Published online Mar 26, 2026. doi: 10.4330/wjc.v18.i3.116217
Revised: November 19, 2025
Accepted: January 12, 2026
Published online: March 26, 2026
Processing time: 138 Days and 11.5 Hours
Diabetes mellitus (DM) and the related sequalae remains one of the most fre
To assess the diagnostic capabilities of the parameters of single lead electrocardiography (ECG) in the diagnosis of DM utilizing machine learning model.
A single center study involved 629 participants with vs without DM. All the study participants passed transthoracic echocardiography, fasting blood glucose mea
The cluster-stratified analysis revealed that the model performed best in cluster 4 (patients with high DM prevalence and significant comorbidities), achieving a sensitivity of 75%, specificity of 83%, and an area under the curve of 88%.
This study demonstrates that a phenotype-stratified approach is crucial for effective ECG-based DM screening. By identifying a specific clinical profile (cluster 4: High comorbidity burden with preserved cardiac function), we developed a model that accurately detects DM from a single-lead ECG. This phenotype-specific strategy overcomes the confounding effect of cardiovascular disease, moving beyond one-size-fits-all algorithms towards a precise and clinically viable tool for non-invasive DM detection in high-risk populations.
Core Tip: This study introduces a crucial paradigm shift for non-invasive diabetes detection using electrocardiography. Instead of a one-size-fits-all model, we employed phenotypic clustering to disentangle the confounding effects of cardiovascular disease. We identified a specific patient profile (cluster 4: High diabetes prevalence with significant but non-severe comorbidities) where a machine learning model, analyzing single-lead electrocardiography features like T-wave morphology and atrial conduction, achieves optimal and clinically viable performance (area under the curve: 0.88). This proves that diabetes-specific cardiac “whispers” are detectable, but only with a precision medicine approach that tailors diagnostics to distinct clinical phenotypes.
- Citation: Karbovskaya AD, Marzoog BA, Stroeva A, Suvorov A, Chomakhidze P, Gognieva D, Kuznetsova N, Syrkin A, Fadeev VV, Ismailova SM, Poluboyarinova IV, Kopylov P. Machine learning-based detection of diabetes mellitus from single-lead electrocardiography: A phenotype-stratified approach. World J Cardiol 2026; 18(3): 116217
- URL: https://www.wjgnet.com/1949-8462/full/v18/i3/116217.htm
- DOI: https://dx.doi.org/10.4330/wjc.v18.i3.116217
Diabetes mellitus (DM) represents one of the most significant global public health challenges of the 21st century, with prevalence rates continuing to rise alarmingly[1]. According to the International Diabetes Federation, approximately 589 million adults were living with diabetes in 2024, a figure projected to reach 853 million by 2050, disproportionately affecting low- and middle-income countries[2]. This epidemic carries devastating human and economic costs, driving increased morbidity and mortality through microvascular complications (retinopathy, nephropathy, neuropathy) and macrovascular sequelae [cardiovascular disease (CVD), stroke, peripheral artery disease][3-5]. Critically, an estimated 44% of diabetes cases remain undiagnosed globally, representing a substantial missed opportunity for early intervention that could prevent or delay debilitating complications[6]. The current diagnostic paradigm relies on invasive, resource-intensive, and often inaccessible methods - fasting plasma glucose, oral glucose tolerance tests, and glycated hemoglobin (HbA1c) - which suffer from limitations including cost, requirement for venous blood sampling, limited availability in resource-poor settings, and variable reliability (e.g., HbA1c influenced by hemoglobinopathies or anemia). Consequently, there is an urgent, unmet need for novel, non-invasive, scalable, and cost-effective screening tools capable of facilitating widespread early detection, particularly in underserved populations and primary care settings.
Electrocardiography (ECG), a ubiquitous, non-invasive, and low-cost technology, has emerged as a promising candidate for such a tool. The convergence of single-lead ECG devices and sophisticated machine learning (ML) repre
However, translating the potential of ECG-based DM detection into clinical reality faces a fundamental methodological challenge: The confounding influence of prevalent CVD. Diabetes rarely exists in isolation; it is a potent risk factor for and frequently coexists with conditions such as hypertension, coronary artery disease, heart failure, and arrhythmias like atrial fibrillation[11-13]. These CVD states themselves induce profound changes in the ECG signal, often overshadowing or mimicking the subtler alterations potentially attributable to DM per se. Consequently, models trained to detect DM using ECG features in mixed populations risk identifying underlying CVD rather than diabetes specifically, acting as a surrogate marker for a correlated outcome rather than the target pathology. This confounding effect significantly undermines the specificity and clinical utility of proposed ECG-based DM diagnostic algorithms and represents a critical barrier to their real-world implementation. Previous studies attempting ECG-based DM detection have often inadequa
Therefore, to advance the field beyond this significant limitation, a more nuanced approach is required - one that explicitly disentangles the electrophysiological signature of DM from the background noise of prevalent CVD. Phenoty
This study aimed to overcome the critical confounding influence of CVD on ECG-based DM detection by employing phenotypic clustering. Specifically, we sought to: (1) Identify distinct clinical phenotypes within a diverse patient cohort using comprehensive cardiovascular and demographic characteristics; (2) Determine the cluster(s) where DM exerts the most discernible influence on the single-lead ECG signal; and (3) Develop and validate cluster-specific ML models opti
A prospective observational non-randomized cross-sectional study was conducted at the I.M. Sechenov First Moscow State Medical University (Sechenov University), Moscow, Russia. The study protocol was approved by the Local Ethical Committee of Sechenov University (approval No. 19-23). The study was also registered on the ClinicalTrials.gov website (ID: NCT04788342). An informed written consent obtained from the study participants.
The study comprised two stages: The first stage involved the consecutive enrollment of 200 patients into the main sample according to the inclusion criteria specified below and the development of an algorithm for detecting DM using single-lead ECG. The second stage involved the additional enrollment of 100 patients into the test sample using the same inclusion criteria as the first stage, for the purpose of analyzing the effectiveness of the developed algorithm. A total of 629 patients’ parameters were included in the study. The proportion of patients with DM in both the training and test samples was required to exceed 30%.
Patients from University Clinical Hospital number 2 undergoing outpatient or inpatient examination and treatment, including in the endocrinology department with a diagnosis of DM, were included in the study. All patients reviewed and signed written informed consent to participate. The inclusion and exclusion criteria are presented in the Table 1.
| Criterion type | Description |
| Inclusion | Patient age over 18 years; agreement to participate in the study |
| Exclusion | Significant QRS complex morphology changes (such as bundle branch block and ventricular extrasystole); poor quality electrocardiography recorded from the fingers (Parkinson’s disease, tremor of any origin, mental disorders). Unsatisfactory quality of electrocardiography and/or photoplethysmography; withdrawal of consent for further participation in the study |
Clinical examination: Collection of complaints, auscultation, palpation, percussion, assessment of edema syndrome, measurement of heart rate and blood pressure level. Medical history analysis: Collection of cardiological history and DM history. The study adjusted medication use, including beta-blockers, angiotensin-converting enzyme inhibitors, sodium-glucose cotransporter 2 inhibitors, that could easily be responsible for the observed ECG changes.
Transthoracic echocardiography: Performed using a GE Vivid 5 device. Assessment of cardiac chamber volumes, wall thickness, diastolic and systolic function status, and detection of valvular apparatus and major cardiac vessel dysfunction according to current guidelines[14]. Coronary artery disease diagnosis obtained from the anamneses of the patient and the drugs that take for it is management.
Standard 12-lead ECG recording: Performed using a SCHILLER AT-5 device. Determination of temporal and amplitude ECG parameters, cardiac electrical axis direction, and assessment of cardiac cycle morphology.
Capillary blood glucose analysis: Fasting blood glucose level measured twice within 1-3 days, in a seated position, adhering to the following requirements: Analysis performed after a 12-hour fast; abstinence from alcohol 2-3 days prior; avoidance of stress or heavy physical exertion; refraining from tooth brushing, smoking, or consumption of fatty foods, sugary drinks, or confectionery products the day before. The diagnostic thresholds for DM are a fasting plasma glucose of 126 mg/dL (7 mmol/L) or higher and the presence of DM history. The type of the DM is classified based on the history of the patients, and the drug they take to it is management.
Single-lead ECG and photoplethysmography recording: Recorded using the Qardio-Qvark® device (LLC “L Card”, Moscow, Russia; registered with the Federal Service for Surveillance in Healthcare No. RZN 2019/8124 dated February 15, 2019) immediately after each blood glucose analysis. Recordings lasted 1 minute, performed at rest in a seated position in a blind manner. The recorder has a smartphone case form factor and features 2 sensors for recording ECG and photoplethysmography. Recordings were transmitted to a cloud-based platform for analysis using proprietary ML algorithms.
The Qardio-Qvark® embedded algorithms perform mathematical calculations obtained from the single lead ECG (lead I). Actually, these calculations are mathematical formulas and of importance to the mathematicians who build this algorithm. The algorithm shows 47 parameters that based on some mathematical features obtained from the single lead ECG, can be found on our previous publication[15]. The algorithm also, can predict, some disease, but further investigations are done to enhance its diagnostic accuracy[15].
Statistical analysis was performed using the R programming language (v4.3). For quantitative variables, the distribution type (using the Shapiro-Wilk test), mean, standard deviation, median, interquartile range (IQR), minimum, and maxi
Comparative analysis for normally distributed quantitative variables was performed using Welch’s t-test (for 2 groups) or analysis of variance (for more than 2 groups) with subsequent pairwise group comparisons; for non-normally distributed quantitative variables, the Mann-Whitney U test (for 2 groups) or the Kruskal-Wallis test (for more than 2 groups) was used. Comparative analysis of categorical and qualitative variables was performed using Pearson’s χ2 test, and, when not applicable, Fisher’s exact test. When adjusting for multiple comparisons, the Holm method was used.
Since MD is a background condition for many CVDs, there may be an issue of correlation between the presence of diabetes and changes observed on the ECG. To reduce the influence of confounding factors, the clinical and demographic data were clustered (using Ward’s method, with prior normalization of quantitative variables and binarization of categorical variables). These data were not used in model training, but were used to determine the corresponding cardiovascular phenotype of each individual. Firstly, this approach was aimed at identifying clusters of individuals without CVDs, or with mild stages of such diseases. Secondly, the prevalence of diabetes in each cluster was assessed.
The clusters were then used to attempt to build descriptive models based on sex, age, smoking status, rhythm disturbances during recording, and the single-lead ECG data itself. For validation, a specific cluster would be used, while all others would be used for training. In this way, it would be possible to test the feasibility of diagnosing diabetes among different cardiovascular phenotypes, as well as to identify factors with high importance that recur in two or more models.
In other words, validation would be performed for different phenotypes, and technically, instead of typical cross-validation, validation on clusters would be used. The training pipeline included normalization of quantitative variables and binarization of categorical variables. Model performance was evaluated on the validation cluster using area under the curve (AUC), sensitivity, specificity, positive and negative predictive values (with the optimal threshold determined by the Youden index on the training data). The input features intent to cluster data includes clinical features and the 47 features of the single lead ECG. The optimal number of clusters was determined by evaluating the dendrogram structure and the average silhouette width, with the goal of maximizing clinical interpretability and internal cohesion.
Prior to model training, a structured preprocessing pipeline was applied to the input features, which included both clinical variables and the 47 single-lead ECG parameters. Missing data were first assessed; variables with more than 5% missing values were excluded from the analysis, while those with fewer missing values were imputed using the median for continuous variables and the mode for categorical variables. Outliers in continuous ECG parameters were identified using the IQR method, where values less than Q1 - 1.5 × IQR or greater than Q3 + 1.5 × IQR were Winsorized (i.e., capped at the 5th and 95th percentiles) to reduce their influence without discarding data. Finally, all continuous variables were normalized to a mean of zero and a standard deviation of one to ensure features were on a comparable scale for the gradient boosting algorithm.
The 47 parameters derived from the single-lead ECG signal encompass a range of temporal, amplitude, and morphological features. For clarity, the physiological basis and calculation of key parameters discussed in the results are defined here: T-wave flatness index (Tfi): A quantitative measure of T-wave morphology, calculated as the ratio of the T-wave’s area to its amplitude. A lower value indicates a flatter, more rounded T-wave, which is associated with altered ventricular repolarization heterogeneity. Corrected QT (QTc) interval: The QT interval adjusted for heart rate using Bazett’s formula (QTc = QT/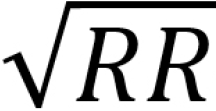 ), representing the duration of ventricular repolarization. QRS energy in band 4 (QRSE4): The spectral energy of the QRS complex within a specific high-frequency band (typically 40-100 Hz), reflecting the integrity and velocity of ventricular depolarization. High-frequency QRS (HFQRS): The root mean square voltage of the QRS complex in a high-frequency band (typically 150-250 Hz), sensitive to changes in myocardial conduction properties. Ventricular activation time (VAT): The interval from the onset of the QRS complex to the peak of the R-wave, representing the initial depolarization time of the ventricles. P-wave start time (Pst) and P-wave finish time: The onset and offset timings of the P-wave, defining the total duration of atrial depolarization.
), representing the duration of ventricular repolarization. QRS energy in band 4 (QRSE4): The spectral energy of the QRS complex within a specific high-frequency band (typically 40-100 Hz), reflecting the integrity and velocity of ventricular depolarization. High-frequency QRS (HFQRS): The root mean square voltage of the QRS complex in a high-frequency band (typically 150-250 Hz), sensitive to changes in myocardial conduction properties. Ventricular activation time (VAT): The interval from the onset of the QRS complex to the peak of the R-wave, representing the initial depolarization time of the ventricles. P-wave start time (Pst) and P-wave finish time: The onset and offset timings of the P-wave, defining the total duration of atrial depolarization.
A nested, cluster-stratified validation framework was employed to ensure robust model evaluation and mitigate overfitting. The overall dataset was first divided into the four predefined phenotypic clusters. For each iteration of the outer validation loop, one entire cluster was held out as the validation set. The remaining three clusters were combined to form the training set.
Within this training set, a 5-fold cross-validation process was implemented for hyperparameter tuning of the gradient boosting machine model. The hyperparameters yielding the best average performance across these 5 folds were then used to retrain the model on the entire training set (all three clusters). This final model was subsequently evaluated on the held-out validation cluster to estimate its generalizable performance on a distinct phenotypic group. This process was repeated four times, with each cluster serving as the validation set once, and the performance metrics were reported for each validation cluster individually.
The comparative features of the two groups of participants, with and without DM represented in the table below (Table 2). Due to the previously identified target variable likely reflects ECG alterations associated with underlying CVD, with DM serving as a comorbid factor (though this requires verification, which is methodologically challenging). Effec
| Factor | Index | Without DM | With DM | Р value |
| Age (years) | 56.1 ± 17.1 | 63.7 ± 10.6 | 0.000 | |
| Sex | Female | 177 (38.6) | 83 (48.5) | 0.032 |
| Left ventricle diastolic dysfunction | No | 333 (72.7) | 74 (43.3) | 0.000 |
| Yes | 125 (27.3) | 97 (56.7) | ||
| Left ventricle diastolic dysfunction_1 stage | No | 340 (74.2) | 84 (49.1) | 0.000 |
| Yes | 118 (25.8) | 87 (50.9) | ||
| Left ventricle diastolic dysfunction 2_3 stage | No | 418 (91.3) | 132 (77.2) | 0.000 |
| Yes | 40 (8.7) | 39 (22.8) | ||
| Hypertension degree | No | 190 (41.5) | 40 (23.4) | 0.000 |
| 1st degree | 39 (8.5) | 25 (14.6) | ||
| 2nd degree | 140 (30.6) | 50 (29.2) | ||
| 3rd degree | 89 (19.4) | 56 (32.7) | ||
| Aterial fibrillation at the registration moment (from the single lead ECG) | No | 435 (95.0) | 162 (94.7) | 1.000 |
| Yes | 23 (5.0) | 9 (5.3) | ||
| Ischemic heart disease | No | 361 (78.8) | 113 (66.1) | 0.001 |
| 1st functional class | 68 (14.8) | 48 (28.1) | ||
| 2nd functional class | 29 (6.3) | 10 (5.8) | ||
| Disturbance of cardiac rhythm and conduction | No | 443 (96.7) | 162 (94.7) | 0.092 |
| 1 | 11 (2.4) | 3 (1.8) | ||
| 2 | 3 (0.7) | 5 (2.9) | ||
| 3 | 1 (0.2) | 1 (0.6) | ||
| Chronic heart failure | No | 407 (89.6) | 150 (88.2) | 0.324 |
| 1st stage | 7 (1.5) | 0 (0.0) | ||
| 2nd stage | 22 (4.8) | 10 (5.9) | ||
| 3rd stage | 17 (3.7) | 9 (5.3) | ||
| 4th stage | 1 (0.2) | 1 (0.6) | ||
| Aortic valve insufficiency | No | 351 (76.6) | 147 (86.0) | 0.022 |
| 1st stage | 95 (20.7) | 23 (13.5) | ||
| 2nd stage | 12 (2.6) | 1 (0.6) | ||
| Aortic valve stenosis | No | 443 (96.7) | 163 (95.9) | 0.753 |
| 1st stage | 9 (2.0) | 5 (2.9) | ||
| 2nd stage | 4 (0.9) | 2 (1.2) | ||
| 3rd stage | 2 (0.4) | 0 (0.0) | ||
| Mitral valve insufficiency | No | 134 (29.5) | 105 (61.4) | 0.000 |
| 1st stage | 274 (60.4) | 60 (35.1) | ||
| 2nd stage | 39 (8.6) | 6 (3.5) | ||
| 3rd stage | 7 (1.5) | 0 (0.0) | ||
| Mitral valve stenosis | No | 454 (99.1) | 166 (97.1) | 0.053 |
| 1st stage | 3 (0.7) | 5 (2.9) | ||
| 2nd stage | 1 (0.2) | 0 (0.0) | ||
| Ejection fraction | 59.5 ± 8.6 | 58.1 ± 8.0 | 0.005 | |
| Ejection fraction > 55 | No | 73 (15.9) | 42 (24.6) | 0.018 |
| Yes | 385 (84.1) | 129 (75.4) | ||
| Ejection fraction < 40 | No | 436 (95.2) | 164 (95.9) | 0.870 |
| Yes | 22 (4.8) | 7 (4.1) | ||
| Ejection fraction < 30 | No | 453 (98.9) | 169 (98.8) | 1.000 |
| Yes | 5 (1.1) | 2 (1.2) | ||
| End diastolic volume (mL) | 93.8 ± 36.2 | 99.9 ± 46.7 | 0.632 | |
| Left atrial volume index | 34.9 ± 11.1 | 36.7 ± 8.1 | 0.000 | |
| Smoking | No | 371 (81.0) | 154 (90.1) | 0.009 |
| Yes | 87 (19.0) | 17 (9.9) | ||
| SDLA | 11.0 ± 5.2 | 10.4 ± 2.7 | 0.176 | |
| HFNoise | 60.7 ± 28.3 | 66.0 ± 25.5 | 0.003 | |
| RR | 822.0 ± 156.2 | 811.2 ± 141.7 | 0.533 | |
| TpTe | 89.4 ± 20.3 | 94.3 ± 29.7 | 0.487 | |
| VAT | 38.8 ± 11.1 | 41.6 ± 13.4 | 0.020 | |
| QTc | 425.8 ± 44.4 | 447.0 ± 52.4 | 0.000 | |
| QT_TQ | 1.0 ± 0.4 | 1.0 ± 0.4 | 0.001 | |
| HFQRS | 15919.9 ± 199679.8 | 10.3 ± 8.5 | 0.989 | |
| HFSNR | 2.5 ± 2.3 | 2.1 ± 1.8 | 0.149 | |
| JA | -24.9 ± 57.7 | -35.9 ± 81.8 | 0.054 | |
| J80A | 1504.0 ± 31942.7 | -3.8 ± 92.9 | 0.025 | |
| TA | 185.0 ± 153.9 | 151.2 ± 162.2 | 0.006 | |
| QRSenergy | 616.1 ± 557.9 | 728.6 ± 870.9 | 0.117 | |
| Tenergy | 108.4 ± 101.3 | 111.7 ± 113.7 | 0.926 | |
| Tpenergy | 77.0 ± 86.1 | 73.6 ± 89.1 | 0.218 | |
| Sbeta | 0.9 ± 0.5 | 1.0 ± 0.9 | 0.003 | |
| Beta | 0.8 ± 0.4 | 0.9 ± 0.5 | 0.000 | |
| QRS11energy | 539.0 ± 692.3 | 713.6 ± 1339.7 | 0.075 | |
| QRS12energy | 809.2 ± 1129.4 | 1074.5 ± 1901.7 | 0.077 | |
| QRS2energy | 226.5 ± 301.4 | 305.8 ± 551.4 | 0.042 | |
| QRSE1 | 4730.9 ± 94779.9 | 2707.4 ± 34733.0 | 0.004 | |
| QRSE2 | 1303.9 ± 25310.4 | 164.5 ± 290.3 | 0.040 | |
| QRSE3 | 224.6 ± 202.5 | 266.1 ± 323.5 | 0.177 | |
| QRSE4 | 230.8 ± 189.8 | 238.7 ± 194.6 | 0.454 | |
| TE1 | 45.2 ± 57.1 | 54.0 ± 72.5 | 0.263 | |
| TE2 | 2548.4 ± 53784.0 | 31.7 ± 27.2 | 0.186 | |
| TE3 | 3621.5 ± 77261.8 | 9.5 ± 9.2 | 0.019 | |
| TE4 | 10275.1 ± 134000.1 | 5.2 ± 5.1 | 0.031 | |
| QRSw | 88.7 ± 23.5 | 92.3 ± 25.9 | 0.078 | |
| PAn | 67.3 ± 37.6 | 60.5 ± 40.1 | 0.081 | |
| Pan_1 | -1.4 ± 17.6 | -1.6 ± 10.3 | 0.981 | |
| RA | 903.0 ± 414.2 | 916.9 ± 432.6 | 0.794 | |
| SA | 180.0 ± 155.1 | 198.7 ± 209.0 | 0.632 | |
| Pst | 217.6 ± 121.4 | 144.2 ± 123.3 | 0.000 | |
| Pfi | 77.3 ± 77.0 | 101.5 ± 79.9 | 0.000 | |
| QRSst | 165.6 ± 85.6 | 190.7 ± 88.4 | 0.000 | |
| QRSfi | 436.2 ± 117.0 | 371.1 ± 169.8 | 0.000 | |
| Tfi | 5.5 ± 170.3 | 293.1 ± 301.6 | 0.000 | |
| PpeakP | -67.5 ± 55.2 | -7.2 ± 71.5 | 0.000 | |
| PpeakP | 107.9 ± 81.7 | 80.7 ± 106.0 | 0.000 | |
| Rpeak | 138.4 ± 92.7 | 197.8 ± 94.6 | 0.000 | |
| Speak | 176.0 ± 109.5 | 223.3 ± 102.9 | 0.000 | |
| Tpeak | 375.6 ± 106.9 | 453.2 ± 123.7 | 0.000 | |
| Tons | 329.7 ± 105.4 | 403.1 ± 118.7 | 0.000 | |
| Toffs | 416.8 ± 113.5 | 493.4 ± 126.5 | 0.000 | |
| RonsF | 34.1 ± 4.7 | 33.0 ± 5.8 | 0.030 | |
| RoffsF | 33.2 ± 6.0 | 32.1 ± 6.8 | 0.078 | |
| SDNN | 2776.7 ± 58807.6 | 25.9 ± 19.1 | 0.030 |
To address this issue a methodological framework performed through the identification of phenotypic subgroups stratified by DM status. Were the entire dataset was clustered using the following baseline clinical and demographic variables - representing all available parameters (notably limited in scope): Age, sex, diastolic dysfunction, hypertension, atrial fibrillation, coronary artery disease, conduction disorders, chronic heart failure, valve stenosis/insufficiency, ejection fraction, end-diastolic volume, obesity, DM, smoking status, and pulmonary artery systolic pressure.
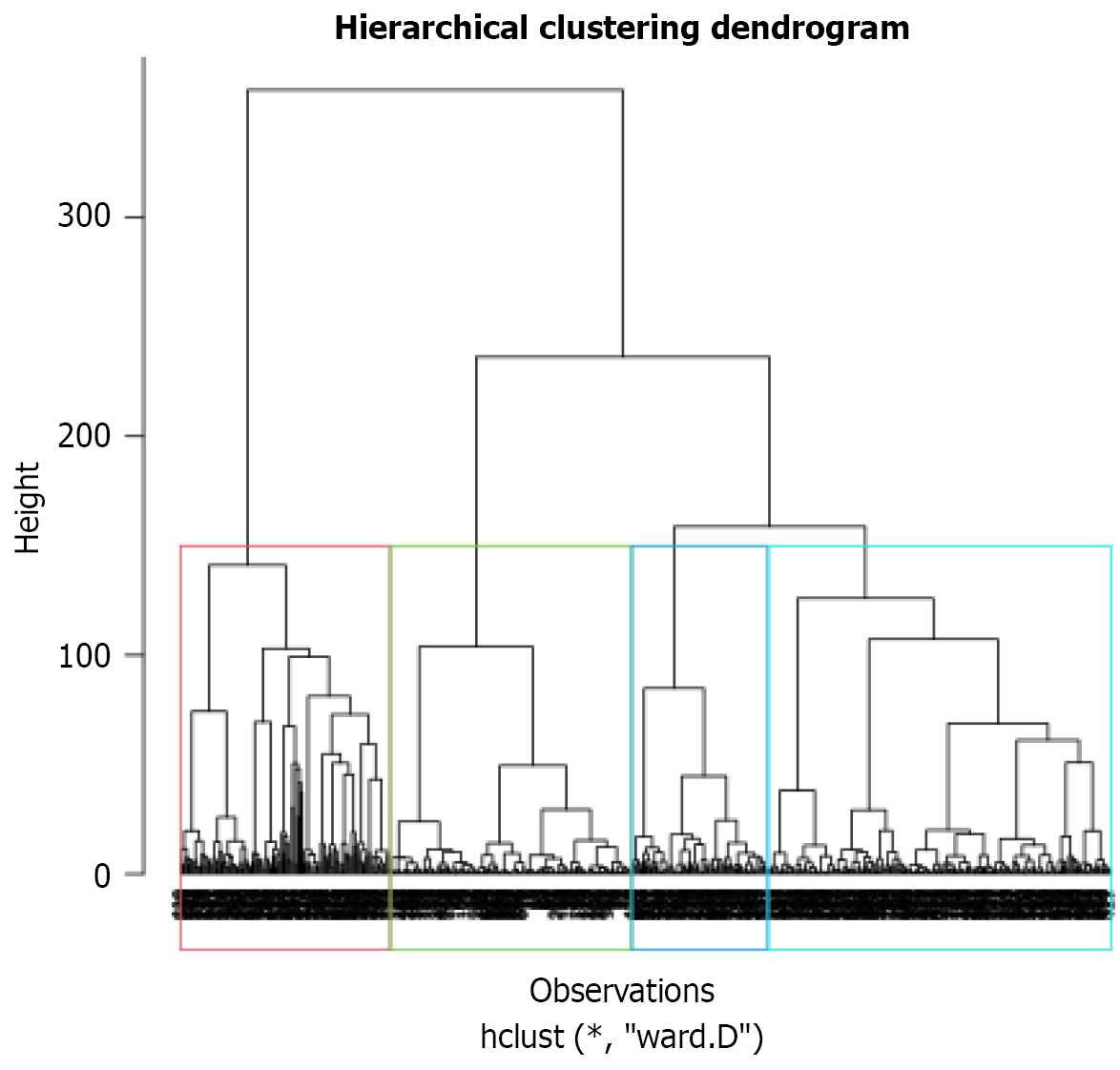
Four distinct clusters emerged: Cluster 1: Younger, predominantly healthy individuals. Cluster 2: Older, predominantly healthy individuals. Clusters 3 and 4: Patients with significant comorbidities. These clusters define four phenotypic patient profiles. The average silhouette width for this 4-cluster solution was 0.32, indicating a reasonable and meaningful structure in the data (Figure 1). Comparative analyses and pairwise statistics are detailed in the below table, for full inter-cluster analysis in Table 3 and Supplementary Table 1.
| Factor | Statistic | Cluster 1 | Cluster 2 | Cluster 3 | Cluster 4 | P value |
| Age (years) | 43.1 ± 16.2 | 62.1 ± 12.6 | 64.8 ± 12.2 | 64.8 ± 10.3 | 0.000 | |
| Sex | Female | 72 (44.2) | 112 (48.3) | 53 (37.3) | 23 (25.0) | 0.001 |
| Male | 91 (55.8) | 120 (51.7) | 89 (62.7) | 69 (75.0) | ||
| Left ventricle diastolic dysfunction | No | 146 (89.6) | 166 (71.6) | 51 (35.9) | 44 (47.8) | 0.000 |
| Yes | 17 (10.4) | 66 (28.4) | 91 (64.1) | 48 (52.2) | ||
| Atrial fibrillation at the time of single lead ECG registration | No | 163 (100.0) | 226 (97.4) | 119 (83.8) | 89 (96.7) | 0.000 |
| Yes | 0 (0.0) | 6 (2.6) | 23 (16.2) | 3 (3.3) | ||
| Ejection fraction (%) | 61.4 ± 5.7 | 61.5 ± 5.1 | 52.9 ± 12.1 | 58.7 ± 7.7 | 0.000 | |
| End diastolic volume (mL) | 81.7 ± 24.4 | 87.7 ± 24.0 | 113.1 ± 51.4 | 101.0 ± 42.3 | 0.000 | |
| Left atrial volume index | 30.3 ± 5.6 | 34.4 ± 7.8 | 42.3 ± 15.3 | 36.1 ± 6.9 | 0.000 | |
| Diabetes mellitus | No | 131 (80.4) | 163 (70.3) | 108 (76.1) | 56 (60.9) | 0.005 |
| Yes | 32 (19.6) | 69 (29.7) | 34 (23.9) | 36 (39.1) | ||
| Smoking | No | 137 (84.0) | 200 (86.2) | 114 (80.3) | 74 (80.4) | 0.398 |
| Yes | 26 (16.0) | 32 (13.8) | 28 (19.7) | 18 (19.6) | ||
| Pulmonary artery systolic pressure | 10.0 ± 0.0 | 10.0 ± 0.0 | 13.6 ± 9.3 | 10.0 ± 0.0 | 0.000 | |
| Hypertension_0 | No | 0 (0.0) | 211 (90.9) | 113 (79.6) | 75 (81.5) | 0.000 |
| Yes | 163 (100.0) | 21 (9.1) | 29 (20.4) | 17 (18.5) | ||
| Hypertension_1 | No | 163 (100.0) | 178 (76.7) | 138 (97.2) | 86 (93.5) | 0.000 |
| Yes | 0 (0.0) | 54 (23.3) | 4 (2.8) | 6 (6.5) | ||
| Hypertension_2 | No | 163 (100.0) | 100 (43.1) | 102 (71.8) | 74 (80.4) | 0.000 |
| Yes | 0 (0.0) | 132 (56.9) | 40 (28.2) | 18 (19.6) | ||
| Hypertension_3 | No | 163 (100.0) | 207 (89.2) | 73 (51.4) | 41 (44.6) | 0.000 |
| Yes | 0 (0.0) | 25 (10.8) | 69 (48.6) | 51 (55.4) | ||
| Ischemic heart disease_0 | No | 0 (0.0) | 1 (0.4) | 63 (44.4) | 91 (98.9) | 0.000 |
| Yes | 163 (100.0) | 231 (99.6) | 79 (55.6) | 1 (1.1) | ||
| Ischemic heart disease_1 | No | 163 (100.0) | 231 (99.6) | 93 (65.5) | 26 (28.3) | 0.000 |
| Yes | 0 (0.0) | 1 (0.4) | 49 (34.5) | 66 (71.7) | ||
| Ischemic heart disease_2 | No | 163 (100.0) | 232 (100.0) | 128 (90.1) | 67 (72.8) | 0.000 |
| Yes | 0 (0.0) | 0 (0.0) | 14 (9.9) | 25 (27.2) | ||
| Disturbance of cardiac rhythm and conduction_0 | No | 0 (0.0) | 0 (0.0) | 24 (16.9) | 0 (0.0) | 0.000 |
| Yes | 163 (100.0) | 232 (100.0) | 118 (83.1) | 92 (100.0) | ||
| Disturbance of cardiac rhythm and conduction_1 | No | 163 (100.0) | 232 (100.0) | 128 (90.1) | 92 (100.0) | 0.000 |
| Yes | 0 (0.0) | 0 (0.0) | 14 (9.9) | 0 (0.0) | ||
| Disturbance of cardiac rhythm and conduction_2 | No | 163 (100.0) | 232 (100.0) | 134 (94.4) | 92 (100.0) | 0.000 |
| Yes | 0 (0.0) | 0 (0.0) | 8 (5.6) | 0 (0.0) | ||
| Disturbance of cardiac rhythm and conduction_3 | No | 163 (100.0) | 232 (100.0) | 140 (98.6) | 92 (100.0) | 0.069 |
| Yes | 0 (0.0) | 0 (0.0) | 2 (1.4) | 0 (0.0) | ||
| Chronic heart failure_0 | No | 0 (0.0) | 0 (0.0) | 67 (48.6) | 0 (0.0) | 0.000 |
| Yes | 163 (100.0) | 231 (100.0) | 71 (51.4) | 92 (100.0) | ||
| Chronic heart failure_1 | No | 163 (100.0) | 231 (100.0) | 131 (94.9) | 92 (100.0) | 0.000 |
| Yes | 0 (0.0) | 0 (0.0) | 7 (5.1) | 0 (0.0) | ||
| Chronic heart failure_2 | No | 163 (100.0) | 231 (100.0) | 106 (76.8) | 92 (100.0) | 0.000 |
| Yes | 0 (0.0) | 0 (0.0) | 32 (23.2) | 0 (0.0) | ||
| Chronic heart failure_3 | No | 163 (100.0) | 231 (100.0) | 112 (81.2) | 92 (100.0) | 0.000 |
| Yes | 0 (0.0) | 0 (0.0) | 26 (18.8) | 0 (0.0) | ||
| Chronic heart failure_4 | No | 163 (100.0) | 231 (100.0) | 136 (98.6) | 92 (100.0) | 0.063 |
| Yes | 0 (0.0) | 0 (0.0) | 2 (1.4) | 0 (0.0) | ||
| Aortic valve stenosis_0 | No | 0 (0.0) | 55 (23.7) | 59 (41.5) | 17 (18.5) | 0.000 |
| Yes | 163 (100.0) | 177 (76.3) | 83 (58.5) | 75 (81.5) | ||
| Aortic valve insufficiency_1 | No | 163 (100.0) | 177 (76.3) | 96 (67.6) | 75 (81.5) | 0.000 |
| Yes | 0 (0.0) | 55 (23.7) | 46 (32.4) | 17 (18.5) | ||
| Aortic valve insufficiency_2 | No | 163 (100.0) | 232 (100.0) | 129 (90.8) | 92 (100.0) | 0.000 |
| Yes | 0 (0.0) | 0 (0.0) | 13 (9.2) | 0 (0.0) | ||
| Aortic valve stenosis_0 | No | 0 (0.0) | 0 (0.0) | 22 (15.6) | 0 (0.0) | 0.000 |
| Yes | 163 (100.0) | 232 (100.0) | 119 (84.4) | 92 (100.0) | ||
| Aortic valve stenosis_1 | No | 163 (100.0) | 232 (100.0) | 127 (90.1) | 92 (100.0) | 0.000 |
| Yes | 0 (0.0) | 0 (0.0) | 14 (9.9) | 0 (0.0) | ||
| Aortic valve stenosis_2 | No | 163 (100.0) | 232 (100.0) | 135 (95.7) | 92 (100.0) | 0.000 |
| Yes | 0 (0.0) | 0 (0.0) | 6 (4.3) | 0 (0.0) | ||
| Aortic valve stenosis_3 | No | 163 (100.0) | 232 (100.0) | 139 (98.6) | 92 (100.0) | 0.078 |
| Yes | 0 (0.0) | 0 (0.0) | 2 (1.4) | 0 (0.0) | ||
| Mitral valve insufficiency_0 | No | 59 (36.2) | 145 (62.5) | 121 (87.7) | 61 (66.3) | 0.000 |
| Yes | 104 (63.8) | 87 (37.5) | 17 (12.3) | 31 (33.7) | ||
| Mitral valve insufficiency_1 | No | 104 (63.8) | 102 (44.0) | 52 (37.7) | 33 (35.9) | 0.000 |
| Yes | 59 (36.2) | 130 (56.0) | 86 (62.3) | 59 (64.1) | ||
| Mitral valve insufficiency_2 | No | 163 (100.0) | 217 (93.5) | 110 (79.7) | 90 (97.8) | 0.000 |
| Yes | 0 (0.0) | 15 (6.5) | 28 (20.3) | 2 (2.2) | ||
| Mitral valve insufficiency_3 | No | 163 (100.0) | 232 (100.0) | 131 (94.9) | 92 (100.0) | 0.000 |
| Yes | 0 (0.0) | 0 (0.0) | 7 (5.1) | 0 (0.0) | ||
| Mitral valve stenosis_0 | No | 0 (0.0) | 0 (0.0) | 9 (6.3) | 0 (0.0) | 0.000 |
| Yes | 163 (100.0) | 232 (100.0) | 133 (93.7) | 92 (100.0) | ||
| Mitral valve stenosis_1 | No | 163 (100.0) | 232 (100.0) | 134 (94.4) | 92 (100.0) | 0.000 |
| Yes | 0 (0.0) | 0 (0.0) | 8 (5.6) | 0 (0.0) | ||
| Mitral valve stenosis_2 | No | 163 (100.0) | 232 (100.0) | 141 (99.3) | 92 (100.0) | 0.356 |
| Yes | 0 (0.0) | 0 (0.0) | 1 (0.7) | 0 (0.0) |
Four gradient boosting models were trained using cluster-stratified cross-validation: Training: Three clusters. Validation: The rest cluster. Features: Age, sex, atrial fibrillation at recording, smoking status, single-lead ECG parameters. The models performance metrics represented in Table 4. Key observations include optimal DM detection occurred in cluster 1 (younger/healthy cohort), cluster 3 exhibited the poorest performance, and the cluster 4 model was selected as the most balanced (Figure 1). The top 5 important features for each cluster with the importance coefficient in the diagnosis of DM represented below, for full feature importance in Table 5 and Supplementary Tables 2-5.
| Validation cluster | Area under the receiver operating characteristic curve | Sensitivity | Specificity | Positive predictive value | Negative predictive value | Threshold |
| Cluster 1 | 0.947 | 0.969 | 0.781 | 0.948 | 0.862 | 0.350 |
| Cluster 2 | 0.835 | 0.816 | 0.725 | 0.875 | 0.625 | 0.221 |
| Cluster 3 | 0.589 | 0.398 | 0.941 | 0.956 | 0.330 | 0.131 |
| Cluster 4 | 0.880 | 0.750 | 0.833 | 0.875 | 0.682 | 0.164 |
| Cluster 1; feature (importance coefficient) | Cluster 2; feature (importance coefficient) | Cluster 3; feature (importance coefficient) | Cluster 4; feature (importance coefficient) |
| Tfi (0.203) | QRSfi (0.206) | Tfi (0.593) | Tfi (0.235) |
| HFQRS (0.043) | Age (0.120) | QTc (0.053) | Age (0.076) |
| RR (0.041) | Toffs (0.106) | Age (0.049) | PpeakN (0.051) |
| Beta (0.033) | QRSE1 (0.053) | Sbeta (0.020) | Rpeak (0.043) |
| QRSE4 (0.032) | QTc (0.040) | QRSE4 (0.019) | Pst (0.035) |
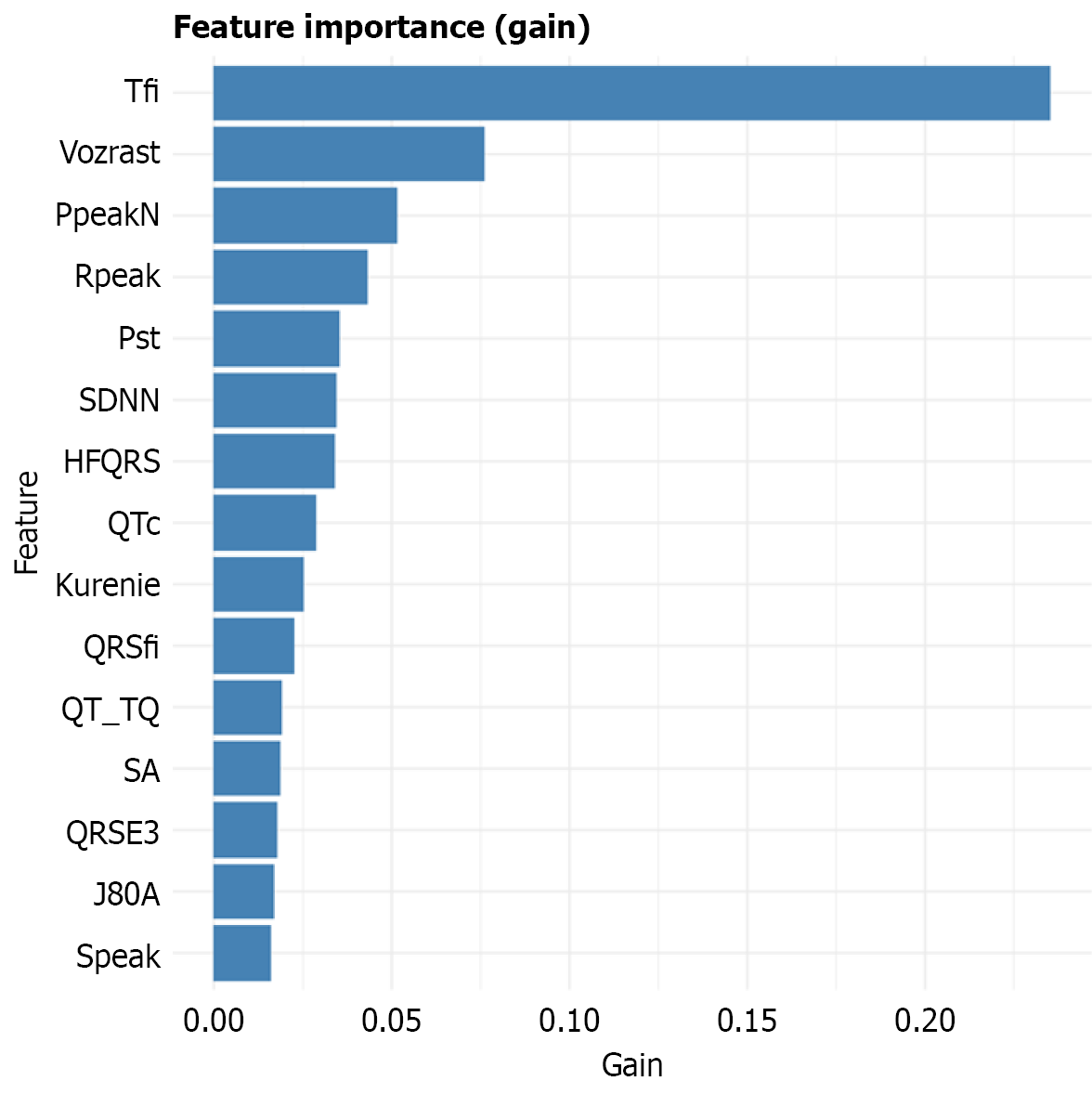
Tfi reflecting impaired ventricular repolarization, is the top feature in clusters 1 (young/healthy), 3 (severe CVD), and 4 (high-DM). Its importance is high in clusters 1 and 4 where DM drives changes, but low in cluster 3 where CVD dominates (Figure 2). Age highlights age-dependent remodeling and is important in cluster 2 (older/healthy), but less relevant in clusters 3 and 4 due to comorbidities. QTc, indicating repolarization issues, shows moderate importance in cluster 2 but is confounded by CVD in cluster 3. QRSE4, suggesting conduction alterations from fibrosis, is key in low-CVD cluster 1 but masked in CVD-heavy cluster 3.
The recurrence of features like Tfi and QTc confirms their link to DM pathophysiology (e.g., neuropathy, fibrosis). However, their diagnostic power is phenotype-dependent, rendering universal screening models ineffective. Stratification is essential - for example, Tfi + atrial features [P-wave amplitude (PpeakN), Pst] achieve optimal performance (AUC: 0.88) in cluster 4.
Tfi is a universal DM biomarker but is clinically reliable only when DM (not CVD) primarily drives ECG changes. Cluster 4 (high DM, moderate CVD) is ideal for ECG-based DM screening due to clear DM-specific feature expression. Unique features add critical insights: Atrial changes (PpeakN, Pst) in cluster 4 imply early DM impact, while QRS complex morphology dominance in cluster 2 may mimic DM effects.
The present study employed an innovative clustering approach to address a fundamental methodological challenge in detecting DM via single-lead ECG: The confounding influence of underlying CVD. Our initial analysis revealed that ECG alterations likely reflected broader CVD pathology rather than DM-specific signatures, as DM frequently presents as a comorbid condition rather than an isolated factor. This necessitated a paradigm shift beyond conventional DM vs non-DM comparisons. By clustering the entire cohort based on comprehensive clinical and demographic profiles, we identified four distinct phenotypic subgroups: Younger healthy individuals (cluster 1), older healthy individuals (cluster 2), and two groups with significant comorbidities but divergent profiles (clusters 3 and 4). This stratification proved crucial for disentangling the complex interplay between DM and CVD and for developing targeted diagnostic algorithms.
Cluster 4 emerged as the most informative subgroup for investigating DM-specific ECG signatures. While both clusters 3 and 4 comprised patients with substantial comorbidity burdens, cluster 4 was distinguished by the highest prevalence of DM (39.1%), a predominance of males (75.0%), a relatively preserved mean ejection fraction (58.7% ± 7.7%), and moderate levels of left ventricular diastolic dysfunction (52.2%) and valvular pathologies (e.g., mitral stenosis categories). Crucially, compared to cluster 3, cluster 4 exhibited less pronounced cardiac structural remodeling (lower mean end-diastolic volume: 101.0 ± 42.3 mL vs 113.1 ± 51.4 mL) and a lower prevalence of severe heart failure (chronic heart failure_2/3/4). This profile suggests cluster 4 represents patients where DM is a major contributing factor within a context of significant, but not maximally severe, CVD, potentially allowing DM-related electrophysiological changes to manifest more distinctly than in cluster 3, where advanced structural heart disease may dominate the ECG signal.
The cluster-stratified model validation further underscored the significance of phenotype-specific analysis and the relevance of cluster 4. While the model trained on cluster 1 (younger/healthy) achieved the highest AUC (0.947), its clinical utility for DM screening in broader, higher-risk populations is likely limited. Conversely, the model for cluster 3 performed poorly (AUC: 0.589), likely because severe coexisting CVD pathologies masked or overwhelmed any specific DM-related ECG features. The cluster 4 model, however, demonstrated a balanced and robust performance (AUC: 0.880, sensitivity: 0.750, specificity: 0.833, positive predictive value: 0.875, negative predictive value: 0.682), making it the most suitable candidate for real-world application in a comorbid population. Feature importance analysis within cluster 4 revealed that Tfi was the most influential parameter, followed by age, PpeakN, R-wave amplitude, and Pst. This contrasts sharply with other clusters; for instance, QRS complex morphology was dominant in cluster 2, while Tfi was also top in cluster 3 but with drastically different performance, reinforcing the context-dependence of ECG feature interpretation. The prominence of repolarization (Tfi) and atrial activity (PpeakN, Pst) features in cluster 4 aligns with known DM pathophysiology, including autonomic neuropathy and subtle conduction system alterations.
Our findings, which highlight the importance of repolarization (Tfi) and atrial conduction (PpeakN, Pst) features, resonate with - yet significantly extend - previous research on ECG alterations in DM. Traditional 12-lead ECG studies have long reported associations between DM and repolarization abnormalities, but with considerable inconsistency. For instance, while some studies in relatively healthy diabetic cohorts have noted flattened T-waves and altered repolarization heterogeneity[16,17], others in more advanced disease states have reported prolonged QRS duration and QTc intervals[18,19]. This inconsistency likely stems from the confounding effect of varying degrees of comorbid CVD, which our phenotypic stratification explicitly addresses. Studies that attempted to link QTc prolongation to DM often found it was more pronounced in patients with complications or severe hypoglycemia[20,21], suggesting it may be a marker of advanced disease rather than early detection. In contrast, our model identified Tfi - a more nuanced measure of T-wave morphology - as a top feature in a comorbid but structurally compensated cohort (cluster 4), suggesting it may capture subtler, earlier repolarization changes before they manifest as overt QTc prolongation. Similarly, the significance of atrial parameters like PpeakN and Pst in our model hints at early electropathology related to diabetic autonomic neuropathy impacting the atria, a dimension less frequently explored in conventional 12-lead analyses. Therefore, our work moves beyond these historical inconsistencies by demonstrating that when analyzed with ML and appropriate phenotypic context, single-lead ECG can distill a coherent DM-specific signature from previously deemed ‘non-specific’ findings.
Beyond the established parameters like QTc and T-wave morphology, our model identified several high-dimensional features, such as HFQRS and QRSE4, as important discriminators. While these lack direct traditional ECG correlates, their importance can be interpreted within the known pathophysiology of DM. HFQRS are known to reflect the integrity of ventricular depolarization; their reduction is associated with slowed conduction velocity and fragmentation of the electrical wavefront, often caused by myocardial fibrosis or ischemia. QRSE4 similarly captures alterations in the VAT sequence. In the context of DM, chronic hyperglycemia promotes the accumulation of advanced glycation end-products and reactive oxygen species, leading to diffuse interstitial fibrosis and electrical remodeling of the myocardium. This subclinical cardiomyopathy manifests as conduction heterogeneity, which ML models can detect as changes in the high-frequency energy and complexity of the QRS complex. Thus, the prominence of HFQRS and QRSE4 in our models provides a plausible, data-driven link to the fibrotic and conductive tissue changes that constitute the diabetic heart’s electrophysiological substrate, often preceding overt systolic dysfunction.
While the model for cluster 1 achieved the highest AUC (0.947), its exceptional performance must be interpreted within its clinical context. Cluster 1 comprises younger, predominantly healthy individuals with a low burden of CVD. In this low-prevalence, low-complexity population, the clinical utility of a DM screening tool is inherently limited, as the pre-test probability of disease is low and the incremental value over routine care is questionable. The model’s near-perfect performance in this cohort, while methodologically validating our approach, may also be less generalizable to real-world screening scenarios that typically target older, higher-risk populations. In contrast, we deliberately selected the cluster 4 model (AUC: 0.880) as optimal because it addresses a far more pressing clinical need: Accurate DM identification within a complex, comorbid patient profile where traditional risk factors are entangled and the pre-test probability is high. The robust performance of the cluster 4 model in this challenging context - characterized by a high prevalence of DM (39.1%), hypertension, and other structural heart diseases - demonstrates its potential for tangible clinical impact. It is precisely in these complex patients, often encountered in cardiology and primary care settings, that a rapid, non-invasive tool for detecting occult DM or pre-diabetes would be most valuable for guiding further diagnostic testing and intensifying preventive management.
These findings carry significant implications. Firstly, they strongly argue against a “one-size-fits-all” approach to ECG-based DM detection. Phenotypic stratification, as achieved through clustering, is essential to account for the modifying effects of age and comorbid CVD on the ECG. Secondly, cluster 4 appears to encapsulate a patient phenotype where DM exerts a detectable, modulatory influence on the ECG amidst prevalent CVD. The robust performance of the cluster 4-specific model suggests that single-lead ECG holds promise for DM identification within this specific, clinically relevant subgroup characterized by significant comorbidity burden where DM is a prominent factor. Future research should focus on validating this cluster 4 model in external cohorts with similar phenotypic profiles and exploring longitudinal outcomes. Refining the feature set, potentially incorporating additional photoplethysmography-derived parameters or serial measurements, may further enhance performance. Ultimately, this phenotype-targeted strategy offers a more realistic and potentially effective pathway towards implementing ECG-based DM screening tools in complex clinical settings.
The changes in the ECG in patients with type 2 DM on the 12-lead ECG considered not specific. Consistent research findings indicate that patients with DM without cardiovascular complications exhibit several electrocardiographic abnor
Notably, the ECG-only model’s performance is striking given the historical view that ECG changes in DM are “non-specific”. Prior studies using 12-lead ECGs reported inconsistent alterations (e.g., QRS shortening vs prolongation, variable QT dynamics), often attributed to advanced cardiovascular complications[24-27]. Our results challenge this paradigm, indicating that single-lead ECG biomarkers - when analyzed via ML - can detect subclinical autonomic dysfunction before overt symptoms arise. This aligns with recent evidence linking DM to prolonged VAT and repolarization heterogeneity (Tfi), driven by hyperglycemia-induced fibrosis and ion channel remodeling[28].
The ECG parameters predictive in our model reflect early cardiac autonomic neuropathy, a well-established complication of DM[15,29,30]. For instance, Tfi and reduced Tpeak correlate with sympathetic denervation, diminishing T-wave amplitude[31,32]. Similarly, increased QRSE4 signifies aberrant conduction velocity due to myocardial collagen deposition, while VAT prolongation aligns with delayed depolarization from hyperglycemia-induced sodium channel dysfunction. These findings support the hypothesis that cardiac electrophysiological changes precede structural damage, positioning ECG as a viable biomarker for early DM detection. Our results corroborate Kittnar’s observations of repolarization abnormalities in DM patients without CVD, while extending them to single-lead applications[1].
Feature repetition underscores shared DM pathophysiology, but diagnostic utility is context-dependent. Phenotypic clustering (e.g., prioritizing cluster 4) prevents misattributing CVD-driven ECG changes to DM, enabling accurate screening. Future work should validate findings externally and explore causal relationships through longitudinal studies.
The Qardio-Qvark® device can be used as a screening tool for early detection of DM. The advantages of using of this tool include cost effective, easy to use and can be implemented in the outpatient department as well as in-patients department in addition to at home use. The patient can record the single lead ECG during one minute and get the results immediately and if some alterations detected in the obtained results that generated by the algorithms implemented in the device, further, the results can be sent to the responsible physician and consulted accordingly. Moreover, the device is currently taught to diagnose several conditions such as, heart failure in both types systolic and diastolic types, ischemic heart disease, arrythmias, and diastolic dysfunction. Therefore, it is more comfortable to do general population screening for multiple organs from a single minute recording a single lead ECG.
Several limitations of this study must be acknowledged. First, the single-center design at a tertiary university hospital in Moscow and the homogeneous Russian cohort limit the generalizability of our findings. The patient population, clinical practices, and genetic background may differ from other healthcare settings and ethnic groups, potentially affecting the performance of our phenotype-stratified model. External validation in multi-center, international, and more diverse populations is essential to confirm the robustness and transportability of our approach.
Second, the representativeness of our sample must be considered. As a hospital-based cohort, our participants likely have a higher burden of comorbidities and more advanced disease stages than the general population or a primary care setting. This spectrum bias means our model, particularly the high-performing cluster 4 algorithm, is optimized for a comorbid, high-prevalence population and may not perform as well in a healthier, community-based screening context.
Third, our methodology is inherently tied to the specific hardware and software of the Qardio-Qvark® device. The 47 ECG parameters used for model training are derived from proprietary algorithms. This device-specificity means our model is not directly applicable to raw ECG signals from other single-lead or standard 12-lead ECG systems without significant recalibration and validation. Furthermore, while we excluded recordings with poor quality, the finger-based form factor remains susceptible to motion artifacts and noise, which could influence parameter extraction.
Fourth, the clustering was based on available clinical and demographic variables; unmeasured confounders (e.g., detailed socioeconomic status, specific dietary habits, or genetic predispositions) may influence phenotypic stratification and introduce residual confounding. Fifth, the cross-sectional design precludes assessment of temporal ECG changes or DM progression. Finally, while the overall sample size is substantial, the subsequent division into four clusters resulted in modest subgroup sizes (e.g., cluster 4, n = 92), which may reduce the statistical power for cluster-specific analyses and increase the risk of overfitting, despite our use of robust cross-validation techniques. Further investigations required to confirm the clinical validity of the built ML model and external validation is the cornerstone for future studies.
This study demonstrates that a phenotype-stratified approach is crucial for effective ECG-based DM detection, with the model for cluster 4 (high DM prevalence, significant comorbidities) achieving robust performance (AUC: 0.880). This confirms that single-lead ECG can accurately identify DM within a specific, clinically relevant high-risk phenotype where DM is a prominent driver of electrophysiological changes.
The clinical implementability of this strategy is a key advantage. By utilizing a low-cost, portable device that requires only a one-minute recording, our approach moves beyond proof-of-concept towards a practical screening solution. This makes it particularly suitable for resource-limited primary care settings, community health programs, and even remote patient monitoring. It can act as a high-throughput triage tool, enabling healthcare workers to identify at-risk individuals efficiently and direct confirmatory testing (like HbA1c or fasting plasma glucose) more strategically. By prioritizing a high-risk phenotype, this strategy maximizes the positive predictive value of screening, offering a scalable and cost-effective method to improve early DM detection in diverse healthcare environments. The need for future multi-center validation is important to confirm the clinical validity of our findings.
| 1. | Pan C, Cao B, Fang H, Liu Y, Zhang S, Luo W, Wu Y. Global burden of diabetes mellitus 1990-2021: epidemiological trends, geospatial disparities, and risk factor dynamics. Front Endocrinol (Lausanne). 2025;16:1596127. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 16] [Article Influence: 16.0] [Reference Citation Analysis (0)] |
| 2. | International Diabetes Federation. IDF Diabetes Atlas 2025. [cited 3 August 2025]. Available from: https://diabetesatlas.org/resources/idf-diabetes-atlas-2025/. |
| 3. | Islam K, Islam R, Nguyen I, Malik H, Pirzadah H, Shrestha B, Lentz IB, Shekoohi S, Kaye AD. Diabetes Mellitus and Associated Vascular Disease: Pathogenesis, Complications, and Evolving Treatments. Adv Ther. 2025;42:2659-2678. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 42] [Cited by in RCA: 40] [Article Influence: 40.0] [Reference Citation Analysis (0)] |
| 4. | Wu H, Norton V, Cui K, Zhu B, Bhattacharjee S, Lu YW, Wang B, Shan D, Wong S, Dong Y, Chan SL, Cowan D, Xu J, Bielenberg DR, Zhou C, Chen H. Diabetes and Its Cardiovascular Complications: Comprehensive Network and Systematic Analyses. Front Cardiovasc Med. 2022;9:841928. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 31] [Article Influence: 7.8] [Reference Citation Analysis (1)] |
| 5. | Lu Y, Wang W, Liu J, Xie M, Liu Q, Li S. Vascular complications of diabetes: A narrative review. Medicine (Baltimore). 2023;102:e35285. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 101] [Cited by in RCA: 72] [Article Influence: 24.0] [Reference Citation Analysis (1)] |
| 6. | Sacks DB, Arnold M, Bakris GL, Bruns DE, Horvath AR, Kirkman MS, Lernmark A, Metzger BE, Nathan DM; National Academy of Clinical Biochemistry; Evidence-Based Laboratory Medicine Committee of the American Association for Clinical Chemistry. Guidelines and recommendations for laboratory analysis in the diagnosis and management of diabetes mellitus. Diabetes Care. 2011;34:e61-e99. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 345] [Cited by in RCA: 318] [Article Influence: 21.2] [Reference Citation Analysis (1)] |
| 7. | Jabara M, Kose O, Perlman G, Corcos S, Pelletier MA, Possik E, Tsoukas M, Sharma A. Artificial Intelligence-Based Digital Biomarkers for Type 2 Diabetes: A Review. Can J Cardiol. 2024;40:1922-1933. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 10] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 8. | Mackenzie SC, Sainsbury CAR, Wake DJ. Diabetes and artificial intelligence beyond the closed loop: a review of the landscape, promise and challenges. Diabetologia. 2024;67:223-235. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 72] [Cited by in RCA: 37] [Article Influence: 18.5] [Reference Citation Analysis (0)] |
| 9. | Shapiro MR, Tallon EM, Brown ME, Posgai AL, Clements MA, Brusko TM. Leveraging artificial intelligence and machine learning to accelerate discovery of disease-modifying therapies in type 1 diabetes. Diabetologia. 2025;68:477-494. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 8] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 10. | Sriram RD, Reddy SSK. Artificial Intelligence and Digital Tools: Future of Diabetes Care. Clin Geriatr Med. 2020;36:513-525. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 27] [Cited by in RCA: 16] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 11. | Khokhar PB, Gravino C, Palomba F. Advances in artificial intelligence for diabetes prediction: insights from a systematic literature review. Artif Intell Med. 2025;164:103132. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 14] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 12. | Guan Z, Li H, Liu R, Cai C, Liu Y, Li J, Wang X, Huang S, Wu L, Liu D, Yu S, Wang Z, Shu J, Hou X, Yang X, Jia W, Sheng B. Artificial intelligence in diabetes management: Advancements, opportunities, and challenges. Cell Rep Med. 2023;4:101213. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 210] [Cited by in RCA: 106] [Article Influence: 35.3] [Reference Citation Analysis (0)] |
| 13. | Maheshwari S, Kalia A, Tewari J, Tewari A, Srivastava A, Dantu R, Sachan AK, Verma N, Maheshwari A. Artificial intelligence for diabetes management - a review. J Diabetes Metab Disord Control. 2025;12:24-32. [DOI] [Full Text] |
| 14. | Lancellotti P, Zamorano JL, Habib G, Badano L. The EACVI Textbook of Echocardiography. 2nd ed. Oxford: Oxford University Press, 2016. |
| 15. | Marzoog BA, Chomakhidze P, Gognieva D, Silantyev A, Suvorov A, Abdullaev M, Mozzhukhina N, Filippova DA, Kostin SV, Kolpashnikova M, Ershova N, Ushakov N, Mesitskaya D, Kopylov P. Development and validation of a machine learning model for diagnosis of ischemic heart disease using single-lead electrocardiogram parameters. World J Cardiol. 2025;17:104396. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 5] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 16. | Isaksen JL, Sivertsen CB, Jensen CZ, Graff C, Linz D, Ellervik C, Jensen MT, Jørgensen PG, Kanters JK. Electrocardiographic markers in patients with type 2 diabetes and the role of diabetes duration. J Electrocardiol. 2024;84:129-136. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 5] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 17. | Isaksen JL, Graff C, Ellervik C, Jensen JS, Andersen HU, Rossing P, Kanters JK, Jensen MT. Type 1 diabetes is associated with T-wave morphology changes. The Thousand & 1 Study. J Electrocardiol. 2018;51:S72-S77. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 11] [Cited by in RCA: 9] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 18. | Lipponen JA, Karjalainen PA, Tarvainen MP, Kemppainen J, Mikkola H, Kärki T, Laitinen T. Continuous analysis of repolarization characteristics during insulin induced hypoglycemia. Proceedings of the International Conference on Bio-inspired Systems and Signal Processing; 2011 Jan 26-29; Rome, Italy. |
| 19. | Mezquita-Raya P, Reyes-García R, de Torres-Sánchez A, Matarín MG, Cepero-García D, Pérez de Isla L. Electrical changes during hypoglycaemia in patients with type 1 and type 2 diabetes and high cardiovascular risk. Diabetes Res Clin Pract. 2018;138:44-46. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 20. | Fitzpatrick C, Chatterjee S, Seidu S, Bodicoat DH, Ng GA, Davies MJ, Khunti K. Association of hypoglycaemia and risk of cardiac arrhythmia in patients with diabetes mellitus: A systematic review and meta-analysis. Diabetes Obes Metab. 2018;20:2169-2178. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 44] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 21. | Stahi T, Kaminer K, Shavit I, Nussinovitch U. Diabetes without Overt Cardiac Disease Is Associated with Markers of Abnormal Repolarization: A Case-Control Study. Life (Basel). 2022;12:1173. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 22. | Kittnar O. Electrocardiographic changes in diabetes mellitus. Physiol Res. 2015;64:S559-S566. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 28] [Cited by in RCA: 21] [Article Influence: 1.9] [Reference Citation Analysis (0)] |
| 23. | Chávez-González E, Calero YME, Harrichand S, Mensah EB. QRS and QT Interval Modifications in Patients with Type 2 Diabetes Mellitus. Curr Health Sci J. 2022;48:270-276. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 24. | Giunti S, Bruno G, Lillaz E, Gruden G, Lolli V, Chaturvedi N, Fuller JH, Veglio M, Cavallo-Perin P; EURODIAB IDDM Complications Study Group. Incidence and risk factors of prolonged QTc interval in type 1 diabetes: the EURODIAB Prospective Complications Study. Diabetes Care. 2007;30:2057-2063. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 41] [Cited by in RCA: 35] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 25. | Ninkovic VM, Ninkovic SM, Miloradovic V, Stanojevic D, Babic M, Giga V, Dobric M, Trenell MI, Lalic N, Seferovic PM, Jakovljevic DG. Prevalence and risk factors for prolonged QT interval and QT dispersion in patients with type 2 diabetes. Acta Diabetol. 2016;53:737-744. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 75] [Cited by in RCA: 66] [Article Influence: 6.6] [Reference Citation Analysis (4)] |
| 26. | Cha SA, Yun JS, Lim TS, Kang YG, Lee KM, Song KH, Yoo KD, Park YM, Ko SH, Ahn YB. Baseline-Corrected QT (QTc) Interval Is Associated with Prolongation of QTc during Severe Hypoglycemia in Patients with Type 2 Diabetes Mellitus. Diabetes Metab J. 2016;40:463-472. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 11] [Article Influence: 1.1] [Reference Citation Analysis (0)] |
| 27. | Pickham D, Flowers E, Drew BJ. Hyperglycemia is associated with corrected QT prolongation and mortality in acutely ill patients. J Cardiovasc Nurs. 2014;29:264-270. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 27] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 28. | Vaykshnorayte MA, Ovechkin AO, Azarov JE. The effect of diabetes mellitus on the ventricular epicardial activation and repolarization in mice. Physiol Res. 2012;61:363-370. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 7] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 29. | Marzoog B. Breathomics Detect the Cardiovascular Disease: Delusion or Dilution of the Metabolomic Signature. Curr Cardiol Rev. 2024;20:e020224226647. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 12] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 30. | I.M. Sechenov First Moscow State Medical University. Volatilome and Single-Lead Electrocardiogram Optimize Ischemic Heart Disease Diagnosis Using Machine Learning Models. [accessed 2025 Agu 25]. In: ClinicalTrials.gov [Internet]. Bethesda (MD): U.S. National Library of Medicine. Available from: https://clinicaltrials.gov/study/NCT06181799 ClinicalTrials.gov Identifier: NCT06181799. |
| 31. | Candia JC, Centurión OA, Alderete JF, Torales JM, Aquino NJ, Miño LM, Scavenius KE, García LB, Cáceres C, Sequeira OJ, Chávez CO, Martínez JE, Lovera OA, Galeano EJ. Relationship of the T-wave Tpeak-Tend interval with conduction system disorders in arterial hypertension. Arch Cardiol Mex. 2023;93:69-76. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 1] [Article Influence: 0.3] [Reference Citation Analysis (0)] |
| 32. | Holkeri A, Eranti A, Haukilahti MAE, Kerola T, Kenttä TV, Noponen K, Seppänen T, Rissanen H, Heliövaara M, Knekt P, Junttila MJ, Huikuri HV, Aro AL. Prognostic significance of flat T-waves in the lateral leads in general population. J Electrocardiol. 2021;69:105-110. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6] [Cited by in RCA: 5] [Article Influence: 1.0] [Reference Citation Analysis (0)] |