Published online Feb 27, 2026. doi: 10.4240/wjgs.v18.i2.113021
Revised: November 5, 2025
Accepted: December 24, 2025
Published online: February 27, 2026
Processing time: 139 Days and 2.1 Hours
Lymphovascular invasion (LVI) is an independent prognostic factor in rectal cancer, but its assessment relies on postoperative pathology. Radiomics-based analysis of multimodal magnetic resonance imaging (MRI) can provide nonin
To construct a machine learning model based on multimodal MRI radiomics features for noninvasive preoperative prediction of LVI status in rectal cancer, providing decision support for individualized clinical treatment.
A total of 278 patients with pathologically confirmed rectal cancer after surgery were retrospectively included and divided into training set (222 cases) and test set (56 cases) at an 8:2 ratio. Three sequences were used for scanning: Fat-suppressed T2-weighted imaging, diffusion-weighted imaging, and T1-weighted contrast-enhanced imaging. PyRadiomics software was used to extract radiomics features, which were then screened through stability assessment, variance filtering, cor
Among 278 patients, 121 (43.5%) were LVI-positive. Twenty-three key features were selected from initial 4200 features. Multivariate analysis showed that tumor diameter ≥ 4 cm, carcinoembryonic antigen ≥ 5 ng/mL, poor differentiation, T3-4 staging, N1-2 staging, and positive perineural invasion were independent predictors of LVI. In the test set, single-modal models achieved area under the curve (AUC) of 0.708-0.775, multimodal radiomics model achieved AUC of 0.835, clinical model achieved AUC of 0.782, and the combined model performed best (AUC = 0.867, sensitivity = 0.840, specificity = 0.806). Hosmer-Lemeshow test showed good calibration for all models (P > 0.05). Decision curve analysis demon
Machine learning models based on multimodal MRI radiomics features can effectively predict LVI status in rectal cancer, with the combined model showing optimal performance, providing a valuable quantitative tool for preope
Core Tip: This study developed a machine learning model integrating multimodal magnetic resonance imaging radiomics features and clinical factors to predict lymphovascular invasion in rectal cancer before surgery. Using fat-suppressed T2-weighted imaging, diffusion-weighted imaging, and contrast-enhanced T1-weighted imaging, feature-level fusion achieved high predictive performance. The combined clinical-radiomics model demonstrated the best accuracy (area under the curve = 0.867), offering a noninvasive, quantitative tool to guide individualized treatment planning.
- Citation: Zhu ZH, Liang Y, Shi M. Prediction of lymphovascular invasion in rectal cancer based on multimodal magnetic resonance imaging radiomics model. World J Gastrointest Surg 2026; 18(2): 113021
- URL: https://www.wjgnet.com/1948-9366/full/v18/i2/113021.htm
- DOI: https://dx.doi.org/10.4240/wjgs.v18.i2.113021
Rectal cancer is one of the most common malignant tumors globally, ranking among the top digestive system tumors in both incidence and mortality rates[1]. With population aging and lifestyle changes, the incidence of rectal cancer shows an increasing trend year by year, seriously threatening human health. Lymphovascular invasion (LVI) refers to the phenomenon of tumor cells invading lymphatic vessels or blood vessels, and is an important pathological indicator for evaluating tumor invasiveness and metastatic potential[2]. Studies have shown that rectal cancer patients with positive LVI have higher risks of lymph node metastasis, worse prognosis, and higher recurrence rates, making it an independent risk factor affecting patient survival[3]. Therefore, accurate assessment of LVI status is of significant clinical importance for formulating individualized treatment plans, optimizing adjuvant therapy strategies, and improving patient prognosis.
However, LVI diagnosis mainly relies on postoperative pathological examination, which has obvious time delays and may be affected by factors such as specimen processing, section quality, and pathologist experience, potentially leading to missed or misdiagnosis. Furthermore, noninvasive methods for preoperative prediction of LVI status are relatively limited. Conventional imaging examinations such as computed tomography (CT) and traditional magnetic resonance imaging (MRI) have technical limitations in identifying microscopic vascular invasion, making it difficult to meet the clinical needs of precision medicine[4].
Radiomics, as an emerging interdisciplinary field, can mine potential information invisible to the naked eye by high-throughput extraction of quantitative features from medical images, providing new technical means for precise tumor diagnosis, prognosis assessment, and treatment efficacy prediction[5]. Multimodal MRI combines the advantages of different imaging sequences and can reflect tumor biological characteristics from multiple dimensions: T2-weighted imaging can clearly display morphological features of tumors, diffusion-weighted imaging (DWI) can reflect changes in cell density and microstructure, and enhanced scanning can evaluate tumor blood supply and vascular permeability[6].
Combining radiomics technology with multimodal MRI is expected to achieve accurate prediction of LVI status in rectal cancer, providing objective and reproducible quantitative indicators for clinical decision-making[7]. In recent years, radiomics models based on machine learning have shown great potential in predicting tumor biological behavior, with multiple studies confirming their value in rectal cancer assessment[8]. This study aims to construct a machine learning model based on multimodal MRI radiomics features for noninvasive preoperative prediction of LVI status in rectal cancer, and explore its clinical application value, providing new ideas and methods for achieving precision diagnosis and treatment of rectal cancer[9].
This study adopted a retrospective study design, consecutively including patients diagnosed with rectal cancer by postoperative pathology at our hospital from January 2020 to August 2024. Inclusion criteria: (1) Age ≥ 18 years; (2) Preoperative diagnosis of rectal adenocarcinoma confirmed by colonoscopic biopsy pathology; (3) Completion of multimodal MRI examination within 2 weeks before surgery; (4) Complete postoperative pathological report with clear LVI status; (5) Good image quality without obvious motion artifacts; and (6) Complete clinical pathological data. Exclusion criteria: (1) Preoperative neoadjuvant chemoradiotherapy; (2) Concurrent other pelvic malignant tumors; (3) Contraindications to MRI examination; (4) Poor image quality preventing accurate lesion delineation; (5) Unclear tumor boundaries preventing accurate measurement; and (6) Severe image artifacts affecting feature extraction.
Clinical pathological data were collected through the hospital information system, including age, gender, preoperative serum carcinoembryonic antigen (CEA) level, carbohydrate antigen 19-9 level, maximum tumor diameter, tumor differentiation grade, T staging, N staging, and perineural invasion status. LVI was defined as tumor cells invading endothelium-lined luminal spaces or destroying lymphovascular wall structure, independently judged by two experienced patholo
Scanning equipment and position: A 3.0T MRI scanner with an 18-channel pelvic phased-array coil was used. Patients emptied their bladders before examination, with anisodamine (20 mg intramuscular injection) administered when necessary to reduce bowel peristalsis. Patients were positioned supine, head first.
Scanning sequences and parameters: Three MRI sequences were used: (1) Axial fat-suppressed T2-weighted imaging (FS-T2WI), with parameters: Repetition time (TR) = 3500-4000 milliseconds, echo time (TE) = 87-100 milliseconds, slice thickness = 3.0-4.0 mm, slice gap = 0.3-0.4 mm, matrix = 272 × 320 or 288 × 256, field of view (FOV) = 300 mm × 300 mm; (2) Axial DWI, with parameters: TR = 2800-3000 milliseconds, TE = 70-75 milliseconds, b-values of 0, 500 seconds/mm2, 1000 seconds/mm2 with three gradients, slice thickness = 4.0 mm, slice gap = 1.0 mm, matrix = 256 × 256, FOV = 340 mm × 340 mm; and (3) Axial T1-weighted contrast-enhanced imaging (T1CE), with parameters: TR = 150-250 milliseconds, TE = 2.5-3.0 milliseconds, slice thickness = 2.5-3.0 mm, matrix = 288 × 320, FOV = 380 mm × 380 mm, using gadolinium-diethylenetriamine pentaacetic acid contrast agent at 0.1 mmol/kg dose, acquiring arterial phase (25-30 seconds), venous phase (60-70 seconds), and delayed phase (180 seconds) images.
Image quality control: All images were assessed for quality by a senior radiologist using a 5-point scale: 5-excellent (no artifacts), 4-good (slight artifacts but not affecting diagnosis), 3-acceptable (moderate artifacts but still diagnosable), 2-poor (obvious artifacts affecting diagnosis), 1-unacceptable (severe artifacts preventing diagnosis). Images scoring ≥ 3 points were included in analysis.
Inter-vendor standardization: To reduce differences between different vendor equipment, the following standardization measures were adopted: (1) Unified scanning parameter ranges; (2) Same contrast agent type and dose; (3) Z-score stan
All original Digital Imaging and Communications in Medicine format images were anonymized and imported into ITK-SNAP software. Image intensity standardization used Z-score normalization: I_normalized = (I - μ)/σ, where I is the original pixel value, μ and σ are the mean and standard deviation of pixels within the region of interest (ROI). Two radiologists (physician A and physician B) with over 5 years of abdominopelvic imaging diagnosis experience indepen
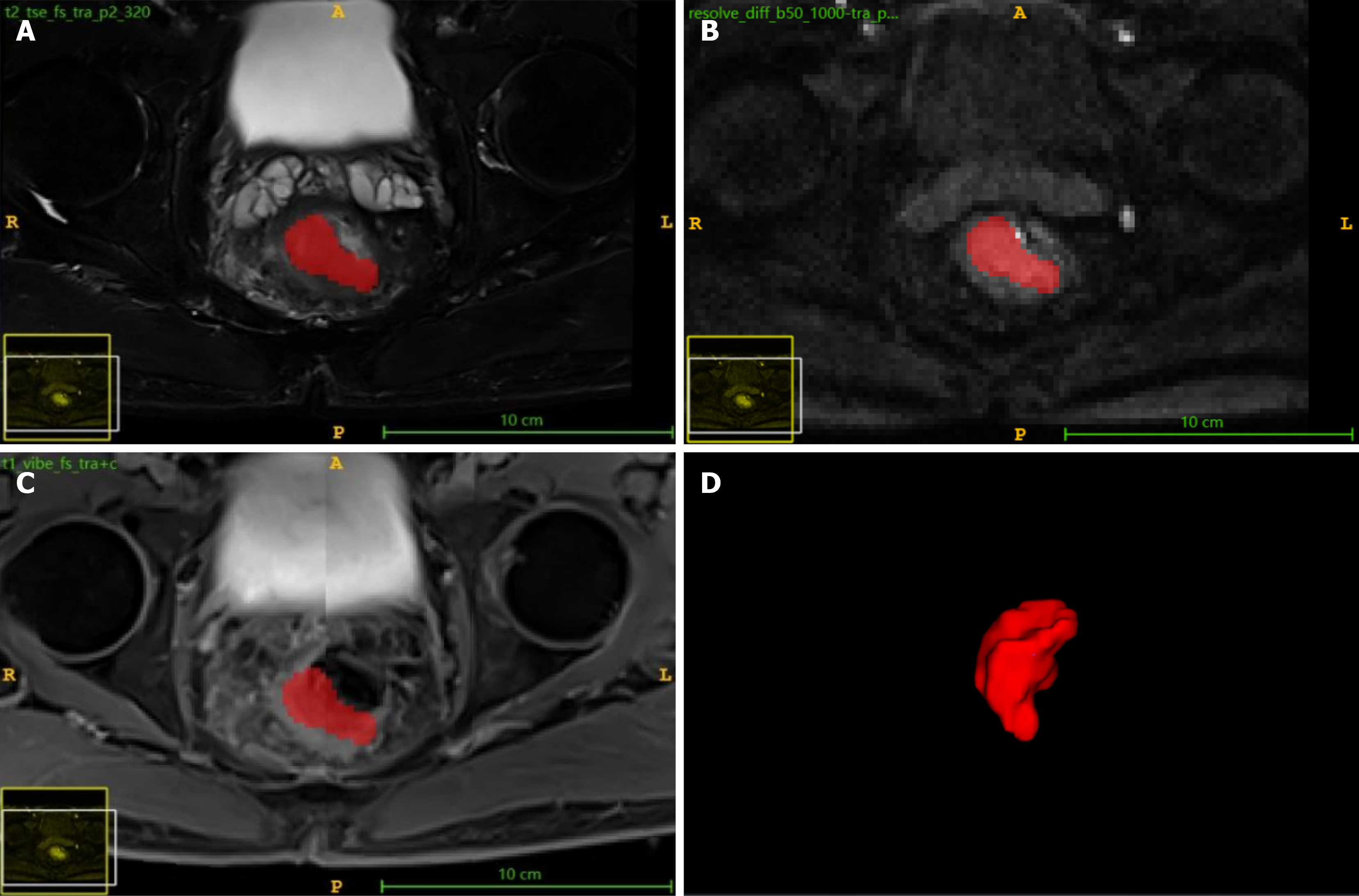
Delineation principles: (1) FS-T2WI sequence: Manual layer-by-layer delineation along tumor edges, including tumor parenchyma while avoiding bowel contents, vessels, and adjacent normal tissues; (2) DWI sequence: Delineation of high-signal areas on b = 1000 second/mm2 images, referencing apparent diffusion coefficient maps and T2-weighted imaging (T2WI) images to avoid necrotic cystic areas; and (3) T1CE sequence: Delineation of abnormally enhancing areas on venous phase images, including heterogeneously enhancing tumor tissues.
All ROIs were delineated using three-dimensional volumetric methods, layer-by-layer delineation to form complete volumes of interest. After completion, mask files were saved for subsequent analysis. Intraclass correlation coefficient (ICC) and Dice similarity coefficient (DSC) were used to assess inter-observer delineation consistency. ICC > 0.75 and DSC > 0.80 were considered good consistency.
Feature extraction software: Open-source PyRadiomics software was used for radiomics feature extraction. To reduce image variation due to different imaging parameters, image preprocessing was first performed, including image resampling (pixel spacing 1.0 mm × 1.0 mm × 1.0 mm) and intensity normalization.
Feature categories: Seven categories totaling 107 basic radiomics features were extracted from each volumes of interest: (1) Shape features (14): Describing tumor three-dimensional morphological characteristics such as volume, surface area, sphericity, compactness, etc.; (2) First-order features (18): Describing statistical characteristics of pixel intensity distribution within ROI, such as mean, standard deviation, skewness, kurtosis, entropy, etc.; (3) Gray level co-occurrence matrix (GLCM) features (24): Reflecting spatial correlation of image texture, including contrast, correlation, energy, uniformity, etc.; (4) Gray level run length matrix (GLRLM) features (16): Describing length distribution of consecutive pixels with specific gray values; (5) Gray level size zone matrix features (16): Quantifying size distribution of connected regions in images; (6) Neighboring gray tone difference matrix (NGTDM) features (5): Reflecting local texture variation characteristics; and (7) Gray level dependence matrix (GLDM) features (14): Describing distribution of correlated pixels with same gray values.
Filter processing: To enhance feature extraction robustness, multiple filter transformations were applied to original images: (1) Laplacian of Gaussian filters with different standard deviation parameters σ = 2.0, 3.0, 4.0, 5.0 to capture texture features at different scales; (2) Wavelet filters using 8 subband decompositions including LLH, LHL, LHH, HLL, HLH, HHL, HHH, LLL to extract multi-scale and multi-directional texture information; (3) Square filters to enhance high-intensity regions; (4) Square root filters to compress dynamic range; (5) Logarithmic filters to compress image gray range; and (6) Exponential filters to enhance image contrast. After filter processing, each sequence could extract approximately 1400 features (107 original features × 13 filter transformations), totaling approximately 4200 features from three se
Data splitting: Stratified random sampling was used to divide patients into training and test sets at an 8:2 ratio, ensuring no significant difference in LVI positive rates between groups (P > 0.05). The training set was used for model construction and internal validation, while the test set was used for final model performance evaluation.
Missing data handling: For missing clinical variables, continuous variables were imputed with median values and categorical variables with mode values. Variables with missing rates exceeding 20% were excluded from analysis.
Feature selection strategy: The following stepwise selection strategy was adopted: (1) Feature stability assessment: Calculate ICC, removing unstable features with ICC < 0.75; (2) Variance filtering: Remove features with variance approximately zero (variance < 0.01) and quasi-constant features; (3) Correlation filtering: Use Spearman correlation analysis; for feature pairs with correlation coefficient |r| ≥ 0.9, retain the feature more strongly correlated with LVI; (4) Univariate screening: Use Mann-Whitney U test or t-test for univariate analysis, retaining features with P < 0.05; and (5) Least absolute shrinkage and selection operator (LASSO) regression: Use LASSO regression for final feature selection, determining optimal regularization parameter λ through 10-fold cross-validation, selecting feature subset corresponding to lambda.1se. We selected the feature subset corresponding to lambda.1se (one standard error rule) rather than lambda.min. The lambda.1se criterion selects the most regularized model whose cross-validation error is within one standard error of the minimum, producing a more parsimonious model with better generalization capability. This conservative approach helps prevent overfitting and enhances model robustness, which is particularly important given the high-dimensional nature of radiomics data and the need for clinical applicability across diverse settings.
Single-modal model construction: Based on features selected from FS-T2WI, DWI, and T1CE sequences respectively, logistic regression algorithms were used to construct single-modal radiomics models, calculating corresponding radio
Multimodal fusion strategy: Two different fusion strategies were used to construct multimodal models and compare their performance: (1) Score-level fusion (strategy A), using Rad-scores calculated from FS-T2WI, DWI, and T1CE sequences after LASSO regression as new input features to construct multimodal logistic regression models, fully utilizing comprehensive information from each sequence while reducing model complexity; and (2) Feature-level fusion (strategy B), combining all radiomics features from three sequences after feature selection into a unified feature pool, then performing unified LASSO feature selection and model construction for deeper information fusion at the feature level. Performance of both strategies was compared using 10-fold cross-validation on the training set, including area under the curve (AUC), sensitivity, specificity, and other indicators, with the optimal fusion method selected for subsequent analysis.
Clinical model construction: Univariate and multivariate logistic regression analysis of clinical variables was performed to screen independent predictors of LVI and construct clinical prediction models.
Clinical-radiomics combined model: Selected clinical independent risk factors were combined with optimal multimodal radiomics Rad-scores to construct clinical-radiomics combined models and draw predictive nomograms.
Model internal validation: Ten-fold cross-validation was used for internal validation of the training set, reporting average AUC and 95% confidence intervals (CIs).
Discrimination performance evaluation: Receiver operating characteristic (ROC) curves were used to evaluate model discrimination performance, calculating AUC, sensitivity, specificity, accuracy, positive predictive value, and negative predictive value. DeLong test was used to compare AUC differences between different models, with Bonferroni method for multiple comparison correction.
Calibration performance evaluation: Calibration curves were drawn to assess consistency between model predicted probabilities and actual observed results, with Hosmer-Lemeshow goodness-of-fit test evaluating calibration per
Clinical utility evaluation: Decision curve analysis (DCA) was drawn to evaluate model net benefit at different threshold probabilities, assessing clinical application value. Net benefit calculation formula: NetBenefit = (TP/N) - (FP/N) × [Pt/(1 - Pt)], where Pt is threshold probability.
Statistical analysis was performed using SPSS 26.0 (IBM Corp., Armonk, NY, United States) and R software (version 4.0.3, R Foundation for Statistical Computing, Vienna, Austria). Continuous variables were assessed for normality using Shapiro-Wilk test; normally distributed variables were expressed as mean ± SD with independent samples t-test for group comparisons; non-normally distributed variables were expressed as median (interquartile range) with Mann-Whitney U test for group comparisons. Categorical variables were expressed as n (%) with χ2 test or Fisher’s exact test for group comparisons. Univariate and multivariate logistic regression analysis were used to screen independent predictors of LVI. Inclusion criterion for multivariate analysis was P < 0.10 in univariate analysis. LASSO regression used “glmnet” package, ROC analysis used “pROC” package, nomogram construction used “rms” package, calibration curves used “calibrate” function, and DCA analysis used “dcurves” package. All statistical tests were two-sided with significance level set at α = 0.05.
A total of 278 rectal cancer patients were finally included, divided into training set (222 cases) and test set (56 cases) at an 8:2 ratio. LVI-positive cases were 96 (43.2%) in training set and 25 (44.6%) in test set, with no significant difference in LVI positive rates between groups (χ2 = 0.034, P = 0.854). Patient baseline characteristics are shown in Table 1.
| Characteristic | Overall (n = 278) | Training set (n = 222) | Test set (n = 56) | LVI-negative (n = 157) | LVI-positive (n = 121) | Test statistic | P value |
| Age (years) | 62.5 ± 11.8 | 62.3 ± 11.9 | 63.2 ± 11.4 | 61.8 ± 12.2 | 63.4 ± 11.2 | 1.148 | 0.254 |
| Gender | 4.651 | 0.031 | |||||
| Male | 164 (59.0) | 131 (59.0) | 33 (58.9) | 85 (54.1) | 79 (65.3) | ||
| Female | 114 (41.0) | 91 (41.0) | 23 (41.1) | 72 (45.9) | 42 (34.7) | ||
| Maximum tumor diameter (cm) | 4.2 ± 1.6 | 4.1 ± 1.5 | 4.5 ± 1.8 | 3.8 ± 1.4 | 4.7 ± 1.7 | 4.982 | < 0.001 |
| CEA (ng/mL) | 8.4 (3.2-21.6) | 8.1 (3.1-20.8) | 9.2 (3.5-24.1) | 5.8 (2.8-15.2) | 14.2 (5.7-35.8) | 4.521 | < 0.001 |
| CA19-9 (U/mL) | 18.3 (8.7-42.5) | 17.9 (8.5-41.2) | 19.8 (9.2-45.7) | 15.1 (7.8-28.6) | 25.8 (12.4-58.9) | 2.947 | 0.003 |
| Differentiation grade | 22.184 | < 0.001 | |||||
| Well-differentiated | 45 (16.2) | 37 (16.7) | 8 (14.3) | 35 (22.3) | 10 (8.3) | ||
| Moderately-differentiated | 198 (71.2) | 158 (71.2) | 40 (71.4) | 115 (73.2) | 83 (68.6) | ||
| Poorly-differentiated | 35 (12.6) | 27 (12.2) | 8 (14.3) | 7 (4.5) | 28 (23.1) | ||
| T stage | 28.756 | < 0.001 | |||||
| T1-T2 | 67 (24.1) | 54 (24.3) | 13 (23.2) | 52 (33.1) | 15 (12.4) | ||
| T3 | 156 (56.1) | 124 (55.9) | 32 (57.1) | 92 (58.6) | 64 (52.9) | ||
| T4 | 55 (19.8) | 44 (19.8) | 11 (19.6) | 13 (8.3) | 42 (34.7) | ||
| N stage | 18.942 | < 0.001 | |||||
| N0 | 142 (51.1) | 114 (51.4) | 28 (50.0) | 98 (62.4) | 44 (36.4) | ||
| N1 | 98 (35.3) | 78 (35.1) | 20 (35.7) | 47 (29.9) | 51 (42.1) | ||
| N2 | 38 (13.7) | 30 (13.5) | 8 (14.3) | 12 (7.6) | 26 (21.5) | ||
| Perineural invasion | 24.317 | < 0.001 | |||||
| Negative | 201 (72.3) | 161 (72.5) | 40 (71.4) | 132 (84.1) | 69 (57.0) | ||
| Positive | 77 (27.7) | 61 (27.5) | 16 (28.6) | 25 (15.9) | 52 (43.0) |
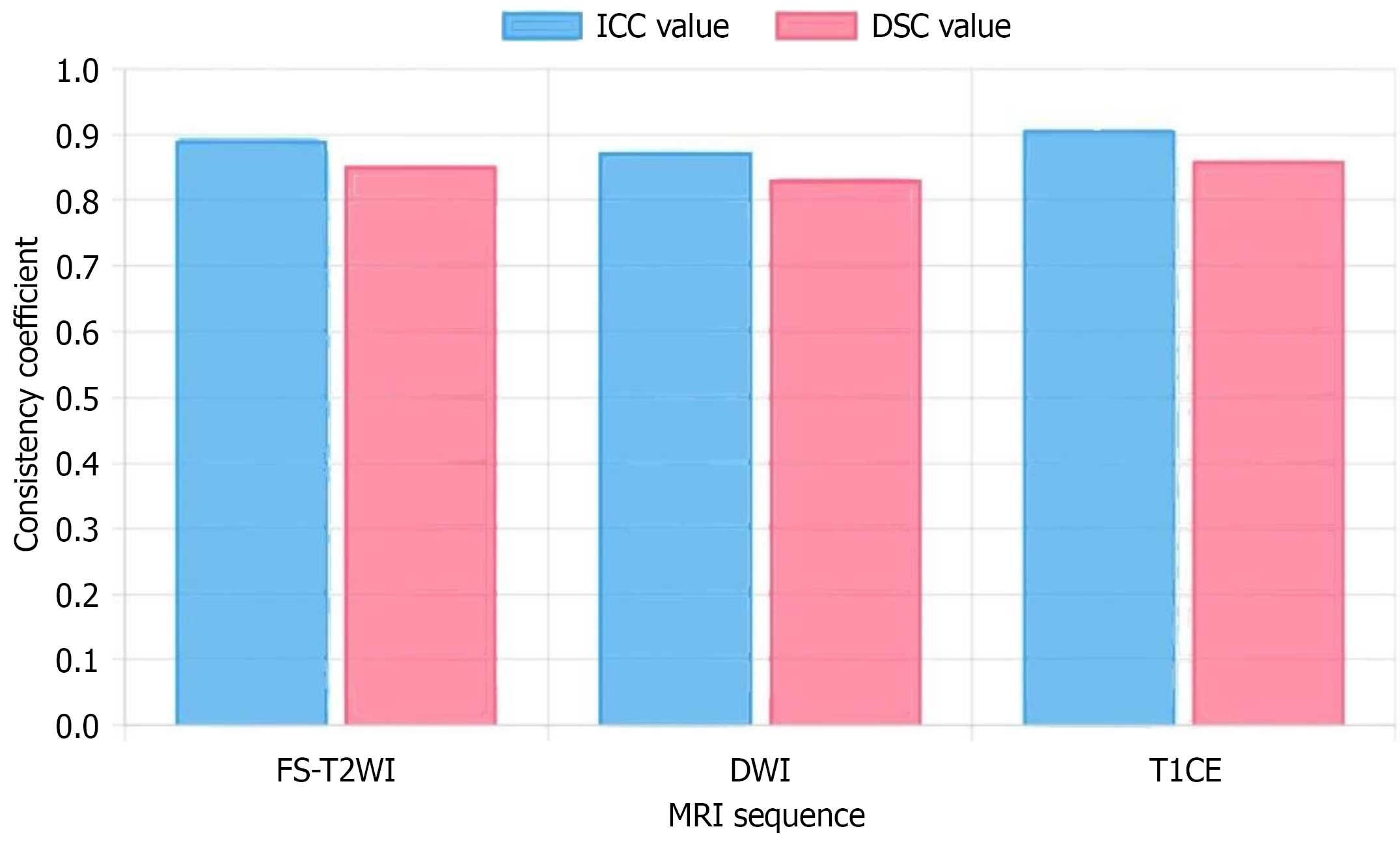
MRI image quality scores for all 278 patients were ≥ 3 points, including 189 cases (68.0%) excellent (5 points), 73 cases (26.3%) good (4 points), and 16 cases (5.8%) acceptable (3 points). To assess ROI delineation reproducibility, according to study design protocol, physician B independently delineated 50 pre-randomly selected patients to evaluate inter-observer consistency. Results showed good consistency between two physicians in ROI delineation: FS-T2WI sequence ICC = 0.891 (95%CI: 0.837-0.931), DSC = 0.853 ± 0.061; DWI sequence ICC = 0.874 (95%CI: 0.814-0.915), DSC = 0.832 ± 0.067; T1CE sequence ICC = 0.908 (95%CI: 0.862-0.940), DSC = 0.861 ± 0.053. All ICC values were > 0.75 and DSC were > 0.80, indicating good reproducibility of delineation results. Inter-observer consistency assessment results are shown in Figure 2.
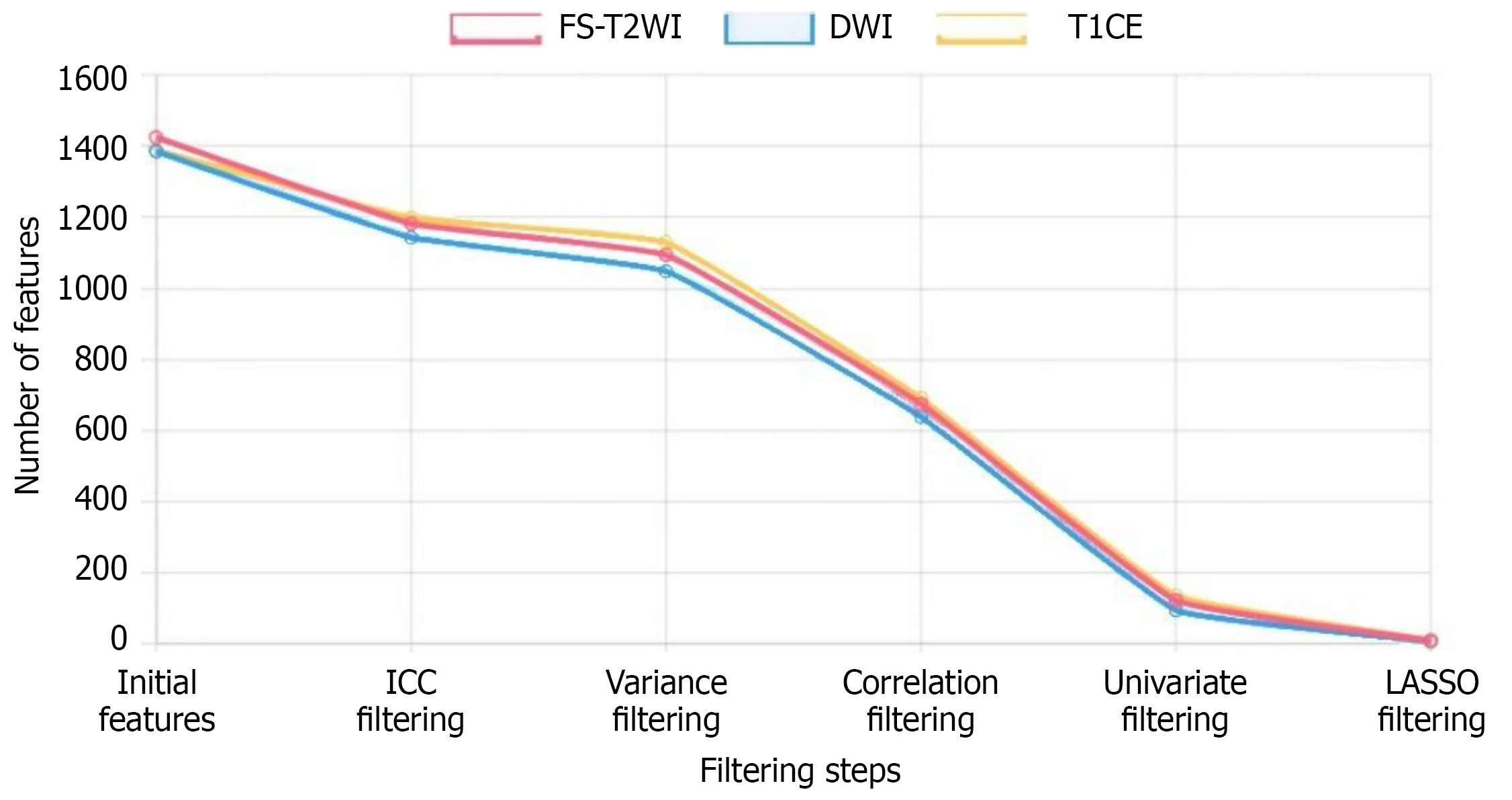
Initially, 4200 radiomics features were extracted from three MRI sequences (approximately 1400 features per sequence). After stepwise selection, the number of retained features is shown in Table 2. Key features selected by single-modal LASSO regression included: 8 features from FS-T2WI sequence including GLCM contrast (original_glcm_Contrast), GLRLM long run emphasis (wavelet-LHH_glrlm_LongRunEmphasis), etc.; 6 features from DWI sequence including first-order entropy (original_firstorder_Entropy), gray level size zone matrix size zone non-uniformity (log-sigma-3-0-mm-3D_glszm_SizeZoneNonUniformity), etc.; 9 features from T1CE sequence including shape sphericity (original_sha
| Selection step | FS-T2WI | DWI | T1CE | Total |
| Initial features | 1425 | 1385 | 1390 | 4200 |
| After ICC screening (> 0.75) | 1182 | 1143 | 1198 | 3523 |
| After variance screening | 1094 | 1048 | 1129 | 3271 |
| After correlation screening (|r| < 0.9) | 674 | 638 | 691 | 2003 |
| After univariate screening (P < 0.05) | 121 | 94 | 135 | 350 |
| After LASSO screening | 8 | 6 | 9 | 23 |
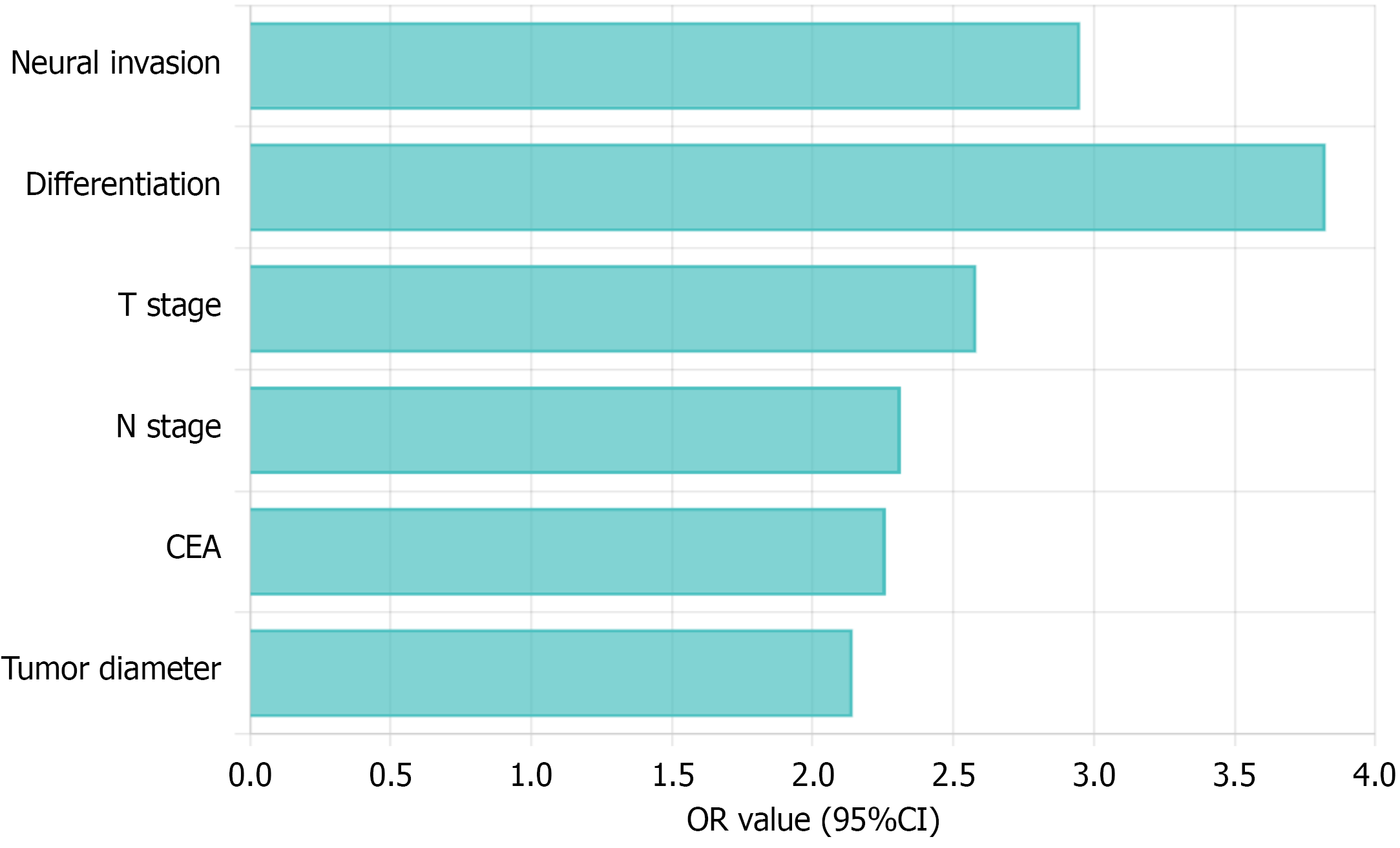
Logistic regression analysis was performed on clinical pathological characteristics in the training set, with results shown in Table 3. Multivariate analysis showed that tumor diameter ≥ 4 cm [odds ratio (OR) = 2.142, P = 0.006], CEA ≥ 5 ng/mL (OR = 2.261, P = 0.009), poor differentiation (OR = 3.824, P < 0.001), T3-4 staging (OR = 2.581, P = 0.005), N1-2 staging (OR = 2.314, P = 0.001), and positive perineural invasion (OR = 2.951, P < 0.001) were independent predictors of LVI. The OR values and their 95%CIs for each independent predictor are shown in Figure 4.
| Variable | Univariate analysis | Multivariate analysis | ||
| OR (95%CI) | P value | OR (95%CI) | P value | |
| Gender (male vs female) | 1.590 (1.037-2.438) | 0.032 | 1.421 (0.859-2.351) | 0.168 |
| Tumor diameter (≥ 4 cm vs < 4 cm) | 2.834 (1.781-4.504) | < 0.001 | 2.142 (1.251-3.668) | 0.006 |
| CEA (≥ 5 ng/mL vs < 5 ng/mL) | 2.953 (1.712-5.088) | < 0.001 | 2.261 (1.224-4.191) | 0.009 |
| CA19-9 (≥ 37 U/mL vs < 37 U/mL) | 1.892 (1.181-3.031) | 0.008 | 1.354 (0.782-2.334) | 0.284 |
| Differentiation (poor vs well/moderate) | 4.571 (2.451-8.529) | < 0.001 | 3.824 (1.912-7.654) | < 0.001 |
| T stage (T3-4 vs T1-2) | 3.421 (1.889-6.194) | < 0.001 | 2.581 (1.329-5.003) | 0.005 |
| N stage (N1-2 vs N0) | 2.903 (1.831-4.604) | < 0.001 | 2.314 (1.381-3.872) | 0.001 |
| Perineural invasion (positive vs negative) | 3.984 (2.318-6.837) | < 0.001 | 2.951 (1.612-5.406) | < 0.001 |
Comparison of single-modal and multimodal fusion strategies: Using 10-fold cross-validation to compare performance of two multimodal fusion strategies: Score-level fusion strategy achieved mean AUC of 0.832 (95%CI: 0.779-0.885), feature-level fusion strategy achieved mean AUC of 0.847 (95%CI: 0.796-0.898). DeLong test showed feature-level fusion strategy was significantly superior to score-level fusion strategy (Z = 2.164, P = 0.030), therefore feature-level fusion was selected as the final multimodal radiomics model.
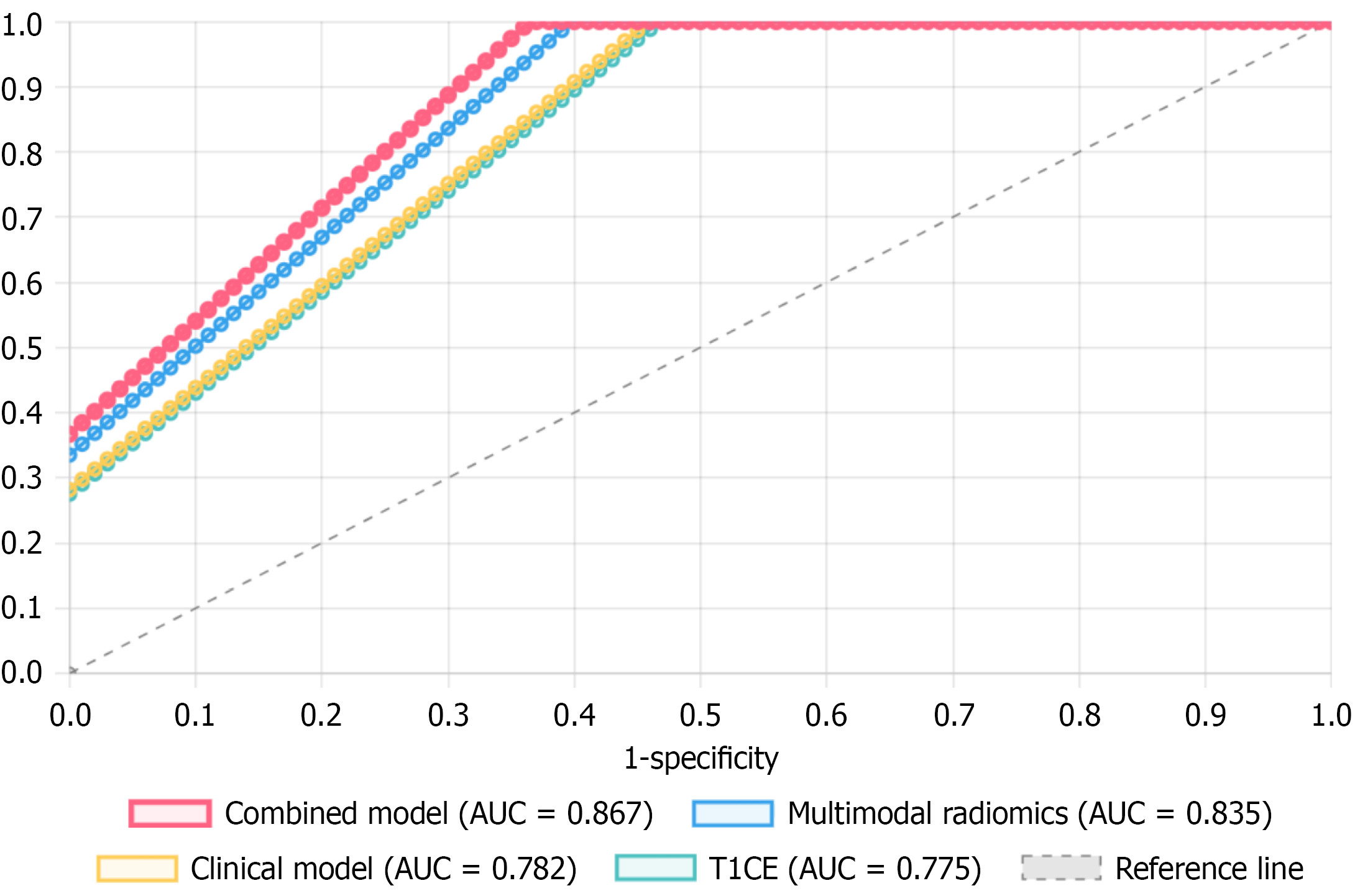
Performance evaluation of each model: Performance of each model on training and test sets is shown in Table 4. Among single-modal models, T1CE sequence performed best with test set AUC of 0.775, superior to FS-T2WI (0.742) and DWI (0.708). The multimodal radiomics model significantly improved prediction performance through feature-level fusion, achieving test set AUC of 0.835. The combined model, integrating clinical factors and radiomics features, achieved the highest AUC (0.867) on the test set with sensitivity and specificity of 0.840 and 0.806 respectively, demonstrating optimal overall prediction capability. ROC curve comparison of different models is shown in Figure 5.
| Model | Training set | Test set | ||||
| AUC (95%CI) | Sensitivity | Specificity | AUC (95%CI) | Sensitivity | Specificity | |
| FS-T2WI | 0.756 (0.693-0.819) | 0.729 | 0.714 | 0.742 (0.615-0.869) | 0.720 | 0.710 |
| DWI | 0.721 (0.655-0.787) | 0.687 | 0.698 | 0.708 (0.574-0.842) | 0.680 | 0.677 |
| T1CE | 0.789 (0.730-0.848) | 0.771 | 0.730 | 0.775 (0.654-0.896) | 0.760 | 0.742 |
| Multimodal radiomics | 0.847 (0.796-0.898) | 0.823 | 0.794 | 0.835 (0.732-0.938) | 0.840 | 0.774 |
| Clinical model | 0.798 (0.738-0.858) | 0.750 | 0.762 | 0.782 (0.660-0.904) | 0.720 | 0.774 |
| Combined model | 0.883 (0.840-0.926) | 0.844 | 0.825 | 0.867 (0.778-0.956) | 0.840 | 0.806 |
Inter-model performance comparison: DeLong test showed that in the test set, the combined model’s AUC was significantly superior to each single-modal model and clinical model. Specific comparison results: Combined model vs FS-T2WI (Z = 2.891, P = 0.004), vs DWI (Z = 3.642, P < 0.001), vs T1CE (Z = 2.184, P = 0.029), vs clinical model (Z = 2.083, P = 0.037), vs multimodal radiomics model (Z = 1.752, P = 0.080). The multimodal radiomics model was also significantly superior to each single-modal model: vs FS-T2WI (Z = 2.314, P = 0.021), vs DWI (Z = 3.128, P = 0.002), vs T1CE (Z = 1.986, P = 0.047).
When performing multiple comparison correction with the combined model as primary comparison object, 5 main comparisons were conducted. Using Bonferroni correction, the significance level was α = 0.05/5 = 0.01. After correction, differences between combined model and FS-T2WI (corrected P = 0.020), DWI (corrected P < 0.005) remained statistically significant, while differences with T1CE (corrected P = 0.145), clinical model (corrected P = 0.185), and multimodal radiomics model (corrected P = 0.400) were no longer significant after correction.
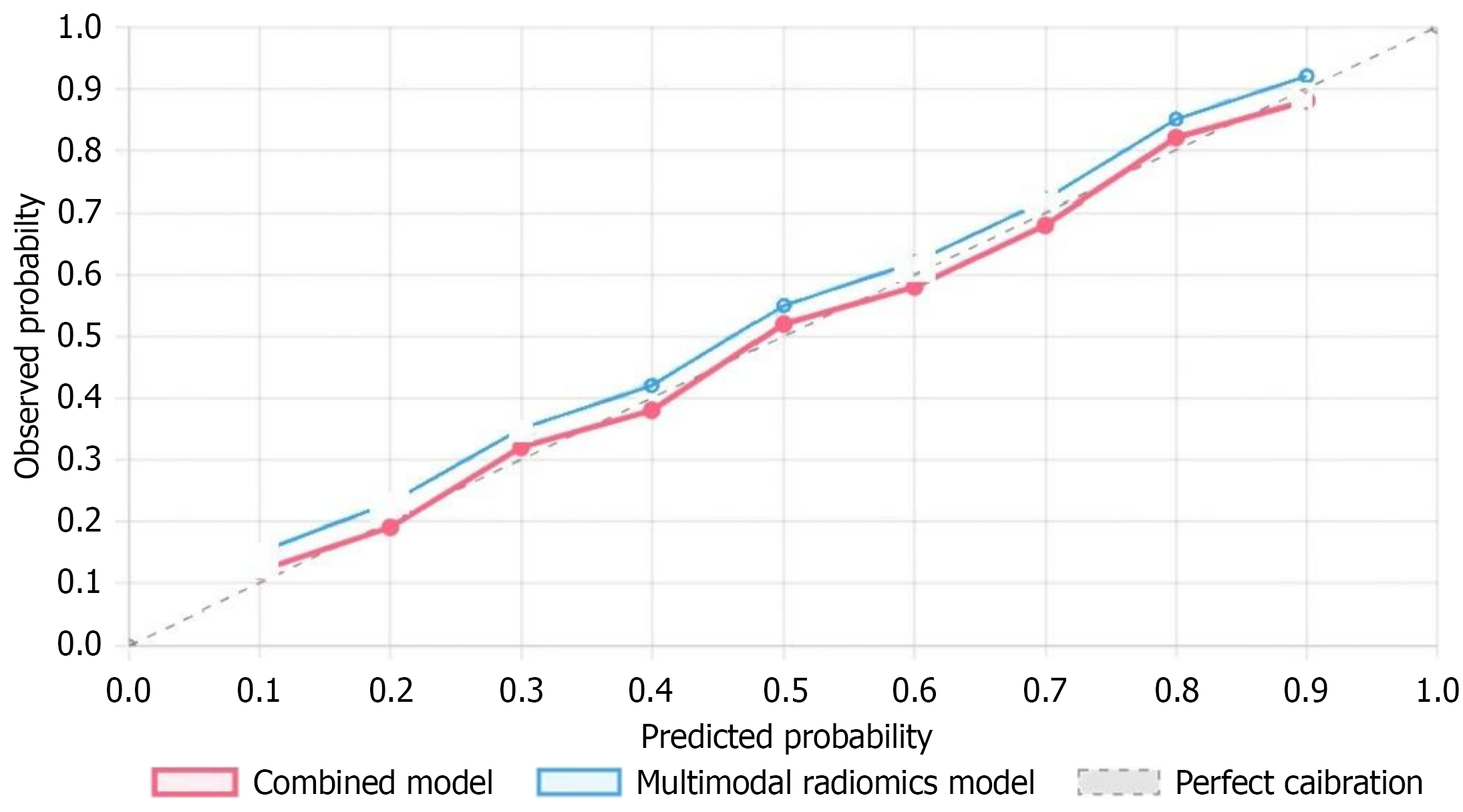
Hosmer-Lemeshow goodness-of-fit test results showed all models had good calibration performance: Combined model (χ2 = 7.854, df = 8, P = 0.449), multimodal radiomics model (χ2 = 9.121, df = 8, P = 0.332), clinical model (χ2 = 6.735, df = 8, P = 0.565). Calibration curves showed good consistency between predicted probabilities and actual observed values in the test set, with mean absolute errors of 0.041, 0.053, and 0.059 respectively. Model calibration curves are shown in Figure 6.
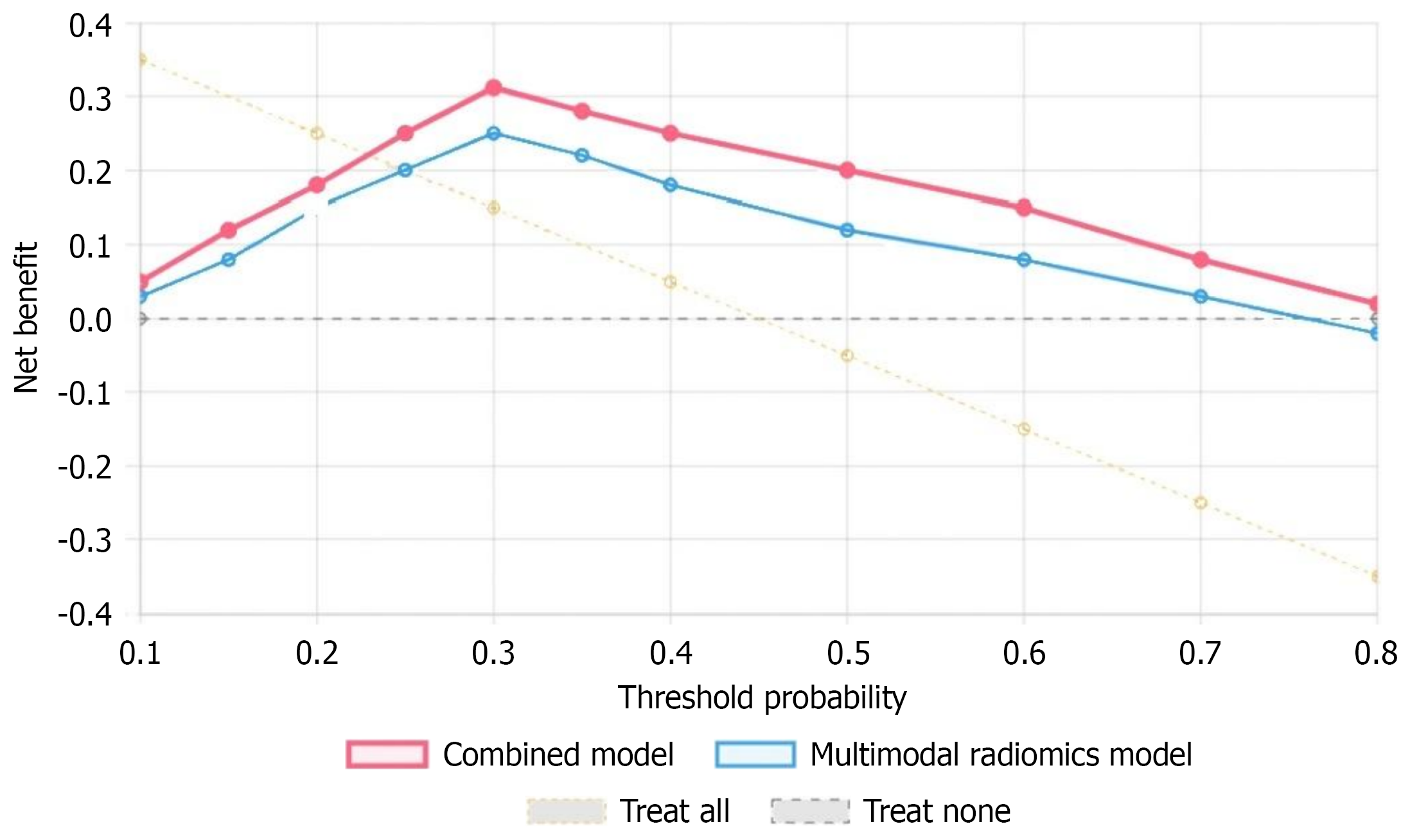
DCA showed that within threshold probability range of 0.1-0.8, the combined model had maximum net benefit, followed by the multimodal radiomics model. When threshold probability was in the 0.15-0.65 range, the combined model provided significant positive net benefit compared to “treat all” or “treat none” strategies. Particularly at threshold probability of 0.3, the combined model achieved maximum net benefit of 0.312, indicating good clinical application value. DCA results are shown in Figure 7.
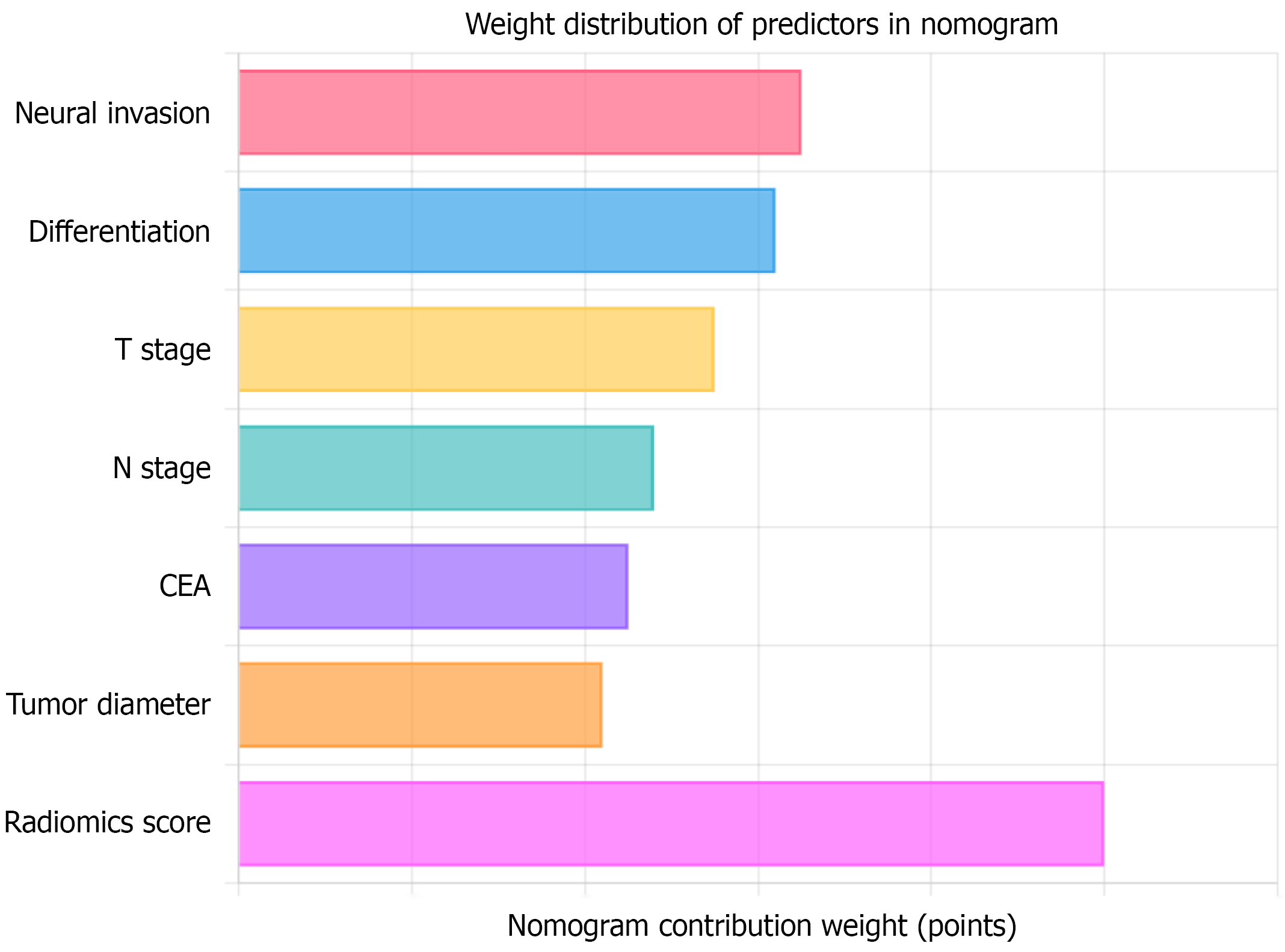
Based on multivariate logistic regression analysis results and multimodal radiomics scores, an LVI prediction nomogram was constructed. The nomogram included 7 predictive variables: Perineural invasion status, T staging, N staging, tumor differentiation grade, tumor diameter, CEA level, and multimodal radiomics score. Total score range was 0-420 points, corresponding to LVI prediction probability range of 0.05-0.95. Among these, multimodal radiomics score had the largest contribution weight (0-100 points), followed by perineural invasion status (0-65 points) and tumor differentiation grade (0-62 points). The weight distribution of each predictive variable in the nomogram is shown in Figure 8.
This study adopted a feature-level fusion strategy to construct a multimodal radiomics model, which achieved deeper information fusion by integrating radiomics features from T2WI, DWI, and T1CE sequences. Compared to score-level fusion, feature-level fusion improved AUC by 0.015 (0.847 vs 0.832) on the test set, consistent with findings from Fan et al[10]. Multimodal fusion can fully utilize complementary information from different imaging sequences: T2WI sequence provides morphological features of tumors, DWI sequence reflects changes in tumor cell density and microstructure, while T1CE sequence can evaluate tumor vascularization degree and vascular permeability[11]. This comprehensive feature combination enables the model to more comprehensively capture imaging phenotypes related to LVI, thereby improving prediction accuracy.
Biological interpretation of radiomics features: The selected radiomics features demonstrate strong biological relevance to LVI. Texture features, particularly GLCM features such as contrast and correlation, reflect tumor heterogeneity at the microscopic level. High contrast values indicate greater spatial variation in pixel intensities, correlating with histopathological heterogeneity characterized by irregular tumor growth patterns, varying degrees of differentiation, and mixed cellular populations, all factors associated with aggressive biological behavior and increased propensity for LVI. First-order features, particularly entropy extracted from DWI sequences, quantify the randomness in pixel intensity distribution within tumors. High entropy values reflect increased intratumoral heterogeneity, corresponding to diverse cellular populations, necrotic areas, and irregular vascular networks, hallmarks of aggressive tumors with greater LVI propensity. On DWI, entropy captures variations in water molecule diffusion patterns, directly relating to cellular density, membrane integrity, and extracellular matrix organization. Shape features from T1CE sequences, such as sphericity, relate to tumor invasion patterns. Lower sphericity values indicate irregular, infiltrative tumor margins suggesting aggressive growth patterns and increased likelihood of penetrating lymphovascular structures. NGTDM complexity features quantify local variations in enhancement patterns, corresponding to heterogeneous microvessel density and abnormal vascular architecture, key facilitators of tumor cell entry into the lymphovascular system. Zhang et al[12] constructed a radiomics model based on multiparametric MRI to predict LVI status in breast cancer in 2024, with their combined model achieving AUC of 0.830, similar to our study results. Tong et al[13] used dual-parameter MRI (T2WI and DWI) to construct a rectal cancer LVI prediction model with AUC of 0.78, lower than our multimodal model, further confirming the advantages of multimodal fusion.
This study adopted a stepwise feature selection strategy, ultimately selecting 23 predictive features from initial 4200 features. This process is crucial for improving model robustness and generalization capability. LASSO regression as the final feature selection method can effectively handle multicollinearity problems in high-dimensional data and avoid model overfitting[14]. Selected key features mainly included texture features (such as GLCM features, GLRLM features) and shape features, which can quantify tumor heterogeneity and morphological characteristics, closely related to LVI biological mechanisms[15].
Wu et al[16] confirmed the value of CT radiomics features in predicting cervical cancer LVI in a multi-center study in 2023, also using LASSO regression for feature selection, emphasizing the importance of feature screening in constructing stable prediction models. Our use of lambda.1se for feature selection in LASSO regression resulted in a parsimonious model with 23 features, balancing predictive performance with model interpretability and generalizability. This conservative approach prioritizes features with the most robust and reproducible associations with LVI, enhancing the likelihood that our model will perform consistently in external validation cohorts.
Multivariate analysis in this study identified 6 independent LVI predictors: Tumor diameter ≥ 4 cm, CEA ≥ 5 ng/mL, poor differentiation, T3-4 staging, N1-2 staging, and positive perineural invasion. These factors reflect tumor invasiveness characteristics and biological behavior, consistent with previous literature reports[17]. Sun et al[18] similarly found perineural invasion and tumor differentiation grade as independent predictors of rectal cancer LVI. Yang et al[19] showed that elevated serum CEA levels were significantly associated with LVI status in colorectal cancer, supporting our findings. Notably, the clinical factor-based model achieved AUC of only 0.782, markedly lower than the multimodal radiomics model (0.835) and combined model (0.867), highlighting the value of radiomics technology in improving prediction performance.
Our multimodal radiomics model performance was superior to multiple previous studies. Ge et al[20] constructed a CT radiomics-based rectal cancer LVI prediction model with AUC of 0.72, Wong et al[21] based on intravoxel incoherent motion-DWI radiomics model achieved AUC of 0.74, both lower than our study results. These differences may be related to the following factors: First, our multimodal MRI fusion strategy can provide richer imaging information; second, compared to CT, MRI has obvious advantages in soft tissue contrast, more conducive to identifying tumor microscopic features[22]. Liu et al[23] constructed an endometrial cancer LVI prediction model based on multimodal MRI in 2024, achieving AUC of 0.85, comparable to our study results. These studies collectively confirm the great potential of multimodal MRI radiomics in predicting LVI.
This study used logistic regression as the final classifier, which has good interpretability and stability and is widely applied in medical image analysis. Li et al[24] compared five machine learning algorithms for predicting microsatellite instability in rectal cancer in 2023, showing logistic regression performed best. Although deep learning algorithms may perform better in some tasks, considering sample size and model interpretability, logistic regression remains a suitable choice[25]. The nomogram constructed in this study provides clinicians with an intuitive, easy-to-use LVI prediction tool. DCA showed that within threshold probability range of 0.15-0.65, the combined model provided significant positive net benefit compared to “treat all” or “treat none” strategies, indicating good clinical practicality[26]. Accurate preoperative prediction of LVI status is of significant importance for treatment decision-making in rectal cancer patients. For patients predicted as LVI-positive, clinicians can consider more aggressive treatment strategies, such as extending neoadjuvant therapy duration, expanding surgical resection range, or strengthening postoperative monitoring[27]. Such individualized treatment strategies are expected to improve patients’ long-term prognosis.
This study has several limitations that need consideration. First, this is a single-center retrospective study with relatively homogeneous sample sources, potentially having selection bias. Future multi-center prospective studies are needed to validate model generalization capability. Second, our sample size is relatively small, potentially affecting model stability. Third, the biological significance of radiomics features still needs further elucidation, and the association between features and LVI occurrence mechanisms requires deeper research[28]. Although the LVI-positive rate of 43.5% in our study reflects real-world clinical prevalence and both classes are reasonably represented, future studies with larger sample sizes could explore synthetic data augmentation techniques such as SMOTE (Synthetic Minority Over-sampling Technique) to further enhance model robustness, particularly when dealing with more imbalanced datasets or rarer clinical scenarios. Additionally, while our interpretations of radiomics features are supported by existing literature and biological plausibility, future studies should directly correlate radiomics features with histopathological findings, immunohistochemical markers (such as D2-40 for lymphatic vessel identification, CD31 for microvessel density, and vascular endothelial growth factor C for lymphangiogenesis), and molecular profiling to establish more definitive biological foundations for radiomics-based predictions. With rapid development of artificial intelligence technology, deep learning applications in medical image analysis have broad prospects[29]. Future research can explore end-to-end LVI prediction models based on deep learning to further improve prediction accuracy. Additionally, combining multi-omics data such as genomics and proteomics to construct more comprehensive prediction models is expected to achieve true precision medicine[30].
In conclusion, this study successfully constructed a rectal cancer LVI prediction model based on multimodal MRI radiomics features. This model demonstrates good prediction performance and clinical application value, providing new technical means for precision diagnosis and treatment of rectal cancer.
| 1. | Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. CA Cancer J Clin. 2023;73:17-48. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12841] [Cited by in RCA: 11237] [Article Influence: 3745.7] [Reference Citation Analysis (4)] |
| 2. | Zhang L, Deng Y, Liu S, Zhang W, Hong Z, Lu Z, Pan Z, Wu X, Peng J. Lymphovascular invasion represents a superior prognostic and predictive pathological factor of the duration of adjuvant chemotherapy for stage III colon cancer patients. BMC Cancer. 2023;23:3. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 14] [Article Influence: 4.7] [Reference Citation Analysis (3)] |
| 3. | Sun Q, Liu T, Liu P, Luo J, Zhang N, Lu K, Ju H, Zhu Y, Wu W, Zhang L, Fan Y, Liu Y, Li D, Zhu Y, Liu L. Perineural and lymphovascular invasion predicts for poor prognosis in locally advanced rectal cancer after neoadjuvant chemoradiotherapy and surgery. J Cancer. 2019;10:2243-2249. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 53] [Cited by in RCA: 47] [Article Influence: 6.7] [Reference Citation Analysis (0)] |
| 4. | Yadav A, Kumar A. Artificial intelligence in rectal cancer: What is the future? Artif Intell Cancer. 2023;4:11-22. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 5. | Liu NJ, Liu MS, Tian W, Zhai YN, Lv WL, Wang T, Guo SL. The value of machine learning based on CT radiomics in the preoperative identification of peripheral nerve invasion in colorectal cancer: a two-center study. Insights Imaging. 2024;15:101. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 13] [Reference Citation Analysis (0)] |
| 6. | Delli Pizzi A, Chiarelli AM, Chiacchiaretta P, d'Annibale M, Croce P, Rosa C, Mastrodicasa D, Trebeschi S, Lambregts DMJ, Caposiena D, Serafini FL, Basilico R, Cocco G, Di Sebastiano P, Cinalli S, Ferretti A, Wise RG, Genovesi D, Beets-Tan RGH, Caulo M. MRI-based clinical-radiomics model predicts tumor response before treatment in locally advanced rectal cancer. Sci Rep. 2021;11:5379. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 78] [Cited by in RCA: 68] [Article Influence: 13.6] [Reference Citation Analysis (0)] |
| 7. | Koh DM, Papanikolaou N, Bick U, Illing R, Kahn CE Jr, Kalpathi-Cramer J, Matos C, Martí-Bonmatí L, Miles A, Mun SK, Napel S, Rockall A, Sala E, Strickland N, Prior F. Artificial intelligence and machine learning in cancer imaging. Commun Med (Lond). 2022;2:133. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 267] [Cited by in RCA: 149] [Article Influence: 37.3] [Reference Citation Analysis (0)] |
| 8. | Zhang Y, Liu J, Wu C, Peng J, Wei Y, Cui S. Preoperative Prediction of Microsatellite Instability in Rectal Cancer Using Five Machine Learning Algorithms Based on Multiparametric MRI Radiomics. Diagnostics (Basel). 2023;13:269. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 12] [Reference Citation Analysis (0)] |
| 9. | Tripathi S, Tabari A, Mansur A, Dabbara H, Bridge CP, Daye D. From Machine Learning to Patient Outcomes: A Comprehensive Review of AI in Pancreatic Cancer. Diagnostics (Basel). 2024;14:174. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 7] [Cited by in RCA: 18] [Article Influence: 9.0] [Reference Citation Analysis (2)] |
| 10. | Fan Y, Chen M, Huang H, Zhou M. Predicting lymphovascular invasion in rectal cancer: evaluating the performance of golden-angle radial sparse parallel MRI for rectal perfusion assessment. Sci Rep. 2023;13:8453. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 4] [Reference Citation Analysis (0)] |
| 11. | Xing X, Li D, Peng J, Shu Z, Zhang Y, Song Q. A combinatorial MRI sequence-based radiomics model for preoperative prediction of microsatellite instability status in rectal cancer. Sci Rep. 2024;14:11760. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 9] [Reference Citation Analysis (0)] |
| 12. | Zhang C, Zhou P, Li R, Li Z, Ouyang A. Prediction of lymphovascular invasion in invasive breast cancer based on clinical-MRI radiomics features. BMC Med Imaging. 2024;24:277. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 13. | Tong P, Sun D, Chen G, Ni J, Li Y. Biparametric magnetic resonance imaging-based radiomics features for prediction of lymphovascular invasion in rectal cancer. BMC Cancer. 2023;23:61. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 23] [Reference Citation Analysis (0)] |
| 14. | Wang Y, Bai G, Huang M, Chen W. Machine learning model based on enhanced CT radiomics for the preoperative prediction of lymphovascular invasion in esophageal squamous cell carcinoma. Front Oncol. 2024;14:1308317. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 7] [Article Influence: 3.5] [Reference Citation Analysis (0)] |
| 15. | Xu X, Zhang HL, Liu QP, Sun SW, Zhang J, Zhu FP, Yang G, Yan X, Zhang YD, Liu XS. Radiomic analysis of contrast-enhanced CT predicts microvascular invasion and outcome in hepatocellular carcinoma. J Hepatol. 2019;70:1133-1144. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 604] [Cited by in RCA: 570] [Article Influence: 81.4] [Reference Citation Analysis (7)] |
| 16. | Wu Y, Wang S, Chen Y, Liao Y, Yin X, Li T, Wang R, Luo X, Xu W, Zhou J, Wang S, Bu J, Zhang X. A Multicenter Study on Preoperative Assessment of Lymphovascular Space Invasion in Early-Stage Cervical Cancer Based on Multimodal MR Radiomics. J Magn Reson Imaging. 2023;58:1638-1648. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 17] [Article Influence: 5.7] [Reference Citation Analysis (0)] |
| 17. | Ma Y, Xu X, Lin Y, Li J, Yuan H. An integrative clinical and CT-based tumoral/peritumoral radiomics nomogram to predict the microsatellite instability in rectal carcinoma. Abdom Radiol (NY). 2024;49:783-790. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 5] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 18. | Sun ZG, Chen SX, Sun BL, Zhang DK, Sun HL, Chen H, Hu YW, Zhang TY, Han ZH, Wu WX, Hou ZY, Yao L, Jie JZ. Important role of lymphovascular and perineural invasion in prognosis of colorectal cancer patients with N1c disease. World J Gastroenterol. 2025;31:102210. [PubMed] [DOI] [Full Text] |
| 19. | Yang Y, Wei H, Fu F, Wei W, Wu Y, Bai Y, Li Q, Wang M. Preoperative prediction of lymphovascular invasion of colorectal cancer by radiomics based on 18F-FDG PET-CT and clinical factors. Front Radiol. 2023;3:1212382. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 13] [Reference Citation Analysis (0)] |
| 20. | Ge YX, Xu WB, Wang Z, Zhang JQ, Zhou XY, Duan SF, Hu SD, Fei BJ. Prognostic value of CT radiomics in evaluating lymphovascular invasion in rectal cancer: Diagnostic performance based on different volumes of interest. J Xray Sci Technol. 2021;29:663-674. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 6] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 21. | Wong C, Liu T, Zhang C, Li M, Zhang H, Wang Q, Fu Y. Preoperative detection of lymphovascular invasion in rectal cancer using intravoxel incoherent motion imaging based on radiomics. Med Phys. 2024;51:179-191. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 13] [Reference Citation Analysis (0)] |
| 22. | Li H, Chai L, Pu H, Yin LL, Li M, Zhang X, Liu YS, Pang MH, Lu T. T2WI-based MRI radiomics for the prediction of preoperative extranodal extension and prognosis in resectable rectal cancer. Insights Imaging. 2024;15:57. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 13] [Reference Citation Analysis (0)] |
| 23. | Liu D, Huang J, Zhang Y, Shen H, Wang X, Huang Z, Chen X, Qiao Z, Hu C. Multimodal MRI-based radiomics models for the preoperative prediction of lymphovascular space invasion of endometrial carcinoma. BMC Med Imaging. 2024;24:252. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 7] [Reference Citation Analysis (0)] |
| 24. | Li Z, Zhang J, Zhong Q, Feng Z, Shi Y, Xu L, Zhang R, Yu F, Lv B, Yang T, Huang C, Cui F, Chen F. Development and external validation of a multiparametric MRI-based radiomics model for preoperative prediction of microsatellite instability status in rectal cancer: a retrospective multicenter study. Eur Radiol. 2023;33:1835-1843. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 31] [Cited by in RCA: 28] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 25. | Yardimci AH, Kocak B, Sel I, Bulut H, Bektas CT, Cin M, Dursun N, Bektas H, Mermut O, Yardimci VH, Kilickesmez O. Radiomics of locally advanced rectal cancer: machine learning-based prediction of response to neoadjuvant chemoradiotherapy using pre-treatment sagittal T2-weighted MRI. Jpn J Radiol. 2023;41:71-82. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 25] [Reference Citation Analysis (0)] |
| 26. | Liang M, Cai Z, Zhang H, Huang C, Meng Y, Zhao L, Li D, Ma X, Zhao X. Machine Learning-based Analysis of Rectal Cancer MRI Radiomics for Prediction of Metachronous Liver Metastasis. Acad Radiol. 2019;26:1495-1504. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 63] [Article Influence: 9.0] [Reference Citation Analysis (5)] |
| 27. | Li ZF, Kang LQ, Liu FH, Zhao M, Guo SY, Lu S, Quan S. Radiomics based on preoperative rectal cancer MRI to predict the metachronous liver metastasis. Abdom Radiol (NY). 2023;48:833-843. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 10] [Reference Citation Analysis (1)] |
| 28. | Jiang Y, Zeng Y, Zuo Z, Yang X, Liu H, Zhou Y, Fan X. Leveraging multimodal MRI-based radiomics analysis with diverse machine learning models to evaluate lymphovascular invasion in clinically node-negative breast cancer. Heliyon. 2024;10:e23916. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 14] [Reference Citation Analysis (0)] |
| 29. | Bibault JE, Giraud P, Housset M, Durdux C, Taieb J, Berger A, Coriat R, Chaussade S, Dousset B, Nordlinger B, Burgun A. Deep Learning and Radiomics predict complete response after neo-adjuvant chemoradiation for locally advanced rectal cancer. Sci Rep. 2018;8:12611. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 80] [Cited by in RCA: 135] [Article Influence: 16.9] [Reference Citation Analysis (5)] |
| 30. | Zhang YP, Zhang XY, Cheng YT, Li B, Teng XZ, Zhang J, Lam S, Zhou T, Ma ZR, Sheng JB, Tam VCW, Lee SWY, Ge H, Cai J. Artificial intelligence-driven radiomics study in cancer: the role of feature engineering and modeling. Mil Med Res. 2023;10:22. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 96] [Reference Citation Analysis (0)] |
Open Access: This article is an open-access article that was selected by an in-house editor and fully peer-reviewed by external reviewers. It is distributed in accordance with the Creative Commons Attribution NonCommercial (CC BY-NC 4.0) license, which permits others to distribute, remix, adapt, build upon this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See: https://creativecommons.org/Licenses/by-nc/4.0/