Published online Mar 24, 2026. doi: 10.5306/wjco.v17.i3.117055
Revised: December 20, 2025
Accepted: February 10, 2026
Published online: March 24, 2026
Processing time: 110 Days and 17.8 Hours
PIGU has been proposed as a potential oncogene in bladder cancer (BLCA); however, experimental evidence regarding PIGU expression in BLCA remains lacking, necessitating further investigation into its functions and underlying mechanisms.
To explore the expression of PIGU in BLCA and to understand its function and underlying mechanisms.
This study assessed PIGU protein expression in BLCA tissues via immunohistochemistry. We integrated 14 high-throughput datasets from TCGA, GEO, and Array Express to evaluate PIGU’s mRNA expression levels and diagnostic efficacy. siRNA-knockdown cell lines were established using the PIGU-overexpressing HT-1376 BLCA cell line. Proliferation capacity was assessed via CCK-8 assays, while apoptosis and cell cycle distribution were evaluated using flow cytometry. PIGU’s potential mechanism in BLCA were explored through differential gene enrichment analysis, protein interaction network analysis, and immune infiltration analysis.
PIGU expression was significantly upregulated in 121 BLCA and 32 non-BLCA samples at the protein level (10.65 ± 2.14 vs 5.56 ± 2.68, P < 0.0001) and in 1065 BLCA and 170 non-BLCA samples at the mRNA level (standardized mean difference = 0.74, 95% confidence interval: 0.07-1.41). Elevated PIGU expression correlated with poorer prognosis in BLCA patients, making it an independent prognostic factor. PIGU knockdown inhibited BLCA cell proliferation (P < 0.05), increased apoptosis, and caused cell cycle arrest at the G1 phase. PIGU-regulated genes are enriched in cell cycle pathways, with PCNA and MCM2 serving as hub genes. High PIGU expression was as
PIGU is upregulated in BLCA and has prognostic value. It promotes tumor progression by regulating the cell cycle and immune microenvironment, showing potential as an effective therapeutic target for BLCA.
Core Tip: This study indicated that PIGU expression is markedly increased in bladder cancer at both the mRNA and protein levels and that this elevation serves as an independent predictor of unfavorable clinical outcomes. In vitro experiments further revealed that PIGU knockdown suppresses cell proliferation and induces apoptosis accompanied by G1-phase arrest, while mechanistic analyses indicated that PIGU facilitates tumor progression by regulating key cell cycle-related genes and modulating the immune microenvironment. These results indicate that PIGU could function not only as an independent prognostic indicator but also as a promising candidate for therapeutic targets.
- Citation: Li SH, Liu ZS, Chen YJ, Li JD, Wei L, Chen YY, Zhang W, Huang ZG, Chen ZD, Chen G, Wei DM, Mo ZN, Deng LL. PIGU overexpression and regulation of cellular biological functions in bladder cancer. World J Clin Oncol 2026; 17(3): 117055
- URL: https://www.wjgnet.com/2218-4333/full/v17/i3/117055.htm
- DOI: https://dx.doi.org/10.5306/wjco.v17.i3.117055
Bladder cancer (BLCA) is the ninth most common malignancy worldwide and carries substantial incidence and mortality burdens[1]. Approximately 90% of cases arise from the urothelial cells lining the bladder and are classified as urothelial carcinomas[2], whose current treatment modalities include surgery, chemotherapy, radiotherapy, and immunotherapy. Although low-grade tumors generally have nonlethal outcomes, most high-grade tumors tend to progress to invasive disease, metastasis, and ultimately death[3]. Therefore, elucidating the novel molecular mechanisms underlying BLCA pathogenesis is essential for identifying alternative therapeutic strategies to improve patient survival.
PIGU, also known as CDC91 L1 or GAB1, is a key subunit of the glycosylphosphatidylinositol transferase (GPI-T) complex, which mediates the attachment of GPI anchors to proteins-an essential process involved in cancer progression[4]. PIGU has been reported to play diverse roles across multiple cancer types; however, its characterization in BLCA remains limited. Previous studies using PCR and northern blotting showed PIGU overexpression in four BLCA cell lines, and 22 of 61 primary BLCA specimens (36%) exhibited higher expression compared with normal tissues[5]. In contrast, another PCR-based study detected elevated PIGU mRNA expression in only one of eight BLCA cell lines and found no meaningful predictive value in 14 normal bladder tissues or 42 BLCA cases[6]. Further contradictory evidence emerged in 2008, when PIGU was found to promote promoting bladder tumorigenesis. The work used reverse transcription PCR (RT-PCR) and immunohistochemistry (IHC) on 73 BLCA and 14 normal urothelial samples, revealing that 30.1% (22/73) of tumors exhibited PIGU mRNA overexpression[7].
Given these inconsistent findings, larger-scale studies are required to clarify PIGU expression patterns in BLCA. Moreover, the underlying mechanisms in BLCA remain poorly understood. In this study, we evaluated PIGU protein expression in BLCA using IHC and integrated high-throughput datasets to assess its transcriptional expression. We further performed a series of in vitro functional assays following PIGU knockdown in the HT-1376 BLCA cell line to determine PIGU’s role in regulating cell proliferation, apoptosis, and associated molecular mechanisms. These results collectively address the limitations of prior studies and provide new insights into PIGU’s contribution to BLCA initiation and progression.
BLCA tissue microarrays BLC1021 and BLC481 were provided by Fanpu Biotech, Inc. (Guilin, China). An additional three pairs of BLCA and adjacent tumor tissue biopsies were obtained from the First Affiliated Hospital of Guangxi Medical University. Immunohistochemical staining was performed on 121 BLCA tissue samples and 32 non-BLCA tissue samples to assess PIGU protein levels. Following formalin fixation, dehydration, and paraffin embedding, IHC staining was performed according to manufacturer guidelines. Two on-site pathologists assessed staining intensity and the proportion of positively stained cells. Staining intensity was graded from 0 (no staining) to 3 (intense staining), while cell proportion was categorized from 0 (< 5%) to 4 (76%-100%). The final IHC score was calculated by multiplying the two scores, representing the degree of PIGU protein expression. Subsequently, further analysis was conducted to assess the correlation between clinical-pathological parameters and PIGU protein expression.
Global microarray and RNA-sequencing (RNA-seq) datasets were collected from multiple databases to analyze PIGU mRNA expression in BLCA. These databases included TCGA, GEO, and Array Express. Search terms were “BLCA” and “gene or mRNA”. Inclusion criteria were: Human samples, containing both BLCA and non-BLCA samples, ≥ 3 samples per group, and presence of PIGU expression data. Data sets were merged and batch effects were removed using the Combat function in the R sva package. TCGA data underwent log2 (TPM + 1) normalization. The final study included a total of 14 data sets from 8 high-throughput cohorts (1065 BLCA samples and 170 non-BLCA samples). After extracting PIGU expression data from BLCA and non-BLCA groups, analyses were performed using student’s t-test. Scatter plots were generated with GraphPad Prism 9.0.0 to visualize expression level distributions, and receiver operating characteristic (ROC) curves were plotted to evaluate the diagnostic accuracy of PIGU expression in distinguishing BLCA from non-BLCA tissues.
To enhance the reliability of results, we performed a meta-analysis of all data using Stata v14.0 by calculating the standardized mean difference (SMD) values for PIGU. A random-effects model was applied when substantial heterogeneity was detected, defined as I² values of 50% or higher accompanied by a P value below 0.05. An SMD > 0 and a 95% confidence interval (CI) not containing zero indicated PIGU overexpression in BLCA. Publication bias was assessed using the Egger and Begg tests.
In Stata v14.0, we plotted a summary ROC (SROC) curve using true positive, false negative, false positive, and true negative values from each dataset. The area under the curve (AUC) reflected the discriminatory ability of PIGU mRNA expression between BLCA and non-BLCA tissues. An AUC between 0.7 and 0.9 was considered moderate discrimination. Publication bias was assessed using Deek’s funnel plot. PIGU RNA-seq profiles and associated clinical-pathological information were obtained from the TCGA database for 405 BLCA patients. The correlation between PIGU mRNA levels and clinical-pathological parameters was evaluated using t-tests. Survival analysis was performed using the Gene Expression Profile Interaction Analysis database (http://gepia.cancer-pku.cn/) to examine overall survival (OS) and disease-free survival (DFS), with the lowest 25% representing the low group and the highest 25% representing the high group[8]. Results were presented using log-rank tests and Kaplan-Meier curves. Univariate and multivariate Cox regression models were applied to estimate hazard ratios (HR) and to assess the prognostic value of PIGU along with various clinicopathological characteristics in BLCA.
RPMI 1640 medium (Catalog No. L210KJ) and MEM medium (Catalog No. L510KJ) were purchased from Shanghai Yuanpei Biotechnology Co., Ltd.; FBS (Catalog No. RL09136) was purchased from Shanghai Ruibosi Biotechnology Co., Ltd.; penicillin-streptomycin dual antibiotics were purchased from Shanghai Bioengineering Co., Ltd.; DMSO (Catalog No. RNBK4714) was purchased from Sigma-Aldrich (Shanghai) Trading Co., Ltd. HRP-labeled goat anti-mouse secondary antibody (Catalog No. ZB2305) was purchased from Beijing Zhongshan Jinqiao Co., Ltd. PIGU antibody (Catalog No. Ab192255) purchased from Abcam; RIPA lysis buffer (Catalog No. P0013B) purchased from Beyotime; Lipo8000™ (Catalog No. C0533) purchased from Shanghai Biyun Tian Biotechnology Co., Ltd.; ECL chemiluminescent substrate (Catalog No. ECL-0011) purchased from Beijing Dingguo Company; human bladder carcinoma cell lines EJ-1, 5637, J82, and HT-1376 obtained from Cell Type Culture Collection Co., Ltd. were cultured in MEM or RPMI-1640 medium supplemented with 100 mL/L FBS and 10 mL/L antibiotic-antimycotic solution. Cells were maintained at 37 °C in a 50 mL/L CO2 incubator and passaged or replenished every 2-3 days based on cell density.
Experimental groups were assigned as follows: Group A (EJ-1 cells), group B (5637 cells), group C (J82 cells), and group D (HT-1376 cells). After collecting cells from each group, they were lysed using RIPA lysis buffer containing PMSF. Following centrifugation, the supernatant was collected and samples were stored at -80 °C. Protein samples were mixed with 5 × SDS loading buffer, denatured by heating in a water bath for 5 minutes, and centrifuged to remove impurities. Subsequently, polyacrylamide gels were prepared. The separating gel concentration was selected based on the target protein size, followed by gel loading and cooling. Electrophoresis was conducted at 80 V for approximately 30 minutes in the stacking gel, followed by 120 V for about 60 minutes in the resolving gel, until the bromophenol blue tracking dye migrated close to the lower edge of the gel. Proteins are transferred to a PVDF membrane using transfer buffer to equilibrate the membrane and complete the transfer. After washing the membrane with TBST, non-specific binding sites are blocked. The membrane is then incubated with appropriately diluted primary and secondary antibodies. Finally, visualization is achieved using ECL chemiluminescent substrate, and results are captured using a chemiluminescence imager.
First, seed HT-1376 cells into a six-well plate and culture until 80% confluence. Prior to transfection, replace each well with 2 mL fresh complete medium. For transfection, prepare clean centrifuge tubes containing 125 μL basal medium mixed with 100 pmol siRNA and 4 μL Lipo8000™ transfection reagent per well. Allow the mixture to stabilize at room temperature for 6 hours. Dispense 125 μL of the Lipo8000™ transfection reagent-siRNA mixture uniformly into each well of the 6-well plate. After transfection, continue culturing for 48 hours, then collect cell samples from each group for subsequent RT-PCR analysis. Total cellular RNA was extracted using the Trizol extraction kit. Primers used: PIGU, forward 5′-ACAACACCCTCATTGCTTTCTT-3′, reverse 5′-GATTACCACTAGGCTTCCCACA-3′; and GAPDH, forward 5′-GGAGCGAGATCCCTCCAAAAT-3′, reverse 5′-GGCTGTTGTCATACTTCTCATGG-3′. After normalization using the endogenous reference GAPDH, relative gene expression levels were calculated using the 2-∆∆Cq method to analyze gene expression differences between groups.
Total RNA was extracted from cells using TRIzol reagent, and cDNA was synthesized with a first-strand cDNA synthesis kit. Primers were designed and synthesized using Primer Premier 5.0 (Shanghai Sangon Biotech Co., Ltd.), with GAPDH serving as the internal control. The primer sequences were as follows: PIGU: Forward 5′-ACAACACCCTCATTGCTT
According to the experimental design, transfected HT-1376 cells were seeded into 96-well plates (group A: HT-1376 + siNC; group B: HT-1376 + siPIGU) at a concentration of 5 × 104 cells/mL (100 μL per well). At 0, 24, 48, and 72 hours, cell morphology was observed and photographed under a microscope. Subsequently, 10 μL of CCK-8 solution was added to each well, followed by incubation at 37 °C in a 50 mL/L CO2 incubator for 2 hours in the dark. The optical density at 450 nm was measured using a microplate reader to assess cell proliferation.
Cells were cultured according to the experimental groups (group A: HT-1376 + siNC; group B: HT-1376 + siPIGU) following the transfection procedure. After removing the culture medium, cells were washed three times with phosphate-buffered saline (PBS) and digested with EDTA-free trypsin to avoid damage to the cell membrane. Cells were collected by centrifugation at 1000 rpm/minute for 5 minutes at room temperature, washed once with PBS, centrifuged again at 2000 rpm/minute for 5 minutes, and resuspended to obtain (1-5) × 105 cells. The cells were first resuspended in 500 μL of Binding Buffer, then 5 μL of Annexin V-FITC and 5 μL of propidium iodide were added. Following gentle mixing, the samples were incubated at room temperature in the dark for 5-15 minutes, after which apoptosis was assessed using flow cytometry.
Based on the experimental groups (group A: HT-1376 + siNC; group B: HT-1376 + siPIGU), HT-1376 cells were cultured to the logarithmic growth phase, adjusted to a concentration of 2 × 105 cells/mL, and seeded into 6-well plates (2 mL per well). After transfection, cells were cultured for 48 hours, harvested by centrifugation, washed 1-2 times with pre-chilled PBS, and fixed overnight at 4 °C with pre-chilled 700 mL/L ethanol. After fixation, cells were centrifuged at 1000 rpm for 3 minutes, washed three times with PBS, and resuspended in PBS containing RNase A (100 μg/mL). Samples were incubated at 37 °C for 30 minutes, followed by staining with propidium iodide (PI, 50 μg/mL) for 30 minutes in the dark. Finally, cells were washed three times with pre-chilled PBS and analyzed for cell cycle distribution by flow cytometry.
For each included dataset, Pearson correlation coefficients between each gene and PIGU were calculated using R v3.6.3. The following filtering criteria were applied to identify PIGU co-expressed genes: |r| ≥ 0.3; P < 0.05; ≥ 3 replicates. We created the differentially co-expressed gene (DCEG) set by intersecting differentially expressed genes (DEGs) in BLCA with these co-expressed genes. Subsequently, for gene enrichment analysis, including Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) analysis, we employed the clusterProfiler package in R v3.6. Following enrichment analysis, genes were selected from the most significantly enriched pathways for subsequent protein-protein interaction (PPI) network analysis[9]. For this purpose, we utilized the String database (https://string-db.org/) and the GeneMANIA database (https://genemania.org/)[10,11].
We utilized the TIMER server (http://cistrome.dfci.harvard.edu/TIMER/) to assess tumor-specific immune infiltration[12]. Through the TIMER database, we examined the correlation between PIGU expression and immune cell infiltration (CD4+ T cells, CD8+ T cells, neutrophils, and dendritic cells) in BLCA. Using the ESTIMATE algorithm, we estimated the proportions of immune cells and stromal cells in tumor samples based on gene expression data[13]. In this study, we generated immune scores, stroma scores, and ESTIMATE scores for each TCGA_BLCA sample using the ESTIMATE package in R v3.6.3. These scores quantify the immune and stromal cell components within the tumor microenvironment. By correlating these scores with PIGU mRNA expression, we aimed to elucidate the interaction between immune cell infiltration and PIGU expression in BLCA. Additionally, we performed correlation analyses between PIGU and immune checkpoint molecules to investigate potential mechanisms driving PIGU expression and immune infiltration.
Data were analyzed using SPSS v26.0. PIGU expression levels are presented as mean ± SD. Appropriate statistical tests were applied based on data type: Independent t-tests for continuous variables (e.g., PIGU expression levels) and χ2 tests for categorical variables (e.g., immune cell infiltration status). For non-normally distributed data, the Mann-Whitney U test was used. A P value < 0.05 was considered statistically significant.
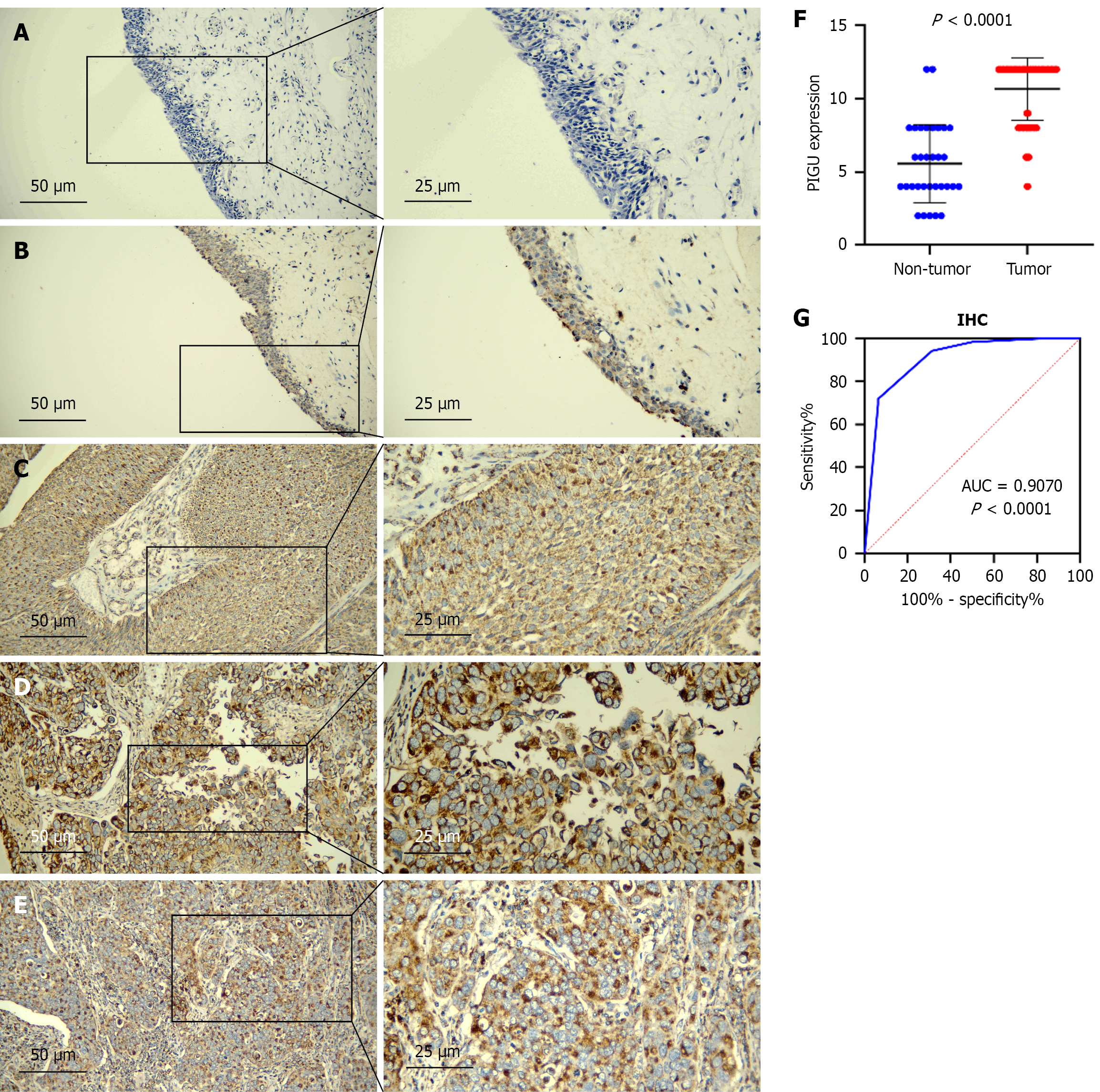
IHC staining revealed that PIGU protein expression intensity was higher in BLCA tissue than in normal tissue (Figure 1A and B). Staining intensity was relatively consistent across different pathological grades (Figure 1C-E). IHC staining scores indicated significantly higher PIGU expression in bladder BLCA tissues compared to normal tissues (10.65 ± 2.14 vs 5.56 ± 2.68, P < 0.0001, Figure 1F). ROC curve analysis demonstrated that PIGU protein strongly distinguished BLCA tissue from non-BLCA tissue (AUC = 0.9070, P < 0.0001, Figure 1G). Additionally, PIGU protein expression was higher in male patients than in female patients (P = 0.012), while no significant differences were observed in other clinical and pathological indicators (Table 1).
| Parameters | Group | Number | mean ± SD | Z | P value |
| Tissue | Tumor | 121 | 10.65 ± 2.14 | -7.858 | P < 0.001a |
| Non-tumor | 32 | 5.56 ± 2.68 | |||
| Gender | Male | 102 | 10.85 ± 1.99 | -2.506 | P = 0.012b |
| Female | 16 | 9.31 ± 2.70 | |||
| Age | < 65 | 60 | 10.58 ± 2.23 | -0.227 | 0.821 |
| ≥ 65 | 58 | 10.71 ± 2.09 | |||
| T stage | T1-T2 | 97 | 10.55 ± 2.24 | -0.958 | 0.338 |
| T3-T4 | 21 | 11.10 ± 1.67 | |||
| N stage | N0 | 111 | 10.59 ± 2.18 | -0.980 | 0.327 |
| N1-N3 | 7 | 11.43 ± 1.51 | |||
| M stage | M0 | 118 | 10.64 ± 2.15 | - | - |
| M1 | 0 | - | |||
| Pathological grade | I-II | 62 | 10.48 ± 2.16 | -1.363 | 0.173 |
| III-IV | 50 | 11.00 ± 1.95 |
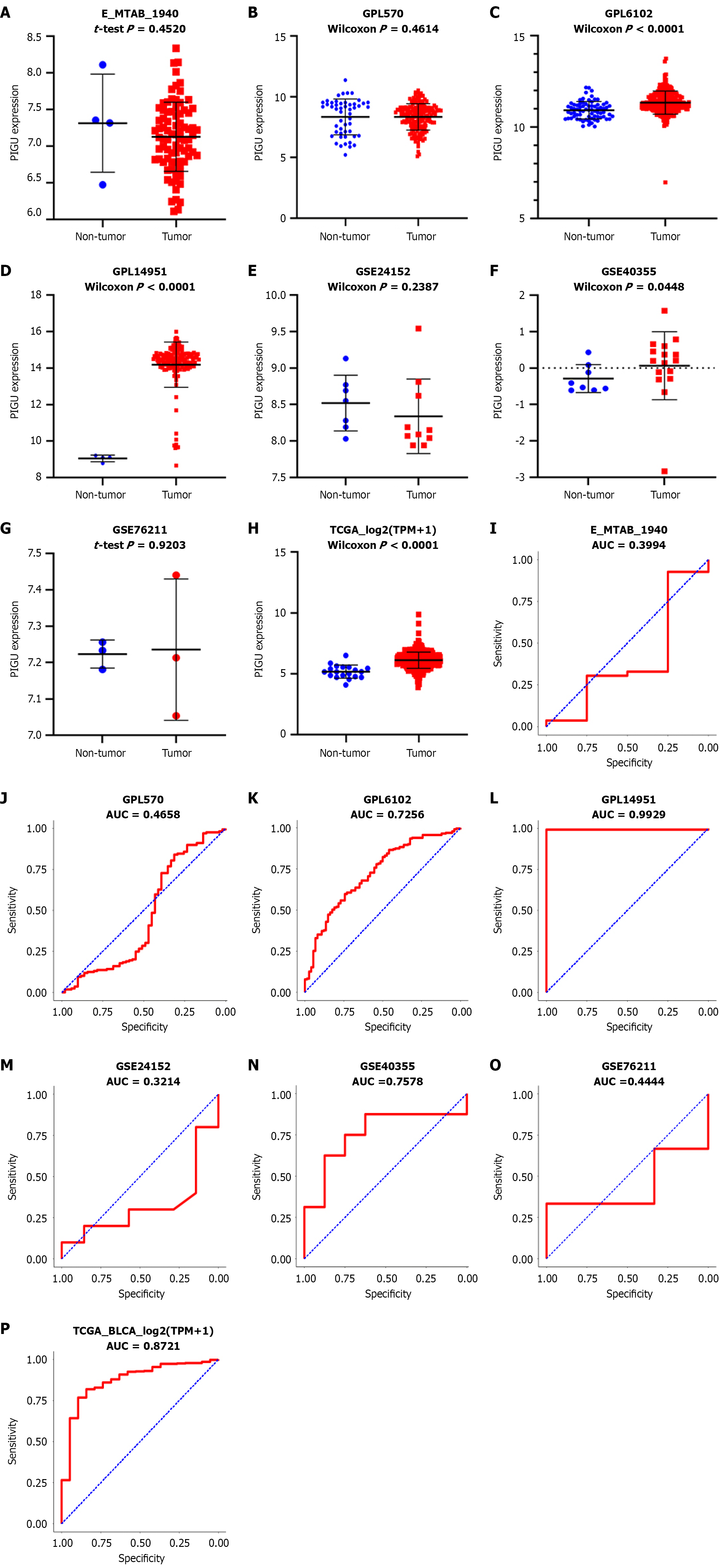
Among the existing BLCA expression datasets (Table 2), GPL6102 (GSE19423, GSE37815, GSE13507), GPL14951 (GSE65635, GSE86411), GSE40355, and TCGA. In these four datasets, PIGU mRNA expression was significantly higher in BLCA tissues than in non-BLCA tissues (P < 0.05). The remaining four datasets-E_MTAB_1940, GPL570 (GSE7476, GSE31684, GSE2109, GSE31189), GSE24152, and GSE76211 (Figure 2A-H) showed no statistically significant differences. ROC curves for each dataset are also presented, with AUC values exceeding 0.7 in four cohorts (Figure 2I-P). Detailed data parameters are shown in Supplementary Figure 1.
| Platform | Study ID | Year | Country | BLCA | Non-BLCA | ||
| n | mean ± SD | n | mean ± SD | ||||
| ArrayExpress | E-MTAB-1940 | 2015 | France | 82 | 7.129 ± 0.471 | 4 | 7.315 ± 0.669 |
| GPL570 | GSE7476 | 2007 | Spain | 169 | 8.348 ± 1.087 | 51 | 8.343 ± 1.480 |
| GSE31684 | 2011 | United States | |||||
| GSE2109 | 2005 | United States | |||||
| GSE31189 | 2013 | United States | |||||
| GPL6102 | GSE19423 | 2010 | South Korea | 231 | 11.35 ± 0.631 | 74 | 10.93 ± 0.475 |
| GSE37815 | 2013 | South Korea | |||||
| GSE13507 | 2010 | South Korea | |||||
| GPL14951 | GSE65635 | 2015 | Russia | 140 | 14.19 ± 1.239 | 4 | 9.053 ± 0.185 |
| GSE86411 | 2016 | United States | |||||
| GPL6791 | GSE24152 | 2010 | United States | 10 | 8.339 ± 0.510 | 7 | 8.520 ± 0.381 |
| GPL13497 | GSE40355 | 2013 | Germany | 16 | 0.066 ± 0.933 | 8 | 0.385 ± 0.045 |
| GPL17586 | GSE76211 | 2017 | China | 3 | 7.235 ± 0.194 | 3 | 7.223 ± 0.038 |
| TCGA | TCGA_BLCA | 2021 | United States | 414 | 6.115 ± 0.673 | 19 | 5.176 ± 0.540 |
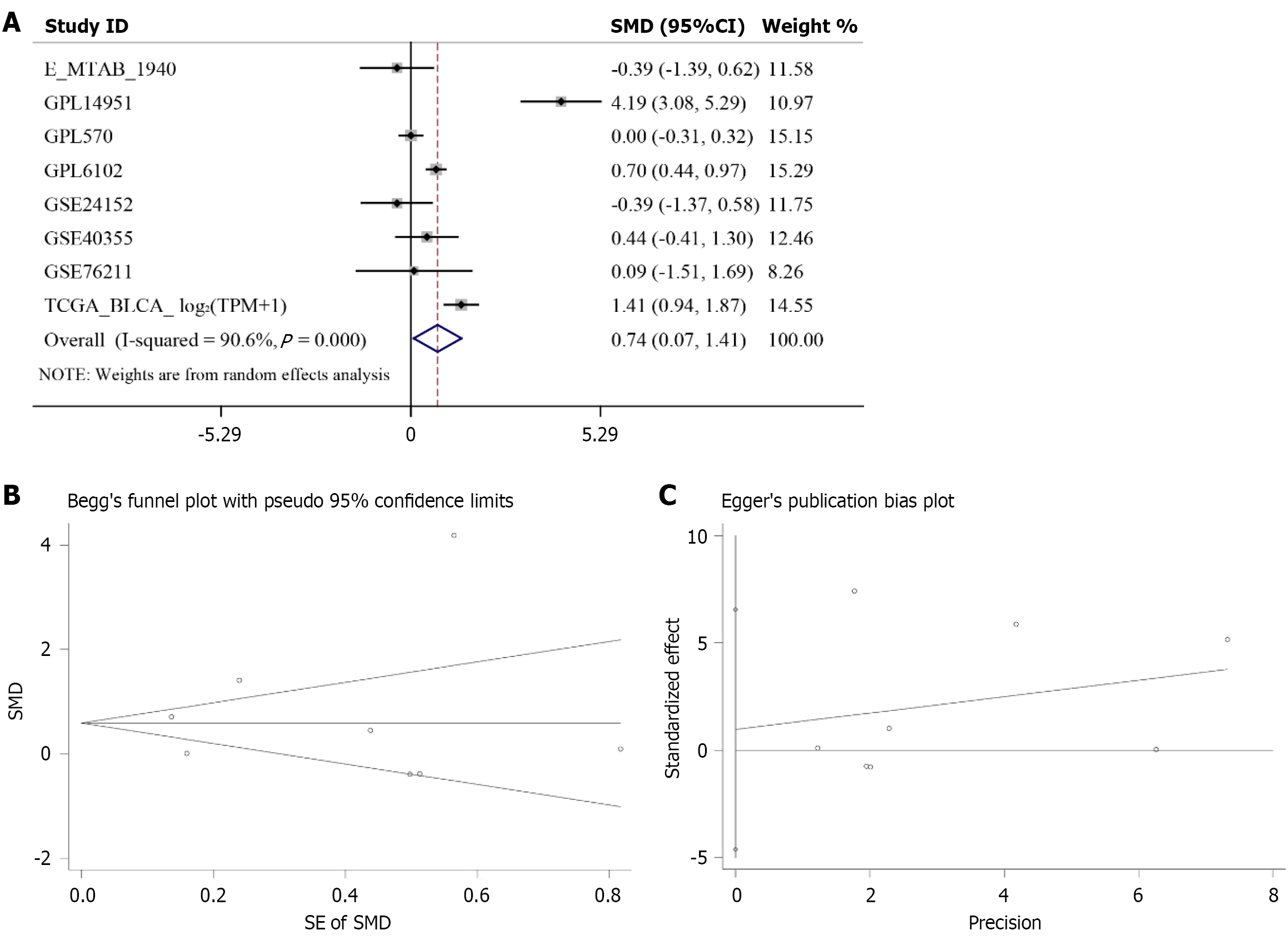
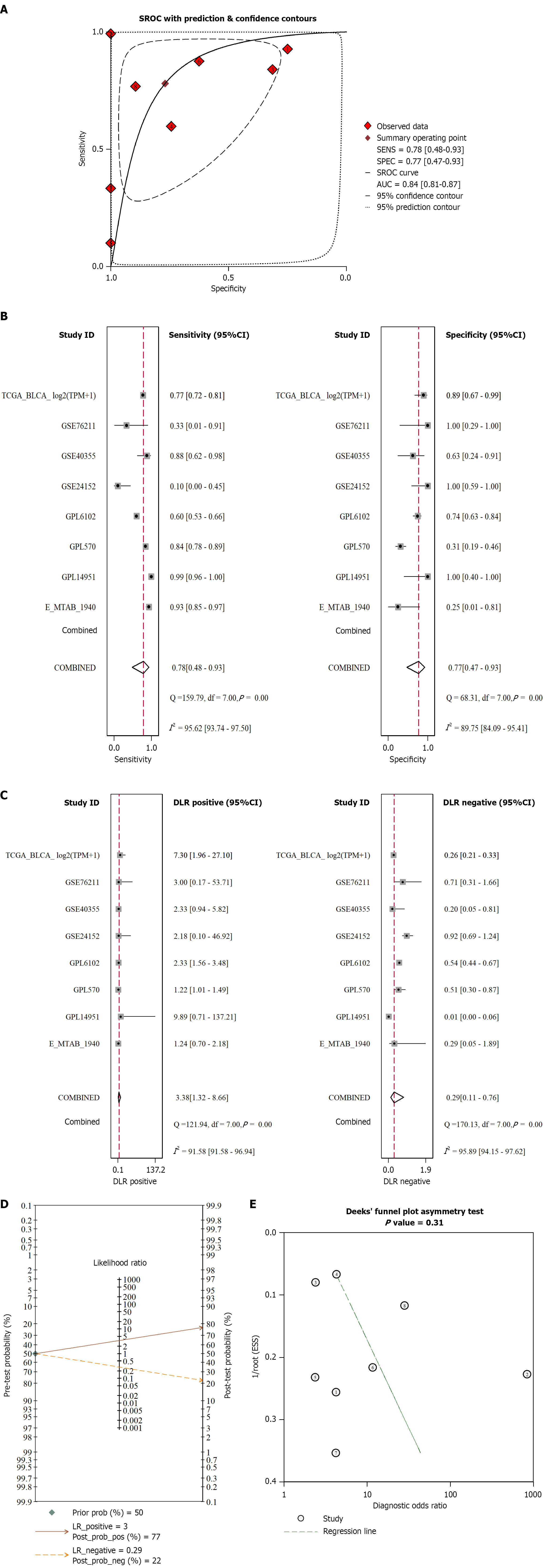
To enhance the reliability of the results, we pooled all included datasets by calculating the SMD values for PIGU expression levels. Given the high heterogeneity (I² = 90.6%, P < 0.001), a random-effects model was selected. Results indicate elevated mRNA levels of PIGU in BLCA tissues (pooled SMD = 0.74, 95%CI: 0.07-1.41, P = 0.0294; Figure 3A). Neither the Begg test (P = 0.902, Figure 3B) nor the Egger test (P = 0.686, Figure 3C) indicated publication bias, suggesting no significant evidence of asymmetry in the included studies. All included data were pooled and an SROC curve was plotted to enhance the reliability of the results. The summary ROC curve analysis demonstrated higher PIGU mRNA expression levels in BLCA tissues compared to non-BLCA tissues. The AUC was 0.84 (95%CI: 0.81-0.87, Figure 4A). Overall sensitivity was 0.78, specificity was 0.77 (Figure 4B), positive likelihood ratio was 3.38, and negative likelihood ratio was 0.29 (Figure 4C), indicating higher accuracy for detecting high PIGU expression in BLCA tissues. This is reflected in the good post-test probability shown by the Fagan plot (Figure 4D). The Deek funnel plot showed no publication bias (P = 0.31, Figure 4E).
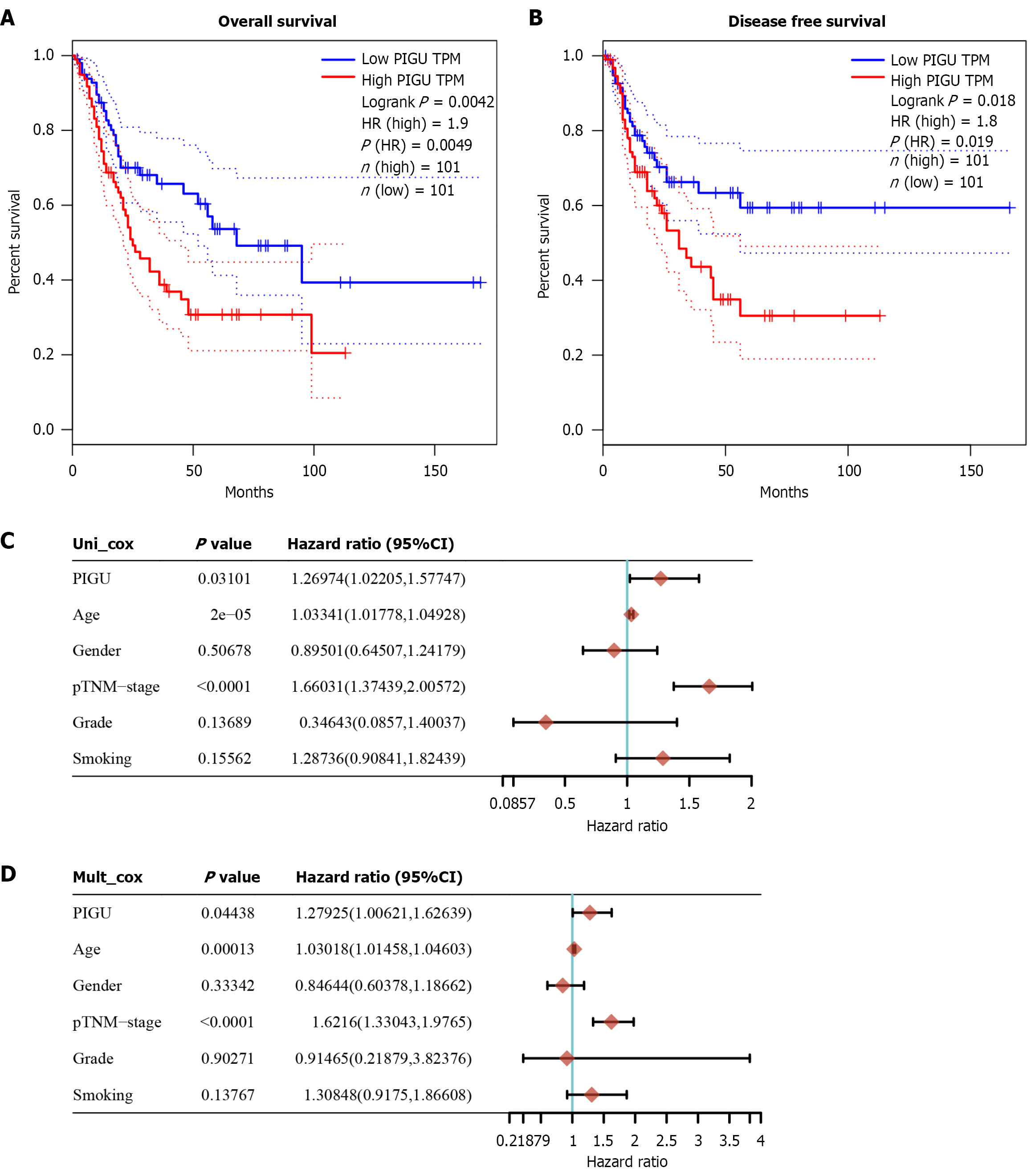
We analyzed the relationship between PIGU expression and clinical-pathological parameters in 406 BLCA patients from the TCGA database. χ2 tests (Supplementary Table 1) and nonparametric rank-sum tests (Supplementary Figure 2) revealed no significant correlation between PIGU mRNA levels and age, sex, survival status, TNM stage, pathological grade, or histological grade. We further investigated the prognostic significance of PIGU expression in BLCA. Survival analysis revealed that high PIGU expression was associated with poorer patient outcomes. The DFS and OS in the high-expression group were significantly shorter than those in the low-expression group: In the OS analysis, the log-rank P value was 0.0042 with an HR of 1.9 (Figure 5A); in the DFS analysis, the log-rank P value was 0.018 with an HR of 1.8 (Figure 5B). Cox regression analysis further demonstrated (Figure 5C and D) that both TNM staging and PIGU mRNA levels were significant independent prognostic factors for BLCA, with higher PIGU expression correlating with poorer prognosis.
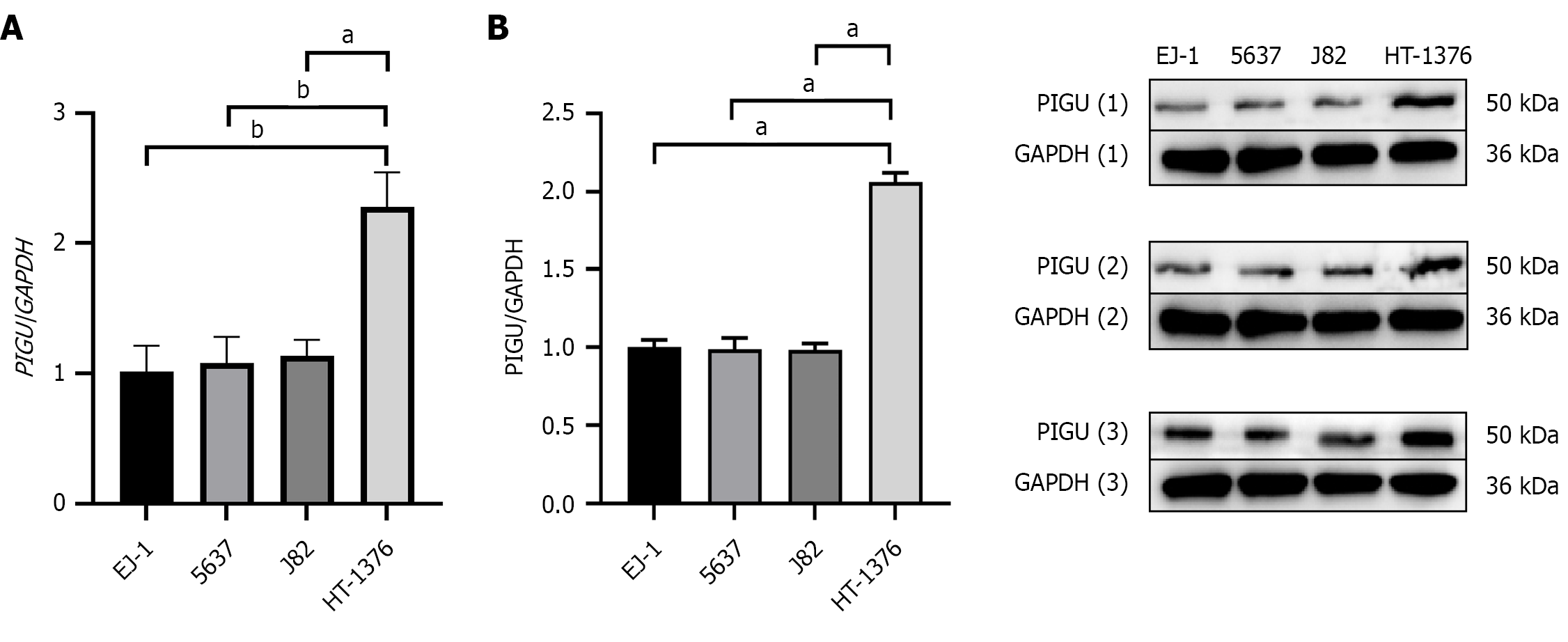
To validate the findings from previous experimental analyses, this study conducted RT-PCR experiments on four BLCA cell lines: EJ-1, 5637, J82, and HT-1376. Results showed that PIGU gene expression in HT-1376 cells was significantly higher than in EJ-1, 5637, and J82 cells (Figure 6A). Further Western blot analysis confirmed that PIGU protein expression was significantly higher in HT-1376 cells than in EJ-1, 5637, and J82 cells (Figure 6B). This indicates that the PIGU gene is highly expressed in BLCA cells, with HT-1376 showing the highest expression in the four BLCA cell lines studied. Consequently, HT-1376 cells were selected for subsequent in vitro experiments.
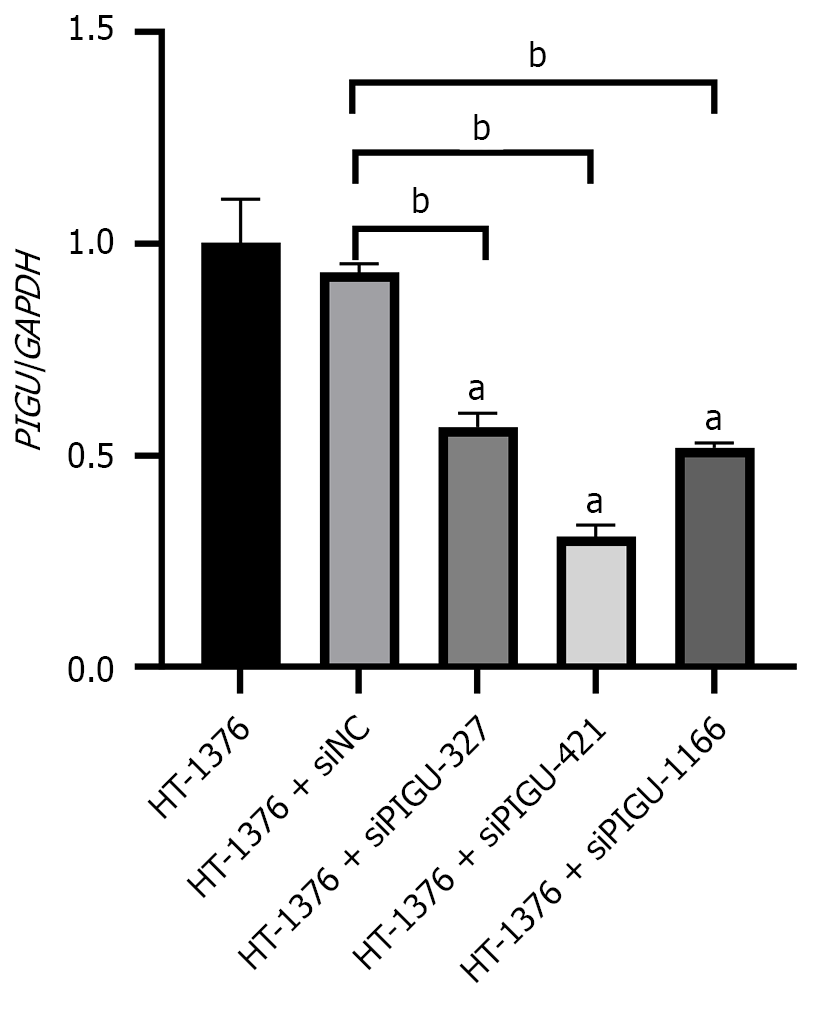
To investigate the role of PIGU in the development and progression of BLCA, this study performed cell transfection to silence PIGU in the HT-1376 cell line. A control group was established using HT-1376 cells transfected with empty lentivirus. The efficiency of silencing PIGU in BLCA cells was assessed using qRT-PCR to detect different knockdown target sequences. Quantitative results showed that compared to the HT-1376 group, no significant changes in PIGU gene expression were observed after transfecting with NC. However, after transfecting with PIGU silencing vectors, PIGU gene expression exhibited varying degrees of downregulation, with the most pronounced effect observed for PIGU-421 silencing (Figure 7). Therefore, this sequence was selected for subsequent experiments.
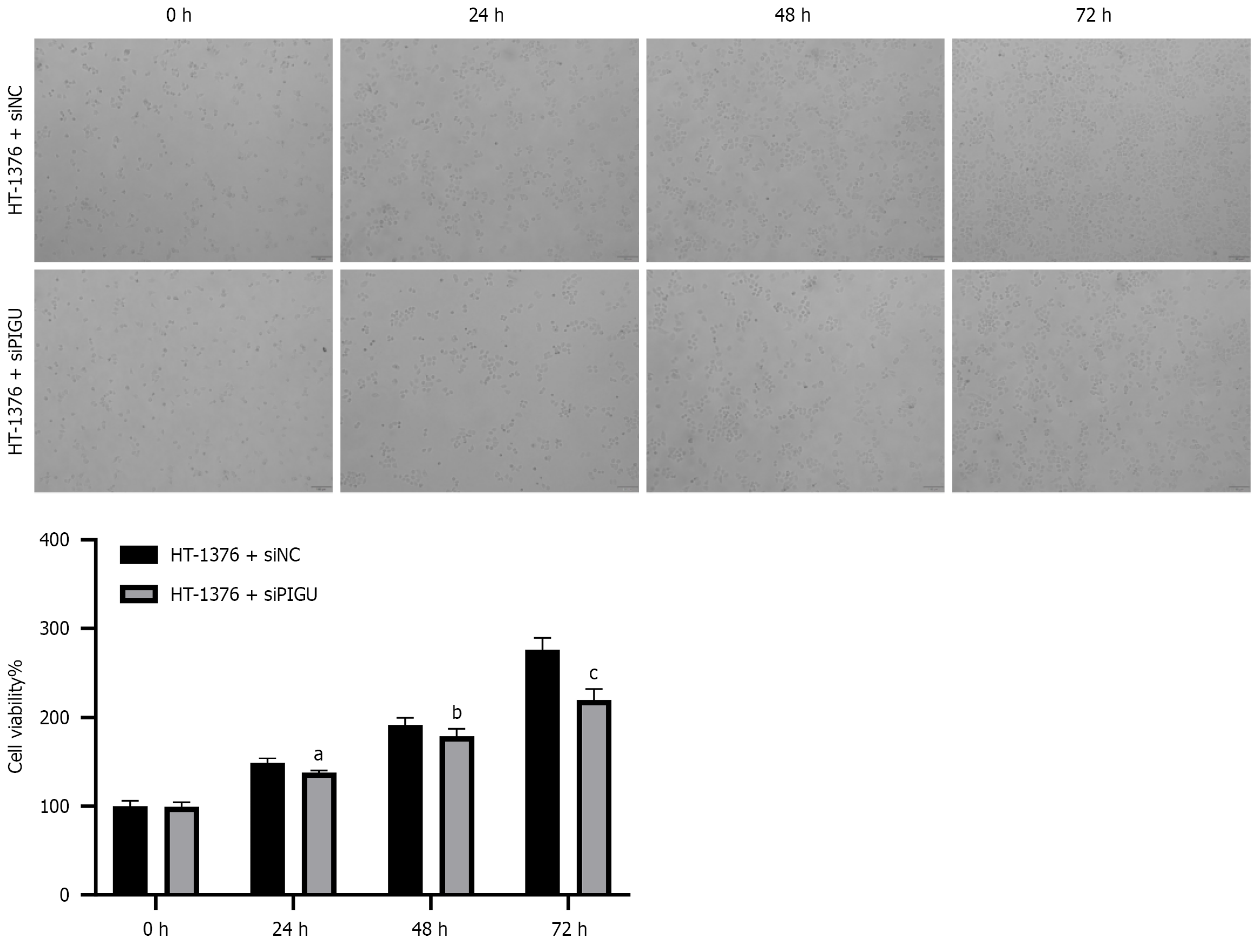
CCK-8 assay revealed that compared to the HT-1376 + si-NC group, PIGU silencing significantly reduced HT-1376 proliferation rates at the same time points, with statistically significant differences observed at 24, 48, and 72 hours (Figure 8). This indicates that PIGU can influence the proliferative capacity of HT-1376 cells. The effect on cell proliferation at 48 hours was particularly significant. Subsequent experiments were therefore conducted at the 48-hour time point.
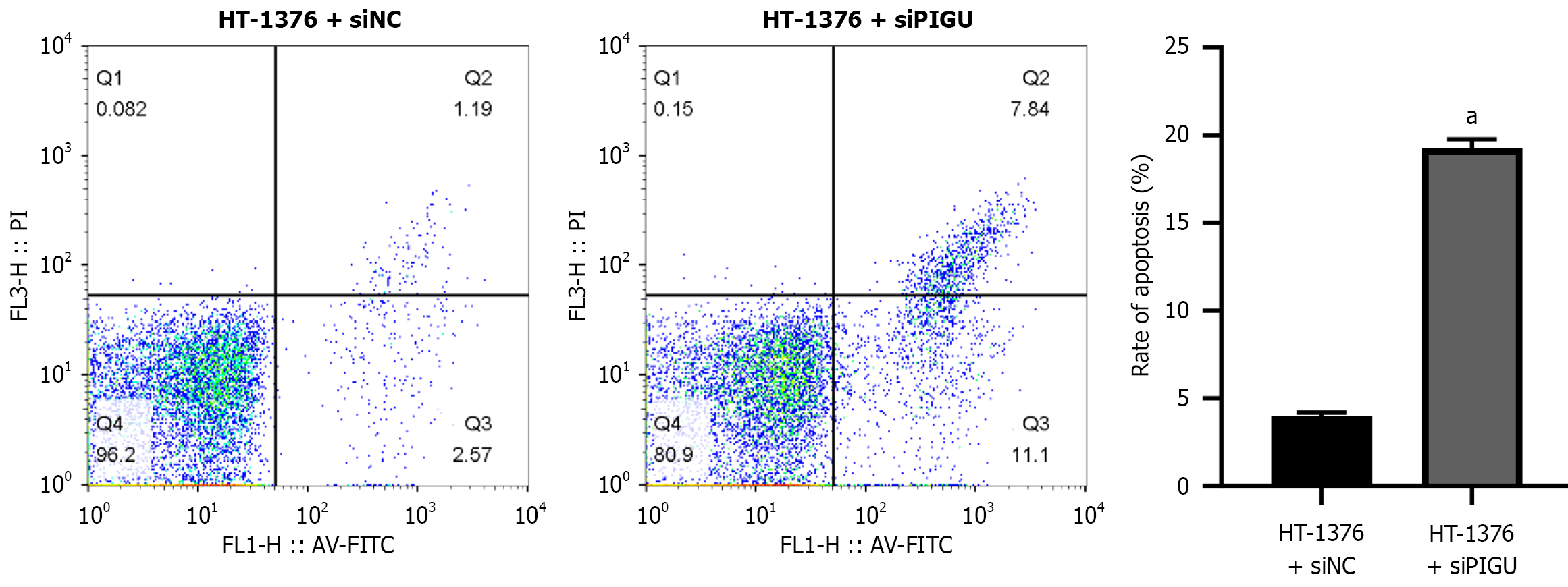
This study quantitatively analyzed the effect of silencing PIGU on BLCA cell apoptosis using flow cytometry. An experiment was set up with a PIGU-silenced group (HT-1376 + siPIGU) and a negative control group (HT-1376 + siNC). FITC/PI double staining was used to distinguish cell populations at different stages of apoptosis. Results revealed significantly increased proportions of early-stage apoptotic cells (Q3) and late-stage apoptotic cells (Q2) in the PIGU-silenced group compared to the control group (Figure 9), indicating markedly elevated apoptosis rates in HT-1376 cells following PIGU silencing (Figure 9). This finding suggests that PIGU silencing may affect cell survival by enhancing apoptosis in BLCA cells.
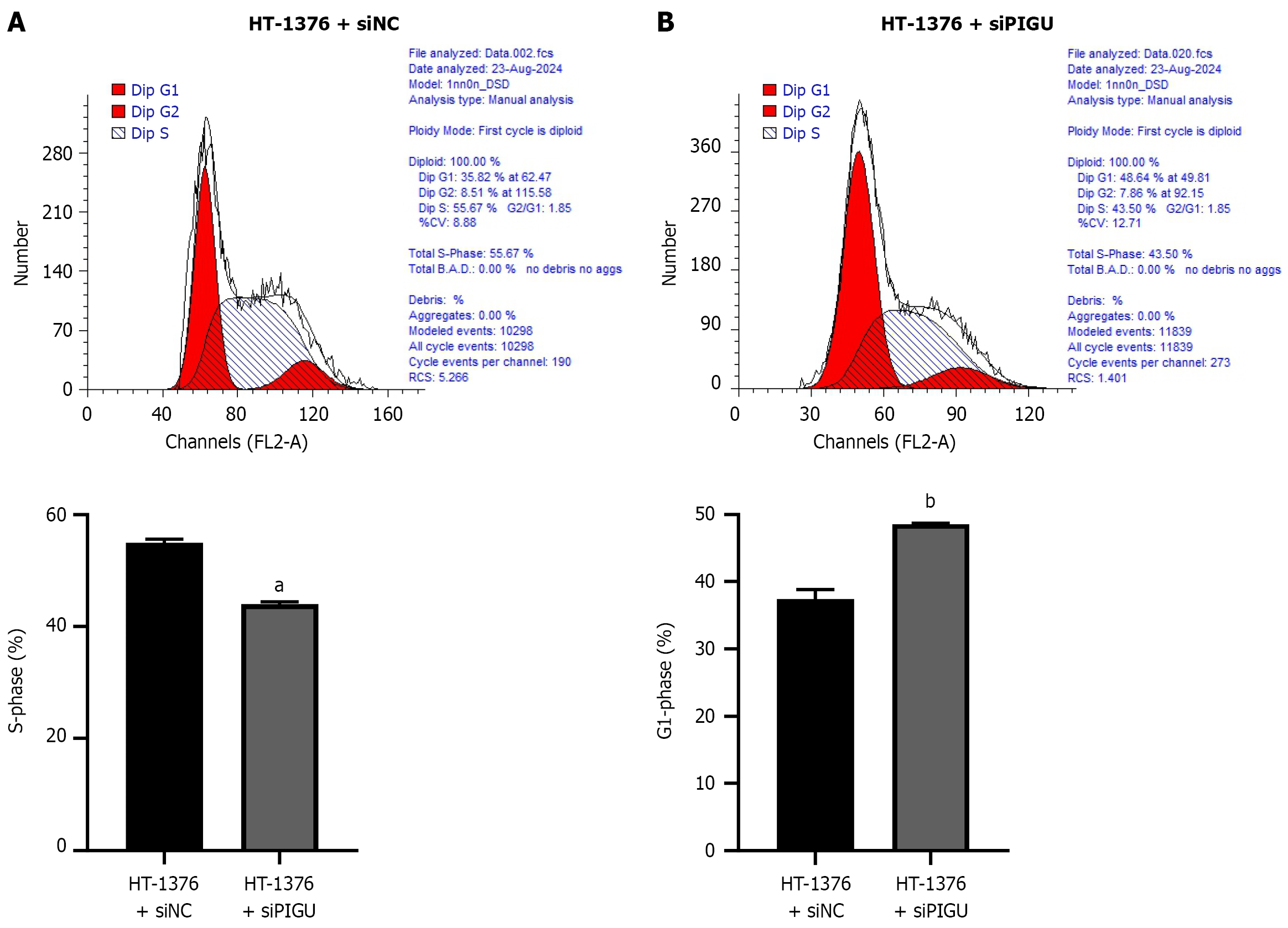
This study further employed flow cytometry to analyze the impact of PIGU silencing on the cell cycle distribution of BLCA cells. In the HT-1376 cell line, the cell cycle characteristics of the PIGU-silenced group (HT-1376 + siPIGU) were compared with those of the empty vector control group (HT-1376 + siNC). Results showed that compared to the HT-1376 + siNC group, the siPIGU group exhibited a reduced proportion of S-phase cells and a significantly increased proportion of G1-phase cells (Figure 10). This indicates that PIGU knockdown weakens the proliferative capacity of BLCA cells, leading to cell arrest at the G1 phase.
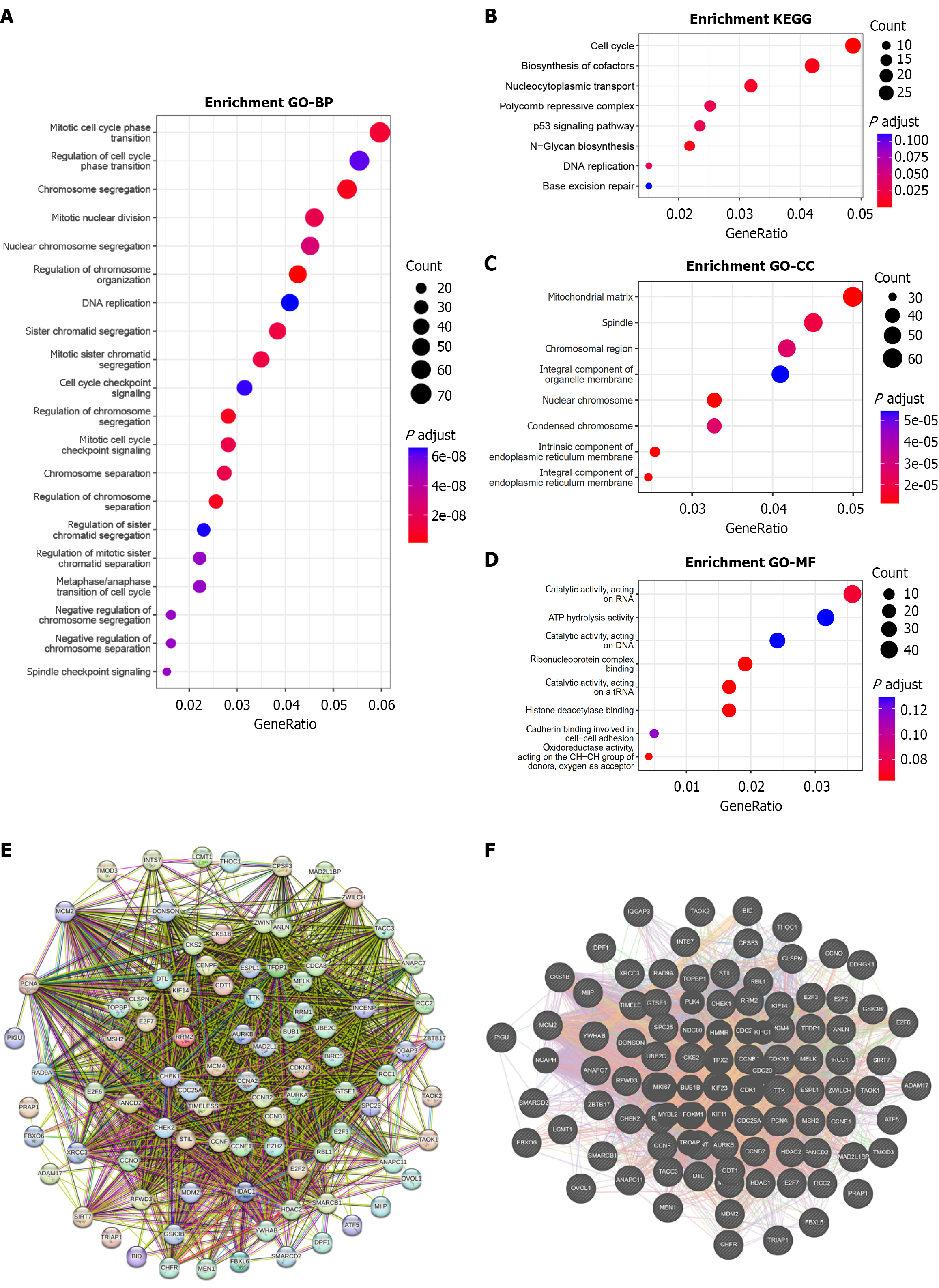
Based on the DCEG screening criteria, a total of 1271 DCEGs were identified (1250 upregulated and 21 downregulated). GO and KEGG enrichment analyses revealed that these genes are primarily involved in mitotic nuclear division, chromosome segregation, and other cell cycle-related biological processes (Figure 11A). KEGG pathway analysis further showed that the cell cycle pathway exhibited the most significant enrichment (Figure 11B). Moreover, enriched molecular functions and cellular components were also closely related to the cell cycle, including nuclear chromosomes, chromosomal regions, and catalytic activity acting on DNA (Figure 11C and D). Therefore, further investigation of these genes participating in cell cycle pathways is warranted (Supplementary Table 2). After removing duplicates, 90 unique genes were retained. PPI network analysis revealed that PIGU interacts with these genes (Figure 11E and F).
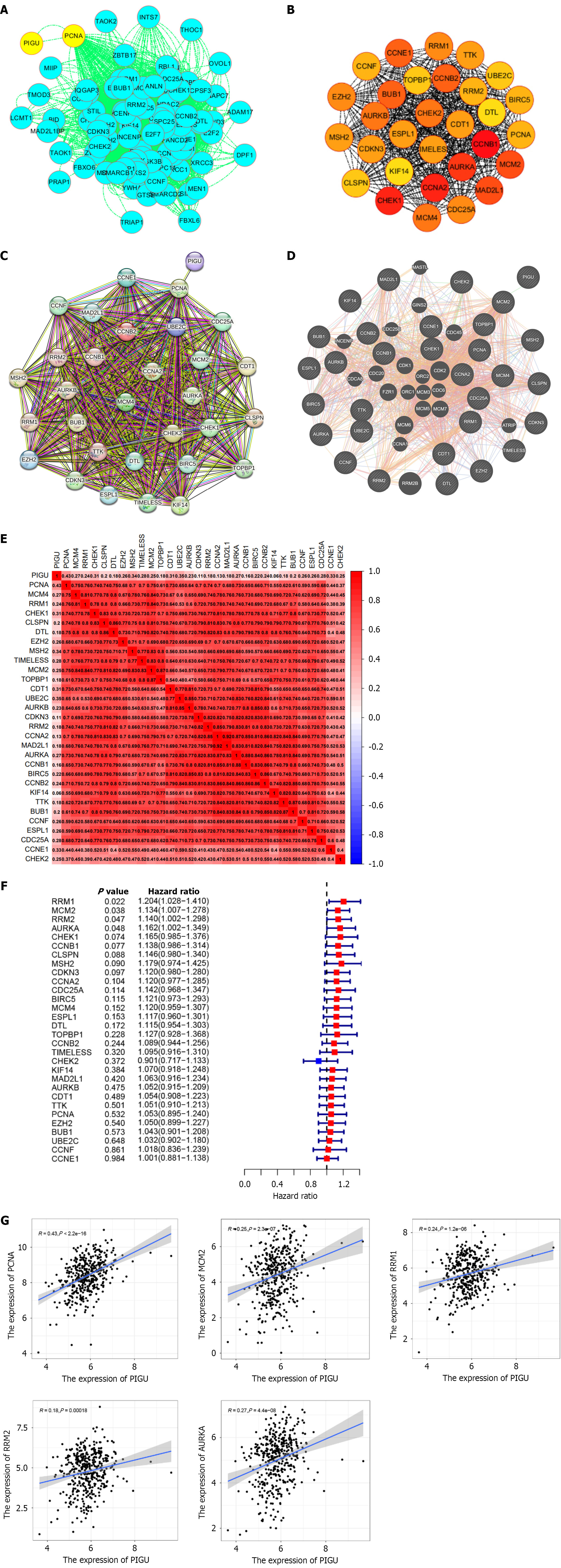
This study further visualized genes interacting with PIGU using Cytoscape (Figure 12A). The cytoHubba module algorithm identified 30 hub genes with the strongest protein interaction capabilities (Figure 12B). Analysis of the PPI network for these genes revealed direct protein interactions between PIGU and PCNA, as well as direct genetic interactions with MCM2. This indicates that PIGU influences cell cycle processes by interacting with PCNA and MCM2 (Figure 12C and D). Heatmap analysis demonstrated significant positive correlations between these hub genes and PIGU expression (Figure 12E). Cox regression analysis identified all 29 genes except CHEK2 as risk factors for BLCA, with RRM1, MCM2, RRM2, and AURKA exhibiting P values < 0.05 (Figure 12F). Further analysis demonstrated significant positive correlations between PIGU and PCNA, MCM2, RRM1, RRM2, and AURKA, with PCNA exhibiting the highest correlation coefficient (Figure 12G).
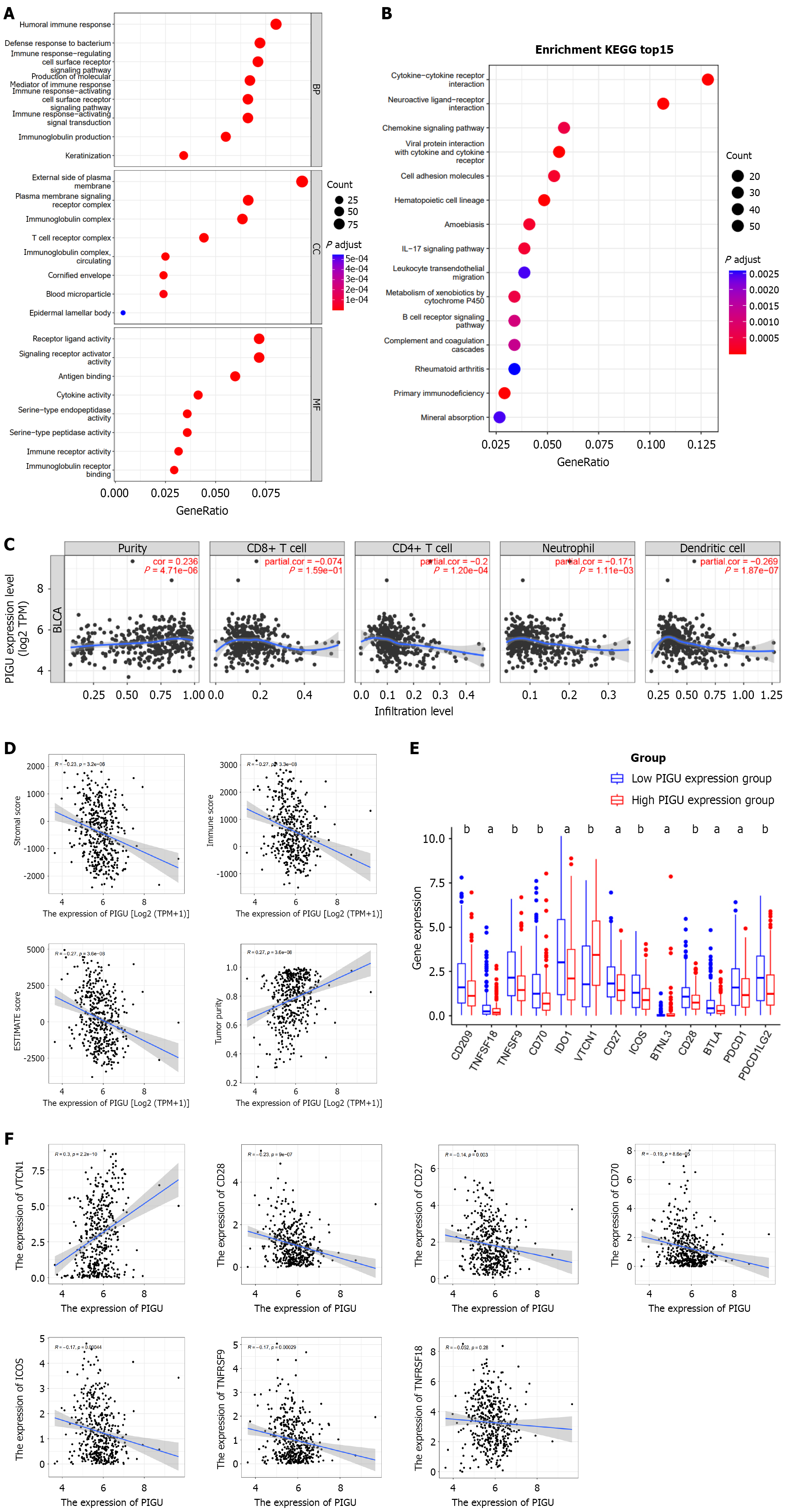
Based on PIGU expression levels, we divided BLCA patients into high-expression and low-expression groups and performed GO and KEGG enrichment analyses of DEGs. Results revealed that DEGs associated with immune-related pathways were significantly enriched in both groups (Figure 13A and B), suggesting PIGU expression may also correlate with tumor immune infiltration. Analysis via the TIMER platform indicated that PIGU expression positively correlated with tumor purity while negatively correlating with CD8+ T cells, CD4+ T cells, neutrophils, and dendritic cells (Figure 13C). Analysis using ESTIMATE technology to calculate tumor immune grading and tumor purity (Figure 13D) revealed strong negative correlations between PIGU expression and the stromal score (r = -0.23, P < 0.001), immunoscore (r = -0.27, P < 0.001), and ESTIMATE score (r = -0.27, P < 0.001). This was further supported by enrichment of immune-related pathways in DEGs and negative correlations with immune cell populations. These findings indicate that PIGU expression negatively correlates with tumor immune infiltration while positively correlating with tumor purity. To explore the mechanism by which PIGU influences tumor immune infiltration, we identified 13 PIGU-regulated immune checkpoint genes by intersecting 79 immune checkpoint genes with DEGs from different PIGU expression groups and compared their expression levels[14]. We found significantly reduced expression of CD28, ICOS, TNFRSF9, CD70, CD27, and TNFRSF18 in the high PIGU expression group, all of which are associated with immune activation (Table 3)[13], while VTCN1, associated with immune suppression, showed significantly increased expression (Figure 13E). Corresponding scatter plots also revealed consistent trends: PIGU showed strong negative correlations with CD28, CD27, CD70, ICOS, TNFRSF9, and TNFRSF18 expression, and a positive correlation with VTCN1 expression (Figure 13F).
| Symbol | Type | Role with immunity |
| VTCN1 | Ligand | Inhibit |
| CD28 | Receptor | Activate |
| ICOS | Receptor | Activate |
| TNFRSF9 | Receptor | Activate |
| CD70 | Ligand | Activate |
| CD27 | Receptor | Activate |
| TNFRSF18 | Receptor | Activate |
| CD209 | Ligand | Two side |
| BTNL3 | Ligand | Two side |
| PDCD1 LG2 | Ligand | Two side |
| IDO1 | Ligand | Inhibit |
| PDCD1 | Receptor | Inhibit |
| BTLA | Receptor | Inhibit |
PIGU, one of the five subunits of the GPI-T complex, contributes to the maturation of GPI, which anchors numerous proteins to the cell surface[15]. Although PIGU has been investigated in several cancer types, research on its role in BLCA has remained limited in recent years; thus, additional evidence is necessary to clarify its oncogenic potential and underlying molecular mechanisms. In this study, we integrated bioinformatics analyses with in vitro cellular experiments to comprehensively examine the involvement of PIGU in BLCA development and progression. We first assessed PIGU protein expression in BLCA by performing IHC on 121 BLCA tissue samples and 32 non-BLCA tissue samples. High-throughput datasets and tissue microarray analyses consistently demonstrated elevated PIGU expression at the protein level. To strengthen the robustness of our findings, we conducted a meta-analysis using a random-effects model combined with SMD to integrate multiple BLCA datasets comprising 1065 BLCA cases and 170 non-BLCA cases. This approach enabled the effective aggregation of large-scale transcriptomic data, confirming significant upregulation of PIGU mRNA in BLCA. These results were further validated through in vitro experiments using BLCA cell lines, which indicated that PIGU may promote tumor progression by modulating cell cycle-related pathways and suppressing immune-related pathways.
When combined with multicenter, integrated, high-throughput datasets, IHC analyses strengthened our findings and further confirmed the upregulation of PIGU in BLCA at both the mRNA and protein levels. To enhance the reliability of these results, we performed RT-PCR on four BLCA cell lines and observed significantly elevated PIGU expression in the HT-1376 cell line, corroborating our previous analyses. This observation is consistent with an earlier study by Guo et al[5], who first identified PIGU as a potential oncogene in BLCA and reported high PIGU expression in four BLCA cell lines: SW-780, HTB-1, HTB-4, and HTB-9. However, subsequent studies investigating PIGU expression in BLCA have yielded inconsistent results. For instance, Schultz et al[6] assessed PIGU expression using RT-qPCR in eight BLCA cell lines, 42 BLCA tissue samples, and 14 non-BLCA tissues but found no evidence of PIGU overexpression. This is in stark contrast to the findings of Guo et al[5]. Conversely, another study integrating RT-qPCR and IHC analyses (73 BLCA and 14 non-BLCA tissues) reported elevated PIGU expression at both the mRNA and protein levels[7], supporting Guo et al’s conclusions[5]. Nevertheless, these studies share several limitations, including restricted sample size and heterogeneity, reliance on limited detection methods, and a lack of functional validation. Additionally, none of these studies explored PIGU’s underlying molecular mechanisms. Our study addresses these limitations by integrating multiple high-throughput datasets with IHC data and performing in vitro functional experiments. This comprehensive approach provides compelling evidence for the upregulation of PIGU in BLCA.
Previous studies have demonstrated the clinical significance of PIGU in various cancers, including hepatocellular carcinoma and colorectal cancer[16,17]. To further investigate its significance in BLCA, this study assessed the clinical relevance of PIGU in BLCA. Utilizing existing datasets, we evaluated its potential diagnostic capability via ROC and SROC analyses. Our results indicate that PIGU expression at both the mRNA and protein levels is significantly altered in BLCA tissues compared to noncancerous tissues, suggesting that the expression difference of PIGU holds promise in distinguishing BLCA from non-BLCA tissues.
Furthermore, through survival analysis and univariate and multivariate Cox regression analyses, we determined that PIGU is an independent prognostic factor for BLCA and that elevated PIGU levels were associated with poorer patient outcomes. These findings jointly highlight the potential of PIGU as a prognostic marker for BLCA patients.
PIGU has been investigated in multiple cancer types. PIGU has been shown to regulate cell viability, migration, invasion, and apoptosis, potentially facilitating tumor progression in hepatocellular carcinoma by enhancing immune evasion and activating the NF-κB signaling pathway; conversely, PIGU knockdown suppresses tumor development in hepatocellular carcinoma[18]. Elevated PIGU expression has also been linked to increased motility, epithelial-mesenchymal transition, lung metastasis, and poor prognosis in breast cancer. Similarly, PIGU overexpression in thyroid cancer promotes tumor cell proliferation, migration, and invasion[4].
To explore the molecular mechanisms of PIGU in BLCA, we first silenced PIGU expression in BLCA cells. Cell proliferation assays showed that PIGU depletion significantly reduced proliferative capacity. Moreover, flow cytometry analysis indicated that PIGU knockdown induces apoptosis and causes G1-phase cell cycle arrest. These results suggest that the antiproliferative effect following PIGU silencing may stem from increased apoptosis and impaired cell cycle progression. Enrichment analysis was performed on PIGU and its DCEG to further interpret these observations. The analysis revealed strong enrichment in cell cycle-related pathways in BLCA. The cell cycle is a tightly regulated biological process consisting of four phases-G1, S, G2, and M. The dysregulation of cell cycle-associated proteins can lead to aberrant cell cycle activation, resulting in uncontrolled proliferation and tumorigenesis[19-21]. Moreover, abnormal cell cycle progression is closely linked to other cancer hallmarks, such as metabolic reprogramming and immune evasion, that further drive tumor progression[22].
Further analyses of the enriched pathways identified 30 hub genes with the strongest predicted interactions with PIGU, among which PCNA and MCM2 exhibited direct protein-protein or genetic interactions with PIGU. PCNA is a highly conserved homotrimer that plays an essential role in DNA replication by increasing the processivity of leading-strand synthesis, making it indispensable for DNA replication and cell cycle regulation[23]. Its post-translational modifications are critical for coordinating DNA synthesis and DNA damage tolerance. Moreover, PCNA dysregulation can lead to double-strand DNA breaks, potentially driving mutagenesis and cancer development[24]. MCM2, a core subunit of the eukaryotic replicative helicase complex, is essential for DNA replication initiation and elongation. MCM2 dysfunction can impair DNA replication and cell proliferation, contributing to disease development[25]. Taken together, our findings indicate that PIGU may influence BLCA development by interacting with key regulators of DNA replication, such as PCNA and MCM2, thereby modulating multiple stages of the cell cycle.
Enrichment analyses of differentially DCEGs in high- and low-PIGU expression groups similarly indicated significant enrichment of immune-related pathways in BLCA. Our study revealed a negative correlation between PIGU expression and infiltration levels of dendritic cells, CD8+ T cells, and CD4+ T cells. CD8+ T cells have been widely recognized as key effector cells in antitumor immunity and cancer immunotherapy[26]. CD4+ T cells possess cytotoxic capabilities and can directly eliminate tumor cells while also supporting the cytotoxic activity of CD8+ T cells[27,28]. Dendritic cells are professional antigen-presenting cells that play essential roles in initiating CD4+ T-cell responses and subsequently activating CD8+ T-cell-mediated antitumor effects[29].
Additionally, this study found PIGU expression to be negatively correlated with the overall abundance of immune and stromal cells in the tumor microenvironment. These findings suggest that higher PIGU expression may be associated with reduced immune cell infiltration in BLCA, providing a potential immunological explanation for its tumor-promoting role.
Immune checkpoints are cell surface molecules that deliver activating or inhibitory signals to regulate the initiation, magnitude, and duration of immune responses. In cancer, the dysregulation of these checkpoints is a key mechanism by which tumor cells evade immune surveillance and promote malignant progression[30]. In our study, VTCN1 expression was significantly elevated in the PIGU high-expression group, whereas CD28, ICOS, TNFRSF9, CD70, CD27, and TNFRSF18 levels were markedly reduced. VTCN1 is widely recognized for its immunosuppressive properties, while CD28, ICOS, TNFRSF9, CD70, CD27, and TNFRSF18 are generally associated with immune activation within the tumor microenvironment[14,31]. These observations suggest that PIGU expression may be associated with alterations in immune checkpoint signaling. Given these findings, we hypothesize that high PIGU expression may contribute to reduced immune infiltration in BLCA by promoting VTCN1-mediated immunosuppressive signaling and attenuating the activity of immune activation-related checkpoints.
Overall, we found that PIGU is overexpressed at both the protein and mRNA levels in BLCA tissues and that PIGU regulates BLCA cell proliferation by affecting PCNA and MCM2 in cell cycle-related pathways as well as apoptosis. However, the present study has several limitations. First, the number of non-BLCA tissues included in the work was insufficient compared to the number of BLCA tissues. Second, when analyzing clinical pathological parameters and prognosis, we examined only 405 patients from the TCGA database, resulting in an overly homogeneous patient population. Furthermore, functional experiments were primarily conducted at the in vitro cellular level, and only the HT-1376 cell line was employed for the main experiments, lacking functional controls from other bladder carcinoma cell lines. Another limitation of this study is the lack of in vivo functional validation. Although our conclusions are supported by integrated multi-omics analyses and in vitro functional experiments, in vivo models such as xenograft assays will be required to further substantiate the oncogenic role and therapeutic relevance of PIGU in BLCA. Such investigations will clarify PIGU’s role in BLCA and provide deeper insights into its potential as a therapeutic target.
PIGU plays an oncogenic role in BLCA and serves as a risk factor for poor patient prognosis. Silencing PIGU inhibits BLCA cell proliferation in vitro, promotes apoptosis, and causes cell cycle arrest at the G1 phase. Furthermore, PIGU may promote tumor progression by affecting cell cycle-related pathways and suppressing immune system-related pathways. These findings highlight the importance of PIGU as a potential biomarker for prognosis and treatment in BLCA.
The authors would like to express gratitude to the TCGA, GEO and ArrayExpress public databases for providing open-access research data.
| 1. | Li J, Wang Z, Wang T. Machine-learning prediction of a novel diagnostic model using mitochondria-related genes for patients with bladder cancer. Sci Rep. 2024;14:9282. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 6] [Reference Citation Analysis (0)] |
| 2. | Coelho KBCA, Wosniaki DK, Marin AM, Fabris L, Borges Dos Reis R, Aoki MN, Zanette DL. Urinary mRNA-based biomarkers for non-muscle-invasive bladder cancer: a mini-review. Front Oncol. 2024;14:1441883. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 4] [Reference Citation Analysis (0)] |
| 3. | Zhang QJ, Liu CH, Wang K, Mao XY, Dong Y, Zang MY, Zhang W, Yu QS, Hao L. Hypoxia-induced HIF-1α/VASN promotes bladder cancer progression. Sci Rep. 2025;15:21635. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 4] [Reference Citation Analysis (0)] |
| 4. | Zuo J, Guo H, Hu G, He Y, Tang Y, Li J, Cui Y, Liang S, Chen X, Luo Z, Wang X, Wang X. Silencing of PIGU inhibits the progression of esophageal squamous cell carcinoma through the PI3K/AKT signaling pathway. Sci Rep. 2025;15:31237. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 5. | Guo Z, Linn JF, Wu G, Anzick SL, Eisenberger CF, Halachmi S, Cohen Y, Fomenkov A, Hoque MO, Okami K, Steiner G, Engles JM, Osada M, Moon C, Ratovitski E, Trent JM, Meltzer PS, Westra WH, Kiemeney LA, Schoenberg MP, Sidransky D, Trink B. CDC91L1 (PIG-U) is a newly discovered oncogene in human bladder cancer. Nat Med. 2004;10:374-381. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 65] [Cited by in RCA: 76] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 6. | Schultz IJ, Kiemeney LA, Witjes JA, Schalken JA, Willems JL, Swinkels DW, de Kok JB. CDC91L1 (PIG-U) mRNA expression in urothelial cell carcinomas. Int J Cancer. 2005;116:282-284. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 7] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 7. | Shen YJ, Ye DW, Yao XD, Trink B, Zhou XY, Zhang SL, Dai B, Zhang HL, Zhu Y, Guo Z, Wu G, Nagpal J. Overexpression of CDC91L1 (PIG-U) in bladder urothelial cell carcinoma: correlation with clinical variables and prognostic significance. BJU Int. 2008;101:113-119. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 6] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 8. | Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45:W98-W102. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7704] [Cited by in RCA: 7552] [Article Influence: 839.1] [Reference Citation Analysis (6)] |
| 9. | Wu T, Hu E, Xu S, Chen M, Guo P, Dai Z, Feng T, Zhou L, Tang W, Zhan L, Fu X, Liu S, Bo X, Yu G. clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. Innovation (Camb). 2021;2:100141. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5856] [Cited by in RCA: 6433] [Article Influence: 1286.6] [Reference Citation Analysis (4)] |
| 10. | Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, Pyysalo S, Doncheva NT, Legeay M, Fang T, Bork P, Jensen LJ, von Mering C. The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021;49:D605-D612. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6825] [Cited by in RCA: 5439] [Article Influence: 1087.8] [Reference Citation Analysis (4)] |
| 11. | Warde-Farley D, Donaldson SL, Comes O, Zuberi K, Badrawi R, Chao P, Franz M, Grouios C, Kazi F, Lopes CT, Maitland A, Mostafavi S, Montojo J, Shao Q, Wright G, Bader GD, Morris Q. The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res. 2010;38:W214-W220. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3989] [Cited by in RCA: 3500] [Article Influence: 218.8] [Reference Citation Analysis (4)] |
| 12. | Li T, Fan J, Wang B, Traugh N, Chen Q, Liu JS, Li B, Liu XS. TIMER: A Web Server for Comprehensive Analysis of Tumor-Infiltrating Immune Cells. Cancer Res. 2017;77:e108-e110. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4366] [Cited by in RCA: 4367] [Article Influence: 485.2] [Reference Citation Analysis (3)] |
| 13. | Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, Treviño V, Shen H, Laird PW, Levine DA, Carter SL, Getz G, Stemke-Hale K, Mills GB, Verhaak RG. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7649] [Cited by in RCA: 7129] [Article Influence: 548.4] [Reference Citation Analysis (5)] |
| 14. | Hu FF, Liu CJ, Liu LL, Zhang Q, Guo AY. Expression profile of immune checkpoint genes and their roles in predicting immunotherapy response. Brief Bioinform. 2021;22:bbaa176. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 266] [Article Influence: 44.3] [Reference Citation Analysis (0)] |
| 15. | Zhang H, Su J, Li B, Gao Y, Liu M, He L, Xu H, Dong Y, Zhang XC, Zhao Y. Structure of human glycosylphosphatidylinositol transamidase. Nat Struct Mol Biol. 2022;29:203-209. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 25] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 16. | Cao J, Wang P, Chen J, He X. PIGU overexpression adds value to TNM staging in the prognostic stratification of patients with hepatocellular carcinoma. Hum Pathol. 2019;83:90-99. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 15] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 17. | Zhang M, Wang HZ, Li HO, Zhou YJ, Peng RY, Liu J, Zhao Q. Identification of PIGU as the Hub Gene Associated with KRAS Mutation in Colorectal Cancer by Coexpression Analysis. DNA Cell Biol. 2020;39:1639-1648. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 11] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 18. | Wei X, Yang W, Zhang F, Cheng F, Rao J, Lu L. PIGU promotes hepatocellular carcinoma progression through activating NF-κB pathway and increasing immune escape. Life Sci. 2020;260:118476. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 18] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 19. | Caglar HO, Biray Avci C. Alterations of cell cycle genes in cancer: unmasking the role of cancer stem cells. Mol Biol Rep. 2020;47:3065-3076. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 40] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 20. | Matthews HK, Bertoli C, de Bruin RAM. Cell cycle control in cancer. Nat Rev Mol Cell Biol. 2022;23:74-88. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 490] [Cited by in RCA: 1019] [Article Influence: 254.8] [Reference Citation Analysis (7)] |
| 21. | Suski JM, Braun M, Strmiska V, Sicinski P. Targeting cell-cycle machinery in cancer. Cancer Cell. 2021;39:759-778. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 488] [Cited by in RCA: 413] [Article Influence: 82.6] [Reference Citation Analysis (0)] |
| 22. | Liu J, Peng Y, Wei W. Cell cycle on the crossroad of tumorigenesis and cancer therapy. Trends Cell Biol. 2022;32:30-44. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 337] [Cited by in RCA: 303] [Article Influence: 75.8] [Reference Citation Analysis (5)] |
| 23. | Strzalka W, Ziemienowicz A. Proliferating cell nuclear antigen (PCNA): a key factor in DNA replication and cell cycle regulation. Ann Bot. 2011;107:1127-1140. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 627] [Cited by in RCA: 557] [Article Influence: 37.1] [Reference Citation Analysis (1)] |
| 24. | Zhang S, Zhou T, Wang Z, Yi F, Li C, Guo W, Xu H, Cui H, Dong X, Liu J, Song X, Cao L. Post-Translational Modifications of PCNA in Control of DNA Synthesis and DNA Damage Tolerance-the Implications in Carcinogenesis. Int J Biol Sci. 2021;17:4047-4059. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 39] [Reference Citation Analysis (0)] |
| 25. | Sun Y, Cheng Z, Liu S. MCM2 in human cancer: functions, mechanisms, and clinical significance. Mol Med. 2022;28:128. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 48] [Reference Citation Analysis (0)] |
| 26. | Raskov H, Orhan A, Christensen JP, Gögenur I. Cytotoxic CD8(+) T cells in cancer and cancer immunotherapy. Br J Cancer. 2021;124:359-367. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 168] [Cited by in RCA: 1193] [Article Influence: 198.8] [Reference Citation Analysis (0)] |
| 27. | Borst J, Ahrends T, Bąbała N, Melief CJM, Kastenmüller W. CD4(+) T cell help in cancer immunology and immunotherapy. Nat Rev Immunol. 2018;18:635-647. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1341] [Cited by in RCA: 1204] [Article Influence: 150.5] [Reference Citation Analysis (3)] |
| 28. | Oh DY, Fong L. Cytotoxic CD4(+) T cells in cancer: Expanding the immune effector toolbox. Immunity. 2021;54:2701-2711. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 402] [Cited by in RCA: 348] [Article Influence: 69.6] [Reference Citation Analysis (0)] |
| 29. | Murphy TL, Murphy KM. Dendritic cells in cancer immunology. Cell Mol Immunol. 2022;19:3-13. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 232] [Cited by in RCA: 209] [Article Influence: 52.3] [Reference Citation Analysis (0)] |
| 30. | Zhao B, Li H, Xia Y, Wang Y, Wang Y, Shi Y, Xing H, Qu T, Wang Y, Ma W. Immune checkpoint of B7-H3 in cancer: from immunology to clinical immunotherapy. J Hematol Oncol. 2022;15:153. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 168] [Cited by in RCA: 151] [Article Influence: 37.8] [Reference Citation Analysis (0)] |
| 31. | Podojil JR, Miller SD. Potential targeting of B7-H4 for the treatment of cancer. Immunol Rev. 2017;276:40-51. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 128] [Article Influence: 14.2] [Reference Citation Analysis (3)] |