Published online Mar 25, 2026. doi: 10.5501/wjv.v15.i1.117643
Revised: January 15, 2026
Accepted: February 9, 2026
Published online: March 25, 2026
Processing time: 91 Days and 17.7 Hours
Infections with certain viruses are strong risk factors for specific cancers. The human major histocompatibility complex class I chain-related genes A (MICA) and B (MICB) are polymorphic, non-classical major histocompatibility complex class I genes located within the human leukocyte antigen region. Polymorphisms in these genes have been associated with susceptibility and outcomes of several virus-associated cancers. The underlying mechanisms involve modulation of natural killer cell- and CD8+ T cell-mediated cytotoxicity by disrupting the na
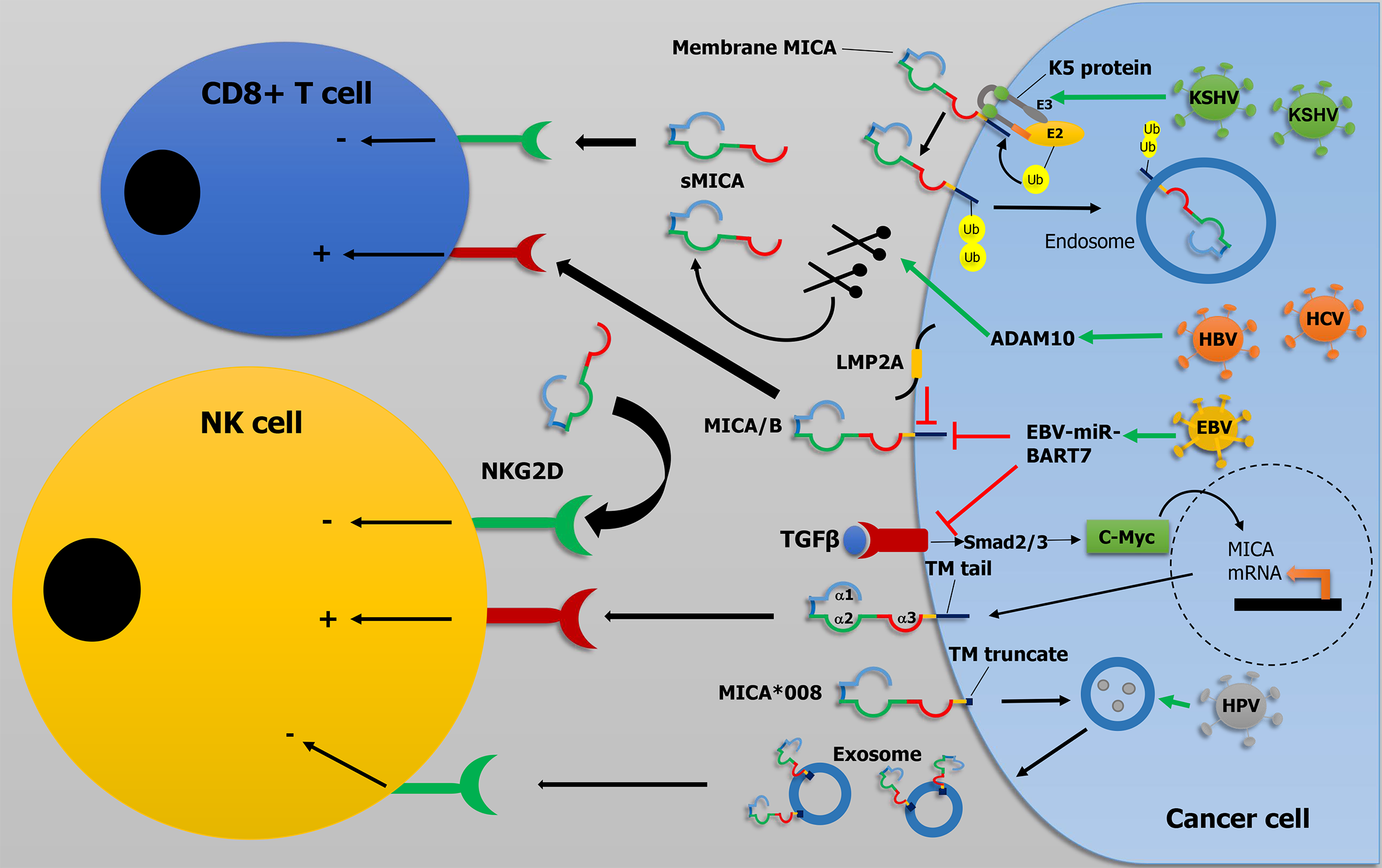
Core Tip: Several oncogenic viruses have evolved multi-layered strategies to subvert the natural killer group 2-member D (NKG2D)-major histocompatibility complex class I chain-related genes A (MICA)/B axis and promote cancer progression. Human papillomavirus drives immune evasion through exosomal release of the truncated MICA*008 variant, which potently downregulates NKG2D and impairs natural killer (NK) cell cytotoxicity. Chronic hepatitis B virus and hepatitis C virus infections create a hypoxic/inflammatory tumor milieu that induces metalloproteases, generating soluble MICA/B that suppresses NKG2D on NK and CD8+ T cells. EBV evades NKG2D-mediated immunity through miR-bamHI-A rightward transcript 7-mediated repression of the transforming growth factor beta 1/cellular myelocytomatosis oncogene/MICA axis and latent membrane protein 2A-driven disruption of stress-induced ligand expression, reinforced by UPR-protein disulfide isomerase inhibition of proper MICA/MICB folding and trafficking. Kaposi’s sarcoma-associated herpesvirus exploits its E3 ubiquitin ligases K3 and K5 to ubiquitinate MICA and other NK-activating ligands such as activation-induced C-type lectin, triggering their endocytic removal and intracellular sequestration to disable multiple NK cell recognition pathways. Thera
- Citation: Bouayad A. Major histocompatibility complex class I chain-related A and B molecules and their potential role in virus-associated cancers. World J Virol 2026; 15(1): 117643
- URL: https://www.wjgnet.com/2220-3249/full/v15/i1/117643.htm
- DOI: https://dx.doi.org/10.5501/wjv.v15.i1.117643
Several human oncogenic viruses have been identified and are responsible for approximately 10%-15% of cancers worldwide[1]. These include human papillomavirus (HPV), hepatitis B and C viruses (HBV/HCV), Epstein-Barr virus (EBV), Kaposi’s sarcoma-associated herpesvirus (KSHV), human T cell lymphotropic virus, and Merkel cell polyomavirus[2]. HPV plays a significant role in the development of cervical carcinoma (CC)[3], while HBV/HCV are major causes of hepatocellular carcinoma (HCC)[4]. EBV is associated with nasopharyngeal carcinoma (NPC), gastric carcinoma, Burkitt lymphoma, and Hodgkin lymphoma[1,5]. KSHV causes Kaposi’s sarcoma[6]. The dynamic interplay between viruses and the tumor microenvironment (TME) shapes the development and progression of virus-associated cancers[7]. Genetic variations in Killer-cell immunoglobulin-like receptors and major histocompatibility complex (MHC) class I genes can modulate natural killer (NK) cell responses and the TME, thereby promoting immune evasion and tumor survival[8].
Among MHC class I genes, polymorphisms and altered expression of MHC class I chain-related genes A (MICA) and B (MICB) have been proposed to contribute to susceptibility to virus-associated cancers[9-12]. MICA and MICB are known nonclassical MHC class I-encoded molecules that play important roles in immune surveillance. Furthermore, many virus-infected tumor cells release soluble MICA (sMICA) proteins to escape NK cell recognition in certain virus-associated cancers, such as HCC[13].
This mini-review summarizes the genetics and biology of MICA and MICB, highlighting their emerging importance in the pathogenesis of virus-associated cancers.
MICA and MICB bind NK group 2-member D (NKG2D) receptor, an activating receptor expressed in NK cells. These genes are located within the human leukocyte antigen (HLA) complex, 46 kb and 141 kb centromeric of the HLA-B locus, respectively[14]. MICA and MICB genes are composed of six exons that encode the leader peptide, three extracellular domains (α1, α2, and α3), a transmembrane segment, and a C-terminal cytoplasmic tail[15]. These highly polymorphic genes exhibit their genetic variation primarily in the α1, α2, and α3 immunoglobulin-like extracellular structural domains, which are encoded by exons 2-4. Current data from the IMGT/HLA database document 589 human MICA alleles and 312 human MICB alleles. Polymorphism of the MICA and MICB genes has been implicated in susceptibility to cancer. Notably, A5 and A9 alleles of MICA may be an important protective factor for cancer in Caucasian populations[16].
MICA and MICB are widely expressed on the surface of tumor cells, including lung, breast, gastric, kidney, ovarian, prostate, colon carcinomas, and melanomas. When expressed on tumor cell surfaces, MICA/B can be recognized by NK cells through the NKG2D receptor, promoting a cytotoxic response that leads to tumor cell lysis. The importance of NKG2D in immune surveillance of tumors is highlighted by the observation that NKG2D-deficient mice are more susceptible to the development of oncogene-induced tumors[17]. Although NKG2D-MICA/B interaction represents a valid strategy to limit tumor progression, tumor cells display escape strategies that subvert the biological function of NKG2D. The underlying mechanisms involve the proteolytic shedding of MICA and other NKG2D ligands, induced by tumor-secreted metalloproteases (matrix metalloproteinase) or secretion in exosomes[18,19]. The released sMICA and soluble MICB can subsequently bind to NKG2D and induce its down-modulation and degradation, subverting NKG2D-dependent effector functions of NK cells and facilitating tumor immune escape[20].
CC and its precursor lesions, cervical intraepithelial neoplasia (CIN), are associated with persistent infection by oncogenic HPV subtypes, particularly HPV-16 and HPV-18, which are responsible for approximately 70% of CC cases. Several factors, including immune response, immunological genes, multiparity, oral contraceptive use, and smoking, appear to influence the mechanisms that determine the regression or progression to CIN and CC[3,21-24].
Among genetic factors, polymorphisms of MICA and MICB have been implicated in the pathogenesis of CC and CIN lesions via their interaction with NKG2D, which may lead either to viral clearance or tumor immune evasion (Figure 1). In a case-control study, von Linsingen et al[10] reported that MICA*008: 01/04 and MICA*018: 01 alleles act as risk factors, whereas MICA*002: 01/020 represents a protective factor against susceptibility to CC and CIN lesions in patients from southern Brazil. Genotyping based on the MICA-129 (Met/Val) dimorphism showed that Val/Val and Val/MICAdel genotypes were more frequent among patients with CIN and CC[10]. The MICA-129 rs1051792 variant, which substitutes Methionine with Valine at position 129, alters the affinity of MICA for the NKG2D receptor and can diminish NK cell activation[25]. Additional genetic studies have identified HPV-related CC-risk variants in the MICA region, including the rs2516448 T allele, which is in strong linkage disequilibrium with the A5.1 frameshift mutation in exon 5 that produces a truncated MICA protein[26], and rs2844511, which is associated with invasive CC and high-grade dysplasia[27]. However, some studies have reported no significant association between MICA polymorphisms and CIN, underscoring population-specific variability[28].
The truncated protein translated from MICA*008 (MICA-A5.1) allele is transported to the apical surface of human cells and released from cells as a membrane-anchored full-length molecule in exosomes that triggers significantly greater NKG2D downregulation and impairs NK cell cytotoxicity (Figure 1)[18].
Despite their strong genomic and protein homology, MICA and MICB are regulated differently in CC-derived cell lines[29]. MICA is generally expressed at higher surface levels than MICB in HPV-positive lines[29], and its expression is significantly upregulated following genotoxic stress, suggesting a stress-inducible role in antiviral immune surveillance[29]. Furthermore, a genome-wide analysis identified rs6938453 in MICA as a protective variant against invasive CC among women with CIN3, further underscoring the complex interplay between MICA expression, polymorphism, and disease progression (Table 1)[30].
| Cancer type | Oncogenic virus | MICA/MICB alterations | Immune evasion mechanisms | Clinical implications |
| Cervical cancer | Human papillomavirus 16/18 | Risk alleles: MICA*008:01, MICA*008:04, and MICA*018:01; protective alleles: MICA*002:01 and MICA*002:02; risk genotypes: MICA-129 Val/Val and Val/MICA-del; truncated MICA*008 | Exosomal release of truncated MICA*008; reduce surface MICA/MICB; reduce NKG2D on NK cells | MICA polymorphisms associated with progression from cervical intraepithelial neoplasia to invasive cancer; reduce invasive risk linked to rs6938453 |
| HCC | HBV/HCV | Reduce MICA/MICB expression; increase sMICA; rs2596542 TT genotype (risk); rs2596538 C/C genotype (increase HCV-HCC risk) | MICA shedding by ADAM10; UPR-mediated reduce MICA/MICB; sMICA-induced NKG2D reduce on NK/CD8+ T cells | Reduce MICA/MICB expression correlates with tumour aggressiveness and poor survival; increase serum sMICA predicts poor prognosis in HBV-HCC |
| Nasopharyngeal carcinoma | EBV | MICA rs2596542 AA genotype (risk); increase MICA protein expression in AA tumors; rs1051792 A allele (increase EBV titers) | EBV-miR-bamHI-A rightward transcript 7 inhibits transforming growth factor beta 1/cellular myelocytomatosis oncogene/MICA; latent membrane protein 2A-induced reduce surface MICA/MICB; UPR-protein disulfide isomerase inhibits proper MICA/MICB folding | AA genotype associated with increase EBV titers and MICA expression; NKG2D rs1049174 LNK/LNK genotype correlates with increase susceptibility |
| Gastric carcinoma | EBV | Variable MICA expression | Latent EBV infection with limited immune recognition | Dihydrotestosterone enhances MICA via nuclear factor-kappa B pathway in cell-line models, boosting NK/T cell cytotoxicity |
| Kaposi’s sarcoma | Kaposi’s sarcoma-associated herpesvirus | MICA*008 allele resistant to viral downregulation | K5 ubiquitination of MICA cytoplasmic tail; endocytic removal and intracellular sequestration of MICA; coordinated targeting of NKG2D (via MICA) and NKp80 (via activation-induced C-type lectin) | MICA*008 allele may influence host susceptibility due to resistance to K5-mediated downregulation |
Chronic HCV and HBV infections account for approximately 80% of HCC cases worldwide[31]. Decreased expression of MICA and MICB is associated with HCC aggressiveness, poor clinical outcomes, and shorter survival times (Table 1)[32,33]. Mechanistically, activation of the unfolded protein response (UPR) downregulates MICA/MICB through post-transcriptional repression and proteasomal degradation, thereby diminishing NK cell–mediated cytotoxicity[33]. This interplay among MICA/B, UPR signaling, and NK cell regulation may constitute a novel immune-evasion pathway in HCC (Figure 1). Restoring membrane MICA expression has therapeutic potential. Histone deacetylase inhibitors, such as suberoylanilide hydroxamic acid, epigenetically upregulate MICA by modulating the miR-17-92 cluster and minichromosome maintenance complex component 7, thereby enhancing the susceptibility of hepatoma cells to NK cell-mediated lysis[34]. Single-cell transcriptomic analyses also show that MICA expression positively correlates with interferon regulatory factor 1 (IRF1) in HCC cells[35]. Consistently, cisplatin treatment or checkpoint kinase 1 inhibition upregulates MICA in an IRF1-dependent manner, acting through a specific IRF-responsive element located at -1756 bp in the MICA promoter[35]. Increased MICA expression may further promote NK cell and CD8+ T cell infiltration into the TME[35]. Additionally, MICA-positive tumor cells influence macrophage phenotype through peroxisome proliferator-activated receptor/enoyl-coA hydratase and 3-hydroxyacyl-coA dehydrogenase (EHHADH)-mediated fatty-acid metabolism. EHHADH enhances fatty-acid uptake and oxidation in macrophages, favoring an M2-like immunosuppressive phe
Genetic polymorphisms in MICA influence susceptibility to HCC in HBV- and HCV-infected individuals. The TT genotype at rs2596542 has been identified as a risk factor for HCC development in patients with HBV or HCV infection, with a stronger effect observed in HCV-infected patients[12]. In addition, the MICA rs2596542 G/G genotype and the rs2596538 C/C genotype are associated with increased risk of HCV-related HCC in a Sicilian population[37]. Gene-gene interaction analyses further suggest that rs2596542 interacts with rs2069705 to increase HCC risk by 1.41-fold, and its interaction with rs2069727 increases risk by 5.56-fold[38]. These findings highlight the importance of MICA genetic va
Chronic HBV and HCV infections generate a hypoxic TME that stabilizes HIF-1α, subsequently inducing hypoxia-responsive metalloproteases such as ADAM10, which mediate proteolytic shedding of membrane-bound MICA to produce sMICA[39-41]. The sMICA downmodulates NKG2D on NK and CD8+ T cells, thereby promoting immune escape in HCC (Figure 1). Retinoids counteract this process by inhibiting ADAM9 and ADAM10 enzymatic activity, leading to reduced sMICA levels[42]. The rs2596542 polymorphism in the MICA promoter region may also regulate sMICA production and influence HCC risk[12]. In HBV-related HCC, the G allele of rs2596542 is associated with elevated sMICA levels, and patients with high serum sMICA exhibit poorer prognosis[13], supporting the utility of sMICA as a predictive biomarker in HBV-induced disease. In contrast, a genome-wide association study reported that the rs2596542 risk allele is associated with lower sMICA levels in HCV-induced HCC[43], suggesting viral-specific differences in the regulation of MICA shedding.
EBV is a human herpesvirus that has been reported to be associated with NPC, gastric carcinoma, Burkitt lymphoma, and Hodgkin lymphoma. In most of these tumors, the virus remains in a latently infected state. During latency, EBV expresses latent membrane protein 2A (LMP2A) along with a few other genes. EBV infection can also be found in normal immunocompetent individuals without triggering an innate immune response[44].
NPC is a rare epithelial carcinoma arising from the nasopharyngeal region. Its pathogenesis is closely linked to EBV infection, although genetic and lifestyle factors also appear to play a role. EBV has evolved several strategies to subvert T cell-mediated and NK cell-mediated cytotoxicity and promote NPC progression (Figure 1). NK cell recognition is compromised by the downregulation of MICA and MICB by EBV-miRNAs[45]. Notably, EBV-encoded microRNA bamHI-A rightward transcript 7 inhibits the transforming growth factor beta 1/cellular myelocytomatosis oncogene/MICA regulatory axis, downregulates MICA, and reduces NK cell cytotoxicity against NPC cells[46]. This immune evasion can be counteracted by synthetic immunomodulatory drugs, such as pomalidomide, which increase MICA expression in EBV-infected Daudi cells, thereby enhancing T cell activation and NK cell-mediated cytotoxicity[47]. LMP2A is another EBV-encoded protein that plays a key role in immune evasion and NPC development (Table 1). Singh and Banerjee[48] demonstrated that LMP2A induces downregulation of MICA and MICB surface expression. UPR activation also contributes to this regulation, as protein disulfide isomerase inhibits proper folding and surface expression of MICA and MICB in LMP2A-expressing cells, thereby further reducing NKG2D-mediated immune recognition[48]. Specific MICA polymorphisms may also influence individual susceptibility to EBV-induced NPC. The AA genotype of MICA rs2596542 was significantly associated with NPC[49]. Tumor specimens from NPC patients with the AA genotype exhibit higher MICA protein expression and higher EBV titers compared with tumors from patients carrying GG or GA genotypes[49]. Higher EBV copy numbers were also observed in tumors with the A allele of MICA rs1051792 (MICA-129 Met/Val) compared with those carrying the G allele[49]. Similarly, the rs1049174 polymorphism in the NKG2D gene, with the LNK/LNK genotype associated with reduced NKG2D receptor expression and diminished NK cell cytotoxicity, correlates with increased susceptibility to NPC[49].
EBV-associated gastric carcinoma (EBVaGC) is a subset of gastric cancers linked to EBV infection. Several therapeutic strategies have been developed to enhance anti-tumor immunity against EBVaGC. Among these, dihydrotestosterone may stimulate MICA-mediated immune responses by activating the nuclear factor-kappa B pathway, leading to increased transcription of target genes, including MICA and EBNA1[50]. These effects enhance interactions with NK and T cells, thereby boosting cytotoxicity against EBVaGC cells.
Accumulating evidence suggests that risk factors for classical Hodgkin lymphoma differ according to tumor EBV status. This potential etiological heterogeneity is not currently reflected in disease classification. A genome-wide association study of classical Hodgkin lymphoma patients identified protective loci in the MHC region, one of which is located adjacent to MICB[51].
Several herpesviruses exploit the ubiquitin system by targeting cell-surface receptors recognized by NK cells and cytotoxic T lymphocytes for internalization, thereby evading immune detection. Notably, KSHV encodes membrane-associated E3 ubiquitin ligases, K3 and K5. K5 downregulates MICA by ubiquitinating lysine residues in its cytoplasmic tail, triggering its redistribution from the plasma membrane to intracellular compartments without accelerating degra
The convergence of immune evasion strategies targeting the NKG2D-MICA/B axis across diverse oncogenic viruses, despite their distinct genomes and life cycles, underscores the fundamental importance of this pathway in anti-tumor immunity. Oncogenic viruses disrupt this mechanism through transcriptional repression (EBV miR-bamHI-A rightward transcript 7), proteolytic shedding mediated by metalloproteases (HBV/HCV), exosomal release of truncated variants (HPV MICA*008), or ubiquitin-mediated internalization (KSHV K5). This convergence highlights the NKG2D-MICA/B axis as a central vulnerability in tumor immune recognition and an attractive target for therapeutic intervention. Strategies restoring NKG2D-mediated immunity, such as blocking sMICA, preventing MICA shedding, or enhancing membrane MICA, may have broad applicability across multiple virus-associated malignancies.
The sMICA facilitates immune evasion by downregulating NKG2D receptor expression and also serves as a valuable prognostic biomarker, particularly in HBV-induced HCC, where elevated serum levels correlate with poor clinical out
| 1. | de Martel C, Georges D, Bray F, Ferlay J, Clifford GM. Global burden of cancer attributable to infections in 2018: a worldwide incidence analysis. Lancet Glob Health. 2020;8:e180-e190. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1982] [Cited by in RCA: 1617] [Article Influence: 269.5] [Reference Citation Analysis (6)] |
| 2. | Schiller JT, Lowy DR. An Introduction to Virus Infections and Human Cancer. Recent Results Cancer Res. 2021;217:1-11. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 110] [Cited by in RCA: 87] [Article Influence: 17.4] [Reference Citation Analysis (1)] |
| 3. | Włoszek E, Krupa K, Skrok E, Budzik MP, Deptała A, Badowska-Kozakiewicz A. HPV and Cervical Cancer-Biology, Prevention, and Treatment Updates. Curr Oncol. 2025;32:122. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 22] [Article Influence: 22.0] [Reference Citation Analysis (0)] |
| 4. | Wen B, Te L, Bai C, Jiang W, Zuo D, Hao Q, Wang J, Ren L. Relative contribution of hepatitis B and C viruses in primary liver cancer in China: A systematic review and meta-analysis. J Infect. 2024;89:106298. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 12] [Article Influence: 6.0] [Reference Citation Analysis (1)] |
| 5. | Young LS, Yap LF, Murray PG. Epstein-Barr virus: more than 50 years old and still providing surprises. Nat Rev Cancer. 2016;16:789-802. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 738] [Cited by in RCA: 669] [Article Influence: 66.9] [Reference Citation Analysis (4)] |
| 6. | Robinson CA, Singh GK, Kleer M, Katsademas T, Castle EL, Boudreau BQ, Corcoran JA. Kaposi's sarcoma-associated herpesvirus (KSHV) utilizes the NDP52/CALCOCO2 selective autophagy receptor to disassemble processing bodies. PLoS Pathog. 2023;19:e1011080. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 12] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 7. | Fernandes Q, Folorunsho OG. Unveiling the nexus: The tumor microenvironment as a strategic frontier in viral cancers. Cytokine. 2025;185:156827. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 4] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 8. | Li S, Dai W, Kam NW, Zhang J, Lee VHF, Ren X, Kwong DL. The Role of Natural Killer Cells in the Tumor Immune Microenvironment of EBV-Associated Nasopharyngeal Carcinoma. Cancers (Basel). 2024;16:1312. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 13] [Article Influence: 6.5] [Reference Citation Analysis (1)] |
| 9. | Goto K, Kato N. MICA SNPs and the NKG2D system in virus-induced HCC. J Gastroenterol. 2015;50:261-272. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 44] [Cited by in RCA: 40] [Article Influence: 3.6] [Reference Citation Analysis (1)] |
| 10. | von Linsingen R, Pinho de França P, de Carvalho NS, Bicalho MDG. MICA and KLRK1 genes and their impact in cervical intraepithelial neoplasia development in the southern Brazilian population. Hum Immunol. 2020;81:249-253. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 4] [Article Influence: 0.7] [Reference Citation Analysis (1)] |
| 11. | Luangtrakool P, Vejbaesya S, Luangtrakool K, Ngamhawornwong S, Apisawes K, Kalayanarooj S, Macareo LR, Fernandez S, Jarman RG, Collins RWM, Cox ST, Srikiatkhachorn A, Rothman AL, Stephens HAF. Major Histocompatibility Complex Class I Chain-Related A and B (MICA and MICB) Gene, Allele, and Haplotype Associations With Dengue Infections in Ethnic Thais. J Infect Dis. 2020;222:840-846. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 6] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 12. | Luo X, Wang Y, Shen A, Deng H, Ye M. Relationship between the rs2596542 polymorphism in the MICA gene promoter and HBV/HCV infection-induced hepatocellular carcinoma: a meta-analysis. BMC Med Genet. 2019;20:142. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 13] [Article Influence: 1.9] [Reference Citation Analysis (1)] |
| 13. | Kumar V, Yi Lo PH, Sawai H, Kato N, Takahashi A, Deng Z, Urabe Y, Mbarek H, Tokunaga K, Tanaka Y, Sugiyama M, Mizokami M, Muroyama R, Tateishi R, Omata M, Koike K, Tanikawa C, Kamatani N, Kubo M, Nakamura Y, Matsuda K. Soluble MICA and a MICA variation as possible prognostic biomarkers for HBV-induced hepatocellular carcinoma. PLoS One. 2012;7:e44743. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 76] [Cited by in RCA: 79] [Article Influence: 5.6] [Reference Citation Analysis (5)] |
| 14. | Al Hadra B, Lukanov TI, Ivanova MI. HLA class I chain-related MICA and MICB genes polymorphism in healthy individuals from the Bulgarian population. Hum Immunol. 2022;83:551-555. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 5] [Article Influence: 1.3] [Reference Citation Analysis (1)] |
| 15. | Machuldova A, Holubova M, Caputo VS, Cedikova M, Jindra P, Houdova L, Pitule P. Role of Polymorphisms of NKG2D Receptor and Its Ligands in Acute Myeloid Leukemia and Human Stem Cell Transplantation. Front Immunol. 2021;12:651751. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 14] [Article Influence: 2.8] [Reference Citation Analysis (1)] |
| 16. | Ji M, Wang J, Yuan L, Zhang Y, Zhang J, Dong W, Peng X. MICA polymorphisms and cancer risk: a meta-analysis. Int J Clin Exp Med. 2015;8:818-826. [PubMed] |
| 17. | Crane CA, Austgen K, Haberthur K, Hofmann C, Moyes KW, Avanesyan L, Fong L, Campbell MJ, Cooper S, Oakes SA, Parsa AT, Lanier LL. Immune evasion mediated by tumor-derived lactate dehydrogenase induction of NKG2D ligands on myeloid cells in glioblastoma patients. Proc Natl Acad Sci U S A. 2014;111:12823-12828. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 158] [Cited by in RCA: 147] [Article Influence: 12.3] [Reference Citation Analysis (1)] |
| 18. | Ashiru O, Boutet P, Fernández-Messina L, Agüera-González S, Skepper JN, Valés-Gómez M, Reyburn HT. Natural killer cell cytotoxicity is suppressed by exposure to the human NKG2D ligand MICA*008 that is shed by tumor cells in exosomes. Cancer Res. 2010;70:481-489. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 346] [Cited by in RCA: 339] [Article Influence: 21.2] [Reference Citation Analysis (1)] |
| 19. | Huang S, Qin Z, Wang F, Kang Y, Ren B. A potential mechanism of tumor immune escape: Regulation and application of soluble natural killer group 2 member D ligands (Review). Oncol Rep. 2024;52:137. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 6] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 20. | Torres N, Regge MV, Secchiari F, Friedrich AD, Spallanzani RG, Raffo Iraolagoitia XL, Núñez SY, Sierra JM, Ziblat A, Santilli MC, Gilio N, Almada E, Lauche C, Pardo R, Domaica CI, Fuertes MB, Madauss KP, Hance KW, Gloger IS, Zylberman V, Goldbaum FA, Zwirner NW. Restoration of antitumor immunity through anti-MICA antibodies elicited with a chimeric protein. J Immunother Cancer. 2020;8:e000233. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 29] [Cited by in RCA: 23] [Article Influence: 3.8] [Reference Citation Analysis (1)] |
| 21. | Luhn P, Walker J, Schiffman M, Zuna RE, Dunn ST, Gold MA, Smith K, Mathews C, Allen RA, Zhang R, Wang S, Wentzensen N. The role of co-factors in the progression from human papillomavirus infection to cervical cancer. Gynecol Oncol. 2013;128:265-270. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 116] [Cited by in RCA: 84] [Article Influence: 6.5] [Reference Citation Analysis (1)] |
| 22. | Qian N, Chen X, Han S, Qiang F, Jin G, Zhou X, Dong J, Wang X, Shen H, Hu Z. Circulating IL-1beta levels, polymorphisms of IL-1B, and risk of cervical cancer in Chinese women. J Cancer Res Clin Oncol. 2010;136:709-716. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 59] [Cited by in RCA: 52] [Article Influence: 3.3] [Reference Citation Analysis (1)] |
| 23. | Ivansson EL, Juko-Pecirep I, Gyllensten UB. Interaction of immunological genes on chromosome 2q33 and IFNG in susceptibility to cervical cancer. Gynecol Oncol. 2010;116:544-548. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 58] [Article Influence: 3.6] [Reference Citation Analysis (1)] |
| 24. | Ades S, Koushik A, Duarte-Franco E, Mansour N, Arseneau J, Provencher D, Gilbert L, Gotlieb W, Ferenczy A, Coutlée F, Roger M, Franco EL; Biomarkers of Cervical Cancer Risk (BCCR) Study Team. Selected class I and class II HLA alleles and haplotypes and risk of high-grade cervical intraepithelial neoplasia. Int J Cancer. 2008;122:2820-2826. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 28] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 25. | Isernhagen A, Malzahn D, Bickeböller H, Dressel R. Impact of the MICA-129Met/Val Dimorphism on NKG2D-Mediated Biological Functions and Disease Risks. Front Immunol. 2016;7:588. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 43] [Article Influence: 4.3] [Reference Citation Analysis (1)] |
| 26. | Chen D, Hammer J, Lindquist D, Idahl A, Gyllensten U. A variant upstream of HLA-DRB1 and multiple variants in MICA influence susceptibility to cervical cancer in a Swedish population. Cancer Med. 2014;3:190-198. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 24] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 27. | Ramachandran D, Schürmann P, Mao Q, Wang Y, Bretschneider LM, Speith LM, Hülse F, Enßen J, Bousset K, Jentschke M, Böhmer G, Strauß HG, Hirchenhain C, Schmidmayr M, Tarbiat J, Runnebaum I, Dürst M, Hein A, Koch M, Ruebner M, Ekici A, Beckmann MW, Fasching PA, Luyten A, Petry KU, Hillemanns P, Dörk T. Association of genomic variants at the human leukocyte antigen locus with cervical cancer risk, HPV status and gene expression levels. Int J Cancer. 2020;147:2458-2468. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 20] [Article Influence: 3.3] [Reference Citation Analysis (1)] |
| 28. | Jumnainsong A, Romphruk AV, Jearanaikoon P, Klumkrathok K, Romphruk A, Luanrattanakorn S, Leelayuwat C. Association of polymorphic extracellular domains of MICA with cervical cancer in northeastern Thai population. Tissue Antigens. 2007;69:326-333. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 17] [Article Influence: 0.9] [Reference Citation Analysis (1)] |
| 29. | Del Toro-Arreola S, Arreygue-Garcia N, Aguilar-Lemarroy A, Cid-Arregui A, Jimenez-Perez M, Haramati J, Barros-Nuñez P, Gonzalez-Ramella O, Del Toro-Arreola A, Ortiz-Lazareno P, Hernandez-Flores G, Bravo-Cuellar A, Daneri-Navarro A, Jave-Suarez LF. MHC class I-related chain A and B ligands are differentially expressed in human cervical cancer cell lines. Cancer Cell Int. 2011;11:15. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 15] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 30. | Bowden SJ, Bodinier B, Kalliala I, Zuber V, Vuckovic D, Doulgeraki T, Whitaker MD, Wielscher M, Cartwright R, Tsilidis KK, Bennett P, Jarvelin MR, Flanagan JM, Chadeau-Hyam M, Kyrgiou M; FinnGen consortium. Genetic variation in cervical preinvasive and invasive disease: a genome-wide association study. Lancet Oncol. 2021;22:548-557. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 85] [Cited by in RCA: 76] [Article Influence: 15.2] [Reference Citation Analysis (0)] |
| 31. | El-Serag HB. Epidemiology of viral hepatitis and hepatocellular carcinoma. Gastroenterology. 2012;142:1264-1273.e1. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2629] [Cited by in RCA: 2552] [Article Influence: 182.3] [Reference Citation Analysis (10)] |
| 32. | Cadoux M, Caruso S, Pham S, Gougelet A, Pophillat C, Riou R, Loesch R, Colnot S, Nguyen CT, Calderaro J, Celton-Morizur S, Guerra N, Zucman-Rossi J, Desdouets C, Couty JP. Expression of NKG2D ligands is downregulated by β-catenin signalling and associates with HCC aggressiveness. J Hepatol. 2021;74:1386-1397. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 58] [Cited by in RCA: 53] [Article Influence: 10.6] [Reference Citation Analysis (4)] |
| 33. | Fang L, Gong J, Wang Y, Liu R, Li Z, Wang Z, Zhang Y, Zhang C, Song C, Yang A, Ting JP, Jin B, Chen L. MICA/B expression is inhibited by unfolded protein response and associated with poor prognosis in human hepatocellular carcinoma. J Exp Clin Cancer Res. 2014;33:76. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 66] [Cited by in RCA: 65] [Article Influence: 5.4] [Reference Citation Analysis (1)] |
| 34. | Yang H, Lan P, Hou Z, Guan Y, Zhang J, Xu W, Tian Z, Zhang C. Histone deacetylase inhibitor SAHA epigenetically regulates miR-17-92 cluster and MCM7 to upregulate MICA expression in hepatoma. Br J Cancer. 2015;112:112-121. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 95] [Cited by in RCA: 99] [Article Influence: 9.0] [Reference Citation Analysis (1)] |
| 35. | Li X, Huang J, Wu Q, Du Q, Wang Y, Huang Y, Cai X, Geller DA, Yan Y. Inhibition of Checkpoint Kinase 1 (CHK1) Upregulates Interferon Regulatory Factor 1 (IRF1) to Promote Apoptosis and Activate Anti-Tumor Immunity via MICA in Hepatocellular Carcinoma (HCC). Cancers (Basel). 2023;15:850. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 11] [Article Influence: 3.7] [Reference Citation Analysis (1)] |
| 36. | Huang J, Teng Y, Yan P, Yang Y, Lin S, Wu Q, Du Q, Li X, Yao M, Li J, Huang Y, Cai X, Geller DA, Yan Y. MICA+ Tumor Cells Modulate Macrophage Phenotype and Function via PPAR/EHHADH-Mediated Fatty Acid Metabolism in Hepatocellular Carcinoma (HCC). Cancers (Basel). 2025;17:2365. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 37. | Augello G, Balasus D, Fusilli C, Mazza T, Emma MR, Giannitrapani L, Agliastro R, Cervello M, Montalto G. Association Between MICA Gene Variants and the Risk of Hepatitis C Virus-Induced Hepatocellular Cancer in a Sicilian Population Sample. OMICS. 2018;22:274-282. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 9] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 38. | Li H, Liu F, Zhu H, Zhou X, Lu J, Chang H, Hu J. Interaction Between Polymorphisms of IFN-γ and MICA Correlated with Hepatocellular Carcinoma. Med Sci Monit. 2016;22:549-553. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12] [Cited by in RCA: 14] [Article Influence: 1.4] [Reference Citation Analysis (1)] |
| 39. | Yoo YG, Oh SH, Park ES, Cho H, Lee N, Park H, Kim DK, Yu DY, Seong JK, Lee MO. Hepatitis B virus X protein enhances transcriptional activity of hypoxia-inducible factor-1alpha through activation of mitogen-activated protein kinase pathway. J Biol Chem. 2003;278:39076-39084. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 150] [Cited by in RCA: 149] [Article Influence: 6.5] [Reference Citation Analysis (1)] |
| 40. | Zhu C, Liu X, Wang S, Yan X, Tang Z, Wu K, Li Y, Liu F. Hepatitis C virus core protein induces hypoxia-inducible factor 1α-mediated vascular endothelial growth factor expression in Huh7.5.1 cells. Mol Med Rep. 2014;9:2010-2014. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 24] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 41. | Kohga K, Takehara T, Tatsumi T, Miyagi T, Ishida H, Ohkawa K, Kanto T, Hiramatsu N, Hayashi N. Anticancer chemotherapy inhibits MHC class I-related chain a ectodomain shedding by downregulating ADAM10 expression in hepatocellular carcinoma. Cancer Res. 2009;69:8050-8057. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 72] [Cited by in RCA: 72] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 42. | Otoyama Y, Arai J, Goto K, Nozawa H, Nakagawa R, Muroyama R, Sugiura I, Nakajima Y, Kajiwara A, Tojo M, Ichikawa Y, Uozumi S, Shimozuma Y, Uchikoshi M, Sakaki M, Kato N, Yoshida H. Retinoids Decrease Soluble MICA Concentration by Inhibiting the Enzymatic Activity of ADAM9 and ADAM10. Anticancer Res. 2021;41:2307-2320. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 10] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 43. | Kumar V, Kato N, Urabe Y, Takahashi A, Muroyama R, Hosono N, Otsuka M, Tateishi R, Omata M, Nakagawa H, Koike K, Kamatani N, Kubo M, Nakamura Y, Matsuda K. Genome-wide association study identifies a susceptibility locus for HCV-induced hepatocellular carcinoma. Nat Genet. 2011;43:455-458. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 312] [Cited by in RCA: 291] [Article Influence: 19.4] [Reference Citation Analysis (6)] |
| 44. | Young LS, Rickinson AB. Epstein-Barr virus: 40 years on. Nat Rev Cancer. 2004;4:757-768. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1776] [Cited by in RCA: 1626] [Article Influence: 73.9] [Reference Citation Analysis (0)] |
| 45. | Nachmani D, Stern-Ginossar N, Sarid R, Mandelboim O. Diverse herpesvirus microRNAs target the stress-induced immune ligand MICB to escape recognition by natural killer cells. Cell Host Microbe. 2009;5:376-385. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 398] [Cited by in RCA: 376] [Article Influence: 22.1] [Reference Citation Analysis (1)] |
| 46. | Wong TS, Chen S, Zhang MJ, Chan JY, Gao W. Epstein-Barr virus-encoded microRNA BART7 downregulates major histocompatibility complex class I chain-related peptide A and reduces the cytotoxicity of natural killer cells to nasopharyngeal carcinoma. Oncol Lett. 2018;16:2887-2892. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 19] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 47. | Davis DA, Shrestha P, Aisabor AI, Stream A, Galli V, Pise-Masison CA, Tagawa T, Ziegelbauer JM, Franchini G, Yarchoan R. Pomalidomide increases immune surface marker expression and immune recognition of oncovirus-infected cells. Oncoimmunology. 2019;8:e1546544. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 36] [Article Influence: 4.5] [Reference Citation Analysis (1)] |
| 48. | Singh S, Banerjee S. Downregulation of HLA-ABC expression through promoter hypermethylation and downmodulation of MIC-A/B surface expression in LMP2A-positive epithelial carcinoma cell lines. Sci Rep. 2020;10:5415. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 20] [Article Influence: 3.3] [Reference Citation Analysis (1)] |
| 49. | Viet NH, Trung NQ, Dong LT, Trung LQ, Espinoza JL. Genetic variants in NKG2D axis and susceptibility to Epstein-Barr virus-induced nasopharyngeal carcinoma. J Cancer Res Clin Oncol. 2021;147:713-723. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 13] [Article Influence: 2.6] [Reference Citation Analysis (1)] |
| 50. | Seo D, Byun H, Cho M, Lee SH, Youn S, Lee J, Jung I, Cho H, Kang H. Dihydrotestosterone Enhances MICA-Mediated Immune Responses to Epstein-Barr Virus-Associated Gastric Carcinoma. Cancers (Basel). 2024;16:3219. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 4] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 51. | Urayama KY, Jarrett RF, Hjalgrim H, Diepstra A, Kamatani Y, Chabrier A, Gaborieau V, Boland A, Nieters A, Becker N, Foretova L, Benavente Y, Maynadié M, Staines A, Shield L, Lake A, Montgomery D, Taylor M, Smedby KE, Amini RM, Adami HO, Glimelius B, Feenstra B, Nolte IM, Visser L, van Imhoff GW, Lightfoot T, Cocco P, Kiemeney L, Vermeulen SH, Holcatova I, Vatten L, Macfarlane GJ, Thomson P, Conway DI, Benhamou S, Agudo A, Healy CM, Overvad K, Tjønneland A, Melin B, Canzian F, Khaw KT, Travis RC, Peeters PH, González CA, Quirós JR, Sánchez MJ, Huerta JM, Ardanaz E, Dorronsoro M, Clavel-Chapelon F, Bueno-de-Mesquita HB, Riboli E, Roman E, Boffetta P, de Sanjosé S, Zelenika D, Melbye M, van den Berg A, Lathrop M, Brennan P, McKay JD. Genome-wide association study of classical Hodgkin lymphoma and Epstein-Barr virus status-defined subgroups. J Natl Cancer Inst. 2012;104:240-253. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 148] [Cited by in RCA: 120] [Article Influence: 8.6] [Reference Citation Analysis (0)] |
| 52. | Thomas M, Wills M, Lehner PJ. Natural killer cell evasion by an E3 ubiquitin ligase from Kaposi's sarcoma-associated herpesvirus. Biochem Soc Trans. 2008;36:459-463. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 29] [Cited by in RCA: 28] [Article Influence: 1.6] [Reference Citation Analysis (1)] |
| 53. | Thomas M, Boname JM, Field S, Nejentsev S, Salio M, Cerundolo V, Wills M, Lehner PJ. Down-regulation of NKG2D and NKp80 ligands by Kaposi's sarcoma-associated herpesvirus K5 protects against NK cell cytotoxicity. Proc Natl Acad Sci U S A. 2008;105:1656-1661. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 147] [Cited by in RCA: 140] [Article Influence: 7.8] [Reference Citation Analysis (1)] |