Published online Apr 27, 2026. doi: 10.4240/wjgs.v18.i4.115798
Revised: December 19, 2025
Accepted: February 9, 2026
Published online: April 27, 2026
Processing time: 154 Days and 1.7 Hours
In pancreatic neuroendocrine neoplasms (pNENs) with liver metastasis, marked upregulation of T-LAK cell-originated protein kinase (TOPK) is associated with poor prognosis.
To elucidate the role of TOPK in pNEN progression and metastasis and to explore the underlying mechanisms. The therapeutic potential of the TOPK inhibitor HI-TOPK-032 was further investigated to inform targeted treatment strategies.
TOPK expression was assessed in tissue samples from 23 patients with pNENs, with and without liver metastasis. TOPK was knocked down in the BON-1 cell line to examine its effects on cell proliferation, epithelial-mesenchymal transition, and activation of the mitogen-activated protein kinase (MAPK) axis. RNA-seq analyzed gene expression changes following TOPK knockdown. The effects of HI-TOPK-032 on pNENs cell proliferation, migration, and invasiveness were also evaluated.
TOPK was significantly upregulated in pNENs with liver metastasis and corre
TOPK plays a critical role in pNEN progression and liver metastasis through the MAPK axis. HI-TOPK-032 exhibits promising antitumor activity by targeting TOPK, suggesting its potential as a therapeutic option for pNENs with liver metastasis. Further in vivo and clinical validation is warranted.
Core Tip: Pancreatic neuroendocrine neoplasms (pNENs) are relatively rare and typically slow growing but may lead to serious health complications. T-LAK cell-derived protein kinase is a serine/threonine kinase originally identified from a cDNA library of lymphokine-activated killer T cells and is overexpressed in multiple malignancies. This study demonstrates that T-LAK cell-derived protein kinase is markedly overexpressed in pNENs tissues with liver metastasis and further clarifies its effects on downstream signaling and the malignant biological behavior of pNENs.
- Citation: Meng M, Chen Q, Yuan M, Guo X, Ren SQ, Yuan CH. T-LAK cell-originated protein kinase promotes tumorigenesis and metastasis of pancreatic neuroendocrine neoplasms via mitogen-activated protein kinase axis activation. World J Gastrointest Surg 2026; 18(4): 115798
- URL: https://www.wjgnet.com/1948-9366/full/v18/i4/115798.htm
- DOI: https://dx.doi.org/10.4240/wjgs.v18.i4.115798
Pancreatic neuroendocrine neoplasms (pNENs) are relatively rare, slow-growing tumors that arise from pancreatic hormone-producing cells. Despite their indolent nature, pHENs can cause serious clinical consequences due to hormone hypersecretion, local tumor growth, and metastatic spread[1]. Liver metastasis represents a prominent clinical feature, occurring in nearly two-thirds of patients at initial presentation[2]. In this context, therapeutic options are limited and generally exhibit reduced responsiveness. Hepatic involvement significantly complicates disease management and is a major determinant of poor prognosis[3]. Investigating the underlying mechanisms is therefore critical for the develop
In multiple cancer types, oncogenesis and tumor progression are driven by coordinated signaling through the Ras/phosphatidylinositol 3-kinase/phosphatase and tensin homologue deleted on chromosome ten/Ak strain transforming and Ras/rapidly accelerated fibrosarcoma 1 (Raf-1)/mitogen-activated protein kinase (MAPK) kinase (MEK)/extracellu
T-LAK cell-originated protein kinase (TOPK) is a serine-threonine kinase originally identified from a cDNA library of lymphokine-activated killer T cells (T-LAK cells). Research has demonstrated that TOPK expression is reacquired during malignant transformation[8]. TOPK is overexpressed in various cancer types (e.g., gastric, non-small cell lung, and esophageal carcinomas), and its expression level correlates positively with tumor aggressiveness[9-11].
In the present study, we identified a significant overexpression of TOPK in pathological tissues from pNENs with liver metastasis. Notably, Yuryev and Wennogle[12] reported in 2003 that TOPK functions as Raf-1-binding protein, identified through yeast two-hybrid screening, suggesting a potential interaction through the cysteine-rich zinc-binding domain. On this basis, we further sought to clarify the effects of TOPK on downstream signaling and its influence on the biological behavior of pNENs.
Clinical information and pathological specimens from patients with pNENs were obtained from Peking University Third Hospital. Ethical approval was granted by the Ethics Committee of Peking University Third Hospital (Approval No. IRB00006761-2016128).
The pNEN cell lines BON-1 and QGP-1 were purchased from Beijing Beina Chuanglian Biotechnology Institute (Beijing, China). The HPDE, Mia-2, and SW-1990 cell lines were obtained from in-house laboratory stocks. All cells were cultured in Roswell Park Memorial Institute (RPMI)-1640 medium supplemented with 10% fetal bovine serum at 37 °C in a humi
The TOPK antibody (rabbit, 16110-1-AP) was purchased from Proteintech Group, Inc. (IL, United States). Antibodies against MEK1/2 (rabbit, #8727), phospho-MEK1/2 (rabbit, #3958), ERK1/2 (rabbit, #9102), phospho-ERK1/2 (rabbit, #4370), ribosomal S6 kinases (RSK) 1/2/3 (rabbit, #9355), phospho-RSK (rabbit, #11989), tubulin (rabbit, #2146), and vimentin (rabbit, #14179) were purchased from Cell Signaling Technology (MA, United States). Glyceraldehyde-3-phosphate dehydrogenase antibody (mouse, ab8245) was obtained from Abcam (Cambridge, United Kingdom), while E-cadherin (mouse, sc-71009) and N-cadherin (mouse, sc-8424) antibodies were sourced from Santa Cruz Biotechnology (TX, United States).
Tumor tissues were fixed in 4% paraformaldehyde and embedded in paraffin. Serial 5-μm sections were mounted on glass slides, deparaffinized in xylene, and rehydrated through graded alcohols, followed by ethylenediaminetetraacetic acid-based antigen retrieval. Endogenous peroxidase activity was quenched with H2O2, and sections were blocked with bovine serum albumin for 30 minutes. After overnight incubation with primary antibodies at 4 °C, sections were washed with phosphate-buffered saline (PBS) and incubated with secondary antibodies for 2 hours at room temperature (RT). Signal development was performed using a diaminobenzidine kit (Invitrogen, CA, United States), followed by counterstaining with Mayer’s hematoxylin. Slides were digitally scanned using an Aperio AT2 scanner (Leica Biosystems Imaging, Inc., CA, United States), and images were captured with a Leica Imaging System Microscope (200 magnification × or 400 × magnification; Leica Microsystems, Inc., CA, United States).
TOPK knockdown in BON-1 cells was achieved using lentiviral particles generated by co-transfecting HEK293T cells with TOPK-targeting short hairpin RNA and packaging plasmids. Viral supernatants were collected 48 hours later, filtered, and used to infect BON-1 in the presence of polybrene (8 μg/mL). Stable TOPK knockdown cells were selected with puromycin (2 μg/mL). Quantitative polymerase chain reaction and western blotting were used to confirm knockdown efficiency.
BON-1 cells were seeded into a 96-well plate and cultured for 24 hours, followed by incubation with HI-TOPK-032-supplemented medium for an additional 48 hours in an IncuCyte ZOOM system (Essen Bioscience, MI, United States). Images were acquired every 3 hours, and confluence was quantified using IncuCyte software.
Cell proliferation was assessed using the cell counting kit-8 (CCK-8) assay. Cells were seeded into a 96-well plate at a density of 3 × 103 cells per well in 200 μL of culture medium. At designated time points over five consecutive days, 10 μL of CCK-8 reagent was added per well. After incubation for 1-2 hours at 37 °C, absorbance was measured at 450 nm using a microplate reader. All experiments were performed in triplicate.
A colony formation assay was performed to assess cell viability. Cells were seeded into a 6-well plate at a density of 1000 cells per well and cultured for 2 weeks. Cells were terminated once individual colonies exceeded 50 cells. After removal of the supernatant and PBS washing, cells were fixed with 4% paraformaldehyde and stained with 1% crystal violet. The number of colonies was then counted.
Cells cultured in 10-cm dishes to approximately 90% confluence were washed with PBS and lysed on ice for 30 minutes using radio immunoprecipitation assay buffer (R0010; Solarbio, Beijing, China) supplemented with protease and pho
For migration assays, cells in logarithmic phase were harvested, trypsinized, washed with PBS, and resuspended in serum-free RPMI-1640 medium containing different concentrations of HI-TOPK-032. BON-1 cells (1.5 × 105 cells in 200 μL) were added to the upper chamber, while the lower chamber contained 600 μL of RPMI-1640 medium supplemented with 20% fetal bovine serum and the corresponding drug concentration. After an appropriate incubation period, cells were fixed with methanol, stained with 0.5% crystal violet, and rinsed with tap water. Non-migrated cells on the upper surface were removed, and membranes were air-dried at RT before imaging and cell counting. Each experiment was performed in triplicate.
For invasion assays, Matrigel was thawed and diluted 1:40 to a final concentration of 50 mg/L. Diluted Matrigel (100 μL) was added to the transwell chambers and allowed to solidify for 1 hour at 37 °C in a 5% CO2 incubator. Subsequent procedures were identical to those used in the migration assay.
Differential gene expression analysis was conducted using the DESeq2 R package (version 1.20.0) with two biological replicates per condition. DESeq2 applies a negative binomial distribution model to identify differentially expressed genes (DEGs). P value were adjusted using the Benjamini-Hochberg method to control the false discovery rate, and genes with adjusted P < 0.05 were considered differentially expressed.
Gene Ontology (GO) enrichment analysis of DEGs was performed using the clusterProfiler R package, which accounts for gene length bias. GO terms with corrected P < 0.05 were considered significantly enriched.
Gene set enrichment analysis was applied to detect coordinated but subtle expression changes between biological conditions. Gene were ranked according to differential expression, and enrichment of predefined gene sets at the top or bottom of the ranked list was evaluated.
Gene set enrichment analysis was performed using the Broad Institute tool (http://www.broadinstitute.org/gsea/index.jsp). GO, Kyoto Encyclopedia of Genes and Genomes (KEGG), Reactome, Disease Ontology, and DisGeNET datasets were analyzed independently.
Protein-protein interaction (PPI) analysis of DEGs was performed using the STRING database to explore known and predicted PPIs and to identify functional interaction networks.
All quantitative experiments were performed at least three times. Statistical analyses were conducted using GraphPad Prism 8 and SPSS version 22.0. Western blot band intensities were quantified using Image J software. Comparison among groups were performed using one-way analysis of variance (ANOVA) followed by LSD-t post hoc tests. Data from at least three independent experiments are presented as mean ± SD. Fisher’s exact test (two-sided) was used where appropriate, and Pearson’s correlation coefficient assessed inter-variable associations. Overall survival (OS) was defined as the interval from treatment initiation to death or last follow-up. Between-group comparisons used Student’s t-test or the Mann-Whitney U test, as appropriate. Continuous variables are presented as mean ± SD or median (95% confidence interval). All tests were two-sided, with significance defined as P < 0.05.
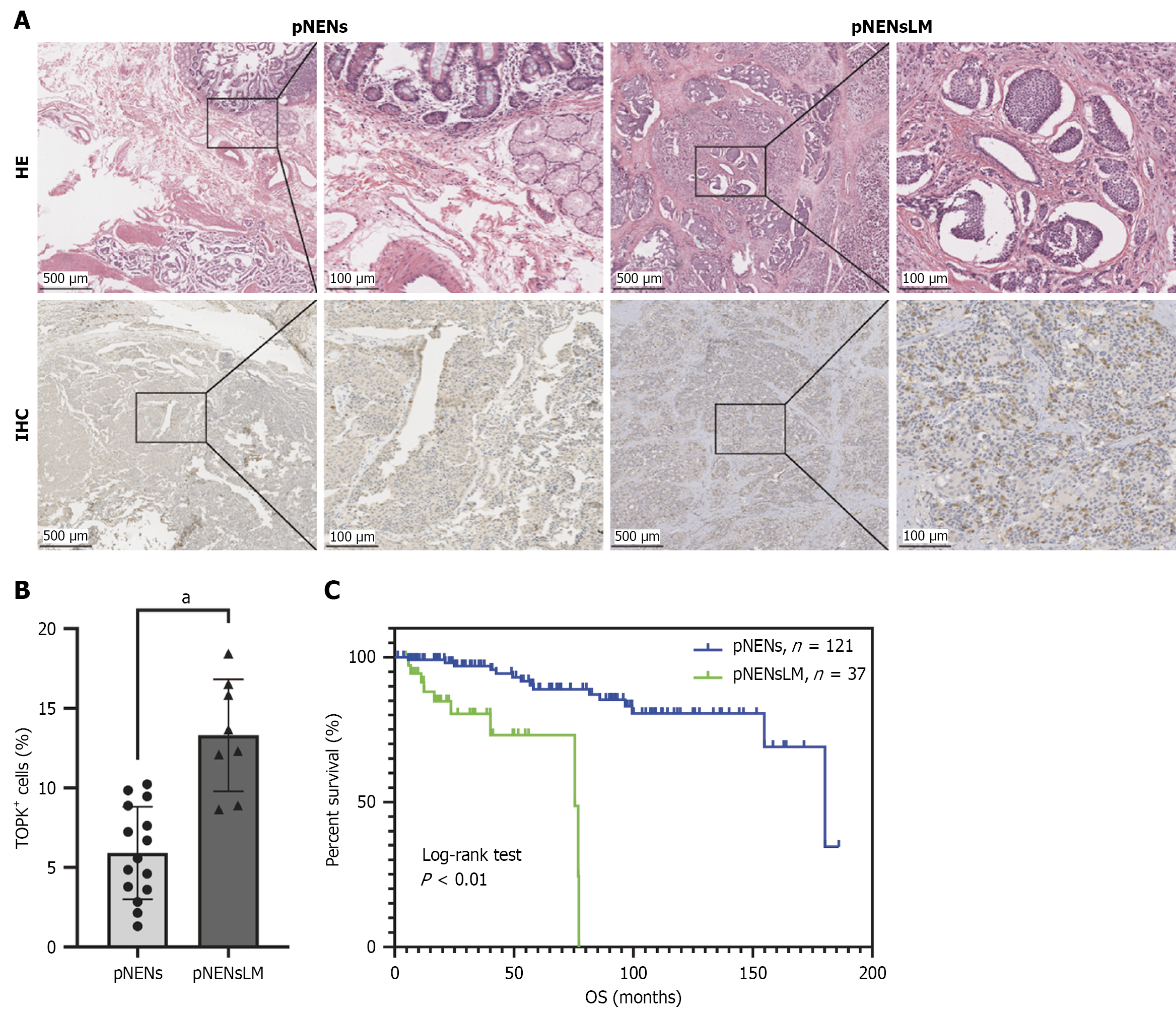
To investigate the clinical relevance of TOPK in pNEN metastasis, tissue samples from 23 patients with or without liver metastasis were analyzed (Figure 1A). The proportion of TOPK-positive cells was significantly higher in the liver metastasis group than in the non-metastatic group (pNENs; P < 0.001), indicating a strong association between TOPK expression and liver metastasis (Figure 1B). In addition, clinical and follow-up data from surgically treated patients at our center between 2010 and 2022 were collected. Kaplan-Meier survival analysis demonstrated that patients with pNENs in the liver metastasis group had significantly shorter OS than those without liver metastasis (Figure 1C).
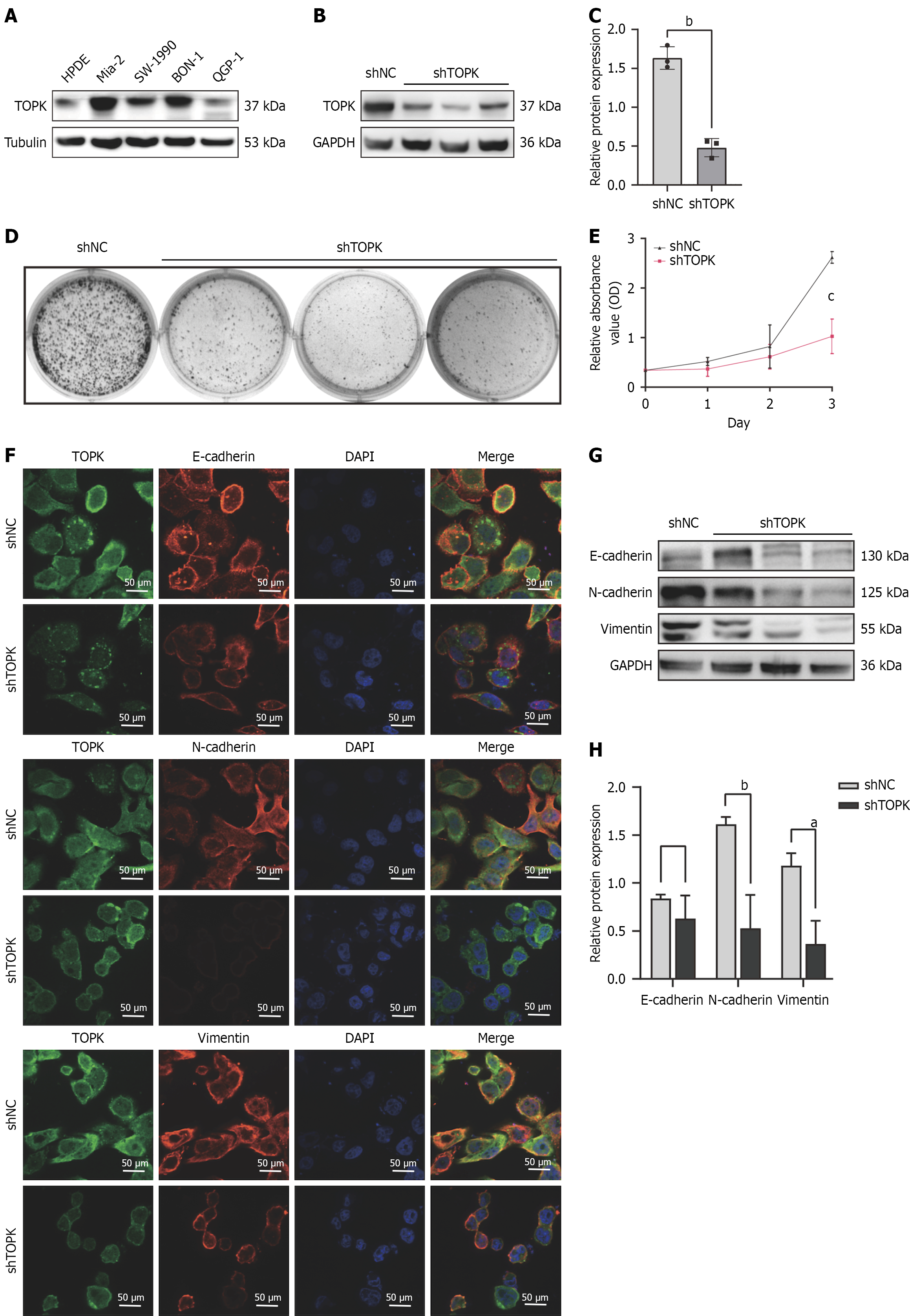
To further clarify the role of TOPK in pNENs, we first examined its expression in HPDE cells, pancreatic ductal adenocarcinoma cell lines (Mia-2 and SW-1990), and pNEN cell lines (BON-1 and QGP-1) using Western blotting. Compared with HPDE cells, TOPK expression was upregulated in Mia-2, SW-1990, and BON-1 cells (Figure 2A). Based on these results, BON-1 cells were selected for subsequent experiments. TOPK was silenced in BON-1 cells using short hairpin RNA, and western blotting confirmed effective suppression of TOPK protein expression 48 hours after transfection (Figure 2B and C).
Cell viability was assessed using a plate colony formation assay. Compared with the short hairpin negative control (shNC) group, TOPK-knockdown BON-1 cells exhibited a markedly reduced colony-forming capacity (Figure 2D). Consistently, CCK-8 assays showed that TOPK knockdown significantly inhibited BON-1 cell proliferation (Figure 2E).
Epithelial-mesenchymal transition (EMT), enables epithelial cells to acquire mesenchymal characteristics and is a key process underlying tumor metastasis and invasion. To determine whether TOPK regulates EMT in pNENs, EMT-related markers were examined following TOPK knockdown. Immunofluorescence analysis revealed decreased expression of N-cadherin and vimentin in short hairpin TOPK (shTOPK)-treated cells (Figure 2F). These findings were further confirmed via western blotting, which showed reduced N-cadherin and vimentin protein levels after TOPK knockdown (Figure 2G and H). Collectively, these results indicates that TOPK contributes to EMT in BON-1 cells, which may partly explain the propensity for liver metastasis in pNENs with high TOPK expression.
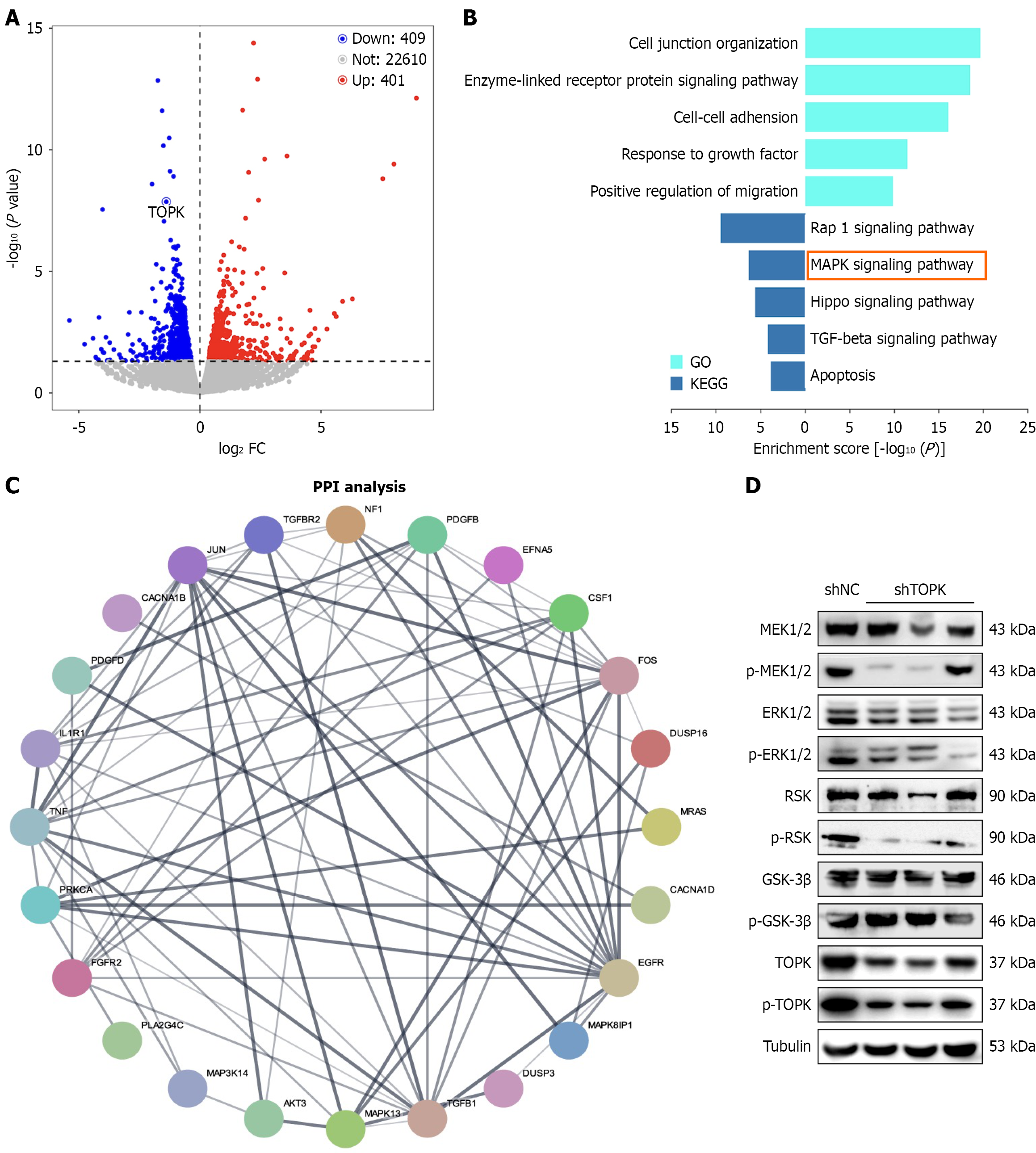
To explore the molecular mechanisms by which TOPK promotes tumor proliferation and metastasis, RNA-seq was performed in shTOPK and shNC groups, with three biological replicates per group. A total of 810 DEGs were identified, including 401 upregulated and 409 downregulated genes in the shTOPK group compared with the shNC group (Figure 3A).
Considering that TOPK is a serine/threonine kinase, enrichment analyses were conducted focusing on the downregulated gene set. The top 20 GO and KEGG terms were identified, and the five most relevant KEGG and GO pathways associated with tumor proliferation and metastasis were highlighted. GO enrichment analysis indicated that downregulated genes were mainly involved in cell junction organization, enzyme-linked receptor protein signaling pathways, cell-cell adhesion, response to growth factors, and positive regulation of cell migration. KEGG pathway analysis revealed significant enrichment in the Ras-proximate-1, MAPK, Hippo, and transforming growth factor-beta signaling pathways, as well as apoptosis (Figure 3B).
Within the MAPK signaling pathway, which was of particular interest, 24 downregulated genes were identified as potentially mediated by TOPK. PPI analysis further demonstrated strong interactions among these genes (Figure 3C).
Consistently, assessment of MAPK signaling activity showed that phosphorylation levels of MEK1/2, ERK1/2, and RSK were significantly reduced following TOPK knockdown compared with the shNC group, whereas glycogen synthase kinase 3β expression remained unchanged (Figure 3D).
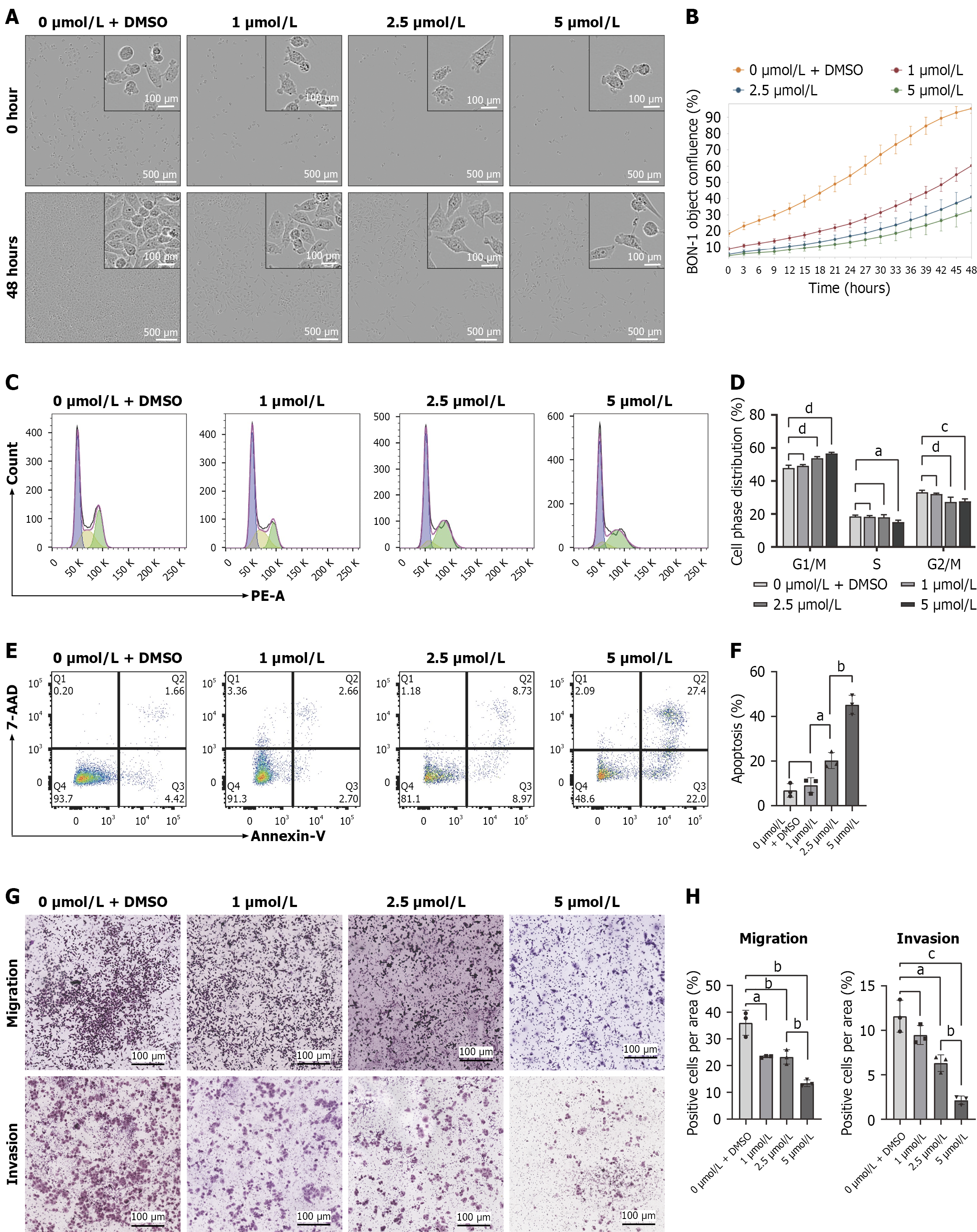
HI-TOPK-032 is a selective inhibitor of TOPK activity. BON-1 cells were treated with varying HI-TOPK-032 concentrations (Figure 4A). Using the IncuCyte system, we observed a significant inhibition of BON-1 cell growth after 48 hours of treatment with 5 μmol/L of HI-TOPK-032 (Figure 4B).
Flow cytometry was performed to assess cell cycle distribution in BON-1 cells following treatment with different HI-TOPK-032 concentrations (Figure 4C). The results showed that HI-TOPK-032 prolonged the G1/M phase and shortened the G2/M phase in BON-1 cells (Figure 4D). The effect of HI-TOPK-032 on BON-1 apoptosis was further evaluated using an Annexin-V/PI apoptosis detection kit and flow cytometry (Figure 4E). Compared with untreated BON-1 cells, HI-TOPK-032-treated cells showed a markedly elevated apoptotic rate (Figure 4F), indicating that HI-TOPK-032 promotes BON-1 apoptosis.
To evaluate the role of HI-TOPK-032 in BON-1 invasion and metastasis, cells were treated with HI-TOPK-032 under conditions of TOPK inhibition (Figure 4G). HI-TOPK-032 treatment significantly reduced cell migration and invasion. Specifically, both migratory and invasive (Figure 4H) capacities were suppressed 24 hours after treatment in a dose-dependent manner. These findings indicate that TOPK contributes to pNEN cell motility and that HI-TOPK-032 effec
pNENs are relatively rare tumors and generally have a better prognosis than other pancreatic malignancies. However, due to the anatomical location of the pancreas and the frequent absence of specific clinical symptoms, many patients present with liver metastases at the time of diagnosis. Liver metastasis significantly limits treatment options and adversely affects OS in patients with pNENs. Although pNENs are typically classified as low-grade malignant tumors, a phenotypic transformation occurs during disease progression that predisposes tumors to liver metastasis. This process appears to be independent of tumor grade, although the underlying mechanisms remain to be fully clarified.
In this study, pNENs cases were stratified according to liver metastasis status and subjected to comparative TOPK expression analysis using immunocytochemistry. To further elucidate the role of TOPK in tumor proliferation and metastasis, we compared TOPK expression across normal pancreatic cells, highly malignant pancreatic ductal adenocarcinoma cell lines, and pNENs cell lines, identifying BON-1 as a suitable model for subsequent experiments.
TOPK knockdown in BON-1 cells resulted in reduced cell viability and decreased expression of metastasis-associated EMT proteins. RNA sequencing revealed distinct transcriptional profiles following TOPK depletion. Given the kinase function of TOPK and its suppressive effects on downstream signaling after knockdown, enrichment analyses focused on downregulated genes. Among pathways closely associated with tumor biology, the MAPK axis emerged as the most prominently affected.
TOPK, a member of the MEK family, plays a central role in the RAS/RAF/MEK/ERK signaling cascade[13]. It is ex
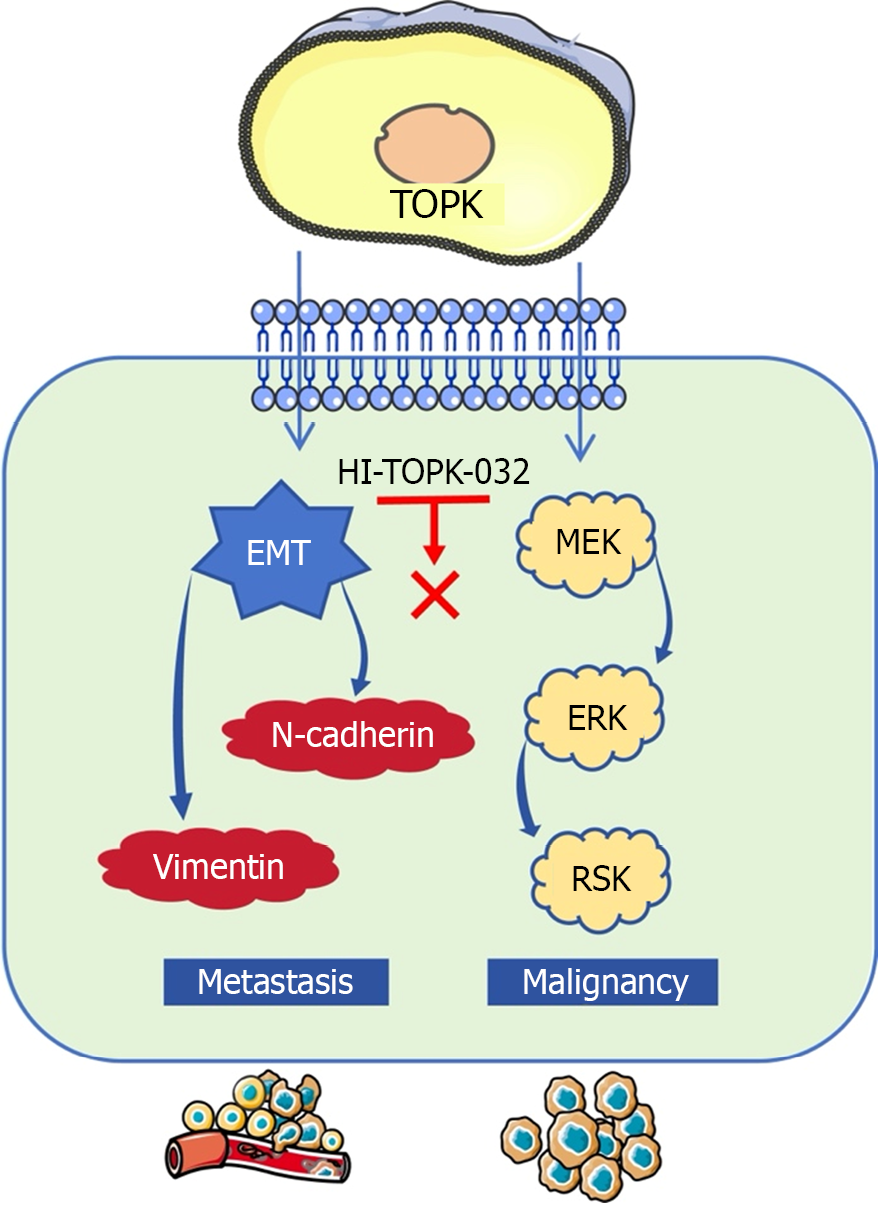
Notably, specific and well-established inhibitors targeting TOPK were used to validate our findings. As anticipated, HI-TOPK-032, a TOPK inhibitor, effectively suppressed BON-1 cell growth and metastatic potential. In pNENs, TOPK functions as a key regulator of the MEK/ERK/RSK axis, an essential intracellular signaling cascade. Activation of this axis involves stepwise phosphorylation of MEK, ERK, and RSK, thereby stimulating downstream signaling. In tumors, this pathway can be aberrantly activated by oncogenes such as BCR-ABL, Flt-3, Kit, and EGFR[17,18]. Consistent with this, our PPI analysis showed that TOPK knockdown resulted in EGFR downregulation in BON-1 cells. Prior studies have linked these gene alterations to apoptosis through post-translational modification of associated molecules (e.g., Bad, Bim, and Mcl-1), thereby influencing cell cycle regulation, apoptosis, and differentiation[19]. Our in vitro findings corroborated these observations. TOPK, a member of the MEK family, has attracted increasing attention for its roles in tumor cell growth, apoptosis, migration, and invasiveness. TOPK upregulation has been reported across multiple cancer types. Functionally, TOPK acts as a MEK, and is critical for catalytic activity during mitosis[20]. Recent evidence further indicates that TOPK regulates mitosis by modulating multiple DNA-binding proteins[21]. Additionally, TOPK is overexpressed in cancer stem cells, where it promotes proliferation and self-renewal, thereby enhancing tumor invasiveness[22,23]. Collectively, these findings suggests that the TOPK-MEK/ERK/RSK interaction is pivotal in pNEN progression and metastasis. By effectively blocking this signaling cascade, HI-TOPK-032 suppresses pNEN cell proliferation and migration (Figure 5). Several TOPK-specific inhibitors have demonstrated promising preclinical efficacy and are therefore anticipated to enter clinical evaluation in the near future[24,25].
Despite these findings, several limitations of this study should be acknowledged. First, the relatively small sample size may limit the reliability of the statistical analyses. Future studies should further evaluate the biological significance of TOPK through in vitro and in vivo experiments and validate its clinical relevance in larger, independent cohorts, po
TOPK plays a pivotal role in pNEN progression and metastasis, with its upregulation correlating with liver metastasis and poor prognosis, highlighting its potential as a therapeutic target. HI-TOPK-032 demonstrates promising efficacy in suppressing aggressive tumor phenotypes by inhibiting TOPK activity. Accordingly, targeting TOPK may represent a novel and viable therapeutic strategy for pNENs, particularly in the presence of liver metastasis. Future studies should focus on the clinical utility of TOPK inhibitors and further elucidate the molecular mechanisms underlying TOPK-driven tumor progression.
| 1. | Howe JR, Merchant NB, Conrad C, Keutgen XM, Hallet J, Drebin JA, Minter RM, Lairmore TC, Tseng JF, Zeh HJ, Libutti SK, Singh G, Lee JE, Hope TA, Kim MK, Menda Y, Halfdanarson TR, Chan JA, Pommier RF. The North American Neuroendocrine Tumor Society Consensus Paper on the Surgical Management of Pancreatic Neuroendocrine Tumors. Pancreas. 2020;49:1-33. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 347] [Cited by in RCA: 300] [Article Influence: 50.0] [Reference Citation Analysis (4)] |
| 2. | Cloyd JM, Ejaz A, Konda B, Makary MS, Pawlik TM. Neuroendocrine liver metastases: a contemporary review of treatment strategies. Hepatobiliary Surg Nutr. 2020;9:440-451. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 55] [Cited by in RCA: 52] [Article Influence: 8.7] [Reference Citation Analysis (0)] |
| 3. | Sonbol MB, Mazza GL, Mi L, Oliver T, Starr J, Gudmundsdottir H, Cleary SP, Hobday T, Halfdanarson TR. Survival and Incidence Patterns of Pancreatic Neuroendocrine Tumors Over the Last 2 Decades: A SEER Database Analysis. Oncologist. 2022;27:573-578. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 44] [Cited by in RCA: 123] [Article Influence: 30.8] [Reference Citation Analysis (0)] |
| 4. | Yan J, Yu S, Jia C, Li M, Chen J. Molecular subtyping in pancreatic neuroendocrine neoplasms: New insights into clinical, pathological unmet needs and challenges. Biochim Biophys Acta Rev Cancer. 2020;1874:188367. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 22] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 5. | Ullah R, Yin Q, Snell AH, Wan L. RAF-MEK-ERK pathway in cancer evolution and treatment. Semin Cancer Biol. 2022;85:123-154. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 14] [Cited by in RCA: 383] [Article Influence: 76.6] [Reference Citation Analysis (1)] |
| 6. | Lim HJ, Crowe P, Yang JL. Current clinical regulation of PI3K/PTEN/Akt/mTOR signalling in treatment of human cancer. J Cancer Res Clin Oncol. 2015;141:671-689. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 128] [Cited by in RCA: 135] [Article Influence: 12.3] [Reference Citation Analysis (0)] |
| 7. | Karkhanis M, Park JI. Sp1 regulates Raf/MEK/ERK-induced p21(CIP1) transcription in TP53-mutated cancer cells. Cell Signal. 2015;27:479-486. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 34] [Article Influence: 3.1] [Reference Citation Analysis (0)] |
| 8. | Han Z, Li L, Huang Y, Zhao H, Luo Y. PBK/TOPK: A Therapeutic Target Worthy of Attention. Cells. 2021;10:371. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 51] [Article Influence: 10.2] [Reference Citation Analysis (0)] |
| 9. | Peng S, Yin Y, Zhang Y, Zhu F, Yang G, Fu Y. FYN/TOPK/HSPB1 axis facilitates the proliferation and metastasis of gastric cancer. J Exp Clin Cancer Res. 2023;42:80. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 9] [Cited by in RCA: 22] [Article Influence: 7.3] [Reference Citation Analysis (0)] |
| 10. | Xiao J, Zhang L, Yi H, Zou L, Mo J, Xue F, Zheng J, Huang Y, Lu H, Wu H, Xue P, Zhang X, He L, Li Z, Pang S, Qiao G, Duan Q, Zhu F. Inhibiting ALK-TOPK signaling pathway promotes cell apoptosis of ALK-positive NSCLC. Cell Death Dis. 2022;13:828. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 7] [Article Influence: 1.8] [Reference Citation Analysis (0)] |
| 11. | Wu W, Xu J, Gao D, Xie Z, Chen W, Li W, Yuan Q, Duan L, Zhang Y, Yang X, Chen Y, Dong Z, Liu K, Jiang Y. TOPK promotes the growth of esophageal cancer in vitro and in vivo by enhancing YB1/eEF1A1 signal pathway. Cell Death Dis. 2023;14:364. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 16] [Reference Citation Analysis (0)] |
| 12. | Yuryev A, Wennogle LP. Novel raf kinase protein-protein interactions found by an exhaustive yeast two-hybrid analysis. Genomics. 2003;81:112-125. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 30] [Cited by in RCA: 33] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 13. | Degirmenci U, Wang M, Hu J. Targeting Aberrant RAS/RAF/MEK/ERK Signaling for Cancer Therapy. Cells. 2020;9:198. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 257] [Cited by in RCA: 418] [Article Influence: 69.7] [Reference Citation Analysis (0)] |
| 14. | Matsumoto S, Abe Y, Fujibuchi T, Takeuchi T, Kito K, Ueda N, Shigemoto K, Gyo K. Characterization of a MAPKK-like protein kinase TOPK. Biochem Biophys Res Commun. 2004;325:997-1004. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 75] [Cited by in RCA: 86] [Article Influence: 4.1] [Reference Citation Analysis (2)] |
| 15. | Hao Y, Baker D, Ten Dijke P. TGF-β-Mediated Epithelial-Mesenchymal Transition and Cancer Metastasis. Int J Mol Sci. 2019;20:2767. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 977] [Cited by in RCA: 896] [Article Influence: 128.0] [Reference Citation Analysis (5)] |
| 16. | Song M, Bode AM, Dong Z, Lee MH. AKT as a Therapeutic Target for Cancer. Cancer Res. 2019;79:1019-1031. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 285] [Cited by in RCA: 610] [Article Influence: 87.1] [Reference Citation Analysis (2)] |
| 17. | Carter BZ, Mak DH, Schober WD, Cabreira-Hansen M, Beran M, McQueen T, Chen W, Andreeff M. Regulation of survivin expression through Bcr-Abl/MAPK cascade: targeting survivin overcomes imatinib resistance and increases imatinib sensitivity in imatinib-responsive CML cells. Blood. 2006;107:1555-1563. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 75] [Cited by in RCA: 83] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 18. | Huang XL, Khan MI, Wang J, Ali R, Ali SW, Zahra QU, Kazmi A, Lolai A, Huang YL, Hussain A, Bilal M, Li F, Qiu B. Role of receptor tyrosine kinases mediated signal transduction pathways in tumor growth and angiogenesis-New insight and futuristic vision. Int J Biol Macromol. 2021;180:739-752. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 62] [Article Influence: 12.4] [Reference Citation Analysis (0)] |
| 19. | Jenkins LJ, Luk IY, Fairlie WD, Lee EF, Palmieri M, Schoffer KL, Tan T, Ng I, Vukelic N, Tran S, Tse JWT, Nightingale R, Alam Z, Chionh F, Iatropoulos G, Ernst M, Afshar-Sterle S, Desai J, Gibbs P, Sieber OM, Dhillon AS, Tebbutt NC, Mariadason JM. Genotype-Tailored ERK/MAPK Pathway and HDAC Inhibition Rewires the Apoptotic Rheostat to Trigger Colorectal Cancer Cell Death. Mol Cancer Ther. 2023;22:52-62. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 18] [Reference Citation Analysis (0)] |
| 20. | Herbert KJ, Ashton TM, Prevo R, Pirovano G, Higgins GS. T-LAK cell-originated protein kinase (TOPK): an emerging target for cancer-specific therapeutics. Cell Death Dis. 2018;9:1089. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 65] [Cited by in RCA: 67] [Article Influence: 8.4] [Reference Citation Analysis (5)] |
| 21. | Rizkallah R, Batsomboon P, Dudley GB, Hurt MM. Identification of the oncogenic kinase TOPK/PBK as a master mitotic regulator of C2H2 zinc finger proteins. Oncotarget. 2015;6:1446-1461. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 24] [Cited by in RCA: 29] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 22. | Matsuo Y, Park JH, Miyamoto T, Yamamoto S, Hisada S, Alachkar H, Nakamura Y. TOPK inhibitor induces complete tumor regression in xenograft models of human cancer through inhibition of cytokinesis. Sci Transl Med. 2014;6:259ra145. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 76] [Cited by in RCA: 106] [Article Influence: 9.6] [Reference Citation Analysis (0)] |
| 23. | Chen D, Cox J, Annam J, Weingart M, Essien G, Rathi KS, Rokita JL, Khurana P, Cuya SM, Bosse KR, Pilgrim A, Li D, Shields C, Laur O, Maris JM, Schnepp RW. LIN28B promotes neuroblastoma metastasis and regulates PDZ binding kinase. Neoplasia. 2020;22:231-241. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 27] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 24. | Roh E, Han Y, Reddy K, Zykova TA, Lee MH, Yao K, Bai R, Curiel-Lewandrowski C, Dong Z. Suppression of the solar ultraviolet-induced skin carcinogenesis by TOPK inhibitor HI-TOPK-032. Oncogene. 2020;39:4170-4182. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 21] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 25. | Hu QF, Gao TT, Shi YJ, Lei Q, Liu ZH, Feng Q, Chen ZJ, Yu LT. Design, synthesis and biological evaluation of novel 1-phenyl phenanthridin-6(5H)-one derivatives as anti-tumor agents targeting TOPK. Eur J Med Chem. 2019;162:407-422. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9] [Cited by in RCA: 14] [Article Influence: 1.8] [Reference Citation Analysis (0)] |