Published online Mar 15, 2026. doi: 10.4239/wjd.v17.i3.115097
Revised: November 18, 2025
Accepted: February 4, 2026
Published online: March 15, 2026
Processing time: 154 Days and 20 Hours
Machine learning (ML) and deep learning (DL) algorithms have been utilised to predict the risk of diabetic kidney disease (DKD) in individuals with type 2 dia
To evaluates the performance of ML-based and DL-based models in predicting DKD risk among T2DM patients.
A systematic search was conducted across five databases. The risk of bias was assessed using the prediction model risk of bias assessment tool. After data extraction, summary point estimates of the area under the receiver operating characteristic curve (AUC) were aggregated. Heterogeneity was evaluated with the I2 statistic and Cochrane Q test, and subgroup analyses were performed to identify potential sources of heterogeneity.
Twelve eligible studies were included. The pooled AUC for the top-performing artificial intelligence models was 0.858 [95% confidence interval (CI): 0.779-0.912], with a prediction interval of 0.480-0.975. Significant heterogeneity was detected (I2 = 99.7%). Studies employing cross-validation methods demonstrated significantly higher diagnostic accuracy (pooled AUC = 0.88; 95%CI: 0.79-0.94) compared to those using simple holdout validation (pooled AUC = 0.77; 95%CI: 0.59-0.89, P = 0.0231). Predictive factors most frequently used for DKD prediction included age, body mass index, estimated glomerular filtration rate, serum creatinine, urinary albumin, glycated hemoglobin, systolic blood pressure, low-density lipoprotein cholesterol, high-density lipoprotein cholesterol, and triglycerides.
ML and DL algorithms exhibit strong performance in predicting DKD in patients with T2DM. However, future research should focus on standardizing model development and validation processes.
Core Tip: Machine learning and deep learning algorithms show great performance in predicting diabetic kidney disease (DKD) among type 2 diabetes mellitus patients. Predictors, such as age, body mass index, estimated glomerular filtration rate, serum creatinine, urinary albumin, glycated hemoglobin, systolic blood pressure, low-density lipoprotein cholesterol, high-density lipoprotein cholesterol, and triglycerides, play significant roles in DKD prediction. Models employing cross-validation methods exhibit superior predictive capability for DKD compared to those using holdout validation approaches.
- Citation: Chen Q, Peng HW, Fu CX, Meng KK, Zhang JB. Machine learning and deep learning in predicting the risk of diabetic kidney disease: A systematic review and meta-analysis. World J Diabetes 2026; 17(3): 115097
- URL: https://www.wjgnet.com/1948-9358/full/v17/i3/115097.htm
- DOI: https://dx.doi.org/10.4239/wjd.v17.i3.115097
Diabetic kidney disease (DKD), a major microvascular complication of type 2 diabetes mellitus (T2DM), is clinically characterized by proteinuria and a rapid decline in estimated glomerular filtration rate (eGFR)[1,2]. Currently, approximately 589 million adults worldwide live with diabetes, a figure projected to increase to 853 million by 2050[3]. Indi
Recent advancements in machine learning (ML) and deep learning (DL) techniques have shown great promise in early disease detection and clinical management[7-9]. These computational methods can effectively integrate and analyze multi-modal data sources, including clinical text, medical imaging, behavioural patterns, and physiological parameters, to uncover complex pathophysiological relationships between DKD and its multifactorial determinants[10-12]. Although previous meta-analyses have evaluated ML applications in DKD research, they have not sufficiently distinguished between predictive and diagnostic modelling paradigms[13-17]. Such methodological confusion may obscure the prac
This study adhered to the Preferred Reporting Items for Systematic Reviews and Meta-Analyses guidelines and was conducted in accordance with the Cochrane Collaboration[18]. The protocol has been registered with PROSPERO (CRD420251080292). Review questions and data extraction were structured using the critical appraisal and data extraction for systematic reviews of prediction modelling studies checklist[19]. All aspects of the protocol, including literature searching, screening, data extraction, quality assessment, and risk of bias evaluation, were performed independently by two reviewers.
A systematic search was conducted across five databases (PubMed, Web of Science, EMBASE, MEDLINE, and the Cochrane Library) up to June 25, 2025. Literature identification and screening followed the PICOTS (population, intervention, comparator, outcome, timing, setting) framework, with inclusion criteria as follows: (1) Population: Patients with confirmed T2DM and eGFR > 60 mL/minute/1.73 m2; (2) Index model: Any models utilising ML or DL algorithms; (3) Comparator model: Not applicable; (4) Outcome: Development of DKD among patients with T2DM; (5) Timing: From T2DM diagnosis onward, including all reported prediction timeframes; and (6) Setting: Both hospitalised and ambulatory care populations. The full search syntax is detailed in Supplementary Table 1.
The risk of bias was assessed using the prediction model risk of bias assessment tool[20], which evaluates four key bias domains: Participants, predictors, outcome, and analysis, with 20 signalling questions. For each domain, the risk was classified as “low”, “high”, or “unclear” based on responses to the signalling questions. The overall risk of bias assessment was derived from evaluations across all domains. Models lacking external validation were automatically considered high risk, regardless of domain-specific assessments.
Data extraction followed the critical appraisal and data extraction for systematic reviews of prediction modelling studies and Transparent Reporting of a multivariable prediction model of Individual Prognosis Or Diagnosis + artificial intelligence checklists, utilising standardised forms[21]. Comprehensive study data extracted included data sources, sample characteristics (size and demographics), outcome definitions, predictors, missing data handling, statistical methods, validation approaches, and performance metrics.
Statistical analyses were performed using R software (version 4.4.2). Continuous variables with normal distribution are reported as mean ± SD, while non-normally distributed data are presented as median (interquartile range). For the meta-analysis of model discrimination performance, area under the receiver operating characteristic (ROC) curve [area under the receiver operating characteristic curve (AUC)], values were pooled using a logit transformation approach. Each AUC value was logit-transformed, and the standard error (SE) was calculated using the delta method. Pooled logit-transformed AUC values and SEs were then used in the meta-analysis. Heterogeneity among studies was assessed using I2 statistics and the Cochrane Q test. If significant heterogeneity was found (I2 > 50% or Q test P < 0.10), a random-effects model was employed for pooling. Subgroup analyses were conducted to explore potential sources of heterogeneity. Sensitivity analyses using the leave-one-out method were performed to evaluate the robustness of the findings. A two-sided P value < 0.05 was considered statistically significant.
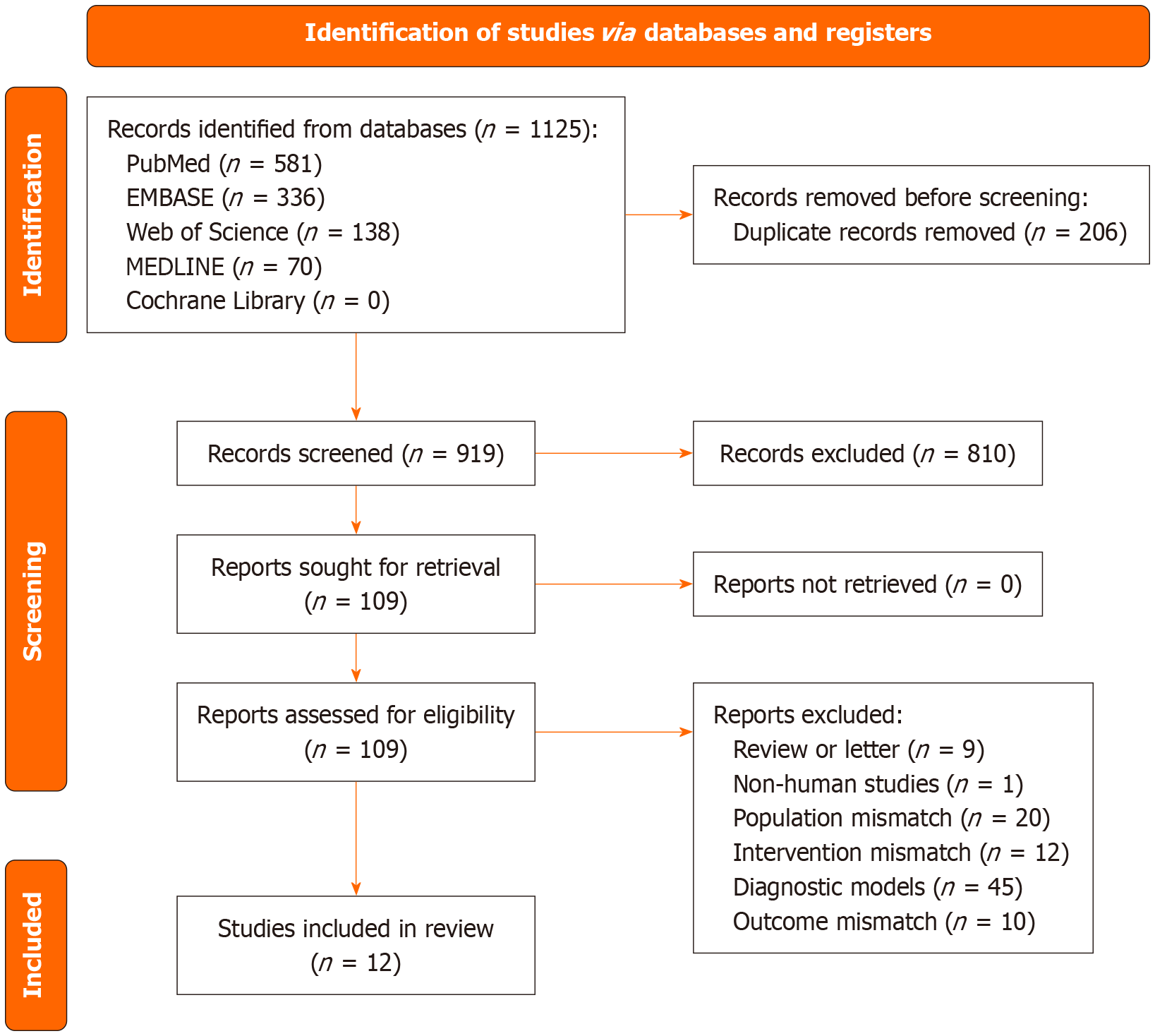
After systematically searching five databases, 1125 studies were initially identified. Duplicate records were removed using EndNote X9 software, resulting in 206 exclusions. Following title and abstract screening, 810 records were excluded. A full-text review of the remaining studies led to the exclusion of an additional 97 records due to the following reasons: Reviews, non-human studies, incompatible demographic characteristics, absence of ML and DL, diagnostic model construction, and mismatched outcomes. Ultimately, 12 studies met all predefined inclusion criteria and were included in the final analysis[22-33]. The study selection process, following Preferred Reporting Items for Systematic Reviews and Meta-Analyses guidelines, is illustrated in Figure 1.
The characteristics of the 12 studies are summarised in Table 1[22-33]. Nine studies obtained data from electronic health records (EHRs)/electronic medical records (EMRs)[22,23,25-27,29-32], one study utilised data from the Action to Control Cardiovascular Risk in Diabetes study[24], another from the Singapore Epidemiology of Eye Diseases study[28], and one study used data from the DApagliflozin Real-World evIdeNce-Renal study[33]. Five studies were single-centre[22,26,29,31,32], while seven were multicentre[23-25,27,28,30,33]. Most studies enrolled T2DM participants who were free from DKD at baseline. For DKD diagnostic criteria, all 12 studies primarily relied on urinary albumin measurements, eGFR, and UACR[22-33].
| Ref. | Country | Data source | Recruitment period | Single-center/multi-center | Sample size | Inclusion criteria | Definition of positive outcome |
| Cho et al[22], 2008 | South Korea | EHRs | 1996-2005 | Single-center | 292 | Confirmed T2DM; age ≥ 18 years; patients underwent up to 20 clinical tests | 20 μg/minute to 200 μg/minute in urinary albumin; no microalbumin or renal failure at diabetes diagnosis; prior evidence of diabetic retinopathy |
| Dagliati et al[23], 2018 | Italy | EHRs | 2012-2016 | Multi-center | 943 | Confirmed T2DM; no pre-existing complications; sufficient follow-up records | eGFR < 60 mL/minute/1.73 m2; UACR=30 mg/gram to 299 mg/gram (≥ 2 morning samples) |
| Rodriguez-Romero et al[24], 2019 | United States | ACCORD dataset | 2001.01-2001.06; 2003.02-2005.10 | Multi-center | 10251 | Confirmed T2DM by ADA criteria; age ≥ 40 with CVD history; age ≥ 55 with high CVD risk; HbA1c 7.5%-9% (multiple drugs); 7.5%-11% (fewer drugs) | Baseline SCr doubling; eGFR < 60 mL/minute/1.73 m2; UACR ≥ 30 mg/gram or 3.4 mg/mmol; UACR ≥ 300 mg/gram or 33.9 mg/mmol; renal failure |
| Allen et al[25], 2022 | United States | EHRs | 2007-2020 | Multi-center | 111046 | Confirmed T2DM by ICD-9/10 codes; age ≥ 18 years; ≥5 years follow-up records; required baseline tests (1-year pre-T2DM); albuminuria/reduced eGFR allowed at baseline; no pre-existing CKD or renal transplant | DKD confirmed by ICD-9/10 codes |
| Dong et al[26], 2022 | China | EMRs | 2008.10-2019.12 | Single-center | 2809 | Confirmed T2DM by ADA criteria; age ≥ 18 years; 3 years follow-up records | UACR > 30 mg/gram; protein excretion rate > 150 mg/24 hours; urine dipstick test ≥ 1+; eGFR < 60 mL/minute/1.73 m2 |
| Nicolucci et al[27], 2022 | Italy | EMRs | NA | Multi-center | 147664 | Confirmed T2DM by ICD-9 CM codes | Confirmed DKD by ICD-9 CM codes; no pre-existing complication at baseline |
| Sabanayagam et al[28], 2023 | Singapore | SEED study | 2004-2011 | Multi-center | 1365 | Random glucose ≥ 11.1 mmol/L; HbA1c ≥ 6.5% (48 mmol/mol); self-reported antidiabetic medication use; diagnosed with diabetes by a physician | eGFR < 60 mL/minute/1.73 m2; ≥ 25% eGFR decline during follow-up |
| Hosseini Sarkhosh et al[29], 2023 | Iran | EHRs | 2012-2021 | Single-center | 1907 | Confirmed T2DM by ADA criteria; 5 years follow-up records; no pre-existing DKD at baseline | UACR ≥ 30 mg/gram; eGFR ≤ 60 mL/minute/1.73 m2 |
| Yun et al[30], 2024 | China | EHRs | NA | Multi-center | 6040 | Confirmed T2DM; age ≥ 18 years; ≥ 7 years follow-up records; ≥ 2 hospital visits per year; no pre-existing DKD at baseline or within 2 years | UACR ≥ 30 mg/gram (≥ 3 months); eGFR < 60 mL/minute/1.73 m2; exclude other kidney diseases |
| Lin et al[31], 2025 | China | EHRs | 2011-2023 | Single-center | 3291 | Confirmed T2DM by ICD-9/10 codes; age ≥ 18 years; at least one hospital visit following the end of the baseline time window | eGFR < 60 mL/minute/1.73 m2; UACR ≥ 30 mg/gram over 3 months; DKD confirmed by ICD-9/10 codes; protein in urine dipstick test ≥ 1+ |
| Zou et al[32], 2025 | China | EHRs | 2014.01-2022.12 | Single-center | 9572 | Confirmed T2DM; no pre-existing DKD; ≥ 3 years follow-up records; exclude glomerular/systemic diseases | UACR ≥ 30 mg/gram in 2/3 tests within 3 months to 6 months; eGFR < 60 mL/minute/1.73 m2 for more than 3 months; renal biopsy was consistent with DKD pathological changes |
| Dei Cas et al[33], 2025 | Italy | DARWIN-Renal | 2015.01-2021.09 | Multi-center | 22379 | Confirmed T2DM; age ≥ 18 years; ≥ 1 year follow-up records | 5 renal disease severity thresholds established by the KDOQI scale based on the eGFR |
The details of prediction model development and validation are presented in Table 2. Of the 12 included studies[22-33], 10 developed models using ML approaches[22-29,31,32], while 2 employed DL methods[30,33]. For missing data imputation, three studies used multiple imputation by chained equations[31-33], three used the random forest (RF) imputation algorithm (missForest)[23,25,26], two applied mean/mode imputation[28,29], and one study filled missing values with an extreme value (-999)[27]. To address class imbalance, four studies used oversampling[23,24,27,32], including three employing synthetic minority over-sampling technique[24,27,32]. Most studies randomly allocated datasets into development and validation sets with varying proportions. Regarding validation methods, most studies utilised holdout or K-fold validation[24,26-28,32,33], except for Cho et al[22] and Dagliati et al[23], who applied leave-one-out cross-validation (LOOCV), and Hosseini Sarkhosh et al[29], who used recursive feature elimination with cross-validation. Four studies performed external validation to assess model generalizability[25,27,29,33].
| Ref. | Number | Algorithm type | Missing data imputation | Oversample | Train-test split | Validation type | Internal validation method | Best model | Prediction horizon, year | AUC (95%CI) |
| Cho et al[22], 2008 | 292 | ML | NA | NA | NA | Internal | LOOCV | SVM | 1 | 0.969 (0.941-0.997) |
| Dagliati et al[23], 2018 | 943 | ML | missForest | Oversampling | NA | Internal | LOOCV | LR | 3 | 0.808 (0.772-0.845) |
| Rodriguez-Romero et al[24], 2019 | 10251 | ML | NA | SMOTE | 66:34 | Internal | CV-10 | RF | 1-1.9 | 0.730 (0.715-0.745) |
| Allen et al[25], 2022 | 111046 | ML | missForest | NA | 7:1 | External | Holdout | XGBoost | 5 | 0.750 (0.734-0.766) |
| Dong et al[26], 2022 | 861 | ML | missForest | NA | 8:2 | Internal | CV-5 | LightGBM | 3 | 0.815 (0.747-0.882) |
| Nicolucci et al[27], 2022 | 147664 | ML | Extra-values imputation | SMOTE | Yes | External | CV-10 | XGBoost | 2 | 0.970 (0.968-0.972) |
| Sabanayagam et al[28], 2023 | 1365 | ML | Mean/mode | NA | 8:2 | Internal | CV-5 | Elastic Net | 6 | 0.851 (0.847-0.856) |
| Hosseini Sarkhosh et al[29], 2023 | 3444 | ML | Mean/mode | NA | 8:2 | External | RFECV | RF | 5 | 0.790 (0.770-0.820) |
| Yun et al[30], 2024 | 6040 | DL | NA | NA | 7:3 | Internal | Holdout | LSTM | 5 | 0.830 (0.807-0.853) |
| Lin et al[31], 2025 | 3291 | ML | MICE | NA | 7:3 | Internal | Holdout | SuperLearner | 2.53 | 0.714 (0.673-0.755) |
| Zou et al[32], 2025 | 12190 | ML | MICE | SMOTE | 8:2 | Internal | CV-5 | LightGBM | 3 | 0.918 (0.906-0.930) |
| Dei Cas et al[33], 2025 | 32379 | DL | MICE | NA | 8:2 | External | CV-5 | RNN | 2 | 0.887 (0.869-0.904) |
Risk of bias assessment revealed that 12 studies[22-33] were rated as having a high risk of bias based on prediction model risk of bias assessment tool (Supplementary Figure 1). Bias in participant selection was identified in studies that extracted data from single-center EHRs or EMRs. In the analytical domain, one study had fewer than 10 events per variable, three failed to handle missing data adequately[22,24,30], and eight studies did not address potential overfitting[22,25,26,28-31,33]. Additionally, five studies did not perform feature selection[24,25,27,30,31].
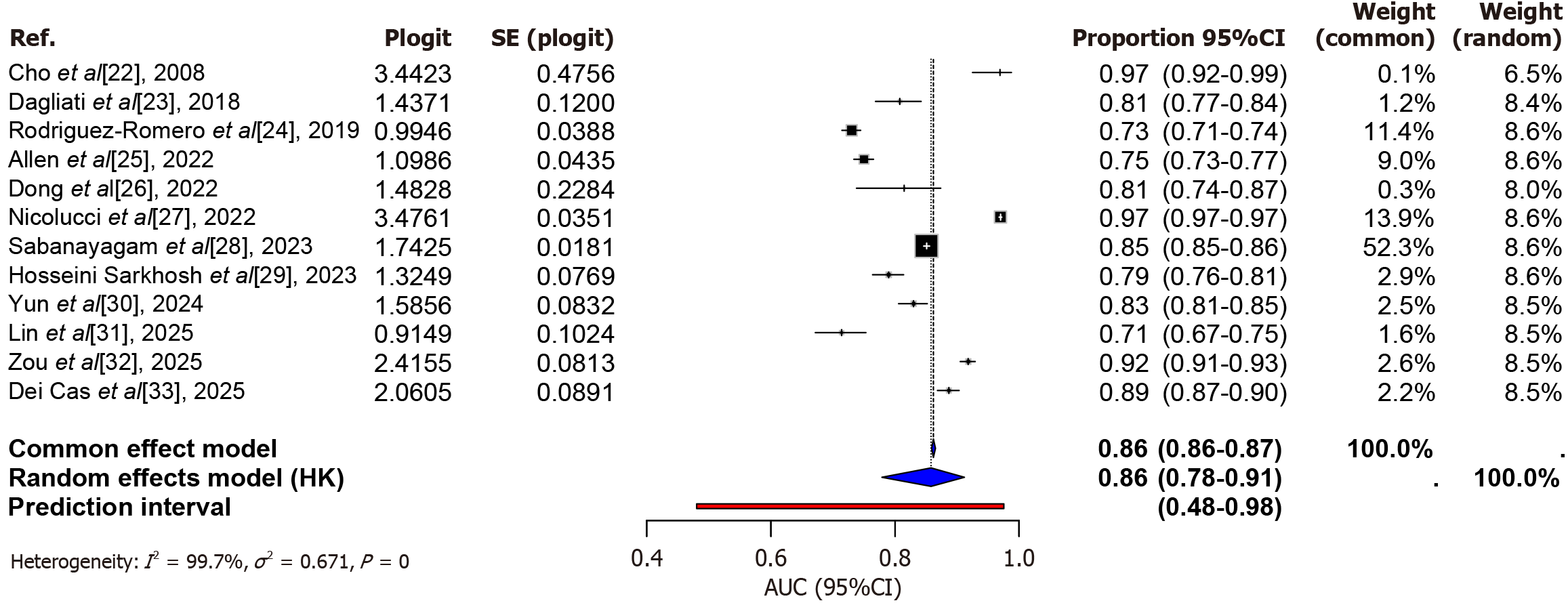
The AUC values across the 12 included studies ranged from 0.714 to 0.970[22-33] (Figure 2). Due to substantial heterogeneity, a random-effects model was employed for synthesising AUC values. After applying the Hartung-Knapp adjustment, the pooled AUC was estimated at 0.858 (95%CI: 0.779-0.912), with a prediction interval spanning (0.480-0.975).
This meta-analysis revealed extremely high heterogeneity in AUC values among the studies (I2 = 99.7%, P = 0, Figure 2). The Baujat plot indicated that Nicolucci et al[27], contributed most to the overall heterogeneity and was identified as a primary source of variation (Supplementary Figure 2A). The Galbraith plot further highlighted that the effect sizes of Nicolucci et al[27] and Rodriguez-Romero et al[24], significantly deviated from the regression line (Supplementary Figure 2B). Although excluding Nicolucci et al[27], slightly reduced the pooled estimate, the change was minimal, and heterogeneity remained largely unchanged (I2 = 98.4%, P < 0.0001). Consequently, Nicolucci et al[27], was retained in the analysis, and subgroup analyses were conducted to explore other potential sources of heterogeneity.
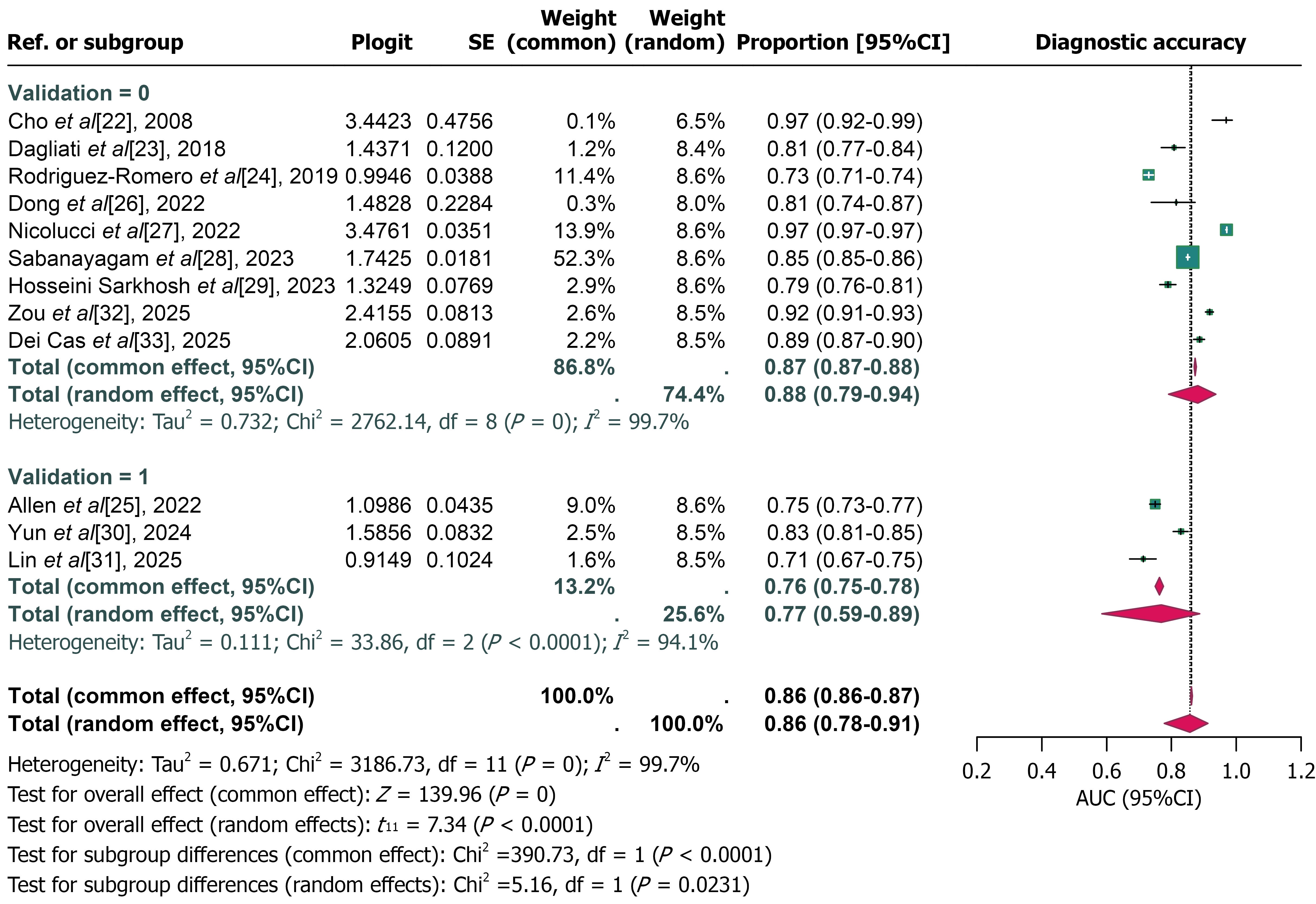
Subgroup analyses were performed based on factors such as region, study type, algorithm type, study center design, use of external validation, and prediction horizon. All subgroup analyses yielded non-significant P value (all P > 0.05; Table 3), suggesting that none of these factors significantly explained the observed heterogeneity. However, when the studies were categorised into holdout and non-holdout groups based on internal validation methods, statistically significant heterogeneity was observed (P = 0.0231, Figure 3). Studies employing cross-validation methods, including k-fold cross-validation, LOOCV, and repeated cross-validation, exhibited markedly superior diagnostic accuracy (pooled AUC = 0.88; 95%CI: 0.79-0.94; Figure 3) compared to those utilising simple holdout validation (pooled AUC = 0.77; 95%CI: 0.59-0.89; Figure 3).
| Subgroup variable | Level | Studies | AUC (95%CI) | I2 | t11 | P value | χ2 | P value |
| Region | Overall | 7.34 | < 0.0001 | 0.01 | 0.9422 | |||
| Asian | 7 | 0.86 (0.74-0.92) | 96.7% | |||||
| Western | 5 | 0.86 (0.78-0.91) | 99.7% | |||||
| Study type | Overall | 12 | 0.86 (0.78-0.91) | 85.8% | 7.34 | < 0.0001 | 0.47 | 0.4951 |
| Prospective study | 3 | 0.90 (0.40-0.99) | 99.7% | |||||
| Retrospective study | 9 | 0.85 (0.74-0.91) | 99.7% | |||||
| Algorithm type | Overall | 7.34 | < 0.0001 | 0.00 | 0.9563 | |||
| ML | 10 | 0.86 (0.76-0.92) | 99.7% | |||||
| DL | 2 | 0.86 (0.23-0.99) | 99.7% | |||||
| Center | Overall | 12 | 0.86 (0.78-0.91) | 99.7% | 7.34 | < 0.0001 | 0.02 | 0.8770 |
| Single-center | 5 | 0.86 (0.66-0.96) | 97.7% | |||||
| Multicenter | 7 | 0.85 (0.78-0.91) | 99.8% | |||||
| Validation | Overall | 12 | 0.86 (0.78-0.91) | 99.7% | 7.34 | < 0.0001 | 0.26 | 0.6107 |
| Internal validation | 8 | 0.84 (0.74-0.91) | 98.5% | |||||
| External validation | 4 | 0.86 (0.78-0.91) | 99.9% | |||||
| Prediction horizon | Overall | 12 | 0.86 (0.78-0.91) | 99.7% | 7.34 | < 0.0001 | 0.92 | 0.3362 |
| < 3 years | 5 | 0.90 (0.64-0.98) | 99.8% | |||||
| ≥ 3 years | 7 | 0.83 (0.77-0.88) | 98.0% |
A leave-one-out sensitivity analysis was conducted (Supplementary Figure 3). Point estimates consistently ranged between 0.82 and 0.88 after iteratively excluding individual studies, indicating minimal variation. The confidence intervals remained similar across all iterations. Egger’s and Begg’s tests were used to assess publication bias, and no significant bias was detected (Supplementary Figure 4). After imputing one study using the Trim-and-Fill method, the pooled AUC was 0.867 (95%CI: 0.796-0.917). The effect size showed only marginal change after imputation, and the width of the confidence interval remained largely unchanged.
A systematic analysis was performed on the characteristic variables integrated into the optimal prediction models developed in 12 studies on DKD[22-33]. These variables were categorised into three primary groups based on their attributes: Demographic characteristics, medical history, and laboratory indicators. Demographic characteristics mainly included age, gender, body mass index (BMI), and diabetes duration (Table 4). Age was incorporated in eight studies, BMI in gender, gender in four, and diabetes duration in three. Medical history variables focused on key risk factor exposures, such as hypertension, smoking, and alcohol consumption. Hypertension history appeared in four studies, smoking in three, and alcohol consumption in two. Laboratory indicators formed the core of the prediction models, covering various dimensions including blood pressure parameters, renal function indicators, glycemic control levels, lipid profiles, complete blood count, and liver function. Glycated hemoglobin (HbA1c) was included in eight studies, low-density lipoprotein (LDL) in seven, eGFR in gender, and systolic blood pressure (SBP) in gender. Additionally, Sabanayagam et al[28], and Dei Cas et al[33], also included antidiabetic medication usage.
| Ref. | Number | Demographic | Past history | Laboratory |
| Cho et al[22], 2008 | 39 | BMI (mean, minimum) | NA | SBP (mean, EST, and initial), DBP (slope and maximum), WBC (initial), WBC (latest), hemoglobin, platelet (slope), platelet (var), platelet (K), platelet (EST), cholesterol (K and EST), AST (K and initial), ALT (initial, minimum, and K), ALP (minimum and latest), creatinine (mean and var), uric acid (var), Na (EST), K (slope and K), triglyceride (EST), HDL-C (var and initial), LDL-C (EST and initial), microalbumin (mean, var, maximum, and minimum) |
| Dagliati et al[23], 2018 | 4 | BMI | Smoking, hypertension | HbA1c |
| Rodriguez-Romero et al[24], 2019 | 18 | Age | NA | UAlb at baseline, eGFR at baseline, UCr at baseline, eGFR from baseline to year 1, trig at baseline, FPG at baseline, CPK at baseline, age, eGFR at month 4, K at baseline, LDL at baseline, Chol at baseline, eGFR at month 12, FPG at month 4, FPG from baseline to year 1, trig from baseline to year 1, UACR at baseline, FPG at month 8 |
| Allen et al[25], 2022 | 15 | Age, gender, BMI | Acute kidney injury, chronic heart failure, smoking, drinking | SBP, DBP, blood urea nitrogen, creatinine, eGFR, cholesterol (HDL and LDL), white cell count |
| Dong et al[26], 2022 | 8 | Age, BMI | NA | Hcy, HbA1c, ALB, eGFR, bicarbonate, LDL |
| Nicolucci et al[27], 2022 | 46 | Gender, age, height, weight, BMI, waist circumference, diabetes duration | NA | SBP, DBP, ankle/brachial index DX, ankle/brachial index SX, fasting blood glucose, blood glucose after breakfast, blood glucose before dinner, blood glucose before lunch, blood glucose at 11:00 pm, blood glucose after lunch, blood glucose after dinner, pre-prandial blood glucose, post-prandial blood glucose, HbA1c, albuminuria, serum creatinine, creatinine clearance, total cholesterol, LDL cholesterol, HDL cholesterol, triglycerides, fibrinogen, GGT, ALT, AST, alkaline phosphatase, amylase, CPK, hemoglobin, platelets, BUN, uric acid, glycosuria, urinary amylase, urinary ketones, urinary potassium, urinary sodium, urine creatinine, urine culture |
| Sabanayagam et al[28], 2023 | 15 | Ethnicity (Malay/Chinese) | Diabetic retinopathy, hypertension | Acetate, diabetic retinopathy, SBP, DHA, GFR-EPI, HbA1c, IDL-CE%, M-HDL-PL%, M-VLDL-PL%, S-HDL-FC%, XL-HDL-CE%, anti-DM meds |
| Hosseini Sarkhosh et al[29], 2023 | 6 | Diabetes duration | Hypertension, CVD | HbA1c, eGFR, ACR |
| Yun et al[30], 2024 | 10 | Age | NA | PP, SBP, variabilities of PP and SBP, Scr, HDL-C, HbA1c, TG, variabilities of HbA1c |
| Lin et al[31], 2025 | 46 | Gender, age | Smoking, drinking | WBC count, neutrophil count, lymph count, mono count, eosinophil count, basophil count, red blood cell count, hemoglobin, hematocrit, mean corpuscular volume, mean corpuscular hemoglobin, mean corpuscular hemoglobin concentration, red cell distribution width-coefficient of variation, platelet, mean platelet volume, platelet distribution width, lymph, mono, eosinophil, basophil, ALT, AST, AST/ALT (AST ALT), total bilirubin, creatinine, Ca, total cholesterol, triglyceride, HDL, LDL, prealbumin, HbA1c, D-Dimer, pH, glucose, billing, ketone, obstetric, nitrogen, urology |
| Zou et al[32], 2025 | 5 | Age | NA | UACR, Cystatin C, eGFR, Neutrophil |
| Dei Cas et al[33], 2025 | 34 | Gender, age, weight, weight-past, BMI, diabetes duration | CKD | SBP, DBP, SBP in the past, DBP in the past, HbA1c, HbA1c in the past, eGFR, eGFR in the past, AER, ACR > 30, AER in the past, and anti-DM Meds |
DKD accounts for approximately 30% to 50% of global end-stage renal disease cases[34]. Therefore, early prediction of DKD is critical for preserving renal function in patients with T2DM. Conventional screening primarily relies on cross-sectional measurements of single or limited biomarkers, such as a spot UACR > 30 mg/gram or an eGFR < 60 mL/minute/1.73 m2[35]. This approach essentially identifies established kidney injury[36]. In contrast, ML and DL models can integrate multi-dimensional longitudinal data. This enables the earlier identification of high-risk individuals before irreversible structural kidney damage occurs. Therefore, multiple research teams develop various DKD prediction models using diverse statistical methods[7]. To evaluate the clinical applicability and reliability of these AI models, 12 eligible studies were included, and a meta-analysis of the highest-performing prediction models was conducted[22-33]. The pooled AUC from internal validation sets was 0.858 (95%CI: 0.779-0.912), indicating that the top-performing models from the included studies demonstrate strong discriminatory ability in predicting DKD, consistent with findings from other meta-analyses[12-17].
The pooled AUC in our analysis exhibited considerable heterogeneity, which may be attributed to variations in internal validation methods used across the included studies. Four distinct validation approaches were employed: K-fold cross-validation, LOOCV, repeated cross-validation, and hold-out validation. Our analysis revealed that studies utilising hold-out validation demonstrated a lower pooled AUC compared to those employing cross-validation. Despite a statistically significant difference in pooled effect estimates between the hold-out and cross-validation groups, substantial heterogeneity remained within both subgroups, suggesting that the validation method accounts for only part of the variability. Leave-one-out sensitivity analyses confirmed that no single study significantly influenced the findings. Furthermore, other subgroup analyses, examining factors such as region, study type, algorithm type, center design, external validation, and prediction horizon revealed no significant differences. The persistent high heterogeneity implies that the outcomes are influenced by complex, multifactorial interactions or unmeasured covariates, such as subtle population differences, variations in model architecture or hyperparameters, or differing clinical outcome definitions. Further investigation with a larger sample of studies, where possible, and individual participant data meta-analysis, is needed to better identify and adjust for these factors.
DKD is a complex condition whose development and progression are driven by a combination of hemodynamic abnormalities and metabolic disturbances. The pathological mechanisms of DKD include hyperglycemia, accumulation of advanced glycation end products, proteinuria, and lipid overload[37,38]. Our analysis identified the core predictive factors for DKD across the 12 included studies as gender, BMI, eGFR, serum creatinine, urinary albumin, HbA1c, SBP, LDL-cholesterol (LDL-C), high-density lipoprotein cholesterol (HDL-C), and triglycerides[22-33]. These factors are critical for the early identification and risk stratification of DKD. Established predictors such as BMI, LDL-C, fasting blood glucose, SBP, and HbA1c have been confirmed as key risk factors for DKD in previous non-ML studies[39,40]. Unlike traditional statistical methods, which rely on clinically pre-defined variable relationships and linear assumptions to test associations with DKD, ML and DL are data-driven. They autonomously learn and capture complex patterns in the data, including nonlinear relationships and interactions among variables. For example, Wu et al[41], demonstrated that in patients with DKD, T2DM diagnosed before age 40 was associated with poorer glycemic control and more severe proteinuria. Other research shows that for every 5 years to 10 years increase in age, the risk of DKD rises by 38%[42]. In the SHapley Additive exPlanation plot presented by Dong et al[26], both age < 40 years and > 60 years were associated with positive SHapley Additive exPlanation values, suggesting that both younger and older age are independent extremes linked to heightened DKD risk. However, this does not imply that complex algorithms are superior to logistic regression. A meta-analysis found that in the field of chronic kidney disease research, the alleged superiority of non-logistic regression models over traditional logistic regression lacks substantial empirical support. This suggests that the value of ML and DL lie in their abilities to capture complex feature relationships, rather than in the complexities of the algorithms themselves. Beyond age, several modifiable factors significantly influence DKD progression. Intensive glycemic control has been shown to reduce DKD risk[43,44]. Hypertension exacerbates renal dysfunction and increases cardiovascular risk, emphasising the importance of blood pressure control in DKD management[45]. Dyslipidemia, particularly elevated LDL-C or reduced HDL-C levels, plays a critical role in DKD development and progression[46,47], Additionally, smoking has been established as an independent risk factor for both the onset and progression of DKD[48,49].
A key strength of this study is that all included studies were based on longitudinal cohorts for constructing prediction models for DKD. In contrast to diagnostic models derived from cross-sectional data, this design more accurately captures the dynamic progression of the disease and the cumulative effects of long-term risk factors, significantly enhancing the reliability and clinical applicability of the predictions.
However, several methodological limitations should be acknowledged: First, the majority of the included studies relied on EHRs or EMRs for retrospective analysis, which inherently introduces a high risk of selection bias and information bias; second, there was substantial heterogeneity in study designs, with most being single-center retrospective cohorts, further increasing the potential for bias; third, many studies had shortcomings in data preprocessing, including inconsistent methods for handling missing values and inadequate attention to class imbalance issues, which may impact the predictive performance and robustness of the models; fourth, the lack of a standardised model evaluation framework led to the use of diverse evaluation metrics across studies, and some studies did not report complete metrics. This limited the synthesis and comparability of results and increased the complexity of interpretation; fifth, our meta-analysis aims to assess the performances of models to predict the risk of early-stage DKD, its findings are primarily applicable to T2DM populations with normal or mildly impaired baseline renal function (eGFR ≥ 60 mL/minute/1.73 m2). Consequently, our findings are not generalizable to patients with a baseline eGFR < 60 mL/minute/1.73 m2, as their disease drivers are more complex, warranting future development of dedicated prediction models; and finally, due to limited access to raw data, an individual patient data meta-analysis could not be conducted, limiting the ability to further adjust for potential confounding factors.
In conclusion, ML and DL algorithms demonstrate strong performance in predicting DKD in patients with T2DM. Key predictors such as age, BMI, eGFR, serum creatinine, urinary albumin, HbA1c, SBP, LDL-C, HDL-C, and triglycerides play significant roles in DKD prediction. Models utilising cross-validation methods show superior predictive capability compared to those employing holdout validation. There is, therefore, a need to standardise disease prediction research. In summary, ML and DL have significant clinical value in predicting DKD, offering an effective solution for early pre
| 1. | Brownrigg JR, Hughes CO, Burleigh D, Karthikesalingam A, Patterson BO, Holt PJ, Thompson MM, de Lusignan S, Ray KK, Hinchliffe RJ. Microvascular disease and risk of cardiovascular events among individuals with type 2 diabetes: a population-level cohort study. Lancet Diabetes Endocrinol. 2016;4:588-597. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 137] [Cited by in RCA: 189] [Article Influence: 18.9] [Reference Citation Analysis (0)] |
| 2. | Jha JC, Banal C, Chow BS, Cooper ME, Jandeleit-Dahm K. Diabetes and Kidney Disease: Role of Oxidative Stress. Antioxid Redox Signal. 2016;25:657-684. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 580] [Cited by in RCA: 509] [Article Influence: 50.9] [Reference Citation Analysis (0)] |
| 3. | Duncan BB, Magliano DJ, Boyko EJ. IDF Diabetes Atlas 11th edition 2025: global prevalence and projections for 2050. Nephrol Dial Transplant. 2025;41:7-9. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 131] [Cited by in RCA: 132] [Article Influence: 132.0] [Reference Citation Analysis (2)] |
| 4. | Drawz P, Rahman M. Chronic kidney disease. Ann Intern Med. 2015;162:ITC1-IT16. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 98] [Cited by in RCA: 176] [Article Influence: 16.0] [Reference Citation Analysis (0)] |
| 5. | Chen J, Zhang Q, Liu D, Liu Z. Exosomes: Advances, development and potential therapeutic strategies in diabetic nephropathy. Metabolism. 2021;122:154834. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 59] [Cited by in RCA: 47] [Article Influence: 9.4] [Reference Citation Analysis (0)] |
| 6. | Levin A, Ahmed SB, Carrero JJ, Foster B, Francis A, Hall RK, Herrington WG, Hill G, Inker LA, Kazancıoğlu R, Lamb E, Lin P, Madero M, McIntyre N, Morrow K, Roberts G, Sabanayagam D, Schaeffner E, Shlipak M, Shroff R, Tangri N, Thanachayanont T, Ulasi I, Wong G, Yang CW, Zhang L, Robinson KA, Wilson L, Wilson RF, Kasiske BL, Cheung M, Earley A, Stevens PE. Executive summary of the KDIGO 2024 Clinical Practice Guideline for the Evaluation and Management of Chronic Kidney Disease: known knowns and known unknowns. Kidney Int. 2024;105:684-701. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 258] [Cited by in RCA: 223] [Article Influence: 111.5] [Reference Citation Analysis (0)] |
| 7. | Jordan MI, Mitchell TM. Machine learning: Trends, perspectives, and prospects. Science. 2015;349:255-260. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8007] [Cited by in RCA: 2170] [Article Influence: 197.3] [Reference Citation Analysis (0)] |
| 8. | van der Laak J, Litjens G, Ciompi F. Deep learning in histopathology: the path to the clinic. Nat Med. 2021;27:775-784. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 776] [Cited by in RCA: 472] [Article Influence: 94.4] [Reference Citation Analysis (0)] |
| 9. | Lin H. Artificial Intelligence with Great Potential in Medical Informatics: A Brief Review. MEDIN. 2024;1:2-9. [DOI] [Full Text] |
| 10. | Obermeyer Z, Emanuel EJ. Predicting the Future - Big Data, Machine Learning, and Clinical Medicine. N Engl J Med. 2016;375:1216-1219. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2898] [Cited by in RCA: 1793] [Article Influence: 179.3] [Reference Citation Analysis (3)] |
| 11. | Esteva A, Robicquet A, Ramsundar B, Kuleshov V, DePristo M, Chou K, Cui C, Corrado G, Thrun S, Dean J. A guide to deep learning in healthcare. Nat Med. 2019;25:24-29. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3854] [Cited by in RCA: 1878] [Article Influence: 268.3] [Reference Citation Analysis (7)] |
| 12. | Li Y, Jin N, Zhan Q, Huang Y, Sun A, Yin F, Li Z, Hu J, Liu Z. Machine learning-based risk predictive models for diabetic kidney disease in type 2 diabetes mellitus patients: a systematic review and meta-analysis. Front Endocrinol (Lausanne). 2025;16:1495306. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 8] [Reference Citation Analysis (0)] |
| 13. | Chen L, Shao X, Yu P. Machine learning prediction models for diabetic kidney disease: systematic review and meta-analysis. Endocrine. 2024;84:890-902. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 11] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 14. | Xu W, Zhou Y, Jiang Q, Fang Y, Yang Q. Risk prediction models for diabetic nephropathy among type 2 diabetes patients in China: a systematic review and meta-analysis. Front Endocrinol (Lausanne). 2024;15:1407348. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 13] [Reference Citation Analysis (0)] |
| 15. | Dai C, Sun X, Xu J, Chen M, Chen W, Li X. The accuracy of Machine learning in the prediction and diagnosis of diabetic kidney Disease: A systematic review and Meta-Analysis. Int J Med Inform. 2025;202:105975. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 5] [Reference Citation Analysis (0)] |
| 16. | Saputro SA, Pattanaprateep O, Pattanateepapon A, Karmacharya S, Thakkinstian A. Prognostic models of diabetic microvascular complications: a systematic review and meta-analysis. Syst Rev. 2021;10:288. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 19] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 17. | Dholariya S, Dutta S, Sonagra A, Kaliya M, Singh R, Parchwani D, Motiani A. Unveiling the utility of artificial intelligence for prediction, diagnosis, and progression of diabetic kidney disease: an evidence-based systematic review and meta-analysis. Curr Med Res Opin. 2024;40:2025-2055. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 5] [Reference Citation Analysis (0)] |
| 18. | Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, Shamseer L, Tetzlaff JM, Akl EA, Brennan SE, Chou R, Glanville J, Grimshaw JM, Hróbjartsson A, Lalu MM, Li T, Loder EW, Mayo-Wilson E, McDonald S, McGuinness LA, Stewart LA, Thomas J, Tricco AC, Welch VA, Whiting P, Moher D. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. 2021;372:n71. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9803] [Reference Citation Analysis (0)] |
| 19. | Moons KG, de Groot JA, Bouwmeester W, Vergouwe Y, Mallett S, Altman DG, Reitsma JB, Collins GS. Critical appraisal and data extraction for systematic reviews of prediction modelling studies: the CHARMS checklist. PLoS Med. 2014;11:e1001744. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1043] [Cited by in RCA: 1449] [Article Influence: 120.8] [Reference Citation Analysis (1)] |
| 20. | Moons KGM, Wolff RF, Riley RD, Whiting PF, Westwood M, Collins GS, Reitsma JB, Kleijnen J, Mallett S. PROBAST: A Tool to Assess Risk of Bias and Applicability of Prediction Model Studies: Explanation and Elaboration. Ann Intern Med. 2019;170:W1-W33. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 667] [Cited by in RCA: 1081] [Article Influence: 154.4] [Reference Citation Analysis (0)] |
| 21. | Collins GS, Moons KGM, Dhiman P, Riley RD, Beam AL, Van Calster B, Ghassemi M, Liu X, Reitsma JB, van Smeden M, Boulesteix AL, Camaradou JC, Celi LA, Denaxas S, Denniston AK, Glocker B, Golub RM, Harvey H, Heinze G, Hoffman MM, Kengne AP, Lam E, Lee N, Loder EW, Maier-Hein L, Mateen BA, McCradden MD, Oakden-Rayner L, Ordish J, Parnell R, Rose S, Singh K, Wynants L, Logullo P. TRIPOD+AI statement: updated guidance for reporting clinical prediction models that use regression or machine learning methods. BMJ. 2024;385:e078378. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1587] [Cited by in RCA: 1389] [Article Influence: 694.5] [Reference Citation Analysis (6)] |
| 22. | Cho BH, Yu H, Kim KW, Kim TH, Kim IY, Kim SI. Application of irregular and unbalanced data to predict diabetic nephropathy using visualization and feature selection methods. Artif Intell Med. 2008;42:37-53. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 94] [Cited by in RCA: 62] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 23. | Dagliati A, Marini S, Sacchi L, Cogni G, Teliti M, Tibollo V, De Cata P, Chiovato L, Bellazzi R. Machine Learning Methods to Predict Diabetes Complications. J Diabetes Sci Technol. 2018;12:295-302. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 148] [Cited by in RCA: 180] [Article Influence: 22.5] [Reference Citation Analysis (0)] |
| 24. | Rodriguez-Romero V, Bergstrom RF, Decker BS, Lahu G, Vakilynejad M, Bies RR. Prediction of Nephropathy in Type 2 Diabetes: An Analysis of the ACCORD Trial Applying Machine Learning Techniques. Clin Transl Sci. 2019;12:519-528. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 36] [Article Influence: 5.1] [Reference Citation Analysis (0)] |
| 25. | Allen A, Iqbal Z, Green-Saxena A, Hurtado M, Hoffman J, Mao Q, Das R. Prediction of diabetic kidney disease with machine learning algorithms, upon the initial diagnosis of type 2 diabetes mellitus. BMJ Open Diabetes Res Care. 2022;10:e002560. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 39] [Article Influence: 9.8] [Reference Citation Analysis (0)] |
| 26. | Dong Z, Wang Q, Ke Y, Zhang W, Hong Q, Liu C, Liu X, Yang J, Xi Y, Shi J, Zhang L, Zheng Y, Lv Q, Wang Y, Wu J, Sun X, Cai G, Qiao S, Yin C, Su S, Chen X. Prediction of 3-year risk of diabetic kidney disease using machine learning based on electronic medical records. J Transl Med. 2022;20:143. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 57] [Article Influence: 14.3] [Reference Citation Analysis (0)] |
| 27. | Nicolucci A, Romeo L, Bernardini M, Vespasiani M, Rossi MC, Petrelli M, Ceriello A, Di Bartolo P, Frontoni E, Vespasiani G. Prediction of complications of type 2 Diabetes: A Machine learning approach. Diabetes Res Clin Pract. 2022;190:110013. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 23] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
| 28. | Sabanayagam C, He F, Nusinovici S, Li J, Lim C, Tan G, Cheng CY. Prediction of diabetic kidney disease risk using machine learning models: A population-based cohort study of Asian adults. Elife. 2023;12:e81878. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 35] [Cited by in RCA: 28] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 29. | Hosseini Sarkhosh SM, Hemmatabadi M, Esteghamati A. Development and validation of a risk score for diabetic kidney disease prediction in type 2 diabetes patients: a machine learning approach. J Endocrinol Invest. 2023;46:415-423. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 20] [Reference Citation Analysis (0)] |
| 30. | Yun C, Tang F, Gao Z, Wang W, Bai F, Miller JD, Liu H, Lee Y, Lou Q. Construction of Risk Prediction Model of Type 2 Diabetic Kidney Disease Based on Deep Learning. Diabetes Metab J. 2024;48:771-779. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 5] [Reference Citation Analysis (0)] |
| 31. | Lin X, Liu C, Wang H, Fan X, Li L, Xu J, Li C, Wang Y, Cai X, Peng X. A SuperLearner approach for predicting diabetic kidney disease upon the initial diagnosis of T2DM in hospital. BMC Med Inform Decis Mak. 2025;25:148. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 2] [Reference Citation Analysis (0)] |
| 32. | Zou LX, Wang X, Hou ZL, Sun L, Lu JT. Machine learning algorithms for diabetic kidney disease risk predictive model of Chinese patients with type 2 diabetes mellitus. Ren Fail. 2025;47:2486558. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 8] [Reference Citation Analysis (0)] |
| 33. | Dei Cas D, Di Camillo B, Fadini GP, Sparacino G, Longato E. The impact of clinical history on the predictive performance of machine learning and deep learning models for renal complications of diabetes. Comput Methods Programs Biomed. 2025;268:108812. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 34. | Gross JL, de Azevedo MJ, Silveiro SP, Canani LH, Caramori ML, Zelmanovitz T. Diabetic nephropathy: diagnosis, prevention, and treatment. Diabetes Care. 2005;28:164-176. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1308] [Cited by in RCA: 1128] [Article Influence: 53.7] [Reference Citation Analysis (1)] |
| 35. | He F, Ng Yin Ling C, Nusinovici S, Cheng CY, Wong TY, Li J, Sabanayagam C. Development and External Validation of Machine Learning Models for Diabetic Microvascular Complications: Cross-Sectional Study With Metabolites. J Med Internet Res. 2024;26:e41065. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 11] [Cited by in RCA: 16] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 36. | Kidney Disease: Improving Global Outcomes (KDIGO) Diabetes Work Group. KDIGO 2022 Clinical Practice Guideline for Diabetes Management in Chronic Kidney Disease. Kidney Int. 2022;102:S1-S127. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 997] [Cited by in RCA: 820] [Article Influence: 205.0] [Reference Citation Analysis (1)] |
| 37. | Chen SJ, Lv LL, Liu BC, Tang RN. Crosstalk between tubular epithelial cells and glomerular endothelial cells in diabetic kidney disease. Cell Prolif. 2020;53:e12763. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 125] [Cited by in RCA: 114] [Article Influence: 19.0] [Reference Citation Analysis (0)] |
| 38. | Wang N, Zhang C. Oxidative Stress: A Culprit in the Progression of Diabetic Kidney Disease. Antioxidants (Basel). 2024;13:455. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 146] [Cited by in RCA: 115] [Article Influence: 57.5] [Reference Citation Analysis (1)] |
| 39. | Yuan Y, Sun X, Yue SJ, Liu Y, Wang HW. [Meta-analysis of influencing factors for early diabetic nephropathy]. Jianyan Yixue Yu Linchuang. 2025;22:439-444. |
| 40. | Fenta ET, Eshetu HB, Kebede N, Bogale EK, Zewdie A, Kassie TD, Anagaw TF, Mazengia EM, Gelaw SS. Prevalence and predictors of chronic kidney disease among type 2 diabetic patients worldwide, systematic review and meta-analysis. Diabetol Metab Syndr. 2023;15:245. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 83] [Reference Citation Analysis (0)] |
| 41. | Wu L, Zhao YY, Li MR, Chang DY, Zhao MH, Chen M. Clinical and pathological characteristics of DKD patients with early-onset type 2 diabetes. J Diabetes Complications. 2023;37:108520. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 42. | Jiang W, Wang J, Shen X, Lu W, Wang Y, Li W, Gao Z, Xu J, Li X, Liu R, Zheng M, Chang B, Li J, Yang J, Chang B. Establishment and Validation of a Risk Prediction Model for Early Diabetic Kidney Disease Based on a Systematic Review and Meta-Analysis of 20 Cohorts. Diabetes Care. 2020;43:925-933. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 192] [Cited by in RCA: 177] [Article Influence: 29.5] [Reference Citation Analysis (0)] |
| 43. | Ismail-Beigi F, Craven T, Banerji MA, Basile J, Calles J, Cohen RM, Cuddihy R, Cushman WC, Genuth S, Grimm RH Jr, Hamilton BP, Hoogwerf B, Karl D, Katz L, Krikorian A, O'Connor P, Pop-Busui R, Schubart U, Simmons D, Taylor H, Thomas A, Weiss D, Hramiak I; ACCORD trial group. Effect of intensive treatment of hyperglycaemia on microvascular outcomes in type 2 diabetes: an analysis of the ACCORD randomised trial. Lancet. 2010;376:419-430. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1146] [Cited by in RCA: 1000] [Article Influence: 62.5] [Reference Citation Analysis (0)] |
| 44. | Kirkman MS, Mahmud H, Korytkowski MT. Intensive Blood Glucose Control and Vascular Outcomes in Patients with Type 2 Diabetes Mellitus. Endocrinol Metab Clin North Am. 2018;47:81-96. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 63] [Cited by in RCA: 49] [Article Influence: 6.1] [Reference Citation Analysis (0)] |
| 45. | Schmieder RE, Mann JF, Schumacher H, Gao P, Mancia G, Weber MA, McQueen M, Koon T, Yusuf S; ONTARGET Investigators. Changes in albuminuria predict mortality and morbidity in patients with vascular disease. J Am Soc Nephrol. 2011;22:1353-1364. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 221] [Cited by in RCA: 197] [Article Influence: 13.1] [Reference Citation Analysis (0)] |
| 46. | Russo GT, De Cosmo S, Viazzi F, Pacilli A, Ceriello A, Genovese S, Guida P, Giorda C, Cucinotta D, Pontremoli R, Fioretto P; AMD-Annals Study Group. Plasma Triglycerides and HDL-C Levels Predict the Development of Diabetic Kidney Disease in Subjects With Type 2 Diabetes: The AMD Annals Initiative. Diabetes Care. 2016;39:2278-2287. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 134] [Cited by in RCA: 122] [Article Influence: 12.2] [Reference Citation Analysis (0)] |
| 47. | Su W, Cao R, He YC, Guan YF, Ruan XZ. Crosstalk of Hyperglycemia and Dyslipidemia in Diabetic Kidney Disease. Kidney Dis (Basel). 2017;3:171-180. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 17] [Cited by in RCA: 36] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 48. | Lin YC, Chang YH, Yang SY, Wu KD, Chu TS. Update of pathophysiology and management of diabetic kidney disease. J Formos Med Assoc. 2018;117:662-675. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 433] [Cited by in RCA: 369] [Article Influence: 46.1] [Reference Citation Analysis (1)] |
| 49. | Sawicki PT, Didjurgeit U, Mühlhauser I, Bender R, Heinemann L, Berger M. Smoking is associated with progression of diabetic nephropathy. Diabetes Care. 1994;17:126-131. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 187] [Cited by in RCA: 162] [Article Influence: 5.1] [Reference Citation Analysis (0)] |