Published online Mar 20, 2026. doi: 10.5493/wjem.v16.i1.116383
Revised: December 23, 2025
Accepted: February 11, 2026
Published online: March 20, 2026
Processing time: 125 Days and 4.8 Hours
N6-methyladenosine (m6A) is the most prevalent internal RNA modification in eukaryotic transcripts, and YTH structural domain family (YTHDF) 2 is a principal m6A reader that governs RNA stability and turnover. Accumulating evidence indicates that YTHDF2 exerts context-dependent and sometimes opposing functions across major neurological disorders, including Alzheimer’s disease, Parkinson’s disease, glioblastoma, epilepsy, and experimentally induced cognitive and neuropsychiatric conditions. By selectively promoting decay of m6A-marked transcripts - such as leucine rich repeat and immunoglobulin domain containing 2, axis inhibition protein 1, breast cancer type 1 susceptibility protein associated protein 1, mitogen-activated protein kinase kinase 4, family with sequence similarity 134 member B, and NOD-like receptor family pyrin domain-containing 3 - YTHDF2 modulates diverse processes including amyloid processing, Wnt signaling, ferroptosis, neuroinflammation, metabolic homeo
Core Tip: This mini-review synthesizes evidence that the N6-methyladenosine (m6A) reader YTH structural domain family (YTHDF) 2 exerts context-dependent, often bidirectional effects in neurodegeneration and glioma by directing degradation of m6A-marked transcripts. YTHDF2 is regulated by upstream axes and interacts functionally with m6A writers/erasers. Because its net effect depends on cell type, brain region, disease stage and target dosage, YTHDF2 constitutes both a mechanistic biomarker and a selectively tractable therapeutic target; advancing translation will require cell/region-resolved targetome mapping, longitudinal dynamics, and testing of conditional genetics, small molecules and antisense oligonucleotides.
- Citation: Liu YM, Liu M, Yuan H, Li B. Double-edged role of N6-methyladenosine reader YTH structural domain family protein 2 in neurological disorders: Molecular mechanisms and translational prospects. World J Exp Med 2026; 16(1): 116383
- URL: https://www.wjgnet.com/2220-315x/full/v16/i1/116383.htm
- DOI: https://dx.doi.org/10.5493/wjem.v16.i1.116383
YTH structural domain family (YTHDF) 2, a member of the YTH-domain protein family, functions as a principal N6-methyladenosine (m6A) “reader” that specifically recognizes m6A-modified mRNAs via its C-terminal YTH domain[1-3]. Predominantly cytoplasmic and enriched at the nuclear periphery, YTHDF2’s subcellular localization is tightly linked to mRNA metabolism[2]. Its expression and stability are regulated at multiple levels; for example, epidermal growth factor receptor (EGFR)/SRC proto-oncogene, non-receptor tyrosine kinase (SRC)/extracellular signal-regulated kinase (ERK) signaling phosphorylates YTHDF2 on serine 39 and threonine 381, markedly increasing protein stability[1]. Mechanistically, YTHDF2 governs gene expression by promoting decay of m6A-marked transcripts: It recruits deadenylase complexes (e.g., carbon catabolite repression 4-negative on TATA-less complex) or ribonuclease assemblies (e.g., ri
YTHDF2 influences multiple signaling and metabolic pathways[5]. In the Wnt/β-catenin cascade, YTHDF2 binds m6A sites on adenomatous polyposis coli and glycogen synthase kinase 3 beta mRNAs, promoting their degradation and thereby activating Wnt signaling; this mechanism enhances glioblastoma cell migration and invasion in model systems[6]. YTHDF2 also targets mitogen-activated protein kinase kinase 4 (MAP2K4) mRNA for decay, which attenuates the stress-activated protein kinase kinase 1-c-Jun N-terminal kinase-c-Jun Proto-oncogene pro-inflammatory axis and reduces manganese-induced astrocytic inflammation[2]. At the metabolic level, YTHDF2 promotes degradation of liver X receptor alpha and human immunodeficiency virus type I enhancer binding protein 2 transcripts, perturbing liver X receptor α-dependent cholesterol homeostasis[1]. During central nervous system development and neural stem-cell differentiation, YTHDF2 is essential: Its loss disrupts progenitor proliferation and differentiation[1].
Emerging evidence links YTHDF2 to neurodegenerative and neurological disorders[7]. In Parkinson’s disease (PD) models, YTHDF2 regulates breast cancer type 1 susceptibility protein associated protein-1 (BAP1) mRNA stability and thereby modulates ferroptosis neuronal death[8]. In epilepsy models, YTHDF2 controls nuclear receptor subfamily 4 group a member 2 (NR4A2) mRNA stability[4]. In other neuropathic contexts, YTHDF2 cooperates with methyltransferase like (METTL) 3 to influence pri-miR-150 processing and maturation[9]. Collectively, by recognizing and promoting degradation of diverse m6A-marked transcripts involved in development, metabolism, inflammation, and cell survival, YTHDF2 serves as a multifaceted regulator of neuronal homeostasis and disease pathogenesis.
Several recent reviews have summarized the biological functions of the m6A reader YTHDF2 and its involvement in central nervous system disorders. Notably, a comprehensive review published in Biochemical Pharmacology provided an important overview of YTHDF2-associated regulatory mechanisms and therapeutic potential across central nervous system diseases[7]. Building on this foundation, the present mini-review offers complementary insights by adopting a disease-context-anchored framework, integrating recent primary studies published in 2024-2025, and proposing a dose-target dependency model to reconcile the bidirectional effects of YTHDF2 observed across different cell types, brain regions, and disease stages. By focusing on representative neurological disorders and induced models, this review aims to highlight mechanistic convergence and divergence and to outline actionable strategies for precision modulation of the m6A-YTHDF2 axis.
Reports on YTHDF2 in Alzheimer’s disease (AD) are inconsistent, likely reflecting differences in cell type, disease stage, and regional brain context. In one set of findings, METTL3 expression and global m6A levels are increased in AD brain and mouse models[9]. Enhanced m6A on leucine-rich repeat and leucine rich repeat and immunoglobulin domain containing 2 (LINGO2) mRNA promotes YTHDF2 binding and degradation of LINGO2 transcripts, reducing LINGO2 protein. Loss of LINGO2 weakens its interaction with beta-secretase 1, thereby favoring amyloid precursor protein-beta-secretase 1 cleavage, increasing soluble amyloid precursor protein β and amyloid-β production, and accelerating amyloid-β deposition, neuroinflammation and cognitive decline in that model[9].
By contrast, machine-learning analyses implicate YTHDF2 together with other m6A regulators (e.g., fat mass and obesity-associated protein [FTO], leucine rich pentatricopeptide repeat containing) in Notch pathway modulation[10]. Those studies report reduced FTO, increased neurogenic locus notch homolog protein 1 (Notch1) m6A, and heightened Notch1 pathway activity with consequent tumor necrosis factor alpha and interleukin (IL)-1β release from dendritic cells after FTO inhibition. Combining these observations with reported YTHDF2 downregulation suggests a plausible scenario in which loss of YTHDF2-mediated repression could de-repress Notch1 or other pro-inflammatory transcripts, thereby amplifying innate immune responses in certain contexts[10] - a hypothesis that requires direct experimental testing.
Multi-region analyses further emphasize spatial heterogeneity: Hippocampal YTHDF2 expression is negatively correlated with Braak stage, implying that reduced YTHDF2 in hippocampus may promote AD progression by dysregulating genes linked to synaptic plasticity, whereas YTHDF2 upregulation in regions such as the entorhinal cortex associates with immune-regulatory imbalance, consistent with region-specific roles in inflammation[11]. Moreover, although not yet studied directly in AD, evidence that METTL3 and YTHDF2 cooperatively regulate pri-miR-150 processing indicates that YTHDF2 can affect noncoding RNA biology and thereby exert context-dependent effects on disease-relevant pathways[12].
In sum, current data indicate that YTHDF2 can both accelerate AD pathology (for example, via YTHDF2-dependent LINGO2 degradation and consequent amyloid-β accumulation) and, when downregulated, permit increased expression of pro-inflammatory effectors (for example, Notch1-linked signaling). The net effect of YTHDF2 in AD thus depends on cell type, regional expression, disease stage, and the balance of vulnerable vs protective target transcripts. Resolving these complexities will require cell- and region-specific manipulations (conditional knockout/overexpression), combined with transcriptome-wide m6A mapping and functional assays of cognition and neuroinflammation.
PD is pathologically defined by loss of dopaminergic neurons in the substantia nigra and accumulation of α-synuclein[13]. Recent studies indicate that m6A RNA modification and its reader YTHDF2 participate at multiple levels in PD-relevant processes, including neuronal survival, ferroptosis, and neuroinflammation. First, YTHDF2 can promote Wnt/β-catenin signaling by targeting the negative regulator axis inhibition protein 1 (AXIN1): YTHDF2 recognizes m6A sites on AXIN1 mRNA and accelerates its degradation, thereby lowering AXIN1 protein, restoring β-catenin activity, and upregulating downstream pro-survival genes such as cyclin D1 and c-Myc[14,15]. In PD mouse models, YTHDF2 downregulation is associated with increased AXIN1 expression and Wnt suppression, which promotes neuronal apoptosis; conversely, YTHDF2 overexpression is predicted to be neuroprotective, although direct in vivo validation remains limited.
Second, YTHDF2 influences ferroptosis via regulation of BAP1 mRNA stability: In 1-methyl-4-phenylpyridinium-treated cellular and murine PD models, FTO upregulation increases BAP1 expression, while YTHDF2 normally limits BAP1 by degrading its m6A -modified transcript. Loss of YTHDF2 therefore stabilizes BAP1 mRNA, activates the p53-solute carrier family 7 member 11 axis, and promotes neuronal ferroptosis, exacerbating neurodegeneration[8].
Third, YTHDF2 modulates neuroinflammation: Manganese exposure markedly reduces YTHDF2 Levels, which impairs degradation of MAP2K4 mRNA, causing MAP2K4 accumulation and activation of the c-Jun N-terminal kinase-c-JUN pro-inflammatory cascade with increased IL-1β and tumor necrosis factor alpha release[2]. This finding is consistent with a model in which YTHDF2 provides negative feedback on inflammatory signaling.
Finally, a METTL14’-m6A-YTHDF2 axis may regulate α-synuclein homeostasis: Several in vitro and in vivo observations report reduced m6A marking of α-synuclein mRNA in PD substantia nigra accompanied by increased transcript and protein levels; METTL14 overexpression promotes m6A modification and, via a YTHDF2-dependent mechanism, enhances α-synuclein mRNA degradation, suggesting this axis helps limit pathological protein accumulation[16].
Collectively, these data portray YTHDF2 as a multifunctional regulator in PD: It can support neuroprotection by promoting Wnt signaling and restraining ferroptosis and inflammation, whereas YTHDF2 deficiency - whether induced by environmental toxins such as manganese or other insults - may disrupt these protective programs and accelerate neurodegeneration. Future studies should map YTHDF2 targets and dynamics across cell types (neurons, astrocytes, microglia) and define its interplay with m6A writers/erasers (METTL14, FTO) to evaluate the therapeutic potential of modulating the YTHDF2 axis in PD.
Glioblastoma is an aggressive brain tumor with very poor prognosis[17]. Accumulating evidence indicates that m6A RNA modification and its reader YTHDF2 play important roles in glioblastoma metabolism and progression, and may represent therapeutic targets. Several studies report oncogenic functions for YTHDF2 in glioblastoma. Fang et al[3] found that YTHDF2 protein is markedly upregulated in glioblastoma tissue relative to adjacent non-tumor brain and is enriched in glioblastoma-derived cancer stem cells (GSCs); YTHDF2 knockdown suppresses proliferation in glioma cell lines, supporting a pro-tumorigenic role[1].
Mechanistically, the EGFR-SRC-ERK pathway stabilizes YTHDF2: Phosphorylation of YTHDF2 at serine 39 and threonine 381 prolongs its half-life and increases protein abundance. Elevated YTHDF2 in turn promotes degradation of m6A-marked transcripts such as liver X receptor alpha and human immunodeficiency virus type I enhancer binding protein 2, thereby disrupting liver X receptor α-dependent cholesterol homeostasis, causing intracellular cholesterol accumulation and adversely affecting patient outcome[1]. Upstream transcriptional control also contributes: Protein arginine methyltransferase 6 (PRMT6) expression rises with glioma grade, and cyclin dependent kinase 9 (CDK9) recruits PRMT6 to the YTHDF2 promoter, forming a PRMT6-CDK9-YTHDF2 activation axis[6].
Downstream, YTHDF2 binds and destabilizes adenomatous polyposis coli and glycogen synthase kinase 3 beta mRNAs, leading to activation of Wnt/β-catenin signaling[6]. Wnt activation is tightly linked to enhanced glioblastoma cell migration, invasion, epithelial-mesenchymal transition, and overall malignancy. Immunogenomic analyses further show that YTHDF2-high tumors display increased infiltration of multiple immune cell types (B cells, CD8+ T cells, macrophages) and upregulation of immune-checkpoint molecules including programmed cell death 1 ligand 1 and cytotoxic T-lymphocyte associated protein 4, suggesting a role for YTHDF2 in modulating the tumor immune microenvironment and promoting immune evasion[18].
In summary, YTHDF2 appears to drive glioblastoma progression by recognizing and degrading m6A-marked transcripts that regulate cholesterol homeostasis, Wnt signaling, and immune regulation[1,6,18]. Dysregulation of YTHDF2 perturbs cellular homeostasis, enhances invasiveness, and reshapes the microenvironment. Future work should dissect YTHDF2’s subtype-specific and microenvironment-specific functions in glioblastoma and evaluate whether targeting YTHDF2 itself or its upstream activation pathways (e.g., EGFR-SRC-ERK, PRMT6-CDK9) or the broader m6A regulatory network offers therapeutic benefit.
Seizure activity substantially perturbs neuronal autophagy, oxidative stress responses, and apoptosis. Recent evidence suggests that the m6A reader YTHDF2 may contribute to epileptic pathology by regulating the autophagy-related gene family with sequence similarity 134 member B (FAM134B). Cai et al[19] showed that YTHDF2 binds m6A sites on FAM134B mRNA and promotes its degradation in porcine adipogenesis, reducing FAM134B expression[19]. Complementary work by Xie et al[20] in epilepsy models demonstrated that FAM134B overexpression enhances neuronal autophagy, reduces apoptosis, and increases endoplasmic-reticulum Ca2+ stores to alleviate endoplasmic reticulum stress and neuronal injury. By analogy, downregulation of YTHDF2 could elevate FAM134B levels and thereby augment protective autophagy - an attractive hypothesis that requires direct in vitro and in vivo validation.
Moreover, cross-talk between the m6A “eraser” FTO and YTHDF2 appears relevant to seizure-related pathways: Traumatic brain injury-associated reductions in FTO increase m6A on NR4A2 mRNA, promoting YTHDF2-mediated degradation and disease progression[4]; similarly, FTO downregulation can raise m6A on nuclear factor erythroid-2-related factor 2 (Nrf2) mRNA, rendering it more susceptible to YTHDF2-dependent decay and weakening neuronal antioxidant defenses[21]. Collectively, these data implicate an m6A-YTHDF2 regulatory axis - acting on FAM134B, NR4A2, Nrf2, and likely other transcripts - that modulates autophagy, antioxidative responses, and neuronal survival in epileptogenesis. Future work should test this model using YTHDF2 knockout/knockdown, target-mRNA stability assays, and behavioral phenotyping to assess therapeutic potential.
Several animal studies have implicated YTHDF2 in experimentally induced cognitive impairment and other neuropsychiatric phenotypes. For example, in a rat model of high homocysteine (Hcy)-induced cognitive dysfunction, YTHDF2 reduces NLRP3 mRNA stability in an m6A-dependent manner, thereby suppressing microglial pyroptosis and neuroinflammation and improving cognitive performance[22]. Concretely, Hcy exposure decreased YTHDF2 expression in the hippocampus and in human microglia clone 3 microglial cells - an observation consistent with hippocampal datasets from AD patients. Treatment with betaine upregulated YTHDF2 and significantly ameliorated deficits in Y-maze and novel-object-recognition assays[22]. Mechanistically, YTHDF2 knockdown fully reversed betaine’s inhibitory effect on the NLRP3-caspase-1-GSDMD pyroptotic cascade: In YTHDF2-silenced microglia, betaine failed to suppress pathway activation, resulting in restored secretion of IL-1β and IL-18, increased membrane rupture, and elevated lactate dehydrogenase release[22]. Transmission-electron microscopy likewise showed that YTHDF2 deficiency abolished betaine’s protection against Hcy-induced membrane blebbing and morphological damage in microglia. Together, these data identify YTHDF2 as a critical mediator of betaine-dependent protection in the Hcy model. Future work should evaluate YTHDF2 in additional inducible models of cognitive and psychiatric dysfunction (e.g., chronic stress, primary neuroinflammation) to define the generality of its role across neuropsychiatric disorders.
Although direct evidence linking YTHDF2 to amyotrophic lateral sclerosis (ALS) remains limited, several recent studies provide mechanistic clues suggesting its involvement in ALS-relevant pathogenic pathways. At the upstream regulatory level, circadian rhythm dysfunction - an emerging feature in ALS - has been shown to disrupt RNA metabolic ho
More direct evidence places YTHDF2 downstream in ALS-associated toxic pathways. A hallmark of ALS pathology is the cytoplasmic accumulation of the RNA-binding protein TAR DNA-binding protein 43 (TDP-43), whose RNA substrates are closely associated with m6A modification[24]. Analyses of postmortem ALS spinal cord tissue reveal widespread RNA hypermethylation and abnormal accumulation of YTHDF2 in motor neurons, with spatial overlap between YTHDF2 enrichment and TDP-43 pathology[24]. Functional studies further demonstrate that YTHDF2 acts as a critical m6A reader that amplifies TDP-43-mediated neurotoxicity: Knockdown of YTHDF2 in ALS-related mutant human induced pluripotent stem Cell-derived neurons significantly prolongs neuronal survival, while genetic depletion of YTHDF2 in primary rodent neurons attenuates neurotoxicity induced by TDP-43 overexpression. Conversely, YTHDF2 overexpression alone is sufficient to induce neuronal toxicity[24].
Together, these findings position YTHDF2 as a context-dependent regulatory modifier that links upstream circadian disruption to downstream RNA toxicity pathways in ALS, rather than as a primary causal factor.
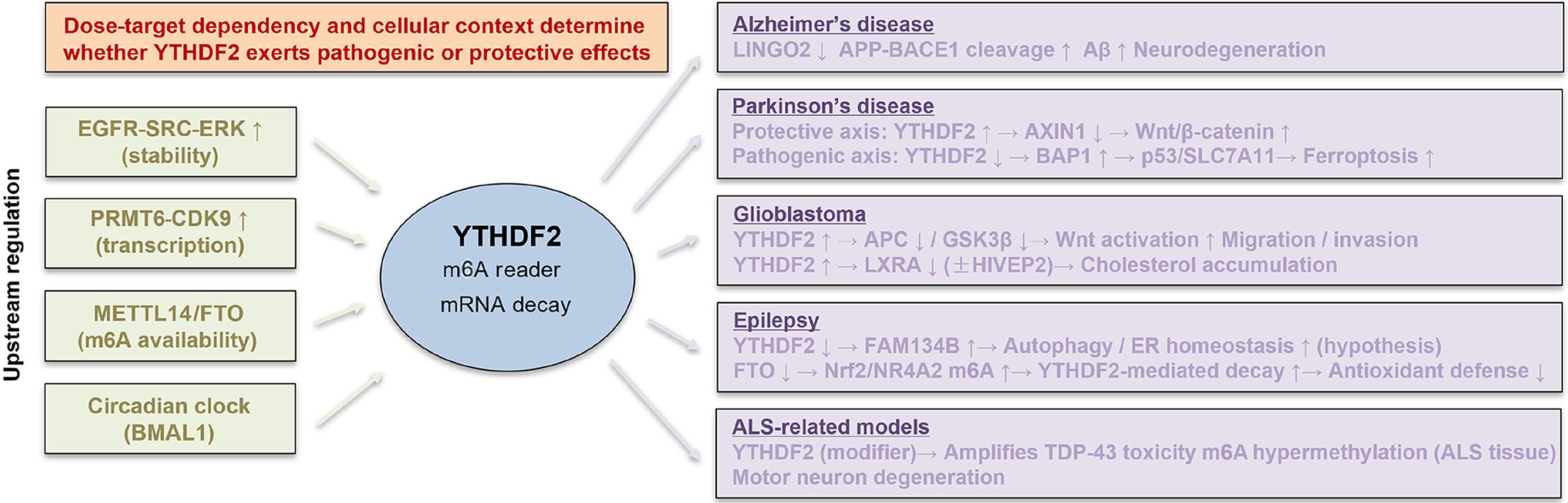
In summary, YTHDF2 is a central m6A reader with context-dependent, sometimes opposing actions in the nervous system (Figure 1). Depending on cell type, brain region, disease stage, and the specific mRNA targets involved, YTHDF2 can either accelerate pathology (for example, by promoting amyloid-β production, enhancing tumor growth, or amplifying inflammation) or exert protective effects (for example, by supporting neuronal survival, suppressing pyroptosis, or limiting tumor progression). Current evidence in AD, PD, glioblastoma, epilepsy, and Hcy-induced cognitive impairment is heterogeneous and incomplete; definitive mechanistic dissection requires combination studies in cells and animals. Future research should map the spatiotemporal expression and targetome of YTHDF2 in neurons, glia, and infiltrating immune cells, using conditional loss-of-function and gain-of-function models, RNA-stability assays, and comprehensive phenotyping. Given YTHDF2’s position within multiple key pathways, its utility as a biomarker or therapeutic target warrants careful evaluation and may open new avenues for diagnosis and treatment of neurological disease. Distinct from prior general overviews, this mini-review emphasizes how disease context, cellular identity, and target dosage collectively determine the net impact of YTHDF2, and integrates emerging evidence into a unifying framework with direct implications for experimental design and translational intervention.
| 1. | Malovic E, Ealy A, Miller C, Jang A, Hsu PJ, Sarkar S, Rokad D, Goeser C, Hartman AK, Zhu A, Palanisamy B, Zenitsky G, Jin H, Anantharam V, Kanthasamy A, He C, Kanthasamy AG. Epitranscriptomic reader YTHDF2 regulates SEK1(MAP2K4)-JNK-cJUN inflammatory signaling in astrocytes during neurotoxic stress. iScience. 2024;27:110619. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 7] [Article Influence: 3.5] [Reference Citation Analysis (1)] |
| 2. | Seigneurin-Berny D, Karczewski C, Delaforge E, Yaacoub K, Gaspar Litholdo C Jr, Favory JJ, Ringkjøbing Jensen M, Bousquet-Antonelli C, Verdel A. ECT2 peptide sequences outside the YTH domain regulate its m(6)A-RNA binding. RNA Biol. 2024;21:1-13. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 3] [Article Influence: 1.5] [Reference Citation Analysis (1)] |
| 3. | Fang R, Chen X, Zhang S, Shi H, Ye Y, Shi H, Zou Z, Li P, Guo Q, Ma L, He C, Huang S. EGFR/SRC/ERK-stabilized YTHDF2 promotes cholesterol dysregulation and invasive growth of glioblastoma. Nat Commun. 2021;12:177. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 183] [Cited by in RCA: 237] [Article Influence: 47.4] [Reference Citation Analysis (1)] |
| 4. | Xiao M, Wang X, Xiao E, Ming Q. Overexpression of FTO alleviates traumatic brain injury induced posttraumatic epilepsy by upregulating NR4A2 expression via m6A demethylation. Funct Integr Genomics. 2025;25:17. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 4] [Article Influence: 4.0] [Reference Citation Analysis (1)] |
| 5. | Wang JY, Lu AQ. The biological function of m6A reader YTHDF2 and its role in human disease. Cancer Cell Int. 2021;21:109. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 96] [Article Influence: 19.2] [Reference Citation Analysis (1)] |
| 6. | Yu P, Xu T, Ma W, Fang X, Bao Y, Xu C, Huang J, Sun Y, Li G. PRMT6-mediated transcriptional activation of ythdf2 promotes glioblastoma migration, invasion, and emt via the wnt-β-catenin pathway. J Exp Clin Cancer Res. 2024;43:116. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 28] [Reference Citation Analysis (1)] |
| 7. | Song L, Liu H, Yang W, Yin H, Wang J, Guo M, Yang Z. Biological functions of the m6A reader YTHDF2 and its role in central nervous system disorders. Biochem Pharmacol. 2024;230:116576. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 9] [Article Influence: 4.5] [Reference Citation Analysis (1)] |
| 8. | Li Z, Chen X, Xiang W, Tang T, Gan L. m6A Demethylase FTO-Mediated Upregulation of BAP1 Induces Neuronal Ferroptosis via the p53/SLC7A11 Axis in the MPP(+)/MPTP-Induced Parkinson's Disease Model. ACS Chem Neurosci. 2025;16:405-416. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 9] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 9. | Zhang L, Zhao X, Wang J, Jin Y, Gong M, Ye Y, Li P. METTL3 suppresses neuropathic pain via modulating N6-methyladenosine-dependent primary miR-150 processing. Cell Death Discov. 2022;8:80. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 34] [Article Influence: 8.5] [Reference Citation Analysis (1)] |
| 10. | Qiao Y, Mei Y, Xia M, Luo D, Gao L. The role of m6A modification in the risk prediction and Notch1 pathway of Alzheimer's disease. iScience. 2024;27:110235. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 10] [Article Influence: 5.0] [Reference Citation Analysis (1)] |
| 11. | Liu Z, Xia Q, Zhao X, Zheng F, Xiao J, Ge F, Wang D, Gao X. The Landscape of m6A Regulators in Multiple Brain Regions of Alzheimer's Disease. Mol Neurobiol. 2023;60:5184-5198. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 15] [Reference Citation Analysis (1)] |
| 12. | Zhao X, Ma C, Sun Q, Huang X, Qu W, Chen Y, Liu Z, Bao A, Sun B, Yang Y, Li X. Mettl3 regulates the pathogenesis of Alzheimer's disease via fine-tuning Lingo2. Mol Psychiatry. 2025;30:4047-4063. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 5] [Reference Citation Analysis (1)] |
| 13. | Lotharius J, Brundin P. Pathogenesis of Parkinson's disease: dopamine, vesicles and alpha-synuclein. Nat Rev Neurosci. 2002;3:932-942. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 890] [Cited by in RCA: 916] [Article Influence: 38.2] [Reference Citation Analysis (1)] |
| 14. | Li Y, Sheng H, Ma F, Wu Q, Huang J, Chen Q, Sheng L, Zhu X, Zhu X, Xu M. RNA m(6)A reader YTHDF2 facilitates lung adenocarcinoma cell proliferation and metastasis by targeting the AXIN1/Wnt/β-catenin signaling. Cell Death Dis. 2021;12:479. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 86] [Article Influence: 17.2] [Reference Citation Analysis (1)] |
| 15. | Zhang G, Chen L, Liu J, Jin Y, Lin Z, Du S, Fu Z, Chen T, Qin Y, Sui F, Jiang Y. HIF-1α/microRNA-128-3p axis protects hippocampal neurons from apoptosis via the Axin1-mediated Wnt/β-catenin signaling pathway in Parkinson's disease models. Aging (Albany NY). 2020;12:4067-4081. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 40] [Article Influence: 6.7] [Reference Citation Analysis (1)] |
| 16. | He H, Zhang Q, Liao J, Lei J, Luo M, Huang J, Chen M, Shen Y, Wang J, Xu P, Xiao Y. METTL14 is decreased and regulates m(6) A modification of α-synuclein in Parkinson's disease. J Neurochem. 2023;166:609-622. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 32] [Reference Citation Analysis (1)] |
| 17. | Tamimi AF, Juweid M. Epidemiology and Outcome of Glioblastoma. In: Glioblastoma [Internet]. Brisbane (AU): Codon Publications; 2017-Sep-27. [PubMed] [DOI] [Full Text] |
| 18. | Jiang X, Chen X, Huang X, Wang C, Wang C, Pan C, Cho WC, Nie Z, Pu J, Wang W. DNA methylation-regulated YTHDF2 correlates with cell migration and immune cell infiltration in glioma. Aging (Albany NY). 2022;14:7774-7793. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 4] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 19. | Cai M, Liu Q, Jiang Q, Wu R, Wang X, Wang Y. Loss of m(6) A on FAM134B promotes adipogenesis in porcine adipocytes through m(6) A-YTHDF2-dependent way. IUBMB Life. 2019;71:580-586. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 35] [Article Influence: 5.0] [Reference Citation Analysis (1)] |
| 20. | Xie N, Li Y, Wang C, Lian Y, Zhang H, Li Y, Meng X, Du L. FAM134B Attenuates Seizure-Induced Apoptosis and Endoplasmic Reticulum Stress in Hippocampal Neurons by Promoting Autophagy. Cell Mol Neurobiol. 2020;40:1297-1305. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 24] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 21. | Tian MQ, Li J, Shu XM, Lang CH, Chen J, Peng LY, Lei WT, Yang CJ. The increase of Nrf2 m6A modification induced by FTO downregulation promotes hippocampal neuron injury and aggravates the progression of epilepsy in a rat model. Synapse. 2023;77:e22270. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 13] [Reference Citation Analysis (1)] |
| 22. | Yang ZJ, Huang SY, Zhong KY, Huang WG, Huang ZH, He TT, Yang MT, Wusiman M, Zhou DD, Chen S, Huang BX, Luo XL, Li HB, Zhu HL. Betaine alleviates cognitive impairment induced by homocysteine through attenuating NLRP3-mediated microglial pyroptosis in an m(6)A-YTHDF2-dependent manner. Redox Biol. 2024;69:103026. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 61] [Article Influence: 30.5] [Reference Citation Analysis (1)] |
| 23. | Tam SB, Waldeck NJ, Wright M, Mojsilovic-Petrovic J, Baker EM, Kiskinis E, Bass J, Kalb RG. A role for the spinal cord cholinergic neuron circadian clock in RNA metabolism and mediating ALS disease phenotypes. bioRxiv. 2025;2025.05.06.652480. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 2] [Reference Citation Analysis (1)] |
| 24. | McMillan M, Gomez N, Hsieh C, Bekier M, Li X, Miguez R, Tank EMH, Barmada SJ. RNA methylation influences TDP43 binding and disease pathogenesis in models of amyotrophic lateral sclerosis and frontotemporal dementia. Mol Cell. 2023;83:219-236.e7. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 83] [Cited by in RCA: 76] [Article Influence: 25.3] [Reference Citation Analysis (1)] |