Copyright: ©Author(s) 2026.
World J Virol. Mar 25, 2026; 15(1): 118362
Published online Mar 25, 2026. doi: 10.5501/wjv.v15.i1.118362
Published online Mar 25, 2026. doi: 10.5501/wjv.v15.i1.118362
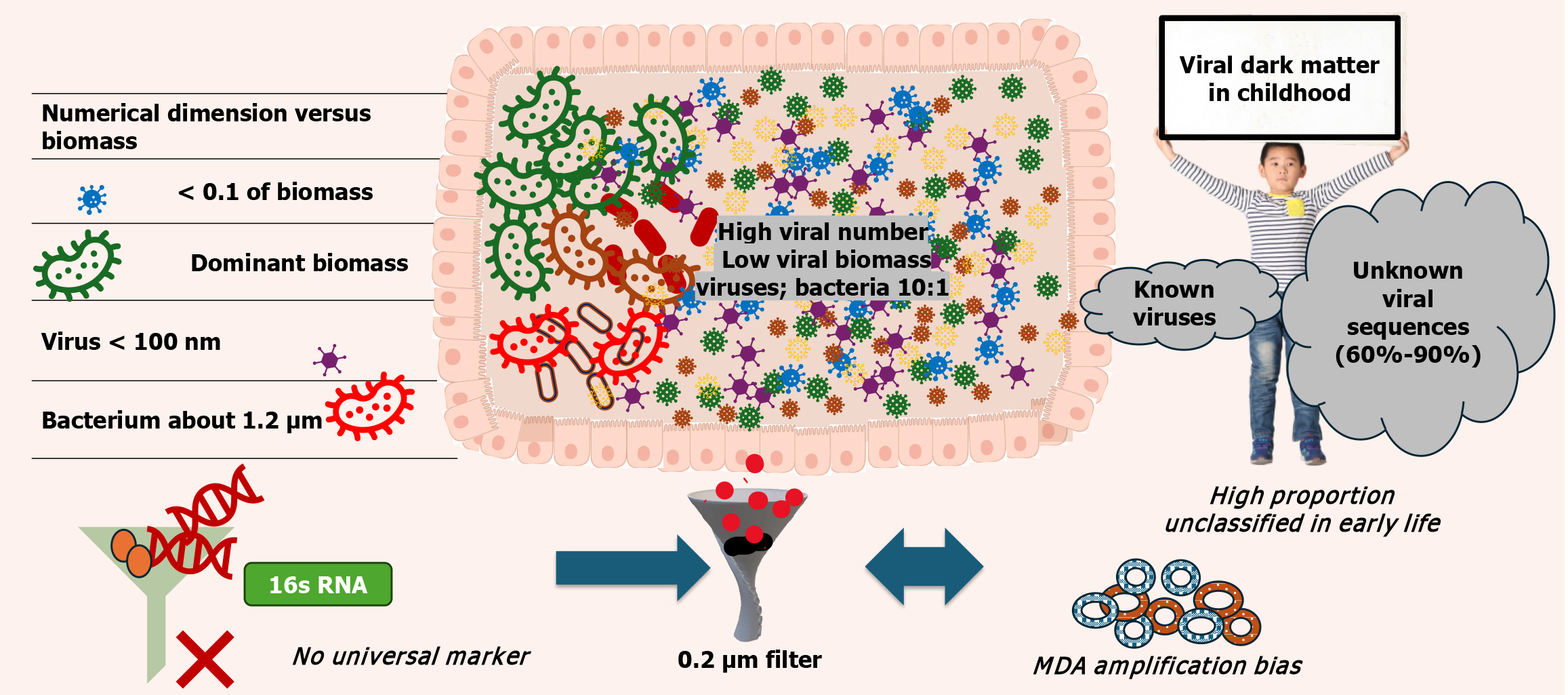
Figure 1 The viral-bacterial paradox in the paediatric gut microbiome.
Despite contributing less than 0.1% of total microbial biomass, viruses-predominantly bacteriophages-numerically dominate the paediatric gut ecosystem, frequently outnumbering bacterial cells by approximately 10:1. Owing to their small size (< 100 nm) and minimal cellular structure, viruses contribute negligibly to overall biomass compared with bacteria (around 1-2 µm), which constitute the bulk of microbial mass. Methodological limitations, including the absence of a universal viral genetic marker, low viral nucleic acid yield, filtration artefacts (e.g., 0.2 µm filters), and amplification biases such as multiple displacement amplification, have historically favored bacterial detection in metagenomic studies. These challenges have resulted in a bacteriocentric view of the gut microbiome and contributed to the underrepresentation of the virome. In paediatric populations, these limitations are further amplified by the highly dynamic nature of early-life viral communities, leading to a large proportion (approximately 60%-90%) of virome sequences remaining unclassified, collectively referred to as viral “dark matter”. Together, these features underscore the numerical dominance, methodological invisibility, and biological significance of the gut virome in early life. MDA: Multiple displacement amplification.
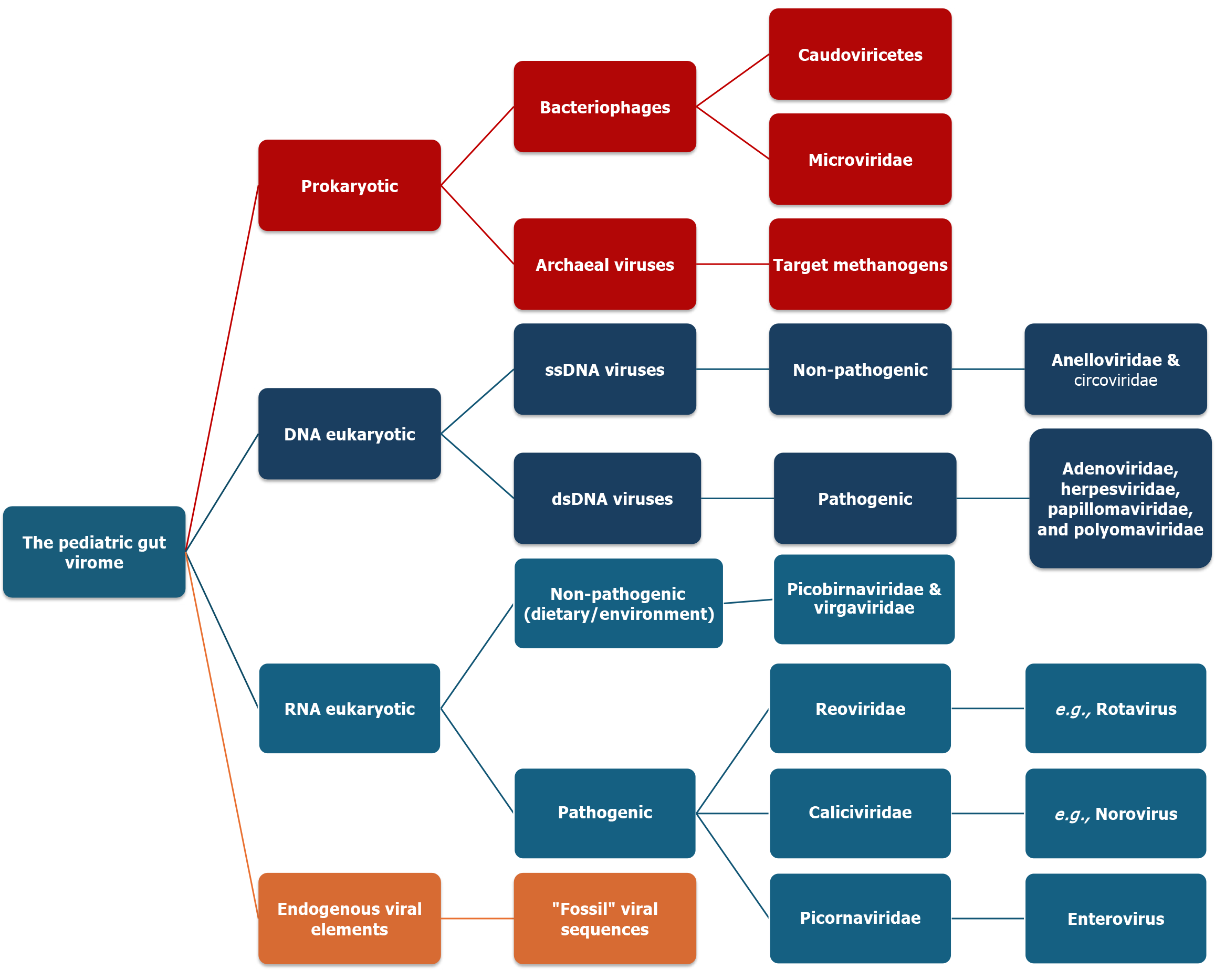
Figure 2 Taxonomy and functional architecture of the paediatric gut virome.
The paediatric gut virome is a multi-kingdom assembly organized into three primary biological domains based on host specificity and genomic integration. Prokaryotic viruses (Bacteriophages and Archaeal viruses): Representing the most abundant fraction in early life, this domain is dominated by the Caudoviricetes (tailed phages) and Microviridae (ssDNA phages). These entities act as “microbial sculptors”, regulating bacterial population densities through lytic predation and facilitating horizontal gene transfer via lysogeny, which drives the evolution of the developing bacteriome. Eukaryotic viruses: This domain comprises both resident host-infecting viruses and transient environmental entities. Host-infecting viruses, such as the Anelloviridae (e.g., torque teno virus), serve as indicators of immune status and are involved in early-life immune priming. Transient viruses, often derived from the diet (e.g., plant-infecting Virgaviridae), represent the interface between environmental exposure and the gut ecosystem. Endogenous viral elements (EVEs): These consist of ancestral viral sequences, including endogenous retroviruses, that are stably integrated into the host genome. In the paediatric context, EVEs are increasingly recognized as active modulators of the innate immune system, contributing to the “molecular memory” that protects the infant against exogenous viral challenges. The hierarchical structure highlights the transition from a phage-rich environment in the neonate to a more diverse, eukaryotic-inclusive ecosystem as the child matures.
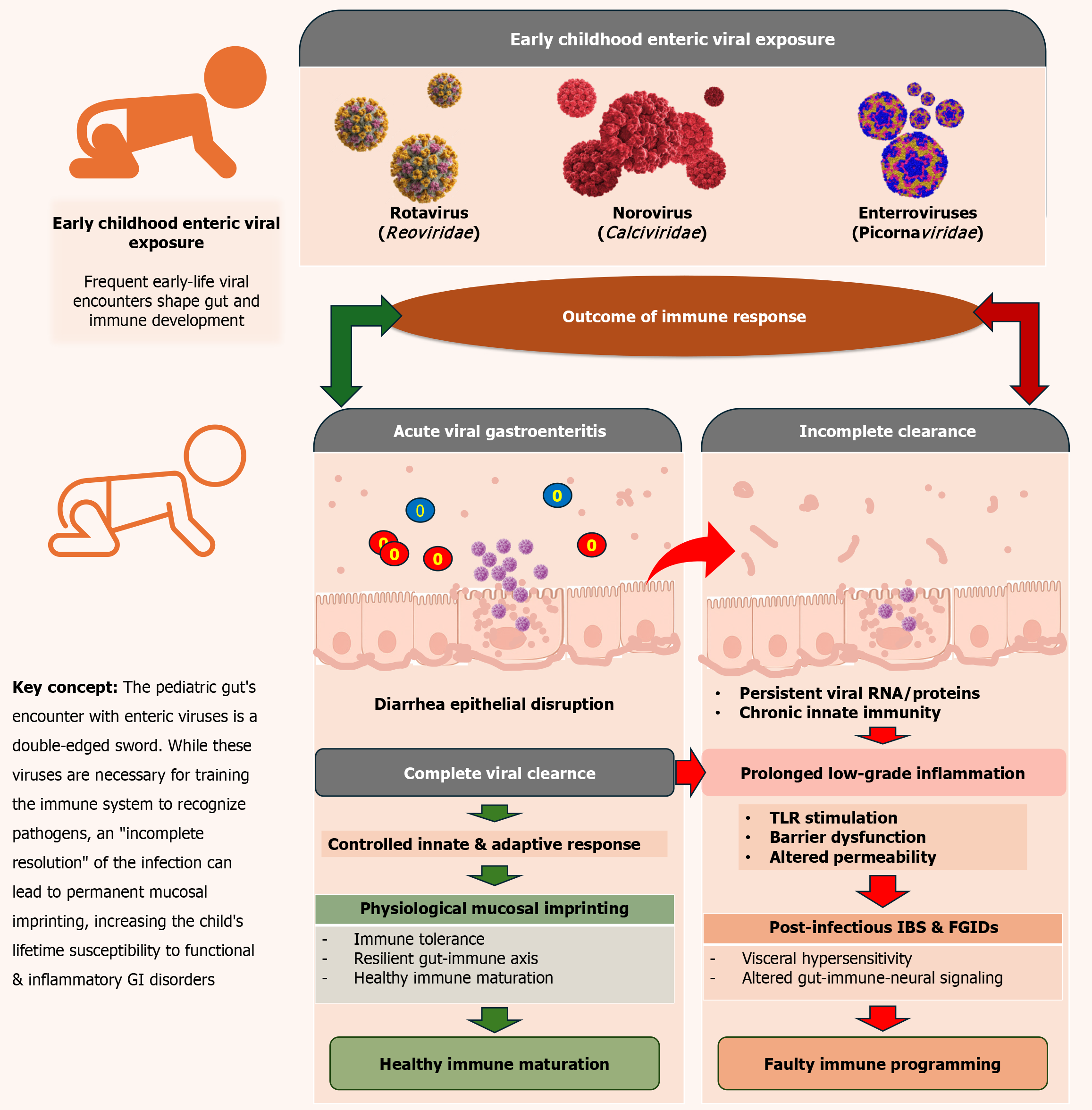
Figure 3 Infectious and post-infectious viral pathways in the paediatric gut.
This schematic illustrates the dual role of early-life enteric viral infections in shaping paediatric gut and immune development. Frequent exposure to common enteric viruses during infancy-such as rotavirus (Reoviridae), norovirus (Caliciviridae), and enteroviruses (Picornaviridae)-often results in acute viral gastroenteritis and transient epithelial disruption. Following infection, immune outcomes diverge depending on the efficiency of viral clearance. Effective immune resolution leads to restoration of epithelial integrity, controlled innate and adaptive immune responses, and physiological mucosal imprinting that supports immune tolerance and healthy immune maturation. In contrast, incomplete viral clearance may result in persistence of viral nucleic acids or proteins within the intestinal mucosa, driving prolonged low-grade inflammation through sustained innate immune activation, barrier dysfunction, and altered intestinal permeability. This maladaptive immune trajectory disrupts gut-immune-neural signaling and promotes visceral hypersensitivity, increasing susceptibility to post-infectious irritable bowel syndrome and other functional gastrointestinal disorders. Overall, the figure highlights the concept that early-life enteric viral infections represent a double-edged sword-essential for immune training, yet capable of inducing long-lasting mucosal imprinting when immune resolution is incomplete, with implications for lifelong gastrointestinal health. IBS: Irritable bowel syndrome; FGID: Functional gastrointestinal disorders; GI: Gastrointestinal.
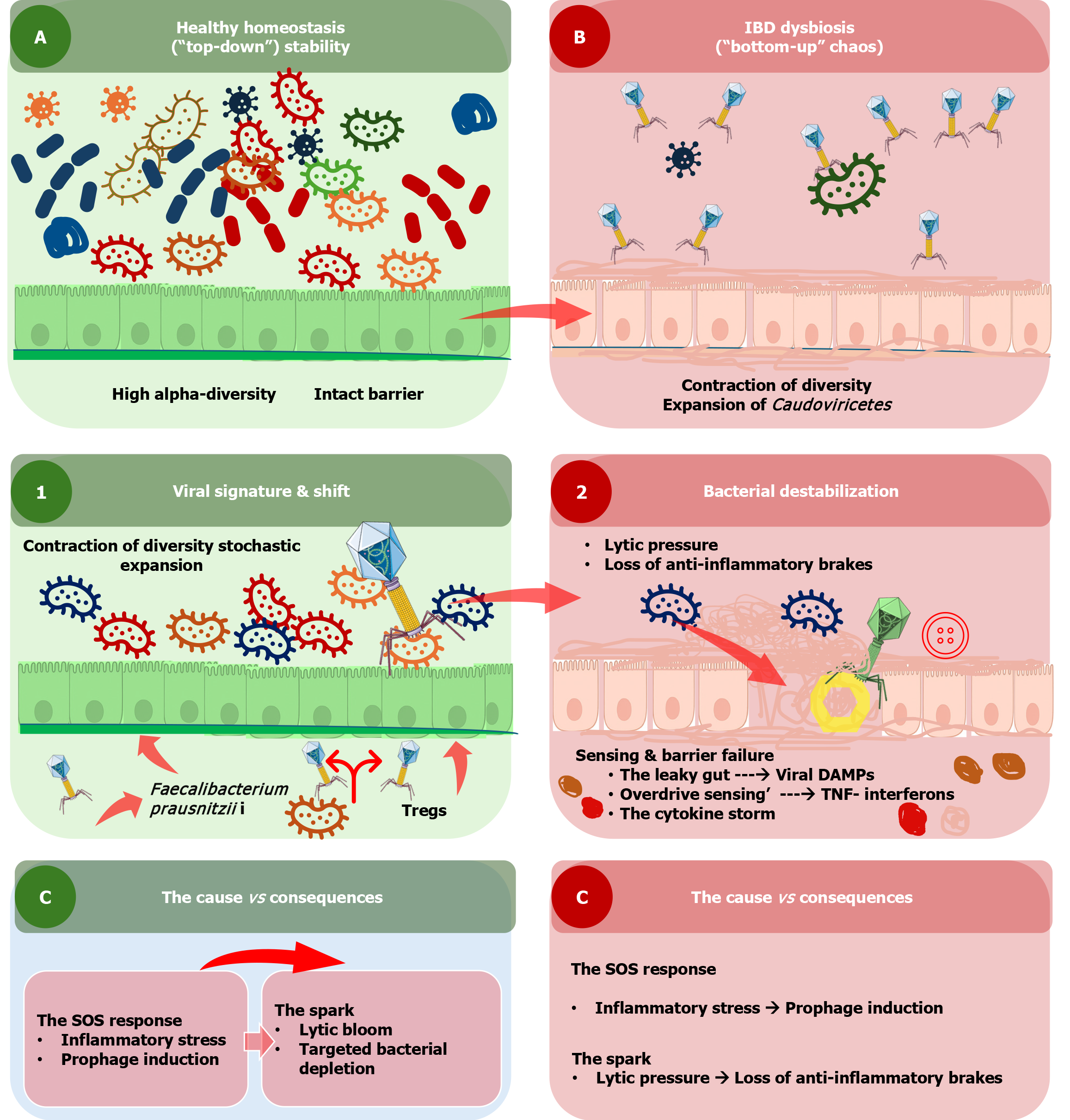
Figure 4 The virome-driven inflammatory cascade in paediatric inflammatory bowel disease.
This schematic illustrates the proposed role of the intestinal virome in maintaining gut homeostasis and driving inflammatory cascades in paediatric inflammatory bowel disease. A: Healthy homeostasis; B: Inflammatory bowel disease dysbiosis; C: The cause vs consequences. IBD: Inflammatory bowel disease.
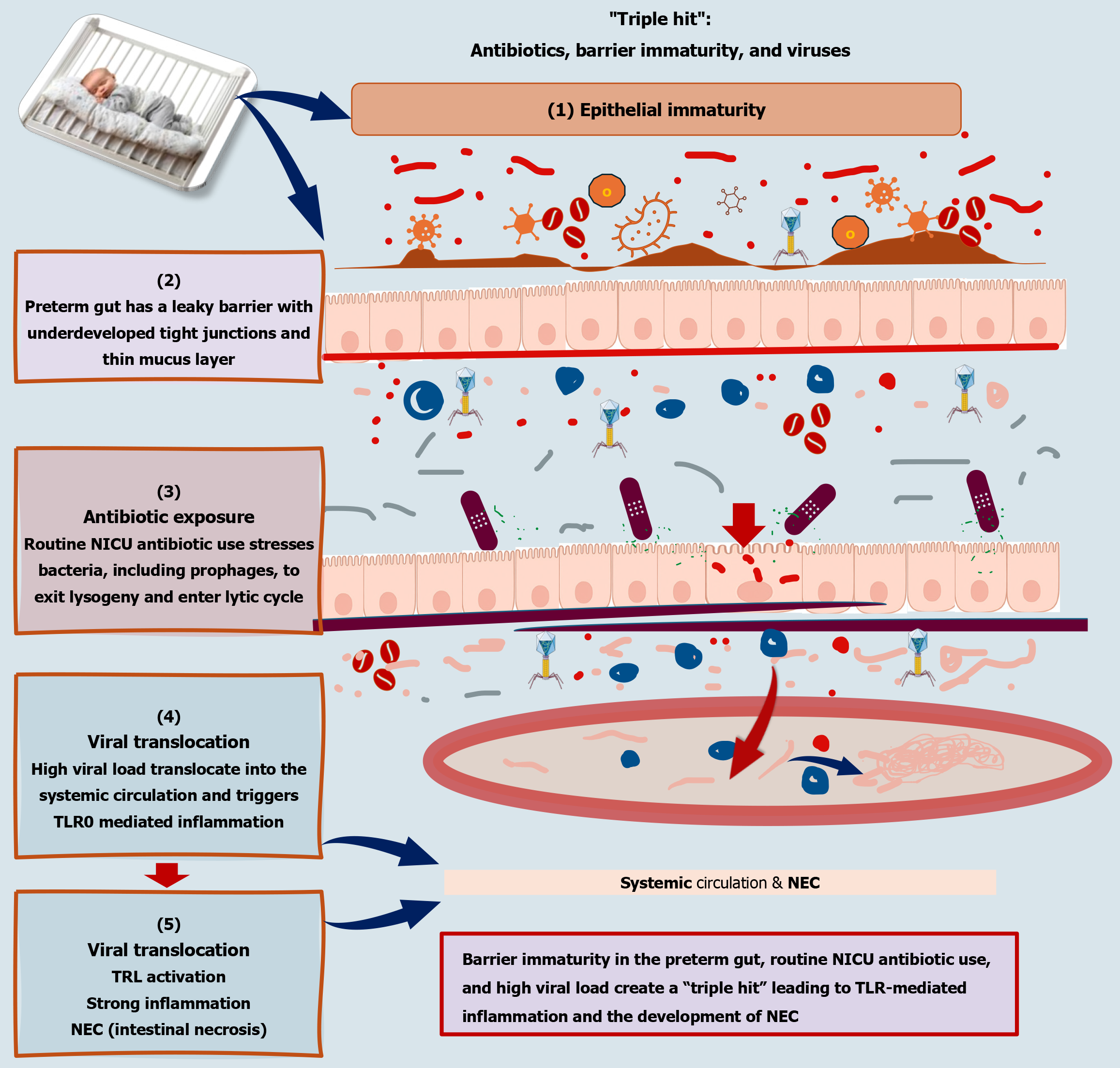
Figure 5 The “triple hit” model of virome-driven necrotizing enterocolitis in preterm infants.
This schematic illustrates the proposed “triple hit” model linking the paediatric gut virome to the pathogenesis of necrotizing enterocolitis (NEC) in preterm infants. The first hit is epithelial immaturity, characterized by an underdeveloped intestinal barrier with defective tight junctions and a thin mucus layer, predisposing the preterm gut to increased permeability. The second hit is antibiotic exposure, commonly encountered in the neonatal intensive care unit, which disrupts the bacterial microbiome and induces prophage activation through bacterial stress responses, leading to bacteriophage lytic cycles and viral blooms. The combination of barrier immaturity and elevated viral burden facilitates the third hit, viral translocation, whereby bacteriophages and viral nucleic acids cross the intestinal barrier into the lamina propria and systemic circulation. This translocation triggers Toll-like receptor-mediated inflammatory cascades, amplifying mucosal and systemic inflammation and ultimately contributing to intestinal injury and necrosis. Collectively, the figure highlights how converging developmental, microbial, and iatrogenic factors interact to drive exaggerated inflammatory responses in NEC. NEC: Necrotizing enterocolitis; NICU: Neonatal intensive care unit; TLR: Toll-like receptors.
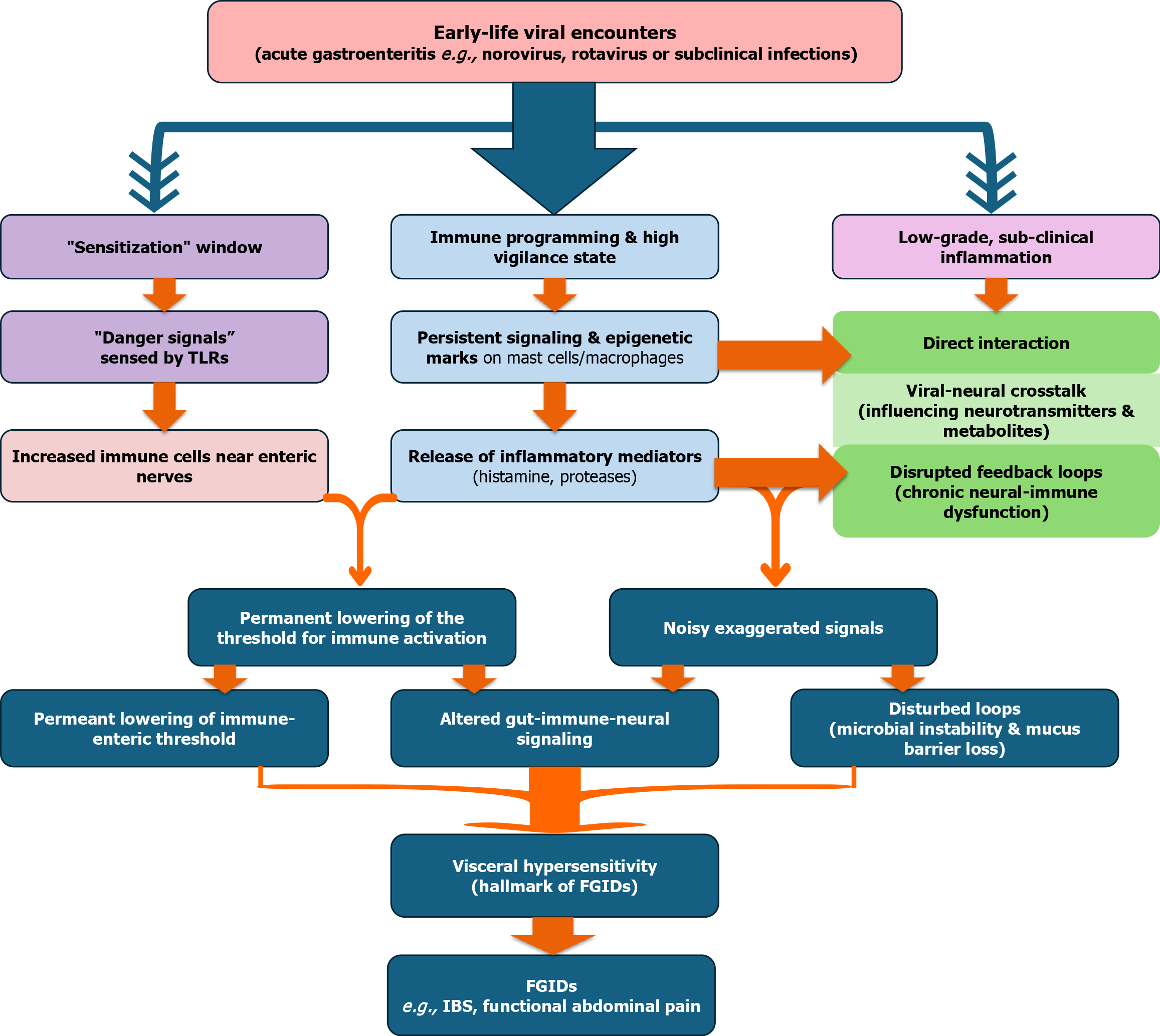
Figure 6 The developmental pathway of functional gastrointestinal disorders via virome-mediated mechanisms.
This flowchart illustrates how early-life viral encounters can lead to chronic functional gastrointestinal disorders (FGIDs) through three interconnected pathways. Immune programming: Viral infections create a “Sensitization” window, leading to persistent signaling via toll-like receptors and epigenetic changes in immune cells, permanently lowering the threshold for immune activation. Low-grade inflammation: Acute insults leave a “molecular hangover” of sub-clinical inflammation. Inflammatory mediators like histamine directly sensitize nociceptors, resulting in peripheral sensitization where the gut becomes hyper-reactive. Altered signaling: Virome disturbances disrupt the gut-immune-neural axis. This involves direct viral-neural crosstalk and disrupted feedback loops due to microbial instability, leading to “noisy” and exaggerated signaling. All three pathways converge to cause visceral hypersensitivity, the hallmark of FGIDs. In a subset of children, a failure to “reset” this axis after the initial infection results in a chronic functional pain disorder, such as post-infectious irritable bowel syndrome. FGID: Functional gastrointestinal disorders; IBS: Irritable bowel syndrome; TLR: Toll-like receptors.
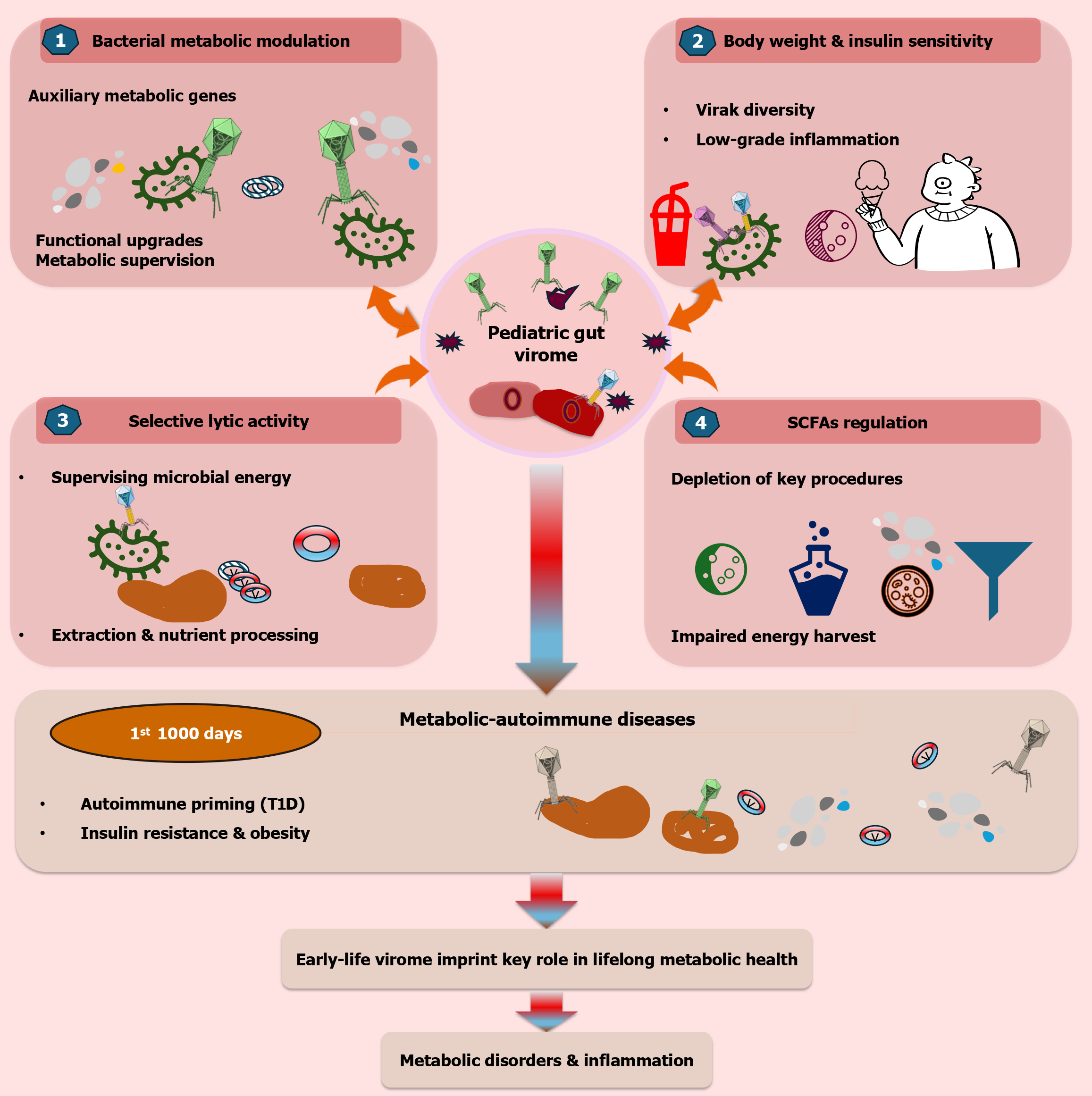
Figure 7 Virome-metabolism-endocrine axis in paediatric health and disease.
This schematic summarizes the central role of the paediatric gut virome in shaping host metabolic and endocrine outcomes across early life. The gut virome-comprising bacteriophages and eukaryotic viruses-acts as a regulatory hub that influences host physiology through multiple interconnected pathways. Bacteriophages modulate bacterial metabolic capacity via horizontal gene transfer of auxiliary metabolic genes and through selective lytic activity, thereby supervising microbial energy extraction and nutrient processing. These virome-driven shifts alter the production of key microbial metabolites, particularly short-chain fatty acids, which are critical for intestinal barrier integrity, immune regulation, and systemic energy homeostasis. Perturbations in early-life virome diversity and composition are associated with low-grade inflammation, altered insulin sensitivity, and changes in body weight regulation. During critical developmental windows-particularly the first 1000 days of life-these effects may contribute to long-term metabolic imprinting, increasing susceptibility to obesity, insulin resistance, and metabolic-autoimmune diseases such as type 1 diabetes. Collectively, the figure highlights the paediatric gut virome as a pivotal determinant linking microbial ecology to lifelong metabolic and endocrine health. T1D: Type 1 diabetes; SCFA: Short-chain fatty acids.
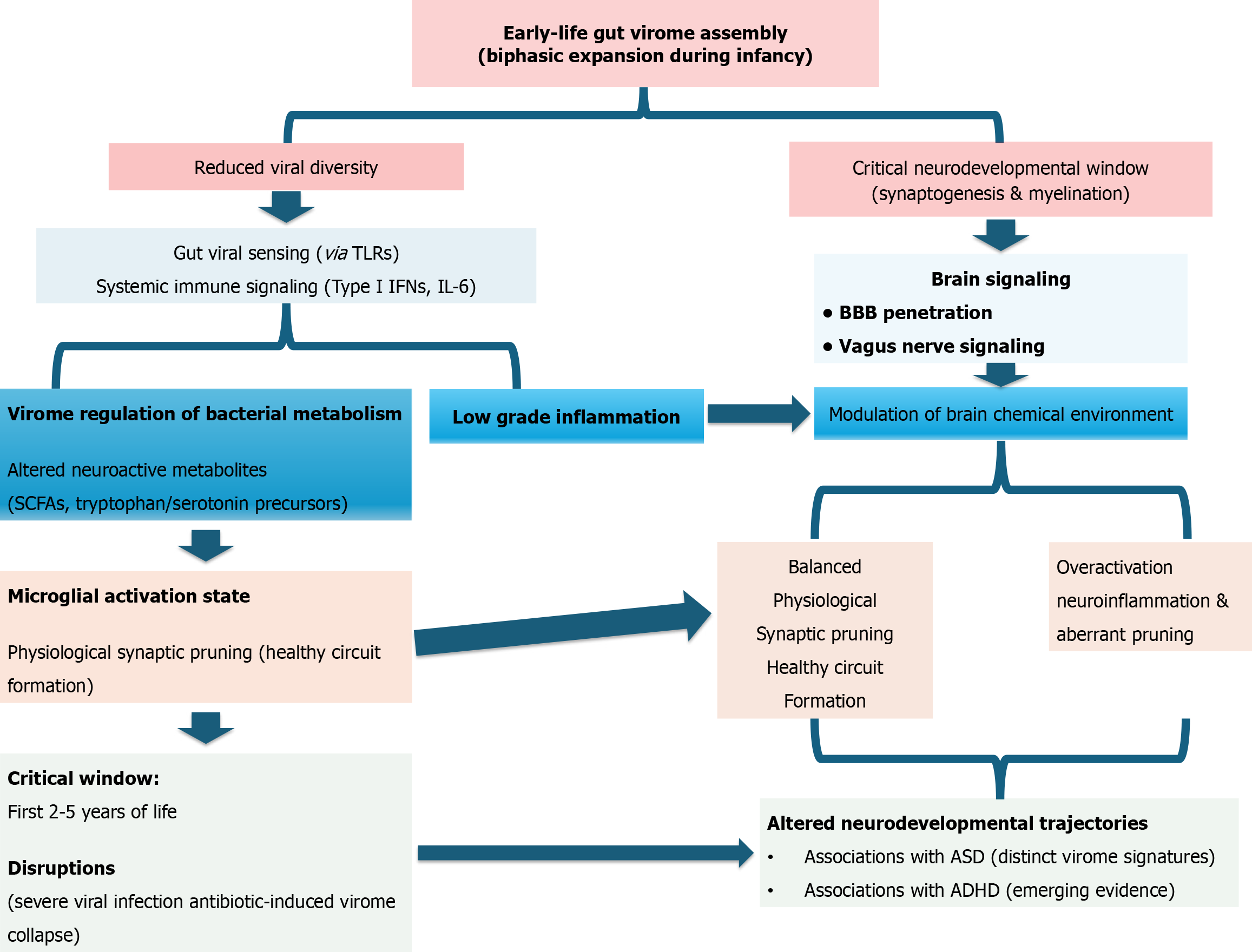
Figure 8 The virome-immune-brain axis in early neurodevelopment.
This single-panel schematic illustrates the proposed role of the gut virome in shaping neurodevelopment through immune and metabolic signaling pathways. During early life, the expanding gut virome interacts with host immune sensors (e.g., toll-like receptors), triggering controlled cytokine responses and type I interferon signaling that can influence the brain directly via the blood-brain barrier or indirectly through vagal pathways. In parallel, bacteriophages modulate bacterial communities and their metabolic outputs, including short-chain fatty acids and neurotransmitter precursors (e.g., tryptophan), thereby altering the neurochemical milieu. These virome-driven immune and metabolic signals regulate microglial priming and synaptic pruning, processes critical for normal brain maturation. Disruption of virome balance during critical developmental windows (approximately the first 2-5 years of life) may lead to excessive neuroinflammation, aberrant synaptic remodeling, and altered neural circuit formation. Emerging evidence links such virome alterations with neurodevelopmental conditions such as autism spectrum disorder and attention-deficit/hyperactivity disorder, although current data support association rather than causation. ASD: Autism spectrum disorder; ADHD; Attention-deficit/hyperactivity disorder; SCFA: Short-chain fatty acids; IL: Interleukin; IFN: Interferon; TLR: Toll-like receptors.
- Citation: Saeed NK, Elbeltagi YM, Al-Beltagi M. Unveiling the viral dimension: The paediatric gut virome as a key modulator of gastrointestinal metabolic, and neurodevelopmental health. World J Virol 2026; 15(1): 118362
- URL: https://www.wjgnet.com/2220-3249/full/v15/i1/118362.htm
- DOI: https://dx.doi.org/10.5501/wjv.v15.i1.118362