Copyright: ©Author(s) 2026.
World J Clin Oncol. Mar 24, 2026; 17(3): 117055
Published online Mar 24, 2026. doi: 10.5306/wjco.v17.i3.117055
Published online Mar 24, 2026. doi: 10.5306/wjco.v17.i3.117055
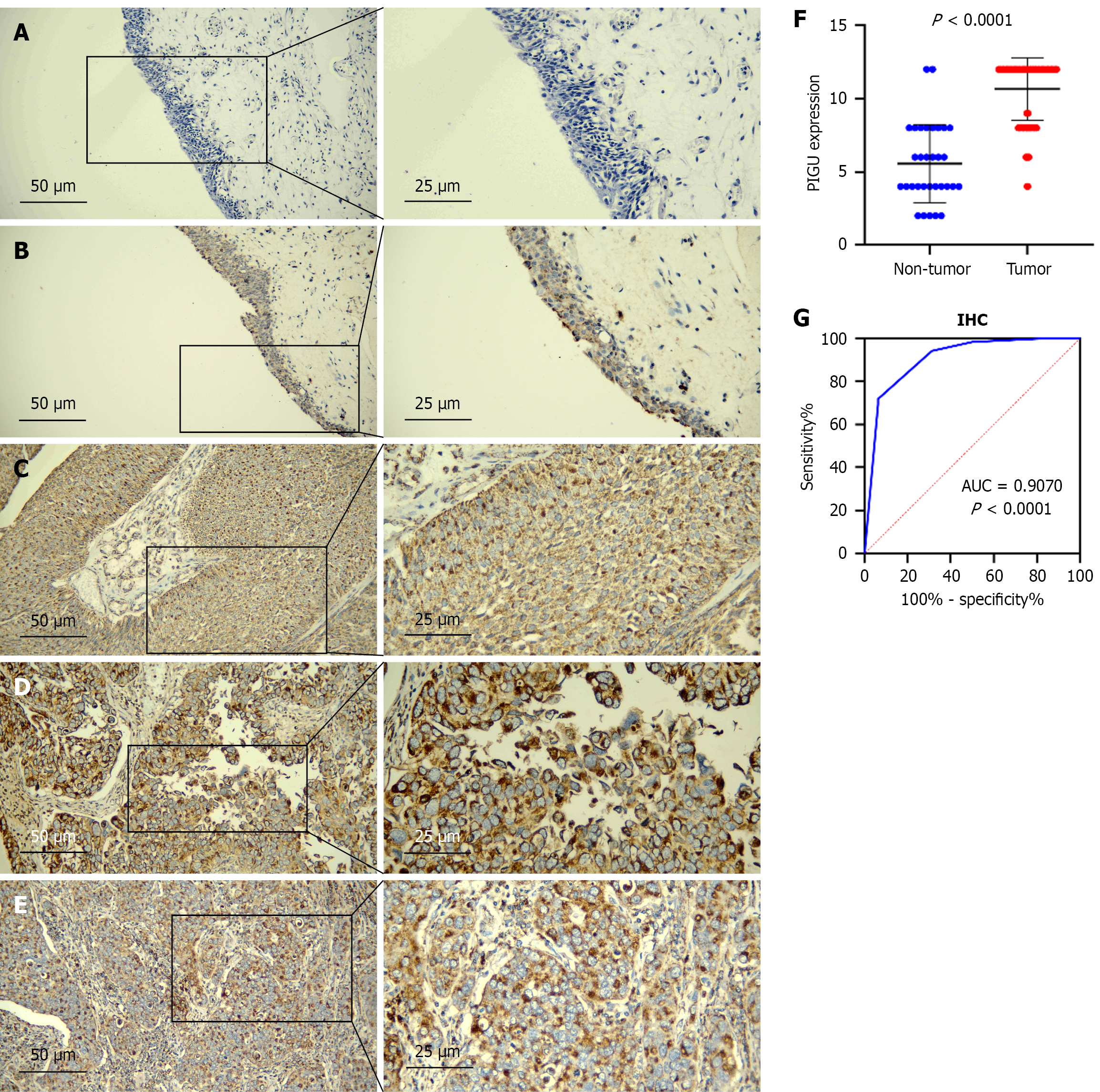
Figure 1 Immunohistochemistry detection of PIGU protein expression in bladder cancer tissues.
PIGU protein expression in para-cancerous tissues. A: Negative, × 200 (left), × 400 (right); B: Weakly positive, × 200 (left), × 400 (right); C-E: PIGU protein expression in bladder cancer (BLCA) tissue, all 12 points, × 200 (left), × 400 (right), pathological grade I-III (C: PIGU protein expression in BLCA tissue, positive, × 200-left, × 400-right); F: Scatter plot of immunohistochemical results; G: Receiver operating characteristic curve. IHC: Immunohistochemistry.
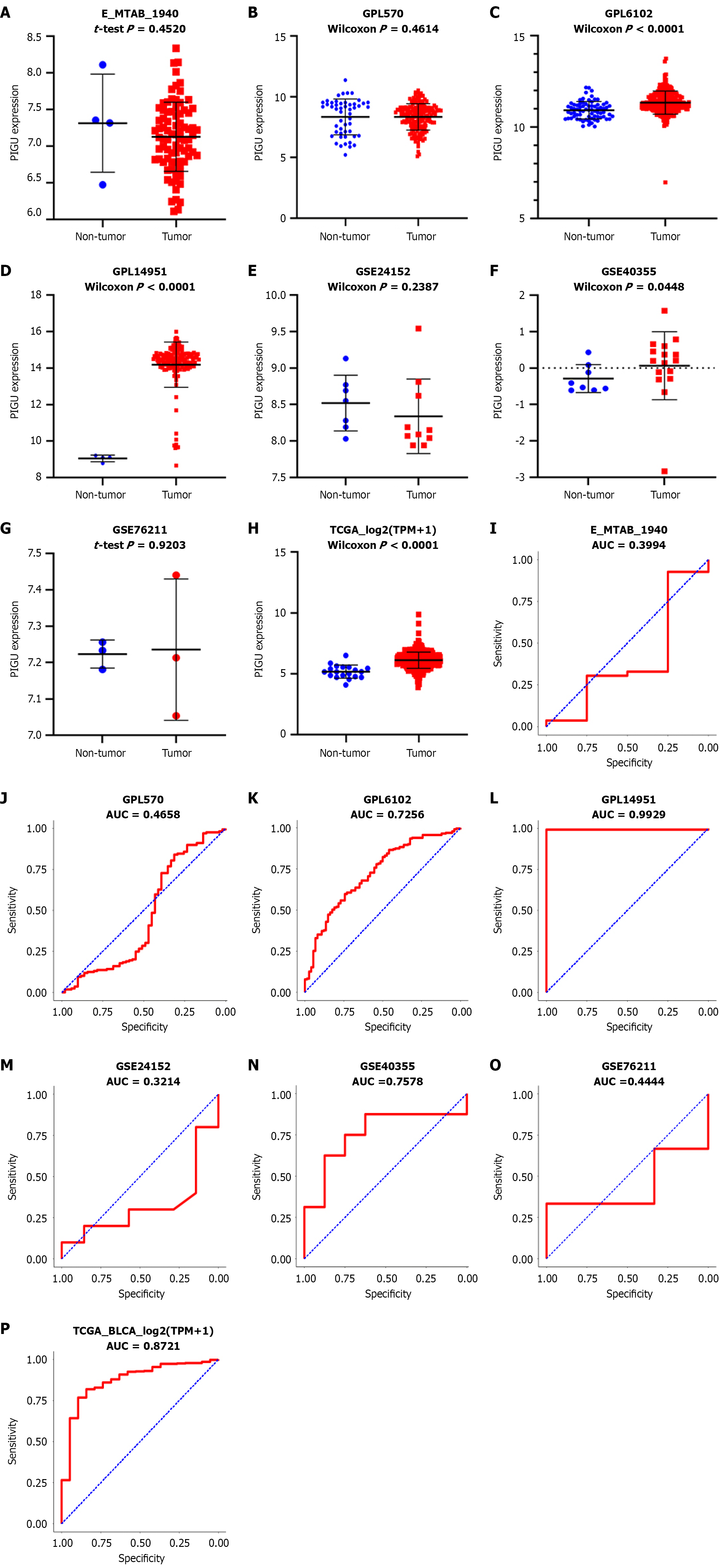
Figure 2 Analysis of all included microarray and RNA-seq datasets.
A-H: Scatter plots of PIGU expression across datasets; I-P: Receiver operating characteristic curves for PIGU in each dataset. AUC: Area under the curve.
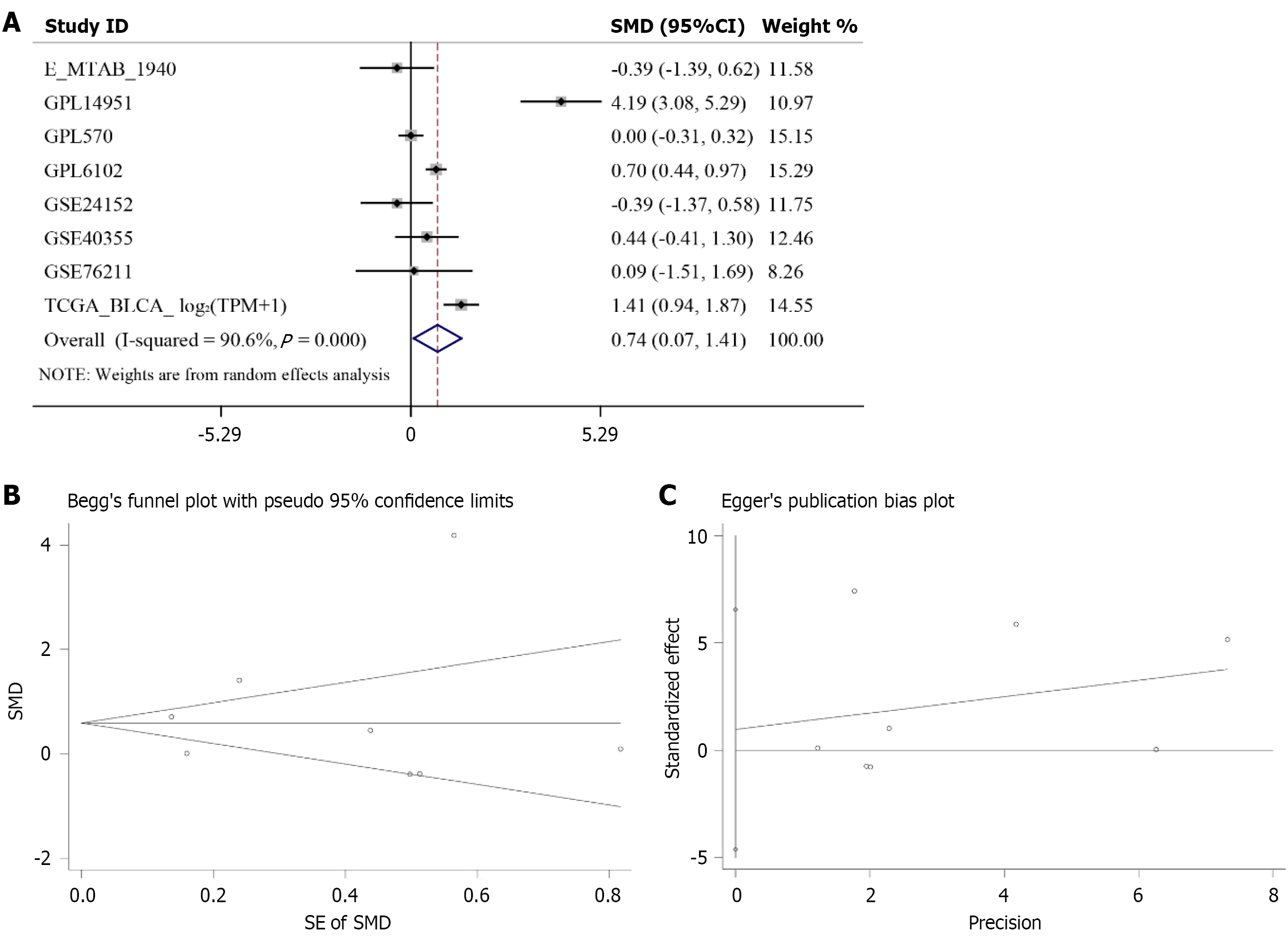
Figure 3 Comprehensive analysis indicates that PIGU mRNA expression is upregulated in bladder cancer.
A: Standardized mean difference forest plot for PIGU; B: Begg test; C: Egger’s publication bias test. SMD: Standardized mean difference.
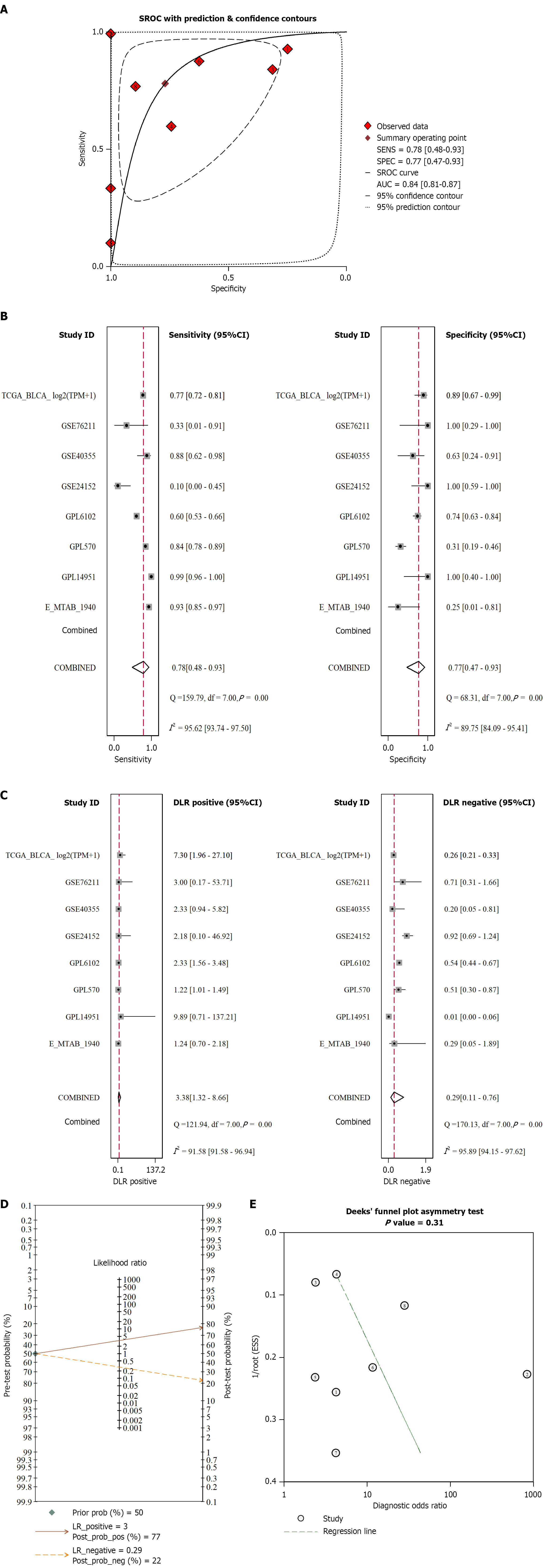
Figure 4 Differential testing of PIGU in bladder cancer and non-bladder cancer tissues.
A: Receiver operating characteristic curve summary based on all included datasets; B: Sensitivity and specificity; C: Positive likelihood ratio and negative likelihood ratio; D: Fagan plot; E: Deek’s publication bias test. AUC: Area under the curve; SROC: Summary receiver operating characteristic.
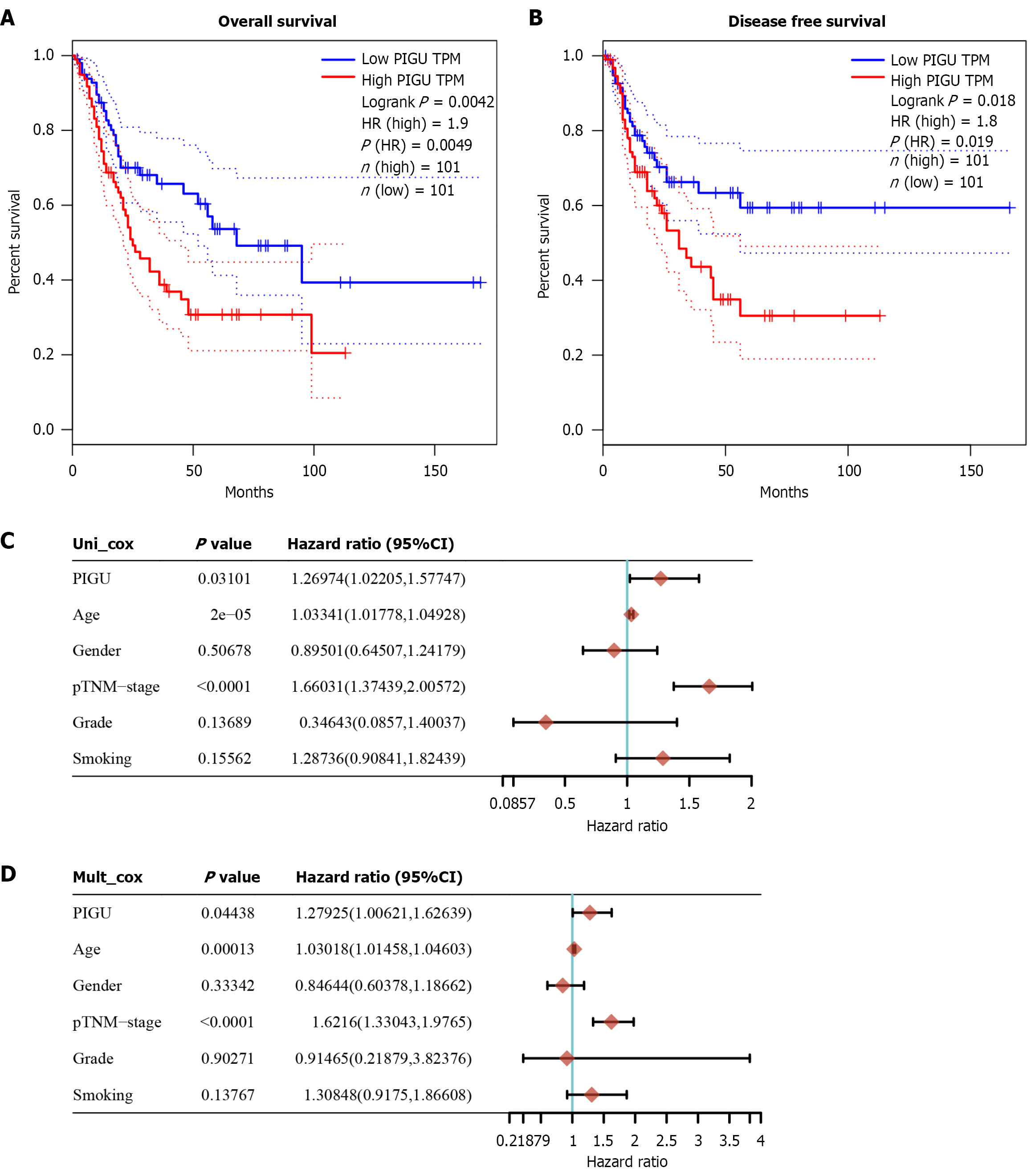
Figure 5 Prognostic value of PIGU mRNA expression levels in bladder cancer.
A and B: Kaplan-Meier curves for overall survival and disease-free survival in bladder cancer patients with high- and low-expression PIGU groups, where the bottom 25% represents the low-expression group and the top 25% represents the high-expression group; C: Forest plot of univariate Cox regression analysis; D: Forest plot of multivariate Cox regression analysis.
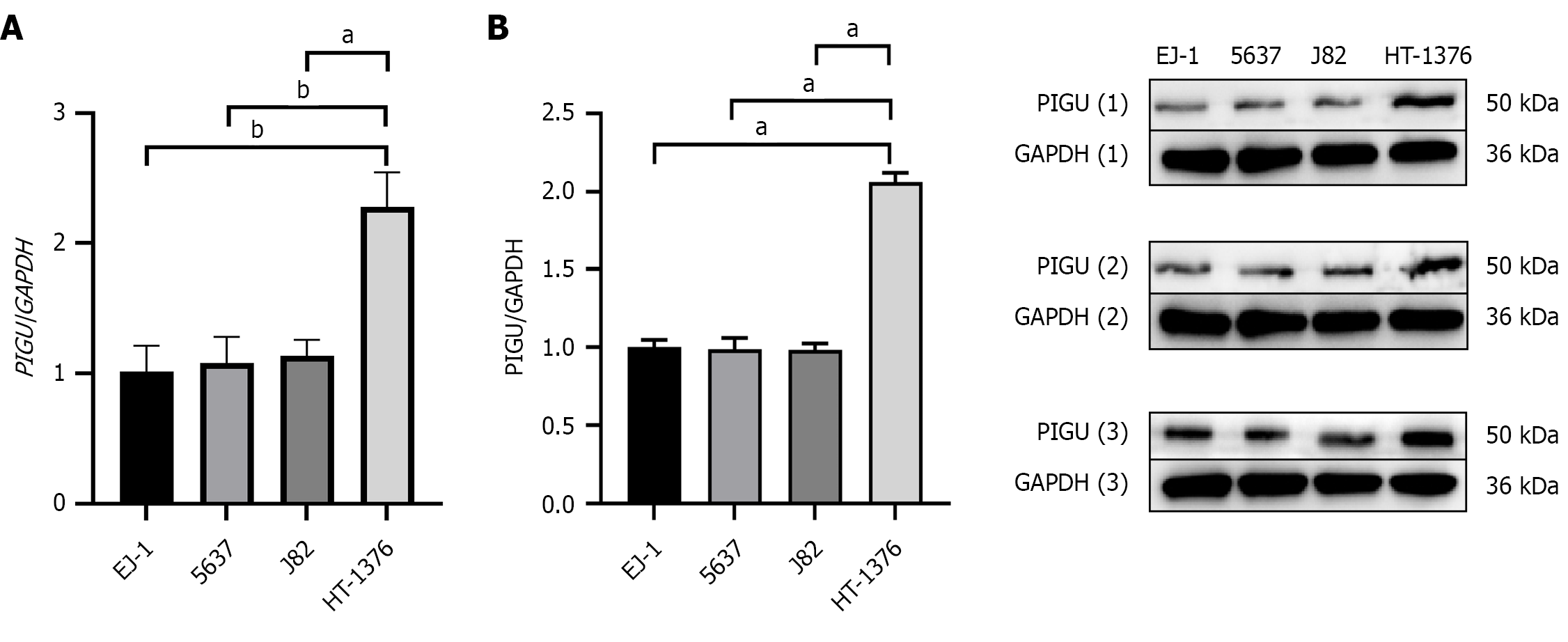
Figure 6 Relative expression levels of PIGU in EJ-1, 5637, J82, and HT-1376 bladder cancer cells.
A: Relative expression levels of the PIGU gene; B: Relative expression levels of PIGU protein. aP < 0.001, bP < 0.0001.
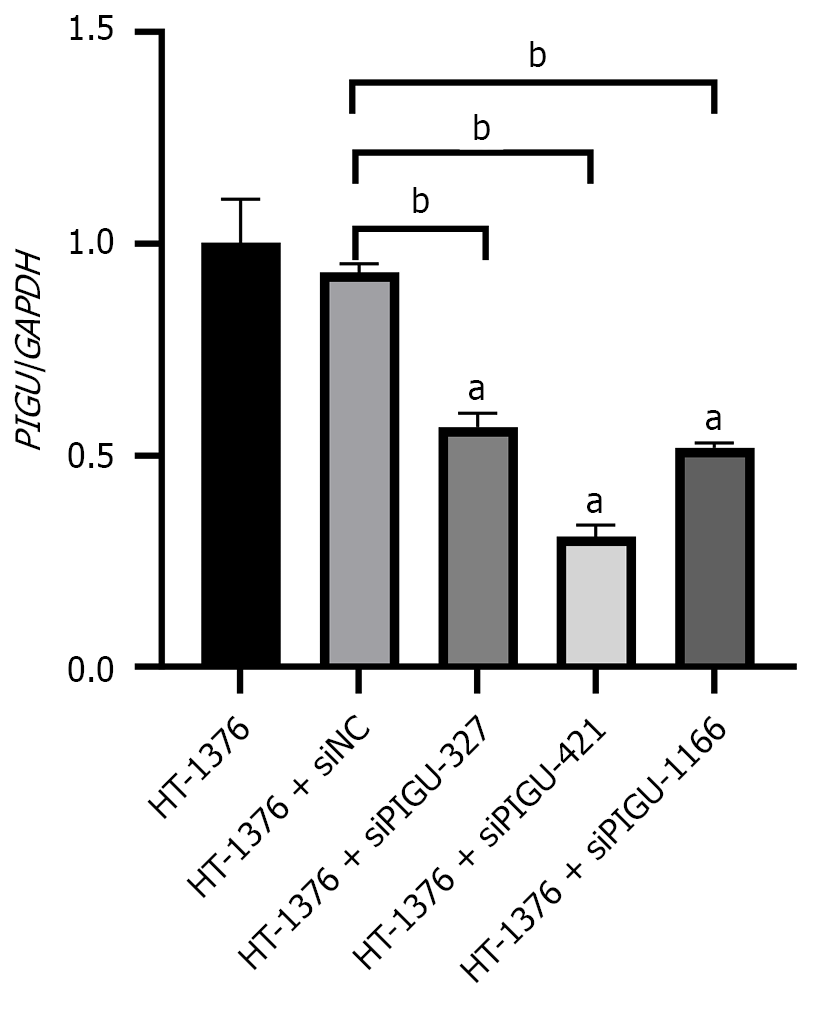
Figure 7 Relative expression levels of PIGU in bladder cancer cells infected with silencing and empty vector lentiviruses.
aP < 0.0001 vs group HT-1376, bP < 0.0001 vs group HT-1376 + siNC.
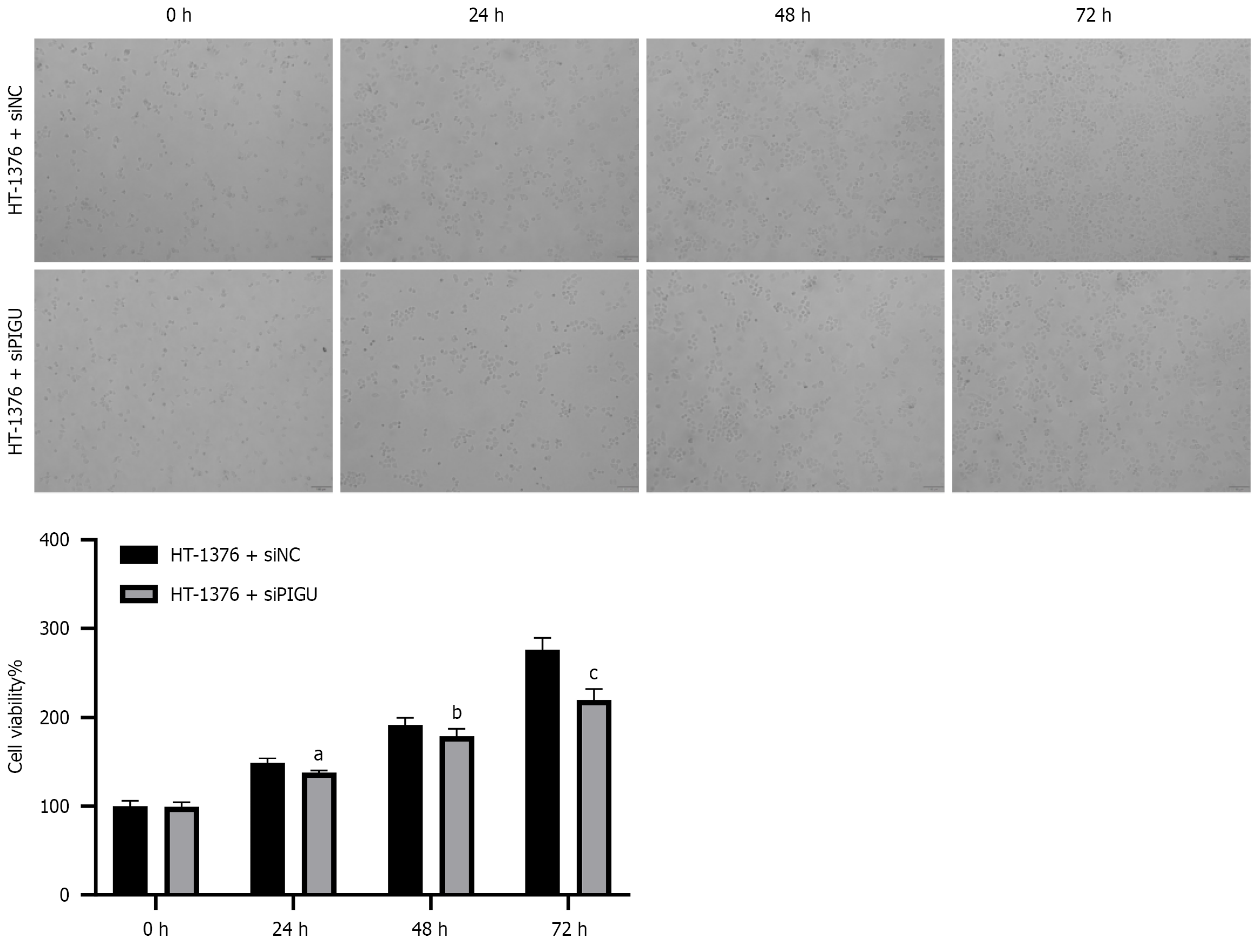
Figure 8 Effects of silent PIGU on in vitro proliferation of bladder cancer cells.
aP < 0.05, bP < 0.01, cP < 0.0001.
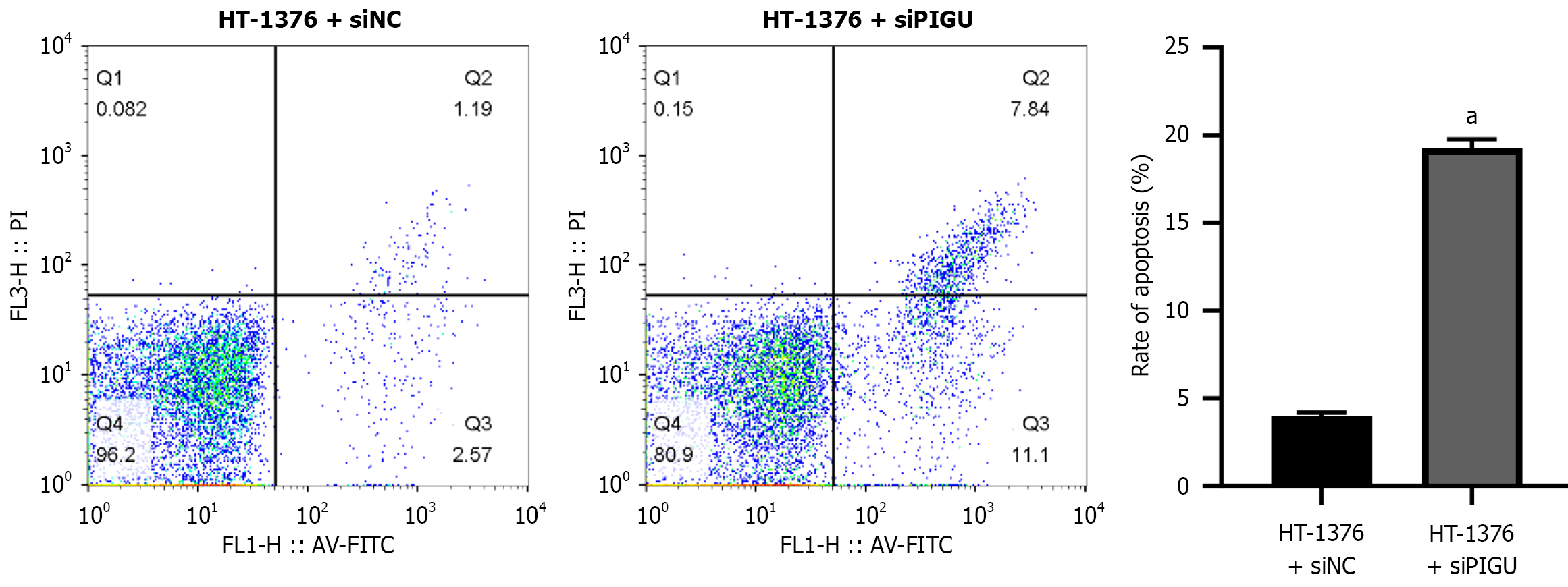
Figure 9 Effect of silencing PIGU on apoptosis in bladder cancer cells.
aP < 0.0001.
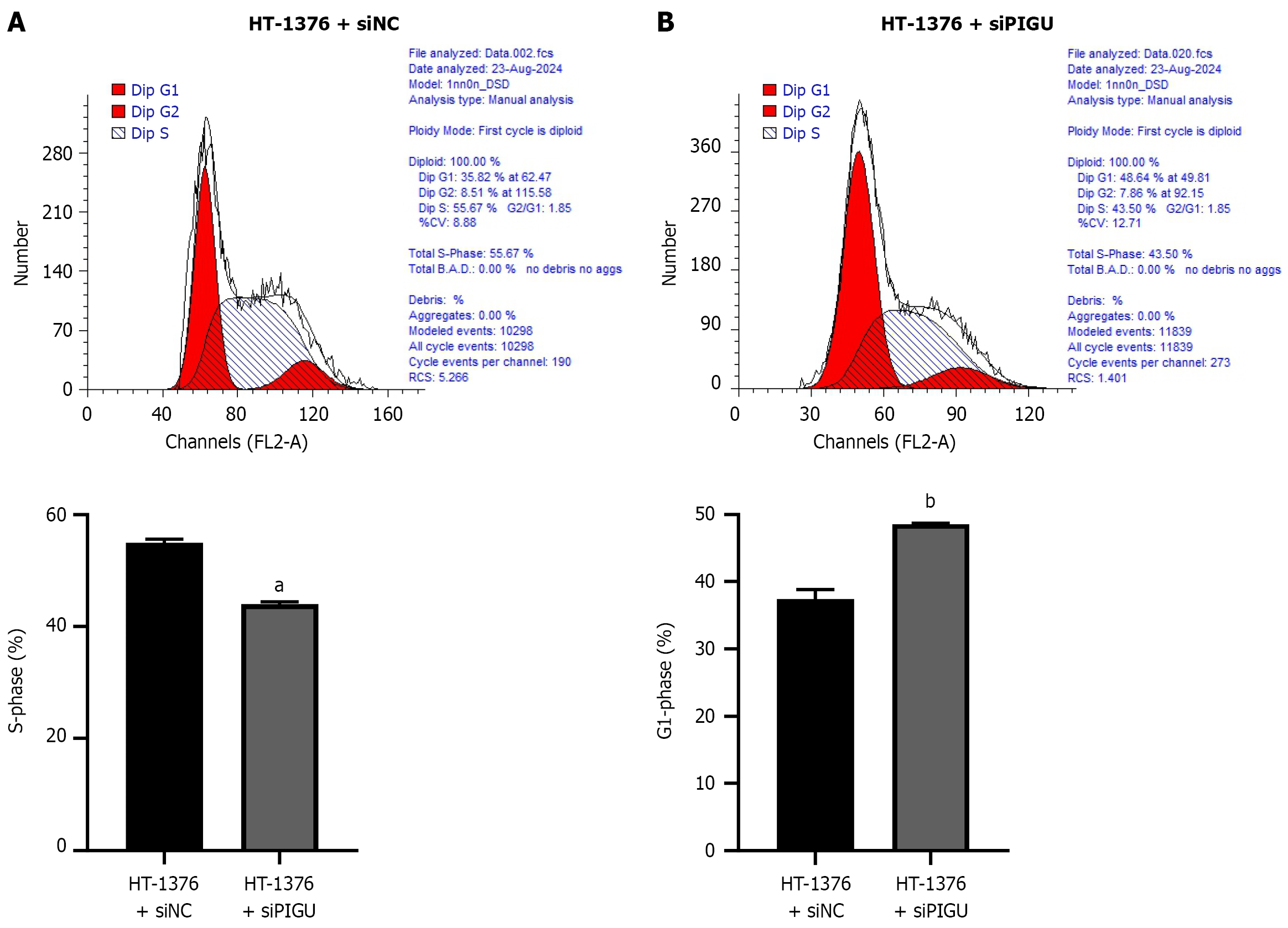
Figure 10 Effects of silencing PIGU on the cell cycle of bladder cancer cells.
A: Experimental results for the HT-1376 + siNC group; B: Experimental results for the HT-1376 + siPIGU group. aP < 0.0001, bP < 0.001.
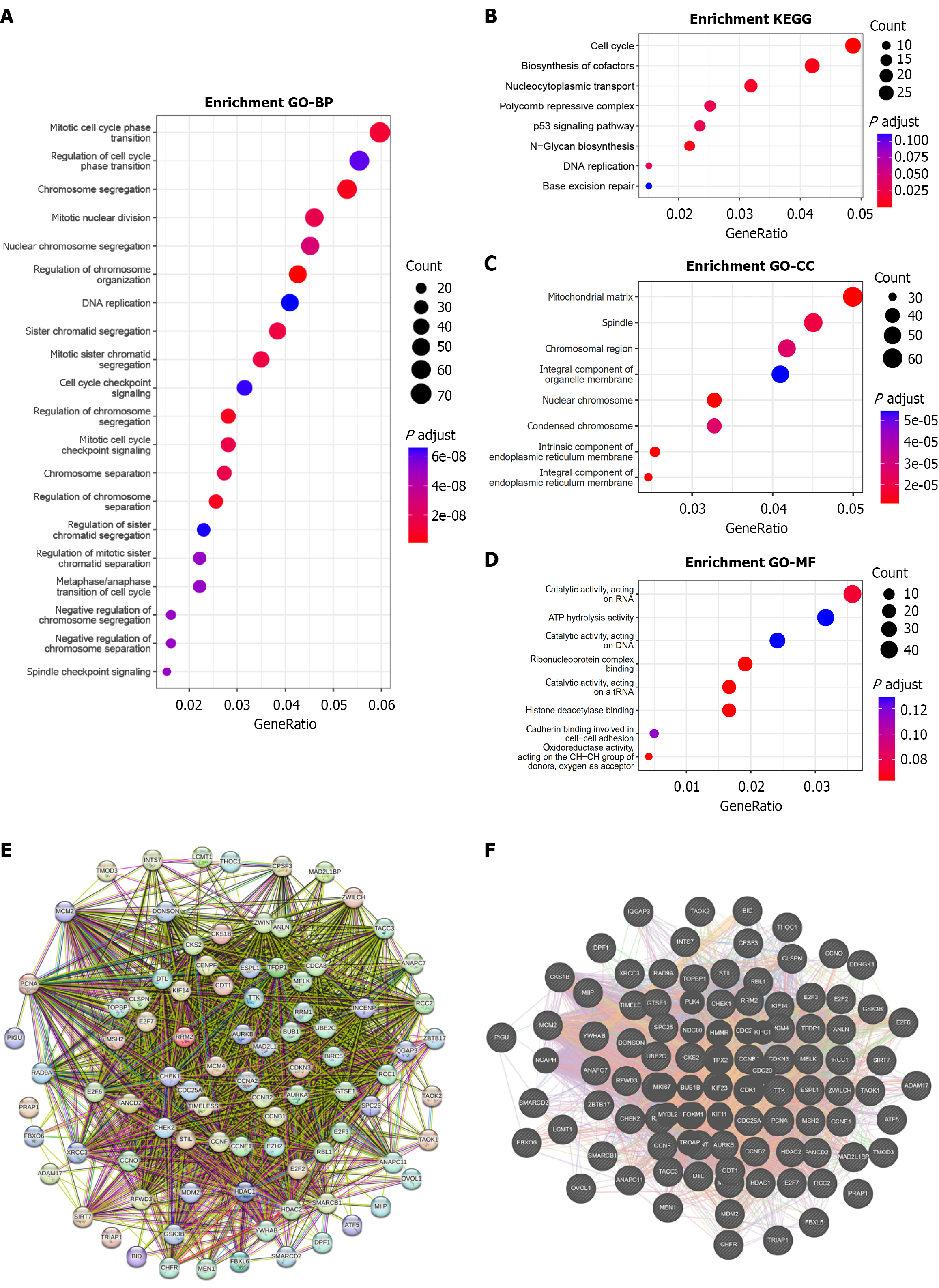
Figure 11 Enrichment analysis of PIGU differentially co-expressed genes and protein-protein interaction network analysis.
A: Biological process in Gene Ontology enrichment analysis; B: Kyoto Encyclopedia of Genes and Genomes enrichment analysis; C and D: Cellular component and molecular function analysis of graphene oxide; E and F: Protein-protein interaction network analysis (E: The String database; F: Gene MANIA database). BP: Biological process; KEGG: Kyoto Encyclopedia of Genes and Genomes; CC: Cellular component; MF: Molecular function.
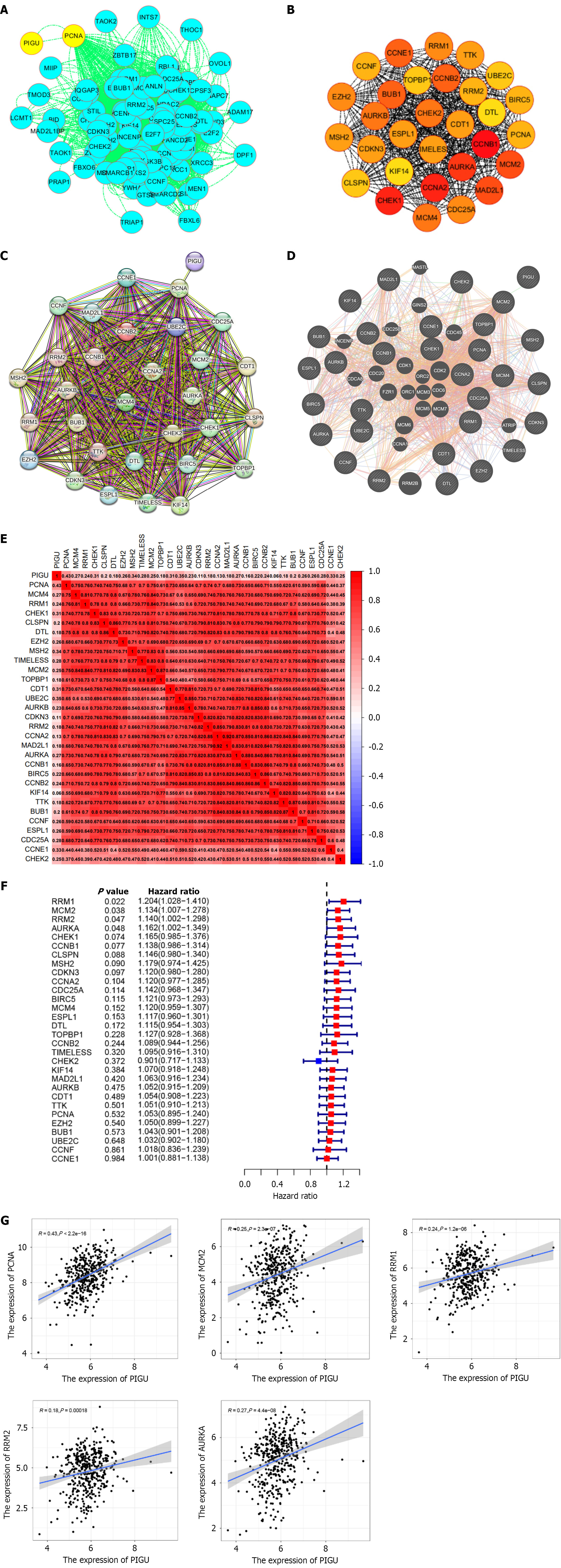
Figure 12 Identification of cell cycle-related hub genes.
A: Protein interaction results from the String database visualized in Cytoscape; B: Hub genes with the strongest protein interactions identified by the Cytoscape algorithm using the cytoHubba module; color shading indicates interaction strength; C and D: Protein-protein interaction network analysis (C: The String database; D: GeneMANIA database); E: Heatmap showing PIGU correlations with 30 hub genes; F: Forest plot of COX regression analysis for 30 hub genes; G: Scatterplot showing PIGU correlations with identified genes PCNA, MCM2, RRM1, RRM2, and AURKA.
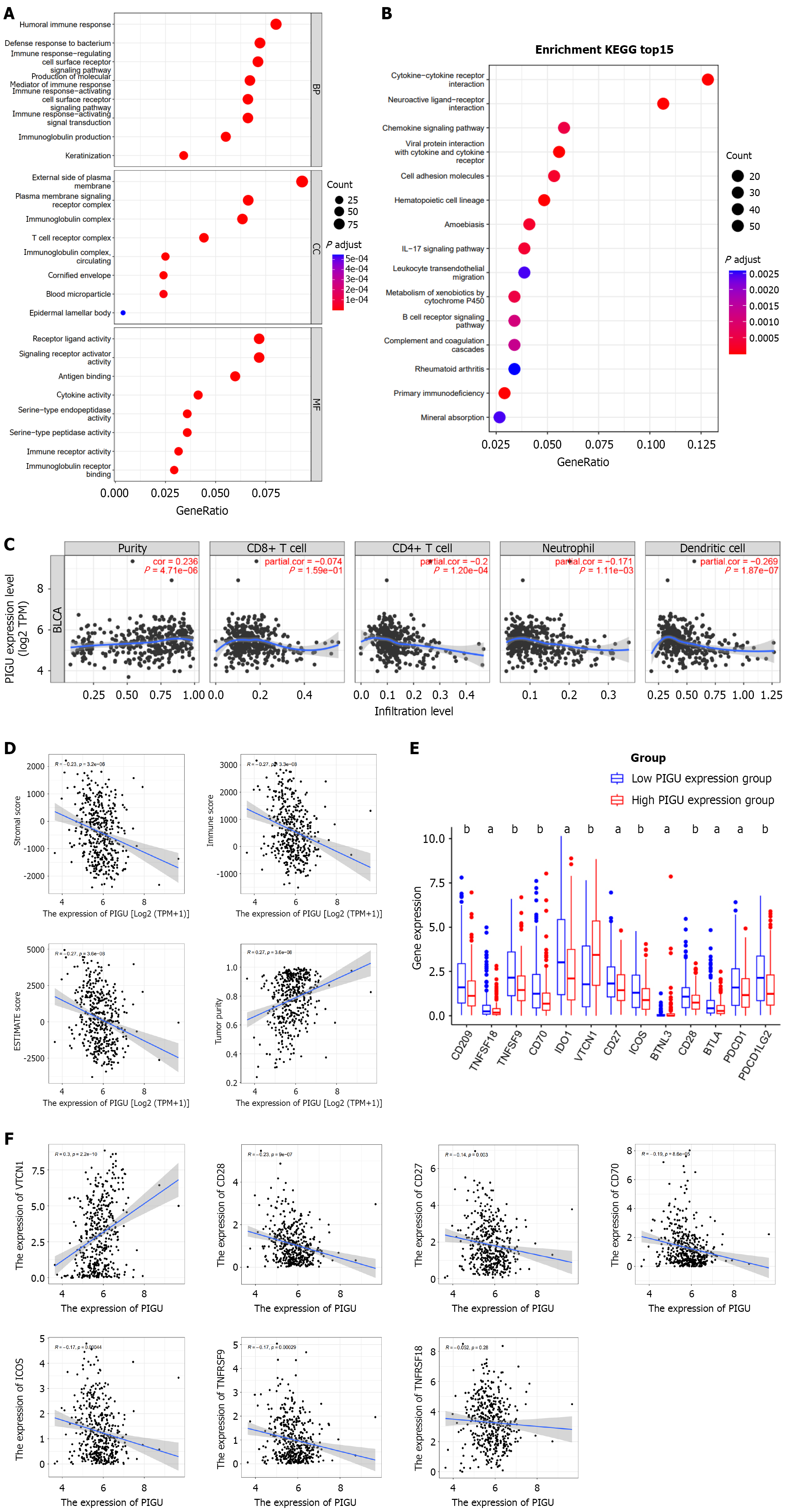
Figure 13 Relationship between PIGU expression and tumor immune infiltration.
A: Gene Ontology enrichment analysis of differentially expressed genes in high- and low-PIGU expression groups; B: Kyoto Encyclopedia of Genes and Genomes enrichment analysis; C: Correlation between PIGU expression and immune cell infiltration levels from the TIMER website; D: Correlation between PIGU expression and stromal score, immune score, ESTIMATE score, and tumor purity; E: Differential expression of 13 immune checkpoint genes in high- and low-PIGU expression groups; F: Correlation scatter plots of PIGU with VTCN1, CD28, CD27, CD70, ICOS, TNFRSF9, and TNFRSF18. aP < 0.01, bP < 0.001.
- Citation: Li SH, Liu ZS, Chen YJ, Li JD, Wei L, Chen YY, Zhang W, Huang ZG, Chen ZD, Chen G, Wei DM, Mo ZN, Deng LL. PIGU overexpression and regulation of cellular biological functions in bladder cancer. World J Clin Oncol 2026; 17(3): 117055
- URL: https://www.wjgnet.com/2218-4333/full/v17/i3/117055.htm
- DOI: https://dx.doi.org/10.5306/wjco.v17.i3.117055