Copyright: ©Author(s) 2026.
World J Virol. Mar 25, 2026; 15(1): 118988
Published online Mar 25, 2026. doi: 10.5501/wjv.v15.i1.118988
Published online Mar 25, 2026. doi: 10.5501/wjv.v15.i1.118988
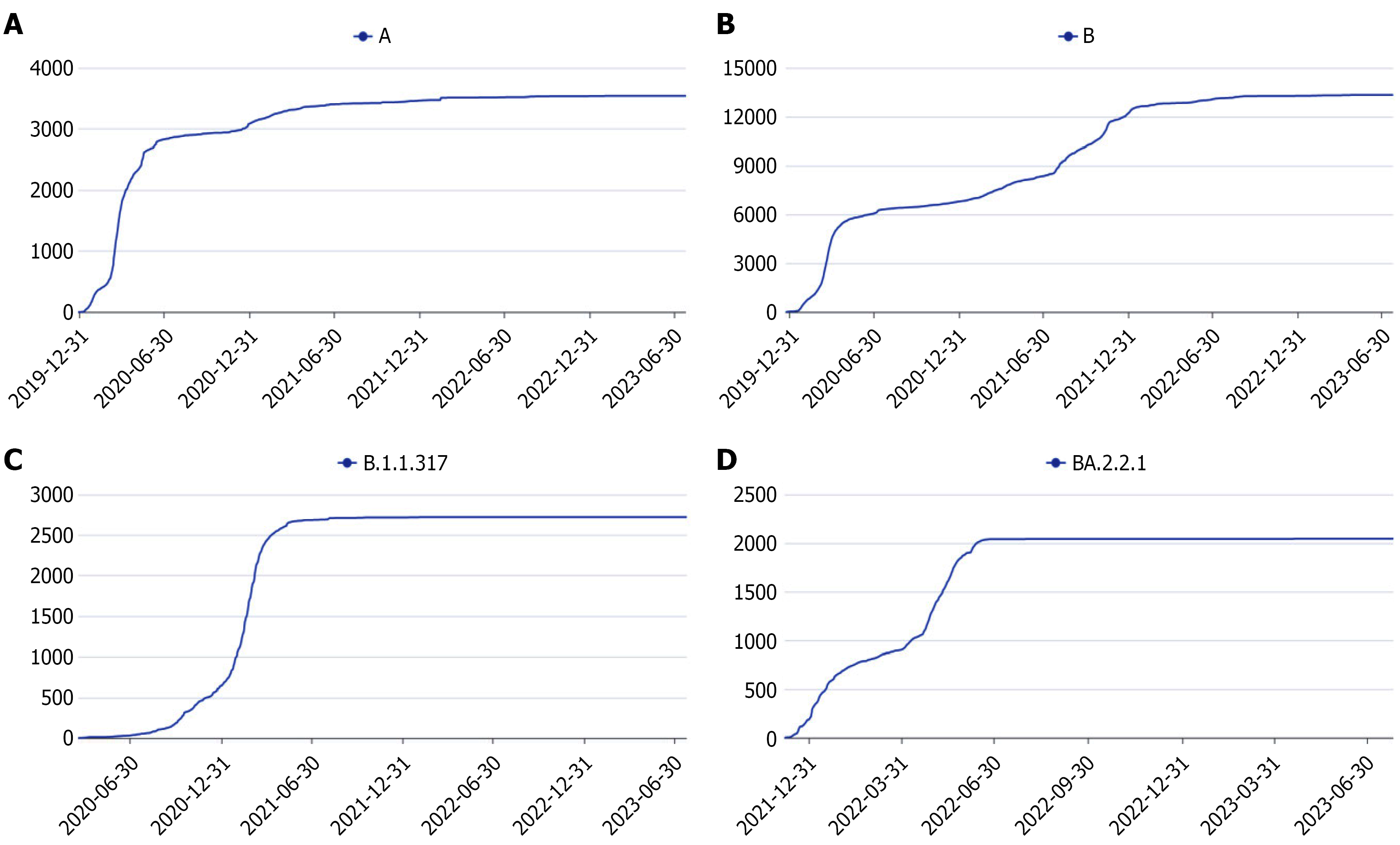
Figure 1 Global submission counts of severe acute respiratory syndrome coronavirus 2 genotypes by time.
A: Type A of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2); B: Type B of SARS-CoV-2; C: Type B.1.1.317 of SARS-CoV-2; D: Type BA.2.2.1 of SARS-CoV-2.
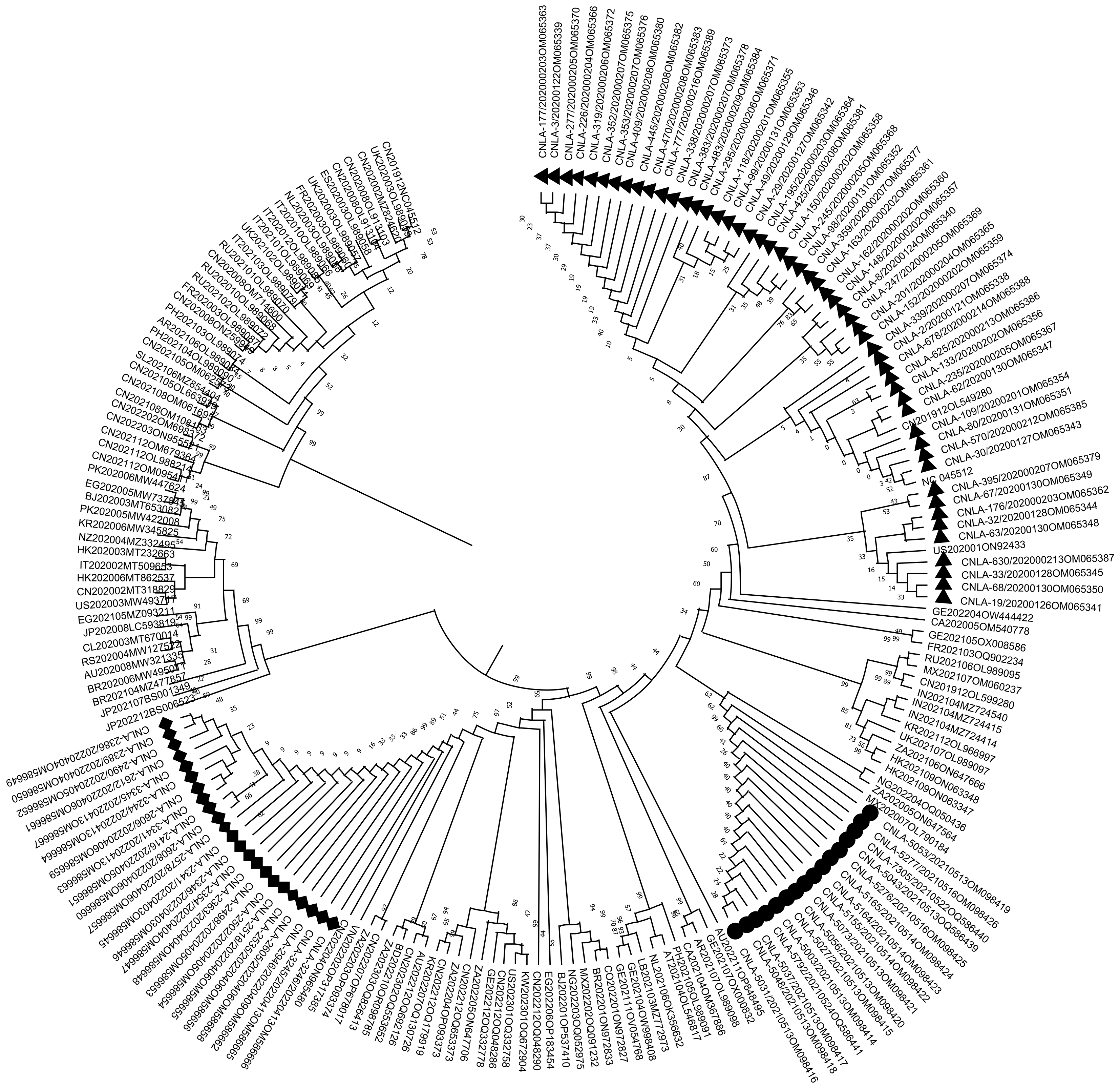
Figure 2 Molecular phylogenetic tree of severe acute respiratory syndrome coronavirus 2 whole genome nucleotide sequences.
The maximum-likelihood phylogenetic tree was constructed based on an alignment of 199 complete genome nucleotide sequences, including local isolates from this study and globally representative reference strains. Branch lengths are scaled to the number of base substitutions per site, and nodal support values from 1000 bootstrap replicates are indicated. The analysis was performed using MEGA X with the maximum composite likelihood model. Key local clusters and their placement relative to major variants (e.g., Delta, Omicron) are highlighted. ▲: 52 sequences of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) whole genome sequences measured in 2020; ●: 16 sequences of SARS-CoV-2 whole genome sequences measured in 2021; ♦: 23 sequences of SARS-CoV-2 whole genome sequences measured in 2022.
- Citation: Chang HW, Yang W, Yu L, Gao DW, Zhang F, Chen ZC, Chen BL, Zhang LM, Zhu R, Zhang Q, Li ZY, Rao JG. Study on the correlation between molecular characteristics of SARS-CoV-2 and its epidemiology and clinical manifestations in Lu’an city. World J Virol 2026; 15(1): 118988
- URL: https://www.wjgnet.com/2220-3249/full/v15/i1/118988.htm
- DOI: https://dx.doi.org/10.5501/wjv.v15.i1.118988