Published online Apr 24, 2026. doi: 10.5306/wjco.v17.i4.118606
Revised: January 27, 2026
Accepted: February 24, 2026
Published online: April 24, 2026
Processing time: 104 Days and 19.8 Hours
Although immunotherapy for colorectal cancer (CRC) has recently gained widespread attention, many patients continue to exhibit inherent or acquired resistance due to a lack of tumor-infiltrating lymphocytes and the poor immunogenicity of cancer cells within an immunosuppressive tumor microenvironment. Advances in high-throughput sequencing and bioinformatics have increasingly highlighted the role of the gut microbiome (GM) in modulating the quantity and phenotypes of innate and/or adaptive immune cells, thereby influencing CRC pathogenesis and the clinical response to immunotherapy. The GM maintains a symbiotic relationship with the host, contributes to protection against opportu
Core Tip: From birth onward, host health is closely linked to the delicate balance of ecological microbiota composition. The microbiota utilizes distinct enzymatic capabilities to metabolize nutrients and produce immunomodulatory byproducts that influence organ function and behavior. A strong and rapid association has been identified between gut microbiome dysbiosis, defined as a disruption in microbial load or diversity, colorectal cancer pathogenesis, and responsiveness to im
- Citation: Ti DD, Liu P, Wu CY, Shi ZM, Guo SM, Gao ZC. Host and gut microbiota crosstalk: A new paradigm for colorectal cancer immunotherapy. World J Clin Oncol 2026; 17(4): 118606
- URL: https://www.wjgnet.com/2218-4333/full/v17/i4/118606.htm
- DOI: https://dx.doi.org/10.5306/wjco.v17.i4.118606
Colorectal cancer (CRC) is the third most prevalent malignancy and the second leading cause of cancer-related mortality worldwide[1]. The rising incidence of CRC at a younger age has been linked to Westernized dietary patterns, sedentary lifestyles, delayed medical diagnosis and treatment, and exposure to colibactin, a genotoxin produced by certain bacterial strains[2,3]. Early-life exposure to colibactin plays a significant mutagenic role, contributing to the increased incidence of CRC in individuals under 50 years of age. CRC is a highly heterogeneous malignant epithelial neoplasm of the intestine, driven by the accumulation of genetic and epigenetic alterations within an immunosuppressive tumor microenvironment (iTME)[4]. Despite substantial progress, the molecular complexity of CRC continues to result in poor clinical outcomes and frequent recurrence following treatment modalities such as surgical resection, neoadjuvant chemoradiotherapy, and immunotherapy. Consequently, there is an urgent need to elucidate the mechanisms underlying CRC’s development of therapeutic resistance, and to develop more effective preventive and therapeutic strategies aimed at improving patient survival and quality of life.
With the rapid advancement of next-generation sequencing (NGS), research on the human microbiome has emerged over the past decade as a promising interdisciplinary field bridging basic and clinical medicine[5,6]. Humans have co-evolved with diverse commensal microflora, including bacteria, fungi, viruses, protozoa, and archaea, forming unique mutualistic holobiont ecosystems within individual hosts. These microbial communities play essential roles in main
The gastrointestinal tract hosts the largest and most diverse microbial community, containing more than 1000 species and approximately 1014 microorganisms, organized in a spatially structured manner[9,12]. Components of the GM exhibit distinct interindividual variability and readily adapt to dynamic environmental and host-derived cues. Although the core GM is 95% conserved across individuals and comprises fewer than 20 genera or species, its collective genetic material generates critical signals that mediate host-microbiota interactions, particularly in the development and regulation of innate and adaptive immune systems[13]. Under homeostasis, the GM prevents colonization by invasive opportunistic pathogens through the production of antagonistic metabolites or enzymes, stimulation of inflammatory cytokine secretion, and promotion of lymphocyte differentiation[14,15]. In response, the host immune system eliminates pathogenic microbes while preserving beneficial microflora to support overall physiological function. In the iTME of CRC, studies have reported a marked increase in the abundance of Fusobacterium and a notable decrease in Ruminococcus, accompanied by protumor neutrophil enrichment, disruption of mucosal immunity, and tumor metastasis[16]. Microbiota-targeted interventions aim to reprogram immune pathways, such as antigen processing and presentation (APP), T-cell receptor signaling, and T-cell activation, to enhance antitumor immune responses[17]. Therefore, it is plausible that manipulation of the GM could serve as a viable clinical adjuvant to conventional CRC immunotherapy. Such interventions may fine-tune the complex crosstalk between host immune components and microbial communities, providing cell-type-specific protection against infection and contributing to overall health improvement.
A strong and rapid association has been identified between GM dysbiosis (GMD), defined as a disruption in microbial load or diversity, CRC pathogenesis, and responsiveness to immunotherapy, due in part to the close anatomical proximity of the intestinal microbiota to the tumor sites[18-20]. The GM operates through distinct biochemical cycles, shaped by its taxonomic composition. In a pathologically hypoxic luminal environment, this ecosystem favors the overgrowth of harmful bacteria and a reduction in probiotic populations, ultimately resulting in an abnormal nutritional supply.
Several core pathogens associated with CRC, including Fusobacterium nucleatum (Fn), colibactin-producing Escherichia coli (CoPEC), and enterotoxigenic Bacteroides fragilis, increase the genotoxicity of intestinal epithelial cells (IECs) and impair barrier integrity, while concurrently inhibiting the growth of antagonistic microbes[21-23]. Following translocation through a compromised intestinal barrier, the conditional pathogen Alcaligenes faecalis has been shown to exacerbate inflammation during colitis and suppress intestinal immune surveillance, ultimately contributing to the transition from inflammation to cancer[24]. Notably, fecal microbiota transplantation (FMT) from CRC patients has been demonstrated to induce intestinal dysplasia, abnormal cellular proliferation, and colonic neoplasia in murine models[25,26]. These findings support a strong association between intestinal dysbiosis and CRC initiation and progression. Various oncogenic stimuli derived from GMD can trigger irreversible cell cycle arrest, referred to as cellular senescence, and dysregulated immune cell activation, both of which contribute to chronic inflammation and tumor development. A novel diagnostic technique for assessing individual intestinal dysbiosis has been developed, offering superior precision and sensitivity for the prevention, diagnosis, personalized immunotherapy, and clinical management of CRC[27].
As a representative approach in cancer immunotherapy, immune checkpoint inhibitors (ICIs) promote the recruitment and expansion of previously inactive or naive T-cell clones within the iTME, thereby restoring their capacity to target and eliminate cancer cells[28]. Despite the therapeutic potential of ICIs as a first-line treatment, the majority of CRC patients exhibit drug resistance due to a deficiency of tumor-infiltrating lymphocytes and the inherently low immunogenicity of cancer cells. Notably, Chinese yams have been shown to remodel GMD in CRC, thereby enhancing ICI efficacy by suppressing tumor-associated macrophage type 2 (TAM2) populations and promoting the infiltration of cytotoxic CD8+ T cells (CTLs)[29]. When the dynamic mutualism between host and microbiota is disrupted, oncomicrobial passengers gain a competitive growth advantage within poorly regulated inflammatory niches that provide a more permissive environment. These accumulated pathogens recruit tumor-infiltrating myeloid cells, enabling evasion of host immune responses. Therefore, identifying strategies to manipulate GMD to enhance antitumor activity and improve responses to ICIs is critical[30,31]. For example, following programmed death-1 (PD-1) blockade, the keystone bacterial species Akkermansia muciniphila (AKK) has been shown to increase the population of CCR9+ CXCR3+ CD4+ central memory T helper cells (Th), positively impacting progression-free survival in cancer patients[32]. There is an urgent need to understand the coordinated roles of specific gut bacteria and their metabolites in CRC immunotherapy, as revealed through both animal and human microbiome studies.
In this review, we summarize current evidence regarding the role of the GM in the occurrence, progression, and treatment of CRC. We highlight key principles of host-microbiota crosstalk that mediate immune regulation of malignant cells, with implications for the development of novel and promising clinical translation strategies.
The mammalian gut functions not only as an organ for nutrient absorption but also as the largest site of peripheral immune defense. In healthy individuals, the symbiotic GM educates the host immune system on tolerance, reactivity, and defense, conferring mutual benefits that support eubiosis in exchange for an anoxic environment and a rich energy supply provided by the host[33]. AKK utilizes intestinal mucin as a unique source of carbon and nitrogen and promotes the proliferation of mucin-secreting goblet cells, potentially contributing to the maintenance of intestinal barrier integrity[34]. The GM promptly expresses antimicrobial peptides, primes innate immune cells through the recognition of microbe-associated molecular patterns (MAMPs), and initiates a pattern recognition receptor-mediated immune response to prevent the penetration, survival, and replication of pathobionts within host cells[34]. Site-specific clustering of gut fungi mediates host-protective immunity and promotes intestinal epithelial repair through CD4+ Th-dependent interleukin (IL)-22 signalling[35]. Consequently, the GM is highly sensitive to environmental factors within the host. As the second-largest gene pool in the human body, GMD is likely to contribute to increased intestinal health risks.
Moreover, the extensive diversity of microbial metabolites profoundly influences host physiology by establishing a complex network within the microbe-host interaction. Microbe-derived short-chain fatty acids (SCFAs), including acetate, propionate, formate, butyrate, lactate, and succinate, are key mediators that directly modulate intestinal barrier function through interactions with both immune and nonimmune cells. Fibrolytic microbes degrade indigestible carbohydrates in the colon, producing SCFAs through fermentation. These small molecules diffuse into immune-associated regions, promoting the generation of anti-inflammatory regulatory T cells (Tregs), a process accompanied by colonic tuft cell expansion and compartmentalized accumulation that supports the maturation and function of the mucosal epithelium[36]. When intestinal and circulating metabolites are altered by inflammatory bowel disease, protective immunity becomes compromised, leading to increased susceptibility to infection due to the dysregulation of dendritic cells (DCs), macrophages (Mφs), and CD4+ T cells[37]. In CRC patients, reduced levels of SCFAs and probiotics, including Roseburia spp., Lactobacillus spp., and Bifidobacterium spp., are commonly observed. Supplementation with SCFAs enhances the immune system’s capacity to cooperate with the efficacy of PD-1 therapy, as SCFAs may initiate programmed cell death signaling and inhibit CRC cell proliferation[38]. These findings indicate that GMD disrupts the metabolic output of gut commensals. Increased awareness of disease-associated metabolomic profiles advances the understanding of interconnected cellular signaling pathways triggered by environmental risk factors and offers promising directions for improving health outcomes and managing complex diseases.
It is well established that the disorganized outer mucus layer of the intestinal barrier provides a permanent habitat for the commensal GM. Mucins, the primary components of mucus, contain a variety of receptors that facilitate bacterial adhesion through glycan interactions. These structures selectively retain beneficial microbiota while inhibiting pathogen attachment[39]. In response to microbial trafficking challenges, the gut initiates robust immune responses at specialized microanatomical sites, such as gut-associated lymphoid tissues. Microfold cells with the gut-associated lymphoid tissues promote microbial antigen transcytosis, facilitate the maturation of secretory immunoglobulin A+ B cells, and support the activation of antigen-specific effector T-cells (Teffs) to maintain mucosal immunity[40]. Segmented filamentous bacteria, the most abundant commensal species in the ileal mucosa, exploit suppressed entry points of microfold cells to mount a strong defense against the survival of invading pathogens[41]. As a regulator of intestinal immunity, the signalling lymphocyte activation molecule family member 4 is imprinted by the GM on T cells, B cells, professional antigen-presenting cells, natural killer cells (NKs), and innate lymphoid cells (ILCs), highlighting the critical role of the GM in immune system maturation and development[42]. In addition, the GM promotes immune tolerance and helps re-establish immune homeostasis by enhancing the cross-differentiation of CD8+ T cells into major histocompatibility complex I+ (MHC-I+) CD4+ T cells, a mechanism that may be relevant in the context of human immunodeficiency virus infection, cancer, and autoimmune disorders[43].
Currently, next-generation probiotics (NGPs), identified through NGS and bioinformatics approaches, exhibit distinct biological properties with potential health benefits and clinical applications. Faecalibacterium prausnitzii, one of the most prevalent anaerobic NGPs in the human colon, plays a protective role by preventing IEC damage, improving paracellular hyperpermeability, and inhibiting the activity of pro-inflammatory effectors[44]. Faecalibacterium prausnitzii also suppresses the enrichment of resident Mφs and supports the self-renewal and differentiation of epithelial progenitor and stem cells through Wnt/β-catenin signaling. Moreover, its supernatant inhibits CRC cell proliferation in a time- and dose-dependent manner[45].
Nontoxigenic Bacteroides fragilis strains, currently regarded as leading NGP candidates, prevent the adherence of opportunistic pathobionts to colonic epithelial cells and enhance mutualistic bacterial diversity[46]. These strains modulate the epithelial CD4+ T-cell compartment by promoting IL-10+ CD4+ T cells and suppressing pro-inflammatory IL-17+ CD4+ T cells in response to physiological stress or damage[47]. Microbe-derived mechanisms have evolved to support host-commensal symbiosis by inhibiting IEC apoptosis and attenuating pathogen-induced Th17 responses.
Recent evidence indicates that AKK abundance is associated with a healthier intestinal environment. AKK reduces circulating levels of pro-inflammatory bacterial lipopolysaccharides linked to increased gut permeability and stimulates the expression of tight junction proteins in IECs[48]. AKK recruits pro-inflammatory Mφs and CTLs into tumor beds, thereby inhibiting CRC progression through tumor necrosis factor-α upregulation and reduced PD-1 expression[49]. Additionally, Lactobacillus and Bifidobacterium species have been extensively studied in chronic inflammation-related disorders and are associated with favorable gastrointestinal tolerance and minimal clinical adverse effects. Lactobacillus and its metabolites enhance DC-mediated IL-10 production, thereby balancing pro-inflammatory and tolerogenic immune responses[50]. A combination of Lactobacilli and Bifidobacteria strains reduces tumor volume by fostering a healthy microbial environment rather than inducing immune-mediated or metabolomic disturbances[51]. Certain druggable bioactive molecules produced by probiotics, such as exopolysaccharides, mobilize CCR6+ CD8+ Teffs from Peyer’s patches to tumor sites, thereby enhancing the tumor-killing efficacy of ICIs[52]. These findings support the development of NGP-based therapeutic strategies, highlighting their potential as medicinal agents rather than mere dietary supplements. Such approaches aim to promote a more stable and interconnected microbial community, restore microbial homeostasis in the host, and alleviate symptoms associated with gastrointestinal diseases.
Using mathematical modeling techniques, approximately 70%-90% of CRC incidence is attributed to environmental factors, including diets low in fiber and high in red meat[53]. Abnormal lipid metabolism is increasingly recognized as a pathological hallmark of CRC. Notably, a high-fat diet (HFD) fails to support healthy regeneration of the intestinal epithelium. Consequently, changes in the relative composition of the GM occur, presenting a causal dilemma for lipid-degrading microbial populations. This microbial shift induces genetic alterations and disturbs the homeostasis of the gut lumen, ultimately promoting pathogenic infection and potentially carcinogenesis[54,55]. Translocation of specific pathobionts, induced by an HFD, is considered an initiating event that compromises immune protection and damages the intestinal epithelial barrier. Dysregulation of innate immune components is observed between DCs and ILC3s, with ILC3s producing IL-22 to mitigate intestinal inflammation associated with HFD-induced gut dysbiosis[56]. Following exposure to an HFD, Mφs are polarized toward the TAM2 phenotype, accompanied by a reduction in both the number and functionality of intratumoral CD8+ T cells. This observation suggests that CRC-associated GMD contributes to immune perturbations in the inflamed gut, potentially playing a key role in facilitating the immune system’s endogenous tumor escape behavior[57].
Mice fed an HFD exhibit an increased abundance of potentially harmful bacterial genera and a decreased presence of beneficial genera, along with associated oncogenic metabolomic abnormalities. These changes result in decreased expression of tight junction proteins, increased intestinal permeability, and enhanced proliferation and transformation of IECs[58]. In addition, expression of farnesoid X receptor, the principal regulator of bile acid synthesis, is reduced in patients with CRC[59]. Bile acid synthesis represents a key metabolic pathway for lipids, glucose, and energy utilization by the GM. When secondary bile acids (sBAs) become excessively enriched due to impaired negative feedback regulation, they can induce uncontrolled inflammation, elevate mitochondrial oxidative stress, and cause solubilization and disruption of cell membranes, ultimately leading to cellular toxicity[60].
Deoxycholic acid, one of the most abundant sBAs in humans, inhibits CD8+ T-cell activation, proliferation, effector molecule production, and cytotoxic function, thereby promoting CRC progression via the calcium signaling pathway[61]. sBAs also downregulate CXCL16 expression on endothelial cells, a chemokine essential for the recruitment of natural killer T cell to sites of inflammation[62]. Furthermore, sBAs interact with secretory immunoglobulin A, whose regulation depends on sBA levels, to enhance the adhesion and biofilm formation of enterotoxigenic Bacteroides fragilis. This process initiates a multistep protumorigenic inflammatory cascade, accelerating myeloid cell-dependent colonic neoplasia[59,63].
IECs integrate signals derived from tumorigenic pathobionts and contribute to host-microbiota mutualism that supports malignant progression. TLRs on the surface of IECs recognize bacterial toxins via MAMPs, leading to increased production of mitochondrial reactive oxygen species, which ultimately accelerates colitis-associated dysplasia and tumorigenesis[64]. IECs infected with Fn exhibit aberrant activation of the extracellular signal-regulated kinase/signal transducer and activator of transcription 3 signaling pathway, which is associated with a somatic mutation in a host RNA helicase and implicated in colorectal tumorigenesis[65]. When GMD disrupts intestinal immune homeostasis, impeding defense against virulent pathobionts and failing to suppress colonic inflammation, IECs promote polarization toward TAM2 and induce Th17 cell responses, thereby initiating a tumorigenic signaling cascade[66]. Additionally, IEC-recruited Tregs enhance the Th1/Th17 ratio by driving the differentiation of naive CD4+ T cells, contributing to protumoral IL-17 production[67].
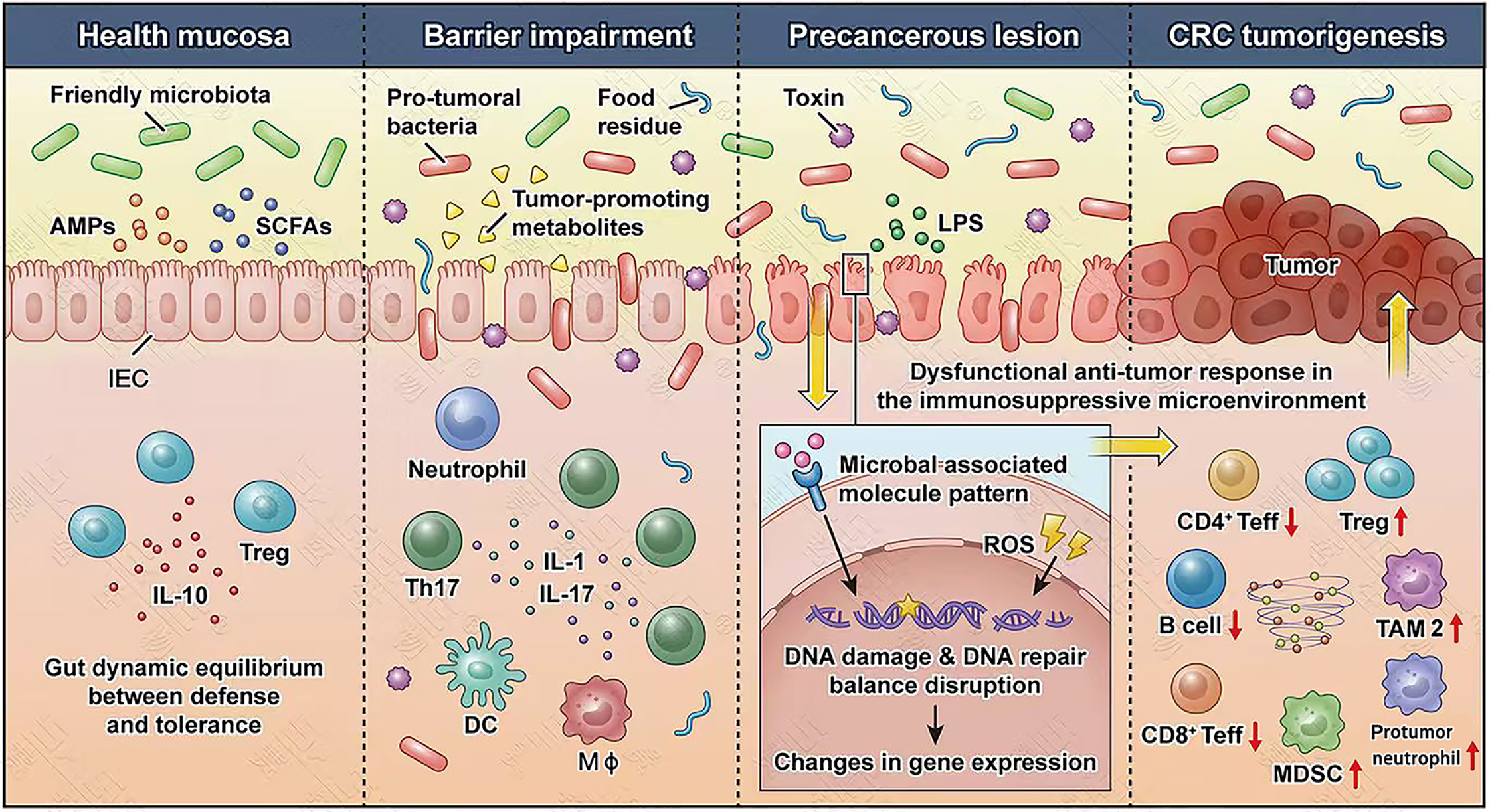
Recent studies have demonstrated that intestinal Fusobacterium exert lineage-specific effects on oncogenesis in CRC[68]. The probiotic Streptococcus thermophilus is notably absent in stool samples from CRC patients, and its metabolite, β-galactosidase, has been shown to promote cellular senescence by impairing energy homeostasis in neoplastic cells[69]. Dysregulation of host-oncomicrobiota interactions, characterized by a competitive growth advantage of pathogenic species, exacerbates mucosal disruption and fosters the development of a unique oncogenic driver iTME. This dysbiosis induces pathogenic mechanisms similar to those driven by oxidative stress, contributing to the transformation of the gut epithelium from a normal state to hyperplasia and, ultimately, to malignancy (Figure 1). The GM primes Mφ bystander potency as an intrinsic mechanism to activate nuclear factor-κB, induce DNA damage, and inhibit DNA repair processes[70]. GMD also promotes IEC tumorigenesis, accompanied by functional exhaustion of DCs and CTLs in CRC[71]. However, aberrant host cell subtypes fail to fully initiate antitumor immune surveillance and eradication due to elevated levels of myeloid-derived suppressor cells (MDSCs), TAM2, and tumor-associated neutrophils with the iTME[72,73]. Therefore, targeted restoration of GMD and associated metabolic reprogramming, such as through fiber-rich diets or NGP treatments, alongside activation of tumor-targeting downstream immune effector cells, may represent promising therapeutic strategies.
Although an increasing number of studies have reported GMD in CRC, identifying specific “oncogenic” microorganisms responsible for CRC initiation and/or progression remains challenging due to individual variability in GM composition. Nonetheless, metagenomic technologies and related approaches allow for the characterization of taxonomic and functional diversity, metabolomic profiles, and molecular mechanisms within perturbed microbial ecosystems (Table 1).
| CRC-associated bacteria | Virulence factors | Proposed carcinogenic mechanisms | Ref. |
| Fn | Fap2 | Inhibited natural killer cell cytotoxicity and tumor-infiltrating lymphocyte activities | [76] |
| FadA, Fap2, RadD, and FomA | Increased bacterial biomass and acute inflammation with pro-tumorigenic potential, tumor-associated macrophage type 2 and regulatory T cell infiltration in the tumor | [75] | |
| Formate | T helper 17 cell-favored proinflammatory profiles and metabolic shift with CRC stemness, invasion and metastasis | [77] | |
| CoPEC | Colibactin | Impaired antitumor T-cell response and procarcinogenic TME leading to immunotherapy resistance | [79] |
| Oncogenic-driven lipid reprogramming for lower tumor immunogenicity and acquired chemoresistance | [80] | ||
| ETBF | BFT | A pro-carcinogenic multi-step inflammatory cascade and nuclear factor-κB-activated myeloid-cell-dependent neoplasia | [63] |
| NTS | AvrA, SopE, SopE2, SopB, and SptP | Oncogenic transformation of pre-transformed cells upon targeting the host’s β-catenin, MAPK, and AKT signaling pathways | [81] |
| Bacterial invasion and intracellular replication | Host cell metabolic transformation associated with high mammalian target of rapamycin activation | [82] |
Fn is one of the most extensively studied CRC-associated microbes, owing to its high abundance in tumor tissues and its association with poor clinical outcomes. Therapeutic strategies targeting Fn have been shown to enhance the immunogenicity of tumor cells and may help to overcome resistance to ICIs[74]. As an opportunistic inhabitant of the oral cavity, Fn adheres to and invades endothelial and epithelial cells, subsequently disseminating through the circulation and becoming enriched at oncogenic sites during infection. In CRC foci, Fn suppresses the activity of tumor-infiltrating lymphocytes and NKs, while promoting the immunosuppressive functions of TAM2, Tregs, and MDSCs[75,76]. Mature Fn-derived formate reprograms the metabolic activity of CRC cells via Th17, thereby enhancing cancer stemness, invasiveness, and resistance to therapy[77]. Although Th17 play a role in host defense against pathobionts, this function is compromised under disrupted immune equilibrium. In contrast, pathogen-driven Th17 responses can increase host susceptibility to adverse disease outcomes and neoplastic transformation[78].
CoPEC is enriched in inflamed and neoplastic lesions and is detected more frequently in human CRC biopsies compared with control samples[79]. Colibactin induces primary cell transformation and precedes the appearance of a distinct mutational signature, including chromosomal abnormalities characteristic of genomic instability. CoPEC contributes to CRC tumorigenesis and chemoresistance by promoting a metabolic energy trade-off under conditions of lipid overload, primarily through the upregulation of reactive oxygen species in cancer cells[80].
Nontyphoidal Salmonella (NTS) infections primarily occur in low- and middle-income countries and are commonly transmitted through contaminated food chains. Mild and repetitive exposure to NTS is considered an environmental risk factor that contributes to the acceleration of oncogenic cell transformation and tumor proliferation[81]. Upon invasion and intracellular replication, NTS hyperactivates the mammalian target of rapamycin pathway, the central regulator of cellular metabolism, thereby promoting cellular transformation and impairing effective host defense responses[82].
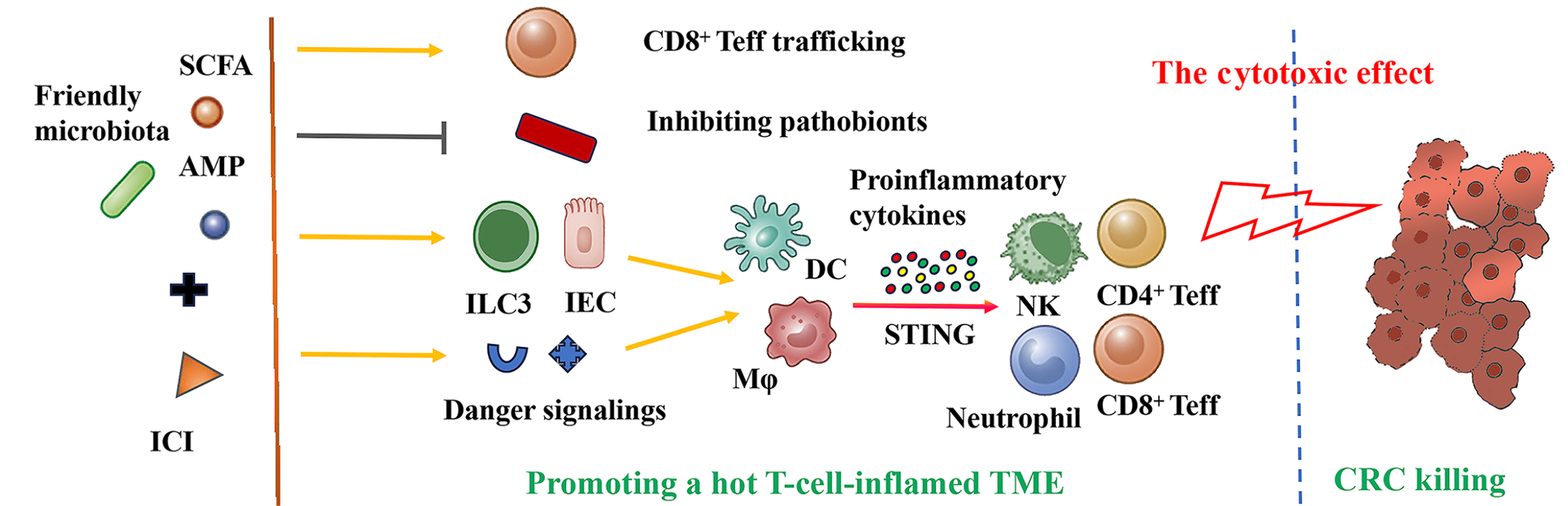
Given that the GM is an essential and intrinsic component of the TME, it is increasingly recognized as a critical regulatory factor due to its interactions with host cells and its influence on interrelated gene expression involved in modulating clinical responses to immunotherapy (Figure 2)[83,84]. Upon physical attachment to the intestinal epithelium, specific GM members shape the T-cell receptor repertoire within intestinal T cells and stimulate diverse T-cell clones to exert either regulatory or pro-inflammatory activities in response to microbiota-derived cognate antigens. Subsequently, GM-produced metabolites penetrate beyond the mucosal barrier and enter host tissues and circulation, where they are sensed by immune cells.
The abundance of probiotics, such as Bifidobacterium and Lacticaseibacillus, is positively correlated with the antitumor effects of ICIs on CRC[85,86]. Probiotic-derived metabolites contribute to enhanced trafficking of CD8+ T cells into the TME, facilitate the transition from “cold” to “warm” and eventually “hot” immune phenotypes, and support intestinal integrity. As a clinically relevant adjuvant, the microbial tryptophan catabolite indole-3-carboxaldehyde not only enhances the therapeutic efficacy of ICIs but also alleviates ICI-induced intestinal toxicity. This process decreases epithelial permeability, suppresses low-grade inflammation, and corrects GMD through activation of the host IL-22 pathway[87].
Distinct immune landscapes have been characterized within the TME of CRC. In a “cold” non-T-cell-inflamed TME, Teff priming, trafficking, and function are inhibited, and APP are silenced. Conversely, in a “hot” T-cell-inflamed TME, there is extensive immune cell infiltration, including clonally expanded memory T-cell populations and exhausted Teffs, along with an abundance of immunosuppressive MDSCs and TAM2[88,89].
Therefore, understanding the immune landscape in CRC is critical for reshaping gut microbiota composition and stimulating the TME to unlock preexisting antitumor immunity and improve the efficacy of ICIs. In a clinical trial, Bifidobacterium longum subsp. longum BB536 enhanced both innate and adaptive immune responses by priming plasmacytoid DCs and sustaining interferon (IFN) production. Tonic IFN signaling subsequently promoted the cytotoxic activity of NK and T cells, as well as neutrophil-mediated phagocytosis of pathogens[90]. When favorable microbiota are phagocytosed by APP, such as immunostimulatory monocytes and DCs, these microbes release IFN through the cyclic GMP-AMP synthase/stimulator of interferon genes (STING) axis, increase APP, and generate a pro-inflammatory environment that facilitates T-cell priming and recruitment[91].
FMT involving a consortium of bacterial species from healthy donors into CRC mouse models collectively increased the expression of CXCL9 and CXCL10, along with other IFN-inducible genes in IECs. Additionally, FMT enhanced
These findings suggest that the enhanced antitumor effects of ICIs are driven by cumulative infiltration and proliferation of Teffs with tumors. They also define a microbial antigen-driven differentiation of immune cells maintained through an IFN-mediated feed-forward loop. Furthermore, enrichment of AKK by a high-fiber diet reprograms the antitumor immunogenic landscape within the iTME via IFN-dependent STING signaling, orchestrates IFN-NK-DC crosstalk, and promotes an optimal response to ICIs in cancer patients[94].
ILC3s are recently recognized innate immune cells that belong to the microbiota-sensing, tissue-resident lymphocyte lineage, characterized by a highly plastic ability to distinguish between pathogens and commensals[56]. Impairment of ILC3s in CRC patients is associated with tumor progression and resistance to ICIs[95]. In coordination with host feeding rhythms, segmented filamentous bacteria interact with the ILC3-IEC immunological circuit to induce high levels of epithelial AMP expression, suggesting that ILC3s are synchronized with host-microbe coexistence to suppress enteric infection[96]. Upon invasion by exogenous pathobionts, increased IL-22 production by ILC3s is induced through tuft cell inflammasome signaling, which plays a critical role in early host defense immunity[97]. The absence of gatekeeper MHCII+ ILC3 disrupts the microbiota-specific CD4+ Teff adaptive immune response, leading to a concomitant increase in Th17/Th1 and promoting pathological intestinal inflammation during GMD[98]. As lymphoid tissue inducers, CCR6+ ILC3s secrete CXCL10 and synergistically enhance the tumor-assault capability of ICIs by increasing the infiltration of CD4+ and CD8+ T cells into the TME[99]. Further investigation is needed to elucidate the significance of ILC3-gut microbiota mutualism in reshaping a T cell-inflamed TME and mediating protective antitumor immunity.
In the context of ICI therapy, the capacity of CRC cells to prime and amplify antigen-presenting cell-dependent innate sensing and T cell-mediated tumor-killing responses is critical. Tumor cells acquire heterogeneous phenotypes through abnormal APP expression or by hijacking immune signaling pathways to evade dynamic immune pressures within the iTME. Due to their highly commensal genomes, members of the GM, such as Enterococcus hirae, may inadvertently mimic tumor antigens while escaping self-tolerance mechanisms[100]. Subsequently, a bacteriophage-encoded, MHC-restricted epitope from Enterococcus hirae is cross-presented by DCs, leading to activation of memory IFN-γ+ CD8+ and CD4+ T cells that specifically target and eliminate tumor cells. Microbe-specific T cells are frequently detected within tumor sites, where they exhibit unique functions by recognizing tumor antigens, mediating cross-reactivity, and attacking neoplastic tissues[101]. One gut microbial metabolite, methylglyoxal, promotes immunogenic cell death and facilitates the recruitment of CD8+ T cells and NKs via STING signaling, thereby enhancing the abscopal effect of cancer therapy[102]. Growing evidence indicates that the GM interacts with the intestinal immune system and modulates the pro- and antitumorigenic balance within the TME. Enterococcus secretes antigen A via MAMPs, initiating antitumor immunity through activation of the key inflammatory nuclear factor-κB signaling pathway in ICI-responsive patients[103]. If specific microorganisms and their metabolites amplify host immunosurveillance through the release of “danger” signals, they may serve as novel adjuvants to enhance tumor antigenicity, thereby contributing directly to neoantigens generation capable of refreshing the immune system and cold-to-hot tumor transition. This shift has the potential to increase immune vigilance, stimulate the release of immunostimulatory chemotactic cytokines, promote effector immune cell infiltration, and enhance the remedial activity of ICIs, ultimately triggering a cascade of favorable biological processes that support immune-mediated tumor clearance.
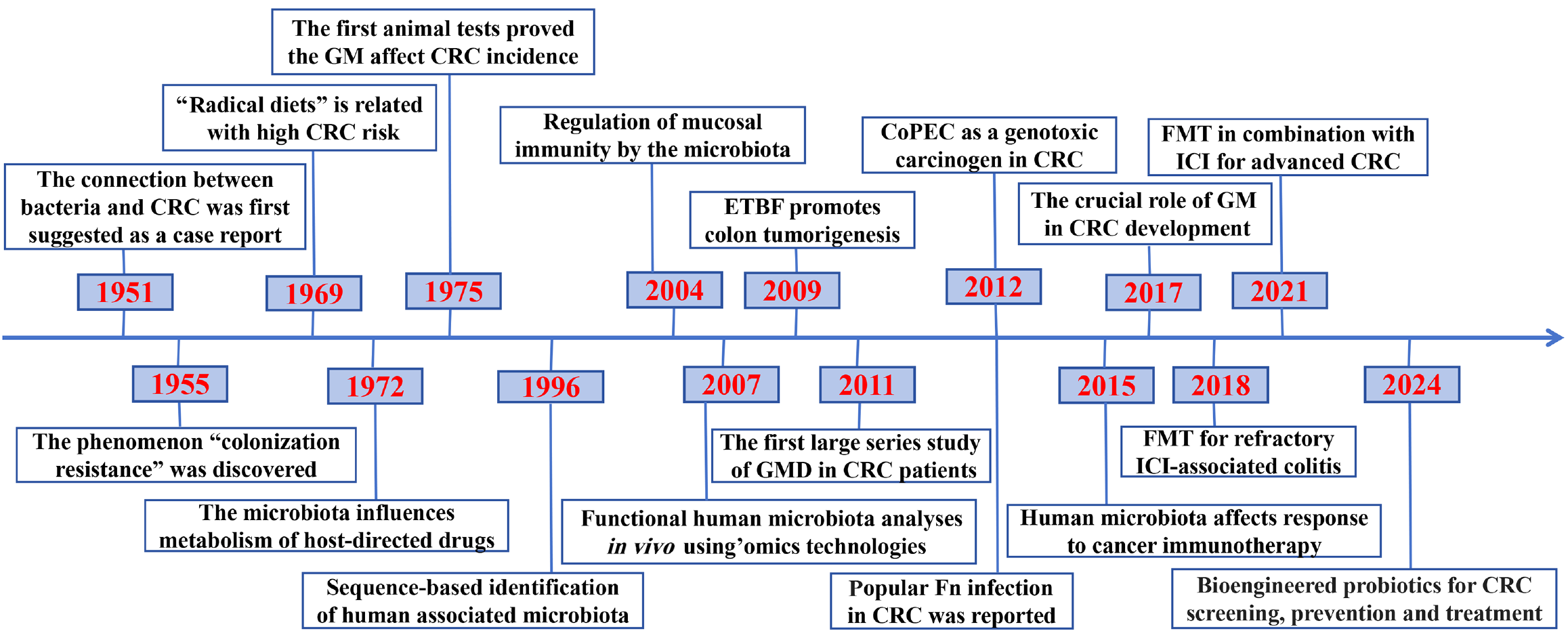
The widespread adoption of NGS has led to substantial progress in elucidating the functional and mechanistic roles of GMD in CRC progression and suboptimal responses to immunotherapy. While existing studies have characterized the biological effects of MAMP-pattern recognition receptor signaling triggered by CRC-associated oncomicrobes and metabolites, reshaping the microbial community represents a promising strategy to enhance the antitumor efficacy of ICIs through targeted adjuvant interventions. Nevertheless, microbiome-based technologies remain in the early stages of clinical translation. Human clinical trials must be designed with rigorous biological, methodological, and logistical frameworks (Figure 3). Furthermore, it is essential to establish personalized and standardized approaches for therapeutic microbial modulation to ensure consistent safety and efficacy, thereby facilitating the integration of microbiome-based therapies into multidisciplinary clinical practice.
Emerging clinical strategies to manipulate the gut microbiota, such as FMT, dietary interventions, and probiotics, suggest a higher probability of treatment response, improved survival, and potentially reduced toxicities in ICIs-treated patients. For example, the Mediterranean diet, riched in plant-based foods and low in processed foods and red meat, promotes the growth of beneficial SCFA-producing bacterial populations and enhances intestinal barrier integrity. Notably, microbiome-metabolome-host interactions facilitated by precision nutritional interventions may increase CD8+ T cell cytotoxicity and reduce infiltration of MDSCs, thereby alleviating ICI resistance[104].
Advances in multi-omics technologies have enabled the identification of dynamic host-GM mutualism and the establishment of tissue-specific protein-microbe associations that reflect pathophysiological signatures of CRC. These insights pave the way for precision, integrative, microbiota-directed therapies aimed at improving clinical outcomes and quality of life for cancer patients. Despite these developments, the capacity of the regulatory network between the gut microbiota and immune cells to effectively enhance CRC treatment remains incompletely understood[105,106]. In line with this concept, reciprocal beneficial or antagonistic interactions among microbial populations provide distinct causal insights that can be integrated into individualized GM profiles via host immune responses. Additionally, reliable single-copy protein-coding marker genes have been used to preliminarily characterize the diversity of microbial phylotypes[107]. Taken together, these findings support the notion that microbe-based interventions, operating through health-oriented and adaptable mechanisms, may become essential components of antitumor therapy in the future. particularly when an immunogenic, “hot” tumor microenvironment is successfully activated.
| 1. | Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71:209-249. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76817] [Cited by in RCA: 69808] [Article Influence: 13961.6] [Reference Citation Analysis (51)] |
| 2. | Díaz-Gay M, Dos Santos W, Moody S, Kazachkova M, Abbasi A, Steele CD, Vangara R, Senkin S, Wang J, Fitzgerald S, Bergstrom EN, Khandekar A, Otlu B, Abedi-Ardekani B, de Carvalho AC, Cattiaux T, Penha RCC, Gaborieau V, Chopard P, Carreira C, Cheema S, Latimer C, Teague JW, Mukeriya A, Zaridze D, Cox R, Albert M, Phouthavongsy L, Gallinger S, Malekzadeh R, Niavarani A, Miladinov M, Erić K, Milosavljevic S, Sangrajrang S, Curado MP, Aguiar S, Reis RM, Reis MT, Romagnolo LG, Guimarães DP, Holcatova I, Kalvach J, Vaccaro CA, Piñero TA, Świątkowska B, Lissowska J, Roszkowska-Purska K, Huertas-Salgado A, Shibata T, Shiba S, Sangkhathat S, Chitapanarux T, Roshandel G, Ashton-Prolla P, Damin DC, de Oliveira FH, Humphreys L, Lawley TD, Perdomo S, Stratton MR, Brennan P, Alexandrov LB. Geographic and age variations in mutational processes in colorectal cancer. Nature. 2025;643:230-240. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 113] [Cited by in RCA: 94] [Article Influence: 94.0] [Reference Citation Analysis (0)] |
| 3. | Aries V, Crowther JS, Drasar BS, Hill MJ, Williams RE. Bacteria and the aetiology of cancer of the large bowel. Gut. 1969;10:334-335. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 244] [Cited by in RCA: 227] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 4. | Yu H, Yang R, Li M, Li D, Xu Y. The role of Treg cells in colorectal cancer and the immunotherapy targeting Treg cells. Front Immunol. 2025;16:1574327. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 8] [Reference Citation Analysis (0)] |
| 5. | Klaassens ES, de Vos WM, Vaughan EE. Metaproteomics approach to study the functionality of the microbiota in the human infant gastrointestinal tract. Appl Environ Microbiol. 2007;73:1388-1392. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 145] [Cited by in RCA: 110] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 6. | Wilson KH, Blitchington RB. Human colonic biota studied by ribosomal DNA sequence analysis. Appl Environ Microbiol. 1996;62:2273-2278. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 355] [Cited by in RCA: 320] [Article Influence: 10.7] [Reference Citation Analysis (0)] |
| 7. | Peppercorn MA, Goldman P. The role of intestinal bacteria in the metabolism of salicylazosulfapyridine. J Pharmacol Exp Ther. 1972;181:555-562. [PubMed] |
| 8. | Bashir ME, Louie S, Shi HN, Nagler-Anderson C. Toll-like receptor 4 signaling by intestinal microbes influences susceptibility to food allergy. J Immunol. 2004;172:6978-6987. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 292] [Cited by in RCA: 264] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 9. | De Luca R, Arrè V, Nardone S, Incerpi S, Giannelli G, Trivedi P, Anastasiadou E, Negro R. Gastrointestinal microbiota and inflammasomes interplay in health and disease: a gut feeling. Gut. 2025;75:161-175. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 13] [Article Influence: 13.0] [Reference Citation Analysis (0)] |
| 10. | Pressley SR, McGill AS, Luu B, Atsumi S. Recent Advances in the Microbial Production of Human Milk Oligosaccharides. Curr Opin Food Sci. 2024;57:101154. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 8] [Reference Citation Analysis (0)] |
| 11. | Guiomar de Almeida Brasiel P, Cristina Potente Dutra Luquetti S, Dutra Medeiros J, Otavio do Amaral Corrêa J, Barbosa Ferreira Machado A, Paula Boroni Moreira A, Novaes Rocha V, Teodoro de Souza C, do Carmo Gouveia Peluzio M. Kefir modulates gut microbiota and reduces DMH-associated colorectal cancer via regulation of intestinal inflammation in adulthood offsprings programmed by neonatal overfeeding. Food Res Int. 2022;152:110708. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 16] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 12. | Gyriki D, Nikolaidis CG, Bezirtzoglou E, Voidarou C, Stavropoulou E, Tsigalou C. The gut microbiota and aging: interactions, implications, and interventions. Front Aging. 2025;6:1452917. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 40] [Cited by in RCA: 39] [Article Influence: 39.0] [Reference Citation Analysis (0)] |
| 13. | Sanna S, Kurilshikov A, van der Graaf A, Fu J, Zhernakova A. Challenges and future directions for studying effects of host genetics on the gut microbiome. Nat Genet. 2022;54:100-106. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 92] [Article Influence: 23.0] [Reference Citation Analysis (0)] |
| 14. | Freter R. The fatal enteric cholera infection in the guinea pig, achieved by inhibition of normal enteric flora. J Infect Dis. 1955;97:57-65. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 123] [Cited by in RCA: 105] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 15. | Ma Z, Zuo T, Frey N, Rangrez AY. A systematic framework for understanding the microbiome in human health and disease: from basic principles to clinical translation. Signal Transduct Target Ther. 2024;9:237. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 123] [Article Influence: 61.5] [Reference Citation Analysis (2)] |
| 16. | Li Q, Xiao Y, Han L, Luo W, Dai W, Fang H, Wang R, Xu Y, Cai S, Goel A, Bai F, Cai G. Microbiome dysbiosis, neutrophil recruitment and mesenchymal transition of mesothelial cells promotes peritoneal metastasis of colorectal cancer. Nat Cancer. 2025;6:493-510. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 29] [Article Influence: 29.0] [Reference Citation Analysis (0)] |
| 17. | Xu Y, Gao Z, Liu J, Yang Q, Xu S. Role of gut microbiome in suppression of cancers. Gut Microbes. 2025;17:2495183. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 12] [Reference Citation Analysis (0)] |
| 18. | Masaadeh AH, Eletrebi M, Parajuli B, De Jager N, Bosch DE. Human colitis-associated colorectal carcinoma progression is accompanied by dysbiosis with enriched pathobionts. Gut Microbes. 2025;17:2479774. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 11] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 19. | Sobhani I, Tap J, Roudot-Thoraval F, Roperch JP, Letulle S, Langella P, Corthier G, Tran Van Nhieu J, Furet JP. Microbial dysbiosis in colorectal cancer (CRC) patients. PLoS One. 2011;6:e16393. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 698] [Cited by in RCA: 623] [Article Influence: 41.5] [Reference Citation Analysis (10)] |
| 20. | Lucas C, Barnich N, Nguyen HTT. Microbiota, Inflammation and Colorectal Cancer. Int J Mol Sci. 2017;18:1310. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 274] [Cited by in RCA: 244] [Article Influence: 27.1] [Reference Citation Analysis (6)] |
| 21. | Castellarin M, Warren RL, Freeman JD, Dreolini L, Krzywinski M, Strauss J, Barnes R, Watson P, Allen-Vercoe E, Moore RA, Holt RA. Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res. 2012;22:299-306. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1757] [Cited by in RCA: 1577] [Article Influence: 112.6] [Reference Citation Analysis (6)] |
| 22. | Wu S, Rhee KJ, Albesiano E, Rabizadeh S, Wu X, Yen HR, Huso DL, Brancati FL, Wick E, McAllister F, Housseau F, Pardoll DM, Sears CL. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nat Med. 2009;15:1016-1022. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1513] [Cited by in RCA: 1381] [Article Influence: 81.2] [Reference Citation Analysis (7)] |
| 23. | Arthur JC, Perez-Chanona E, Mühlbauer M, Tomkovich S, Uronis JM, Fan TJ, Campbell BJ, Abujamel T, Dogan B, Rogers AB, Rhodes JM, Stintzi A, Simpson KW, Hansen JJ, Keku TO, Fodor AA, Jobin C. Intestinal inflammation targets cancer-inducing activity of the microbiota. Science. 2012;338:120-123. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1978] [Cited by in RCA: 1749] [Article Influence: 124.9] [Reference Citation Analysis (17)] |
| 24. | Zheng J, Zhou C, Li Z, Jin X, Zou Y, Bai S, Zheng H, Ling W, Zhao Y, Wang Y, Zhang R, Liu Z, Lu L. Alcaligenes faecalis promotes colitis to colorectal cancer transition through IgA+ B cell suppression and vinculin acetylation. Gut Microbes. 2025;17:2473511. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 14] [Reference Citation Analysis (0)] |
| 25. | Chao X, Lei Z, Hongqin L, Ziwei W, Dechuan L, Weidong D, Lu X, Haitao C, Bo Z, Haixing J, Qinghua Y. Faeces from malnourished colorectal cancer patients accelerate cancer progression. Clin Nutr. 2022;41:632-644. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 17] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 26. | Weisburger JH, Reddy BS, Narisawa T, Wynder EL. Germ-free status and colon tumor induction by N-methyl-N'-nitro-N-nitrosoguanidine. Proc Soc Exp Biol Med. 1975;148:1119-1121. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 32] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 27. | Derosa L, Iebba V, Silva CAC, Piccinno G, Wu G, Lordello L, Routy B, Zhao N, Thelemaque C, Birebent R, Marmorino F, Fidelle M, Messaoudene M, Thomas AM, Zalcman G, Friard S, Mazieres J, Audigier-Valette C, Sibilot DM, Goldwasser F, Scherpereel A, Pegliasco H, Ghiringhelli F, Bouchard N, Sow C, Darik I, Zoppi S, Ly P, Reni A, Daillère R, Deutsch E, Lee KA, Bolte LA, Björk JR, Weersma RK, Barlesi F, Padilha L, Finzel A, Isaksen ML, Escudier B, Albiges L, Planchard D, André F, Cremolini C, Martinez S, Besse B, Zhao L, Segata N, Wojcik J, Kroemer G, Zitvogel L. Custom scoring based on ecological topology of gut microbiota associated with cancer immunotherapy outcome. Cell. 2024;187:3373-3389.e16. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 141] [Article Influence: 70.5] [Reference Citation Analysis (6)] |
| 28. | Zhang M, Liu C, Tu J, Tang M, Ashrafizadeh M, Nabavi N, Sethi G, Zhao P, Liu S. Advances in cancer immunotherapy: historical perspectives, current developments, and future directions. Mol Cancer. 2025;24:136. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 125] [Reference Citation Analysis (0)] |
| 29. | Zhang G, Pan J, Xu X, Nie S, Lu L, Jing Y, Yang F, Ji G, Xu H. Chinese yam polysaccharide enhances anti-PD-1 immunotherapy in colorectal cancer through alterations in the gut microbiota and metabolites. Int J Biol Macromol. 2025;310:143323. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 15] [Reference Citation Analysis (0)] |
| 30. | Sivan A, Corrales L, Hubert N, Williams JB, Aquino-Michaels K, Earley ZM, Benyamin FW, Lei YM, Jabri B, Alegre ML, Chang EB, Gajewski TF. Commensal Bifidobacterium promotes antitumor immunity and facilitates anti-PD-L1 efficacy. Science. 2015;350:1084-1089. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3408] [Cited by in RCA: 3116] [Article Influence: 283.3] [Reference Citation Analysis (12)] |
| 31. | Kang YB, Cai Y. Faecal microbiota transplantation enhances efficacy of immune checkpoint inhibitors therapy against cancer. World J Gastroenterol. 2021;27:5362-5375. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 20] [Cited by in RCA: 32] [Article Influence: 6.4] [Reference Citation Analysis (0)] |
| 32. | Routy B, Le Chatelier E, Derosa L, Duong CPM, Alou MT, Daillère R, Fluckiger A, Messaoudene M, Rauber C, Roberti MP, Fidelle M, Flament C, Poirier-Colame V, Opolon P, Klein C, Iribarren K, Mondragón L, Jacquelot N, Qu B, Ferrere G, Clémenson C, Mezquita L, Masip JR, Naltet C, Brosseau S, Kaderbhai C, Richard C, Rizvi H, Levenez F, Galleron N, Quinquis B, Pons N, Ryffel B, Minard-Colin V, Gonin P, Soria JC, Deutsch E, Loriot Y, Ghiringhelli F, Zalcman G, Goldwasser F, Escudier B, Hellmann MD, Eggermont A, Raoult D, Albiges L, Kroemer G, Zitvogel L. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science. 2018;359:91-97. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4719] [Cited by in RCA: 4276] [Article Influence: 534.5] [Reference Citation Analysis (4)] |
| 33. | Wallrapp A, Chiu IM. Neuroimmune Interactions in the Intestine. Annu Rev Immunol. 2024;42:489-519. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 56] [Cited by in RCA: 54] [Article Influence: 27.0] [Reference Citation Analysis (0)] |
| 34. | Gao F, Cheng C, Li R, Chen Z, Tang K, Du G. The role of Akkermansia muciniphila in maintaining health: a bibliometric study. Front Med (Lausanne). 2025;12:1484656. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 18] [Reference Citation Analysis (0)] |
| 35. | Leonardi I, Gao IH, Lin WY, Allen M, Li XV, Fiers WD, De Celie MB, Putzel GG, Yantiss RK, Johncilla M, Colak D, Iliev ID. Mucosal fungi promote gut barrier function and social behavior via Type 17 immunity. Cell. 2022;185:831-846.e14. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 52] [Cited by in RCA: 240] [Article Influence: 60.0] [Reference Citation Analysis (0)] |
| 36. | Krause FF, Mangold KI, Ruppert AL, Leister H, Hellhund-Zingel A, Lopez Krol A, Pesek J, Watzer B, Winterberg S, Raifer H, Binder K, Kinscherf R, Walker A, Nockher WA, Taudte RV, Bertrams W, Schmeck B, Kühl AA, Siegmund B, Romero R, Luu M, Göttig S, Bekeredjian-Ding I, Steinhoff U, Schütz B, Visekruna A. Clostridium sporogenes-derived metabolites protect mice against colonic inflammation. Gut Microbes. 2024;16:2412669. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 40] [Reference Citation Analysis (0)] |
| 37. | Edwards M, Brockmann L. Microbiota-dependent modulation of intestinal anti-inflammatory CD4(+) T cell responses. Semin Immunopathol. 2025;47:23. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 7] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 38. | Thulasinathan B, Suvilesh KN, Maram S, Grossmann E, Ghouri Y, Teixeiro EP, Chan J, Kaif JT, Rachagani S. The impact of gut microbial short-chain fatty acids on colorectal cancer development and prevention. Gut Microbes. 2025;17:2483780. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 36] [Article Influence: 36.0] [Reference Citation Analysis (4)] |
| 39. | Li M, Ding Y, Wei J, Dong Y, Wang J, Dai X, Yan J, Chu F, Zhang K, Meng F, Ma J, Zhong W, Wang B, Gao Y, Yang R, Liu X, Su X, Cao H. Gut microbiota metabolite indole-3-acetic acid maintains intestinal epithelial homeostasis through mucin sulfation. Gut Microbes. 2024;16:2377576. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 109] [Cited by in RCA: 97] [Article Influence: 48.5] [Reference Citation Analysis (0)] |
| 40. | Kulkarni DH, Newberry RD. Antigen Uptake in the Gut: An Underappreciated Piece to the Puzzle? Annu Rev Immunol. 2025;43:571-588. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 11] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 41. | Lai NY, Musser MA, Pinho-Ribeiro FA, Baral P, Jacobson A, Ma P, Potts DE, Chen Z, Paik D, Soualhi S, Yan Y, Misra A, Goldstein K, Lagomarsino VN, Nordstrom A, Sivanathan KN, Wallrapp A, Kuchroo VK, Nowarski R, Starnbach MN, Shi H, Surana NK, An D, Wu C, Huh JR, Rao M, Chiu IM. Gut-Innervating Nociceptor Neurons Regulate Peyer's Patch Microfold Cells and SFB Levels to Mediate Salmonella Host Defense. Cell. 2020;180:33-49.e22. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 182] [Cited by in RCA: 287] [Article Influence: 47.8] [Reference Citation Analysis (0)] |
| 42. | Cabinian A, Sinsimer D, Tang M, Jang Y, Choi B, Laouar Y, Laouar A. Gut symbiotic microbes imprint intestinal immune cells with the innate receptor SLAMF4 which contributes to gut immune protection against enteric pathogens. Gut. 2018;67:847-859. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 26] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 43. | Lui JB, McGinn LS, Chen Z. Gut microbiota amplifies host-intrinsic conversion from the CD8 T cell lineage to CD4 T cells for induction of mucosal immune tolerance. Gut Microbes. 2016;7:40-47. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 9] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 44. | Martín R, Rios-Covian D, Huillet E, Auger S, Khazaal S, Bermúdez-Humarán LG, Sokol H, Chatel JM, Langella P. Faecalibacterium: a bacterial genus with promising human health applications. FEMS Microbiol Rev. 2023;47:fuad039. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 447] [Cited by in RCA: 363] [Article Influence: 121.0] [Reference Citation Analysis (0)] |
| 45. | Dikeocha IJ, Al-Kabsi AM, Chiu HT, Alshawsh MA. Faecalibacterium prausnitzii Ameliorates Colorectal Tumorigenesis and Suppresses Proliferation of HCT116 Colorectal Cancer Cells. Biomedicines. 2022;10:1128. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 72] [Reference Citation Analysis (4)] |
| 46. | Chan JL, Wu S, Geis AL, Chan GV, Gomes TAM, Beck SE, Wu X, Fan H, Tam AJ, Chung L, Ding H, Wang H, Pardoll DM, Housseau F, Sears CL. Non-toxigenic Bacteroides fragilis (NTBF) administration reduces bacteria-driven chronic colitis and tumor development independent of polysaccharide A. Mucosal Immunol. 2019;12:164-177. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 51] [Cited by in RCA: 80] [Article Influence: 11.4] [Reference Citation Analysis (0)] |
| 47. | Round JL, Lee SM, Li J, Tran G, Jabri B, Chatila TA, Mazmanian SK. The Toll-like receptor 2 pathway establishes colonization by a commensal of the human microbiota. Science. 2011;332:974-977. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1398] [Cited by in RCA: 1251] [Article Influence: 83.4] [Reference Citation Analysis (4)] |
| 48. | Aja E, Zeng A, Gray W, Connelley K, Chaganti A, Jacobs JP. Health Effects and Therapeutic Potential of the Gut Microbe Akkermansia muciniphila. Nutrients. 2025;17:562. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 26] [Reference Citation Analysis (0)] |
| 49. | Wang L, Tang L, Feng Y, Zhao S, Han M, Zhang C, Yuan G, Zhu J, Cao S, Wu Q, Li L, Zhang Z. A purified membrane protein from Akkermansia muciniphila or the pasteurised bacterium blunts colitis associated tumourigenesis by modulation of CD8(+) T cells in mice. Gut. 2020;69:1988-1997. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 542] [Cited by in RCA: 493] [Article Influence: 82.2] [Reference Citation Analysis (4)] |
| 50. | Liu HY, Li S, Ogamune KJ, Yuan P, Shi X, Ennab W, Ahmed AA, Kim IH, Hu P, Cai D. Probiotic Lactobacillus johnsonii Reduces Intestinal Inflammation and Rebalances Splenic Treg/Th17 Responses in Dextran Sulfate Sodium-Induced Colitis. Antioxidants (Basel). 2025;14:433. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 13] [Reference Citation Analysis (0)] |
| 51. | Niechcial A, Schwarzfischer M, Wawrzyniak P, Determann M, Pöhlmann D, Wawrzyniak M, Gueguen E, Walker MR, Morsy Y, Atrott K, Wilmink M, Linzmeier L, Spalinger MR, Holowacz S, Leblanc A, Scharl M. Probiotic Administration Modulates Gut Microbiota and Suppresses Tumor Growth in Murine Models of Colorectal Cancer. Int J Mol Sci. 2025;26:4404. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 7] [Reference Citation Analysis (0)] |
| 52. | Di Luccia B, Colonna M. Precision Probiotic Medicine to Improve ICB Immunotherapy. Cancer Discov. 2022;12:1189-1190. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 18] [Reference Citation Analysis (3)] |
| 53. | Berbecka M, Berbecki M, Gliwa AM, Szewc M, Sitarz R. Managing Colorectal Cancer from Ethology to Interdisciplinary Treatment: The Gains and Challenges of Modern Medicine. Int J Mol Sci. 2024;25:2032. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 7] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 54. | Yo S, Matsumoto H, Gu T, Sasahira M, Oosawa M, Handa O, Umegaki E, Shiotani A. Exercise Affects Mucosa-Associated Microbiota and Colonic Tumor Formation Induced by Azoxymethane in High-Fat-Diet-Induced Obese Mice. Microorganisms. 2024;12:957. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 6] [Reference Citation Analysis (0)] |
| 55. | Mccoy WC, Mason JM 3rd. Enterococcal endocarditis associated with carcinoma of the sigmoid; report of a case. J Med Assoc State Ala. 1951;21:162-166. [PubMed] |
| 56. | Xiong L, Diwakarla S, Chatzis R, Artaiz O, Macowan M, Zhang S, Garnham A, Morgan PK, Mellett NA, Meikle PJ, Lancaster GI, Marsland BJ, Nutt SL, Seillet C. Acute exposure to high-fat diet impairs ILC3 functions and gut homeostasis. Immunity. 2025;58:1185-1200.e8. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 31] [Article Influence: 31.0] [Reference Citation Analysis (0)] |
| 57. | Wunderlich CM, Ackermann PJ, Ostermann AL, Adams-Quack P, Vogt MC, Tran ML, Nikolajev A, Waisman A, Garbers C, Theurich S, Mauer J, Hövelmeyer N, Wunderlich FT. Obesity exacerbates colitis-associated cancer via IL-6-regulated macrophage polarisation and CCL-20/CCR-6-mediated lymphocyte recruitment. Nat Commun. 2018;9:1646. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 66] [Cited by in RCA: 122] [Article Influence: 15.3] [Reference Citation Analysis (0)] |
| 58. | Sun J, Shi L, Xu F, Sun H, Liu Y, Sun J, Zhou Q. Naringenin Inhibits Colorectal Cancer associated with a High-Fat Diet through Modulation of Gut Microbiota and IL-6/STAT3 Pathway. J Microbiol Biotechnol. 2025;35:e2412029. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 6] [Reference Citation Analysis (0)] |
| 59. | Guo S, Peng Y, Lou Y, Cao L, Liu J, Lin N, Cai S, Kang Y, Zeng S, Yu L. Downregulation of the farnesoid X receptor promotes colorectal tumorigenesis by facilitating enterotoxigenic Bacteroides fragilis colonization. Pharmacol Res. 2022;177:106101. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 28] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 60. | Režen T, Rozman D, Kovács T, Kovács P, Sipos A, Bai P, Mikó E. The role of bile acids in carcinogenesis. Cell Mol Life Sci. 2022;79:243. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 236] [Article Influence: 59.0] [Reference Citation Analysis (4)] |
| 61. | Cong J, Liu P, Han Z, Ying W, Li C, Yang Y, Wang S, Yang J, Cao F, Shen J, Zeng Y, Bai Y, Zhou C, Ye L, Zhou R, Guo C, Cang C, Kasper DL, Song X, Dai L, Sun L, Pan W, Zhu S. Bile acids modified by the intestinal microbiota promote colorectal cancer growth by suppressing CD8(+) T cell effector functions. Immunity. 2024;57:876-889.e11. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 112] [Cited by in RCA: 200] [Article Influence: 100.0] [Reference Citation Analysis (2)] |
| 62. | Ma C, Han M, Heinrich B, Fu Q, Zhang Q, Sandhu M, Agdashian D, Terabe M, Berzofsky JA, Fako V, Ritz T, Longerich T, Theriot CM, McCulloch JA, Roy S, Yuan W, Thovarai V, Sen SK, Ruchirawat M, Korangy F, Wang XW, Trinchieri G, Greten TF. Gut microbiome-mediated bile acid metabolism regulates liver cancer via NKT cells. Science. 2018;360:eaan5931. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1249] [Cited by in RCA: 1180] [Article Influence: 147.5] [Reference Citation Analysis (6)] |
| 63. | Chung L, Thiele Orberg E, Geis AL, Chan JL, Fu K, DeStefano Shields CE, Dejea CM, Fathi P, Chen J, Finard BB, Tam AJ, McAllister F, Fan H, Wu X, Ganguly S, Lebid A, Metz P, Van Meerbeke SW, Huso DL, Wick EC, Pardoll DM, Wan F, Wu S, Sears CL, Housseau F. Bacteroides fragilis Toxin Coordinates a Pro-carcinogenic Inflammatory Cascade via Targeting of Colonic Epithelial Cells. Cell Host Microbe. 2018;23:203-214.e5. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 486] [Cited by in RCA: 446] [Article Influence: 55.8] [Reference Citation Analysis (4)] |
| 64. | Burgueño JF, Fritsch J, González EE, Landau KS, Santander AM, Fernández I, Hazime H, Davies JM, Santaolalla R, Phillips MC, Diaz S, Dheer R, Brito N, Pignac-Kobinger J, Fernández E, Conner GE, Abreu MT. Epithelial TLR4 Signaling Activates DUOX2 to Induce Microbiota-Driven Tumorigenesis. Gastroenterology. 2021;160:797-808.e6. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 79] [Article Influence: 15.8] [Reference Citation Analysis (0)] |
| 65. | Zhu H, Li M, Bi D, Yang H, Gao Y, Song F, Zheng J, Xie R, Zhang Y, Liu H, Yan X, Kong C, Zhu Y, Xu Q, Wei Q, Qin H. Fusobacterium nucleatum promotes tumor progression in KRAS p.G12D-mutant colorectal cancer by binding to DHX15. Nat Commun. 2024;15:1688. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 28] [Cited by in RCA: 54] [Article Influence: 27.0] [Reference Citation Analysis (0)] |
| 66. | Yang Y, Li L, Xu C, Wang Y, Wang Z, Chen M, Jiang Z, Pan J, Yang C, Li X, Song K, Yan J, Xie W, Wu X, Chen Z, Yuan Y, Zheng S, Yan J, Huang J, Qiu F. Cross-talk between the gut microbiota and monocyte-like macrophages mediates an inflammatory response to promote colitis-associated tumourigenesis. Gut. 2020;70:1495-1506. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 99] [Cited by in RCA: 130] [Article Influence: 21.7] [Reference Citation Analysis (4)] |
| 67. | Geis AL, Fan H, Wu X, Wu S, Huso DL, Wolfe JL, Sears CL, Pardoll DM, Housseau F. Regulatory T-cell Response to Enterotoxigenic Bacteroides fragilis Colonization Triggers IL17-Dependent Colon Carcinogenesis. Cancer Discov. 2015;5:1098-1109. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 94] [Cited by in RCA: 158] [Article Influence: 14.4] [Reference Citation Analysis (3)] |
| 68. | Bi D, Zhu Y, Gao Y, Li H, Zhu X, Wei R, Xie R, Cai C, Wei Q, Qin H. Profiling Fusobacterium infection at high taxonomic resolution reveals lineage-specific correlations in colorectal cancer. Nat Commun. 2022;13:3336. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 32] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 69. | Li Q, Hu W, Liu WX, Zhao LY, Huang D, Liu XD, Chan H, Zhang Y, Zeng JD, Coker OO, Kang W, Ng SSM, Zhang L, Wong SH, Gin T, Chan MTV, Wu JL, Yu J, Wu WKK. Streptococcus thermophilus Inhibits Colorectal Tumorigenesis Through Secreting β-Galactosidase. Gastroenterology. 2021;160:1179-1193.e14. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 190] [Article Influence: 38.0] [Reference Citation Analysis (4)] |
| 70. | Wang X, Huycke MM. Colorectal cancer: role of commensal bacteria and bystander effects. Gut Microbes. 2015;6:370-376. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 36] [Article Influence: 3.6] [Reference Citation Analysis (4)] |
| 71. | Lavoie S, Chun E, Bae S, Brennan CA, Gallini Comeau CA, Lang JK, Michaud M, Hoveyda HR, Fraser GL, Fuller MH, Layden BT, Glickman JN, Garrett WS. Expression of Free Fatty Acid Receptor 2 by Dendritic Cells Prevents Their Expression of Interleukin 27 and Is Required for Maintenance of Mucosal Barrier and Immune Response Against Colorectal Tumors in Mice. Gastroenterology. 2020;158:1359-1372.e9. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 85] [Article Influence: 14.2] [Reference Citation Analysis (1)] |
| 72. | Min S, Zhang Y, Zhang H, Liu Q. Gut microbiota drives cancer evolution and therapy resistance. Mol Cancer. 2026;25:39. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 73. | Hu ZJ, Zhu HR, Jin YJ, Liu P, Yu XW, Zhang YR. Correlation between gut microbiota and tumor immune microenvironment: A bibliometric and visualized study. World J Clin Oncol. 2025;16:101611. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 74. | Ding T, Chen Q, Liu H, Zhang H, Sun Y, Zhao L, Gao Y, Wei Q. Single-cell RNA sequencing analysis reveals the distinct features of colorectal cancer with or without Fusobacterium nucleatum infection in PD-L1 blockade therapy. Heliyon. 2024;10:e37511. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 9] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 75. | Kvich L, Fritz BG, Zschach H, Terkelsen T, Raskov H, Høst-Rasmussen K, Jakobsen MR, Gheorghe AG, Gögenur I, Bjarnsholt T. Biofilms and core pathogens shape the tumor microenvironment and immune phenotype in colorectal cancer. Gut Microbes. 2024;16:2350156. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 19] [Reference Citation Analysis (0)] |
| 76. | Gur C, Ibrahim Y, Isaacson B, Yamin R, Abed J, Gamliel M, Enk J, Bar-On Y, Stanietsky-Kaynan N, Coppenhagen-Glazer S, Shussman N, Almogy G, Cuapio A, Hofer E, Mevorach D, Tabib A, Ortenberg R, Markel G, Miklić K, Jonjic S, Brennan CA, Garrett WS, Bachrach G, Mandelboim O. Binding of the Fap2 protein of Fusobacterium nucleatum to human inhibitory receptor TIGIT protects tumors from immune cell attack. Immunity. 2015;42:344-355. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1238] [Cited by in RCA: 1131] [Article Influence: 102.8] [Reference Citation Analysis (4)] |
| 77. | Ternes D, Tsenkova M, Pozdeev VI, Meyers M, Koncina E, Atatri S, Schmitz M, Karta J, Schmoetten M, Heinken A, Rodriguez F, Delbrouck C, Gaigneaux A, Ginolhac A, Nguyen TTD, Grandmougin L, Frachet-Bour A, Martin-Gallausiaux C, Pacheco M, Neuberger-Castillo L, Miranda P, Zuegel N, Ferrand JY, Gantenbein M, Sauter T, Slade DJ, Thiele I, Meiser J, Haan S, Wilmes P, Letellier E. The gut microbial metabolite formate exacerbates colorectal cancer progression. Nat Metab. 2022;4:458-475. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 72] [Cited by in RCA: 242] [Article Influence: 60.5] [Reference Citation Analysis (5)] |
| 78. | Wu L, Hollinshead KER, Hao Y, Au C, Kroehling L, Ng C, Lin WY, Li D, Silva HM, Shin J, Lafaille JJ, Possemato R, Pacold ME, Papagiannakopoulos T, Kimmelman AC, Satija R, Littman DR. Niche-Selective Inhibition of Pathogenic Th17 Cells by Targeting Metabolic Redundancy. Cell. 2020;182:641-654.e20. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 42] [Cited by in RCA: 102] [Article Influence: 17.0] [Reference Citation Analysis (0)] |
| 79. | Dubinsky V, Dotan I, Gophna U. Carriage of Colibactin-producing Bacteria and Colorectal Cancer Risk. Trends Microbiol. 2020;28:874-876. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 55] [Article Influence: 9.2] [Reference Citation Analysis (3)] |
| 80. | de Oliveira Alves N, Dalmasso G, Nikitina D, Vaysse A, Ruez R, Ledoux L, Pedron T, Bergsten E, Boulard O, Autier L, Allam S, Motreff L, Sauvanet P, Letourneur D, Kashyap P, Gagnière J, Pezet D, Godfraind C, Salzet M, Lemichez E, Bonnet M, Najjar I, Malabat C, Monot M, Mestivier D, Barnich N, Yadav P, Fournier I, Kennedy S, Mettouchi A, Bonnet R, Sobhani I, Chamaillard M. The colibactin-producing Escherichia coli alters the tumor microenvironment to immunosuppressive lipid overload facilitating colorectal cancer progression and chemoresistance. Gut Microbes. 2024;16:2320291. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 54] [Article Influence: 27.0] [Reference Citation Analysis (0)] |
| 81. | van Elsland DM, Duijster JW, Zhang J, Stévenin V, Zhang Y, Zha L, Xia Y, Franz E, Sun J, Mughini-Gras L, Neefjes J. Repetitive non-typhoidal Salmonella exposure is an environmental risk factor for colon cancer and tumor growth. Cell Rep Med. 2022;3:100852. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 16] [Reference Citation Analysis (0)] |
| 82. | Stévenin V, Coipan CE, Duijster JW, van Elsland DM, Voogd L, Bigey L, van Hoek AHAM, Wijnands LM, Janssen L, Akkermans JJLL, Neefjes-Borst A, Franz E, Mughini-Gras L, Neefjes J. Multi-omics analyses of cancer-linked clinical salmonellae reveal bacterial-induced host metabolic shift and mTOR-dependent cell transformation. Cell Rep. 2024;43:114931. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 3] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 83. | Zhou P, Zhang W, Liao Y, Zhai Z, Lu S, Jiang S, Hu W, Gong W. Colorectal cancer: the immune microenvironment and the gut microbiota - new perspectives, challenges, and opportunities. Int J Surg. 2025;111:5363-5386. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 6] [Reference Citation Analysis (0)] |
| 84. | Wang Y, Wiesnoski DH, Helmink BA, Gopalakrishnan V, Choi K, DuPont HL, Jiang ZD, Abu-Sbeih H, Sanchez CA, Chang CC, Parra ER, Francisco-Cruz A, Raju GS, Stroehlein JR, Campbell MT, Gao J, Subudhi SK, Maru DM, Blando JM, Lazar AJ, Allison JP, Sharma P, Tetzlaff MT, Wargo JA, Jenq RR. Fecal microbiota transplantation for refractory immune checkpoint inhibitor-associated colitis. Nat Med. 2018;24:1804-1808. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 642] [Cited by in RCA: 609] [Article Influence: 76.1] [Reference Citation Analysis (6)] |
| 85. | Zhang SL, Han B, Mao YQ, Zhang ZY, Li ZM, Kong CY, Wu Y, Chen GQ, Wang LS. Lacticaseibacillus paracasei sh2020 induced antitumor immunity and synergized with anti-programmed cell death 1 to reduce tumor burden in mice. Gut Microbes. 2022;14:2046246. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 72] [Article Influence: 18.0] [Reference Citation Analysis (4)] |
| 86. | Nan K, Zhong Z, Yue Y, Shen Y, Zhang H, Wang Z, Zhuma K, Yu B, Fu Y, Wang L, Sun X, Qu M, Chen Z, Guo M, Zhang J, Chu Y, Liu R, Miao C. Fasting-mimicking diet-enriched Bifidobacterium pseudolongum suppresses colorectal cancer by inducing memory CD8(+) T cells. Gut. 2025;74:775-786. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 34] [Cited by in RCA: 27] [Article Influence: 27.0] [Reference Citation Analysis (1)] |
| 87. | Renga G, Nunzi E, Pariano M, Puccetti M, Bellet MM, Pieraccini G, D'Onofrio F, Santarelli I, Stincardini C, Aversa F, Riuzzi F, Antognelli C, Gargaro M, Bereshchenko O, Ricci M, Giovagnoli S, Romani L, Costantini C. Optimizing therapeutic outcomes of immune checkpoint blockade by a microbial tryptophan metabolite. J Immunother Cancer. 2022;10:e003725. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 111] [Article Influence: 27.8] [Reference Citation Analysis (0)] |
| 88. | Wang Y, Jia J, Wang F, Fang Y, Yang Y, Zhou Q, Yuan W, Gu X, Hu J, Yang S. Pre-metastatic niche: formation, characteristics and therapeutic implication. Signal Transduct Target Ther. 2024;9:236. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 155] [Cited by in RCA: 152] [Article Influence: 76.0] [Reference Citation Analysis (4)] |
| 89. | Ma SX, Li L, Cai H, Guo TK, Zhang LS. Therapeutic challenge for immunotherapy targeting cold colorectal cancer: A narrative review. World J Clin Oncol. 2023;14:81-88. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 6] [Cited by in RCA: 8] [Article Influence: 2.7] [Reference Citation Analysis (1)] |
| 90. | Li Y, Arai S, Kato K, Iwabuchi S, Iwabuchi N, Muto N, Motobayashi H, Ebihara S, Tanaka M, Hashimoto S. The Potential Immunomodulatory Effect of Bifidobacterium longum subsp. longum BB536 on Healthy Adults through Plasmacytoid Dendritic Cell Activation in the Peripheral Blood. Nutrients. 2023;16:42. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 13] [Reference Citation Analysis (0)] |
| 91. | Si W, Liang H, Bugno J, Xu Q, Ding X, Yang K, Fu Y, Weichselbaum RR, Zhao X, Wang L. Lactobacillus rhamnosus GG induces cGAS/STING- dependent type I interferon and improves response to immune checkpoint blockade. Gut. 2022;71:521-533. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 48] [Cited by in RCA: 252] [Article Influence: 63.0] [Reference Citation Analysis (0)] |
| 92. | Wang X, Fang Y, Liang W, Wong CC, Qin H, Gao Y, Liang M, Song L, Zhang Y, Fan M, Liu C, Lau HC, Xu L, Li X, Song W, Wang J, Wang N, Yang T, Mo M, Zhang X, Fang J, Liao B, Sung JJY, Yu J. Fusobacterium nucleatum facilitates anti-PD-1 therapy in microsatellite stable colorectal cancer. Cancer Cell. 2024;42:1729-1746.e8. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 169] [Cited by in RCA: 145] [Article Influence: 72.5] [Reference Citation Analysis (0)] |
| 93. | Lee PJ, Hung CM, Yang AJ, Hou CY, Chou HW, Chang YC, Chu WC, Huang WY, Kuo WC, Yang CC, Lin KI, Hung KH, Chang LC, Lee KY, Kuo HP, Lu KM, Lai HC, Kuo ML, Chen WJ. MS-20 enhances the gut microbiota-associated antitumor effects of anti-PD1 antibody. Gut Microbes. 2024;16:2380061. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 22] [Reference Citation Analysis (0)] |
| 94. | Lam KC, Araya RE, Huang A, Chen Q, Di Modica M, Rodrigues RR, Lopès A, Johnson SB, Schwarz B, Bohrnsen E, Cogdill AP, Bosio CM, Wargo JA, Lee MP, Goldszmid RS. Microbiota triggers STING-type I IFN-dependent monocyte reprogramming of the tumor microenvironment. Cell. 2021;184:5338-5356.e21. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 214] [Cited by in RCA: 477] [Article Influence: 95.4] [Reference Citation Analysis (13)] |
| 95. | Goc J, Lv M, Bessman NJ, Flamar AL, Sahota S, Suzuki H, Teng F, Putzel GG; JRI Live Cell Bank, Eberl G, Withers DR, Arthur JC, Shah MA, Sonnenberg GF. Dysregulation of ILC3s unleashes progression and immunotherapy resistance in colon cancer. Cell. 2021;184:5015-5030.e16. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 176] [Article Influence: 35.2] [Reference Citation Analysis (5)] |
| 96. | Brooks JF 2nd, Behrendt CL, Ruhn KA, Lee S, Raj P, Takahashi JS, Hooper LV. The microbiota coordinates diurnal rhythms in innate immunity with the circadian clock. Cell. 2021;184:4154-4167.e12. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 89] [Cited by in RCA: 169] [Article Influence: 33.8] [Reference Citation Analysis (0)] |
| 97. | Churchill MJ, Pandeya A, Bauer R, Christopher T, Krug S, Honodel R, Smita S, Warner L, Mooney BM, Gibson AR, Mitchell PS, Tait Wojno ED, Rauch I. Enteric tuft cell inflammasome activation drives NKp46+ILC3 IL22 via PGD2 and inhibits Salmonella. J Exp Med. 2025;222:e20230803. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 9] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 98. | Hepworth MR, Monticelli LA, Fung TC, Ziegler CG, Grunberg S, Sinha R, Mantegazza AR, Ma HL, Crawford A, Angelosanto JM, Wherry EJ, Koni PA, Bushman FD, Elson CO, Eberl G, Artis D, Sonnenberg GF. Innate lymphoid cells regulate CD4+ T-cell responses to intestinal commensal bacteria. Nature. 2013;498:113-117. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 704] [Cited by in RCA: 633] [Article Influence: 48.7] [Reference Citation Analysis (4)] |
| 99. | Bruchard M, Geindreau M, Perrichet A, Truntzer C, Ballot E, Boidot R, Racoeur C, Barsac E, Chalmin F, Hibos C, Baranek T, Paget C, Ryffel B, Rébé C, Paul C, Végran F, Ghiringhelli F. Recruitment and activation of type 3 innate lymphoid cells promote antitumor immune responses. Nat Immunol. 2022;23:262-274. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 77] [Article Influence: 19.3] [Reference Citation Analysis (3)] |
| 100. | Fluckiger A, Daillère R, Sassi M, Sixt BS, Liu P, Loos F, Richard C, Rabu C, Alou MT, Goubet AG, Lemaitre F, Ferrere G, Derosa L, Duong CPM, Messaoudene M, Gagné A, Joubert P, De Sordi L, Debarbieux L, Simon S, Scarlata CM, Ayyoub M, Palermo B, Facciolo F, Boidot R, Wheeler R, Boneca IG, Sztupinszki Z, Papp K, Csabai I, Pasolli E, Segata N, Lopez-Otin C, Szallasi Z, Andre F, Iebba V, Quiniou V, Klatzmann D, Boukhalil J, Khelaifia S, Raoult D, Albiges L, Escudier B, Eggermont A, Mami-Chouaib F, Nistico P, Ghiringhelli F, Routy B, Labarrière N, Cattoir V, Kroemer G, Zitvogel L. Cross-reactivity between tumor MHC class I-restricted antigens and an enterococcal bacteriophage. Science. 2020;369:936-942. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 115] [Cited by in RCA: 316] [Article Influence: 52.7] [Reference Citation Analysis (0)] |
| 101. | Zitvogel L, Kroemer G. Cross-reactivity between microbial and tumor antigens. Curr Opin Immunol. 2022;75:102171. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 31] [Reference Citation Analysis (0)] |
| 102. | Zhou H, Wang L, Lin Z, Jiang C, Chen X, Wang K, Liu L, Shao L, Pan J, Li J, Zhang D, Wu J. Methylglyoxal from gut microbes boosts radiosensitivity and radioimmunotherapy in rectal cancer by triggering endoplasmic reticulum stress and cGAS-STING activation. J Immunother Cancer. 2023;11:e007840. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 33] [Article Influence: 11.0] [Reference Citation Analysis (0)] |
| 103. | Griffin ME, Espinosa J, Becker JL, Luo JD, Carroll TS, Jha JK, Fanger GR, Hang HC. Enterococcus peptidoglycan remodeling promotes checkpoint inhibitor cancer immunotherapy. Science. 2021;373:1040-1046. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 51] [Cited by in RCA: 289] [Article Influence: 57.8] [Reference Citation Analysis (3)] |
| 104. | Jani CT, Edwards K, Bhanushali C, Zheng X, Salazar AS, Lopes G, Watson DC. Leveraging beneficial microbiome-immune interactions via probiotic use in cancer immunotherapy. Front Immunol. 2025;16:1713382. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 4] [Article Influence: 4.0] [Reference Citation Analysis (5)] |
| 105. | Calabrò A, Drommi F, De Pasquale C, Navarra G, Carrega P, Bonaccorsi I, Ferlazzo G, Campana S. Microbiota and plasticity of antigen-presenting ILC3s: impact on antitumor immune response. Gut Microbes. 2024;16:2390135. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 7] [Reference Citation Analysis (0)] |
| 106. | Gurbatri CR, Radford GA, Vrbanac L, Im J, Thomas EM, Coker C, Taylor SR, Jang Y, Sivan A, Rhee K, Saleh AA, Chien T, Zandkarimi F, Lia I, Lannagan TRM, Wang T, Wright JA, Kobayashi H, Ng JQ, Lawrence M, Sammour T, Thomas M, Lewis M, Papanicolas L, Perry J, Fitzsimmons T, Kaazan P, Lim A, Stavropoulos AM, Gouskos DA, Marker J, Ostroff C, Rogers G, Arpaia N, Worthley DL, Woods SL, Danino T. Engineering tumor-colonizing E. coli Nissle 1917 for detection and treatment of colorectal neoplasia. Nat Commun. 2024;15:646. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 153] [Cited by in RCA: 129] [Article Influence: 64.5] [Reference Citation Analysis (0)] |
| 107. | Yang C, Su Q, Tang M, Luo S, Zheng H, Zhang X, Zhou X. Amplicon Sequencing of Single-Copy Protein-Coding Genes Reveals Accurate Diversity for Sequence-Discrete Microbiome Populations. Microbiol Spectr. 2022;10:e0210521. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 5] [Reference Citation Analysis (0)] |