Published online Apr 28, 2026. doi: 10.4329/wjr.v18.i4.118196
Revised: January 21, 2026
Accepted: February 4, 2026
Published online: April 28, 2026
Processing time: 119 Days and 1 Hours
This letter to the editor discusses a recent multi-institutional study that developed a noninvasive deep learning-based radiomics score derived from preoperative magnetic resonance imaging (MRI) to predict event-free survival in pediatric hepatoblastoma. The original study by Yang and Li published in World Journal of Radiology, leveraged convolutional neural networks to extract high-dimensional features from T1 and T2 sequences, the researchers developed an integrated nomogram that combines these imaging signatures with traditional markers like alpha-fetoprotein and the pretreatment extension of disease stage. This model significantly out
Core Tip: Accurate preoperative risk stratification remains challenging in pediatric hepatoblastoma. This study demonstrates that a magnetic resonance imaging-based deep learning radiomics score predicts event-free survival and refines risk stratification beyond conventional clinical factors. Integration with pretreatment extension of disease stage and alpha-fetoprotein improves prognostic accuracy, supporting noninvasive, imaging-driven decision-making for individualized.
- Citation: Chowdhury U, Mahajan AA, Kavitha MS, Rajendran RL, Gangadaran P, Ahn BC. Letter to the Editor: Magnetic resonance imaging-based deep learning radiomics for preoperative risk stratification in pediatric hepatoblastoma. World J Radiol 2026; 18(4): 118196
- URL: https://www.wjgnet.com/1949-8470/full/v18/i4/118196.htm
- DOI: https://dx.doi.org/10.4329/wjr.v18.i4.118196
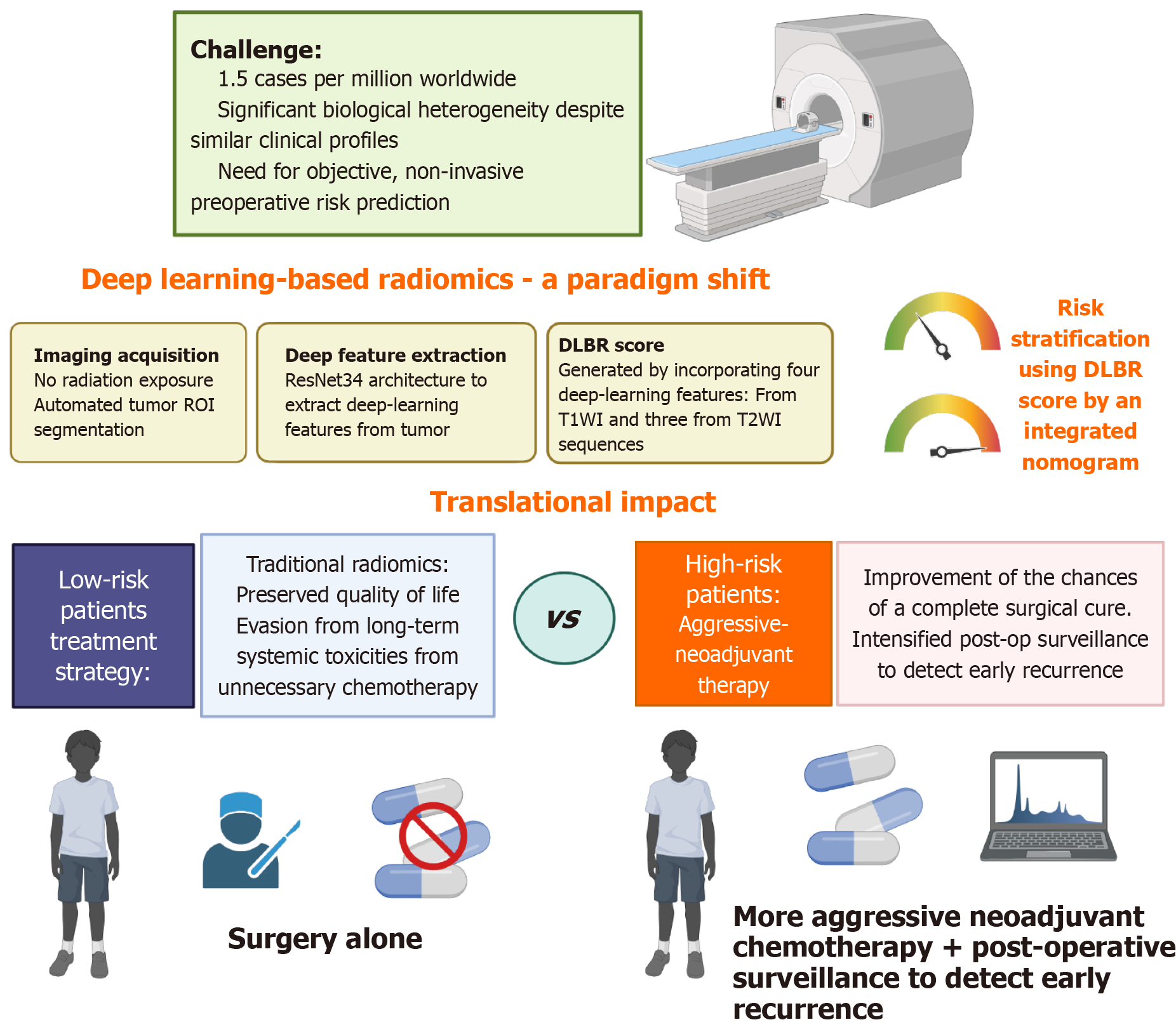
The original study by Yang and Li[1] published in World Journal of Radiology, leveraged convolutional neural networks (CNNs) to extract high-dimensional features from T1 and T2 sequences, the researchers developed an integrated nomogram that combines these imaging signatures with traditional markers like alpha-fetoprotein and the pretreatment extension of disease stage. Hepatoblastoma represents the most prevalent primary hepatic malignancy in the pediatric population, occurring at a rate of approximately 1.5 cases per million worldwide[2]. While complete surgical resection of primary lesions is the established first-line treatment for early-stage cases without distant metastasis, these tumors exhibit significant biological heterogeneity that leads to widely divergent survival outcomes even among patients with similar clinical profiles[3]. To address this, current protocols utilize the pretreatment extension of disease (PRETEXT) system to guide personalized therapeutic strategies. However, this system relies heavily on qualitative imaging assessment which is often prone to unsatisfactory accuracy and a tendency toward over-staging when evaluated by the naked eye[4]. Furthermore, existing risk-stratification models frequently rely on “low-latitude” clinical data and qualitative assessments while ignoring the “high-latitude” features hidden within imaging data that could better reflect tumor heterogeneity. Consequently, there is an urgent clinical need to develop objective, noninvasive tools that can identify high-risk patients before surgery, allowing clinicians to determine if intensive neoadjuvant chemotherapy is required to improve long-term prognosis[5].
Deep learning-based radiomics (DLBR) represents a paradigm shift from traditional handcrafted features by utilizing advanced CNNs to automatically decode high-throughput quantitative data[6]. While conventional radiomics relies on predefined mathematical descriptors, DLBR captures complex hierarchical and spatial patterns often invisible to the human eye, directly addressing the biological ambiguity of standard imaging[1].
In pediatric populations, magnetic resonance imaging (MRI) is the preferred modality for radiomic analysis because it provides highly detailed soft-tissue contrast and multi-planar imaging capabilities without exposing children to ionizing radiation. This multiparametric approach-leveraging both T1 and T2 characteristic that allows for a more comprehensive evaluation of tumor morphology than computed tomography scans.
Network: This study employed a ResNet34 architecture to extract 512 deep learning (DL) features from tumor regions of interest (ROIs) (Figure 1).
Automation: ROIs were segmented automatically using unsupervised clustering-based algorithms, such as simple-linear-iterative-clustering superpixels and fuzzy c-means clustering, ensuring consistent feature derivation across the cohort. These automated contours were then visually verified by two experienced radiologists who were blinded to all patient clinical outcomes and survival data to prevent observer bias.
Biological signatures: These high-level features are uniquely equipped to interpret intratumoral heterogeneity, providing a quantitative bridge to qualitative descriptors like boundary irregularity and internal septations. This mapping ensures that the DLBR score is not merely a mathematical output but a reflection of the tumor’s physical morphology, which is often difficult for the human eye to consistently quantify. Manual intervention was kept to a minimum, primarily serving to confirm the exclusion of peritumoral vessels or liver parenchyma. The high level of agreement between the automated system and radiologists was reflected in a mean dice coefficient of 0.906, indicating that the DLBR score is derived from highly stable and objective spatial data[1].
Clinical value: By mapping these intricate patterns, DLBR provides a more objective biological signature of the malignancy, enabling precise risk stratification where qualitative visual assessments by radiologists might fail[1].
From the initial 512 DL features extracted via ResNet34, feature selection was refined using ComBat compensation technology to filter out inconsistent variables across different MRI equipment[1]. This ensured that only robust, high-latitude features were retained for final score construction.
While DL features are often considered ‘black-box’ components, our study utilized Spearman correlations to map these abstract variables to established radiological descriptors (Table 1). For example, features derived from T1WI sequences were found to correlate strongly with border irregularity and margin sharpness. Conversely, T2WI-derived features were indicative of intratumoral texture heterogeneity and signal variations, reflecting the underlying biological diversity of the lesion[1].
| DL feature class | MRI sequence | Radiological interpretation |
| Border features | T1-weighted (T1WI) | Correlates with margin sharpness and lesion boundary irregularity |
| Texture features | T2-weighted (T2WI) | Reflects intratumoral heterogeneity and signal intensity variations |
| Internal features | T2-weighted (T2WI) | Associated with the presence of internal septations or lobulations |
The final DLBR score was generated by incorporating four essential deep-learning features in which one derived from T1WI and three from T2WI sequences.
DLBR score acted as an independent predictor of event-free survival (EFS) in both the training and external testing cohorts (P < 0.001).
Using a cutoff score of 0.0, the model accurately divided patients into low-risk and high-risk groups with significantly different mean survival times.
Spearman correlations suggested these abstract DL features correlate with clinical descriptors such as intratumoral texture heterogeneity, lesion boundary irregularity, and internal septations.
The final DLBR score utilized a cutoff of 0.0, derived from the median value of the training set, to categorize patients into high-risk and low-risk groups. To evaluate the model’s reliability, we utilized an external testing cohort for va
A significant hurdle in medical artificial intelligence is scanner effects. The variations in results caused by different hardware and imaging protocols across hospitals were mitigated by the following steps.
All images were resampled to 1 mm isotropic voxels and underwent Nyul standardization for intensity harmonization.
The researchers used ComBat compensation technology to filter out inconsistent features and retain only those that were robust across different MRI equipment and scanning parameters.
To ensure accuracy, tumor regions were segmented using unsupervised clustering-based algorithms (SLIC-S and FCM) and visually verified by experienced radiologists.
The data for this multi-institutional study were acquired from multiple clinical centers using various MRI scanners, including both 1.5T and 3.0T field strengths. This introduced significant imaging heterogeneity in terms of signal-to-noise ratios and sequence parameters. To standardize these variations, all images were resampled to 1 mm isotropic voxels and underwent Nyul intensity standardization. Furthermore, ComBat harmonization was specifically employed to remove ‘center effects’ arising from different hardware vendors and scanning protocols, ensuring the deep-learning features remained stable and representative of tumor biology rather than technical noise.
To ensure data quality, a strict inclusion criterion was applied where only patients with complete T1WI, T2WI, and clinical marker data (AFP and PRETEXT) were analyzed. Furthermore, the proportional hazards assumption for our survival models was tested and met, ensuring that the hazard ratios reported remained constant over the follow-up period.
The study’s most effective tool was the integrated clinical-DL nomogram, which combined the DLBR score with AFP concentration and the PRETEXT stage.
Decision-curve analysis showed that the integrated model provided a higher net benefit for clinical decision-making than using clinical models alone, particularly within the threshold probability range of 0.1 to 0.8. This indicates that within this range, using the nomogram to guide treatment leads to better patient outcomes than standard staging pro
Clinically, the integration of DLBR features helps mitigate the risks associated with misclassification. For instance, a false positive result might suggest treatment escalation (e.g., aggressive neoadjuvant therapy) for a patient who could have been cured by surgery alone, potentially increasing toxicity. Conversely, a false negative could lead to treatment de-escalation, risking early recurrence in high-risk patients. The superior C-index (0.696) and lower Integrated Brier Score of our model suggest a significant reduction in these predictive errors compared to traditional PRETEXT staging[1].
A primary limitation of this study is its retrospective nature and the relatively small size of the external validation cohort (n = 32)[1]. The current performance metrics are indicative of the model’s potential but do not yet establish clinical readiness for independent decision-making.
While the DLBR score demonstrates potential for future translation into routine practice, its current application remains a research tool pending prospective confirmation. Because this score is derived from preoperative MRI, it provides a critical window for clinicians to optimize treatment strategies based on the tumor’s specific biological signature (Table 2).
| Stage | Process applied | Purpose and clinical benefit |
| Normalization | Resampling to 1 mm isotropic voxels | Ensures all images have a consistent spatial resolution regardless of the original scanner settings |
| Intensity alignment | Nyul standardization | Harmonizes signal intensities across different T1 and T2 sequences to reduce “brightness” variations |
| Clustering | Simple-linear-iterative-clustering superpixels and fuzzy c-means clustering | Unsupervised algorithms that automatically group voxels to isolate hyperintense tumor regions |
| Validation | Manual radiologist review | Experts verified the automatic contours, achieving a high mean dice coefficient of 0.906 |
| Heterogeneity management | Multi-vendor, multi-field (1.5T/3.0T) | Accounts for real-world variation in scanner hardware and field strengths |
| Batch correction | ComBat harmonization | Removes “center effects” or scanner-specific noise to ensure the DLBR score is stable across different hospitals |
Patients identified as high-risk by the DLBR score-even if they appear low-risk via traditional staging could be considered for more aggressive neoadjuvant therapy or dose-dense chemotherapy to improve the chances of a complete surgical cure. Furthermore, these patients may require intensified postoperative surveillance to detect early recurrence[7].
Conversely, low-risk patients with favorable DLBR signatures might safely undergo surgery alone, thereby sparing them from unnecessary chemotherapy and its associated long-term systemic toxicities, such as hearing or heart damage[8].
To ensure the generalizability and reliability of the DLBR score in various clinical settings, future research must prioritize validation within larger prospective cohorts. Such studies will be essential to confirm the model’s stability across diverse patient populations and different institutional imaging protocols. Research will also explore the model’s ability to guide high-stakes decisions, such as determining candidacy for liver transplantation in complex cases. By identifying tumors with highly aggressive biological signatures that are unlikely to respond to neoadjuvant chemotherapy, the DLBR score could serve as a critical decision-support tool for early referral to transplant programs. This integration into real-world clinical workflows would enhance the model's impact by providing objective data for cases that are currently prone to subjective qualitative assessment
The development of an MRI-based DLBR score marks a significant advancement in the objective assessment of pediatric hepatoblastoma[1]. By translating complex imaging data into a usable clinical score, this study provides a reliable, noninvasive method to predict EFS and stratify patient risk. When integrated into a clinical nomogram, this approach offers pediatricians a powerful tool to facilitate personalized treatment and improve clinical outcomes for children with this rare malignancy. When integrated into a clinical nomogram, this approach offers pediatricians a powerful tool to facilitate personalized treatment, ranging from chemotherapy adjustment to assisting in high-stakes decisions like liver transplantation, ultimately improving clinical outcomes for children with this rare malignancy. While this study establishes a robust framework for MRI-based risk stratification, ongoing multi-center prospective validation will be critical to cementing its role as a universal standard in pediatric oncology.
| 1. | Yang YH, Li Y. Magnetic resonance imaging-based deep-learning radiomics score for survival prediction and risk stratification in pediatric hepatoblastoma receiving surgical resection. World J Radiol. 2025;18:115503. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in CrossRef: 1] [Cited by in RCA: 1] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 2. | Yang Y, Si J, Zhang K, Li J, Deng Y, Wang F, Liu H, He L, Chen X. Identification of high-risk hepatoblastoma in the CHIC risk stratification system based on enhanced CT radiomics features. Dig Liver Dis. 2025;57:1802-1809. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 3. | DeRenzi AD, Bowen A. A Case Report and a Review of Pediatric Hepatoblastoma. HCA Healthc J Med. 2023;4:377-382. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 2] [Reference Citation Analysis (0)] |
| 4. | Roebuck DJ, Aronson D, Clapuyt P, Czauderna P, de Ville de Goyet J, Gauthier F, Mackinlay G, Maibach R, McHugh K, Olsen OE, Otte JB, Pariente D, Plaschkes J, Childs M, Perilongo G; International Childrhood Liver Tumor Strategy Group. 2005 PRETEXT: a revised staging system for primary malignant liver tumours of childhood developed by the SIOPEL group. Pediatr Radiol. 2007;37:123-32; quiz 249. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 289] [Cited by in RCA: 215] [Article Influence: 11.3] [Reference Citation Analysis (4)] |
| 5. | Meyers RL, Maibach R, Hiyama E, Häberle B, Krailo M, Rangaswami A, Aronson DC, Malogolowkin MH, Perilongo G, von Schweinitz D, Ansari M, Lopez-Terrada D, Tanaka Y, Alaggio R, Leuschner I, Hishiki T, Schmid I, Watanabe K, Yoshimura K, Feng Y, Rinaldi E, Saraceno D, Derosa M, Czauderna P. Risk-stratified staging in paediatric hepatoblastoma: a unified analysis from the Children's Hepatic tumors International Collaboration. Lancet Oncol. 2017;18:122-131. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 354] [Cited by in RCA: 310] [Article Influence: 34.4] [Reference Citation Analysis (6)] |
| 6. | Avanzo M, Wei L, Stancanello J, Vallières M, Rao A, Morin O, Mattonen SA, El Naqa I. Machine and deep learning methods for radiomics. Med Phys. 2020;47:e185-e202. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 502] [Cited by in RCA: 412] [Article Influence: 68.7] [Reference Citation Analysis (0)] |
| 7. | Zhang P, Yao W, Li Z, Fan Y, Du X, Wang B, Zhang F, Hou J, Su Q. Radiomics for predicting sensitivity to neoadjuvant chemotherapy in osteosarcoma: current status and advances. Oncol Rev. 2025;19:1633211. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 2] [Reference Citation Analysis (0)] |
| 8. | He L, Li Z, Chen X, Huang Y, Yan L, Liang C, Liu Z. A radiomics prognostic scoring system for predicting progression-free survival in patients with stage IV non-small cell lung cancer treated with platinum-based chemotherapy. Chin J Cancer Res. 2021;33:592-605. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 14] [Article Influence: 2.8] [Reference Citation Analysis (0)] |