the paediatric gut virome and host metabolism represents a rapidly evolving frontier in paediatric gastroenterology and endocrinology. Beyond its role in shaping microbial ecology, the virome exerts a sustained influence on host metabolic programming through a dynamic virome-bacteriome-host axis. This interaction operates via both direct genetic contributions to microbial metabolism and indirect immune-mediated effects that in
Virome-driven modulation of bacterial metabolic pathways: Bacteriophages are not passive passengers within the gut ecosystem; rather, they function as active metabolic regulators of bacterial communities. Through HGT, phages introduce AMGs that can rewire bacterial metabolic capabilities[142]. Phage-encoded AMGs may enhance bacterial survival and metabolic efficiency under stress conditions common in the infant gut, such as fluctuating oxygen tension, dietary transitions, and immune pressure. These genes can enable bacteria to metabolize complex polysaccharides, resist oxidative stress, or adapt to inflammatory environments[143]. By selectively lysing specific bacterial populations, phages exert top-down control over microbial metabolic output. This “kill-the-winner” or “piggyback-the-winner” dynamic determines which metabolic pathways dominate at a given developmental stage, effectively supervising nutrient flux, energy extraction, and metabolite availability to the host[144]. Collectively, these processes position the virome as a metabolic gatekeeper, shaping not only bacterial composition but also the biochemical signals delivered to the developing host[145].
Associations with obesity and insulin resistance: Accumulating evidence links early-life virome configurations with long-term metabolic phenotypes, including obesity and insulin resistance. Paediatric cohorts have demonstrated that children with obesity often harbor a less diverse gut virome, characterized by the expansion of specific bacteriophage families and reduced viral richness[146]. This mirrors patterns observed in bacterial dysbiosis and suggests that viral diversity may serve as an early biomarker of metabolic risk. Alterations in phage populations targeting key bacterial phyla (e.g., Bacteroidetes vs Firmicutes) have been associated with disrupted glucose metabolism and chronic low-grade inflammation[147]. This inflammatory milieu contributes to impaired insulin signaling and may precede the onset of overt metabolic disease. Importantly, these associations appear strongest when virome alterations occur during critical developmental windows, reinforcing the concept of early-life metabolic imprinting[148].
Viral influences on SCFA production: SCFAs-including acetate, propionate, and butyrate-are central mediators of host energy balance, immune regulation, and epithelial integrity. The virome influences SCFA availability by regulating bacterial producers[149]. Phages modulate the abundance and functional capacity of key SCFA-producing taxa, such as Bifidobacterium and Faecalibacterium prausnitzii. Balanced phage activity supports metabolic resilience, whereas excessive lytic pressure may lead to depletion of these beneficial organisms. Reduced SCFA production is associated with impaired intestinal barrier integrity, altered lipid metabolism, and dysregulated appetite signaling[149]. In paediatric populations, such disturbances may contribute to increased energy harvest efficiency, adiposity, and susceptibility to metabolic inflammation. Thus, virome-mediated shifts in SCFA dynamics represent a plausible mechanistic link between early viral ecology and later metabolic outcomes[150].
The paediatric virome and T1D: T1D is the most extensively studied metabolic-autoimmune condition in relation to the gut virome, with evidence supporting both direct viral effects and indirect immune-mediated pathways. Classical models implicate eukaryotic viruses-particularly enteroviruses such as coxsackievirus B-in the initiation of beta-cell autoimmunity. Proposed mechanisms include direct pancreatic infection, bystander inflammation, and molecular mimicry between viral and beta-cell antigens[151]. Emerging data suggest a complementary role for bacteriophages. In genetically susceptible children, increased intestinal permeability may allow phage to translocate into the systemic circulation[152]. These viral particles and nucleic acids can activate innate immune pathways (e.g., TLR signaling), fostering a pro-inflammatory immune tone that lowers the threshold for autoimmune activation against pancreatic beta cells[153]. Much of the current human evidence remains associative, derived from longitudinal birth cohorts such as TEDDY. However, animal models increasingly support a mechanistic link, demonstrating that virome-induced gut inflammation can accelerate progression from islet autoimmunity to overt diabetes[154].
The paediatric gut virome does far more than process dietary substrates or eliminate bacterial competitors-it actively programs the host’s metabolic and endocrine trajectory. By regulating bacterial metabolism, shaping SCFA availability, influencing immune tone, and potentially contributing to autoimmune risk, the virome emerges as a critical determinant of long-term metabolic health[155]. The viral landscape established during the first 1000 days of life may therefore represent a modifiable window for preventing obesity, insulin resistance, and autoimmune endocrine disorders[156].
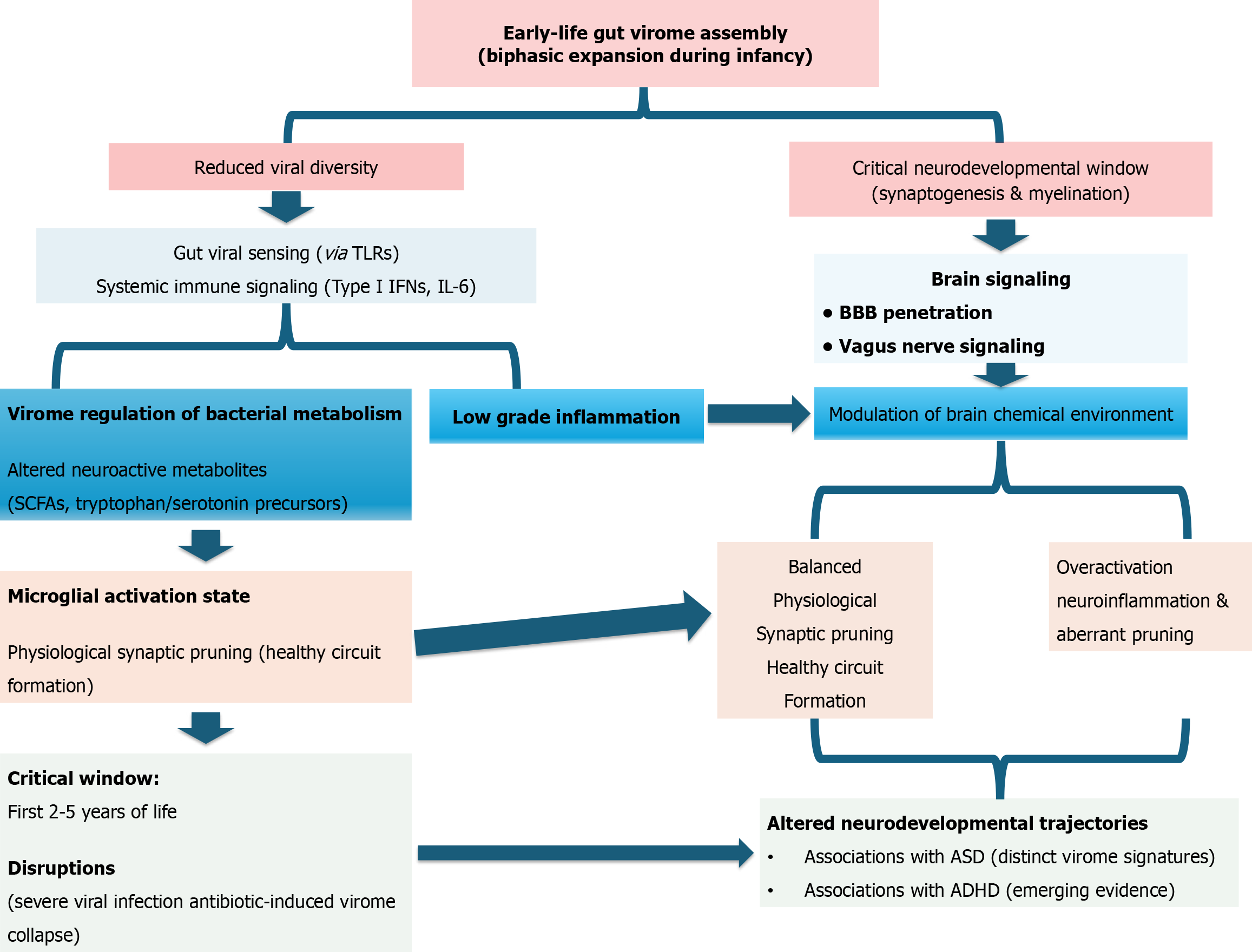
The “gut-brain axis” has traditionally focused on bacterial metabolites. However, the virome is now emerging as a critical anchor in this bidirectional communication network. Because the most rapid period of brain development-synaptogenesis and myelination-coincides with the “biphasic” expansion of the gut virome, this period represents a unique window of neuro-immunological programming[157].
The virome doesn’t communicate with the brain in isolation; it utilizes the immune system as a primary conduit. Viral sensing in the gut (via TLRs) leads to the production of systemic cytokines (e.g., type I interferons, interleukin-6). These molecules can cross the blood-brain barrier or signal via the vagus nerve to influence brain function (the cytokine relay)[158]. By regulating bacterial populations that produce neurotransmitter precursors (like tryptophan for serotonin) or SCFAs, phages indirectly modulate the chemical environment of the developing brain (metabolic intermediaries)[159].
Microglia are the resident immune cells of the brain, responsible for “synaptic pruning” the removal of weak neural connections to streamline brain function. Low-level, “tonic” signaling from gut viruses helps set the microglial activation threshold (the “priming” signal)[160]. If the gut virome is dysbiotic (e.g., a massive lytic bloom), the resulting systemic inflammatory signals can drive microglial overactivation. This leads to neuroinflammation and aberrant synaptic pruning, which are hallmarks of several neurodevelopmental disorders[161].
Research into the virome signatures of children with neurodevelopmental challenges is a rapidly growing field. Children with autism spectrum disorder (ASD) often exhibit significant gastrointestinal comorbidities. Metagenomic studies have identified a distinct viral “fingerprint” in ASD cohorts, often characterized by reduced phage diversity and overrepresentation of specific eukaryotic viral families compared to neurotypical siblings[162]. While data is more limited, preliminary studies suggest that early-life viral-bacterial imbalances may correlate with the hyper-inflammatory states sometimes associated with attention-deficit/hyperactivity disorder (ADHD) symptoms. Imbalances in viral/bacterial compositions (dysbiosis) in early life, often linked to inflammation, can affect brain development and are associated with increased ADHD risk and symptoms, potentially by disrupting the gut-brain axis, increasing systemic inflammation, and altering neurotransmitters, though the specific microbial profiles and mechanisms are still being explored[163,164].
The timing of virome assembly is paramount. There is a “critical window” during the first 2 to 5 years of life where the brain is most sensitive to environmental inputs. The peak of eukaryotic viral expansion (around age 2) mirrors a period of intense cognitive and language development (coincidental peaks)[165]. Disruptions during this window—such as severe viral infections or antibiotic-induced virome collapse-may lead to permanent “rewiring” of neural circuits, potentially increasing the risk of long-term neurodevelopmental or psychiatric conditions[166] (Figure 8).
It is imperative to maintain scientific caution when distinguishing between association and causality. Currently, we cannot state that a specific virome signature “causes” ASD or ADHD. These are associations. It remains unclear whether virome changes drive neurodevelopmental shifts or whether lifestyle/dietary factors associated with these conditions (e.g., restrictive eating in ASD) drive virome changes. The gut virome acts as a “remote control” for brain health. By modulating systemic immune tone and microglial activity during critical early-life windows, gut viruses contribute to the delicate process of neural architecture development[79]. While current evidence is associative, the virome-immune-brain axis offers a provocative new target for understanding and potentially supporting neurodevelopment. Importantly, observed associations between early-life virome patterns and neurodevelopmental outcomes must be interpreted in the context of potential confounders, including diet composition, antibiotic exposure, mode of feeding, environmental microbial load, and socioeconomic factors, all of which independently shape the gut ecosystem and neuroimmune development.
The dramatic rise in pediatric allergic and autoimmune conditions over the last half-century has long been linked to the “hygiene hypothesis”. However, the virome offers a modern refinement of this theory: It is not just a lack of bacteria, but a lack of viral-driven immune education that may underlie the modern epidemic of “atopic” and “auto-aggressive” immune systems[167].
The original hygiene hypothesis suggested that cleaner environments lead to less microbial exposure, resulting in an “untrained” immune system[168]. Through a viromic lens, this is expanded to include the “old friends” hypothesis. Early-life exposure to a diverse array of bacteriophages is essential for keeping bacterial populations in check and pro
Newborns are naturally “Th2-skewed”, a state that favors allergic-type responses. For a healthy transition, the immune system must be “skewed” toward a Th1/Treg dominant state. Viral nucleic acids (DNA/RNA) sensed via TLR3, 7, and 9 are powerful drivers of Th1 differentiation[171]. This shift is essential for effective anti-pathogen defense and for counteracting the Th2 allergic bias. Healthy viral-bacterial interactions promote the expansion of Tregs[172]. These cells act as the “police” of the immune system, preventing it from overreacting to pollen, dust, or the host’s own tissues. Without suf
The virome’s influence is not confined to the gut; it is a major player in the gut-lung axis, influencing the development of asthma and allergic rhinitis. Cytokines and activated immune cells (like dendritic cells) “trained” by viruses in the gut travel through the lymphatic system to the lungs[174]. By shaping the production of SCFAs in the gut, the virome indirectly supports the integrity of the lung’s mucosal barrier. Longitudinal studies have shown that children with higher gut viral diversity in infancy have a significantly lower risk of developing asthma by school age, suggesting that the “gut virome signature” can predict respiratory health years in advance[175].
The mechanisms driving allergic diseases (Th2 dominance/Lack of Tregs) often overlap with the roots of autoimmunity, such as the previously discussed T1D. Both conditions stem from the immune system’s failure to “reset” or “regulate” itself after an initial stimulus[176]. The lack of general viral diversity can predispose the system to such triggers by failing to build a robust regulatory infrastructure[177].
In the context of allergy and autoimmunity, the pediatric virome acts as a rheostat for immune sensitivity. It provides the signals needed to shift the host away from an allergic (Th2) bias toward a regulated (Treg/Th1) state. The modern “depleted” virome may be a primary driver of the rising rates of asthma, allergies, and autoimmune conditions in the pediatric population[178].
As our understanding of the paediatric virome transitions from descriptive to mechanistic, the focus shifts toward clinical application (Table 7). The virome represents an untapped frontier for precision paediatrics, offering new avenues for early diagnosis and highly specific therapeutic interventions that could minimize the “collateral damage” often associated with broad-spectrum treatments.
| Modality | Strategy | Primary advantage |
| Biomarkers | Viral “Fingerprinting” | Early detection before symptoms or bacterial shifts. AI-driven “Early Warning Systems” for high-risk neonates |
| Phage therapy | Targeting pathobionts | Highly specific; preserves the “good” microbiome. Routine “precision editing” for pathobiont-driven IBD |
| FVT | Sterile filtrate transfer | Safer than whole-fecal transplants for children. FVT is a primary “reset” for metabolic and immune health |
| Nutrition | Indirect modulation | Scalable and low risk for long-term health |
Traditional diagnostics often identify disease only after significant tissue damage or an inflammatory cascade has begun. The virome offers a “pre-symptomatic” window, allowing for a shift from reactive to proactive medicine. Because the virome often shifts before significant bacterial dysbiosis or clinical symptoms occur, it can serve as a “biological early-warning system[179]”. In conditions such as NEC or T1D, specific phage blooms or eukaryotic viral infections can be detected in stool samples weeks before clinical onset[180]. Stool-based virome sequencing offers a non-invasive way to monitor high-risk infants (e.g., preterm neonates in the NICU) to intervene before irreversible tissue damage occurs. Viral signatures may help clinicians distinguish between different disease phenotypes (e.g., CD vs UC) that may appear similar but require different therapeutic approaches[181,182].
The virome is highly sensitive to environmental changes. Monitoring viral diversity during treatment (e.g., following a course of biologics in IBD) can serve as a real-time indicator of mucosal healing or treatment failure[108]. Baseline virome composition may eventually predict which children will respond to specific therapies, such as dietary interventions or fecal microbiota transplantation (FMT), sparing others the risks of trial-and-error medicine[86]. Baseline virome composition may eventually predict which children will respond to specific therapies, such as dietary interventions or FMT, sparing others the risks of trial-and-error medicine. Intellectual honesty requires acknowledging the hurdles[183]. These include the potential for rapid bacterial resistance, the risk of phages carrying ARGs via transduction, and the possibility that the host immune system neutralizes the phages before they reach their targets[184].
We are moving toward an era of “ecological engineering”, where the virome is no longer just observed but actively modulated to restore health. Unlike broad-spectrum antibiotics that act like “carpet bombing”, phage therapy acts as a “sniper”. Phages can be engineered or selected to target specific pathobionts-bacteria that are normally harmless but become pathogenic under certain conditions[185]. A primary target is adherent-invasive Escherichia coli (AIEC), which is heavily implicated in paediatric CD. By deploying phages that specifically lyse AIEC, clinicians could reduce inflammation without disrupting the beneficial commensal bacteria[186]. Current probiotics often fail because they are “cleared” by indigenous bacteriophages. “Virome-aware” probiotics involve selecting or engineering bacterial strains that are naturally resistant to the phages present in the patient’s gut. Combining specific prebiotics (to feed the bacteria) with phages (to kill competitors) creates a “fortified” niche that allows beneficial microbes to engraft more successfully in the paediatric gut[182].
FMT has shown success in treating Clostridium difficile, but concerns remain about the transfer of antibiotic resistance genes or the introduction of unintended bacterial pathogens. fecal virome transplantation (FVT) uses a sterile-filtered liquid containing only the viral fraction (mostly phages) and microbial metabolites, with all bacteria removed[187]. This approach minimizes the risk of introducing “bad” bacteria or metabolic disorders into a child’s developing gut. Pre
Indirectly shaping the virome through traditional methods remains a highly accessible clinical strategy. Certain fibers and “good” bacteria can alter the gut’s metabolic environment, which, in turn, influences which phages are induced from their bacterial hosts[188]. High-fat or high-sugar diets have been shown to trigger prophage induction (the “SOS response”)[189]. Clinical dietary interventions could potentially prevent these harmful viral “blooms” and maintain a more stable, diverse virome.
As we stand on the threshold of a new era in paediatric gastroenterology and immunology, it is clear that our un
| Step | Challenge | Impact on pediatric research |
| Sample collection | Low volume (especially from neonates) | Limits the ability to perform multiple enrichment steps |
| Viral enrichment | Distinguishing “free” viruses from prophages | Makes it hard to tell if a virus is active or just dormant in a bacterium |
| Sequencing | High cost of deep metagenomics | Reduces the sample size of studies, often leading to “underpowered” results |
| Bioinformatics | Lack of universal markers | Prevents the easy, high-throughput identification possible with 16S sequencing |
| Research frontier | Current status | Future goal |
| Dark matter | 90% uncharacterized | Comprehensive viral reference catalogs |
| Study design | Cross-sectional “snapshots” | Multi-decade longitudinal tracking. |
| Mechanisms | Purely associative | Proof-of-concept in organoids and GF models |
| Therapeutics | Experimental/compassionate | Phase III clinical trials in paediatric IBD/NEC |
Perhaps the greatest bottleneck in viromics is the sheer volume of unidentified genetic sequences, often referred to as viral dark matter. In typical paediatric stool metagenomes, up to 90% of viral sequences cannot be mapped to any known reference database (the annotation gap)[190]. This means we are effectively “blind” to most of the viral community. Even when sequences are identified, we often lack knowledge of their host range or metabolic function. Future research must prioritize the development of de novo assembly algorithms and high-throughput culturing techniques to “illuminate” this dark matter.
Most current studies are cross-sectional, providing only a “snapshot” of a highly dynamic process. We need large-scale, international birth cohorts that track infants from birth through adolescence. This is the only way to determine if early-life “viral scars” truly lead to adult-onset diseases. Future studies must integrate viromics with transcriptomics, proteomics, and metabolomics to see not just who is there, but what they are doing to the host in real-time.
The lack of standardized protocols makes it difficult to compare results across different laboratories. Differences in stool storage, viral enrichment (e.g., polyethylene glycol precipitation vs ultra-centrifugation), and DNA/RNA extraction (wet-lab variability) can lead to vastly different “versions” of the same virome[191]. We also need “gold standard” pipelines for viral identification and ecological diversity calculations. Creating open-access, shared databases (similar to the human microbiome project) is essential for scientific reproducibility.
The field is currently “association-rich” but “mechanistically-lean”. We must move beyond observing that “virus X is present in disease Y”. The use of human intestinal organoids (“gut-on-a-chip”) and Germ-Free mice colonized with specific viral fractions is crucial to prove that a virus causes a specific immune or metabolic shift[192]. Using bacterial CRISPR spacers to track past viral infections provides a historical record of “predatory encounters”, offering a more mechanistic view of how phages shape the microbiome[193].
Manipulating a child’s virome-whether through FVT or phage therapy-raises unique ethical questions that must be addressed by bioethicists and clinicians alike. Unlike adults, a child’s immune system and brain are in a state of rapid development[194]. We do not yet know the multi-decadal consequences of early-life virome editing. As viromic “fingerprints” are highly unique, protecting the genomic privacy of paediatric participants in large-scale databases is a paramount concern. As these therapies emerge, we must ensure they do not become “boutique” treatments available only to high-income populations, further widening the health gap.
Despite rapid advances in sequencing technologies, paediatric virome research remains in an early exploratory phase. Future studies should prioritize longitudinal birth cohorts with repeated virome sampling to establish temporal causality rather than cross-sectional associations. Integration of viromics with multi-omics platforms-including metagenomics, metabolomics, transcriptomics, and immune profiling-will be essential to unravel functional host-virome interactions.
Experimental models capable of disentangling viral causality from bacterial confounding effects are urgently needed, particularly in diseases such as NEC, IBD, and autoimmune disorders. Additionally, controlled interventional studies evaluating phage-based modulation, virome-preserving antibiotic strategies, and maternal-infant virome transmission represent promising avenues for preventive paediatric medicine.
Finally, standardization of virome sequencing pipelines, reference databases, and analytical frameworks is critical to ensure reproducibility and meaningful inter-study comparisons.
Translation of paediatric virome research into clinical practice faces substantial regulatory and ethical challenges. Unlike conventional probiotics or antibiotics, bacteriophages and virome-based interventions are biologically dynamic entities capable of evolution, HGT, and ecological disruption. This complexity complicates safety assessment, dosing standardization, and long-term risk evaluation-particularly in children.
Regulatory frameworks for phage therapy remain fragmented globally, with limited paediatric-specific guidance. Concerns regarding phage-encoded virulence or antibiotic resistance genes further underscore the need for rigorous genomic screening and post-intervention surveillance. In addition, the absence of standardized reference viromes and normative paediatric datasets limits the clinical interpretability of virome-based diagnostics.
Ethical considerations unique to paediatrics-including informed consent, long-term monitoring, and intervention during critical developmental windows-must be carefully addressed before virome-targeted therapies can be widely implemented.
The exploration of the paediatric gut virome represents one of the most significant shifts in our understanding of early-life development. We are moving away from a binary view of viruses as either “pathogenic” or “absent” toward a sophisticated ecological model in which the virome serves as the primary architect of human health. The evidence synthesized in this review demonstrates that the gut virome is not a passive collection of biological entities. Instead, it is an active biological modulator that exerts “top-down” control over bacterial communities and “bottom-up” influence on the host immune system. From the lytic blooms that drive NEC and IBD to the subtle viral-neural crosstalk that shapes functional gut disorders, the virome is a constant, dynamic force in the intestinal microenvironment.
The first 1000 days of life constitute a “priming window” during which the virome is at its most volatile and influential. The transition from the low-diversity, milk-oriented virome of the neonate to the complex, phage-dominated landscape of the toddler is a high-stakes successional process. Disruptions during this period-whether through iatrogenic insults like antibiotics or environmental factors-can leave “molecular scars” that predispose children to a lifetime of metabolic, autoimmune, and neurodevelopmental challenges. For the clinician, the virome offers a new set of tools for precision medicine. The ability to use viral signatures as pre-symptomatic biomarkers and the potential for “sniper-like” phage therapies represent the future of paediatric care. However, the virome’s reach extends beyond the gut; its influence on the gut-brain, gut-lung, and gut-pancreas axes suggests that viromics will soon become central to neonatology, endocrinology, and developmental neurology. The “bacteriocentic” era of microbiome research must now evolve into an integrated “multi-kingdom” approach. We cannot fully understand the bacteriome without accounting for the predatory pressure of phages, nor can we understand host immunity without acknowledging the constant stimulus of the eukaryotic virome. The paediatric virome is the “silent programmer” of the developing human. As we continue to illuminate the viral dark matter and refine our therapeutic toolkits, our ability to cultivate a healthy viral ecosystem will likely become one of the most powerful interventions in 21st century paediatric medicine.
| 1. | Runge S, Rosshart SP. The Mammalian Metaorganism: A Holistic View on How Microbes of All Kingdoms and Niches Shape Local and Systemic Immunity. Front Immunol. 2021;12:702378. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 24] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 2. | Kinsella CM, Deijs M, Becker C, Broekhuizen P, van Gool T, Bart A, Schaefer AS, van der Hoek L. Host prediction for disease-associated gastrointestinal cressdnaviruses. Virus Evol. 2022;8:veac087. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 16] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 3. | Cesanelli F, Scarvaglieri I, De Francesco MA, Alberti M, Salvi M, Tiecco G, Castelli F, Quiros-roldan E. The Human Virome in Health and Its Remodeling During HIV Infection and Antiretroviral Therapy: A Narrative Review. Microorganisms. 2025;14:50. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 5] [Article Influence: 5.0] [Reference Citation Analysis (1)] |
| 4. | Graham DB, Xavier RJ. Conditioning of the immune system by the microbiome. Trends Immunol. 2023;44:499-511. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 61] [Article Influence: 20.3] [Reference Citation Analysis (1)] |
| 5. | Kennedy EA, Holtz LR. Gut virome in early life: origins and implications. Curr Opin Virol. 2022;55:101233. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 14] [Article Influence: 3.5] [Reference Citation Analysis (1)] |
| 6. | Yi C, Chen J, She X. The emerging role of the gut virome in necrotizing enterocolitis. Heliyon. 2024;10:e30496. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 3] [Article Influence: 1.5] [Reference Citation Analysis (1)] |
| 7. | Kotsiri I, Xanthi M, Domazinaki CM, Magiorkinis E. The Role of Viral Infections in the Immunopathogenesis of Type 1 Diabetes Mellitus: A Narrative Review. Biology (Basel). 2025;14:981. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 8. | Feng Z, Burgermeister E, Philips A, Zuo T, Wen W. The gut virome in association with the bacteriome in gastrointestinal diseases and beyond: roles, mechanisms, and clinical applications. Precis Clin Med. 2025;8:pbaf010. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 12] [Article Influence: 12.0] [Reference Citation Analysis (1)] |
| 9. | Zhang Y, Wang R. The human gut phageome: composition, development, and alterations in disease. Front Microbiol. 2023;14:1213625. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 22] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 10. | Berman JJ. Viruses. Taxonomic Guide to Infectious Diseases. 2nd edition. Academic Press, 2019: 263-319. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 5] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 11. | Louten J. Virus Structure and Classification. Essential Human Virology. Boston: Academic Press, 2016: 19-29. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 55] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 12. | Hillary LS, Knotts TA, Adams SH, Ali MR, Olm MR, Emerson JB. DNA extraction and virome processing methods strongly influence recovered human gut viral community characteristics. bioRxiv. 2025;2025.11.25.690293. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 13. | Yang SY, Han SM, Lee JY, Kim KS, Lee JE, Lee DW. Advancing Gut Microbiome Research: The Shift from Metagenomics to Multi-Omics and Future Perspectives. J Microbiol Biotechnol. 2025;35:e2412001. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 40] [Article Influence: 40.0] [Reference Citation Analysis (0)] |
| 14. | Janda JM, Abbott SL. 16S rRNA gene sequencing for bacterial identification in the diagnostic laboratory: pluses, perils, and pitfalls. J Clin Microbiol. 2007;45:2761-2764. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1634] [Cited by in RCA: 1167] [Article Influence: 61.4] [Reference Citation Analysis (0)] |
| 15. | Zheng N, Yu HL, Zhang BJ, Wang D, Ji YL, Dai LL, Li W, Li SH, Hu ZL, Zheng YS. Metagenomic next-generation sequencing-based characterization of the viral spectrum in clinical pulmonary and peripheral blood samples of patients. Front Cell Infect Microbiol. 2025;15:1562965. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 16. | Wei Y, Zhou C. Bacteriophages: a double-edged sword in the gastrointestinal tract. Front Microbio. 2024;3:1450523. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 5] [Article Influence: 2.5] [Reference Citation Analysis (1)] |
| 17. | Koonin EV, Yutin N. The crAss-like Phage Group: How Metagenomics Reshaped the Human Virome. Trends Microbiol. 2020;28:349-359. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 84] [Cited by in RCA: 61] [Article Influence: 10.2] [Reference Citation Analysis (1)] |
| 18. | Yang Y, Nagata T. Viral Production in Seawater Filtered Through 0.2-μm Pore-Size Filters: A Hidden Biogeochemical Cycle in a Neglected Realm. Front Microbiol. 2021;12:774849. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 2] [Article Influence: 0.4] [Reference Citation Analysis (1)] |
| 19. | Hahn MW. Broad diversity of viable bacteria in ‘sterile’ (0.2 microm) filtered water. Res Microbiol. 2004;155:688-691. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 86] [Cited by in RCA: 55] [Article Influence: 2.5] [Reference Citation Analysis (1)] |
| 20. | Ospino MC, Engel K, Ruiz-Navas S, Binns WJ, Doxey AC, Neufeld JD. Evaluation of multiple displacement amplification for metagenomic analysis of low biomass samples. ISME Commun. 2024;4:ycae024. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 9] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 21. | Kim KH, Chang HW, Nam YD, Roh SW, Kim MS, Sung Y, Jeon CO, Oh HM, Bae JW. Amplification of uncultured single-stranded DNA viruses from rice paddy soil. Appl Environ Microbiol. 2008;74:5975-5985. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 151] [Cited by in RCA: 126] [Article Influence: 7.0] [Reference Citation Analysis (1)] |
| 22. | Cryan JF, O’Riordan KJ, Cowan CSM, Sandhu KV, Bastiaanssen TFS, Boehme M, Codagnone MG, Cussotto S, Fulling C, Golubeva AV, Guzzetta KE, Jaggar M, Long-Smith CM, Lyte JM, Martin JA, Molinero-Perez A, Moloney G, Morelli E, Morillas E, O’Connor R, Cruz-Pereira JS, Peterson VL, Rea K, Ritz NL, Sherwin E, Spichak S, Teichman EM, van de Wouw M, Ventura-Silva AP, Wallace-Fitzsimons SE, Hyland N, Clarke G, Dinan TG. The Microbiota-Gut-Brain Axis. Physiol Rev. 2019;99:1877-2013. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4131] [Cited by in RCA: 3446] [Article Influence: 492.3] [Reference Citation Analysis (4)] |
| 23. | Liang G, Zhao C, Zhang H, Mattei L, Sherrill-Mix S, Bittinger K, Kessler LR, Wu GD, Baldassano RN, DeRusso P, Ford E, Elovitz MA, Kelly MS, Patel MZ, Mazhani T, Gerber JS, Kelly A, Zemel BS, Bushman FD. The stepwise assembly of the neonatal virome is modulated by breastfeeding. Nature. 2020;581:470-474. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 273] [Cited by in RCA: 215] [Article Influence: 35.8] [Reference Citation Analysis (0)] |
| 24. | García-López R, Pérez-Brocal V, Moya A. Beyond cells - The virome in the human holobiont. Microb Cell. 2019;6:373-396. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 19] [Article Influence: 2.7] [Reference Citation Analysis (1)] |
| 25. | Wylie KM, Wylie TN, Cahill AG, Macones GA, Tuuli MG, Stout MJ. The vaginal eukaryotic DNA virome and preterm birth. Am J Obstet Gynecol. 2018;219:189.e1-189.e12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 80] [Cited by in RCA: 67] [Article Influence: 8.4] [Reference Citation Analysis (1)] |
| 26. | Shamash M, Maurice CF. Phages in the infant gut: a framework for virome development during early life. ISME J. 2022;16:323-330. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 69] [Cited by in RCA: 59] [Article Influence: 14.8] [Reference Citation Analysis (2)] |
| 27. | Kim G, Bae J, Kim MJ, Kwon H, Park G, Kim SJ, Choe YH, Kim J, Park SH, Choe BH, Shin H, Kang B. Delayed Establishment of Gut Microbiota in Infants Delivered by Cesarean Section. Front Microbiol. 2020;11:2099. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 80] [Cited by in RCA: 64] [Article Influence: 10.7] [Reference Citation Analysis (1)] |
| 28. | Guo Y, Liu Y, Xu S, Zhang R, Yu Z, He W. Phage diversity in human breast milk: a systematic review. Eur J Pediatr. 2025;184:334. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 4] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 29. | Moubareck CA. Human Milk Microbiota and Oligosaccharides: A Glimpse into Benefits, Diversity, and Correlations. Nutrients. 2021;13:1123. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 128] [Cited by in RCA: 93] [Article Influence: 18.6] [Reference Citation Analysis (11)] |
| 30. | Virgin HW. The virome in mammalian physiology and disease. Cell. 2014;157:142-150. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 530] [Cited by in RCA: 435] [Article Influence: 36.3] [Reference Citation Analysis (1)] |
| 31. | Meaden S, Westra ER, Fineran PC. Phage defence-system abundances vary across environments and increase with viral density. Philos Trans R Soc Lond B Biol Sci. 2025;380:20240069. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 4] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 32. | Abd-Allah IM, El-Housseiny GS, Yahia IS, Aboshanab KM, Hassouna NA. Rekindling of a Masterful Precedent; Bacteriophage: Reappraisal and Future Pursuits. Front Cell Infect Microbiol. 2021;11:635597. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 20] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 33. | Mukhopadhya I, Segal JP, Carding SR, Hart AL, Hold GL. The gut virome: the ‘missing link’ between gut bacteria and host immunity? Therap Adv Gastroenterol. 2019;12:1756284819836620. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 157] [Cited by in RCA: 129] [Article Influence: 18.4] [Reference Citation Analysis (5)] |
| 34. | Yao Y, Cai X, Ye Y, Wang F, Chen F, Zheng C. The Role of Microbiota in Infant Health: From Early Life to Adulthood. Front Immunol. 2021;12:708472. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 216] [Cited by in RCA: 171] [Article Influence: 34.2] [Reference Citation Analysis (1)] |
| 35. | Bushman F, Liang G. Assembly of the virome in newborn human infants. Curr Opin Virol. 2021;48:17-22. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 23] [Article Influence: 4.6] [Reference Citation Analysis (1)] |
| 36. | Breitbart M, Haynes M, Kelley S, Angly F, Edwards RA, Felts B, Mahaffy JM, Mueller J, Nulton J, Rayhawk S, Rodriguez-Brito B, Salamon P, Rohwer F. Viral diversity and dynamics in an infant gut. Res Microbiol. 2008;159:367-373. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 296] [Cited by in RCA: 253] [Article Influence: 14.1] [Reference Citation Analysis (1)] |
| 37. | Fulci V, Stronati L, Cucchiara S, Laudadio I, Carissimi C. Emerging Roles of Gut Virome in Pediatric Diseases. Int J Mol Sci. 2021;22:4127. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 29] [Article Influence: 5.8] [Reference Citation Analysis (1)] |
| 38. | Lim ES, Zhou Y, Zhao G, Bauer IK, Droit L, Ndao IM, Warner BB, Tarr PI, Wang D, Holtz LR. Early life dynamics of the human gut virome and bacterial microbiome in infants. Nat Med. 2015;21:1228-1234. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 582] [Cited by in RCA: 495] [Article Influence: 45.0] [Reference Citation Analysis (1)] |
| 39. | Ronan V, Yeasin R, Claud EC. Childhood Development and the Microbiome-The Intestinal Microbiota in Maintenance of Health and Development of Disease During Childhood Development. Gastroenterology. 2021;160:495-506. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 222] [Cited by in RCA: 184] [Article Influence: 36.8] [Reference Citation Analysis (1)] |
| 40. | Ogilvie LA, Jones BV. The human gut virome: a multifaceted majority. Front Microbiol. 2015;6:918. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 177] [Cited by in RCA: 144] [Article Influence: 13.1] [Reference Citation Analysis (0)] |
| 41. | Barthes K, Rousset F, Wein T. Viral effectors trigger innate immunity across the tree of life. Philos Trans R Soc Lond B Biol Sci. 2025;380:20240077. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 1] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 42. | Fernández-Pato A, Sinha T, Garmaeva S, Gulyaeva A, Kuzub N, Roux S, Fu J, Kurilshikov A, Zhernakova A. Early-life development of the gut virome and plasmidome: A longitudinal study in cesarean-born infants. Cell Rep. 2025;44:115731. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 8] [Article Influence: 8.0] [Reference Citation Analysis (1)] |
| 43. | Mpakosi A, Sokou R, Theodoraki M, Iacovidou N, Cholevas V, Tsantes AG, Liakou AI, Drogari-Apiranthitou M, Kaliouli-Antonopoulou C. The Role of Infant and Early Childhood Gut Virome in Immunity and the Triggering of Autoimmunity-A Narrative Review. Diagnostics (Basel). 2025;15:413. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 5] [Article Influence: 5.0] [Reference Citation Analysis (1)] |
| 44. | Kurilovich E, Geva-Zatorsky N. Effects of bacteriophages on gut microbiome functionality. Gut Microbes. 2025;17:2481178. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 24] [Article Influence: 24.0] [Reference Citation Analysis (0)] |
| 45. | Singha B, Rawat BS, Venkataraman R, Nair T, Rosenn EH, Soni V. Gut microbiome associated dysbiosis: Limited regimens and expanding horizons of phage therapy. Asp Mol Med. 2023;2:100029. [DOI] [Full Text] |
| 46. | Gulyaeva A, Garmaeva S, Kurilshikov A, Vich Vila A, Riksen NP, Netea MG, Weersma RK, Fu J, Zhernakova A. Diversity and Ecology of Caudoviricetes Phages with Genome Terminal Repeats in Fecal Metagenomes from Four Dutch Cohorts. Viruses. 2022;14:2305. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 15] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 47. | Yang P, Zhang H, Yin L, Chen J, Chen Y, Yang H, Liu Q, Zhang W. Genetic diversity of Microviridae phages in the human respiratory tract. Front Cell Infect Microbiol. 2025;15:1629120. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 48. | Webb B, Rakibuzzaman A, Ramamoorthy S. Torque teno viruses in health and disease. Virus Res. 2020;285:198013. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 57] [Article Influence: 9.5] [Reference Citation Analysis (1)] |
| 49. | Freer G, Maggi F, Pifferi M, Di Cicco ME, Peroni DG, Pistello M. The Virome and Its Major Component, Anellovirus, a Convoluted System Molding Human Immune Defenses and Possibly Affecting the Development of Asthma and Respiratory Diseases in Childhood. Front Microbiol. 2018;9:686. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 98] [Cited by in RCA: 81] [Article Influence: 10.1] [Reference Citation Analysis (1)] |
| 50. | Dong Y, Fan S, Zhu L, Sharshov K, Wang W. Viromic Insights into Gut RNA Virus Diversity Among Three Corvid Species. Viruses. 2025;17:1508. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 51. | Makimaa H, Ingle H, Baldridge MT. Enteric Viral Co-Infections: Pathogenesis and Perspective. Viruses. 2020;12:904. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 53] [Cited by in RCA: 51] [Article Influence: 8.5] [Reference Citation Analysis (1)] |
| 52. | Alotaibi M, Al-Khalaifah H, Bouhoudan A. Global Research Trends on Major Pathogenic Enteric Viruses (1990-2024): A Bibliometric Analysis of Epidemiology, Transmission, and Public Health Impact. Pathogens. 2025;14:938. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 2] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 53. | Baquero F, Martínez JL, F Lanza V, Rodríguez-Beltrán J, Galán JC, San Millán A, Cantón R, Coque TM. Evolutionary Pathways and Trajectories in Antibiotic Resistance. Clin Microbiol Rev. 2021;34:e0005019. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 259] [Cited by in RCA: 175] [Article Influence: 35.0] [Reference Citation Analysis (1)] |
| 54. | Sutton TDS, Hill C. Gut Bacteriophage: Current Understanding and Challenges. Front Endocrinol (Lausanne). 2019;10:784. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 169] [Cited by in RCA: 134] [Article Influence: 19.1] [Reference Citation Analysis (1)] |
| 55. | Thingstad TF. Elements of a theory for the mechanisms controlling abundance, diversity, and biogeochemical role of lytic bacterial viruses in aquatic systems. Limnol Oceanogr. 2000;45:1320-1328. [DOI] [Full Text] |
| 56. | Roughgarden J. Lytic/Lysogenic Transition as a Life-History Switch. Virus Evol. 2024;10:veae028. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 18] [Article Influence: 9.0] [Reference Citation Analysis (1)] |
| 57. | Brüssow H, Canchaya C, Hardt WD. Phages and the evolution of bacterial pathogens: from genomic rearrangements to lysogenic conversion. Microbiol Mol Biol Rev. 2004;68:560-602, table of contents. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1395] [Cited by in RCA: 1161] [Article Influence: 52.8] [Reference Citation Analysis (1)] |
| 58. | Lathakumari RH, Vajravelu LK, Gopinathan A, Vimala PB, Panneerselvam V, Ravi SSS, Thulukanam J. The gut virome and human health: From diversity to personalized medicine. Eng Microbiol. 2025;5:100191. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 11] [Article Influence: 11.0] [Reference Citation Analysis (1)] |
| 59. | Dapa T, Ramiro RS, Pedro MF, Gordo I, Xavier KB. Diet leaves a genetic signature in a keystone member of the gut microbiota. Cell Host Microbe. 2022;30:183-199.e10. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 91] [Cited by in RCA: 73] [Article Influence: 18.3] [Reference Citation Analysis (1)] |
| 60. | Castledine M, Buckling A. Critically evaluating the relative importance of phage in shaping microbial community composition. Trends Microbiol. 2024;32:957-969. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 41] [Article Influence: 20.5] [Reference Citation Analysis (1)] |
| 61. | Zhang S, Easwaran M, Elafify M, Mahmoud AA, Wang X, Ahn J. Prophages and their interactions with lytic phages in the human gut microbiota and their impact on microbial diversity, gut health, and disease. Appl Environ Microbiol. 2025;91:e0189925. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 7] [Article Influence: 7.0] [Reference Citation Analysis (1)] |
| 62. | Logan AC, Katzman MA, Balanzá-Martínez V. Natural environments, ancestral diets, and microbial ecology: is there a modern “paleo-deficit disorder”? Part I. J Physiol Anthropol. 2015;34:1. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 20] [Article Influence: 1.8] [Reference Citation Analysis (1)] |
| 63. | Carroll-Portillo A, Lin HC. Bacteriophage and the Innate Immune System: Access and Signaling. Microorganisms. 2019;7:625. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 69] [Cited by in RCA: 62] [Article Influence: 8.9] [Reference Citation Analysis (1)] |
| 64. | Podlacha M, Gaffke L, Grabowski Ł, Mantej J, Grabski M, Pierzchalska M, Pierzynowska K, Węgrzyn G, Węgrzyn A. Bacteriophage DNA induces an interrupted immune response during phage therapy in a chicken model. Nat Commun. 2024;15:2274. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 42] [Cited by in RCA: 37] [Article Influence: 18.5] [Reference Citation Analysis (1)] |
| 65. | Dogra SK, Kwong Chung C, Wang D, Sakwinska O, Colombo Mottaz S, Sprenger N. Nurturing the Early Life Gut Microbiome and Immune Maturation for Long Term Health. Microorganisms. 2021;9:2110. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 83] [Cited by in RCA: 64] [Article Influence: 12.8] [Reference Citation Analysis (1)] |
| 66. | Joiner KL, Baljon A, Barr J, Rohwer F, Luque A. Impact of bacteria motility in the encounter rates with bacteriophage in mucus. Sci Rep. 2019;9:16427. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 41] [Cited by in RCA: 30] [Article Influence: 4.3] [Reference Citation Analysis (1)] |
| 67. | Martín Giménez VM, Modrego J, Gómez-Garre D, Manucha W, de Las Heras N. Gut Microbiota Dysbiosis in COVID-19: Modulation and Approaches for Prevention and Therapy. Int J Mol Sci. 2023;24:12249. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 28] [Cited by in RCA: 22] [Article Influence: 7.3] [Reference Citation Analysis (1)] |
| 68. | Koonin EV, Dolja VV, Krupovic M. The healthy human virome: from virus-host symbiosis to disease. Curr Opin Virol. 2021;47:86-94. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 62] [Article Influence: 12.4] [Reference Citation Analysis (1)] |
| 69. | Khokhar RK, Nashwan AJ. Gut virome and its emerging role in inflammatory bowel disease. World J Methodol. 2025;15:100534. [PubMed] [DOI] [Full Text] |
| 70. | Dodi G, Attanasi M, Di Filippo P, Di Pillo S, Chiarelli F. Virome in the Lungs: The Role of Anelloviruses in Childhood Respiratory Diseases. Microorganisms. 2021;9:1357. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 29] [Article Influence: 5.8] [Reference Citation Analysis (1)] |
| 71. | Taylor LJ, Keeler EL, Bushman FD, Collman RG. The enigmatic roles of Anelloviridae and Redondoviridae in humans. Curr Opin Virol. 2022;55:101248. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 42] [Article Influence: 10.5] [Reference Citation Analysis (1)] |
| 72. | Okumura R, Takeda K. The role of the mucosal barrier system in maintaining gut symbiosis to prevent intestinal inflammation. Semin Immunopathol. 2024;47:2. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 40] [Article Influence: 20.0] [Reference Citation Analysis (0)] |
| 73. | Douadi C, Lamy-Besnier Q, Theodorou I, Schiettekatte O, Sbardella Y, Brot L, Costantini PE, Saporetti R, Danielli A, Calvaresi M, De Paepe M, Sokol H, Garcia-Weber D, Carrière V, Thenet S, De Sordi L. Differential translocation of bacteriophages across the intestinal barrier in health and Crohn’s disease. Cell Rep. 2025;45:116726. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 3] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 74. | Thompson MR, Kaminski JJ, Kurt-Jones EA, Fitzgerald KA. Pattern recognition receptors and the innate immune response to viral infection. Viruses. 2011;3:920-940. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 719] [Cited by in RCA: 649] [Article Influence: 43.3] [Reference Citation Analysis (5)] |
| 75. | Górski A, Jończyk-Matysiak E, Łusiak-Szelachowska M, Międzybrodzki R, Weber-Dąbrowska B, Borysowski J. Bacteriophages targeting intestinal epithelial cells: a potential novel form of immunotherapy. Cell Mol Life Sci. 2018;75:589-595. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 30] [Cited by in RCA: 26] [Article Influence: 3.3] [Reference Citation Analysis (1)] |
| 76. | Gavkare AM, Nanaware NL, Sonar MN, Dhotre SV, Mumbre SS, Nagoba BS. Gut microbiome and viral infections: A hidden nexus for immune protection. World J Virol. 2025;14:111912. [PubMed] [DOI] [Full Text] |
| 77. | Managlia E, Yan X, De Plaen IG. Intestinal Epithelial Barrier Function and Necrotizing Enterocolitis. Newborn (Clarksville). 2022;1:32-43. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 18] [Article Influence: 4.5] [Reference Citation Analysis (1)] |
| 78. | Tiamani K, Luo S, Schulz S, Xue J, Costa R, Khan Mirzaei M, Deng L. The role of virome in the gastrointestinal tract and beyond. FEMS Microbiol Rev. 2022;46:fuac027. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 49] [Cited by in RCA: 41] [Article Influence: 10.3] [Reference Citation Analysis (0)] |
| 79. | Cao Z, Sugimura N, Burgermeister E, Ebert MP, Zuo T, Lan P. The gut virome: A new microbiome component in health and disease. EBioMedicine. 2022;81:104113. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 245] [Cited by in RCA: 203] [Article Influence: 50.8] [Reference Citation Analysis (0)] |
| 80. | Lange O, Proczko-Stepaniak M, Mika A. Short-Chain Fatty Acids-A Product of the Microbiome and Its Participation in Two-Way Communication on the Microbiome-Host Mammal Line. Curr Obes Rep. 2023;12:108-126. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 110] [Cited by in RCA: 93] [Article Influence: 31.0] [Reference Citation Analysis (5)] |
| 81. | Wang X, Wang S, Huang M, He Y, Guo S, Yang K, Wang N, Sun T, Yang H, Yang T, Xu Y, Shen Q, Friman V-P, Wei Z. Phages enhance both phytopathogen density control and rhizosphere microbiome suppressiveness. mBio. 2024;15:e0301623. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 40] [Cited by in RCA: 22] [Article Influence: 11.0] [Reference Citation Analysis (1)] |
| 82. | Duranti S, Lugli GA, Mancabelli L, Armanini F, Turroni F, James K, Ferretti P, Gorfer V, Ferrario C, Milani C, Mangifesta M, Anzalone R, Zolfo M, Viappiani A, Pasolli E, Bariletti I, Canto R, Clementi R, Cologna M, Crifò T, Cusumano G, Fedi S, Gottardi S, Innamorati C, Masè C, Postai D, Savoi D, Soffiati M, Tateo S, Pedrotti A, Segata N, van Sinderen D, Ventura M. Maternal inheritance of bifidobacterial communities and bifidophages in infants through vertical transmission. Microbiome. 2017;5:66. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 290] [Cited by in RCA: 256] [Article Influence: 28.4] [Reference Citation Analysis (0)] |
| 83. | Azam AH, Tan XE, Veeranarayanan S, Kiga K, Cui L. Bacteriophage Technology and Modern Medicine. Antibiotics (Basel). 2021;10:999. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 39] [Cited by in RCA: 35] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 84. | Ding M, Ross RP, Dempsey E, Li B, Stanton C. Infant gut microbiome reprogramming following introduction of solid foods (weaning). Gut Microbes. 2025;17:2571428. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 8] [Article Influence: 8.0] [Reference Citation Analysis (3)] |
| 85. | Zhong P, Li Q, Zhang Y, Guo C, Abdelsattar MM, Bi Y. Precision microbial regulation: Strategies for modulating GIT microbiota for host health. iMetaOmics. 2025;2:e54. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 5] [Article Influence: 5.0] [Reference Citation Analysis (1)] |
| 86. | Liang G, Gao H, Bushman FD. The pediatric virome in health and disease. Cell Host Microbe. 2022;30:639-649. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 25] [Article Influence: 6.3] [Reference Citation Analysis (1)] |
| 87. | Crane JK, Alvarado CL, Sutton MD. Role of the SOS Response in the Generation of Antibiotic Resistance In Vivo. Antimicrob Agents Chemother. 2021;65:e0001321. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 66] [Cited by in RCA: 57] [Article Influence: 11.4] [Reference Citation Analysis (1)] |
| 88. | Torres-Barceló C. The disparate effects of bacteriophages on antibiotic-resistant bacteria. Emerg Microbes Infect. 2018;7:168. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 132] [Cited by in RCA: 127] [Article Influence: 15.9] [Reference Citation Analysis (1)] |
| 89. | Dong TS, Gupta A. Influence of Early Life, Diet, and the Environment on the Microbiome. Clin Gastroenterol Hepatol. 2019;17:231-242. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 185] [Cited by in RCA: 147] [Article Influence: 21.0] [Reference Citation Analysis (5)] |
| 90. | Odiase E, Frank DN, Young BE, Robertson CE, Kofonow JM, Davis KN, Berman LM, Krebs NF, Tang M. The Gut Microbiota Differ in Exclusively Breastfed and Formula-Fed United States Infants and are Associated with Growth Status. J Nutr. 2023;153:2612-2621. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 38] [Article Influence: 12.7] [Reference Citation Analysis (1)] |
| 91. | Kim AH, Armah G, Dennis F, Wang L, Rodgers R, Droit L, Baldridge MT, Handley SA, Harris VC. Enteric virome negatively affects seroconversion following oral rotavirus vaccination in a longitudinally sampled cohort of Ghanaian infants. Cell Host Microbe. 2022;30:110-123.e5. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 50] [Cited by in RCA: 48] [Article Influence: 12.0] [Reference Citation Analysis (1)] |
| 92. | Cianci R, Caldarelli M, Brani P, Bosi A, Ponti A, Giaroni C, Baj A. Role of the Virome in Vaccine-Induced Immunization. Vaccines (Basel). 2025;13:895. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 93. | Stefanetti G, Kasper DL. Impact of the Host Microbiome on Vaccine Responsiveness: Lessons Learned and Future Perspective. Biochemistry. 2022;61:2849-2855. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 8] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 94. | D’Agata AL, Wu J, Welandawe MKV, Dutra SVO, Kane B, Groer MW. Effects of early life NICU stress on the developing gut microbiome. Dev Psychobiol. 2019;61:650-660. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 32] [Article Influence: 4.6] [Reference Citation Analysis (1)] |
| 95. | Liddicoat C, Waycott M, Weinstein P. Environmental Change and Human Health: Can Environmental Proxies Inform the Biodiversity Hypothesis for Protective Microbial-Human Contact? BioScience. 2016;66:1023-1034. [DOI] [Full Text] |
| 96. | Orenstein R. Gastroenteritis, Viral. In: Kuipers FJ, editor. Encyclopedia of Gastroenterology. 2nd edition. Oxford: Academic Press, 2020: 652-657. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 6] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 97. | Wang R, Lan C, Benlagha K, Camara NOS, Miller H, Kubo M, Heegaard S, Lee P, Yang L, Forsman H, Li X, Zhai Z, Liu C. The interaction of innate immune and adaptive immune system. MedComm (2020). 2024;5:e714. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 184] [Cited by in RCA: 161] [Article Influence: 80.5] [Reference Citation Analysis (1)] |
| 98. | Thabane M, Marshall JK. Post-infectious irritable bowel syndrome. World J Gastroenterol. 2009;15:3591-3596. [PubMed] [DOI] [Full Text] |
| 99. | Zuniga EI, Macal M, Lewis GM, Harker JA. Innate and Adaptive Immune Regulation During Chronic Viral Infections. Annu Rev Virol. 2015;2:573-597. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 140] [Cited by in RCA: 133] [Article Influence: 12.1] [Reference Citation Analysis (1)] |
| 100. | Pensabene L, Talarico V, Concolino D, Ciliberto D, Campanozzi A, Gentile T, Rutigliano V, Salvatore S, Staiano A, Di Lorenzo C; Post-Infectious Functional Gastrointestinal Disorders Study Group of Italian Society for Pediatric Gastroenterology, Hepatology and Nutrition. Postinfectious functional gastrointestinal disorders in children: a multicenter prospective study. J Pediatr. 2015;166:903-7.e1. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 40] [Article Influence: 3.6] [Reference Citation Analysis (1)] |
| 101. | Gyriki D, Nikolaidis C, Stavropoulou E, Bezirtzoglou I, Tsigalou C, Vradelis S, Bezirtzoglou E. Exploring the Gut Microbiome’s Role in Inflammatory Bowel Disease: Insights and Interventions. J Pers Med. 2024;14:507. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 29] [Cited by in RCA: 22] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 102. | Hetta HF, Ahmed R, Ramadan YN, Fathy H, Khorshid M, Mabrouk MM, Hashem M. Gut virome: New key players in the pathogenesis of inflammatory bowel disease. World J Methodol. 2025;15:92592. [PubMed] [DOI] [Full Text] |
| 103. | Zhao D, Zou B, Do QL, Wu SK, Shen Y, Yang Y, Kang JX, Su KP, Wang B. Circadian rhythms and gut microbiota Dysbiosis: emerging gut-brain axis pathways in insomnia pathophysiology and Therapeutics. Brain Behav Immun. 2026;132:106203. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 2] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 104. | Figliuolo da Paz VR, Jamwal DR, Kiela PR. Intestinal Regulatory T Cells. Adv Exp Med Biol. 2021;1278:141-190. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 14] [Article Influence: 2.8] [Reference Citation Analysis (1)] |
| 105. | Gogokhia L, Buhrke K, Bell R, Hoffman B, Brown DG, Hanke-Gogokhia C, Ajami NJ, Wong MC, Ghazaryan A, Valentine JF, Porter N, Martens E, O’Connell R, Jacob V, Scherl E, Crawford C, Stephens WZ, Casjens SR, Longman RS, Round JL. Expansion of Bacteriophages Is Linked to Aggravated Intestinal Inflammation and Colitis. Cell Host Microbe. 2019;25:285-299.e8. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 538] [Cited by in RCA: 462] [Article Influence: 66.0] [Reference Citation Analysis (7)] |
| 106. | Ullah K, Kanwar R, Ali S, Uddin S, Izza I, Ahmad I, Alzahrani KJ, Qadeer A, Chu CT, Chen CC. Gut Virome and Aging: Phage-Driven Microbial Stability and Immune Modulation. Aging Dis. 2025. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 5] [Article Influence: 5.0] [Reference Citation Analysis (1)] |
| 107. | Qv L, Mao S, Li Y, Zhang J, Li L. Roles of Gut Bacteriophages in the Pathogenesis and Treatment of Inflammatory Bowel Disease. Front Cell Infect Microbiol. 2021;11:755650. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 34] [Cited by in RCA: 25] [Article Influence: 5.0] [Reference Citation Analysis (1)] |
| 108. | Wu Y, Cheng R, Lin H, Li L, Jia Y, Philips A, Zuo T, Zhang H. Gut virome and its implications in the pathogenesis and therapeutics of inflammatory bowel disease. BMC Med. 2025;23:183. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 13] [Article Influence: 13.0] [Reference Citation Analysis (1)] |
| 109. | Ullah H, Arbab S, Tian Y, Chen Y, Liu CQ, Li Q, Li K. Crosstalk between gut microbiota and host immune system and its response to traumatic injury. Front Immunol. 2024;15:1413485. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 60] [Cited by in RCA: 48] [Article Influence: 24.0] [Reference Citation Analysis (0)] |
| 110. | Tun HM, Peng Y, Massimino L, Sin ZY, Parigi TL, Facoetti A, Rahman S, Danese S, Ungaro F. Gut virome in inflammatory bowel disease and beyond. Gut. 2024;73:350-360. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 65] [Cited by in RCA: 56] [Article Influence: 28.0] [Reference Citation Analysis (0)] |
| 111. | Borowitz SM. The epidemiology of inflammatory bowel disease: Clues to pathogenesis? Front Pediatr. 2022;10:1103713. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 86] [Cited by in RCA: 68] [Article Influence: 22.7] [Reference Citation Analysis (1)] |
| 112. | Lin PW, Nasr TR, Stoll BJ. Necrotizing enterocolitis: recent scientific advances in pathophysiology and prevention. Semin Perinatol. 2008;32:70-82. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 239] [Cited by in RCA: 211] [Article Influence: 11.7] [Reference Citation Analysis (4)] |
| 113. | Shah SA, Deng L, Thorsen J, Pedersen AG, Dion MB, Castro-Mejía JL, Silins R, Romme FO, Sausset R, Jessen LE, Ndela EO, Hjelmsø M, Rasmussen MA, Redgwell TA, Leal Rodríguez C, Vestergaard G, Zhang Y, Chawes B, Bønnelykke K, Sørensen SJ, Bisgaard H, Enault F, Stokholm J, Moineau S, Petit MA, Nielsen DS. Expanding known viral diversity in the healthy infant gut. Nat Microbiol. 2023;8:986-998. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 137] [Cited by in RCA: 96] [Article Influence: 32.0] [Reference Citation Analysis (1)] |
| 114. | Erickson I, Tung J, Schwartz DJ. Consequences of host-microbiome interactions in preterm infants. Infect Immun. 2025;93:e0050124. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 7] [Article Influence: 7.0] [Reference Citation Analysis (1)] |
| 115. | Kaelin EA, Rodriguez C, Hall-Moore C, Hoffmann JA, Linneman LA, Ndao IM, Warner BB, Tarr PI, Holtz LR, Lim ES. Longitudinal gut virome analysis identifies specific viral signatures that precede necrotizing enterocolitis onset in preterm infants. Nat Microbiol. 2022;7:653-662. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 79] [Cited by in RCA: 57] [Article Influence: 14.3] [Reference Citation Analysis (1)] |
| 116. | Land WG. The Role of Damage-Associated Molecular Patterns in Human Diseases: Part I - Promoting inflammation and immunity. Sultan Qaboos Univ Med J. 2015;15:e9-e21. [PubMed] |
| 117. | Scheese DJ, Sodhi CP, Hackam DJ. New insights into the pathogenesis of necrotizing enterocolitis and the dawn of potential therapeutics. Semin Pediatr Surg. 2023;32:151309. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 22] [Cited by in RCA: 21] [Article Influence: 7.0] [Reference Citation Analysis (1)] |
| 118. | Sajankila N, Wala SJ, Ragan MV, Volpe SG, Dumbauld Z, Purayil N, Mihi B, Besner GE. Current and future methods of probiotic therapy for necrotizing enterocolitis. Front Pediatr. 2023;11:1120459. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 13] [Article Influence: 4.3] [Reference Citation Analysis (1)] |
| 119. | Mani S, Hazra S, Hagan J, Sisson A, Nair J, Pammi M. Viral Infections and Neonatal Necrotizing Enterocolitis: A Meta-analysis. Pediatrics. 2023;152:e2022060876. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 23] [Article Influence: 7.7] [Reference Citation Analysis (1)] |
| 120. | Gujral N, Freeman HJ, Thomson AB. Celiac disease: prevalence, diagnosis, pathogenesis and treatment. World J Gastroenterol. 2012;18:6036-6059. [PubMed] [DOI] [Full Text] |
| 121. | Llorente Pelayo S, Palacios Sánchez M, Docio Pérez P, Gutiérrez Buendía D, Peña Sainz-Pardo E, Vega Santa-Cruz B, García Calatayud S. [Infections in early life as risk factor for coeliac disease]. An Pediatr (Engl Ed). 2021;94:293-300. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 3] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 122. | Bouziat R, Hinterleitner R, Brown JJ, Stencel-Baerenwald JE, Ikizler M, Mayassi T, Meisel M, Kim SM, Discepolo V, Pruijssers AJ, Ernest JD, Iskarpatyoti JA, Costes LM, Lawrence I, Palanski BA, Varma M, Zurenski MA, Khomandiak S, McAllister N, Aravamudhan P, Boehme KW, Hu F, Samsom JN, Reinecker HC, Kupfer SS, Guandalini S, Semrad CE, Abadie V, Khosla C, Barreiro LB, Xavier RJ, Ng A, Dermody TS, Jabri B. Reovirus infection triggers inflammatory responses to dietary antigens and development of celiac disease. Science. 2017;356:44-50. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 427] [Cited by in RCA: 356] [Article Influence: 39.6] [Reference Citation Analysis (5)] |
| 123. | Brown JJ, Jabri B, Dermody TS. A viral trigger for celiac disease. PLoS Pathog. 2018;14:e1007181. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 42] [Cited by in RCA: 35] [Article Influence: 4.4] [Reference Citation Analysis (1)] |
| 124. | Cecilio LA, Bonatto MW. The prevalence of HLA DQ2 and DQ8 in patients with celiac disease, in family and in general population. Arq Bras Cir Dig. 2015;28:183-185. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 31] [Article Influence: 2.8] [Reference Citation Analysis (1)] |
| 125. | Cardoso-Silva D, Delbue D, Itzlinger A, Moerkens R, Withoff S, Branchi F, Schumann M. Intestinal Barrier Function in Gluten-Related Disorders. Nutrients. 2019;11:2325. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 110] [Cited by in RCA: 90] [Article Influence: 12.9] [Reference Citation Analysis (1)] |
| 126. | Oikarinen M, Puustinen L, Lehtonen J, Hakola L, Simell S, Toppari J, Ilonen J, Veijola R, Virtanen SM, Knip M, Hyöty H. Enterovirus Infections Are Associated With the Development of Celiac Disease in a Birth Cohort Study. Front Immunol. 2020;11:604529. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 34] [Cited by in RCA: 30] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 127. | Valitutti F, Mennini M, Monacelli G, Fagiolari G, Piccirillo M, Di Nardo G, Di Cara G. Intestinal permeability, food antigens and the microbiome: a multifaceted perspective. Front Allergy. 2024;5:1505834. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 9] [Article Influence: 9.0] [Reference Citation Analysis (1)] |
| 128. | Cohen R, Mahlab-Guri K, Atali M, Elbirt D. Viruses and celiac disease: what do we know ? Clin Exp Med. 2023;23:2931-2939. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 12] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 129. | Brodin P. Immune-microbe interactions early in life: A determinant of health and disease long term. Science. 2022;376:945-950. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 142] [Cited by in RCA: 129] [Article Influence: 32.3] [Reference Citation Analysis (1)] |
| 130. | Limbri LF, Wilson TG, Oliver MR. Prevalence of irritable bowel syndrome and functional abdominal pain disorders in children with inflammatory bowel disease in remission. JGH Open. 2022;6:818-823. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 7] [Article Influence: 1.8] [Reference Citation Analysis (1)] |
| 131. | Mahesan A, Kamila G, Gulati S. Pediatric Functional Gastrointestinal Disorders: Pathophysiology, Diagnosis and Management. J Transl Gastroenterol. 2024;2:101-108. [RCA] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 3] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 132. | Bendiks M, Kopp MV. The relationship between advances in understanding the microbiome and the maturing hygiene hypothesis. Curr Allergy Asthma Rep. 2013;13:487-494. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 60] [Cited by in RCA: 53] [Article Influence: 4.1] [Reference Citation Analysis (1)] |
| 133. | Planz O, Kircheis R. Editorial: The role of toll-like receptors and their related signaling pathways in viral infection and inflammation. Front Immunol. 2024;15:1363958. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.5] [Reference Citation Analysis (1)] |
| 134. | Bouin M, Lupien F, Riberdy M, Boivin M, Plourde V, Poitras P. Intolerance to visceral distension in functional dyspepsia or irritable bowel syndrome: an organ specific defect or a pan intestinal dysregulation? Neurogastroenterol Motil. 2004;16:311-314. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 64] [Cited by in RCA: 58] [Article Influence: 2.6] [Reference Citation Analysis (5)] |
| 135. | Šojat D, Volarić M, Keškić T, Volarić N, Cerovečki V, Trtica Majnarić L. Putting Functional Gastrointestinal Disorders within the Spectrum of Inflammatory Disorders Can Improve Classification and Diagnostics of These Disorders. Biomedicines. 2024;12:702. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 6] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 136. | Li XL, Megdadi M, Quadri HS. Interaction between gut virome and microbiota on inflammatory bowel disease. World J Methodol. 2025;15:100332. [PubMed] [DOI] [Full Text] |
| 137. | Mhanna A, Martini N, Hmaydoosh G, Hamwi G, Jarjanazi M, Zaifah G, Kazzazo R, Haji Mohamad A, Alshehabi Z. The correlation between gut microbiota and both neurotransmitters and mental disorders: A narrative review. Medicine (Baltimore). 2024;103:e37114. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 104] [Cited by in RCA: 79] [Article Influence: 39.5] [Reference Citation Analysis (1)] |
| 138. | Sasso JM, Ammar RM, Tenchov R, Lemmel S, Kelber O, Grieswelle M, Zhou QA. Gut Microbiome-Brain Alliance: A Landscape View into Mental and Gastrointestinal Health and Disorders. ACS Chem Neurosci. 2023;14:1717-1763. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 171] [Cited by in RCA: 117] [Article Influence: 39.0] [Reference Citation Analysis (1)] |
| 139. | Mina S, Daher S, Jamal Elddin S, Mina N, Nasreddine W. Post-infectious irritable bowel syndrome in a paediatric population: first data in a Middle Eastern country. BMC Pediatr. 2025;25:531. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 140. | Delanote J, Correa Rojo A, Wells PM, Steves CJ, Ertaylan G. Systematic identification of the role of gut microbiota in mental disorders: a TwinsUK cohort study. Sci Rep. 2024;14:3626. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 6] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 141. | Vincent W. The Role of Gut Virome in the Overall Population Health. Sci Insights. 2025;47:1967-1977. [DOI] [Full Text] |
| 142. | Kirk D, Costeira R, Visconti A, Khan Mirzaei M, Deng L, Valdes AM, Menni C. Bacteriophages, gut bacteria, and microbial pathways interplay in cardiometabolic health. Cell Rep. 2024;43:113728. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 12] [Article Influence: 6.0] [Reference Citation Analysis (1)] |
| 143. | Shrivastav K, Nasser H, Ikeda T, Nema V. Possible Crosstalk and Alterations in Gut Bacteriome and Virome in HIV-1 Infection and the Associated Comorbidities Related to Metabolic Disorder. Viruses. 2025;17:990. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 4] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 144. | Thurber RV, Payet JP, Thurber AR, Correa AM. Virus-host interactions and their roles in coral reef health and disease. Nat Rev Microbiol. 2017;15:205-216. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 160] [Cited by in RCA: 96] [Article Influence: 10.7] [Reference Citation Analysis (1)] |
| 145. | Chica Cardenas LA, Leonard MM, Baldridge MT, Handley SA. Gut virome dynamics: from commensal to critical player in health and disease. Nat Rev Gastroenterol Hepatol. 2026;23:126-144. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 8] [Article Influence: 8.0] [Reference Citation Analysis (1)] |
| 146. | Liu L, Li M, Qin Y, Liu Y, Li M, Lian B, Guo R, Xiao Y, Yin C. Childhood obesity and insulin resistance is correlated with gut microbiome serum protein: an integrated metagenomic and proteomic analysis. Sci Rep. 2025;15:21436. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 147. | Chen Q, Ma X, Li C, Shen Y, Zhu W, Zhang Y, Guo X, Zhou J, Liu C. Enteric Phageome Alterations in Patients With Type 2 Diabetes. Front Cell Infect Microbiol. 2020;10:575084. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 39] [Cited by in RCA: 35] [Article Influence: 7.0] [Reference Citation Analysis (1)] |
| 148. | Guo S. Insulin signaling, resistance, and the metabolic syndrome: insights from mouse models into disease mechanisms. J Endocrinol. 2014;220:T1-T23. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 247] [Cited by in RCA: 348] [Article Influence: 29.0] [Reference Citation Analysis (2)] |
| 149. | Hays KE, Pfaffinger JM, Ryznar R. The interplay between gut microbiota, short-chain fatty acids, and implications for host health and disease. Gut Microbes. 2024;16:2393270. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 275] [Cited by in RCA: 220] [Article Influence: 110.0] [Reference Citation Analysis (0)] |
| 150. | Federici S, Nobs SP, Elinav E. Phages and their potential to modulate the microbiome and immunity. Cell Mol Immunol. 2021;18:889-904. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 200] [Cited by in RCA: 164] [Article Influence: 32.8] [Reference Citation Analysis (1)] |
| 151. | Lemos JRN, Hirani K, von Herrath M. Immunological and virological triggers of type 1 diabetes: insights and implications. Front Immunol. 2023;14:1326711. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 47] [Cited by in RCA: 36] [Article Influence: 18.0] [Reference Citation Analysis (1)] |
| 152. | Sausset R, Petit MA, Gaboriau-Routhiau V, De Paepe M. New insights into intestinal phages. Mucosal Immunol. 2020;13:205-215. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 196] [Cited by in RCA: 161] [Article Influence: 26.8] [Reference Citation Analysis (1)] |
| 153. | Duan T, Du Y, Xing C, Wang HY, Wang RF. Toll-Like Receptor Signaling and Its Role in Cell-Mediated Immunity. Front Immunol. 2022;13:812774. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 791] [Cited by in RCA: 668] [Article Influence: 167.0] [Reference Citation Analysis (0)] |
| 154. | Morse ZJ, Simister RL, Crowe SA, Horwitz MS, Osborne LC. Virus induced dysbiosis promotes type 1 diabetes onset. Front Immunol. 2023;14:1096323. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 18] [Article Influence: 6.0] [Reference Citation Analysis (1)] |
| 155. | Luo Y, Li M, Luo D, Tang B. Gut Microbiota: An Important Participant in Childhood Obesity. Adv Nutr. 2025;16:100362. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 36] [Cited by in RCA: 25] [Article Influence: 25.0] [Reference Citation Analysis (0)] |
| 156. | Zepeda-Ríos PA, Cárdenas-Villarreal VM, Castro-Sifuentes D, Guevara-Valtier MC. Interventions to prevent obesity in the first 1000 days: scoping review. Rev Cuid. 2024;15:e3558. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.5] [Reference Citation Analysis (1)] |
| 157. | Muhammad F, Fan B, Wang R, Ren J, Jia S, Wang L, Chen Z, Liu XA. The Molecular Gut-Brain Axis in Early Brain Development. Int J Mol Sci. 2022;23:15389. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 41] [Cited by in RCA: 31] [Article Influence: 7.8] [Reference Citation Analysis (1)] |
| 158. | Neil JA, Cadwell K. The Intestinal Virome and Immunity. J Immunol. 2018;201:1615-1624. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 107] [Cited by in RCA: 90] [Article Influence: 11.3] [Reference Citation Analysis (1)] |
| 159. | Chen Y, Xu J, Chen Y. Regulation of Neurotransmitters by the Gut Microbiota and Effects on Cognition in Neurological Disorders. Nutrients. 2021;13:2099. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 724] [Cited by in RCA: 570] [Article Influence: 114.0] [Reference Citation Analysis (11)] |
| 160. | Volpedo G, Riva A, Nobili L, Zara F, Ravizza T, Striano P. Gut-immune-brain interactions during neurodevelopment: from a brain-centric to a multisystem perspective. BMC Med. 2025;23:263. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 8] [Article Influence: 8.0] [Reference Citation Analysis (1)] |
| 161. | Solanki R, Karande A, Ranganathan P. Emerging role of gut microbiota dysbiosis in neuroinflammation and neurodegeneration. Front Neurol. 2023;14:1149618. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 146] [Cited by in RCA: 115] [Article Influence: 38.3] [Reference Citation Analysis (1)] |
| 162. | Yuan M, Wang Q, Lu Y, Xu P, Pan C, Zhang W, Lu H. Comparison of gut viral communities between autism spectrum disorder and healthy children. Front Cell Infect Microbiol. 2025;15:1660970. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 163. | Bekdas M, Tufan AE, Hakyemez IN, Tas T, Altunhan H, Demircioglu F, Kısmet E; Abant Izzet Baysal University Faculty of Medicine. Subclinical immune reactions to viral infections may correlate with child and adolescent diagnosis of attention-deficit/hyperactivity disorder: a preliminary study from Turkey. Afr Health Sci. 2014;14:439-445. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 12] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 164. | Cassidy-Bushrow AE, Sitarik AR, Johnson CC, Johnson-Hooper TM, Kassem Z, Levin AM, Lynch SV, Ownby DR, Phillips JM, Yong GJM, Wegienka G, Straughen JK. Early-life gut microbiota and attention deficit hyperactivity disorder in preadolescents. Pediatr Res. 2023;93:2051-2060. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 25] [Article Influence: 8.3] [Reference Citation Analysis (1)] |
| 165. | Vatanen T, Jabbar KS, Ruohtula T, Honkanen J, Avila-Pacheco J, Siljander H, Stražar M, Oikarinen S, Hyöty H, Ilonen J, Mitchell CM, Yassour M, Virtanen SM, Clish CB, Plichta DR, Vlamakis H, Knip M, Xavier RJ. Mobile genetic elements from the maternal microbiome shape infant gut microbial assembly and metabolism. Cell. 2022;185:4921-4936.e15. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 155] [Cited by in RCA: 135] [Article Influence: 33.8] [Reference Citation Analysis (1)] |
| 166. | Marano G, Sfratta G, Marzo EM, Cozzo G, Abate F, Traversi G, Mazza O, Capristo E, Gaetani E, Mazza M. The Pediatric Microbiota-Gut-Brain Axis: Implications for Neuropsychiatric Development and Intervention. Children (Basel). 2025;12:1561. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 3] [Article Influence: 3.0] [Reference Citation Analysis (5)] |
| 167. | Lerner A, Arleevskaya M, Schmiedl A, Matthias T. Microbes and Viruses Are Bugging the Gut in Celiac Disease. Are They Friends or Foes? Front Microbiol. 2017;8:1392. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 60] [Cited by in RCA: 46] [Article Influence: 5.1] [Reference Citation Analysis (1)] |
| 168. | Gupta V, Kumar R, Sood U, Singhvi N. Reconciling Hygiene and Cleanliness: A New Perspective from Human Microbiome. Indian J Microbiol. 2020;60:37-44. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 15] [Article Influence: 2.5] [Reference Citation Analysis (1)] |
| 169. | Rook GAW. The old friends hypothesis: evolution, immunoregulation and essential microbial inputs. Front Allergy. 2023;4:1220481. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 68] [Cited by in RCA: 45] [Article Influence: 15.0] [Reference Citation Analysis (1)] |
| 170. | Guarner F, Bourdet-Sicard R, Brandtzaeg P, Gill HS, McGuirk P, van Eden W, Versalovic J, Weinstock JV, Rook GA. Mechanisms of disease: the hygiene hypothesis revisited. Nat Clin Pract Gastroenterol Hepatol. 2006;3:275-284. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 236] [Cited by in RCA: 180] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 171. | Kong LZ, Kim SM, Wang C, Lee SY, Oh SC, Lee S, Jo S, Kim TD. Understanding nucleic acid sensing and its therapeutic applications. Exp Mol Med. 2023;55:2320-2331. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 60] [Cited by in RCA: 52] [Article Influence: 17.3] [Reference Citation Analysis (1)] |
| 172. | Conrad ML, Barrientos G, Cai X, Mukherjee S, Das M, Stephen-Victor E, Harb H. Regulatory T cells and their role in allergic disease. Allergy. 2025;80:77-93. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 24] [Article Influence: 24.0] [Reference Citation Analysis (1)] |
| 173. | Kalinski P, Moser M. Consensual immunity: success-driven development of T-helper-1 and T-helper-2 responses. Nat Rev Immunol. 2005;5:251-260. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 65] [Cited by in RCA: 67] [Article Influence: 3.2] [Reference Citation Analysis (1)] |
| 174. | Frati F, Salvatori C, Incorvaia C, Bellucci A, Di Cara G, Marcucci F, Esposito S. The Role of the Microbiome in Asthma: The Gut⁻Lung Axis. Int J Mol Sci. 2018;20:123. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 231] [Cited by in RCA: 197] [Article Influence: 24.6] [Reference Citation Analysis (2)] |
| 175. | Verma A, Bhagchandani T, Rai A, Nikita, Sardarni UK, Bhavesh NS, Gulati S, Malik R, Tandon R. Short-Chain Fatty Acid (SCFA) as a Connecting Link between Microbiota and Gut-Lung Axis-A Potential Therapeutic Intervention to Improve Lung Health. ACS Omega. 2024;9:14648-14671. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 65] [Cited by in RCA: 59] [Article Influence: 29.5] [Reference Citation Analysis (0)] |
| 176. | Burzyn D, Kuswanto W, Kolodin D, Shadrach JL, Cerletti M, Jang Y, Sefik E, Tan TG, Wagers AJ, Benoist C, Mathis D. A special population of regulatory T cells potentiates muscle repair. Cell. 2013;155:1282-1295. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1098] [Cited by in RCA: 1012] [Article Influence: 77.8] [Reference Citation Analysis (5)] |
| 177. | Lauring AS. Within-Host Viral Diversity: A Window into Viral Evolution. Annu Rev Virol. 2020;7:63-81. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 86] [Cited by in RCA: 70] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 178. | Galeana-Cadena D, Gómez-García IA, Lopez-Salinas KG, Irineo-Moreno V, Jiménez-Juárez F, Tapia-García AR, Boyzo-Cortes CA, Matías-Martínez MB, Jiménez-Alvarez L, Zúñiga J, Camarena A. Winds of change a tale of: asthma and microbiome. Front Microbiol. 2023;14:1295215. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 13] [Article Influence: 4.3] [Reference Citation Analysis (1)] |
| 179. | Handley SA. The virome: a missing component of biological interaction networks in health and disease. Genome Med. 2016;8:32. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 49] [Cited by in RCA: 42] [Article Influence: 4.2] [Reference Citation Analysis (1)] |
| 180. | Talavera-Barber MM, Sánchez PJ, Conces M, Kaptsan I, Everhart K, Leber A, Malleske DT, Moallem M, Panesso-Gómez S, Shimamura M. Molecular testing for gastrointestinal pathogens in intestinal tissue of infants with necrotizing enterocolitis or spontaneous intestinal perforation. J Perinatol. 2024;44:1755-1761. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.5] [Reference Citation Analysis (1)] |
| 181. | Naing Z, Rayner B, Killikulangara A, Vunnam K, Leach S, McIver CJ, Scott GM, Craig ME, Lui K, Rawlinson WD. Prevalence of viruses in stool of premature neonates at a neonatal intensive care unit. J Paediatr Child Health. 2013;49:E221-E226. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 17] [Article Influence: 1.3] [Reference Citation Analysis (1)] |
| 182. | Akhtar N, Jain C, Chauhan SB, Singh I. Gut virus axis: Unravelling viral contribution to gut microbiota dysbiosis and translational medicine in inflammatory bowel disease. Med Microecol. 2025;26:100148. [DOI] [Full Text] |
| 183. | Becerra A, Iša P, Gutiérrez-Escolano AL, Velázquez FR, Torres J, Arias CF, Estrada-Garcia T. Differential virome composition and richness between children’s diarrheagenic stools kept at ultra-low temperatures for long-term. J Infect Dev Ctries. 2023;17:93-101. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 2] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 184. | Zhao L, Li X, Cong H, Yu B. Bacteriophage antimicrobial therapy: From mechanism exploration to biomedical applications. Biomed Pharmacother. 2025;193:118502. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 4] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 185. | Cui L, Watanabe S, Miyanaga K, Kiga K, Sasahara T, Aiba Y, Tan XE, Veeranarayanan S, Thitiananpakorn K, Nguyen HM, Wannigama DL. A Comprehensive Review on Phage Therapy and Phage-Based Drug Development. Antibiotics (Basel). 2024;13:870. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 89] [Cited by in RCA: 77] [Article Influence: 38.5] [Reference Citation Analysis (1)] |
| 186. | Conte MP, Longhi C, Marazzato M, Conte AL, Aleandri M, Lepanto MS, Zagaglia C, Nicoletti M, Aloi M, Totino V, Palamara AT, Schippa S. Adherent-invasive Escherichia coli (AIEC) in pediatric Crohn’s disease patients: phenotypic and genetic pathogenic features. BMC Res Notes. 2014;7:748. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 79] [Cited by in RCA: 68] [Article Influence: 5.7] [Reference Citation Analysis (4)] |
| 187. | Wortelboer K, de Jonge PA, Scheithauer TPM, Attaye I, Kemper EM, Nieuwdorp M, Herrema H. Phage-microbe dynamics after sterile faecal filtrate transplantation in individuals with metabolic syndrome: a double-blind, randomised, placebo-controlled clinical trial assessing efficacy and safety. Nat Commun. 2023;14:5600. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 42] [Article Influence: 14.0] [Reference Citation Analysis (1)] |
| 188. | Zhang Y, Sharma S, Tom L, Liao YT, Wu VCH. Gut Phageome-An Insight into the Role and Impact of Gut Microbiome and Their Correlation with Mammal Health and Diseases. Microorganisms. 2023;11:2454. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 26] [Article Influence: 8.7] [Reference Citation Analysis (1)] |
| 189. | Zhao F, Wang J. Another piece of puzzle for the human microbiome: the gut virome under dietary modulation. J Genet Genomics. 2024;51:983-996. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 7] [Article Influence: 3.5] [Reference Citation Analysis (1)] |
| 190. | Santiago-Rodriguez TM, Hollister EB. Unraveling the viral dark matter through viral metagenomics. Front Immunol. 2022;13:1005107. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 70] [Cited by in RCA: 59] [Article Influence: 14.8] [Reference Citation Analysis (1)] |
| 191. | Wang X, Ma T, Chen Z, Liu Y, Wang K, Liu G, Li K, Chen T, Zhang G, Zhang W, Zhang B. Review of Methods for Studying Viruses in the Environment and Organisms. Viruses. 2025;17:86. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 2] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 192. | Wang Q, Guo F, Jin Y, Ma Y. Applications of human organoids in the personalized treatment for digestive diseases. Signal Transduct Target Ther. 2022;7:336. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 61] [Cited by in RCA: 45] [Article Influence: 11.3] [Reference Citation Analysis (1)] |
| 193. | Hidalgo-Cantabrana C, Sanozky-Dawes R, Barrangou R. Insights into the Human Virome Using CRISPR Spacers from Microbiomes. Viruses. 2018;10:479. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 15] [Article Influence: 1.9] [Reference Citation Analysis (1)] |
| 194. | Bilbo SD, Schwarz JM. The immune system and developmental programming of brain and behavior. Front Neuroendocrinol. 2012;33:267-286. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 513] [Cited by in RCA: 450] [Article Influence: 32.1] [Reference Citation Analysis (0)] |