Published online Mar 24, 2025. doi: 10.5306/wjco.v16.i3.94091
Revised: October 17, 2024
Accepted: November 25, 2024
Published online: March 24, 2025
Processing time: 315 Days and 23.4 Hours
Hepatocellular carcinoma (HCC) is a difficult cancer to manage due to its highly invasive and metastatic nature.
To investigate the molecular function of transmembrane channel-like 5 (TMC5) in vitro and in vivo, with the objective of identifying novel diagnosis and treatment targets for HCC.
The expression of TMC in cancer and normal tissues, along with its correlation with HCC prognosis, was analyzed using the GENT2, GEPIA database, and Human Protein Atlas. COX analysis was conducted to assess the relationship between TMC5 expression and overall survival in TCGA-LIHC patients. Further experiments were conducted to investigate the effect of TMC5 in cancer pro
Bioinformatics revealed that TMC5 expression was generally higher in tumors than in normal tissues, and its expression was associated with poorer patient survival outcomes. TMC5 expression in HCC tissues and cells was consistent with the results of the bioinformatics analysis. Suppression of TMC5 expression re
Our study provides compelling evidence that TMC5 is highly expressed in HCC and drives cancer progression through the activation of EMT-mediated invasion. TMC5 could represent a valuable molecular target for the diagnosis and treatment of HCC.
Core Tip: In this study, transmembrane channel-like 5 (TMC5), which is highly expressed in hepatocellular carcinoma (HCC) and associated with increased invasive migration ability, is proposed for the first time as a potential marker for HCC. Bioinformatics analysis and in vivo and in vitro experiments confirmed that TMC5 regulates epithelial-mesenchymal transition to promote invasive migration in HCC. TMC5 might be a valuable molecular target for HCC diagnosis and treatment.
- Citation: Li J, Wang ZY, Jin Y, Xu J, Ya YJ, Wan TQ, Li X, Wang X. Transmembrane channel-like 5 drives hepatocellular carcinoma progression by regulating epithelial-mesenchymal transition. World J Clin Oncol 2025; 16(3): 94091
- URL: https://www.wjgnet.com/2218-4333/full/v16/i3/94091.htm
- DOI: https://dx.doi.org/10.5306/wjco.v16.i3.94091
Hepatocellular carcinoma (HCC) is the most prevalent form of neoplasia, accounting for approximately 90% of primary liver cancer cases[1]. Currently, it ranks as the third leading cause of cancer-related deaths worldwide and the sixth most frequently diagnosed cancer[2]. World Health Organization projects more than 1 million deaths from liver cancer in 2023[3]. Most cases of HCC occur in patients with underlying liver disease, with risk factors including infection with hepatitis B or C viruses, excessive alcohol consumption, metabolic liver disease, and exposure to dietary toxins. HCC is a stealthy and aggressive disease, with most patients diagnosed in the mid-to-late stage, often exhibiting varying degrees of in
Epithelial-mesenchymal transition (EMT) is a critical cellular phenotype in the invasion and metastasis of HCC, playing a role in all phases of tumor progression, including tumor initiation, distant dissemination, tumor cell migration, intravascular metastasis, and malignant progression. EMT is a process in which epithelial cells acquire a mesenchymal phenotype through undergoing phenotypic and genotypic transformation[5]. During EMT, epithelial cells lose their polarity, functional adherens junctions, and detach from the basement membrane. EMT involves the reprogramming of epithelial gene expression. Snail is a prominent EMT transcription factor, which suppresses the expression of the cell surface protein E-cadherin and upregulates mesenchymal proteins, including N-cadherin and Vimentin proteins[6]. As EMT progresses, the transcription of junction proteins is repressed, which stabilizes the loss of epithelial junctions[7,8]. These EMT markers are associated with invasion, metastasis, and poor prognosis[9].
Transmembrane channel-like (TMC) 5 is a member of the TMC gene family, which comprises eight members (TMC1 to TMC8). Previous studies have demonstrated the critical involvement of the TMC family in human cancer. Research in
In this study, we systematically examined the impact of TMC5 expression on the prognosis of various human cancers. Specifically, TMC5 expression was analyzed in HCC tissues and cell lines. Through functional assays and an orthotopic HCC mouse model, we investigated the functional role of TMC5 in HCC and the underlying molecular mechanisms. Our findings reveal the mechanisms by which TMC5 influences HCC progression and may provide a promising therapeutic target for the treatment of HCC.
TMC5 expression differences between cancer and normal tissues, as well as its correlation with HCC prognosis, were analyzed using bioinformatics. TMC5 expression profiles were retrieved from the GENT2 (http://gent2.appex.kr/gent2/) and GEPIA (https://gepia2.cancer-pku.cn) databases, with GENT2 encompassing two microarray platforms, GPL570 and GPL96. Spatial expression of TMC5 protein in normal and tumor tissues was investigated using the Human Protein Atlas (HPA) database (https://www.proteinatlas.org/). The correlation between TMC5 expression and patient prognosis was analyzed using data from the TCGA-LIHC dataset. Correlations between clinical and prognostic characteristics, including gender, age, stage, and grade, and overall survival were evaluated using the log-rank test and Cox reg
Paraffin-embedded tumor and normal tissues were collected from 10 HCC patients following histological and clinical diagnosis at The First People's Hospital of Yunnan Province. All samples were used after obtaining informed consent. The ethics committee of the First People's Hospital of Yunnan Province approved all protocols according to the Declaration of Helsinki.
Human HCC cell lines with high metastatic potential, MHCC97-LM3 (BNCC359345), and low metastatic potential, MHCC97 L (BNCC337741), were sourced from BeNa Culture Collection (China), while normal liver cell line LO2 (HL-7702) was acquired from Procell Life Science & Technology Co., Ltd (China). All cells were cultured in Dulbecco's Modified Eagle Medium (DMEM; 10829, Gibco, Grand Island, NY, United States) supplemented with 10% (v/v) fetal bovine serum (FBS; 1902417, Gibco, Grand Island, NY, United States) at 37 °C under 5% CO2.
Lentiviruses expressing three short hairpin RNAs (shRNAs) and an overexpression lentivirus (OV) of TMC5, along with short hairpin negative control (shNC or vector), were designed and synthesized by GenePharma (China). The interfering vector was pGMLV-SC7 RNAi, with the following sequences: ShNC (TTCTCCGAACGTGTCACGT), shRNA1 (GCCT GTCGGAAATTCTGAATT), shRNA2 (GGACTTCACTGTCACTCATGA), shRNA3 (GCAACTGATCACAAGTC TTGG). The overexpression vector was anti-CMV-H_TMC5-PGK-Blasticidin. MHCC97-LM3 cells were infected with shNC and TMC5-shRNAs, while vector and OV-TMC5 were transfected into MHCC97 L cells using Lipofectamine 2000 (11668027, Invitrogen, Carlsbad, CA, United States). To establish cell lines with stable expression, cells (1 × 105 cells/well) were seeded into a 12-well plate and transfected with shRNAs and overexpression lentivirus at a multiplicity of infection of approximately 10. Following a 48-hour transfection period, the efficiency of the fluorescently labeled lentiviral infection was assessed using fluorescence microscopy. When the infection efficiency reached 80%, puromycin was then used to select for stably transfected cells. Western blot and qPCR analysis were performed to confirm the efficiency of knockdown and overexpression.
Approximately 500 cells per well were plated onto 6-well plates. After a culture period of 2 weeks, the formed colonies were fixed in 4% paraformaldehyde for 30 minutes and subsequently stained with 0.1% crystal violet for 10 minutes, then washed with phosphate buffered saline (PBS). The colonies were counted using Image J software.
In the transwell invasion assays, the transwell chambers were coated with 50 μL of Matrigel and incubated at 37 °C for 30 minutes. Cells (1 × 105 cells/mL) per well were incubated in 500 μL of serum-free DMEM and seeded a 24-well transwell chamber with an 8 μm pore size (Corning, China), and 500 µL of medium containing 10% FBS was added to the lower chamber. After incubation for 48 hours at 37 °C, the cells that migrated through the lower chamber were fixed in 4% paraformaldehyde and stained with 0.5% crystal violet. Subsequently, cells were observed under light microscopy and counted using Image J software.
In the wound-healing assays, cells (2.5 × 104 cells/mL) per well were seeded into ibidi culture-inserts, with both the left and right wells incubated for 24 hours. When the cells had reached 95% confluence, the ibidi culture-insert was removed. After washing with PBS, the cells were cultured for 24 hours in DMEM supplemented with varying concentrations of FBS (1%, 5%, and 10%). Cell movement toward the wound was recorded, and wound area measurements were performed using Image J software.
Total RNA from cells was isolated with TriReagent (15596026, Invitrogen, Carlsbad, CA), and reverse-transcribed into cDNA using FastKing RT Kit (With gDNase) from Tiangen Biotech (China). The PCR reaction for the TMC5 gene was performed using Taq Pro Universal SYBR qPCR Master Mix (Vazyme Biotech Co., Ltd, China) under the following PCR cycles: 95 °C for 10 minutes, followed by 40 cycles of 95 °C for 15 seconds, 60 °C for 30 seconds, and 72 °C for 1 minute. Amplification of GAPDH served as an internal control. The target primers were as follows: TMC5 forward: AATAACTGGTCTGAGGAA, TMC5 reverse: CTGGAACATCTGGATAAC, GAPDH forward: TTGCCCTCAACGACCACTTT, GAPDH reverse: TGGTCCAGGGGTCTTACTCC. The relative mRNA expression level was calculated using the 2−ΔΔCt method.
Total protein samples were extracted from cells using RIPA buffer (Beyotime, China), and 60 μg of protein per well were separated by 10% SDS-PAGE gels and electroblotted onto a PVDF membrane. The membranes were then blocked with 5% nonfat dry milk and treated with primary antibodies: anti-TMC5 (1:2000; PA5-95696, Invitrogen, Carlsbad, CA, United States), anti-snail (1:2000; PA5-85493, Invitrogen, Carlsbad, CA, United States), anti-E-cadherin (1:1000; MA5-12547, Invitrogen, Carlsbad, CA, United States), anti-N-Cadherin (1:2000; PA5-19486, Invitrogen, Carlsbad, CA, United States), anti-Zo-1 (1:1000; PA5-28858, Invitrogen, Carlsbad, CA, United States), and anti-β-actin (1:4000; TA-09, ZSGB-bio, China) overnight at 4 °C. The membranes were washed three times with TBST and subsequently incubated with HRP-labeled secondary antibodies at room temperature for 2 hours. Protein signal was detected using a multi-electroluminescence detection system (Tanon Science & Technology, China) with β-actin as the loading control proteins.
Female BALB/C-(nu/nu) nude mice aged 5-6 weeks and weighting 20-25 g were obtained from Kunming Medical University. All mice were treated in accordance with the guidelines for the care and use of laboratory animals of the National Institutes of Health. The ethics committee of Kunming Medical University approved all protocols according to the ARRIVE. Mice were anesthetized and fixed on a sterilized experiment board. Following sterilization with 70% alcohol, a transverse incision measuring 2 mm in depth and 1 to 1.5 cm in length was made below the xiphoid, perpendicular to the midline. The left lobe of the liver was carefully excised from the peritoneal cavity using a sterile cotton swab. MHCC97 L cells (1 × 106 cells in 0.1 mL PBS) transfected with LV-OV-TMC5, LV-vector, LV-shNC, or LV-sh-TMC5 were injected into the left liver lobe of each mouse.
After three weeks, cancer progression was imaged in the mice by intraperitoneal injection of 10 μL/g of D-fluorescein. The mice were anesthetized with an intraperitoneal injection of 10 μL/g of pentobarbital solution. After imaging, the experimental animals were sacrificed using an overdose of 2% sodium pentobarbital (0.5 mL). Animals were dissected with medical scissors and tweezers to observe the lungs, liver, and other organs, which were excised for the assessment of biological changes and subsequent immunohistochemistry (IHC).
For the IHC staining, liver tissue samples were embedded in paraffin and sectioned into 5-μm-thick sections. These sections were baked at 60 °C for 30 minutes, deparaffinized in xylene, and rehydrated through a graded ethanol series to distilled water. The sections were then placed in citrate buffer for antigen retrieval and subjected to microwave for 20 minutes. The sections were treated with 3% H2O2 in methanol to quench endogenous peroxidase activity. Then, 5% sheep serum albumin was applied to block non-specific binding. Anti-E-cadherin antibody (1:200; ab231303, Abcam, Cam
Statistical analysis was conducted using GraphPad version 8.0. Data were expressed as the mean ± SE from at least three independent experiments. t-test was used to compare means between two samples, and ANOVA was used to compare means between multiple groups. Differences were considered statistically significant at P < 0.05.
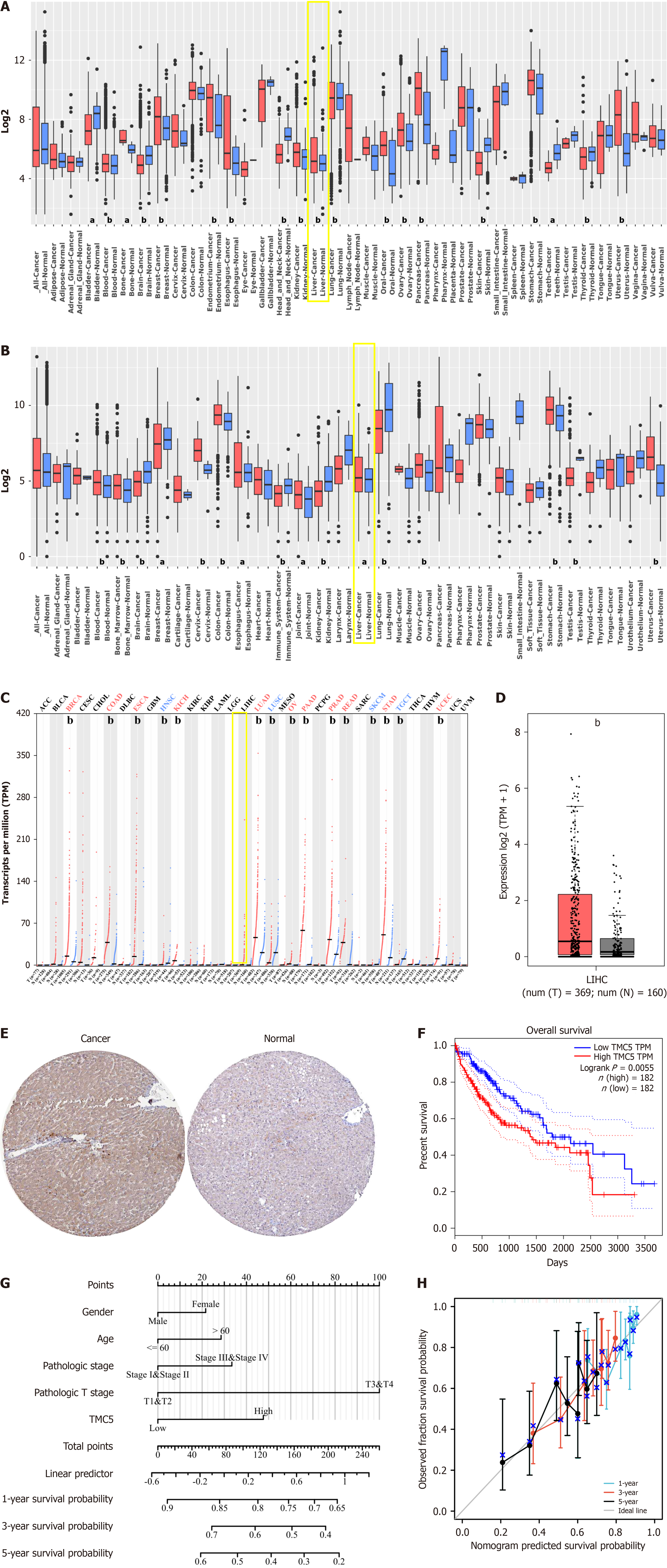
TMC5 expression was analyzed across various human cancers and their corresponding normal tissues using GENT2 and GEPIA databases. The results indicated that TMC5 was expressed in multiple malignancies, including breast, esophageal, ovarian, pancreas, and lung cancers (Figure 1A-C). Specifically, there was a significant upregulation of TMC5 mRNA in HCC tissues compared to normal tissues (Figure 1D). Furthermore, IHC staining from HPA databases revealed higher TMC5 protein levels in HCC compared to normal liver tissues (Figure 1E).
To further assess the clinical significance of TMC5, we performed a correlation analysis between TMC5 expression and patient prognosis. Higher TMC5 expression was progressively correlated with advanced tumor stage and T stage in HCC (Table 1). Both the log-rank and Cox univariate analyses demonstrated that elevated TMC5 expression was associated with a worse prognosis (Figure 1F and Table 2). To better understand the clinical relevance of TMC5, a prognostic no
| Characteristics | Low expression of TMC5 (n = 187) | High expression of TMC5 (n = 187) | P value |
| Gender | 0.036 | ||
| Female | 51 (13.64) | 70 (18.72) | |
| Male | 136 (36.36) | 117 (31.28) | |
| Age | 0.499 | ||
| ≤ 60 | 92 (24.60) | 85 (22.73) | |
| > 60 | 95 (25.40) | 101 (27.01) | |
| Stage | 0.029 | ||
| I | 99 (26.47) | 74 (19.79) | |
| II | 40 (10.70) | 12.57 (12.57) | |
| III | 41 (10.96) | 44 (11.76) | |
| IV | 0 | 5 (1.34) | |
| T stage | 0.023 | ||
| T1 | 101 (27.01) | 82 (34) | |
| T2 | 42 (11.23) | 53 (16.3) | |
| T3 | 39 (10.43) | 41 (10.96) | |
| T4 | 2 (0.53) | 11 (2.94) | |
| N stage | 1.000 | ||
| N0 | 134 (35.83) | 120 (32.09) | |
| N1 | 2 (0.27) | 2 (0.27) | |
| M stage | 0.120 | ||
| M0 | 139 (37.17) | 129 (34.49) | |
| M1 | 0 | 4 (1.07) | |
| Characteristics | Total (n) | Univariate analysis | Multivariate analysis | ||
| HR (95%CI) | P value | HR (95%CI) | P value | ||
| Gender | 373 | ||||
| Female | 121 | 0.79 (0.56-1.13) | 0.200 | ||
| Male | 252 | ||||
| Age | 373 | ||||
| ≤ 60 | 177 | 1.21 (0.85-1.71) | 0.295 | ||
| > 60 | 196 | ||||
| Stage | 349 | ||||
| I | 173 | 1.42 (0.868-2.312) | 0.164 | 0.994 | |
| II | 86 | ||||
| III | 85 | 2.734 (1.792-4.172) | < 0.001 | 4.803 (0.641-35.997) | 0.127 |
| IV | 5 | 5.597 (1.726-18.148) | 0.004 | 0.999 | |
| T stage | 370 | ||||
| T1 | 183 | 1.431 (0.902-2.268) | 0.128 | 0.994 | |
| T2 | 94 | ||||
| T3 | 80 | 2.674 (1.761-4.060) | < 0.001 | 0.673 (0.091-4.997) | 0.699 |
| T4 | 13 | 5.386 (2.690-10.784) | < 0.001 | 1.091 (0.120-9.930) | 0.938 |
| N stage | 258 | ||||
| N0 | 254 | 2.03 (0.50-8.28) | 0.324 | ||
| N1 | 4 | ||||
| M stage | 272 | ||||
| M0 | 268 | 4.077 (1.281-12.973) | 0.017 | ||
| M1 | 4 | ||||
| TMC5 | 373 | ||||
| Low | 187 | 1.499 (1.059-2.122) | 0.023 | 1.210 (0.772-1.895) | 0.406 |
| High | 186 | ||||
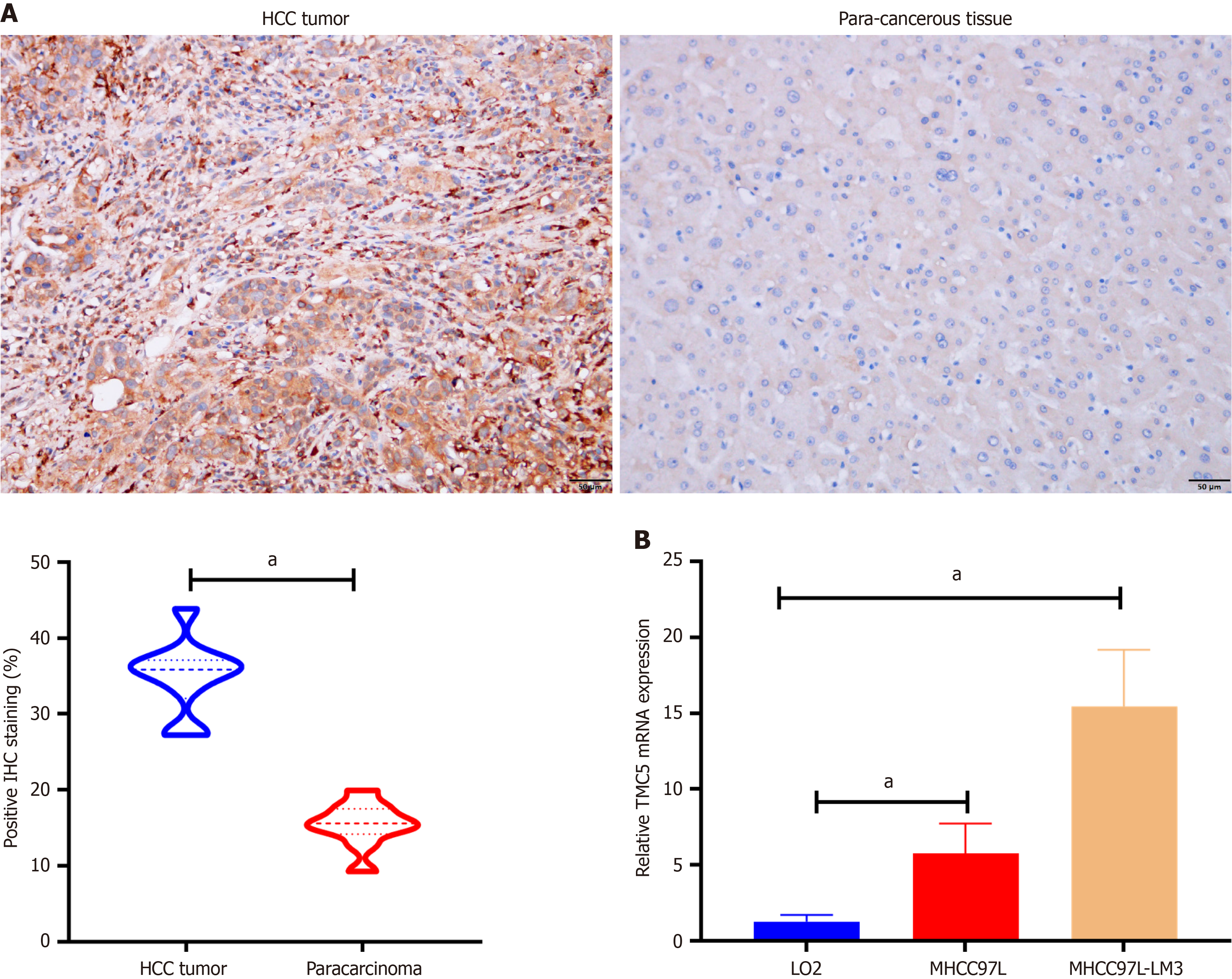
To investigate the correlation between TMC5 and HCC, we conducted IHC to examine TMC expression in 10 pairs of HCC and matched para-cancerous tissues. The results indicated that TMC5 expression was significantly stronger in tumor tissue compared to para-cancerous tissues (Figure 2A). Additionally, we evaluated TMC5 expression in HCC cell lines, finding that TMC5 expression was increased in low-metastatic HCC cell line MHCC-97H, while the high-metastatic HCC cell line MHCC97-LM3 demonstrated particularly high TMC5 expression levels (Figure 2B). These findings indicate that TMC5 may play an important role in the development of HCC.
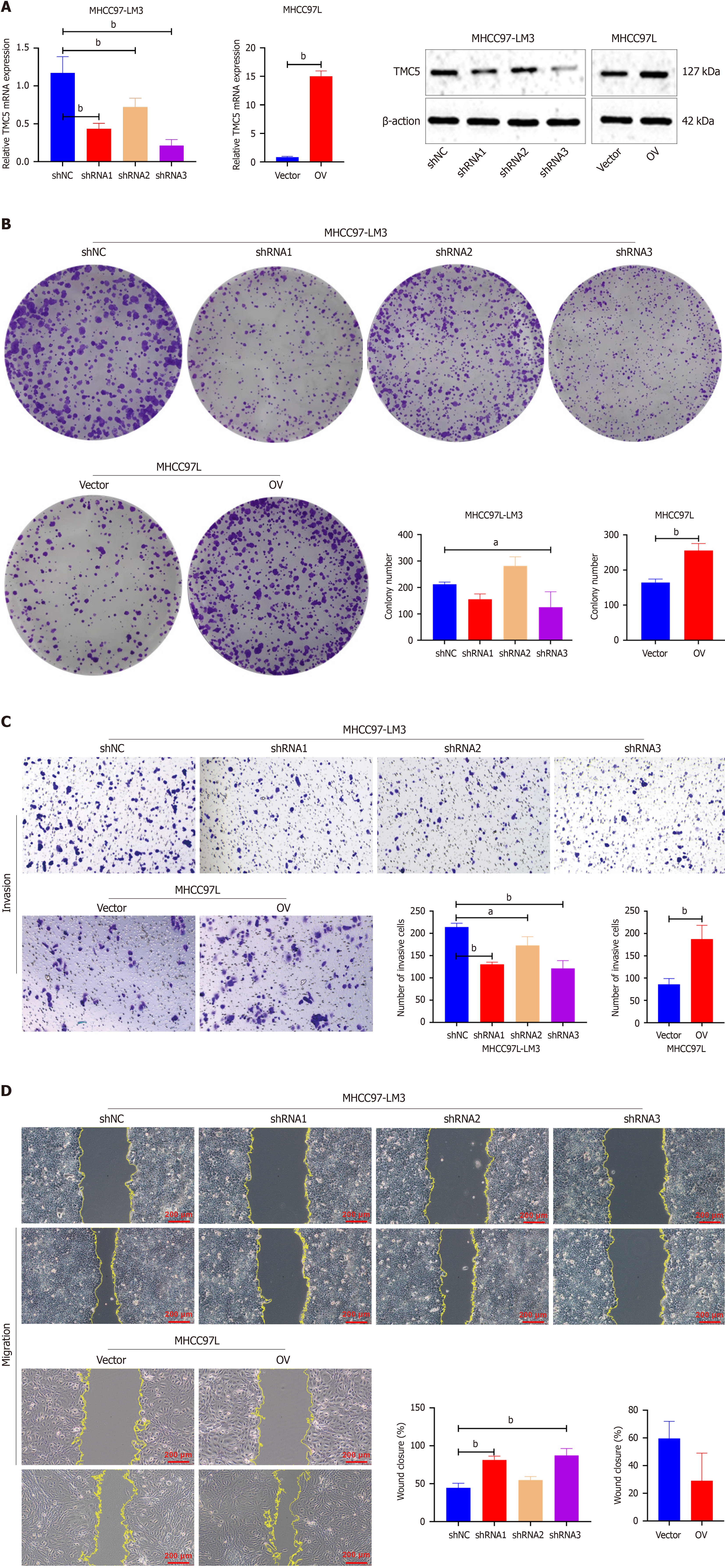
Based on TMC5 expression in HCC cells, three LV-TMC5-shRNAs were infected into MHCC97-LM3 cells to knock down TMC5, and LV-OV-TMC5 was transfected into MHCC-97H to overexpress TMC5. The efficacy of TMC5 knockdown and overexpression was confirmed by qRT-PCR and Western blot (Figure 3A). LV-TMC5-shRNA3 resulted in a significant reduction of the TMC5 expression in MHCC97-LM3 cells, while LV-TMC5-shRNA1 and LV-TMC5-shRNA2 slightly reduced TMC5 expression. In contrast, LV-OV-TMC5 induced a significant increase in TMC5 expression in MHCC-97H cells. Colony formation, transwell invasion and wound healing assays demonstrated that TMC5 expression influenced the proliferative, migratory, and invasive abilities of HCC cells. Colony formation assays showed that TMC5 depletion in MHCC97-LM3 corresponded to reduced cell proliferative, while overexpressing TMC5 in MHCC-97H noticeably increased cell proliferation (Figure 3B). In addition, transwell invasion assay and wound healing analysis revealed that TMC5 knockdown significantly weakened the capability of migration and invasion in MHCC97-LM3 as compared with that of shNC (Figure 3C). Overexpression of TMC5 in MHCC97H enhanced the migration and invasion ability of MHCC-97H as compared with the control empty vector-transfected cell (Figure 3D). Thus, TMC5 promotes proliferation, mi
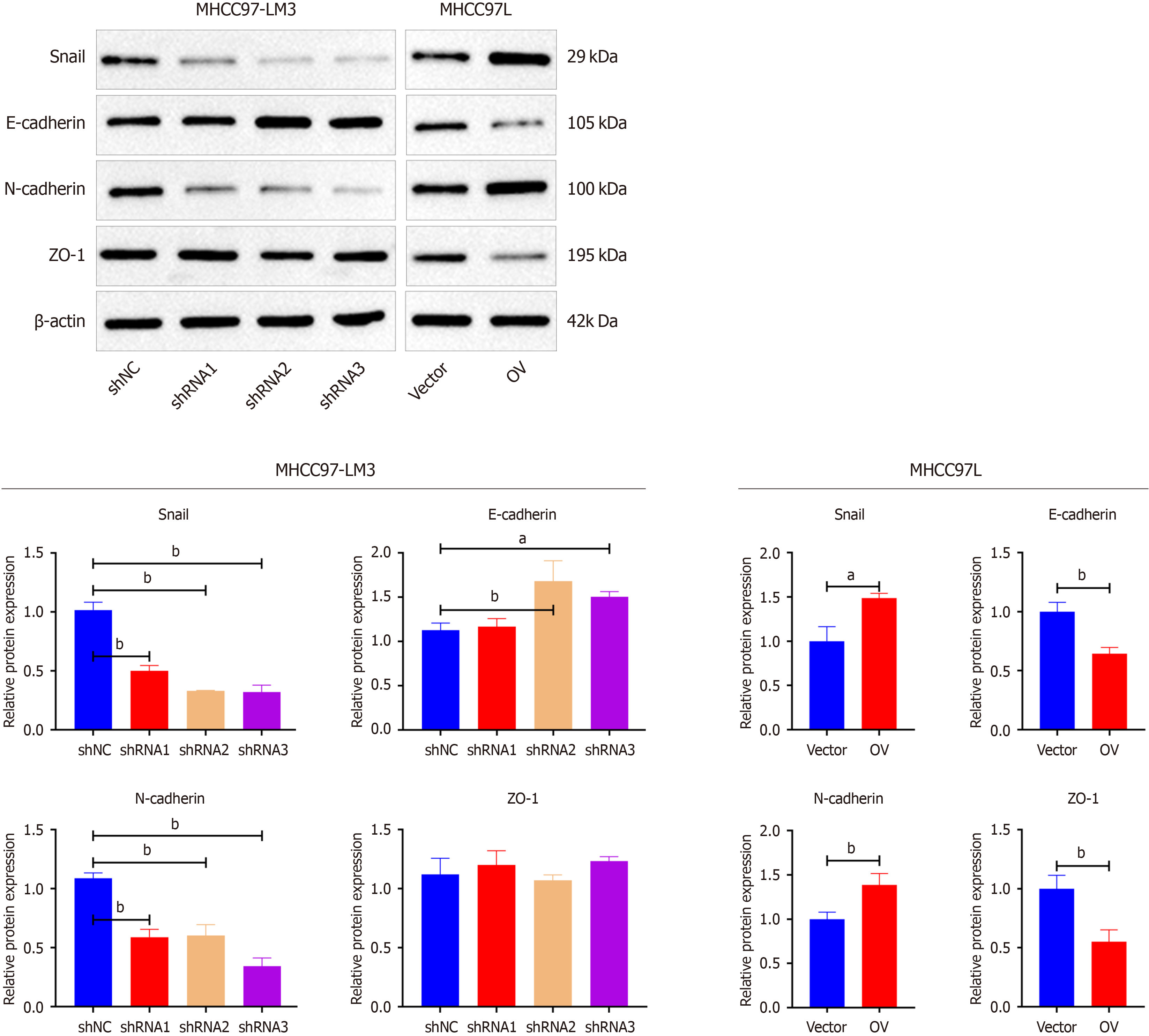
To elucidate the mechanisms by which TMC5 regulates HCC proliferation, migration and invasion, we performed western blotting analysis of protein expression. Knockdown of TMC5 in MHCC97-LM3 cell resulted in a corresponding downregulation of EMT transcription factor Snail and mesenchymal marker N-cadherin, along with an upregulation of the epithelial markers E-cadherin and ZO-1 (Figure 4). Conversely, increased expression levels of Snail and N-cadherin, along with decreased expression levels of the E-cadherin and ZO-1, were observed in MHCC-97H after upregulation of TMC5 expression (Figure 4). These results suggest that TMC5 plays a potential role in regulating EMT, and further highlight its potential as a therapeutic target for cancer treatment.
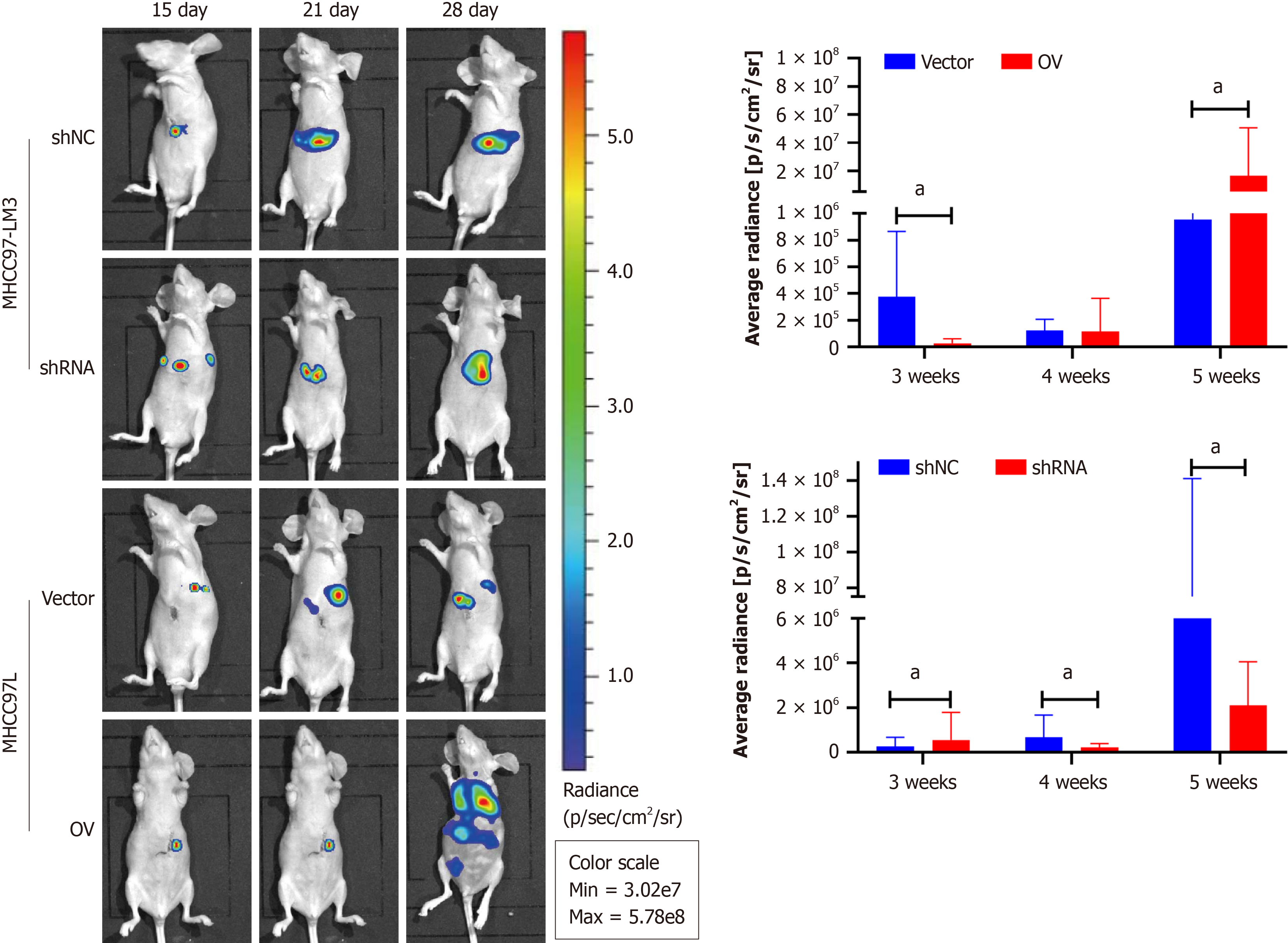
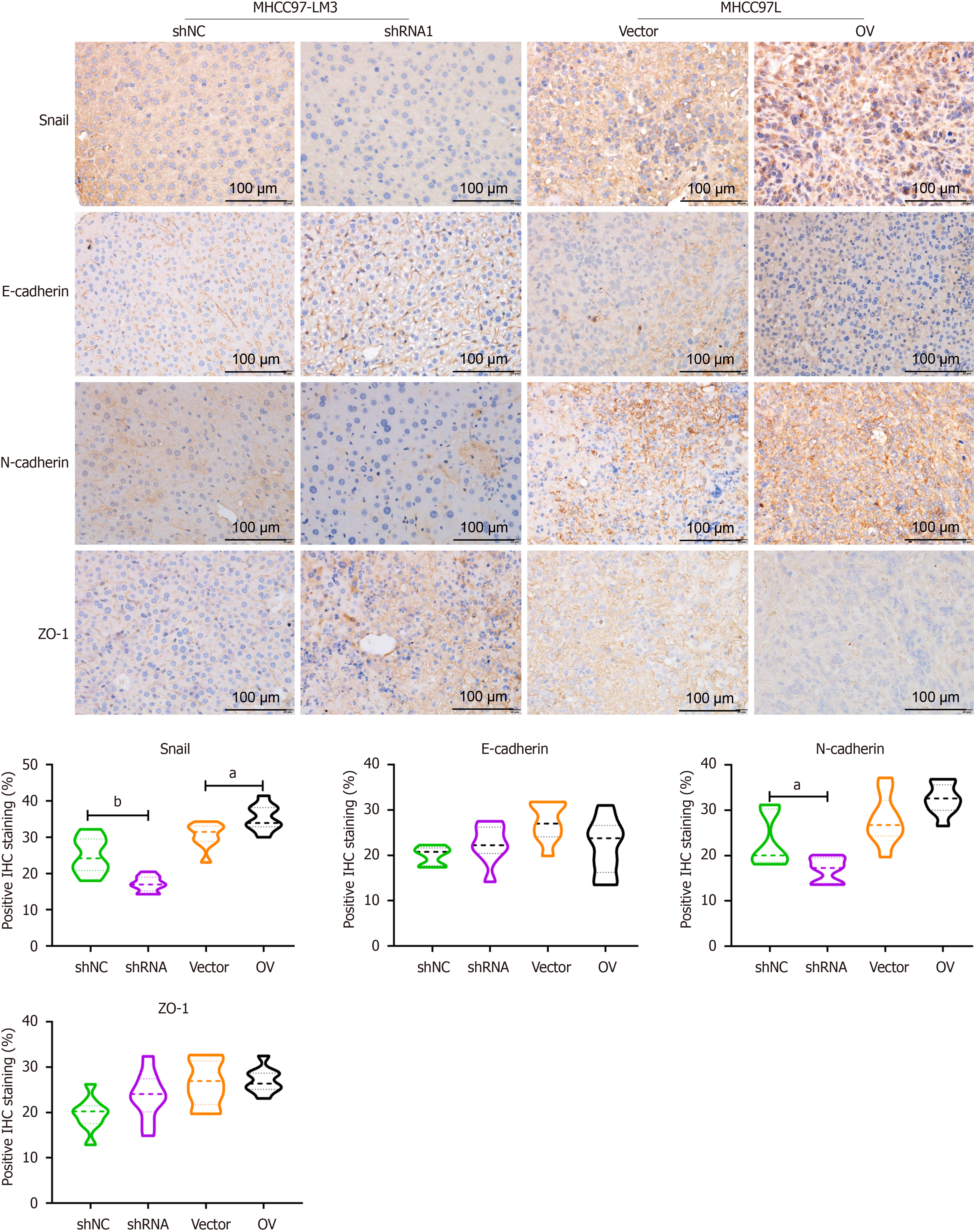
To investigate whether TMC5 promotes tumor metastasis in vivo, we injected MHCC97-LM3 cells with stable TMC5 knockdown and MHCC-97H cells with stable TMC5 overexpression into the left lobe of nude mice, using cells carrying an empty vector as a control. After two weeks, we found that, compared to hepatic tumors initiated by empty vector-expressing cells, tumors initiated by TMC5-overexpressing cells produced significantly more metastatic nodules in the lung and liver. In vivo imaging analysis confirmed the presence of both liver tumors and metastatic lung tumors in the mice (Figure 5). To further explore the correlation between TMC5 expression and EMT, IHC staining was used to identify proteins associated with EMT in liver samples of mice (Figure 6). The results indicate that liver tissue from mice injected with MHCC-97H overexpressing TMC5 exhibited higher positive staining for Snail and N-cadherin, while positive staining for E-cadherin and ZO-1 was lower compared to the vector group. Conversely, liver tissue from mice receiving MHCC97-LM3 with TMC5 knockdown showed that TMC5 inhibition reduced the positive staining area of Snail and N-cadherin, while increasing the positive staining area of E-cadherin and ZO-1. These findings suggest that TMC5 overexpression significantly enhances the development of both intrahepatic and distal pulmonary metastatic tumors.
TMC5 is observed to be differentially expressed in various cancers. Previous studies have demonstrated that TMC5 is highly expression in lung cancer[18,23], liver cancer[23,24], ICC[16,23], oral squamous cell carcinoma[19,20], and prostate cancer[17,23]. Additionally, TMC5 could serve as a sensitive and specific marker to distinguish LUAD from LUSC[19]. The role of TMC5 in PCa has been shown to PCa cell proliferation by regulating the cell cycle[17]. The expression level of TMC5 has also been correlated with a poor prognosis in LUSC, AML, and renal clear cell carcinoma[20,21,25]. Previous studies have only revealed that TMC5 is highly expressed in HCC. However, the prognostic value and function of TMC5 in HCC are unknown. In this study, HCC samples were immunohistochemically stained for TMC5, and the results showed higher TMC5 expression in HCC tissues compared to adjacent tissues, with a positive correlation between intracellular expression and metastatic potential. Notably, high expression of TMC5 predicted a possible reduction in survival in HCC patients, and the TMC5-associated nomogram model predicted HCC survival outcomes that were similar to the actual results. This suggests that TMC5 is an effective prognostic biomarker for HCC. The results of univariate COX analysis showed that TMC5 was significantly associated with the prognosis of HCC patients, but the results of multivariate COX analysis were not statistically significant. This suggests that TMC5 is not an independent prognostic factor in HCC and that its effect is masked or attenuated by other prognostic variables (age, gender, and tumor stage). Furthermore, TMC5 significantly enhanced cells proliferation, migration, invasion and metastasis in both in vitro and in vivo models. Taken together, our study demonstrates that TMC5 plays an oncogenic role in HCC.
EMT plays a crucial role in HCC tumor metastasis. EMT was first observed in mouse model of HCC and was found to result from the functional synergy between oncogenic H-Ras and TGF-β[26-29]. In a cohort of 323 patients with HCC, downregulation of E-cadherin and nuclear translocation of β-catenin, both of which are associated with microvascular invasion and metastasis, were identified[30]. Collectively, EMT plays a critical role in initiating the metastatic cascade in tumors, and its upregulation is essential for promoting metastatic growth[31]. During EMT, tumor cells lose their apical-basal polarity, leading to decreased E-cadherin expression and increased N-cadherin expression, which reduces cell adhesion and enhances motility. TMC5 has emerged as a key regulator of HCC aggressiveness and metastasis. In line with these results, TMC5 expressed at higher levels corresponding to more aggressive HCC cell. In this study, we observed that TMC5 is required to induce mesenchymal markers and suppress epithelial markers in malignant HCC cells. These findings confirm the involvement of EMT in HCC metastases and provide new insights into the role of TMC5 in HCC progression, though its precise regulatory mechanism remains unclear. These results suggest a mechanism through which TMC5 contributes to HCC cell invasion and metastasis.
Abnormal expression of the genes within the TMC protein family has been observed in several different types of cancer. TMCs are ion channels that transport various ions and are involved in development and progression of disease[32]. Mechanistic studies of TMC family members have focused exclusively on TMC1 and TMC2, which are critical for auditory conduction by contributing to mechano transduction currents in hair cells, regulating the resting membrane potential and background Na-leak conductance[33]. TMC4 acts as an anion channel, facilitating Cl- currents that help generate action potentials in taste cells in response to salt stimulation[34]. Other proteins in the same family may have regulatory role in tumorigenesis, but the exact mechanisms have yet to be determined. TMC5 may function as a multi-functional intracellular ion channel protein. TMC5 regulates cell differentiation, migration, and proliferation by acting as a transporter that imports anterior gradient proteins from cells and mediates outward-inward signal transduction events[35,36]. Mutations and altered expression of ion channels, pumps, and binding proteins in cancer cells facilitate EMT, which facilitates proliferation and malignancy[37]. Therefore, we hypothesize that TMC5 regulates EMT via mediating ion transport. However, further research is needed to elucidate its molecular mechanism in HCC development.
In summary, we reviewed the role of TMC5 in HCC, including its expression profile, prognostic value, biological functions, and regulatory mechanisms. The common overexpression of TMC5 in HCC renders it a valuable prognostic indicator, but not an independent prognostic factor. TMC5 is highly expressed in HCC and promotes the activation of EMT, contributing to pro-tumorigenic activity. As a significant oncogene, TMC5 has emerged as a potential target for therapeutic intervention strategies in HCC.
| 1. | Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87-108. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18589] [Cited by in RCA: 21017] [Article Influence: 1910.6] [Reference Citation Analysis (2)] |
| 2. | Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin. 2021;71:209-249. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76817] [Cited by in RCA: 69509] [Article Influence: 13901.8] [Reference Citation Analysis (40)] |
| 3. | Villanueva A. Hepatocellular Carcinoma. N Engl J Med. 2019;380:1450-1462. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3748] [Cited by in RCA: 3433] [Article Influence: 490.4] [Reference Citation Analysis (3)] |
| 4. | Chen Z, Xie H, Hu M, Huang T, Hu Y, Sang N, Zhao Y. Recent progress in treatment of hepatocellular carcinoma. Am J Cancer Res. 2020;10:2993-3036. [PubMed] |
| 5. | Chen T, You Y, Jiang H, Wang ZZ. Epithelial-mesenchymal transition (EMT): A biological process in the development, stem cell differentiation, and tumorigenesis. J Cell Physiol. 2017;232:3261-3272. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 251] [Cited by in RCA: 442] [Article Influence: 49.1] [Reference Citation Analysis (0)] |
| 6. | Batlle E, Sancho E, Francí C, Domínguez D, Monfar M, Baulida J, García De Herreros A. The transcription factor snail is a repressor of E-cadherin gene expression in epithelial tumour cells. Nat Cell Biol. 2000;2:84-89. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2178] [Cited by in RCA: 2070] [Article Influence: 79.6] [Reference Citation Analysis (0)] |
| 7. | Lamouille S, Xu J, Derynck R. Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol. 2014;15:178-196. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6736] [Cited by in RCA: 6509] [Article Influence: 542.4] [Reference Citation Analysis (1)] |
| 8. | De Craene B, Berx G. Regulatory networks defining EMT during cancer initiation and progression. Nat Rev Cancer. 2013;13:97-110. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2059] [Cited by in RCA: 1979] [Article Influence: 152.2] [Reference Citation Analysis (0)] |
| 9. | Nieto MA, Cano A. The epithelial-mesenchymal transition under control: global programs to regulate epithelial plasticity. Semin Cancer Biol. 2012;22:361-368. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 194] [Cited by in RCA: 205] [Article Influence: 14.6] [Reference Citation Analysis (0)] |
| 10. | Castro FA, Ivansson EL, Schmitt M, Juko-Pecirep I, Kjellberg L, Hildesheim A, Gyllensten UB, Pawlita M. Contribution of TMC6 and TMC8 (EVER1 and EVER2) variants to cervical cancer susceptibility. Int J Cancer. 2012;130:349-355. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 31] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 11. | Madeleine MM, Carter JJ, Johnson LG, Wipf GC, Davis C, Berg D, Nelson K, Daling JR, Schwartz SM, Galloway DA. Risk of squamous cell skin cancer after organ transplant associated with antibodies to cutaneous papillomaviruses, polyomaviruses, and TMC6/8 (EVER1/2) variants. Cancer Med. 2014;3:1440-1447. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 19] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 12. | Lu P, Ding Q, Ding S, Fan Y, Li X, Tian D, Liu M. Transmembrane channel-like protein 8 as a potential biomarker for poor prognosis of hepatocellular carcinoma. Mol Clin Oncol. 2017;7:244-248. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 5] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 13. | Lin B, Wang S, Yao Y, Shen Y, Yang H. Comprehensive co-expression analysis reveals TMC8 as a prognostic immune-associated gene in head and neck squamous cancer. Oncol Lett. 2021;22:498. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 12] [Article Influence: 2.4] [Reference Citation Analysis (1)] |
| 14. | Cheng Y, Wang K, Geng L, Sun J, Xu W, Liu D, Gong S, Zhu Y. Identification of candidate diagnostic and prognostic biomarkers for pancreatic carcinoma. EBioMedicine. 2019;40:382-393. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 80] [Cited by in RCA: 92] [Article Influence: 13.1] [Reference Citation Analysis (0)] |
| 15. | Aushev VN, Gopalakrishnan K, Teitelbaum SL, Parada H Jr, Santella RM, Gammon MD, Chen J. Tumor expression of environmental chemical-responsive genes and breast cancer mortality. Endocr Relat Cancer. 2019;26:843-851. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 16] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 16. | Subrungruanga I, Thawornkunob C, Chawalitchewinkoon-Petmitrc P, Pairojkul C, Wongkham S, Petmitrb S. Gene expression profiling of intrahepatic cholangiocarcinoma. Asian Pac J Cancer Prev. 2013;14:557-563. [PubMed] |
| 17. | Zhang W, Wang S, Zhang X, Liu K, Song J, Leng X, Luo R, Ran L. Transmembrane Channel-Like 5 (TMC5) promotes prostate cancer cell proliferation through cell cycle regulation. Biochimie. 2019;165:115-122. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 16] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 18. | Arroyo M, Larrosa R, Gómez-Maldonado J, Cobo MÁ, Claros MG, Bautista R. Expression-based, consistent biomarkers for prognosis and diagnosis in lung cancer. Clin Transl Oncol. 2020;22:1867-1874. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 13] [Article Influence: 2.2] [Reference Citation Analysis (0)] |
| 19. | Zhan C, Yan L, Wang L, Sun Y, Wang X, Lin Z, Zhang Y, Shi Y, Jiang W, Wang Q. Identification of immunohistochemical markers for distinguishing lung adenocarcinoma from squamous cell carcinoma. J Thorac Dis. 2015;7:1398-1405. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 42] [Reference Citation Analysis (0)] |
| 20. | Xiao J, Lu X, Chen X, Zou Y, Liu A, Li W, He B, He S, Chen Q. Eight potential biomarkers for distinguishing between lung adenocarcinoma and squamous cell carcinoma. Oncotarget. 2017;8:71759-71771. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 33] [Cited by in RCA: 53] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 21. | Huang R, Liao X, Li Q. Identification and validation of potential prognostic gene biomarkers for predicting survival in patients with acute myeloid leukemia. Onco Targets Ther. 2017;10:5243-5254. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 38] [Cited by in RCA: 62] [Article Influence: 6.9] [Reference Citation Analysis (0)] |
| 22. | Yusenko MV, Kovacs G. Identifying CD82 (KAI1) as a marker for human chromophobe renal cell carcinoma. Histopathology. 2009;55:687-695. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 36] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 23. | Zhang H, Zhang X, Xu W, Wang J. TMC5 is Highly Expressed in Human Cancers and Corelates to Prognosis and Immune Cell Infiltration: A Comprehensive Bioinformatics Analysis. Front Mol Biosci. 2021;8:810864. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 8] [Reference Citation Analysis (0)] |
| 24. | Pan L, Fang J, Chen MY, Zhai ST, Zhang B, Jiang ZY, Juengpanich S, Wang YF, Cai XJ. Promising key genes associated with tumor microenvironments and prognosis of hepatocellular carcinoma. World J Gastroenterol. 2020;26:789-803. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 21] [Cited by in RCA: 31] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 25. | Tang W, Shi Z, Zhu Y, Shan Z, Jiang A, Wang A, Chen M, Bao Y, Ju G, Xu W, Wang J. Comprehensive analysis of the prognosis and immune infiltration of TMC family members in renal clear cell carcinoma. Sci Rep. 2023;13:11668. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 5] [Reference Citation Analysis (0)] |
| 26. | Fischer AN, Fuchs E, Mikula M, Huber H, Beug H, Mikulits W. PDGF essentially links TGF-beta signaling to nuclear beta-catenin accumulation in hepatocellular carcinoma progression. Oncogene. 2007;26:3395-3405. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 117] [Cited by in RCA: 113] [Article Influence: 5.9] [Reference Citation Analysis (0)] |
| 27. | Gotzmann J, Huber H, Thallinger C, Wolschek M, Jansen B, Schulte-Hermann R, Beug H, Mikulits W. Hepatocytes convert to a fibroblastoid phenotype through the cooperation of TGF-beta1 and Ha-Ras: steps towards invasiveness. J Cell Sci. 2002;115:1189-1202. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 158] [Cited by in RCA: 149] [Article Influence: 6.2] [Reference Citation Analysis (0)] |
| 28. | Petz M, Them NC, Huber H, Mikulits W. PDGF enhances IRES-mediated translation of Laminin B1 by cytoplasmic accumulation of La during epithelial to mesenchymal transition. Nucleic Acids Res. 2012;40:9738-9749. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 38] [Cited by in RCA: 49] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 29. | van Zijl F, Mair M, Csiszar A, Schneller D, Zulehner G, Huber H, Eferl R, Beug H, Dolznig H, Mikulits W. Hepatic tumor-stroma crosstalk guides epithelial to mesenchymal transition at the tumor edge. Oncogene. 2009;28:4022-4033. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 150] [Cited by in RCA: 142] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 30. | Zhu K, Dai Z, Pan Q, Wang Z, Yang GH, Yu L, Ding ZB, Shi GM, Ke AW, Yang XR, Tao ZH, Zhao YM, Qin Y, Zeng HY, Tang ZY, Fan J, Zhou J. Metadherin promotes hepatocellular carcinoma metastasis through induction of epithelial-mesenchymal transition. Clin Cancer Res. 2011;17:7294-7302. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 117] [Cited by in RCA: 115] [Article Influence: 7.7] [Reference Citation Analysis (1)] |
| 31. | Pastushenko I, Blanpain C. EMT Transition States during Tumor Progression and Metastasis. Trends Cell Biol. 2019;29:212-226. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2351] [Cited by in RCA: 2084] [Article Influence: 297.7] [Reference Citation Analysis (0)] |
| 32. | Adiga D, Radhakrishnan R, Chakrabarty S, Kumar P, Kabekkodu SP. The Role of Calcium Signaling in Regulation of Epithelial-Mesenchymal Transition. Cells Tissues Organs. 2022;211:134-156. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 18] [Cited by in RCA: 24] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 33. | Jia Y, Zhao Y, Kusakizako T, Wang Y, Pan C, Zhang Y, Nureki O, Hattori M, Yan Z. TMC1 and TMC2 Proteins Are Pore-Forming Subunits of Mechanosensitive Ion Channels. Neuron. 2020;105:310-321.e3. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 130] [Article Influence: 18.6] [Reference Citation Analysis (0)] |
| 34. | Kasahara Y, Narukawa M, Takeuchi A, Tominaga M, Abe K, Asakura T. Molecular logic of salt taste reception in special reference to transmembrane channel-like 4 (TMC4). J Physiol Sci. 2022;72:31. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 19] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 35. | Richardson MM, Jennings LK, Zhang XA. Tetraspanins and tumor progression. Clin Exp Metastasis. 2011;28:261-270. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 63] [Cited by in RCA: 65] [Article Influence: 4.3] [Reference Citation Analysis (0)] |
| 36. | Obacz J, Takacova M, Brychtova V, Dobes P, Pastorekova S, Vojtesek B, Hrstka R. The role of AGR2 and AGR3 in cancer: similar but not identical. Eur J Cell Biol. 2015;94:139-147. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 44] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 37. | Stewart TA, Yapa KT, Monteith GR. Altered calcium signaling in cancer cells. Biochim Biophys Acta. 2015;1848:2502-2511. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 198] [Cited by in RCA: 264] [Article Influence: 22.0] [Reference Citation Analysis (0)] |
Open-Access: This article is an open-access article that was selected by an in-house editor and fully peer-reviewed by external reviewers. It is distributed in accordance with the Creative Commons Attribution NonCommercial (CC BY-NC 4.0) license, which permits others to distribute, remix, adapt, build upon this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See: https://creativecommons.org/Licenses/by-nc/4.0/