Published online Apr 24, 2026. doi: 10.5306/wjco.v17.i4.115287
Revised: November 19, 2025
Accepted: March 2, 2026
Published online: April 24, 2026
Processing time: 190 Days and 14 Hours
The biological heterogeneity of breast cancer requires robust biomarkers to guide therapy. MicroRNAs (miRNAs) are pivotal post-transcriptional regulators, yet their longitudinal dynamics in matched tissue and plasma during neoadjuvant chemotherapy (NACT) remain poorly defined.
To evaluate the prognostic and predictive value of a panel of tumor-suppressive (miR-34a, -124a, -137) and oncogenic (miR-155, -373) miRNAs pre- and post-NACT.
We performed longitudinal profiling of five miRNAs in matched tumor and plasma from 38 patients [27 hormone receptor (HR)-positive; 11 HR-negative]. The study integrated matched tissue-plasma sampling, HR-stratified evaluation, and long-term survival follow-up. Prognostic associations were assessed via multivariate Cox regression, adjusting for age, HR status, and Ki-67.
High baseline miR-34a and miR-373 independently predicted improved overall survival (P < 0.001). miR-137 was the only miRNA significantly associated with pathological response, showing increased expression in good responders (P = 0.048). While clinical factors, such as older age and HR-positive status, correlated with favorable outcomes, elevated Ki-67 predicted worse overall survival (P = 0.003). NACT significantly modulated profiles, including a shift of miR-124a toward G2 tumors (P = 0.041). Notably, the prognostic value of baseline miRNAs diminished post-treatment, reflecting a convergence of miRNA expression patterns and unstable risk estimates in the post-NACT setting.
Baseline miR-34a and miR-373 are potent prognostic markers, while miR-137 predicts chemosensitivity. Treatment-induced remodeling diminishes the prognostic clarity of post-NACT miRNA landscapes.
Core Tip: This longitudinal study profiles matched tumor and plasma microRNAs dynamics in breast cancer patients before and after neoadjuvant chemotherapy. By integrating tissue-plasma signatures with survival data, stratified by hormone receptor status, we identified miR-34a and miR-373as independent prognostic markers at diagnosis. Furthermore, miR-137 was identified as candidate predictor of chemosensitivity. Our findings reveal that chemotherapy induces significant “molecular remodeling” and expression convergence across the cohort, which diminishes the prognostic clarity of markers post-treatment. Consequently, our data suggest that baseline profiling offers superior prognostic utility. These findings underscore the critical role of temporal context in microRNA-guided precision oncology and highlight the intricate interplay between systemic and intratumoral molecular landscapes.
- Citation: Ryspayeva D, Seyhan AA, Mu K, Liu M, Purcell C, MacDonald WJ, Halytskiy V, Drevytska T, Inomistova M, Khranovska N, Potorocha O, Taran L, Sumkina O, Smolanka Sr I, El-Deiry WS. Longitudinal microRNA profiles in breast cancer tissue and plasma: Associations with hormone receptors, response, and survival. World J Clin Oncol 2026; 17(4): 115287
- URL: https://www.wjgnet.com/2218-4333/full/v17/i4/115287.htm
- DOI: https://dx.doi.org/10.5306/wjco.v17.i4.115287
Breast cancer (BC) remains the most common malignancy among women worldwide and a leading cause of cancer-related morbidity[1]. It accounts for approximately 32% of all new cancer cases in women and represents the leading cause of cancer death in women under 50 years[1]. Early BC treatment is complicated by substantial biological heterogeneity, resulting in variable therapeutic responses across molecular subtypes like hormone receptor (HR)-positive, human epidermal growth factor receptor 2 (HER2)-positive, and triple-negative disease[2]. This diversity limits the effectiveness of uniform treatment strategies and underscores the need for predictive biomarkers to guide therapy selection. Despite advances in diagnostic and prognostic signatures for invasive BC, reliable minimally invasive biomarkers for early detection remain lacking.
MicroRNAs (miRNAs) have emerged as promising diagnostic, prognostic, and therapeutic tools in oncology[3,4]. These short (approximately 22 nucleotide) non-coding RNAs regulate gene expression by binding to complementary sequences in target RNAs, primarily within the 3’ untranslated region of mRNAs, leading to transcriptional and post-transcriptional regulation[5-8]. Aberrant miRNA expression, arising from altered levels, mutations, or defects in biogenesis, represents a key mechanism in cancer initiation and progression[9,10]. Dysregulation of miRNAs can result from both genetic alterations and epigenetic mechanisms, including DNA hypermethylation and hypomethylation, as well as aberrant histone modifications[3,11].
Functionally, cancer-associated miRNAs are broadly categorized into two groups: Oncogenic miRNAs (oncomiRs), which are typically upregulated and promote tumor progression, and tumor-suppressive miRNAs, which inhibit carcinogenesis and are frequently downregulated. Some act in a context-dependent manner, functioning as either oncogenes or tumor suppressors depending on tissue type and disease state[11,12].
Of particular clinical interest, circulating miRNAs are stable in body fluids and can be identified non-invasively, making them attractive biomarkers[13-16]. Concordance between tumor and circulating miRNA profiles supports their tumor-derived origin[17-19]. Several studies have demonstrated that circulating miRNAs often mirror expression patterns observed in primary tumors, suggesting that they are actively secreted by cancer cells through exosomes, microvesicles, or protein-RNA complexes[9,20]. Their stability in plasma and serum further enhances their potential use in diagnosis, prognosis, and treatment monitoring[21,22].
In BC, numerous miRNAs are differentially expressed across molecular subtypes and are implicated in tumor progression, resistance to therapy, and metastasis[12,15,23-31]. Using TargetScan software[32], we identified conserved miRNA binding sites in transcripts of genes critical for BC development and progression. Based on these analyses, miR-34a, miR-124a, miR-137, miR-155, and miR-373 were selected, as their predicted targets are involved in key oncogenic pathways, representing both tumor-suppressive and oncogenic miRNAs with established relevance in BC biology[24,26,28,29]. In this study, we focused on tumor-suppressors such as miR-34a, miR-124a, miR-137, and tumor-promoting oncomiRs such as miR-155 and miR-373, to access their expression in matched tumor, adjacent non-tumor tissue, and plasma samples collected before and after neoadjuvant chemotherapy (NACT). The goal was to clarify their functional significance and evaluate their potential as predictive or prognostic biomarkers.
Our prospective study included 38 patients with a confirmed diagnosis of invasive BC (Table 1). Clinical data and tumor specimens were collected at the National Cancer Institute of Ukraine from 2015 to 2017. Among them, 34 patients provided matched pairs of samples, consisting of pre-chemotherapy biopsies and post-NACT surgical resections (Supplementary Figure 1). The mean age of the patients was 53.5 ± 11.2 years (range: 23-75 years), with a median age of 54 years.
| Characteristics | Primary tumor, n = 38 | Post-NACT, n = 35 | P value | |
| Stage (AJCC) | 0 | 0 | 3 | 0.21 |
| I | 0 | 1 | ||
| IIA | 5 | 7 | ||
| IIB | 15 | 8 | ||
| IIIA | 10 | 8 | ||
| IIIB | 7 | 7 | ||
| IIIC | 1 | 1 | ||
| Category T | 0 | 0 | 3 | 0.54 |
| 1 | 2 | 2 | ||
| 2 | 27 | 21 | ||
| 3 | 2 | 2 | ||
| 4 | 7 | 7 | ||
| Category N | 0 | 4 | 11 | 0.14 |
| 1 | 17 | 11 | ||
| 2 | 16 | 12 | ||
| 3 | 1 | 1 | ||
| Ki-67 index, median (interquartile range) | 25 (15.0-38.8) | 22.5 (12.9-37.85) | 0.645 | |
BC subtypes are classified as luminal A (HR+/HER2-), luminal B (HR+/HER2+), HER2-positive (HR-/HER2+), and triple-negative (ER-/progesterone receptor-/HER2-). In our study, we grouped patients into two categories: HR-positive, which included HR+/HER2- and HR+/HER2+, and HR-negative, which included HR-/HER2+ and triple-negative tumors.
Among the 38 patients (Table 1), 27 were HR-positive and 11 were HR-negative disease. The HR-positive group included luminal A (n = 7), luminal B (n = 13), and luminal B (HER2+) (n = 7). The HR-negative group included triple-negative (n = 7) and HER2-positive (n = 4) subtypes. Patients received 2 cycles to 4 cycles of NACT consisting of combinations of doxorubicin, cyclophosphamide, fluorouracil, and/or taxanes[33]. All patients provided written informed consent, and the study was approved by the local ethics committee.
Pathological response to NACT was evaluated using the Miller-Payne grading (MPG) system, a five-point scale that assesses changes in tumor cellularity between pretreatment biopsy and surgical specimens after neoadjuvant therapy. Grade 1 indicates no response, grades 2-4 represent a partial pathological response, and grade 5 indicates a complete pathological response[33]. For our analysis, grades 4 and 5 were classified as positive treatment response, while grades 2 and 3 were considered a partial response.
Tumor tissue was obtained at diagnostic biopsy and after surgery, with matched non-tumor tissue collected from the corresponding surgical specimens. Peripheral blood was collected at three points: Pre-treatment, post-NACT, and post-treatment (after surgery and adjuvant chemotherapy). Plasma was separated immediately and stored at -70 °C, while tissue samples were stabilized in DNA/RNA shield (Eppendorf) and stored at -70 °C for subsequent miRNA profiling.
Reverse transcription quantitative polymerase chain reaction (RT-qPCR) assays were conducted to measure the expression levels of miR-34a, miR-124a, miR-137, miR-155, and miR-373. Expression analysis was performed on tumor tissue samples collected during biopsy and from the targeted area excised during surgery.
Total RNA, including miRNA, was isolated using the NucleoSpin® miRNA kit (Macherey-Nagel, Germany) following the manufacturer's instructions. RT-qPCR was performed using the 7500 Real-Time Polymerase Chain Reaction System (Applied Biosystems, United States) to evaluate miRNA expression levels. Reactions were prepared with TaqMan® Universal Polymerase Chain Reaction Master Mix (Applied Biosystems, United States), and specific TaqMan® miRNA primers (hsa-miR-34a, mmu-miR-124a, mmu-miR-137, hsa-miR-155, and hsa-miR-373). All reactions were conducted in accordance with the manufacturer’s protocols. MiRNA expression data were analyzed using the comparative Ct method. Expression levels of target miRNAs were normalized to the endogenous control RNU6B (TaqMan® miRNA Control Assay, Applied Biosystems, United States) to account for differences in cDNA input. Relative expression levels of miRNAs were calculated using the ΔCt method, defined as the difference between the Ct value of the target miRNA and that of the endogenous control. Expression was then represented as 2-ΔCt.
Patients were stratified into high- and low-expression groups based on the median expression level of each miRNA. All statistical analyses were performed using R software. Missing data were handled using conditional mean imputation.
The distributions of American Joint Committee on Cancer (AJCC) stage, T category, and N category were compared between the primary tumor and post-NACT groups. Because these variables are ordinal, the Cochran-Armitage trend test was used to evaluate whether a systematic distributional shift (i.e., downstaging) occurred following treatment. This test is appropriate for ordered categorical data and provides complementary information to exact tests and rank-based approaches, particularly in small-sample settings. Continuous variables, such as the Ki-67 index, were analyzed using the Wilcoxon rank-sum test for unpaired comparisons and the Wilcoxon signed-rank test for paired Ki-67 measurements.
Differences in miRNA expression between groups were assessed using the Wilcoxon signed-rank test for paired samples, the Wilcoxon rank-sum test for unpaired samples, or linear mixed-effects models, as appropriate. For visualization purposes, boxplot values were shifted by the minimum value in the dataset to ensure that all plotted values were above zero.
Survival outcomes were analyzed using the Kaplan-Meier method, with differences between groups assessed by the log-rank test. Follow-up duration was estimated using the reverse Kaplan-Meier method, which estimates time to censoring and provides an unbiased measure of follow-up. Survival curves display the estimated survival probabilities over time (X-axis: Months; Y-axis: Probability, 0-1), with tick marks indicating censored observations and shaded areas representing 95% confidence intervals (CIs). Additionally, cumulative hazard functions were estimated using the Nelson-Aalen estimator, showing the accumulation of hazard over time.
Cox proportional hazards regression models were used to evaluate associations between miRNA expression and survival outcomes. Both univariate and multivariate analyses were conducted, with multivariate models adjusted for potential confounders. Final Cox proportional hazards models were selected based on the Akaike Information Criterion and model diagnostics were performed to verify the proportional hazards assumption and overall model fit. Because age violated the proportional hazards assumption, its association with survival was further examined using accelerated failure time (AFT) models. Specifically, Weibull AFT models were fitted to characterize increasing, decreasing, or constant hazard patterns and to provide more accurate estimates of the effect of age on survival. All statistical significance was assessed at a two-sided α = 0.05.
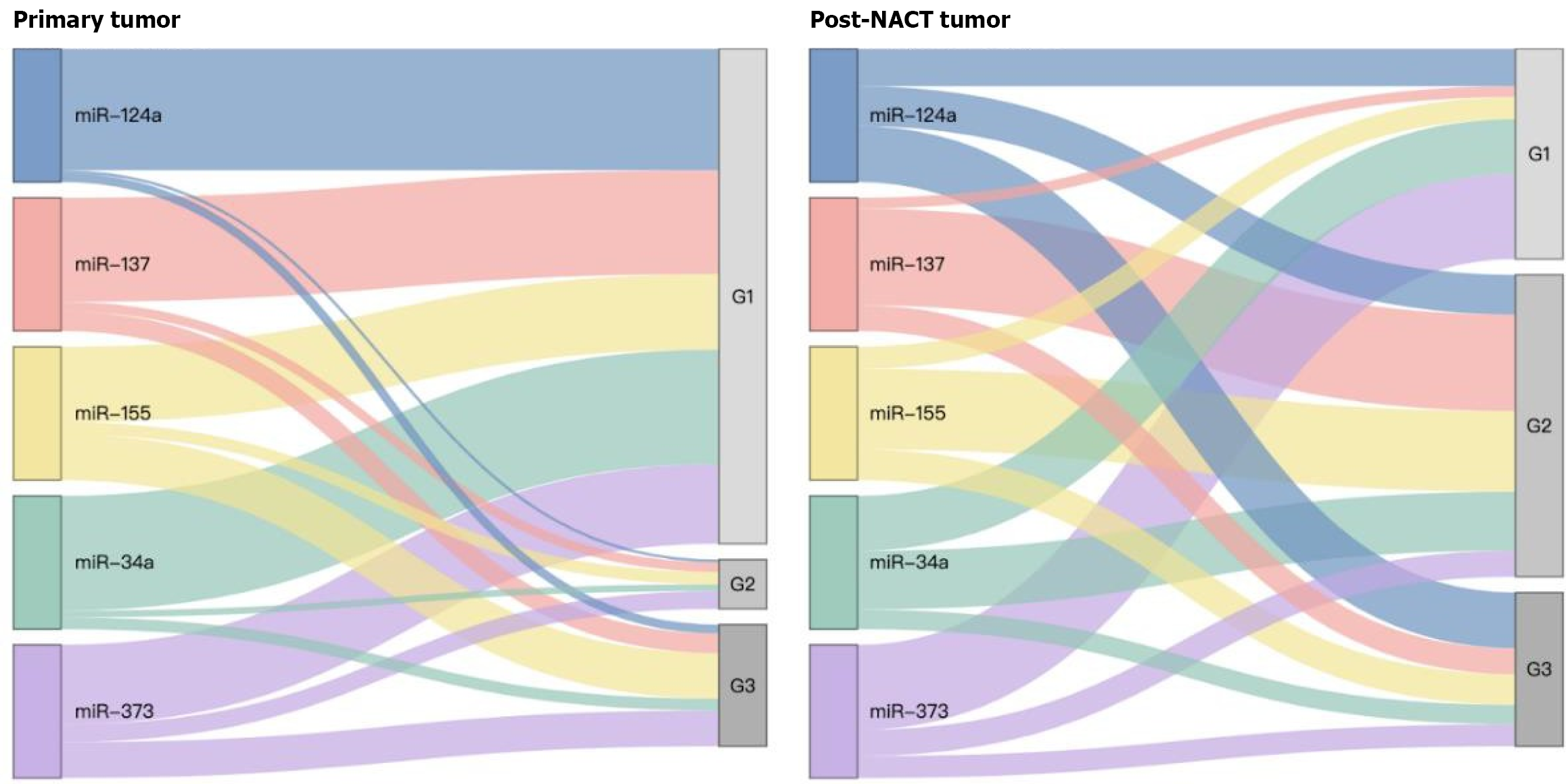
For the Sankey diagrams, median expression values of each miRNA in primary untreated tumor and post-NACT samples were transformed into proportional log2 fold changes, which defined the flow widths and reflected relative expression levels. Diagrams were generated in R using the networkD3, htmlwidgets, dplyr, and tidyr packages to visualize the distribution of selected miRNAs (miR-34a, miR-124a, miR-137, miR-155, and miR-373). Two sets of diagrams were created: One illustrating expression flows across tumor differentiation grades (G1-G3) in primary and post-NACT samples, and another showing distributions across tumor subtypes (HR-positive and HR-negative). For each miRNA-grade or miRNA-subtype pair, median values were normalized so that link widths reflected relative contributions. Nodes on the left represented individual miRNAs, while nodes on the right represented tumor grades or subtypes. Primary and post-NACT diagrams were displayed side by side to highlight treatment-related changes in expression patterns.
NACT was associated with trends, but not statistically significant changes in tumor stage and nodal status (Table 1). Although a shift toward lower AJCC stage and an increased proportion of patients with N0 nodal status was observed after NACT, these differences did not reach statistical significance (Cochran-Armitage test P = 0.1328).
In the overall cohort (Table 1), median Ki-67 expression showed a modest decrease following NACT [25.0 (interquartile range: 15.0-38.8) vs 22.5 (interquartile range: 12.9-37.85)], which was not statistically significant (P = 0.645). Consistent with this finding, analysis of twenty-four paired samples (Supplementary Figure 2) confirmed the absence of a significant difference between pre- and post-treatment Ki-67 levels (P = 0.587).
When stratified by HR status, HR-positive tumors showed relatively lower Ki-67 expression both before and after NACT (mean ± SD: 23.95 ± 16.36 vs 22.48 ± 16.81 respectively), whereas HR-negative tumors consistently demonstrated higher Ki-67 levels (44.24 ± 22.67 vs 44.05 ± 22.35 before and after NACT, respectively), with no significant treatment-associated changes observed within either subgroup.
The Sankey diagram presented in Figure 1 illustrates the distribution of five selected miRNAs (miR-124a, miR-137, miR-155, miR-34a, and miR-373) across tumor grades (G1, G2, and G3) in primary tumors (left) and post-NACT tumors (right). In primary tumors, G1 (well differentiated) predominates for most miRNAs. miR-124a, miR-137, miR-155, and especially miR-34a show strong associations with this grade, while miR-373 demonstrates a more dispersed pattern across all grades, including a notable proportion in G3.
Following NACT, G2 (moderately differentiated) becomes the predominant grade, with miR-137, miR-155, and miR-34a showing strong flows into this category. The flow from miR-124a to G2 increases significantly (adjusted P = 0.041), reflecting a treatment-induced expression shift, whereas its change toward G3 is not significant (adjusted P = 0.305). miR-34a displays a more dispersed pattern across all grades, with increasing flow contributions to G2 and G3. For miR-373, the flow into G3 decreases post-NACT.
Overall, the diagram highlights a treatment-induced redistribution of tumor grades, with reduction of G1 and enrichment of G2 and G3. These findings support the presence of grade-associated miRNA expression patterns, particularly for miR-124a and miR-34a, that may reflect tumor evolution and therapeutic response.
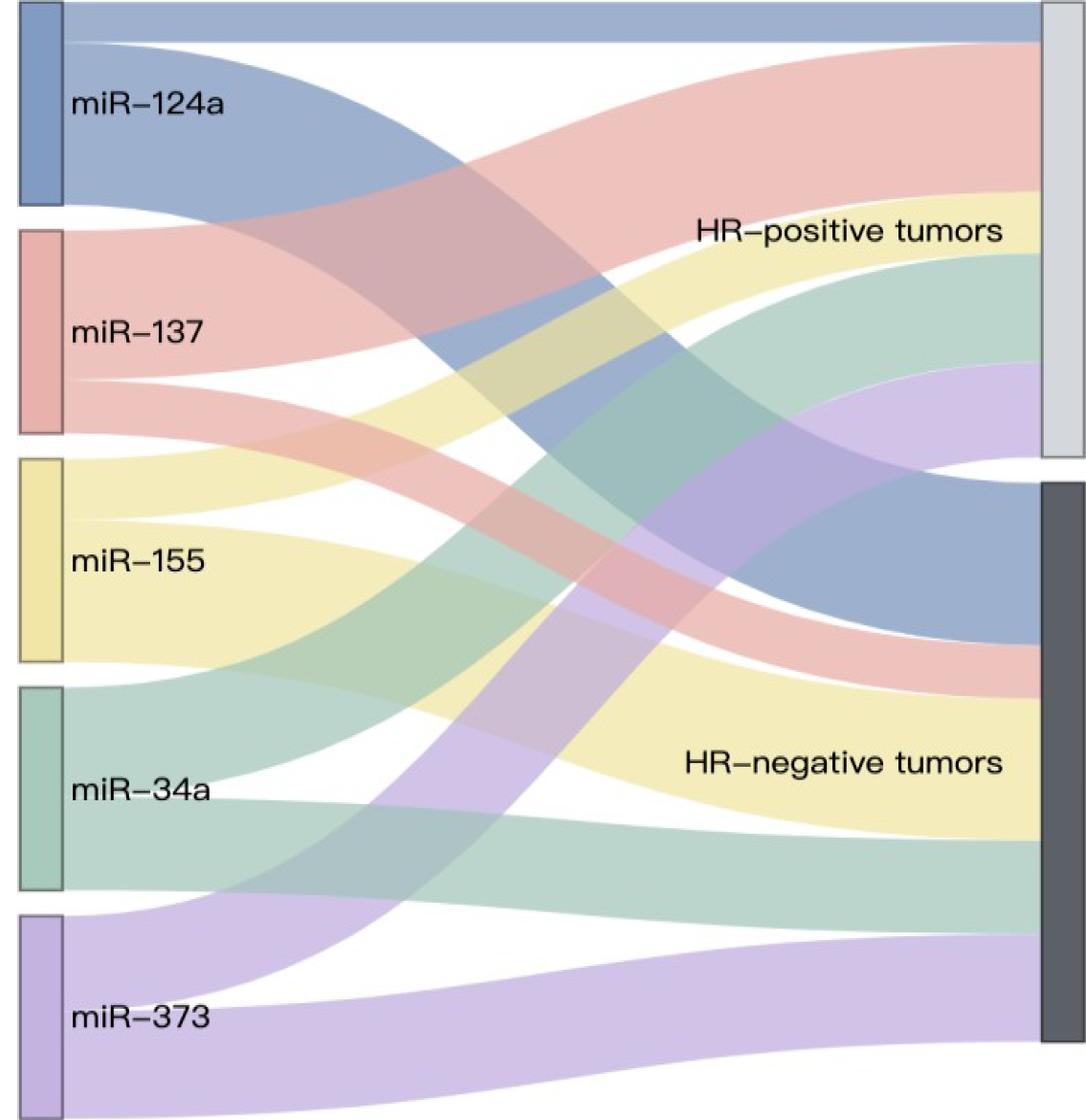
To further explore clinical subtypes, a separate Sankey diagram was constructed using normalized miRNA expression values to compare HR-positive and HR-negative primary tumors (Figure 2). This visualization shows that miR-137 is predominantly expressed in HR-positive tumors, whereas miR-124a and miR-155 are more enriched in HR-negative tumors. miR-34a is relatively evenly distributed, with a slight predominance in HR-positive tumors, while miR-373 is nearly balanced between subtypes, showing a minor shift toward HR-negative disease.
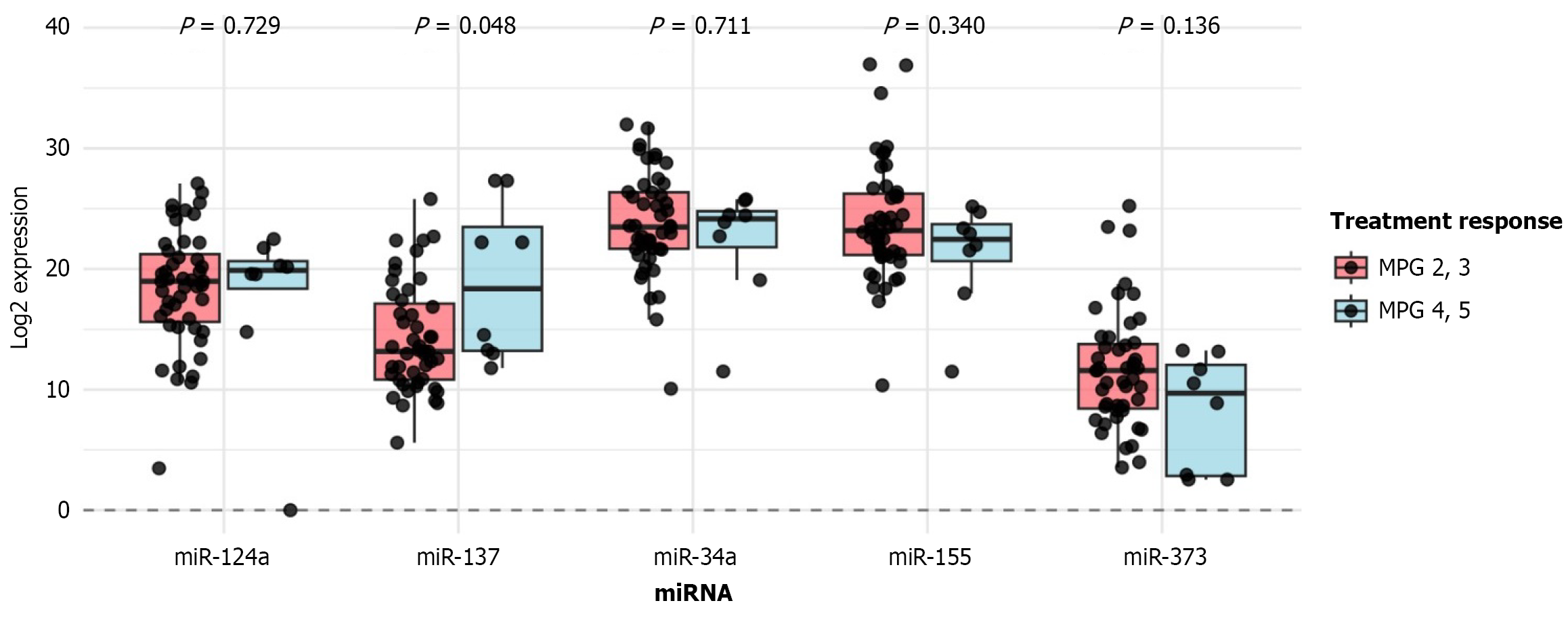
We evaluated the relationship between post-treatment miRNA expression and pathological response to NACT using the MPG system, in which only grades 2, 3, 4, and 5 were observed among our patients. Expression levels of five selected miRNAs (miR-124a, miR-137, miR-34a, miR-155, and miR-373) were compared between patients with partial response (MPG 2-3) and those with near complete or complete response (MPG 4-5), the latter being defined as a positive response.
As shown in the boxplots (Figure 3), only miR-137 demonstrated a statistically significant difference in expression between the two response groups (P = 0.048) with higher median expression observed in patients achieving an MPG score of 4-5. For other miRNAs, no statistically significant associations with treatment response were detected (miR-124a: P = 0.729; miR-34a: P = 0.711; miR-155: P = 0.340; miR-373: P = 0.136). Overall, these results suggest that while miR-137 may be associated with improved pathological response to NACT, the expression patterns of the other tested miRNAs did not show consistent or significance associations with MPG categories.
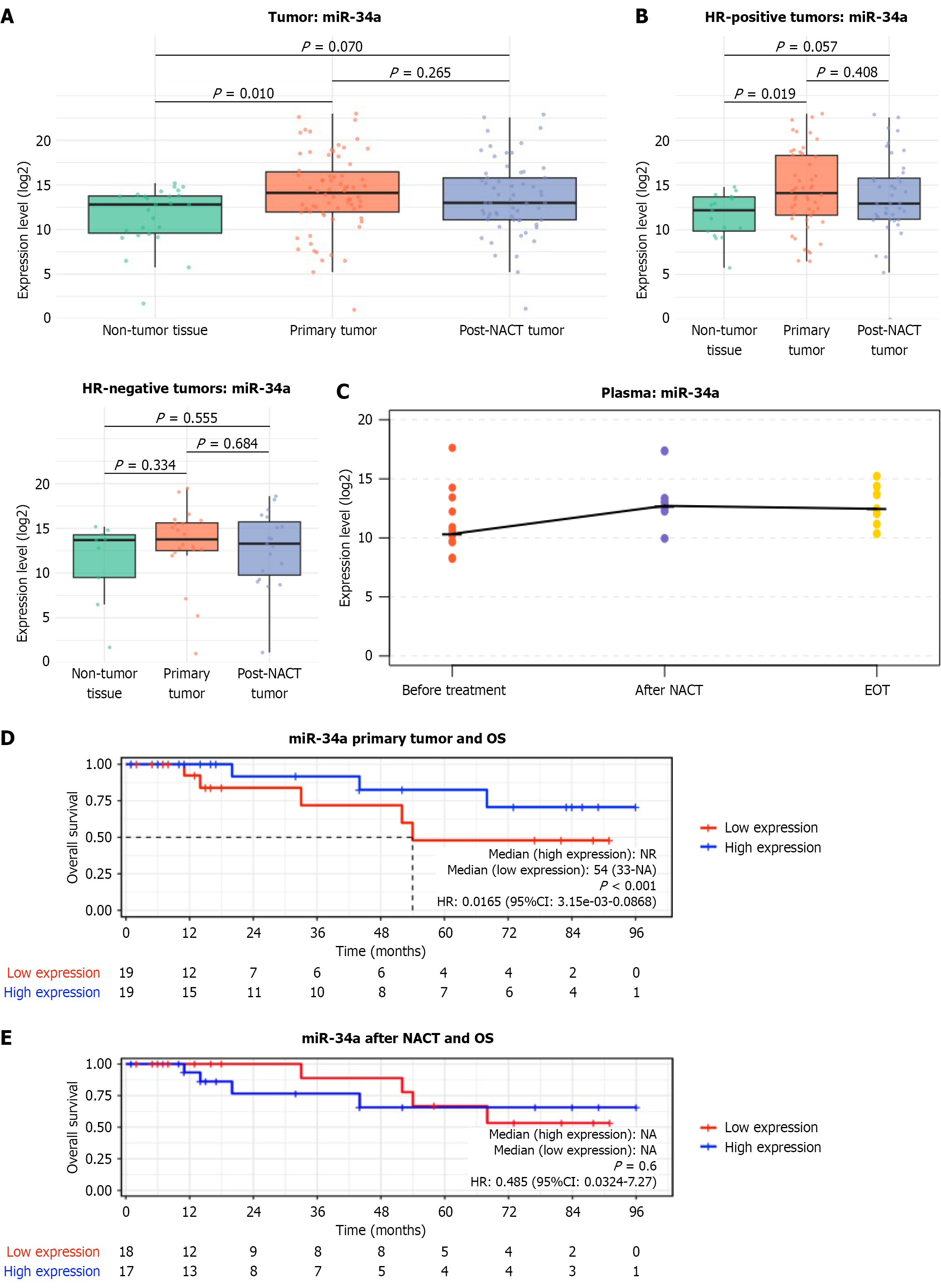
miR-34a (Figure 4A), a well-characterized tumor suppressor miRNA, was significantly upregulated in primary tumor tissue compared to adjacent non-tumor tissue (P = 0.010). Following NACT, miR-34a expression exhibited a downward trend, resulting in levels that were no longer significantly different non-tumor tissue (P = 0.070).
Stratification by HR status revealed that this elevated expression was specific to HR-positive tumors (P = 0.019), whereas HR-negative cases showed negligible differences (Figure 4B), consistent with previous findings[34]. While the post-NACT decline in HR-positive tumors was only borderline significant (P = 0.057), likely due to the limited sample size, the shift in significance within the survival analysis suggests this decrease is clinically meaningful, the change in significance observed in the survival analysis pre- and post-NACT suggests that the decrease in miR-34a expression level is clinically meaningful. Conversely, in blood samples (Figure 4C), chemotherapy induced an elevation in circulating miR-34a levels. These fluctuations, however, warrant cautious interpretation given the small cohort and limited number of events.
To assess the clinical relevance of miR-34a, overall survival (OS) was analyzed according to miR-34a expression levels. Kaplan-Meier analysis showed a separation of survival curves according to miR-34a expression in primary untreated tumors (Figure 4D). Patients with high miR-34a expression in primary breast tumors had improved OS compared with those with low expression. The low-expression group reached a median survival of 54 months, whereas the high-expression group did not reach a median survival, as more than half of the patients remained alive. Censoring and the number at risk at 96 months (0 vs 1) further highlight the survival advantage (hazard ratios = 0.0165, 95%CI: 3.15e-03 to 0.0868, P < 0.001).
After stratification by HR status, this association was attenuated and did not reach statistical significance within individual subgroups (Supplementary Figure 3). Nevertheless, a trend toward improved OS was observed among HR-positive patients with higher miR-34a expression, whereas no clear association was detected in HR-negative patients. The loss of statistical significance after stratification is likely due to reduced sample size and limited statistical power.
Following NACT, the association between miR-34a expression and survival differed from that observed in primary tumors (Figure 4E). In post-NACT tumor samples, miR-34a expression was not significantly associated with OS. A non-significant trend toward improved survival was observed in patients with lower miR-34a expression after NACT (hazard ratios = 0.485, 95%CI: 0.0324-7.27, P = 0.6).
Further stratification of post-NACT tumors by HR status revealed no significant differences in OS between high and low miR-34a expression groups in either HR-positive (P = 0.505) or HR-negative tumors (P = 0.887), indicating that miR-34a expression after NACT does not retain prognostic value.
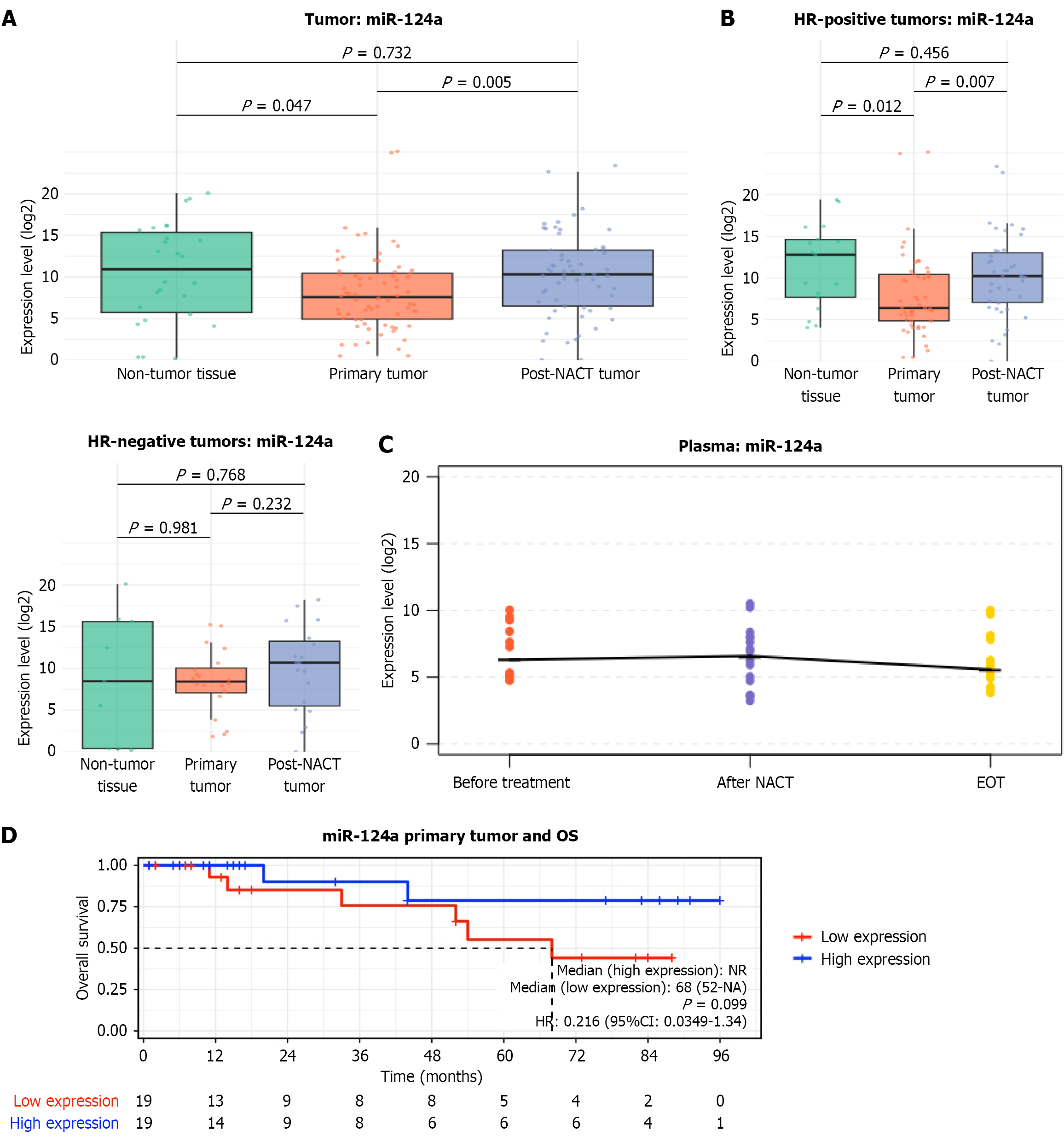
The expression of tumor-suppressor miR-124a in primary tumor tissue (Figure 5A) was significantly decreased compared with the corresponding adjacent normal tissue (P = 0.047), with a more pronounced reduction observed in the HR-positive cohort (P = 0.012; Figure 5B). Following chemotherapy, miR-124a levels were significantly increased in entire cohort (P = 0.005), primarily driven by changes in HR-positive patients (P = 0.007). After NACT, miR-124a expression in these tumors was no longer significantly different from adjacent normal tissue (P = 0.456). HR-negative patients demonstrated a similar trend toward increased expression, but the difference did not reach statistical significance (Figure 5B). In plasma, a trend toward decreased miR-124a was observed after treatment, particularly following surgery and adjuvant chemotherapy, but the changes were not statistically significant, likely due to the limited sample size (Figure 5C).
Kaplan-Meier analysis suggested a trend toward improved OS in patients with high miR-124a expression in primary untreated tumors (Figure 5D). Median OS was not reached in the high-expression group, whereas the low-expression group had a median survival of 68 months, consistent with a favorable survival pattern (hazard ratios = 0.216, 95%CI: 0.0349-1.34; P = 0.099). Although the difference did not reach conventional statistical significance, the hazard ratio indicates an approximately 78% reduction in the risk of death for patients with high miR-124a, supporting a potential protective role. This trend was more pronounced in HR-positive patients (Supplementary Figure 4).
Following NACT, Kaplan-Meier curves for OS showed substantial overlap, with no significant difference between high- and low-expression groups (hazard ratios = 1.67, 95%CI: 0.179-15.5; P = 0.654), and a median OS was not reached in either group (Supplementary Figure 5). The absence of a reached median OS in the low-expression group after NACT suggests a shift in the survival pattern, indicating that NACT may provide a benefit by extending survival in patients with initially low miR-124a expression.
These observations align with the post-NACT redistribution of miR-124a shown in the Sankey diagram (Figure 1), reflecting treatment-induced tumor remodeling or selection. Together, the data indicate that NACT can modulate miR-124a expression and tumor biology, potentially contributing to survival benefit even in patients with low baseline miR-124a levels.
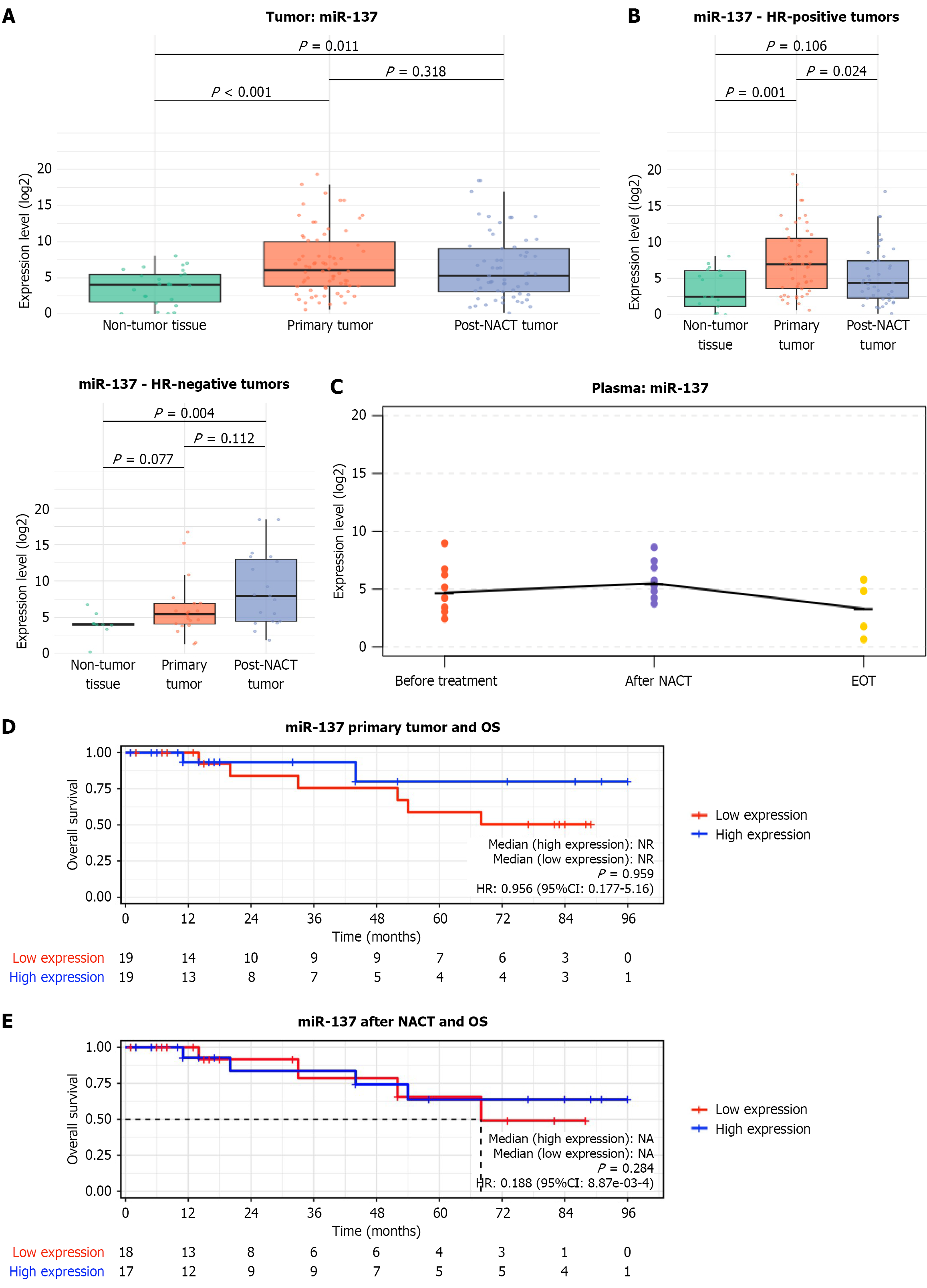
miR-137 expression was significantly increased in primary breast tumor tissue compared with matched non-tumor tissue (P < 0.001; Figure 6A) and remained significantly elevated relative to non-tumor tissue after NACT (P = 0.011), although a decreasing trend was observed following treatment.
Stratification by HR status revealed distinct patterns. In HR-positive tumors, miR-137 expression was significantly reduced after NACT compared with primary tumors (P = 0.024; Figure 6B) and was no longer significantly different from non-tumor tissue (P = 0.106). In contrast, HR-negative tumors exhibited persistently elevated miR-137 expression following NACT, remaining significantly higher than non-tumor tissue P = 0.004), with no significant change between primary and post-NACT tumors (P = 0.112). In plasma (Figure 6C), miR-137 levels showed a trend toward increased expression following NACT, followed by a decline after surgery and adjuvant chemotherapy, ultimately reaching levels below those observed at baseline.
Kaplan-Meier analysis was performed to assess the association between miR-137 expression and OS in patients with primary breast tumors. As shown in Figure 6D, no statistically significant difference in OS existed between the high- and low-expression groups (P = 0.959). Survival curves largely overlapped throughout the follow-up period. The effect estimate was 0.956 with a wide 95%CIs (0.177-5.16), indicating no significant association between miR-137 expression and survival. A median OS was not reached in either group during the observation period.
Stratification by HR status similarly revealed no significant associations (Supplementary Figure 6). In the HR-positive subgroup, a non-significant trend toward improved OS was observed in patients with high miR-137 expression (hazard ratios = 0.533, 95%CI: 0.0963-2.95; P = 0.460). In the HR-negative subgroup, meaningful statistical comparisons were not feasible due to small sample size and limited follow-up, with no patients remaining at risk in the high-expression group beyond 24 months.
After NACT (Figure 6E), Kaplan-Meier curves for high and low miR-137 expression overlapped and crossed multiple times, with no consistent separation between groups. Although the estimated hazard suggested a potential protective effect (hazard ratios = 0.188), this association did not reach statistical significance, indicating no clear survival advantage associated with miR-137 expression in the post-NACT cohort.
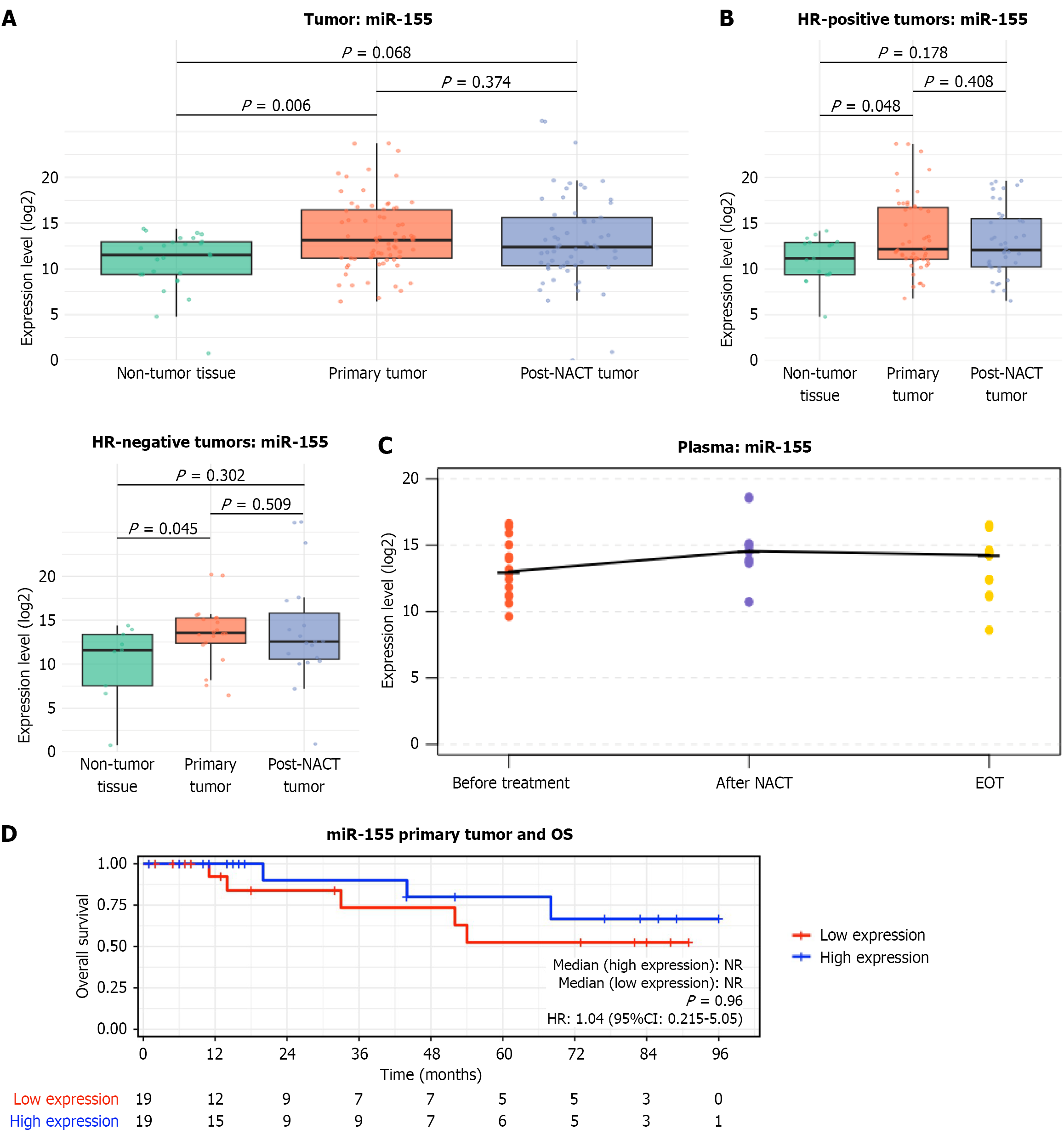
miR-155, an oncogenic miRNA, was significantly overexpressed in primary breast tumor tissue at baseline compared with matched adjacent non-tumor tissue (P = 0.006; Figure 7A). This upregulation was observed in both HR-positive and HR-negative subgroups (P = 0.048 and P = 0.045, respectively; Figure 7B), with a more pronounced increase in HR-negative tumors.
After NACT, miR-155 expression in residual tumor tissue showed a decreasing trend but remained higher than in non-tumor tissue, although this difference did not reach statistical significance (P = 0.068; Figure 7A). Similar patterns were observed in HR-positive and HR-negative subgroups, with no significant differences relative to non-tumor tissue (P = 0.178 and P = 0.302, respectively; Figure 7B).
In plasma, miR-155 (Figure 7C) exhibited a non-significant increase after NACT and remained elevated compared with baseline, with a trend toward reduction at treatment completion. Although both tumor tissue and plasma showed treatment-associated changes in miR-155 levels, the dynamics of plasma did not mirror those observed in tumor tissue.
Stratification of BC patients according to the median tumor miR-155 expression in the primary tumor revealed no statistically significant association with OS (P = 0.960; Figure 7D). The Cox model yielded a relative risk estimate of 1.04 (95%CI: 0.215-5.05) indicating no meaningful difference between groups. The wide CIs reflect limited precision due to the small sample size and low number of events. Visual assessment of the Kaplan-Meier curves suggested a modest separation favoring the high miR-155 group; however, this trend was not statistically significant.
The favorable trend was more pronounced in the HR-positive subgroup (Supplementary Figure 7), supporting the potential prognostic relevance of miR-155 in this specific patient population. Although these associations did not reach statistical significance, the survival curves clearly indicated a favorable trend that warrants further validation in larger cohorts of HR-positive patients.
Following NACT, survival differences were no longer apparent. Notably, as observed with other miRNAs, the effect of NACT appeared to attenuate these survival differences. In post-NACT samples, the survival curves for high and low miR-155 expression became nearly identical (P = 0.999; Supplementary Figure 8). The resulting indeterminate risk estimate (3.14-10; 95%CI: 0.00e-Inf) reflects model instability due to the very low number of events and the near-complete overlap of survival curves in the post-treatment setting, indicating that the model could not reliably estimate relative risk.
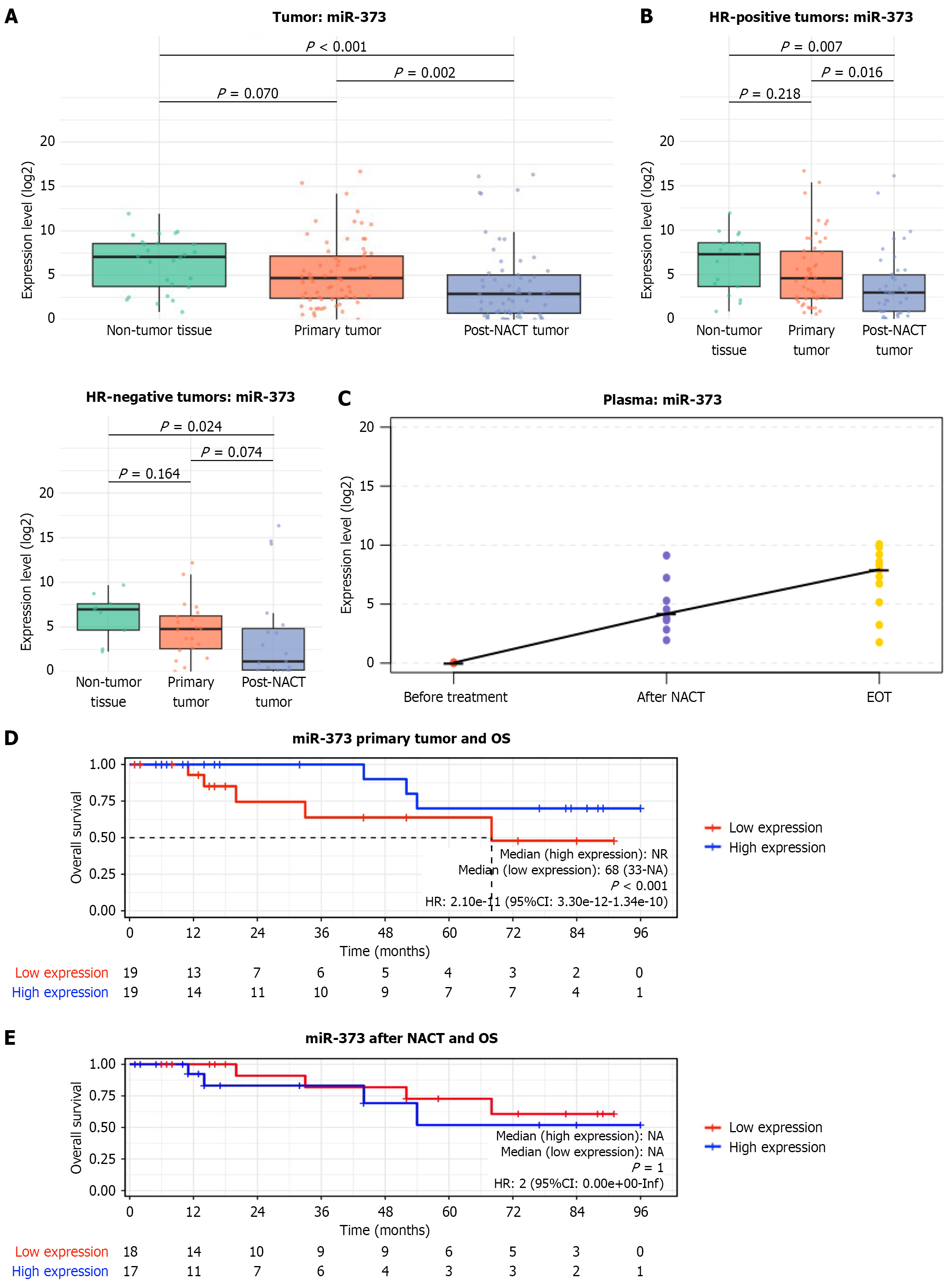
In the overall cohort, miR-373 expression showed a borderline decrease in primary untreated tumors compared with adjacent non-tumor tissue (P = 0.070; Figure 8A). Following NACT, tumor miR-373 levels decreased significantly (P = 0.002). Stratification by HR status revealed a decrease in miR-373 expression following NACT in both subgroups. This reduction was statistically significant in HR-positive tumors (P = 0.016) and reached borderline significance in HR-negative tumors (P = 0.074) (Figure 8B). Furthermore, post-NACT tumor samples exhibited significantly lower miR-373 expression levels compared with non-tumor tissue in both HR-positive (P = 0.007) and HR-negative (P = 0.024) groups. In contrast, plasma miR-373 levels increased during treatment, demonstrating an upward trend across NACT, surgery, and adjuvant chemotherapy (Figure 8C).
To evaluate the prognostic value of miR-373, patients were categorized into two groups based on expression levels in untreated tumors (Figure 8D). Kaplan-Meier analysis demonstrated that high miR-373 expression was associated with significantly improved OS compared to low expression (P < 0.001). The median OS for the low-expression group was 68 months (95%CI: 33-not available), while the high-expression group did not reach median survival during the follow-up period. Cox proportional hazards regression confirmed this favorable association [hazard ratios = 2.10 × 10-11 (95%CI: 3.30 × 10-12 to 1.34 × 10-10] within this cohort.
Subgroup analysis by HR-status revealed more complex patterns (Supplementary Figure 9). Although individual Kaplan-Meier curves for these cohorts did not reach statistical significance (P = 0.999 for HR-negative and P = 0.749 for HR-positive), interaction testing in the Cox model demonstrated that HR status significantly modified the effect of miR-373 (Table 2). In HR-negative patients, high miR-373 expression appeared to correlate with extended survival (median OS not reached) compared to low expression (median OS of 26.5 months). Conversely, in HR-positive patients, the survival benefit of high miR-373 expression was less pronounced and diminished after 50 months.
| Variable | Univariate analysis | Multivariate analysis | ||
| P value | Hazard ratios (95%CI) | P value | Hazard ratios (95%CI) | |
| Age | 0.370 | 0.9707 (0.9096-1.036) | 0.01094 | 0.918 (0.859-0.980) |
| Ki-67 index | 0.381 | 1.015 (0.9818-1.049) | 0.00270 | 1.068 (1.023-1.116) |
| Subtype (HR+ vs HR-) | 0.712 | 0.7369 (0.1456-3.729) | 0.01618 | 0.0759 (0.0093-0.621) |
| miRNA-34a (high vs low) | 0.218 | 0.4061 (0.0968-1.704) | 1.23e-06 | 0.0165 (0.0032-0.0868) |
| miRNA-124a (high vs low) | 0.172 | 0.3272 (0.0659-1.626) | 0.09923 | 0.216 (0.035-1.336) |
| miRNA-137 (high vs low) | 0.344 | 0.4595 (0.0918-2.300) | 0.95869 | 0.956 (0.177-5.163) |
| miRNA-155 (high vs low) | 0.367 | 0.5174 (0.1234-2.169) | 0.96029 | 1.041 (0.215-5.048) |
| miRNA-373 (high vs low) | 0.209 | 0.3975 (0.0944-1.675) | < 2 × 10-16 | 2.10e-11 (3.31e-12 to 1.34e-10) |
| HR+: MiRNA-373 (high vs low) | - | - | < 2 × 10-16 | 6.42e+10 (1.01e+10 to 4.09e+11) |
It is important to note that the options available for HR-negative subgroup analysis (n = 11) are limited by low statistical power. The absence of events in the high-expression arm resulted in infinite CIs and an uninformative P-value (P = 0.999), rendering the numerical hazard ratio unreliable. However, the clear visual divergence in the survival curves suggests a biological trend that warrants investigation in a larger cohort.
These findings suggest that the prognostic impact of miR-373 is subtype-specific, as evidenced by the interaction terms, indicating that HR status modifies the effect of miR-373 expression. Such variability may stem from the paradoxical role of miR-373, which can function as either an oncogene or a tumor suppressor depending on the molecular landscape. These results suggest that miR-373 is not a universal biomarker; rather, its clinical significance is context-dependent and contingent upon HR-status.
Interestingly, the survival advantage associated with baseline miR-373 was no longer observed following NACT, suggesting that treatment intervention alter the miRNA’s baseline prognostic value (Figure 8E). These findings suggest that the prognostic impact of miR-373 may be treatment-dependent and modified by exposure to NACT.
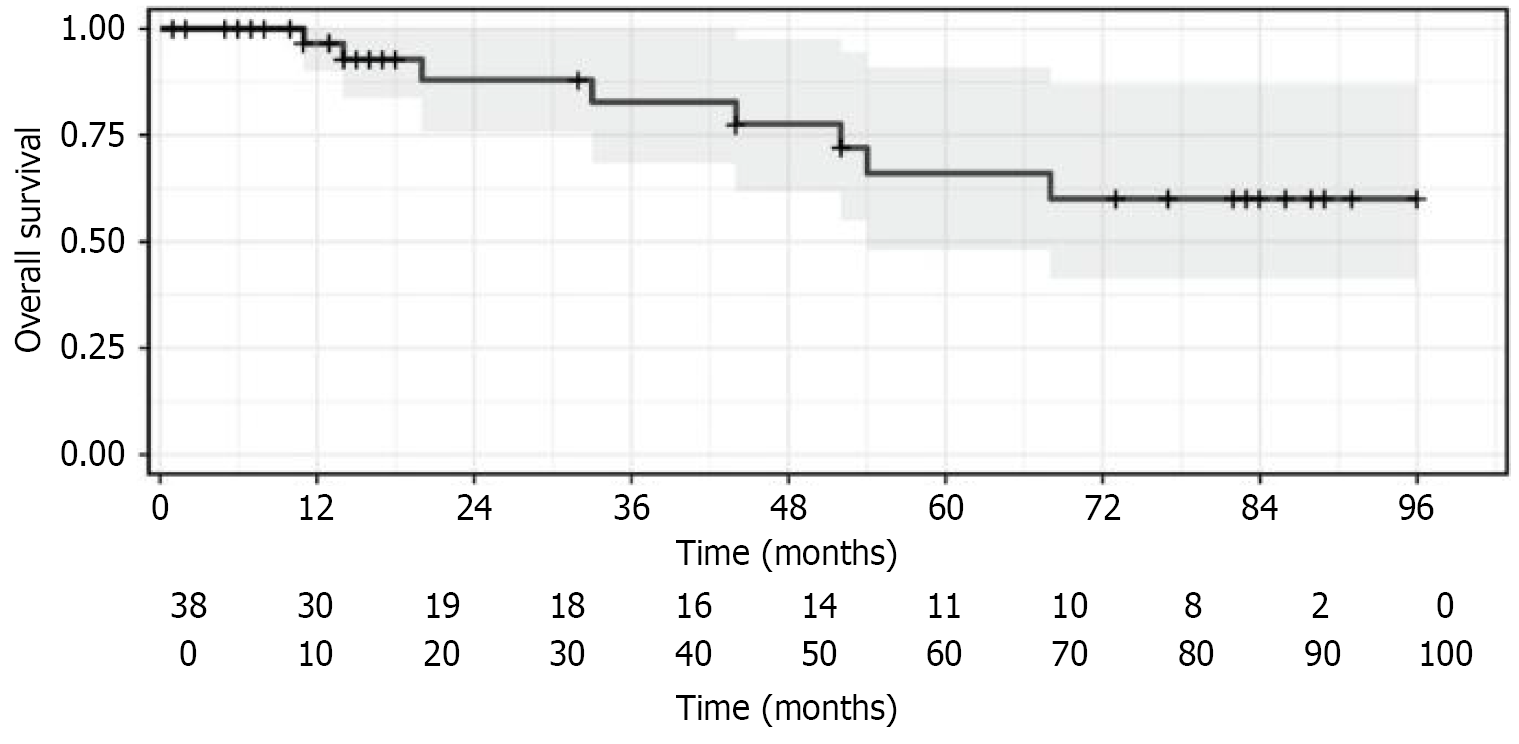
Survival analysis was performed in 38 patients with primary tumors using the Kaplan-Meier method, which accounts for censored data. The survival curve (Figure 9) displays a shaded grey area indicating the 95%CIs for the estimated survival probabilities. A summary table (Table 3) provides the survival proportions with corresponding 95%CIs at yearly intervals.
| Time (months) | Survival proportion | 95% confidences interval |
| 0 | 1.000 | 1.000-1.000 |
| 12 | 0.966 | 0.901-1.000 |
| 24 | 0.880 | 0.759-1.000 |
| 36 | 0.828 | 0.685-1.000 |
| 48 | 0.776 | 0.618-0.975 |
| 60 | 0.661 | 0.480-0.909 |
| 72 | 0.601 | 0.415-0.870 |
| 84 | 0.601 | 0.415-0.870 |
| 96 | 0.601 | 0.415-0.870 |
OS remained above 50% throughout the study period, with a survival probability of 0.601 at the end of follow-up. As the survival curve did not drop below 50%, the median survival time was not reached. The median follow-up duration was 44 months (95%CI: 16 months to 83 months).
Using the Weibull AFT model, the median age in our cohort was 54 years (Supplementary Figure 10). Survival curves were stratified by age groups corresponding to the 25th (47 years), 50th (54 years), and 75th (61 years) percentiles. Survival probability declined over time across all age groups, although the patterns were not uniform. Notably, patients in the younger age group (48 years) demonstrated a steeper decline in survival, suggesting a higher cumulative hazard and consistent with epidemiological data[1]. In contrast, the older age group (61 years) appeared to have more favorable survival, aligning with Cox proportional hazards analysis, in which increased age was identified as a significant protective factor (P = 0.011, Table 2). However, according to the Weibull AFT model, these differences were not statistically significant (P = 0.322).
Cox regression analysis of baseline clinical and molecular variables in untreated primary BC: To identify clinicopathological and molecular factors associated with patient survival, we performed both univariate and multivariate Cox regression analyses (Table 2). In the univariate model, none of the individual parameters reached statistical significance. Clinical factors, including age (P = 0.370), Ki-67 index (P = 0.381), and HR status (P = 0.712), were not significantly associated with survival. Similarly, the expression levels of the five evaluated miRNAs (miRNA-34a, -124a, -137, -155, and -373) did not demonstrate independent prognostic value when analyzed in isolation.
In contrast, the multivariate model, adjusting for all variables, revealed several significant independent predictors. Among clinical factors, increased age (hazard ratios = 0.918, 95%CI: 0.859-0.980; P = 0.011) and HR-positive status (hazard ratios = 0.0759, 95%CI: 0.0093-0.621; P = 0.016) emerged as significant protective factors. Conversely, a higher Ki-67 index was associated with a modest but statistically significant increase in hazard (hazard ratios = 1.068, P = 0.003), indicating an elevated mortality risk. This finding is consistent with the established role of Ki-67 as a marker of tumor proliferation and aggressive biological behavior. The apparent protective effect of age likely reflects that older patients in this cohort more frequently had less aggressive, HR-positive tumors, whereas younger patients more often presented with aggressive HR-negative subtypes.
Regarding miRNA expression, high miR-34a levels emerged as a strong independent predictor of favorable outcome in the overall cohort (hazard ratios = 0.0165, P < 0.001). High miR-373 expression also showed a highly significant association with survival (P < 0.001), however, the overall effect estimate was extremely small (hazard ratios = 2.10 ×
A key finding of this study is the marked shift in prognostic significance when moving from univariate to multivariate analysis. Factors such as age, Ki-67 index, miR-34a and miR-373 failed to reach significance in isolation but emerged as powerful independent predictors upon multivariate adjustment. This phenomenon underscores the presence of complex confounding relationships in the dataset, suggesting that the prognostic value of these miRNAs may be masked by clinical variables, such as HR status, until statistically controlled for in a combined model.
Prognostic value of post-NACT clinicopathological factors and miRNA expression profiles: To evaluate the prognostic significance of clinicopathological factors and miRNA expression levels post-NACT, both univariate and multivariate Cox regression analyses were performed (Table 4). In the univariate analysis, none of the studied variables reached statistical significance as predictors of survival. Specifically, age (P = 0.530), Miller-Payne score (P = 0.792), and molecular subtype (P = 0.897) did not show a correlation with hazard rates. Similarly, the expression levels of all five miRNAs (miRNA-34a, -124a, -137, -155, and -373) were not individually associated with survival outcomes (all P > 0.05).
| Variable | Univariate analysis | Multivariate analysis | ||
| P value | Hazard ratios (95%CI) | P value | Hazard ratios (95%CI) | |
| Age | 0.530 | 0.980 (0.919-1.044) | 0.532 | 0.959 (0.842-1.093) |
| Miller-Payne score (MPG 2, 3 vs 4, 5) | 0.792 | 1.326 (0.162-10.840) | 0.752 | 3.101 (0.003-3457.333) |
| Subtype (HR+ vs HR-) | 0.897 | 1.113 (0.222-5.579) | 0.044 | 0.015 (0.000-0.892) |
| miRNA-34a (high vs low) | 0.827 | 1.169 (0.289-4.728) | 0.600 | 0.485 (0.032-7.266) |
| miRNA-124a (high vs low) | 0.642 | 1.394 (0.344-5.648) | 0.654 | 1.666 (0.179-15.473) |
| miRNA-137 (high vs low) | 0.739 | 0.790 (0.197-3.173) | 0.284 | 0.188 (0.009-3.999) |
| miRNA-155 (high vs low) | 0.974 | 1.023 (0.254-4.127) | 0.999 | 3.143e-10 (0.000-Inf) |
| miRNA-373 (high vs low) | 0.499 | 1.617 (0.401-6.515) | 1.000 | 1.998 (0.000-Inf) |
| HR+: MiRNA-155 (high vs low) | - | - | 0.999 | 7.324e+09 (0.000-Inf) |
| HR+: MiRNA-373 (high vs low) | - | - | 1.000 | 9.290 (0.000-Inf) |
A multivariate Cox model was constructed to adjust for potential confounders. After adjustment, tumor subtype emerged as a significant independent prognostic factor; patients with HR-positive tumors exhibited a significantly lower hazard of an event compared to those with HR-negative tumors (hazard ratios = 0.015; 95%CI: 0.000-0.892; P = 0.044).
Regarding the miRNA profiles in the multivariate model, miRNA-137 showed a trend toward a reduced hazard rate (hazard ratios = 0.188; P = 0.284). While this did not reach statistical significance for survival, the direction of this effect is consistent with our earlier finding that high miR-137 expression is associated with an improved pathological response to NACT (Figure 3), further suggesting a potential protective role.
Conversely, the analyses for miRNA-155 and miRNA-373, particularly within the HR-positive subgroup, yielded unstable estimates with extremely wide CIs and non-significant P values. This likely indicates a lack of sufficient events and small sample sizes within these specific strata, preventing a reliable hazard estimate. These wide CIs, also observed for the Miller-Payne score, represent a study limitation and suggest that the model’s predictive power is constrained by the limited number of events in the post-NACT cohort.
This study uniquely integrates paired profiling of five miRNAs (miRNA-34a, -124a, -137, -155, and -373), representing both tumor-suppressive and oncogenic classes, across matched BC tissue and plasma collected before and after NACT. By combining: (1) Matched tissue-plasma sampling; (2) Pre/post-NACT longitudinal analysis; (3) HR-stratified evaluation; and (4) Survival follow-up, this work addresses a critical gap in translational evidence. Unlike previous studies limited to single-timepoint or single-compartment analyses, our dual-sample design enables direct comparison of intra-tumoral and systemic miRNA regulation within the same individuals. This approach, supported by rigorous sample handling of fresh tissue and immediate plasma processing, identifies candidate miRNAs for future validation as early predictors of chemotherapy sensitivity and long-term outcomes in BC.
In our study, untreated breast tumors exhibited distinct expression patterns aligned with established biology and HR status. The tumor suppressor miR-124a was significantly downregulated in primary tumors. Conversely, another suppressor miR-34a was elevated at baseline, predominantly in HR-positive cases. Given that miR-34a is a direct transcriptional target of p53, its elevation in these tumors likely reflects p-53-mediated activation in a wild-type p53 context.
Regarding oncogenic miRNAs, miR-155 was highly expressed at baseline, particularly in HR-negative tumors, consistent with its role in aggressive tumor phenotypes. Meanwhile, miR-137 and miR-373 exhibited more complex, context-dependent behavior influenced by HR status and potential epigenetic regulation.
NACT induced coordinated, miRNA-specific shifts across the cohort, highlighting the dynamic regulation of miRNAs under therapeutic pressure. During treatment, miR-34a, miR-124a, and miR-155 shifted towards levels observed in non-tumor tissue. This “normalization” suggests an attenuation of stress-response pathways and a treatment-driven reactivation of tumor-suppressive programs.
However, responses were not uniform. miR-137 showed divergent, HR-dependent shifts, normalizing in HR-positive tumor but remaining elevated in HR-negative cases. miR-373 continued to decline post-NACT, likely reflecting the chemotherapy-induced deactivation of invasive programs. Notably, miR-137 emerged as the sole miRNA significantly associated with pathological response, identifying it as a candidate dynamic biomarker for chemosensitivity.
A key finding was that miRNA expression in tumor tissue did not always mirror changes in the plasma. These discrepancies underscore compartment-specific dynamics and likely reflect the underlying heterogeneity of the systemic disease vs the primary tumor.
Survival analyses revealed that baseline miRNA expression carried stronger prognostic value than post-treatment levels. Among the studied cohort, miR-34a and miR-373 retained independent prognostic significance at baseline, with higher expression levels associated with improved OS. Interestingly, this prognostic value weakened or reversed after NACT, reflecting treatment-induced remodeling that diminishes the biological distinctions present at diagnosis and contributes to the convergence of post-treatment miRNA landscapes. Overall, these findings demonstrate that miRNAs in BC are highly dynamic, treatment-responsive, and shaped by tumor subtype, with clear distinctions between untreated and post-NACT states. Below, we describe each miRNA in detail.
miR-34a: In our study, high expression of tumor-suppressor miR-34a in primary tumors was significantly associated with improved OS, supporting its role as a tumor-suppressive miRNA involved in promoting apoptosis, inducing cell cycle arrest, and reducing proliferation[35,36]. This observation is consistent with previous reports linking miR-34a upregulation to increased apoptosis through targeting BCL2 and to enhanced sensitivity to chemotherapy and radiation[26,37-43].
Our results demonstrated significant upregulation of miR-34a in primary breast tumors, predominantly driven by HR-positive cases. Given that miR-34a is a direct transcriptional target of p53, its elevated expression in untreated tumors reflects activation of a p53-mediated cellular stress response[44,45]. Previous studies have shown that of miR-34a overexpression suppresses metastasis by inhibiting facilitates epithelial-mesenchymal transition (EMT) relating factors, such as Notch1, TWIST1, ZEB1 and ZEB2, suggesting that miR-34a upregulation may also occur in association with stress-adaptive or dedifferentiation programs[46,47].
Following NACT, we observed a reduction in tumor miR-34a expression, indicating possible attenuation of the stress response. This decrease may reflect loss of p53 activity, epigenetic silencing of MIR34A, or the selection of resistant subclones with impaired p53-miR-34a signaling[48-52]. Alternatively, chemotherapy may selectively eliminate miR-34a-high tumor cells or promote normalization of cellular regulatory networks. Thus, post-NACT decrease in miR-34a expression likely represents treatment-induced remodeling of tumor cell populations rather than a simple restoration of tumor suppression.
In contrast, circulating miR-34a showed a trend toward increased levels during therapy, although this association was less statistically robust due to the limited sample size and number of events. Similar findings have been reported by others, with treatment-induced upregulation of circulating miR-34a during NACT, particularly in patients with luminal B subtype BC[38]. Conversely, significantly lower expression of miR-34a/34b/34c has been observed in TNBC patients[53], underscoring the heterogeneity of its regulation across subtypes.
The prognostic significance of miR-34a also appeared to differ by HR status. As previously reported, high miR-34a expression has been associated with reduced proliferative activity, lower-grade tumors[34] and overall, less aggressive disease. Consistent with these findings, we observed that high miR-34a expression correlated with improved survival (P < 0.001), an association that was primarily evident in HR-positive patients. Importantly, after NACT, survival curves for high vs low miR-34a expression overlapped, and no significant difference was observed (P = 0.6), suggesting that the prognostic utility of miR-34a diminishes following chemotherapy. Interestingly, we noted a trend toward better survival in patients with lower post-NACT miR-34a expression.
These dynamic changes in miR-34a expression may reflect a treatment-induced reduction driven by tumor adaptation, as supported by diverging patterns of expression observed across tumor differentiation states after NACT (Figure 1). Unlike the classical tumor-suppressor paradigm, our findings suggest that high miR-34a expression in untreated tumors and its subsequent reduction after therapy are more consistent with treatment-induced remodeling of tumor cell populations rather than a straightforward restoration of tumor suppression.
miR-124a: miR-124 is a tumor suppressor frequently silenced through DNA methylation[54-57] and downregulated in multiple cancers, including breast, lung, melanoma, and colon cancer[58-64]. It suppresses oncogenesis by targeting oncogenic transcripts and regulating proliferation, migration, and differentiation, while its loss facilitates EMT, metastasis, and therapy resistance[54,56,59]. In BC, it inhibits proliferation by inducing cell cycle arrest through CD151 suppression[61] and reduced expression has been associated with poor prognosis[57,65,66].
In our cohort, miR-124a was significantly downregulated in primary tumors compared with adjacent normal tissue. After NACT, expression increased across the cohort, driven mainly by HR-positive tumors, restoring levels comparable to normal tissue. This therapy-induced upregulation suggests an anti-tumor effect and aligns with reports of higher miR-124a expression in HR-positive compared with HR-negative BCs[66]. Elevated expression of miR-124a during treatment may therefore help limit metastatic potential, consistent with its proposed role as a metastasis-suppressor miRNA[63].
miR-124a has also been linked to chemosensitivity in other cancers. In colorectal cancer, overexpression of miR-124a enhanced oxaliplatin response by targeting CAPN2, while in glioblastoma, it improved temozolomide efficacy through aurora kinase A suppression[67,68]. These findings support a broader role for miR-124a in modulating chemotherapy response.
By contrast, plasma samples of miR-124a showed only a non-significant trend toward downregulation after treatment, likely reflecting differences in biological origin and secretion dynamics rather than tumor-specific expression. Persistent downregulation of miR-124a may suggest the selection of resistant clones capable of evading apoptosis, as supported by our Sankey diagram (Figure 1), which shows redistribution toward higher-grade tumors after NACT. Since miR-124a targets oncogenes such as STAT3, EZH2, CDK6, CD151, and AURKA, reduced expression could facilitate reactivation of proliferative and migratory pathways, potentially serving as a negative prognostic marker linked to poor therapeutic response or early recurrence.
In survival analysis, patients with low miR-124a expression in primary tissue showed a non-significant trend toward worse OS, consistent with prior studies[65,67,69]. Intriguingly, after NACT, higher tumor expression of miR-124a was paradoxically associated with a worse survival (hazard ratios > 1 in multivariate Cox analysis), suggesting that treatment-driven modulation may alter its prognostic significance.
miR-137: miR-137 displays a context-dependent role across cancers, acting predominantly as a tumor suppressor in TNBC[70,71], colorectal cancer[72], gastric cancer[73-75], cholangiocarcinoma[76], melanoma[77,78], and ovarian cancer[79], although oncogenic activity has been reported in certain contexts[80,81]. In BC, its role is similarly complex, generally suppressing proliferation and migration[30,82], but also showing subtype-specific expression and potential involvement in hormone therapy resistance[83]. Mechanistically, miR-137 targets estrogen-related receptor α, linking it to estrogen-related pathways[84].
Our data reflect the dual behavior of miR-137. In primary tumors, miR-137 was elevated, with further upregulation during NACT in HR-negative cases, whereas HR-positive tumors exhibited post-NACT normalization to non-tumor levels. Persistent overexpression in HR-negative tumors may indicate a treatment-induced suppressive effect, consistent with evidence that miR-137 inhibits proliferation, invasion, migration[30,76], and EMT through Del-1 downregulation[71,85] or modulation of epigenetic regulators such as KDM5 family demethylases[86] and the histone methyltransferase EZH2[87]. Additionally, miR-137 has been reported to enhance chemosensitivity by inhibiting DUSP4[88] and by reducing P-glycoprotein levels through targeting Y-box binding, a known mediator of drug resistance and tumor progression in BC[89].
Conversely, post-NACT normalization in HR-positive tumors likely reflects therapy adaptation rather than increased aggressiveness, with high baseline expression possibly linked to tumor development. Sankey diagram (Figure 2) further supports a strong association between miR-137 and HR-positive tumors, consistent with estrogen receptor-related regulation.
Clinically, reduced miR-137 expression is generally associated with poor survival across cancers[79,90-93]. In our cohort, high miR-137 trended toward improved survival, particularly in HR-positive tumors, though this did not reach significance. After NACT, this effect was attenuated, possibly reflecting systemic therapy-induced modulation or reduced intergroup differences. Collectively, these findings position miR-137 as a dynamic, treatment-responsive biomarker. Its expression appears shaped by both tumor subtype and therapy, suggesting that miR-137 may be more informative for monitoring treatment response than as a static baseline prognostic marker.
miR-155: miR-155 is a well-established oncogenic miRNA (oncomiR) frequently upregulated in solid tumors, including BC[19,24,29]. Elevated expression has been linked to aggressive phenotypes[24,94-96], yet other studies report associations with increased immune infiltration and improved prognosis, underscoring its context-dependent role[97,98]. Functionally, miR-155 promotes angiogenesis by enhancing endothelial and macrophage migration, partly through targeting the von Hippel-Lindau tumor suppressor[94,98].
Elevated miR-155 is also characteristic of basal-like and BRCA1-mutated BC, where loss of BRCA1 impairs histone deacetylase-mediated transcriptional repression and derepresses miR-155[24,95,99]. This places miR-155 within the context of DNA repair deficiency and homologous recombination-related tumors[100]. Evidence from human tumors supports a strong correlation between BRCA1 loss and miR-155 upregulation, with emerging data suggesting that miR-155 may influence sensitivity to poly (ADP-ribose) polymerase inhibitors[15,95].
In our cohort, tumor miR-155 expression was consistent with its oncogenic role. Elevated miR-155 levels were observed in untreated tumor tissues, followed by a decreasing trend after NACT. Although no significant differences were detected between HR-positive and HR-negative tumors, a tendency toward higher expression in HR-negative cases supports previous findings linking miR-155 to more aggressive disease[24]. This observation was further supported by our Sankey diagram analysis (Figures 1 and 2), which demonstrated enrichment of miR-155 in HR-negative and grade 3 tumors, reinforcing its association with aggressive clinicopathological features.
Regarding circulating levels, plasma miR-155 increased during NACT and, although remaining elevated, showed a gradual decline after surgery and during adjuvant chemotherapy - consistent with other findings[101,102]. However, these post-treatment changes did not fully mirror tumor expression, likely reflecting additional contributions from immune and stromal compartments[28,96-98]. While some studies have reported decreased circulating miR-155 in BC patients[28,103,104], others found significantly higher serum levels compared with healthy controls[29,97,102]. Such discrepancies likely reflect heterogeneity in disease stage, tumor subtype, and treatment context[28,97,102,103].
Elevated miR-155 levels observed during chemotherapy may serve as a potential marker of better outcomes, as patients with high tumor miR-155 expression in our cohort showed a trend toward improved OS. This aligns with previous reports suggesting that low miR-155 levels predict early relapse and metastatic disease in BC patients[104-106].
Prognostically, miR-155 has been reported both as an indicator of poor outcome and, conversely, as a protective factor depending on biological context. Wang et al[97] reported that elevated miR-155 expression was linked to improved survival outcomes, while Gasparini et al[100] demonstrated that, in triple-negative BC, increased miR-155 levels were associated with reduced RAD51 expression and better OS. In our study, survival analysis revealed a non-significant trend toward improved OS in patients with high baseline tumor miR-155 expression. When stratified by HR status, HR-positive patients with high miR-155 expression tended to have better outcomes, whereas HR-negative patients exhibited the opposite pattern, with higher expression associated with shorter survival. Although limited by sample size, our findings are broadly consistent with prior observations and support a potential prognostic role for miR-155 in BC that warrants further investigation in larger, well-characterized cohorts.
miR-373: miR-373 was initially identified as a potential oncogene in BC[107], but subsequent studies have revealed context-dependent and sometimes contradictory functions[108-110]. In BC, it has been reported to act either as a tumor promoter or suppressor depending on the cellular and molecular context. On one hand, miR-373 promotes EMT and metastasis via the Wnt/β-catenin pathway or the TXNIP-HIF1α-TWIST axis[107,109,111-113]. On the other hand, particularly in ER-negative BC, the miR-520/373 family functions as a tumor suppressor by inhibiting both nuclear factor-kappa B (NF-κB) and transforming growth factor-beta (TGF-β) signaling pathways through the direct targeting of RELA and TGFBR2, respectively. This dual inhibition prevents the induction of EMT, ultimately suppressing tumor cell intravasation and progression[114]. These opposing roles suggest that miR-373 function is shaped by tumor subtype, dominant signaling networks, and the surrounding microenvironment.
Across malignancies, miR-373 is overexpressed in testicular germ cell tumors[115], BC[112,116,117], gastric cancer[118], and colorectal cancer[119]. Conversely, it is downregulated in ER-negative BC and in cholangiocarcinoma, colon, pancreatic, lung, and ovarian cancers[108,114,120-122].
In our study, miR-373 remained downregulated after NACT in both HR-positive and HR-negative tumors compared with adjacent non-tumor tissue. This pattern may reflect chemotherapy-induced epigenetic repression of the miR-520/373 family[122,123]. Such shifts have been associated with reduced proliferation and enhanced chemosensitivity[124-126]. Although direct BC data are limited, glioblastoma studies similarly link lower miR-373 with improved chemosensitivity, suggesting possible selection against miR-373-high, chemoresistant clones[124,127]. Consistent with these findings, analysis of the Sankey diagram (Figure 1) demonstrated a redistribution toward lower histologic grades following NACT. Additionally, miR-373 downregulation was observed in both good and partial responders (Figure 3), suggesting a reduction of more aggressive tumor cell populations.
Several mechanisms could explain this reduction. NACT may selectively eliminate high-miR-373 subclones, favor resistant clones with intrinsically low levels[27,128-130], or decrease stromal and immune fractions that contribute to bulk tissue miRNA[9,114,131]. Mechanistically, the links of miR-373 with EMT and invasiveness allow two non-exclusive interpretations: (1) Chemotherapy-induced “deactivation” or clonal pruning of invasive programs; and (2) Epigenetic silencing of tumor-suppressive circuits (miR-373 can inhibit pro-metastatic TGF-β and NF-κB signaling). Thus, the observed decline likely integrates tumor-intrinsic and microenvironmental changes rather than reflecting a simple loss of tumor suppression.
By contrast, plasma miR-373 increased after NACT, consistent with compartment-specific dynamics. Tumor cell death can passively release intracellular miRNAs, while active exosomal export may enrich circulating miR-373 even as tumor levels decline. Elevated serum miR-373 has been reported with nodal involvement and HER2/neu overexpression[18,132,133], and a pilot study also observed post-chemotherapy rises[134]. Functionally, increased circulating miR-373 could restore tumor-suppressive activity, particularly in HR-negative disease, by inhibiting TGF-β and NF-κB pathways[114,135]. Supporting this, low serum miR-373 has been linked to poor prognosis in pancreatic cancer[121,136].
Taken together, the paradoxical pattern of miR-373, downregulated in tumor tissue but elevated in plasma, likely reflects a multifactorial process involving epigenetic repression, clonal selection, and compositional changes in tissue, alongside passive release and active secretion into circulation. Clinically, increases in plasma levels of miR-373 may serve as a noninvasive indicator of tumor turnover and treatment response, whereas decreases in tissue levels suggest chemotherapy-mediated modulation of oncogenic programs[18,133,137,138]. The prognostic role of miR-373 is similarly context-dependent. In glioma, low expression predicted worse survival[139], while in oral carcinoma, high expression was associated with poorer outcomes[140]. In pancreatic cancer, low serum miR-373 predicted shorter survival and functioned as an independent prognostic marker[136].
In BC, large miRNA expression panels have been associated with poor survival outcomes[138]; however specific Kaplan-Meier survival data for miR-373 remain limited. In our cohort, high baseline tumor miR-373 expression was associated with improved OS, supporting a potential tumor-suppressive role in untreated disease. Following NACT, survival curves converged, and notably, patients with low post-treatment miR-373 expression demonstrated better OS. This shift is consistent with treatment-associated downregulation of miR-373, suggesting that its prognostic impact is dynamically modulated by therapeutic exposure. Collectively, these findings support a dual and context-dependent role for miR-373, which may function either as a tumor suppressor or as an oncogenic driver depending on tumor subtype, cellular context, and treatment status.
HR status is a well-established predictor of BC outcome. HR-positive tumors are generally associated with less aggressive biology, slower proliferation, and improved prognosis[2]. Consistent with this, our analysis demonstrated that HR-positive patients had a lower risk of death and more favorable OS (hazard ratios = 0.0759, P = 0.016), particularly among older patients (median age 61 years) (hazard ratios = 0.918, P = 0.011). Using the Weibull AFT model, we observed that younger patients (median age 48 years) experienced reduced survival, in line with global epidemiological data showing that younger BC populations often present with more aggressive disease and poorer outcomes[1].
The Ki-67 proliferation index is another important prognostic and predictive marker, reflecting tumor growth activity. Although we did not observe statistically significant changes in Ki-67 expression following NACT, baseline analysis of primary tumors confirmed that higher Ki-67 levels were associated with increased mortality risk (hazard ratios = 1.068, P = 0.003). This emphasizes the value of Ki-67 in identifying aggressive tumors and informing risk stratification.
Collectively, age, HR status, and Ki-67 provide complementary insights into patient prognosis and treatment response. Integrating these clinical parameters with molecular biomarkers, such as miRNAs, may improve predictive accuracy and support personalized therapeutic decision-making in BC management.
Despite the strong biological signals observed, several limitations warrant consideration. The modest sample size and low number of survival events limited statistical power in certain analyses, contributing to extreme point estimates and wide CIs in the Cox regression models. In particular, the substantial overlap of survival curves following NACT impaired the model’s ability to reliably estimate relative hazard in the post-treatment setting.
Additionally, the focused panel of five miRNAs does not encompass the full complexity of the BC miRNome. Validation in larger, multicenter cohorts is necessary to confirm these findings and to determine whether the identified miRNAs function as causal drivers of therapeutic response or as surrogate markers of treatment-induced remodeling. Such studies will be essential to establish the role of miRNA-guided strategies within precision oncology frameworks.
In summary, this study demonstrates that miRNAs represent highly dynamic biomarkers whose clinical relevance in BC is shaped by both HR status and therapeutic pressure. Our longitudinal, dual-compartment approach indicates that baseline molecular profiling provides greater prognostic value than post-treatment assessment.
At baseline, miR-34a and miR-373 emerged as independent prognostic markers. High initial expression, particularly within HR-positive disease, was associated with significantly improved OS (P < 0.001). With respect to treatment response, miR-137 was the only miRNA significantly associated with pathological response, highlighting its potential as a candidate biomarker for real-time monitoring of chemosensitivity.
The treatment-induced remodeling observed after NACT, characterized by convergence of miRNA expression profiles, likely explains the attenuation of prognostic separation and the statistical instability observed in post-treatment survival models. Moreover, the lack of consistent correlation between tissue and plasma miRNA levels suggests that liquid biopsy alone may not fully recapitulate the intratumoral molecular landscape. These dynamic shifts likely reflect broader changes within the tumor microenvironment, including stromal and immune compartments, rather than alterations confined to malignant epithelial cells.
The authors would like to express their gratitude to Professor Joseph W Hogan, Associate Professor of Medicine Dr. Adam J Olszewski, and Associate Professor Vitaly G Gurianov for their insightful critiques and constructive feedback, which significantly enhanced the quality of this work.
| 1. | Siegel RL, Kratzer TB, Giaquinto AN, Sung H, Jemal A. Cancer statistics, 2025. CA Cancer J Clin. 2025;75:10-45. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1828] [Cited by in RCA: 2135] [Article Influence: 2135.0] [Reference Citation Analysis (8)] |
| 2. | Waks AG, Winer EP. Breast Cancer Treatment. JAMA. 2019;321:316. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 150] [Cited by in RCA: 128] [Article Influence: 18.3] [Reference Citation Analysis (0)] |
| 3. | Hussen BM, Hidayat HJ, Salihi A, Sabir DK, Taheri M, Ghafouri-Fard S. MicroRNA: A signature for cancer progression. Biomed Pharmacother. 2021;138:111528. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 339] [Cited by in RCA: 270] [Article Influence: 54.0] [Reference Citation Analysis (0)] |
| 4. | Seyhan AA. Trials and Tribulations of MicroRNA Therapeutics. Int J Mol Sci. 2024;25:1469. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 303] [Cited by in RCA: 230] [Article Influence: 115.0] [Reference Citation Analysis (0)] |
| 5. | Cai Y, Yu X, Hu S, Yu J. A brief review on the mechanisms of miRNA regulation. Genomics Proteomics Bioinformatics. 2009;7:147-154. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 490] [Cited by in RCA: 728] [Article Influence: 45.5] [Reference Citation Analysis (0)] |
| 6. | Khraiwesh B, Arif MA, Seumel GI, Ossowski S, Weigel D, Reski R, Frank W. Transcriptional control of gene expression by microRNAs. Cell. 2010;140:111-122. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 338] [Cited by in RCA: 325] [Article Influence: 20.3] [Reference Citation Analysis (1)] |
| 7. | O'Brien J, Hayder H, Zayed Y, Peng C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front Endocrinol (Lausanne). 2018;9:402. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4388] [Cited by in RCA: 3813] [Article Influence: 476.6] [Reference Citation Analysis (5)] |
| 8. | Nazki FH, Bracken CP. Regulation and dysregulation of microRNA - transcription factor axes in differentiation and neuroblastoma. Cell Mol Life Sci. 2025;82:304. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 9. | Suzuki HI, Katsura A, Matsuyama H, Miyazono K. MicroRNA regulons in tumor microenvironment. Oncogene. 2015;34:3085-3094. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 153] [Cited by in RCA: 169] [Article Influence: 15.4] [Reference Citation Analysis (0)] |
| 10. | Ali Syeda Z, Langden SSS, Munkhzul C, Lee M, Song SJ. Regulatory Mechanism of MicroRNA Expression in Cancer. Int J Mol Sci. 2020;21:1723. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 836] [Cited by in RCA: 737] [Article Influence: 122.8] [Reference Citation Analysis (5)] |
| 11. | Fridrichova I, Zmetakova I. MicroRNAs Contribute to Breast Cancer Invasiveness. Cells. 2019;8:1361. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 61] [Cited by in RCA: 124] [Article Influence: 17.7] [Reference Citation Analysis (0)] |
| 12. | Wong JS, Cheah YK. Potential miRNAs for miRNA-Based Therapeutics in Breast Cancer. Noncoding RNA. 2020;6:29. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 19] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 13. | Huang SK, Luo Q, Peng H, Li J, Zhao M, Wang J, Gu YY, Li Y, Yuan P, Zhao GH, Huang CZ. A Panel of Serum Noncoding RNAs for the Diagnosis and Monitoring of Response to Therapy in Patients with Breast Cancer. Med Sci Monit. 2018;24:2476-2488. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 26] [Cited by in RCA: 39] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 14. | Seyhan AA. Circulating microRNAs as Potential Biomarkers in Pancreatic Cancer-Advances and Challenges. Int J Mol Sci. 2023;24:13340. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 42] [Reference Citation Analysis (8)] |
| 15. | Alavanda C, Dirimtekin E, Mortoglou M, Arslan Ates E, Guney AI, Uysal-Onganer P. BRCA Mutations and MicroRNA Expression Patterns in the Peripheral Blood of Breast Cancer Patients. ACS Omega. 2024;9:17217-17228. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 3] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 16. | de Miranda FS, Slaibi-Filho J, Calasans Dos Santos G, Carmo NT, Kaneto CM, Borin TF, Luiz WB, Gastalho Campos LC. MicroRNA as a promising molecular biomarker in the diagnosis of breast cancer. Front Mol Biosci. 2024;11:1337706. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 10] [Reference Citation Analysis (0)] |
| 17. | Wang F, Zheng Z, Guo J, Ding X. Correlation and quantitation of microRNA aberrant expression in tissues and sera from patients with breast tumor. Gynecol Oncol. 2010;119:586-593. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 261] [Cited by in RCA: 233] [Article Influence: 14.6] [Reference Citation Analysis (0)] |
| 18. | Swellam M, El Magdoub HM, Hassan NM, Hefny MM, Sobeih ME. Potential diagnostic role of circulating MiRNAs in breast cancer: Implications on clinicopathological characters. Clin Biochem. 2018;56:47-54. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 30] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 19. | Grimaldi AM, Nuzzo S, Condorelli G, Salvatore M, Incoronato M. Prognostic and Clinicopathological Significance of MiR-155 in Breast Cancer: A Systematic Review. Int J Mol Sci. 2020;21:5834. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 37] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 20. | Li J, He D, Bi Y, Liu S. The Emerging Roles of Exosomal miRNAs in Breast Cancer Progression and Potential Clinical Applications. Breast Cancer (Dove Med Press). 2023;15:825-840. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 14] [Reference Citation Analysis (0)] |
| 21. | Zhang K, Wang YW, Wang YY, Song Y, Zhu J, Si PC, Ma R. Identification of microRNA biomarkers in the blood of breast cancer patients based on microRNA profiling. Gene. 2017;619:10-20. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 63] [Cited by in RCA: 76] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 22. | Linares-Rodríguez M, Blancas I, Rodríguez-Serrano F. The Predictive Value of Blood-Derived Exosomal miRNAs as Biomarkers in Breast Cancer: A Systematic Review. Clin Breast Cancer. 2025;25:e48-e55.e15. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 9] [Reference Citation Analysis (0)] |
| 23. | Blenkiron C, Goldstein LD, Thorne NP, Spiteri I, Chin SF, Dunning MJ, Barbosa-Morais NL, Teschendorff AE, Green AR, Ellis IO, Tavaré S, Caldas C, Miska EA. MicroRNA expression profiling of human breast cancer identifies new markers of tumor subtype. Genome Biol. 2007;8:R214. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 706] [Cited by in RCA: 733] [Article Influence: 40.7] [Reference Citation Analysis (5)] |
| 24. | Mattiske S, Suetani RJ, Neilsen PM, Callen DF. The oncogenic role of miR-155 in breast cancer. Cancer Epidemiol Biomarkers Prev. 2012;21:1236-1243. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 251] [Cited by in RCA: 228] [Article Influence: 16.3] [Reference Citation Analysis (0)] |
| 25. | Al-Khanbashi M, Caramuta S, Alajmi AM, Al-Haddabi I, Al-Riyami M, Lui WO, Al-Moundhri MS. Tissue and Serum miRNA Profile in Locally Advanced Breast Cancer (LABC) in Response to Neo-Adjuvant Chemotherapy (NAC) Treatment. PLoS One. 2016;11:e0152032. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 40] [Cited by in RCA: 49] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 26. | Li ZH, Weng X, Xiong QY, Tu JH, Xiao A, Qiu W, Gong Y, Hu EW, Huang S, Cao YL. miR-34a expression in human breast cancer is associated with drug resistance. Oncotarget. 2017;8:106270-106282. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 42] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 27. | Hu W, Tan C, He Y, Zhang G, Xu Y, Tang J. Functional miRNAs in breast cancer drug resistance. Onco Targets Ther. 2018;11:1529-1541. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 61] [Cited by in RCA: 95] [Article Influence: 11.9] [Reference Citation Analysis (0)] |
| 28. | Fortis SP, Vaxevanis CK, Mahaira LG, Sofopoulos M, Sotiriadou NN, Dinou A, Arnogiannaki N, Stavropoulos-Giokas C, Thanos D, Baxevanis CN, Perez SA. Serum miRNA-based distinct clusters define three groups of breast cancer patients with different clinicopathological and immune characteristics. Cancer Immunol Immunother. 2019;68:57-70. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 11] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 29. | Canatan D, Sönmez Y, Yılmaz Ö, Çim A, Coşkun HŞ, Sezgin Göksu S, Ucar S, Aktekin MR. MicroRNAs as biomarkers for breast cancer. Acta Biomed. 2021;92:e2021028. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 9] [Reference Citation Analysis (0)] |
| 30. | Wu D, Thompson LU, Comelli EM. MicroRNAs: A Link between Mammary Gland Development and Breast Cancer. Int J Mol Sci. 2022;23:15978. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 11] [Reference Citation Analysis (0)] |
| 31. | Garrido-Cano I, Pattanayak B, Adam-Artigues A, Lameirinhas A, Torres-Ruiz S, Tormo E, Cervera R, Eroles P. MicroRNAs as a clue to overcome breast cancer treatment resistance. Cancer Metastasis Rev. 2022;41:77-105. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 26] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 32. | Agarwal V, Bell GW, Nam JW, Bartel DP. Predicting effective microRNA target sites in mammalian mRNAs. Elife. 2015;4:e05005. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6012] [Cited by in RCA: 5671] [Article Influence: 515.5] [Reference Citation Analysis (4)] |
| 33. | Ryspayeva D, Lyashenko A, Dosenko I, Kostryba O, Koshyk O, Krotevych M, Smolanka I. Predictive factors of pathological response to neoadjuvant chemotherapy in patients with breast cancer. J BUON. 2020;25:168-175. [PubMed] |
| 34. | Tokumaru Y, Katsuta E, Oshi M, Sporn JC, Yan L, Le L, Matsuhashi N, Futamura M, Akao Y, Yoshida K, Takabe K. High Expression of miR-34a Associated with Less Aggressive Cancer Biology but Not with Survival in Breast Cancer. Int J Mol Sci. 2020;21:3045. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 42] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 35. | Fu J, Imani S, Wu MY, Wu RC. MicroRNA-34 Family in Cancers: Role, Mechanism, and Therapeutic Potential. Cancers (Basel). 2023;15:4723. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 65] [Reference Citation Analysis (0)] |
| 36. | Adams BD, Parsons C, Slack FJ. The tumor-suppressive and potential therapeutic functions of miR-34a in epithelial carcinomas. Expert Opin Ther Targets. 2016;20:737-753. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 71] [Cited by in RCA: 95] [Article Influence: 8.6] [Reference Citation Analysis (0)] |
| 37. | Kouhestani SD, Khalili S, Razi A, Aghili M, Moghadam MF. Ectopic expression of miR-34a/-328 sensitizes breast cancer stem cells to gamma rays/doxorubicin by BCL2/ABCG2 targeting. Mol Biol Rep. 2025;52:490. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 3] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 38. | Zhang Z, Zhang H, Li C, Xiang Q, Xu L, Liu Q, Pang X, Zhang W, Zhang H, Zhang S, Duan X, Liu Y, Cui Y. Circulating microRNAs as indicators in the prediction of neoadjuvant chemotherapy response in luminal B breast cancer. Thorac Cancer. 2021;12:3396-3406. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 19] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 39. | Takeda Y, Venkitaraman AR. Micro(mi) RNA-34a targets protein phosphatase (PP)1γ to regulate DNA damage tolerance. Cell Cycle. 2015;14:3830-3841. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 8] [Article Influence: 0.7] [Reference Citation Analysis (0)] |
| 40. | Marques SC, Ranjbar B, Laursen MB, Falgreen S, Bilgrau AE, Bødker JS, Jørgensen LK, Primo MN, Schmitz A, Ettrup MS, Johnsen HE, Bøgsted M, Mikkelsen JG, Dybkær K. High miR-34a expression improves response to doxorubicin in diffuse large B-cell lymphoma. Exp Hematol. 2016;44:238-46.e2. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 52] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 41. | Li QC, Xu H, Wang X, Wang T, Wu J. miR-34a increases cisplatin sensitivity of osteosarcoma cells in vitro through up-regulation of c-Myc and Bim signal. Cancer Biomark. 2017;21:135-144. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 34] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 42. | Chen X, Yin C, Liu QF, Liu JX. Regulation Effect of miR-34a Expression on Radiosensitivity of Lung Adenocarcinoma Cells by Targeting Bcl-2 and CDK4/6 Signaling Pathways. J Cancer Ther. 2022;13:187-198. [DOI] [Full Text] |
| 43. | Zhu W, Liu M, Fan Y, Ma F, Xu N, Xu B. Dynamics of circulating microRNAs as a novel indicator of clinical response to neoadjuvant chemotherapy in breast cancer. Cancer Med. 2018;7:4420-4433. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 19] [Cited by in RCA: 37] [Article Influence: 4.6] [Reference Citation Analysis (0)] |
| 44. | Chang TC, Wentzel EA, Kent OA, Ramachandran K, Mullendore M, Lee KH, Feldmann G, Yamakuchi M, Ferlito M, Lowenstein CJ, Arking DE, Beer MA, Maitra A, Mendell JT. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol Cell. 2007;26:745-752. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1609] [Cited by in RCA: 1549] [Article Influence: 81.5] [Reference Citation Analysis (0)] |
| 45. | Bommer GT, Gerin I, Feng Y, Kaczorowski AJ, Kuick R, Love RE, Zhai Y, Giordano TJ, Qin ZS, Moore BB, MacDougald OA, Cho KR, Fearon ER. p53-mediated activation of miRNA34 candidate tumor-suppressor genes. Curr Biol. 2007;17:1298-1307. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 832] [Cited by in RCA: 857] [Article Influence: 45.1] [Reference Citation Analysis (3)] |
| 46. | Imani S, Wei C, Cheng J, Khan MA, Fu S, Yang L, Tania M, Zhang X, Xiao X, Zhang X, Fu J. MicroRNA-34a targets epithelial to mesenchymal transition-inducing transcription factors (EMT-TFs) and inhibits breast cancer cell migration and invasion. Oncotarget. 2017;8:21362-21379. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 97] [Cited by in RCA: 96] [Article Influence: 10.7] [Reference Citation Analysis (0)] |
| 47. | Ding XM. MicroRNAs: regulators of cancer metastasis and epithelial-mesenchymal transition (EMT). Chin J Cancer. 2014;33:140-147. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 67] [Cited by in RCA: 79] [Article Influence: 6.1] [Reference Citation Analysis (1)] |
| 48. | Tazawa H, Tsuchiya N, Izumiya M, Nakagama H. Tumor-suppressive miR-34a induces senescence-like growth arrest through modulation of the E2F pathway in human colon cancer cells. Proc Natl Acad Sci U S A. 2007;104:15472-15477. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 758] [Cited by in RCA: 740] [Article Influence: 38.9] [Reference Citation Analysis (4)] |
| 49. | Weeraratne SD, Amani V, Neiss A, Teider N, Scott DK, Pomeroy SL, Cho YJ. miR-34a confers chemosensitivity through modulation of MAGE-A and p53 in medulloblastoma. Neuro Oncol. 2011;13:165-175. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 92] [Cited by in RCA: 108] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 50. | Navarro F, Lieberman J. miR-34 and p53: New Insights into a Complex Functional Relationship. PLoS One. 2015;10:e0132767. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 118] [Cited by in RCA: 159] [Article Influence: 14.5] [Reference Citation Analysis (0)] |
| 51. | Deng S, Wang M, Wang C, Zeng Y, Qin X, Tan Y, Liang B, Cao Y. p53 downregulates PD-L1 expression via miR-34a to inhibit the growth of triple-negative breast cancer cells: a potential clinical immunotherapeutic target. Mol Biol Rep. 2023;50:577-587. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 29] [Article Influence: 9.7] [Reference Citation Analysis (0)] |
| 52. | Kong D, Heath E, Chen W, Cher M, Powell I, Heilbrun L, Li Y, Ali S, Sethi S, Hassan O, Hwang C, Gupta N, Chitale D, Sakr WA, Menon M, Sarkar FH. Epigenetic silencing of miR-34a in human prostate cancer cells and tumor tissue specimens can be reversed by BR-DIM treatment. Am J Transl Res. 2012;4:14-23. [PubMed] |
| 53. | Zeng Z, Chen X, Zhu D, Luo Z, Yang M. Low Expression of Circulating MicroRNA-34c is Associated with Poor Prognosis in Triple-Negative Breast Cancer. Yonsei Med J. 2017;58:697-702. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 34] [Cited by in RCA: 44] [Article Influence: 4.9] [Reference Citation Analysis (0)] |
| 54. | Liu Y, Yang Y, Wang X, Yin S, Liang B, Zhang Y, Fan M, Fu Z, Shen C, Han Y, Chen B, Zhang Q. Function of microRNA124 in the pathogenesis of cancer (Review). Int J Oncol. 2024;64:6. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 19] [Reference Citation Analysis (0)] |
| 55. | Gourishetti K, Balaji Easwaran V, Mostakim Y, Ranganath Pai KS, Bhere D. MicroRNA (miR)-124: A Promising Therapeutic Gateway for Oncology. Biology (Basel). 2023;12:922. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 9] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 56. | Braga EA, Fridman MV, Burdennyy AM, Filippova EA, Loginov VI, Pronina IV, Dmitriev AA, Kushlinskii NE. Regulation of the Key Epithelial Cancer Suppressor miR-124 Function by Competing Endogenous RNAs. Int J Mol Sci. 2022;23:13620. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 18] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 57. | Zhou Z, Lv J, Wang J, Yu H, Lu H, Yuan B, Han J, Zhou R, Zhang X, Yang X, Yang H, Lu Q, Li P. Role of MicroRNA-124 as a Prognostic Factor in Multiple Neoplasms: A Meta-Analysis. Dis Markers. 2019;2019:1654780. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 9] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 58. | Han ZB, Yang Z, Chi Y, Zhang L, Wang Y, Ji Y, Wang J, Zhao H, Han ZC. MicroRNA-124 suppresses breast cancer cell growth and motility by targeting CD151. Cell Physiol Biochem. 2013;31:823-832. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 60] [Cited by in RCA: 71] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 59. | Du S, Li H, Sun X, Li D, Yang Y, Tao Z, Li Q, Liu K. MicroRNA-124 inhibits cell proliferation and migration by regulating SNAI2 in breast cancer. Oncol Rep. 2016;36:3259-3266. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 24] [Article Influence: 2.4] [Reference Citation Analysis (0)] |
| 60. | Zhang L, Chen X, Liu B, Han J. MicroRNA-124-3p directly targets PDCD6 to inhibit metastasis in breast cancer. Oncol Lett. 2018;15:984-990. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 19] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 61. | Cha N, Jia B, He Y, Luan W, Bao W, Han X, Gao W, Gao Y. MicroRNA-124 suppresses the invasion and proliferation of breast cancer cells by targeting TFAP4. Oncol Lett. 2021;21:271. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 10] [Reference Citation Analysis (0)] |
| 62. | Yang Q, Wan L, Xiao C, Hu H, Wang L, Zhao J, Lei Z, Zhang HT. Inhibition of LHX2 by miR-124 suppresses cellular migration and invasion in non-small cell lung cancer. Oncol Lett. 2017;14:3429-3436. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 23] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 63. | Moubarak RS, Koetz-Ploch L, Mullokandov G, Gaziel A, de Pablos-Aragoneses A, Argibay D, Kleffman K, Sokolova E, Berwick M, Thomas NE, Osman I, Brown BD, Hernando E. In Vivo miRNA Decoy Screen Reveals miR-124a as a Suppressor of Melanoma Metastasis. Front Oncol. 2022;12:852952. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 3] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 64. | Chen Z, Liu S, Tian L, Wu M, Ai F, Tang W, Zhao L, Ding J, Zhang L, Tang A. miR-124 and miR-506 inhibit colorectal cancer progression by targeting DNMT3B and DNMT1. Oncotarget. 2015;6:38139-38150. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 68] [Cited by in RCA: 75] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 65. | Yan G, Li Y, Zhan L, Sun S, Yuan J, Wang T, Yin Y, Dai Z, Zhu Y, Jiang Z, Liu L, Fan Y, Yang F, Hu W. Decreased miR-124-3p promoted breast cancer proliferation and metastasis by targeting MGAT5. Am J Cancer Res. 2019;9:585-596. [PubMed] |
| 66. | Feng T, Zhang P, Sun Y, Han X, Tong J, Hua Z. Evaluation of the Role of hsa-mir-124 in Predicting Clinical Outcome in Breast Invasive Carcinoma Based on Bioinformatics Analysis. Biomed Res Int. 2020;2020:1839205. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 6] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 67. | Xie XQ, Wang MJ, Li Y, Lei LP, Wang N, Lv ZY, Chen KL, Zhou B, Ping J, Zhou ZG, Sun XF. miR-124 Intensified Oxaliplatin-Based Chemotherapy by Targeting CAPN2 in Colorectal Cancer. Mol Ther Oncolytics. 2020;17:320-331. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 15] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 68. | Qiao W, Guo B, Zhou H, Xu W, Chen Y, Liang Y, Dong B. miR-124 suppresses glioblastoma growth and potentiates chemosensitivity by inhibiting AURKA. Biochem Biophys Res Commun. 2017;486:43-48. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 43] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 69. | Wang LQ, Sun W, Wang Y, Li D, Hu AM. Downregulation of plasma miR-124 expression is a predictive biomarker for prognosis of glioma. Eur Rev Med Pharmacol Sci. 2019;23:271-276. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 4] [Reference Citation Analysis (0)] |
| 70. | Chen F, Luo N, Hu Y, Li X, Zhang K. MiR-137 Suppresses Triple-Negative Breast Cancer Stemness and Tumorigenesis by Perturbing BCL11A-DNMT1 Interaction. Cell Physiol Biochem. 2018;47:2147-2158. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 41] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 71. | Lee SJ, Jeong JH, Kang SH, Kang J, Kim EA, Lee J, Jung JH, Park HY, Chae YS. MicroRNA-137 Inhibits Cancer Progression by Targeting Del-1 in Triple-Negative Breast Cancer Cells. Int J Mol Sci. 2019;20:6162. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 25] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 72. | Balaguer F, Link A, Lozano JJ, Cuatrecasas M, Nagasaka T, Boland CR, Goel A. Epigenetic silencing of miR-137 is an early event in colorectal carcinogenesis. Cancer Res. 2010;70:6609-6618. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 249] [Cited by in RCA: 231] [Article Influence: 14.4] [Reference Citation Analysis (0)] |
| 73. | Cheng Y, Li Y, Liu D, Zhang R, Zhang J. miR-137 effects on gastric carcinogenesis are mediated by targeting Cox-2-activated PI3K/AKT signaling pathway. FEBS Lett. 2014;588:3274-3281. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 36] [Cited by in RCA: 39] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 74. | Chen Q, Chen X, Zhang M, Fan Q, Luo S, Cao X. miR-137 is frequently down-regulated in gastric cancer and is a negative regulator of Cdc42. Dig Dis Sci. 2011;56:2009-2016. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 96] [Cited by in RCA: 104] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 75. | Du Y, Chen Y, Wang F, Gu L. miR-137 plays tumor suppressor roles in gastric cancer cell lines by targeting KLF12 and MYO1C. Tumour Biol. 2016;37:13557-13569. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 39] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 76. | Chen T, Lei S, Zeng Z, Pan S, Zhang J, Xue Y, Sun Y, Lan J, Xu S, Mao D, Guo B. MicroRNA137 suppresses the proliferation, migration and invasion of cholangiocarcinoma cells by targeting WNT2B. Int J Mol Med. 2020;45:886-896. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 8] [Article Influence: 1.3] [Reference Citation Analysis (0)] |
| 77. | Chang X, Zhang H, Lian S, Zhu W. miR-137 suppresses tumor growth of malignant melanoma by targeting aurora kinase A. Biochem Biophys Res Commun. 2016;475:251-256. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 18] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 78. | Luo C, Tetteh PW, Merz PR, Dickes E, Abukiwan A, Hotz-Wagenblatt A, Holland-Cunz S, Sinnberg T, Schittek B, Schadendorf D, Diederichs S, Eichmüller SB. miR-137 inhibits the invasion of melanoma cells through downregulation of multiple oncogenic target genes. J Invest Dermatol. 2013;133:768-775. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 101] [Cited by in RCA: 110] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 79. | Dong P, Xiong Y, Watari H, Hanley SJ, Konno Y, Ihira K, Yamada T, Kudo M, Yue J, Sakuragi N. MiR-137 and miR-34a directly target Snail and inhibit EMT, invasion and sphere-forming ability of ovarian cancer cells. J Exp Clin Cancer Res. 2016;35:132. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 78] [Cited by in RCA: 106] [Article Influence: 10.6] [Reference Citation Analysis (0)] |
| 80. | Su TJ, Ku WH, Chen HY, Hsu YC, Hong QS, Chang GC, Yu SL, Chen JJ. Oncogenic miR-137 contributes to cisplatin resistance via repressing CASP3 in lung adenocarcinoma. Am J Cancer Res. 2016;6:1317-1330. [PubMed] |
| 81. | Ying X, Sun Y, He P. MicroRNA-137 inhibits BMP7 to enhance the epithelial-mesenchymal transition of breast cancer cells. Oncotarget. 2017;8:18348-18358. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 33] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 82. | Lee JM, Cho KW, Kim EJ, Tang Q, Kim KS, Tickle C, Jung HS. A contrasting function for miR-137 in embryonic mammogenesis and adult breast carcinogenesis. Oncotarget. 2015;6:22048-22059. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 10] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 83. | Kudela E, Samec M, Koklesova L, Liskova A, Kubatka P, Kozubik E, Rokos T, Pribulova T, Gabonova E, Smolar M, Biringer K. miRNA Expression Profiles in Luminal A Breast Cancer-Implications in Biology, Prognosis, and Prediction of Response to Hormonal Treatment. Int J Mol Sci. 2020;21:7691. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 35] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 84. | Zhao Y, Li Y, Lou G, Zhao L, Xu Z, Zhang Y, He F. MiR-137 targets estrogen-related receptor alpha and impairs the proliferative and migratory capacity of breast cancer cells. PLoS One. 2012;7:e39102. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 96] [Cited by in RCA: 103] [Article Influence: 7.4] [Reference Citation Analysis (0)] |
| 85. | Chae YS, Baek DW, Lee IH, Lee SJ, Lee RK, Lee J, Jung J, Park H, Jeong J, Kang J, Park J. MicroRNA-137 inhibits cancer progression by targeting DEL-1 in triple negative breast cancer cells, MDA-MB-231. Cancer Res. 2019;79:P6-05. [RCA] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 86. | Denis H, Van Grembergen O, Delatte B, Dedeurwaerder S, Putmans P, Calonne E, Rothé F, Sotiriou C, Fuks F, Deplus R. MicroRNAs regulate KDM5 histone demethylases in breast cancer cells. Mol Biosyst. 2016;12:404-413. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 37] [Cited by in RCA: 33] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 87. | Zhang W, Chen JH, Shan T, Aguilera-Barrantes I, Wang LS, Huang TH, Rader JS, Sheng X, Huang YW. miR-137 is a tumor suppressor in endometrial cancer and is repressed by DNA hypermethylation. Lab Invest. 2018;98:1397-1407. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 47] [Cited by in RCA: 61] [Article Influence: 7.6] [Reference Citation Analysis (0)] |
| 88. | Du F, Yu L, Wu Y, Wang S, Yao J, Zheng X, Xie S, Zhang S, Lu X, Liu Y, Chen W. miR-137 alleviates doxorubicin resistance in breast cancer through inhibition of epithelial-mesenchymal transition by targeting DUSP4. Cell Death Dis. 2019;10:922. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 55] [Cited by in RCA: 57] [Article Influence: 8.1] [Reference Citation Analysis (4)] |
| 89. | Zhu X, Li Y, Shen H, Li H, Long L, Hui L, Xu W. miR-137 restoration sensitizes multidrug-resistant MCF-7/ADM cells to anticancer agents by targeting YB-1. Acta Biochim Biophys Sin (Shanghai). 2013;45:80-86. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 44] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 90. | Gu Q, Zhang J, Hu H, Tan YE, Shi S, Nian Y. Clinical Significance of MiR-137 Expression in Patients with Gastric Cancer After Radical Gastrectomy. PLoS One. 2015;10:e0142377. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 10] [Article Influence: 0.9] [Reference Citation Analysis (0)] |
| 91. | Li N. Low Expression of Mir-137 Predicts Poor Prognosis in Cutaneous Melanoma Patients. Med Sci Monit. 2016;22:140-144. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 18] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 92. | Steponaitiene R, Kupcinskas J, Langner C, Balaguer F, Venclauskas L, Pauzas H, Tamelis A, Skieceviciene J, Kupcinskas L, Malfertheiner P, Link A. Epigenetic silencing of miR-137 is a frequent event in gastric carcinogenesis. Mol Carcinog. 2016;55:376-386. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 45] [Cited by in RCA: 54] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 93. | Peres J, Kwesi-Maliepaard EM, Rambow F, Larue L, Prince S. The tumour suppressor, miR-137, inhibits malignant melanoma migration by targetting the TBX3 transcription factor. Cancer Lett. 2017;405:111-119. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 32] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 94. | Raeisi F, Mahmoudi E, Dehghani-Samani M, Hosseini SSE, Ghahfarrokhi AM, Arshi A, Forghanparast K, Ghazanfari S. Differential Expression Profile of miR-27b, miR-29a, and miR-155 in Chronic Lymphocytic Leukemia and Breast Cancer Patients. Mol Ther Oncolytics. 2020;16:230-237. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 28] [Article Influence: 4.7] [Reference Citation Analysis (0)] |
| 95. | Pasculli B, Barbano R, Fontana A, Biagini T, Di Viesti MP, Rendina M, Valori VM, Morritti M, Bravaccini S, Ravaioli S, Maiello E, Graziano P, Murgo R, Copetti M, Mazza T, Fazio VM, Esteller M, Parrella P. Hsa-miR-155-5p Up-Regulation in Breast Cancer and Its Relevance for Treatment With Poly[ADP-Ribose] Polymerase 1 (PARP-1) Inhibitors. Front Oncol. 2020;10:1415. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 49] [Article Influence: 8.2] [Reference Citation Analysis (0)] |
| 96. | Khalighfard S, Alizadeh AM, Irani S, Omranipour R. Plasma miR-21, miR-155, miR-10b, and Let-7a as the potential biomarkers for the monitoring of breast cancer patients. Sci Rep. 2018;8:17981. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 143] [Cited by in RCA: 121] [Article Influence: 15.1] [Reference Citation Analysis (0)] |
| 97. | Wang J, Wang Q, Guan Y, Sun Y, Wang X, Lively K, Wang Y, Luo M, Kim JA, Murphy EA, Yao Y, Cai G, Fan D. Breast cancer cell-derived microRNA-155 suppresses tumor progression via enhancing immune cell recruitment and antitumor function. J Clin Invest. 2022;132:e157248. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 67] [Article Influence: 16.8] [Reference Citation Analysis (0)] |
| 98. | Kohlhapp FJ, Mitra AK, Lengyel E, Peter ME. MicroRNAs as mediators and communicators between cancer cells and the tumor microenvironment. Oncogene. 2015;34:5857-5868. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 174] [Cited by in RCA: 171] [Article Influence: 15.5] [Reference Citation Analysis (0)] |
| 99. | Chang S, Wang RH, Akagi K, Kim KA, Martin BK, Cavallone L; Kathleen Cuningham Foundation Consortium for Research into Familial Breast Cancer (kConFab), Haines DC, Basik M, Mai P, Poggi E, Isaacs C, Looi LM, Mun KS, Greene MH, Byers SW, Teo SH, Deng CX, Sharan SK. Tumor suppressor BRCA1 epigenetically controls oncogenic microRNA-155. Nat Med. 2011;17:1275-1282. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 163] [Cited by in RCA: 182] [Article Influence: 12.1] [Reference Citation Analysis (0)] |
| 100. | Gasparini P, Lovat F, Fassan M, Casadei L, Cascione L, Jacob NK, Carasi S, Palmieri D, Costinean S, Shapiro CL, Huebner K, Croce CM. Protective role of miR-155 in breast cancer through RAD51 targeting impairs homologous recombination after irradiation. Proc Natl Acad Sci U S A. 2014;111:4536-4541. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 161] [Cited by in RCA: 169] [Article Influence: 14.1] [Reference Citation Analysis (1)] |
| 101. | Sochor M, Basova P, Pesta M, Dusilkova N, Bartos J, Burda P, Pospisil V, Stopka T. Oncogenic microRNAs: miR-155, miR-19a, miR-181b, and miR-24 enable monitoring of early breast cancer in serum. BMC Cancer. 2014;14:448. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 141] [Cited by in RCA: 140] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 102. | Han JG, Jiang YD, Zhang CH, Yang YM, Pang D, Song YN, Zhang GQ. A novel panel of serum miR-21/miR-155/miR-365 as a potential diagnostic biomarker for breast cancer. Ann Surg Treat Res. 2017;92:55-66. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 57] [Cited by in RCA: 87] [Article Influence: 9.7] [Reference Citation Analysis (0)] |
| 103. | Jurkovicova D, Smolkova B, Magyerkova M, Sestakova Z, Kajabova VH, Kulcsar L, Zmetakova I, Kalinkova L, Krivulcik T, Karaba M, Benca J, Sedlackova T, Minarik G, Cierna Z, Danihel L, Mego M, Chovanec M, Fridrichova I. Down-regulation of traditional oncomiRs in plasma of breast cancer patients. Oncotarget. 2017;8:77369-77384. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 25] [Cited by in RCA: 37] [Article Influence: 4.1] [Reference Citation Analysis (0)] |
| 104. | Papadaki C, Thomopoulou K, Monastirioti A, Koronakis G, Papadaki MA, Rounis K, Vamvakas L, Nikolaou C, Mavroudis D, Agelaki S. MicroRNAs Regulating Tumor and Immune Cell Interactions in the Prediction of Relapse in Early Stage Breast Cancer. Biomedicines. 2021;9:421. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 10] [Reference Citation Analysis (0)] |
| 105. | Eichelser C, Flesch-Janys D, Chang-Claude J, Pantel K, Schwarzenbach H. Deregulated serum concentrations of circulating cell-free microRNAs miR-17, miR-34a, miR-155, and miR-373 in human breast cancer development and progression. Clin Chem. 2013;59:1489-1496. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 142] [Cited by in RCA: 150] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 106. | Bašová P, Pešta M, Sochor M, Stopka T. Prediction Potential of Serum miR-155 and miR-24 for Relapsing Early Breast Cancer. Int J Mol Sci. 2017;18:2116. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 29] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 107. | Huang Q, Gumireddy K, Schrier M, le Sage C, Nagel R, Nair S, Egan DA, Li A, Huang G, Klein-Szanto AJ, Gimotty PA, Katsaros D, Coukos G, Zhang L, Puré E, Agami R. The microRNAs miR-373 and miR-520c promote tumour invasion and metastasis. Nat Cell Biol. 2008;10:202-210. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 790] [Cited by in RCA: 731] [Article Influence: 40.6] [Reference Citation Analysis (4)] |
| 108. | Wei F, Cao C, Xu X, Wang J. Diverse functions of miR-373 in cancer. J Transl Med. 2015;13:162. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 61] [Cited by in RCA: 83] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 109. | Chen D, Dang BL, Huang JZ, Chen M, Wu D, Xu ML, Li R, Yan GR. MiR-373 drives the epithelial-to-mesenchymal transition and metastasis via the miR-373-TXNIP-HIF1α-TWIST signaling axis in breast cancer. Oncotarget. 2015;6:32701-32712. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 78] [Cited by in RCA: 100] [Article Influence: 10.0] [Reference Citation Analysis (0)] |
| 110. | van Schooneveld E, Wildiers H, Vergote I, Vermeulen PB, Dirix LY, Van Laere SJ. Dysregulation of microRNAs in breast cancer and their potential role as prognostic and predictive biomarkers in patient management. Breast Cancer Res. 2015;17:21. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 232] [Cited by in RCA: 224] [Article Influence: 20.4] [Reference Citation Analysis (0)] |
| 111. | Rahmani F, Tadayyon Tabrizi A, Hashemian P, Alijannejad S, Rahdar HA, Ferns GA, Hassanian SM, Shahidsales S, Avan A. Role of regulatory miRNAs of the Wnt/ β-catenin signaling pathway in tumorigenesis of breast cancer. Gene. 2020;754:144892. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 24] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 112. | Yan GR, Xu SH, Tan ZL, Liu L, He QY. Global identification of miR-373-regulated genes in breast cancer by quantitative proteomics. Proteomics. 2011;11:912-920. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 68] [Cited by in RCA: 72] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 113. | Tang J, Ahmad A, Sarkar FH. The role of microRNAs in breast cancer migration, invasion and metastasis. Int J Mol Sci. 2012;13:13414-13437. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 112] [Cited by in RCA: 145] [Article Influence: 10.4] [Reference Citation Analysis (2)] |
| 114. | Keklikoglou I, Koerner C, Schmidt C, Zhang JD, Heckmann D, Shavinskaya A, Allgayer H, Gückel B, Fehm T, Schneeweiss A, Sahin O, Wiemann S, Tschulena U. MicroRNA-520/373 family functions as a tumor suppressor in estrogen receptor negative breast cancer by targeting NF-κB and TGF-β signaling pathways. Oncogene. 2012;31:4150-4163. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 212] [Cited by in RCA: 217] [Article Influence: 14.5] [Reference Citation Analysis (3)] |
| 115. | Voorhoeve PM, le Sage C, Schrier M, Gillis AJ, Stoop H, Nagel R, Liu YP, van Duijse J, Drost J, Griekspoor A, Zlotorynski E, Yabuta N, De Vita G, Nojima H, Looijenga LH, Agami R. A genetic screen implicates miRNA-372 and miRNA-373 as oncogenes in testicular germ cell tumors. Cell. 2006;124:1169-1181. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 954] [Cited by in RCA: 941] [Article Influence: 47.1] [Reference Citation Analysis (3)] |
| 116. | O'Bryan S, Dong S, Mathis JM, Alahari SK. The roles of oncogenic miRNAs and their therapeutic importance in breast cancer. Eur J Cancer. 2017;72:1-11. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 84] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 117. | Bakr NM, Mahmoud MS, Nabil R, Boushnak H, Swellam M. Impact of circulating miRNA-373 on breast cancer diagnosis through targeting VEGF and cyclin D1 genes. J Genet Eng Biotechnol. 2021;19:84. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 26] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 118. | Zhang X, Li X, Tan Z, Liu X, Yang C, Ding X, Hu X, Zhou J, Xiang S, Zhou C, Zhang J. MicroRNA-373 is upregulated and targets TNFAIP1 in human gastric cancer, contributing to tumorigenesis. Oncol Lett. 2013;6:1427-1434. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 35] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 119. | Chen Q, Li Y, Lu T, Luo J, Yang L, Zhou Z, Tian Z, Tan S, Liu Q. miR-373 promotes invasion and metastasis of colorectal cancer cells via activating ERK/MAPK pathway. Sci Rep. 2024;14:124. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 14] [Reference Citation Analysis (0)] |
| 120. | Zhang Y, Zhao FJ, Chen LL, Wang LQ, Nephew KP, Wu YL, Zhang S. MiR-373 targeting of the Rab22a oncogene suppresses tumor invasion and metastasis in ovarian cancer. Oncotarget. 2014;5:12291-12303. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 53] [Cited by in RCA: 65] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 121. | Yin QH, Zhou Y, Li ZHY. miR-373 Suppresses Cell Proliferation and Apoptosis via Regulation of SIRT1/PGC-1α/NRF2 Axis in Pancreatic Cancer. Cell J. 2021;23:199-210. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 9] [Reference Citation Analysis (0)] |
| 122. | Seol HS, Akiyama Y, Shimada S, Lee HJ, Kim TI, Chun SM, Singh SR, Jang SJ. Epigenetic silencing of microRNA-373 to epithelial-mesenchymal transition in non-small cell lung cancer through IRAK2 and LAMP1 axes. Cancer Lett. 2014;353:232-241. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 61] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 123. | Shah JA, Khattak S, Rauf MA, Cai Y, Jin J. Potential Biomarkers of miR-371-373 Gene Cluster in Tumorigenesis. Life (Basel). 2021;11:984. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 9] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 124. | Peng T, Wang T, Liu G, Zhou L. Effects of miR-373 Inhibition on Glioblastoma Growth by Reducing Limk1 In Vitro. J Immunol Res. 2020;2020:7671502. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 6] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 125. | Adi Harel S, Bossel Ben-Moshe N, Aylon Y, Bublik DR, Moskovits N, Toperoff G, Azaiza D, Biagoni F, Fuchs G, Wilder S, Hellman A, Blandino G, Domany E, Oren M. Reactivation of epigenetically silenced miR-512 and miR-373 sensitizes lung cancer cells to cisplatin and restricts tumor growth. Cell Death Differ. 2015;22:1328-1340. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 47] [Cited by in RCA: 65] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 126. | Lindholm EM, Ragle Aure M, Haugen MH, Kleivi Sahlberg K, Kristensen VN, Nebdal D, Børresen-Dale AL, Lingjaerde OC, Engebraaten O. miRNA expression changes during the course of neoadjuvant bevacizumab and chemotherapy treatment in breast cancer. Mol Oncol. 2019;13:2278-2296. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 42] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 127. | Li C, Feng S, Chen L. MSC-AS1 knockdown inhibits cell growth and temozolomide resistance by regulating miR-373-3p/CPEB4 axis in glioma through PI3K/Akt pathway. Mol Cell Biochem. 2021;476:699-713. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 36] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 128. | Hu W, Liu Q, Pan J, Sui Z. MiR-373-3p enhances the chemosensitivity of gemcitabine through cell cycle pathway by targeting CCND2 in pancreatic carcinoma cells. Biomed Pharmacother. 2018;105:887-898. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 19] [Cited by in RCA: 32] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 129. | Kutanzi KR, Yurchenko OV, Beland FA, Checkhun VF, Pogribny IP. MicroRNA-mediated drug resistance in breast cancer. Clin Epigenetics. 2011;2:171-185. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 123] [Cited by in RCA: 131] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 130. | Maryam M, Naemi M, Hasani SS. A comprehensive review on oncogenic miRNAs in breast cancer. J Genet. 2021;100:15. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 27] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 131. | Xing Y, Ruan G, Ni H, Qin H, Chen S, Gu X, Shang J, Zhou Y, Tao X, Zheng L. Tumor Immune Microenvironment and Its Related miRNAs in Tumor Progression. Front Immunol. 2021;12:624725. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 60] [Cited by in RCA: 79] [Article Influence: 15.8] [Reference Citation Analysis (1)] |
| 132. | Chen W, Cai F, Zhang B, Barekati Z, Zhong XY. The level of circulating miRNA-10b and miRNA-373 in detecting lymph node metastasis of breast cancer: potential biomarkers. Tumour Biol. 2013;34:455-462. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 117] [Cited by in RCA: 123] [Article Influence: 8.8] [Reference Citation Analysis (0)] |
| 133. | Anas HA, Hassan MA, El-mokhtar MA, Eltybe HA, Hassan HM. Aberrant Expression of Circulating MicroRNA-373 and MicroRNA-660 as Biomarkers for Diagnosis of Breast Cancer in Egyptian Females. Egypt J Med Microbiol. 2025;34:17-25. [DOI] [Full Text] |
| 134. | Müller V, Gade S, Steinbach B, Loibl S, von Minckwitz G, Untch M, Schwedler K, Lübbe K, Schem C, Fasching PA, Mau C, Pantel K, Schwarzenbach H. Changes in serum levels of miR-21, miR-210, and miR-373 in HER2-positive breast cancer patients undergoing neoadjuvant therapy: a translational research project within the Geparquinto trial. Breast Cancer Res Treat. 2014;147:61-68. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 82] [Cited by in RCA: 97] [Article Influence: 8.1] [Reference Citation Analysis (0)] |
| 135. | Eichelser C, Stückrath I, Müller V, Milde-Langosch K, Wikman H, Pantel K, Schwarzenbach H. Increased serum levels of circulating exosomal microRNA-373 in receptor-negative breast cancer patients. Oncotarget. 2014;5:9650-9663. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 225] [Cited by in RCA: 282] [Article Influence: 25.6] [Reference Citation Analysis (0)] |
| 136. | Hua Y, Chen H, Wang L, Wang F, Wang P, Ning Z, Li Y, Liu L, Chen Z, Meng Z. Low serum miR-373 predicts poor prognosis in patients with pancreatic cancer. Cancer Biomark. 2017;20:95-100. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 31] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 137. | Jawad Nayyef H, Hussien Aziz I. The Impact of miRNA-425 and miRNA-373 in the Pathogenesis of Breast Cancer. Arch Breast Cancer. 2025;12:171-180. [DOI] [Full Text] |
| 138. | Samara M, Thodou E, Patoulioti M, Poultsidi A, Thomopoulou GE, Giakountis A. Integrated miRNA Signatures: Advancing Breast Cancer Diagnosis and Prognosis. Biomolecules. 2024;14:1352. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 139. | Jing SY, Jing SQ, Liu LL, Xu LF, Zhang F, Gao JL. Down-expression of miR-373 predicts poor prognosis of glioma and could be a potential therapeutic target. Eur Rev Med Pharmacol Sci. 2017;21:2421-2425. [PubMed] |
| 140. | Tu HF, Chang KW, Cheng HW, Liu CJ. Upregulation of miR-372 and -373 associates with lymph node metastasis and poor prognosis of oral carcinomas. Laryngoscope. 2015;125:E365-E370. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 42] [Article Influence: 3.8] [Reference Citation Analysis (0)] |