Published online Mar 26, 2026. doi: 10.4330/wjc.v18.i3.116115
Revised: December 4, 2025
Accepted: January 19, 2026
Published online: March 26, 2026
Processing time: 140 Days and 21.9 Hours
Current advances in diagnostic and therapeutic strategies remain insufficient to reduce the prevalence and incidence rate of diabetes mellitus (DM).
To investigate any association between single-lead electrocardiography (ECG) parameters and the diagnosis of DM.
A single center study involved participants of Caucasian origin for the period between May 2, 2022 and August 23, 2025 with or without DM and aged ≥ 18 years. All participants participating in the study passed the cardiologist’s, random glucose measurement using a glucometer, single lead-ECG registration (using Cardio-Qvark®) and transthoracic echocardiography. Statistical analysis conducted using the R programming language (version 4.5).
The built logistic regression machine learning model demonstrated diagnostic performance in discriminating (area under the curve) type 1 DM 0.84 (95%CI: 0.76-0.91), type 2 DM 0.69 (95%CI: 0.61-0,76), and healthy control 0.82 (95%CI: 0.76-0.87).
The developed model demonstrates an association between single-lead ECG parameters and diabetes status that can support the clinical identification of indi
Core Tip: This study pioneers a novel, non-invasive screening strategy for diabetes mellitus by leveraging the ubiquity of single-lead electrocardiography. Using a machine learning model, we demonstrate that the diabetic state leaves a distinct electrophysio
- Citation: Karbovskaya AD, Marzoog BA, Stroeva A, Chomakhidze P, Gognieva D, Kuznetsova N, Syrkin A, Fadeev VV, Poluboyarinova IV, Ismailova SM, Suvorov A, Kopylov P. Discriminating diabetes mellitus from single-lead electrocardiography using machine learning and multinomial regression. World J Cardiol 2026; 18(3): 116115
- URL: https://www.wjgnet.com/1949-8462/full/v18/i3/116115.htm
- DOI: https://dx.doi.org/10.4330/wjc.v18.i3.116115
Diabetes mellitus (DM) constitutes a pervasive global health crisis, affecting over 500 million individuals worldwide and serving as a primary driver of significant morbi
The electrocardiography (ECG), a cornerstone of cardiovascular assessment, offers a non-invasive window into cardiac electrophysiology and autonomic nervous system function[19]. In recent years, the proliferation of wearable technology and consumer-grade devices capable of recording medical-grade single-lead ECGs has created an unprecedented oppor
The extraction and interpretation of these subtle, complex patterns from the ECG waveform transcend the capabilities of traditional analysis. This is where the transformative potential of artificial intelligence, specifically machine learning (ML) and deep learning (DL), comes to the fore[26-29]. ML algorithms excel at identifying intricate, multidimensional patterns within large datasets that are often imperceptible to the human eye. Several pioneering studies have demon
While the concept is promising, the specific application of ML to single-lead ECG for differentiating not only diabetes from health but also between its principal types (type 1 and type 2) remains a developing field. The pathophysiological pathways leading to type 1 DM (T1DM) (autoimmune beta-cell destruction) and type 2 DM (T2DM) (insulin resistance and progressive secretory defect) impart distinct, yet poorly characterized, impressions on cardiac electrophysiology. A model that can decipher these unique signatures could revolutionize screening protocols and provide profound insights into the cardiac manifestations of different diabetic states[32,33].
Therefore, this study aims to investigate the association between a comprehensive set of parameters derived from a single-lead ECG and the diagnosis of DM. We hypothesize that ML analysis of single-lead ECG features can identify a unique electrophysiological signature associated with DM and develop a validated model capable of distinguishing between healthy individuals, patients with T1DM, and patients with T2DM. By harnessing the synergy of consumer-grade technology and advanced analytics, this research seeks to contribute to the development of a novel, non-invasive, and accessible tool for the early detection and differentiation of DM.
A single center prospective study conducted at university clinical hospital number 2 for the period between May 2, 2022 and August 23, 2025. The study included participants with vs without DM (both first and second type). Patients were recruited from both outpatient and inpatient settings, including those undergoing examination in the endocrinology department. Recruitment was not limited to patients with optimal signal quality at enrollment; however, single exclu
The main logic of the study was to determine the presence or absence of DM without adjustments for covariates – based solely on ECG data and transformations of average complexes. The rationale is straightforward: The algorithm should work quickly, simply, in a screening concept.
Including numerous traditional cardiology covariates would have taken the algorithm out of the screening framework. Additionally, some covariates can be determined very imprecisely (many patients do not know if they had a myocardial infarction, when it occurred, its location, whether they have heart failure, etc.). This inevitably leads to significant bias due to retrospective analysis and degradation of model quality. Such an approach would no longer qualify as a screening method that could be used with an electrocardiograph.
We classified diabetes status primarily using fasting plasma glucose measurements (threshold ≥ 126 mg/dL or 7 mmol/L) following World Health Organization recommendations, supplemented by documented clinical history and current medication use. While this approach reflects real-world clinical practice in our setting, we acknowledge it does not incorporate the complete diagnostic algorithm recommended by international guidelines (American Diabetes Association/World Health Organization), particularly the systematic use of HbA1c and specialized testing for diabetes subtype classification.
The study was carried out according to the principles of Good Clinical Practice and Helinski declaration. All parti
Patients from University Clinical Hospital Number 2 undergoing outpatient or inpatient examination and treatment, including in the Endocrinology Department with a diagnosis of DM, were included in the study. The inclusion and exclu
Inclusion: (1) Patient age over 18 years; and (2) Agreement to participate in the study.
Exclusion: (1) Significant changes in the morphology of the QRS complex (such as bundle branch block, arterial fibrillation, and ventricular extrasystole); (2) Poor quality ECG recorded from the fingers (Parkinson’s disease, tremor of any origin, mental disorders); (3) Chronic heart failure with reduced ejection fraction < 40%; (4) Chronic coronary syndrome; (5) Infectious diseases at acute stage; (6) Unsatisfactory quality of ECG and/or photoplethysmograph (PPG); and (7) Withdrawal of consent for further participation in the study.
The patients underwent a comprehensive examination.
Clinical examination: (1) Collection of complaints; (2) Auscultation; (3) Palpation; (4) Percussion; (5) Assessment of edema syndrome; (6) Measurement of HR; and (7) Blood pressure (BP) level.
Medical history analysis: (1) Collection of cardiological history; and (2) DM history.
Transthoracic echocardiography: Performed using a GE Vivid 5 device. Assessment of cardiac chamber volumes, wall thickness, diastolic and systolic function status, and detection of valvular apparatus and major cardiac vessel dysfunction according to current guidelines[34].
Standard 12-lead ECG recording: Performed using a SCHILLER AT-5 device. Determination of temporal and amplitude ECG parameters, cardiac electrical axis direction, and assessment of cardiac cycle morphology.
Capillary blood glucose analysis: Fasting blood glucose level measured twice within 1-3 days, in a seated position, adhering to the following requirements: (1) Analysis performed after a 12-hour fast; (2) Abstinence from alcohol 2-3 days prior; (3) Avoidance of stress or heavy physical exertion; and (4) Refraining from tooth brushing, smoking, or consump
Single-channel ECG and PPG recording: Recorded using the Qardio-QVARK device (LLC “L Card”, Moscow, Russia; registered with the Federal Service for Surveillance in Healthcare number RZN 2019/8124) immediately after each blood glucose analysis. Recordings lasted 1 minute, performed at rest in a seated position. The recorder has a smartphone case form factor and features 2 sensors for recording ECG and PPG. Recordings were transmitted to a cloud-based platform for analysis using proprietary ML algorithms.
Statistical analysis was performed using the R programming language (version 4.5). For quantitative variables, the distribution normality was assessed using the Shapiro-Wilk test. Descriptive statistics were calculated, including the mean, standard deviation, median, interquartile range, and minimum and maximum values. For categorical and qualitative variables, proportions (percentages) and absolute frequencies were calculated.
Comparative analysis for independent, normally distributed quantitative variables was performed using Welch’s t-test (for two groups) or one-way analysis of variance (for more than two groups), followed by post-hoc pairwise comparisons. For independent, non-normally distributed quantitative variables, the Mann-Whitney U test (for two groups) or the Kruskal-Wallis test (for more than two groups) was employed.
For related (paired) samples, comparative analysis of normally distributed quantitative variables was conducted using a paired Welch’s t-test (for two groups) or repeated-measures analysis of variance (for more than two groups), again followed by post-hoc pairwise comparisons. For related, non-normally distributed quantitative variables, the Wilcoxon signed-rank test (for two groups) or Fisher’s test (for more than two groups) was used.
To analyze dynamic changes across visits, Cochran’s Q test was applied. Where necessary, pairwise comparisons for these dynamic changes were performed using McNemar’s test. The Holm-Bonferroni method was used to correct for multiple comparisons. A significance level (α) of 0.05 was adopted for all analyses, corresponding to a 5% probability of committing a type I error.
The sample size was sufficient to allocate the majority of the data for building associative models (70% of the data), with a separate hold-out set reserved for subsequent validation (20% of the data).
To assess associations with a quantitative outcome (glycemia), a linear regression model with combined L1 and L2 (Elastic Net) regularization was employed. This approach significantly mitigated the risk of multicollinearity and facilitated the selection of factors contributing most substantially to the outcome’s variance. Quantitative data were normalized prior to analysis. The model’s performance was evaluated on the validation set using the coefficient of determination (R2), mean absolute error, maximum error, and the correlation coefficient between predicted and true values.
To evaluate the diagnostic properties of single-lead ECG in determining the presence of DM and its types [resulting in three classes: (1) No diabetes; (2) T1DM; and (3) T2DM], a multinomial logistic regression model with L1 (least absolute shrinkage and selection operator) regularization was utilized. This algorithm enabled the selection of the strongest predictors, after which associations were assessed through the conventional calculation of odds ratios (OR), using the non-diabetic class as the reference. The model’s performance was evaluated on the validation set using the area under the curve (AUC), sensitivity, specificity, and positive and negative predictive value (NPV).
A total of 743 patients with data on the presence or absence of T1DM or T2DM were included in the study. The control group included 537 (72.3%), T1DM group included 61 (8.2%), and T2DM included 145 (19.5%) participants. Information on random glucose level at the time of single-lead ECG recording was available for 696 participants; control 145 (19.5%), T1DM 535 (99.6%), and T2DM 100 (69%). The general characteristics of the study group are presented in the Supplementary Table 1. No significant differences were observed in key cardiovascular risk factors, namely sex and age, among the included participants. Body mass index (BMI) demonstrated overall statistical significance (P = 0.032), which was primarily driven by a marked difference between non-diabetic patients and those with T2DM (P = 0.024), with the latter group exhibiting higher values. No significant differences were observed between type 1 and type 2 diabetes groups (Supplementary Figure 1).
Statistical analysis of smoking prevalence revealed a significant overall difference (P < 0.001). Both insulin-dependent and non-insulin-dependent diabetic patients were significantly more likely to smoke compared to healthy controls (P = 0.002 and P = 0.001, respectively). However, no substantial differences were found between the two diabetic groups. Blood group distribution showed significant variations across all compared subgroups (P < 0.001). The most pronounced deviation was observed in the type 2 diabetes subgroup, which demonstrated a predominance of blood group AB (44.8% vs 15.7% and 8.2% in other groups, respectively). Systolic BP showed a significant difference in the overall sample (P = 0.005), attributable to lower BP levels in type 2 diabetic patients compared to the control group (P = 0.012; Supplementary Figure 1).
The prevalence of coronary artery disease reached statistical significance (P = 0.005), primarily due to lower disease frequency among type 2 diabetic patients compared to non-diabetic subjects (P = 0.009). The distribution of peak location values showed significant variation (P = 0.004), particularly evidenced by the complete absence of anterior peak cases in the insulin-dependent diabetes group. Finally, glucose levels demonstrated highly significant overall differences (P < 0.001), resulting from substantially elevated values in both diabetic groups compared to healthy controls, while no significant differences were observed between the two diabetic subgroups for this parameter (Supplementary Figure 1). Table of pairwise comparisons for categorical variables (holm correction) represented in Supplementary Table 2.
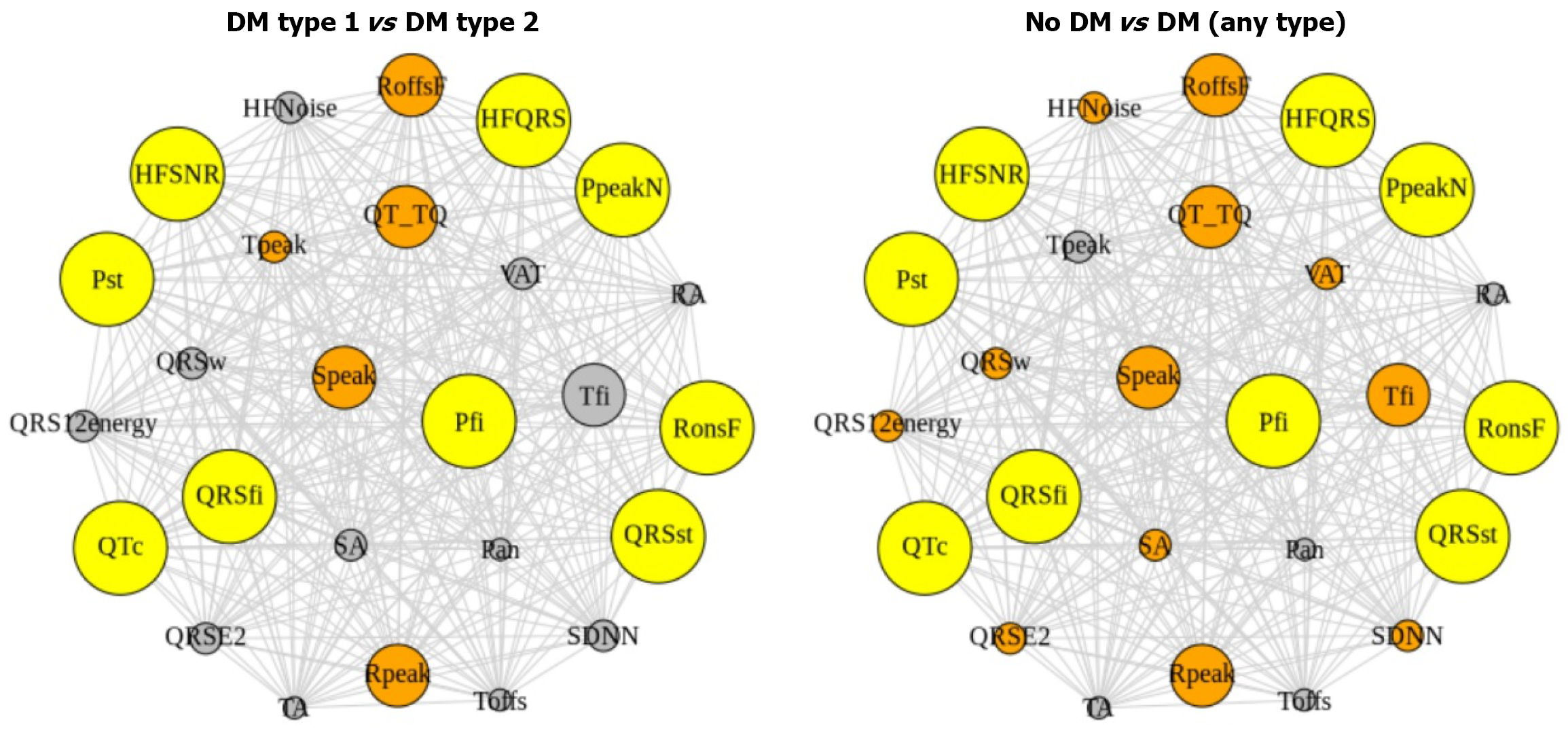
The greatest number of significant differences was identified in the analysis of pulse wave parameters. Statistically significant differences were observed for the majority of the measured indices. Due to distributions that deviated from normality, central tendency and variability are presented as medians with 25% and 75% (interquartile range). Group comparisons for these parameters are presented in the Supplementary Table 3. Pairwise comparisons (adjusted for multiple comparisons using the Holm method) revealed statistically significant differences not only relative to the non-diabetic control group but also between the two types of DM (Table 1). Of particular interest are those factors that demonstrate significant differences between the two types of DM, as well as those that effectively differentiate either type of diabetes from the control group. Such factors are visualized in the graph below (Figure 1). Thus, the factors that demonstrated significant differences between the three classes were prolonged QT interval (QTc), HFQRS, HFSNR, Pst, P-wave dispersion index (Pfi), QRSst, QRSfi, PpeakN, and RonsF.
| Factor | No DM: T1DM | No DM: T2DM | T1DM - T2DM |
| HFNoise | 0.660 | 0.009 | 0.660 |
| Ventricular activation | 0.704 | 0.009 | 0.210 |
| Prolonged QT interval | 0.000 | 0.024 | 0.034 |
| QT_TQ | 0.001 | 0.284 | 0.022 |
| HFQRS | 0.000 | 0.000 | 0.000 |
| HFSNR | 0.000 | 0.000 | 0.000 |
| T-wave amplitude | 0.078 | 0.078 | 0.697 |
| QRS12energy | 0.024 | 0.541 | 0.108 |
| QRSE2 | 0.033 | 0.639 | 0.084 |
| QRSw | 0.084 | 0.006 | 0.878 |
| Pan | 0.078 | 0.128 | 0.457 |
| RA | 0.129 | 0.129 | 0.519 |
| SA | 0.024 | 0.484 | 0.136 |
| Pst | 0.000 | 0.000 | 0.007 |
| P-wave dispersion index | 0.000 | 0.001 | 0.001 |
| QRSst | 0.000 | 0.000 | 0.001 |
| QRSfi | 0.000 | 0.000 | 0.000 |
| T-wave flattening | 0.000 | 0.000 | 0.637 |
| PpeakN | 0.000 | 0.000 | 0.000 |
| Rpeak | 0.000 | 0.721 | 0.001 |
| Speak | 0.001 | 0.546 | 0.016 |
| Tpeak | 0.158 | 0.158 | 0.048 |
| Toffs | 0.192 | 0.192 | 0.051 |
| RonsF | 0.026 | 0.026 | 0.002 |
| RoffsF | 0.004 | 0.188 | 0.002 |
| SDNN | 0.018 | 0.278 | 0.222 |
To identify predictors associated with diabetes and its subtypes, a multinomial logistic regression model was employed. The model was initially adjusted for predefined covariates including age, sex, BMI, and smoking status. Continuous variables were standardized using z-score normalization (mean-centered and scaled by standard deviation). The regression model was trained on a randomly selected subset of 519 participants (70% of the total cohort), with validation performed on the remaining 222 participants (30%). Stratified sampling ensured proportional representation of both type 1 and type 2 diabetes cases across both training and validation sets.
Feature selection was performed using L1-regularized multinomial regression to mitigate multicollinearity risks. Model stability was ensured through 10-fold cross-validation. The following predictors were selected: (1) Smoking status (categorized as never smoked, current smoker, or former smoker); (2) QTc; (3) T-wave amplitude (TA); (4) Pan; (5) RA; (6) SA; (7) Pst; (8) T-wave flattening (Tfi); (9) RoffsF; (10) Age; (11) Pfi; and (12) PpeakN.
These predictors were combined into a single regression equation with estimation of the coefficients, their 95%CI, subsequent exponentiation, and calculation of the OR (Table 2). A multinomial logistic regression model with healthy participants as the reference group, the associations between clinical and electrocardiographic parameters and the likelihood of having either type 1 or type 2 diabetes were analyzed.
| Type 1 DM | Type 2 DM | |||||
| OR | 95%CI | P value | OR | 95%CI | P value | |
| Intercept | 0.016 | 0.005-0.048 | < 0.001 | 1.391 | 1.049-1.845 | 0.022 |
| Smoking | 0.156 | 0.110-0.221 | < 0.001 | 0.662 | 0.100-4.401 | 0.670 |
| Prolonged QT interval | 3.810 | 1.222-11.875 | 0.021 | 0.351 | 0.197-0.624 | < 0.001 |
| T-wave amplitude | 4.033 | 1.819-8.941 | 0.001 | 11.218 | 3.778-33.312 | < 0.001 |
| Pan | 1.537 | 0.917-2.577 | 0.103 | 4.738 | 2.943-7.630 | < 0.001 |
| RA | 1.185 | 0.846-1.659 | 0.324 | 2.668 | 1.505-4.730 | 0.001 |
| SA | 0.947 | 0.556-1.611 | 0.840 | 1.286 | 0.940-1.757 | 0.115 |
| Pst | 0.714 | 0.533-0.957 | 0.024 | 0.682 | 0.442-1.053 | 0.085 |
| T-wave flattening | 0.605 | 0.350-1.046 | 0.072 | 0.972 | 0.716-1.320 | 0.854 |
| RoffsF | 0.954 | 0.710-1.282 | 0.755 | 0.544 | 0.133-2.223 | 0.397 |
| Age | 0.954 | 0.607-1.500 | 0.839 | 0.980 | 0.481-1.997 | 0.956 |
| P-wave dispersion index | 0.725 | 0.540-0.974 | 0.033 | 0.732 | 0.169-3.177 | 0.677 |
| PpeakN | 1.510 | 0.965-2.361 | 0.071 | 1.946 | 0.865-4.376 | 0.108 |
T1DM (compared to healthy controls): The presence of type 1 diabetes was statistically significantly associated with the following factors: (1) Smoking: A reduced likelihood of T1DM was observed among smokers (OR = 0.16, 95%CI: 0.11-0.22, P < 0.001), which may reflect potential confounding factors or behavioral characteristics; (2) Prolonged QTc interval (OR = 3.81, 95%CI: 1.22-11.87, P = 0.021); (3) Increased TA (OR = 4.03, 95%CI: 1.82-8.94, P = 0.001); (4) A decrease in the P-wave terminal force parameter, reflecting atrial electrical activity (OR = 0.71, 95%CI: 0.53-0.96, P = 0.024); (5) A reduction in the Pfi (OR = 0.73, 95%CI: 0.54-0.97, P = 0.033); and (6) A borderline trend towards statistical significance was also identified for the following predictors: Tfi (P = 0.072) and PpeakN (P = 0.071).
T2DM (compared to healthy controls): For patients with T2DM, the following statistically significant predictors were identified: (1) QTc interval: An inverse association was observed: A shorter QTc interval was associated with a higher likelihood of T2DM (OR = 0.35, 95%CI: 0.20-0.62, P < 0.001); (2) TA: Was strongly associated with an increased likelihood of T2DM (OR = 11.22, 95%CI: 3.78-33.31, P < 0.001); (3) Pan parameter, reflecting the spatial characteristics of the P-wave (OR = 4.74, 95%CI: 2.94-7.63, P < 0.001); and (4) RA (OR = 2.67, 95%CI: 1.50-4.73, P = 0.001).
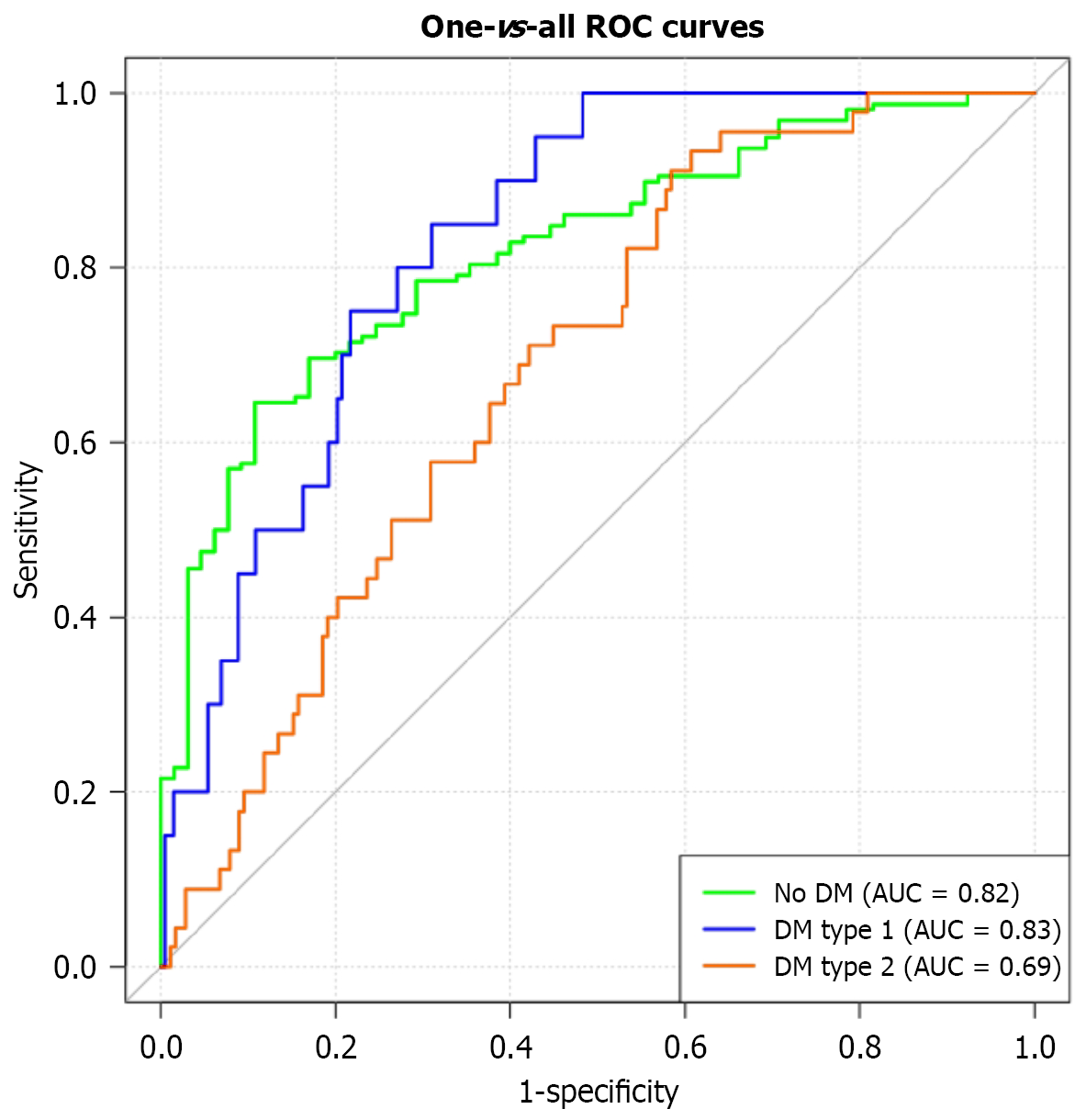
The remaining parameters, including age, Tfi, SA, and RoffsF, did not demonstrate statistically significant differences between the T2DM group and healthy controls. The most stable predictors of diabetes presence (compared to the control group) were abnormalities in QTc interval, TA magnitude, and atrial activity parameters (Pan, RA, Pfi, Pst). Notably, QTc intervals exhibited divergent directional associations: Prolongation was associated with T1DM, while shortening was linked to T2DM. When validated on the holdout dataset of 222 observations, the model demonstrated the following performance metrics (Table 3, Figure 2). In the one-versus-one classification approach, the highest AUC was observed for differentiating between any type of diabetes and non-diabetic conditions, while the poorest performance was achieved in distinguishing type 1 diabetes from type 2 diabetes (type 1 diabetes vs type 2 diabetes) (Table 4).
| Statistic | No DM | Type 1 DM | Type 2 DM |
| Area under the curve | 0.82 (0.76-0.87) | 0.84 (0.76-0.91) | 0.69 (0.61-0.76) |
| Sensitivity | 0.65 (0.57-0.72) | 0.85 (0.68-1.00) | 0.91 (0.81-0.98) |
| Specificity | 0.89 (0.81-0.97) | 0.69 (0.63-0.75) | 0.42 (0.34-0.49) |
| Positive predictive value | 0.94 (0.89-0.98) | 0.21 (0.13-0.32) | 0.28 (0.21-0.36) |
| Negative predictive value | 0.51 (0.42-0.60) | 0.98 (0.95-1.00) | 0.95 (0.89-0.99) |
| Groups comparing | Area under the curve | Lower limit 95%CI | Upper limit 95%CI |
| No DM against T1DM | 0.8981013 | 0.8439558 | 0.9522468 |
| No DM against T2DM | 0.7758087 | 0.7054986 | 0.8461189 |
| T1DM against T2DM | 0.6366667 | 0.4914294 | 0.7819039 |
The validation of the diagnostic model revealed statistically significant differences in predictive performance across patient groups depending on diabetic status. The overall diagnostic performance, evaluated by the AUC, demonstrated strong discriminative ability for the absence of diabetes (AUC = 0.82; 95%CI: 0.76-0.87). The model also showed good performance for T1DM (AUC = 0.84; 95%CI: 0.76-0.91). For T2DM, the discriminative capability was comparatively limited, though the confidence interval did not include 0.5 (AUC = 0.69; 95%CI: 0.61-0.76). These results indicate the current model has restricted applicability specifically for detecting T2DM.
In the non-diabetic group, the model exhibited high specificity (0.89; 95%CI: 0.81-0.97) and high positive predictive value (PPV) (0.94; 95%CI: 0.89-0.98). Among diabetic patients, a pronounced asymmetry in performance characteristics was observed. The model demonstrated high sensitivity in both type 1 (0.85; 95%CI: 0.68-1.00) and type 2 diabetes groups (0.91; 95%CI: 0.81-0.98), coupled with high NPV (0.98 and 0.95, respectively). This supports its potential utility for ruling out pathology in these cohorts. However, the low specificity in the type 2 diabetes group (0.42; 95%CI: 0.34-0.49) resulted in correspondingly low PPV values (0.28; 95%CI: 0.21-0.36), indicating a high rate of false positives for this particular class.
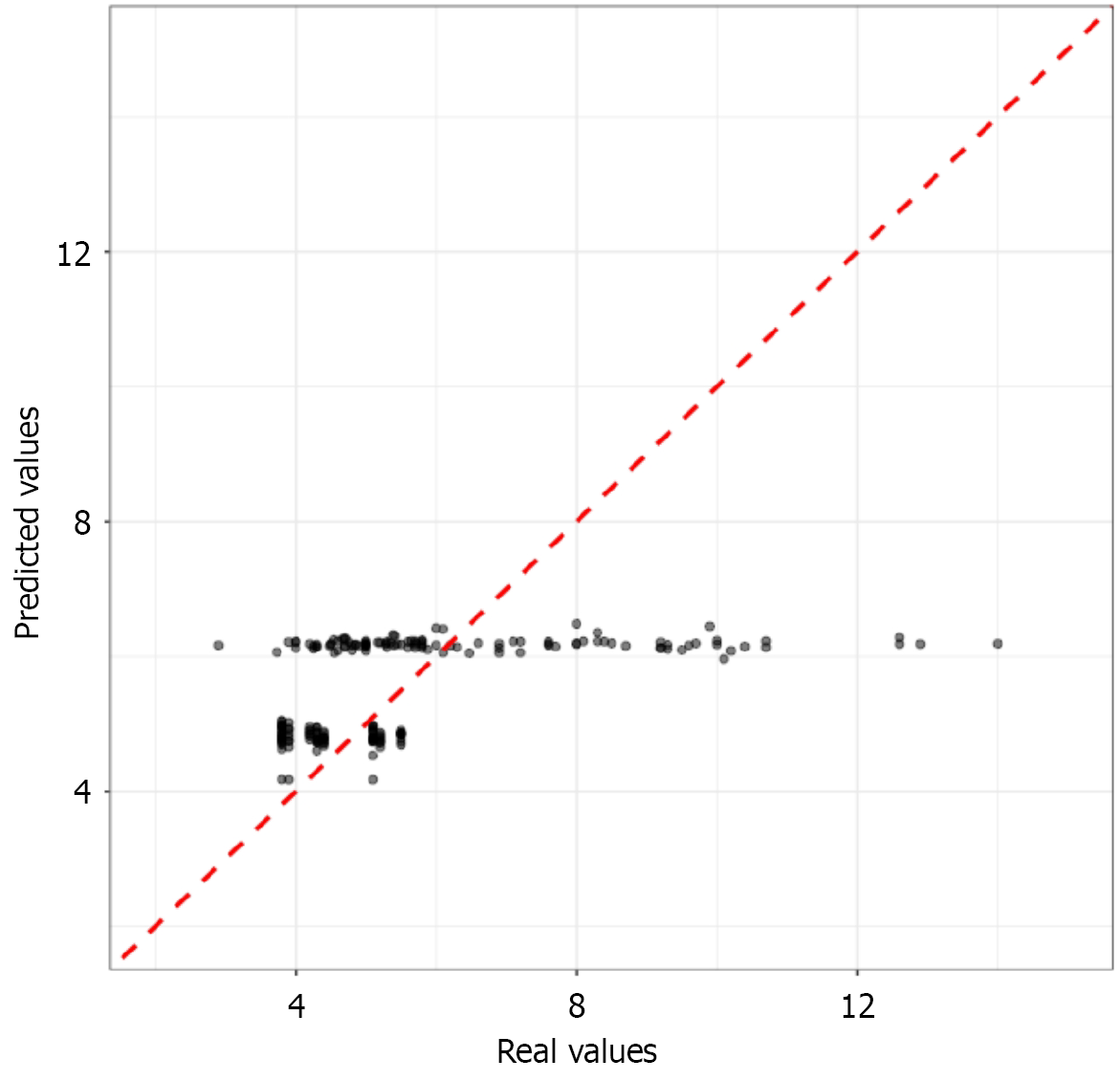
Glycemic level is a more labile parameter than some electrocardiographic-derived features. Nevertheless, an attempt was made to identify associations between glycemic levels and several potential predictors. A linear regression model with combined L1 and L2 regularization (Elastic Net) was employed to identify associations while mitigating multicollinearity risks. The analysis utilized 10-fold cross-validation with mean-centered normalization and scaling by standard deviation for all continuous variables. The Table 5 below presents the coefficient estimates of selected factors from the final model, along with their 95%CI and statistical significance. However, a graphical representation of the data indicated some degree of correspondence between predicted and observed values up to a glycemic level of 5 mmol/L. Beyond this threshold, the predicted values effectively plateaued, exhibiting minimal dependence on the predictor variables (Figure 3). Analysis of standard regression quality metrics revealed suboptimal model performance, characterized by a low coefficient of determination (R2) 0.217, a moderate mean absolute error 1.35, and a substantially elevated maximum error 14.5. There
| Parameter | Estimated coefficient | Lower limit 95%CI | Upper limit 95%CI | Р value |
| Intercept | 5.852 | 4.950 | 6.753 | 0.0000000 |
| HFSNR | -0.058 | -0.159 | 0.043 | 0.261 |
| QRSfi | -0.001 | -0.003 | 0.0009 | 0.253 |
| T-wave flattening | 0.0018 | 0.001 | 0.003 | 0.0000114 |
| PpeakN | -0.0017 | -0.004 | 0.0009 | 0.202 |
The results demonstrate a significant and clinically relevant association between features derived from a single-lead ECG and the diagnosis of DM, utilizing a ML framework. Our findings indicate that the electrophysiological and autonomic nervous system alterations inherent to the diabetic state manifest in measurable, albeit subtle, changes in the ECG signal. These changes are sufficiently distinct to allow for the development of predictive models with promising diagnostic accuracy, particularly for differentiating diabetic from non-diabetic states and for identifying T1DM.
The core finding is the robust performance of the multinomial logistic regression model. The AUC values 0.84 for T1DM and 0.82 for the healthy control group signify a strong discriminative capacity. This performance is noteworthy given the non-invasive, low-cost, and highly accessible nature of single-lead ECG technology, which is increasingly integrated into consumer wearables and portable health devices. The model's high sensitivity (0.85 for T1DM, 0.91 for T2DM) and NPV (NPV > 0.95 for both DM types) are its most compelling attributes from a clinical screening perspective. This suggests that such a tool could be highly effective as a rule-out test, potentially identifying individuals who are very unlikely to have diabetes and thus reducing the need for more invasive or resource-intensive testing in a large portion of the population.
However, the model’s performance for T2DM was notably weaker (AUC = 0.69), characterized by low specificity (0.42) and a consequently high false-positive rate (low PPV of 0.28). This asymmetry in performance highlights a critical nuance. The electrophysiological signature of T2DM, as captured by our selected features, appears to be less distinct or more heterogeneous than that of T1DM. This could be attributed to the broader pathophysiological spectrum of T2DM, which often coexists with a cluster of other metabolic and cardiovascular conditions like hypertension and obesity, as also seen in our cohort’s higher BMI and hypertension prevalence. These comorbid states likely introduce confounding electrophysiological changes that overlap with those caused by hyperglycemia alone, making it challenging for the model to isolate a unique “T2DM fingerprint”. Furthermore, the one-vs-one analysis confirmed the greatest difficulty lies in distinguishing T1DM from T2DM (AUC = 0.64), underscoring that while both types share the common endpoint of hyper
The feature selection process yielded profound insights into the specific ECG parameters most affected by diabetes. The identification of factors like QTc interval, TA, and various P-wave parameters (Pst, Pfi, Pan, RA) points towards diffuse involvement of the heart’s electrical system. The divergent behavior of the QTc interval is particularly fascinating. QTc prolongation was associated with T1DM, which aligns with existing literature on diabetic autonomic neuropathy leading to reduced sympathetic tone and potentially prolonged repolarization. Conversely, QTc shortening was associated with T2DM. This counterintuitive finding may be explained by the different autonomic progression in T2DM, often characterized by an early hyperadrenergic state, or by the complex interplay of ischemia, hypertrophy, and elec
The alterations in P-wave parameters (Pst, Pfi, PpeakN) strongly implicate atrial myopathy in the diabetic process. Diabetes is a known risk factor for atrial fibrillation, and these early subclinical changes in atrial conduction and repolarization, detectable on a simple single-lead ECG, could represent a valuable biomarker for assessing atrial fibrillation risk long before overt arrhythmia occurs. This suggests that the utility of this technology may extend beyond diabetes diagnosis to the stratification of cardiovascular complications within the diabetic population.
In stark contrast to diagnosing the disease state, our attempts to predict real-time glycemic levels using ECG features were unsuccessful. The regression model for glycemia exhibited poor performance (R2 = 0.22, high maximum error), with predictions plateauing beyond 5 mmol/L. This result is, in fact, expected and reinforces a crucial principle: The ECG reflects the end-organ electrophysiological impact of prolonged glycemic exposure and autonomic dysfunction, not the immediate, labile concentration of glucose in the blood. Glycemic levels fluctuate rapidly due to meals, stress, and medication, while the myocardial remodeling and autonomic imbalance that affect ECG parameters develop over months and years. Therefore, the ECG is a biomarker of chronic diabetic disease burden rather than an acute glucometer. This distinction is vital for setting appropriate expectations for this technology.
A DL model, IGRNet, was developed to diagnose prediabetes using 12-lead ECGs, achieving an accuracy of 0.781 and an AUC of 0.777. This suggests potential for non-invasive, real-time prediabetes diagnosis[35]. Another study proposed a novel approach using single-lead ECGs to detect HbA1c levels, achieving an accuracy of 0.9015 and an AUC of 0.9899. This method highlights the potential for non-invasive and fast detection of diabetes-related metrics[36].
The DiaBeats algorithm, using ECG signal data, accurately predicted diabetes and pre-diabetes classes with a precision of 97.1% and an accuracy of 96.8%. This model utilized leads III, aVL, V4, V5, and V6, indicating the significant role of specific ECG leads in diabetes detection[21]. Combining single-lead ECG with other sensors (e.g., glucose, accelerometer) has shown improved accuracy in diabetes prediction. A multisensor combination achieved a prediction accuracy of 98.2%, significantly higher than using single sensors alone[37].
A study utilized multi-resolution analysis of single-lead ECG signals to develop a system for detecting diabetic patients. The system achieved a classification accuracy of 91.5%, demonstrating the effectiveness of ML in identifying diabetes through ECG analysis[38]. Another framework used intrinsic time-scale decomposition and ML to screen for diabetes using single-lead ECG signals, achieving an accuracy of 86.9 %. This approach is suitable for resource-limited environments and highlights the potential for widespread screening[25]. The single lead ECG using Cardio_Qvark studied to diagnose ischemic heart disease and showed an improvement in the diagnostic accuracy comparing to the physical ECG-stress test[39].
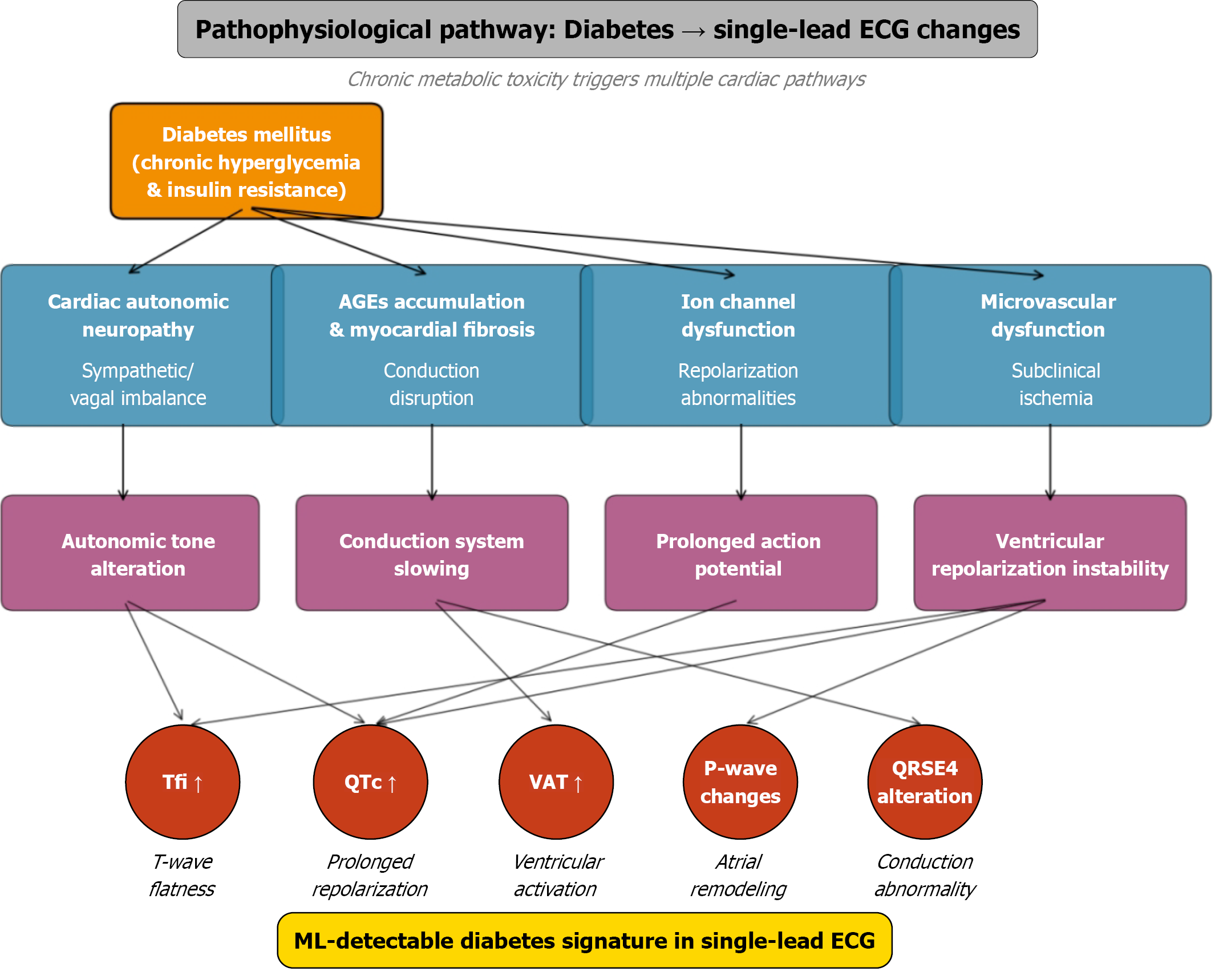
DM induces distinct cardiac alterations through chronic hyperglycemia and insulin resistance, which create a detectable electrophysiological signature on a single-lead ECG. Key pathophysiological pathways include cardiac autonomic neuropathy, which disrupts sympathetic-parasympathetic balance and causes heterogeneous ventricular repolarization, and the accumulation of advanced glycation end-products that promote myocardial fibrosis. These pro
The integration of these subtle changes – manifested as Tfi, QTc, slowed ventricular activation (VAT), and altered P-wave morphology – creates a composite biomarker for diabetes. While individually minor, these features collectively form a distinct pattern that ML models can decode from a single-lead ECG, effectively identifying the integrated cardiac impact of diabetes before overt clinical disease develops (Figure 4).
The integration of these subtle changes – manifested as Tfi, QTc, slowed ventricular activation, and altered P-wave morphology – creates a composite biomarker for diabetes. While individually minor, these features collectively form a distinct pattern that ML models can decode from a single-lead ECG, effectively identifying the integrated cardiac impact of diabetes before overt clinical disease develops.
Our study has several limitations. Firstly, it is a single-center study with a predominantly Caucasian cohort, which may limit the generalizability of our findings to other ethnicities and healthcare settings. Secondly, the imbalance in group sizes, particularly the smaller T1DM group, can affect the stability of the estimates for that subclass. Thirdly, while we adjusted for key confounders like age, sex, and BMI, residual confounding from unmeasured factors (e.g., duration of diabetes, specific medications, presence of subclinical coronary artery disease) is possible. The diagnosis of DM was not solely based on HbA1c or oral glucose tolerance tests in all cases, relying partly on medical history, which could in
Future research directions should include multi-center, multi-ethnic validation studies with larger, balanced cohorts to improve model robustness and generalizability. Incorporating additional data streams from wearables, such as activity level and HR variability over time, could enhance predictive power. Exploring more complex DL architectures that analyze the raw ECG signal directly might capture features invisible to conventional parameterization. Furthermore, longitudinal studies are needed to determine if these ECG changes can predict the future development of diabetes or its complications, positioning single-lead ECG as a tool for risk stratification and prevention, not just diagnosis.
The optimal application of this technology would be in clinical settings where it serves as a decision-support tool to identify patients requiring definitive biochemical evaluation, rather than as a standalone diagnostic or broad population screening method. Our model detects electrophysiological changes associated with established diabetes rather than predicting future development of the disease. It should be viewed as an adjunct to, not a replacement for, established diagnostic criteria including fasting glucose, HbA1c, and oral glucose tolerance testing.
Diabetic state leaves a discernible imprint on the heart’s electrical activity, detectable through ML analysis of a single-lead ECG. While not a replacement for standard glycemic tests, this technology shows potential as a supplementary clinical tool to identify individuals who may benefit from confirmatory glycemic testing, particularly in settings where standard diagnostic approaches are limited or when diabetes status is uncertain. It represents a significant step towards leveraging ubiquitous digital health technology for the early detection and management of one of the world’s most prevalent chronic diseases.
The developed models demonstrate a stronger association between single-lead ECG parameters and the diagnosis of DM. This is likely attributable to the relatively stable and long-term physiological alterations associated with the state of disease. In contrast, single-lead ECG indicators do not adequately account for the values or variance in glycemic fluctuations.
| 1. | Hossain MJ, Al-Mamun M, Islam MR. Diabetes mellitus, the fastest growing global public health concern: Early detection should be focused. Health Sci Rep. 2024;7:e2004. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 652] [Cited by in RCA: 419] [Article Influence: 209.5] [Reference Citation Analysis (8)] |
| 2. | Siam NH, Snigdha NN, Tabasumma N, Parvin I. Diabetes Mellitus and Cardiovascular Disease: Exploring Epidemiology, Pathophysiology, and Treatment Strategies. Rev Cardiovasc Med. 2024;25:436. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 63] [Cited by in RCA: 50] [Article Influence: 25.0] [Reference Citation Analysis (1)] |
| 3. | Conning-Rowland M, Cubbon RM. Molecular mechanisms of diabetic heart disease: Insights from transcriptomic technologies. Diab Vasc Dis Res. 2023;20:14791641231205428. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 8] [Article Influence: 2.7] [Reference Citation Analysis (0)] |
| 4. | Ershova AI, Ivanova AA, Kiseleva AV, Sotnikova EA, Meshkov AN, Drapkina OM. From biobanking to personalized prevention of obesity, diabetes and metabolic syndrome. Cardiovasc Ther Prev. 2022;20:3123. [RCA] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 5. | Toscano A, Barca E, Musumeci O. Update on diagnostics of metabolic myopathies. Curr Opin Neurol. 2017;30:553-562. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 13] [Article Influence: 1.4] [Reference Citation Analysis (1)] |
| 6. | Furuta Y, Tinker RJ, Hamid R, Cogan JD, Ezell KM, Oglesbee D, DeBerardinis RJ, Phillips JA 3rd; Undiagnosed Diseases Network. A review of multiple diagnostic approaches in the undiagnosed diseases network to identify inherited metabolic diseases. Orphanet J Rare Dis. 2024;19:427. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 6] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 7. | Chandra TB, Reddy AS, Adarsh A, Jabbar MA, Jyothi BN. Diabetes Prediction Using Gaussian Naive Bayes and Artificial Neural Network. 2024 International Conference on Distributed Computing and Optimization Techniques (ICDCOT); 2024 Mar 15-16; Bengaluru, India. Bengaluru: IEEE, 2024: 1-5. [DOI] [Full Text] |
| 8. | Viberti G. Thiazolidinediones-benefits on microvascular complications of type 2 diabetes. J Diabetes Complications. 2005;19:168-177. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 14] [Article Influence: 0.7] [Reference Citation Analysis (1)] |
| 9. | Fan W, Wong ND. Diabetes mellitus and macrovascular disease: epidemiology and cardiovascular risk assessment. In: Krentz AJ, Chilton RJ, editors. Cardiovascular Endocrinology and Metabolism: Theory and Practice of Cardiometabolic Medicine. Netherlands: Elsevier, 2023: 11-38. |
| 10. | Simon Machado R, Mathias K, Joaquim L, Willig de Quadros R, Petronilho F, Tezza Rezin G. From diabetic hyperglycemia to cerebrovascular Damage: A narrative review. Brain Res. 2023;1821:148611. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 19] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 11. | LeRoith D. Hyperglycemia, Hypertension, and Dyslipidemia in Type 2 Diabetes Mellitus: Goals for Diabetes Management. Clin Cornerstone. 2008;9:S8-S16. [DOI] [Full Text] |
| 12. | Beckman JA, Creager MA. Vascular Complications of Diabetes. Circ Res. 2016;118:1771-1785. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 344] [Cited by in RCA: 285] [Article Influence: 28.5] [Reference Citation Analysis (0)] |
| 13. | Pedro-Botet J, Chillarón JJ, Benaiges D, Flores-Le Roux JA. [Cardiovascular prevention in diabetes mellitus: A multifactorial challenge]. Clin Investig Arterioscler. 2016;28:154-163. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 5] [Article Influence: 0.5] [Reference Citation Analysis (1)] |
| 14. | Viigimaa M, Sachinidis A, Toumpourleka M, Koutsampasopoulos K, Alliksoo S, Titma T. Macrovascular Complications of Type 2 Diabetes Mellitus. Curr Vasc Pharmacol. 2020;18:110-116. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 313] [Cited by in RCA: 247] [Article Influence: 41.2] [Reference Citation Analysis (4)] |
| 15. | Sone H. Diabetes Mellitus. In: Vasan RS, Sawyer DB, editors. Encyclopedia of Cardiovascular Research and Medicine. Netherlands: Elsevier, 2017: 9-16. |
| 16. | Szuszkiewicz-Garcia MM, Davidson JA. Cardiovascular disease in diabetes mellitus: risk factors and medical therapy. Endocrinol Metab Clin North Am. 2014;43:25-40. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 45] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 17. | Cavagnolli G, Comerlato J, Comerlato C, Renz PB, Gross JL, Camargo JL. HbA(1c) measurement for the diagnosis of diabetes: is it enough? Diabet Med. 2011;28:31-35. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 44] [Cited by in RCA: 35] [Article Influence: 2.3] [Reference Citation Analysis (1)] |
| 18. | Salarizadeh N, Shojai S, Pebdeni AB, Nojoki F, Zargar SJ, Rezaei MH. Classical and new candidate biomarkers for developing biosensors in diagnosing diabetes and prediabetes; past, present and future. In: Barhoum A, Altintas Z. Advanced Sensor Technology: Biomedical, Environmental, and Construction Applications. Netherlands: Elsevier, 2022: 337-381. |
| 19. | Hou R, Dou J, Wu L, Zhang X, Li C, Wang W, Gao Z, Tang X, Yan L, Wan Q, Luo Z, Qin G, Chen L, Ji J, He Y, Wang W, Mu Y, Zheng D. Development and validation of a machine learning-based model to predict isolated post-challenge hyperglycemia in middle-aged and elder adults: Analysis from a multicentric study. Diabetes Metab Res Rev. 2024;40:e3832. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.5] [Reference Citation Analysis (0)] |
| 20. | Angelaki E, Barmparis GD, Fragkiadakis K, Maragkoudakis S, Zacharis E, Plevritaki A, Kampanieris E, Kalomoirakis P, Kassotakis S, Kochiadakis G, Tsironis GP, Marketou ME. Diagnostic performance of single-lead electrocardiograms for arterial hypertension diagnosis: a machine learning approach. J Hum Hypertens. 2025;39:58-65. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 5] [Article Influence: 5.0] [Reference Citation Analysis (1)] |
| 21. | Kulkarni AR, Patel AA, Pipal K V, Jaiswal SG, Jaisinghani MT, Thulkar V, Gajbhiye L, Gondane P, Patel AB, Mamtani M, Kulkarni H. Machine-learning algorithm to non-invasively detect diabetes and pre-diabetes from electrocardiogram. BMJ Innov. 2022;9:32-42. [DOI] [Full Text] |
| 22. | Glass GF, Sudhir A, Pandit AAK. The ECG and Metabolic Abnormalities. In: Brady WJ, Lipinski MJ, Darby AE, Bond MC, Charlton NP, Hudson K, Williamson K, editors. Electrocardiogram in Clinical Medicine. Hoboken: John Wiley and Sons Ltd, 2020: 307-313. |
| 23. | Wald DA. ECG manifestations of selected metabolic and endocrine disorders. Emerg Med Clin North Am. 2006;24:145-157, vii. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 15] [Article Influence: 0.8] [Reference Citation Analysis (0)] |
| 24. | Darby AE. Nontraditional Rhythm Disorders: Dysrhythmias Related to Metabolic and Toxicologic Conditions. In: Brady WJ, Lipinski MJ, Darby AE, Bond MC, Charlton NP, Hudson K, Williamson K, editors. Electrocardiogram in Clinical Medicine. Hoboken: John Wiley and Sons Ltd, 2020: 187-192. |
| 25. | Gupta K, Bajaj V. A Robust Framework for Automated Screening of Diabetic Patient Using ECG Signals. IEEE Sens J. 2022;22:24222-24229. [DOI] [Full Text] |
| 26. | Sun JY, Shen H, Qu Q, Sun W, Kong XQ. The application of deep learning in electrocardiogram: Where we came from and where we should go? Int J Cardiol. 2021;337:71-78. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 24] [Article Influence: 4.8] [Reference Citation Analysis (0)] |
| 27. | Sleiman J, Daher M, Khalil M, Diab A, Kassem A. Biological age estimation from ECG using Artificial Intelligence approaches. 2024 2nd Jordanian International Biomedical Engineering Conference (JIBEC); 2024 Nov 27; Amman, Jordan. Bengaluru: IEEE, 2024: 78-83. |
| 28. | Buyya A, Ogeti T, Suhas G, Kashapogula P, Panigrahy AK. Arrhythmias Classification by using STFT-based Spectrograms, Transfer Learning and Concatenation of features. 2023 4th International Conference for Emerging Technology; 2023 May 26-28; Belgaum, India. Bengaluru: IEEE, 2023. [DOI] [Full Text] |
| 29. | Naik S, Debnath S, Justin V. A Review of Arrhythmia Classification with Artificial Intelligence Techniques: Deep vs Machine Learning. 2021 2nd International Conference for Emerging Technology (INCET); 2021 May 21-23; Belagavi, India. Bengaluru: IEEE, 2021: 1-14. |
| 30. | Bergquist JA, Zenger B, Brundage J, MacLeod RS, Shah R, Ye X, Lyones A, Ranjan R, Tasdizen T, Bunch TJ, Steinberg BA. Comparison of Machine Learning Detection of Low Left Ventricular Ejection Fraction Using Individual ECG Leads. Comput Cardiol (2010). 2023;50:10.22489/cinc.2023.047. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2] [Cited by in RCA: 4] [Article Influence: 1.3] [Reference Citation Analysis (1)] |
| 31. | Kuznetsova N, Sagirova Z, Suvorov A, Dhif I, Gognieva D, Afina B, Poltavskaya M, Sedov V, Chomakhidze P, Kopylov P. A screening method for predicting left ventricular dysfunction based on spectral analysis of a single-channel electrocardiogram using machine learning algorithms. Biomed Signal Process Control. 2023;86:105219. [DOI] [Full Text] |
| 32. | Moussaoui R, Salem O, Mehaoua A. Analysis of Different Machine Learning Models for Diabetes Prediction. 2023 IEEE International Conference on E-health Networking, Application and Services (Healthcom); 2023 Dec 15-17; China. Bengaluru: IEEE, 2023: 43-48. |
| 33. | Wijoseno MR, Permanasari AE, Pratama AR. Machine Learning Diabetes Diagnosis Literature Review. 2023 10th International Conference on Information Technology, Computer, and Electrical Engineering (ICITACEE); 2023 Aug 31-Sep 1; Semarang, Indonesia; Bengaluru: IEEE, 2023: 304-308. |
| 34. | Lancellotti P, Zamorano JL, Habib G, Badano L. The EACVI textbook of echocardiography. Oxford: Oxford University Press, 2017. |
| 35. | Wang L, Mu Y, Zhao J, Wang X, Che H. IGRNet: A Deep Learning Model for Non-Invasive, Real-Time Diagnosis of Prediabetes through Electrocardiograms. Sensors (Basel). 2020;20:2556. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 37] [Cited by in RCA: 19] [Article Influence: 3.2] [Reference Citation Analysis (1)] |
| 36. | Li J, Lu J, Tobore I, Liu Y, Kandwal A, Wang L, Zhou J, Nie Z. Towards noninvasive and fast detection of Glycated hemoglobin levels based on ECG using convolutional neural networks with multisegments fusion and Varied-weight. Expert Syst Appl. 2021;186. [DOI] [Full Text] |
| 37. | Site A, Nurmi J, Lohan ES. Machine-Learning-Based Diabetes Prediction Using Multisensor Data. IEEE Sens J. 2023;23:28370-28377. [DOI] [Full Text] |
| 38. | Jain A, Verma A, Verma AK, Bajaj V. Tunable Q-factor wavelet transform based identification of diabetic patients using ECG signals. Comput Methods Biomech Biomed Engin. 2024;1-10. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 3] [Article Influence: 1.5] [Reference Citation Analysis (0)] |
| 39. | Marzoog BA, Chomakhidze P, Gognieva D, Silantyev A, Suvorov A, Abdullaev M, Mozzhukhina N, Filippova DA, Kostin SV, Kolpashnikova M, Ershova N, Ushakov N, Mesitskaya D, Kopylov P. Development and validation of a machine learning model for diagnosis of ischemic heart disease using single-lead electrocardiogram parameters. World J Cardiol. 2025;17:104396. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 5] [Article Influence: 5.0] [Reference Citation Analysis (0)] |