Copyright: ©Author(s) 2026.
World J Diabetes. Mar 15, 2026; 17(3): 112177
Published online Mar 15, 2026. doi: 10.4239/wjd.v17.i3.112177
Published online Mar 15, 2026. doi: 10.4239/wjd.v17.i3.112177
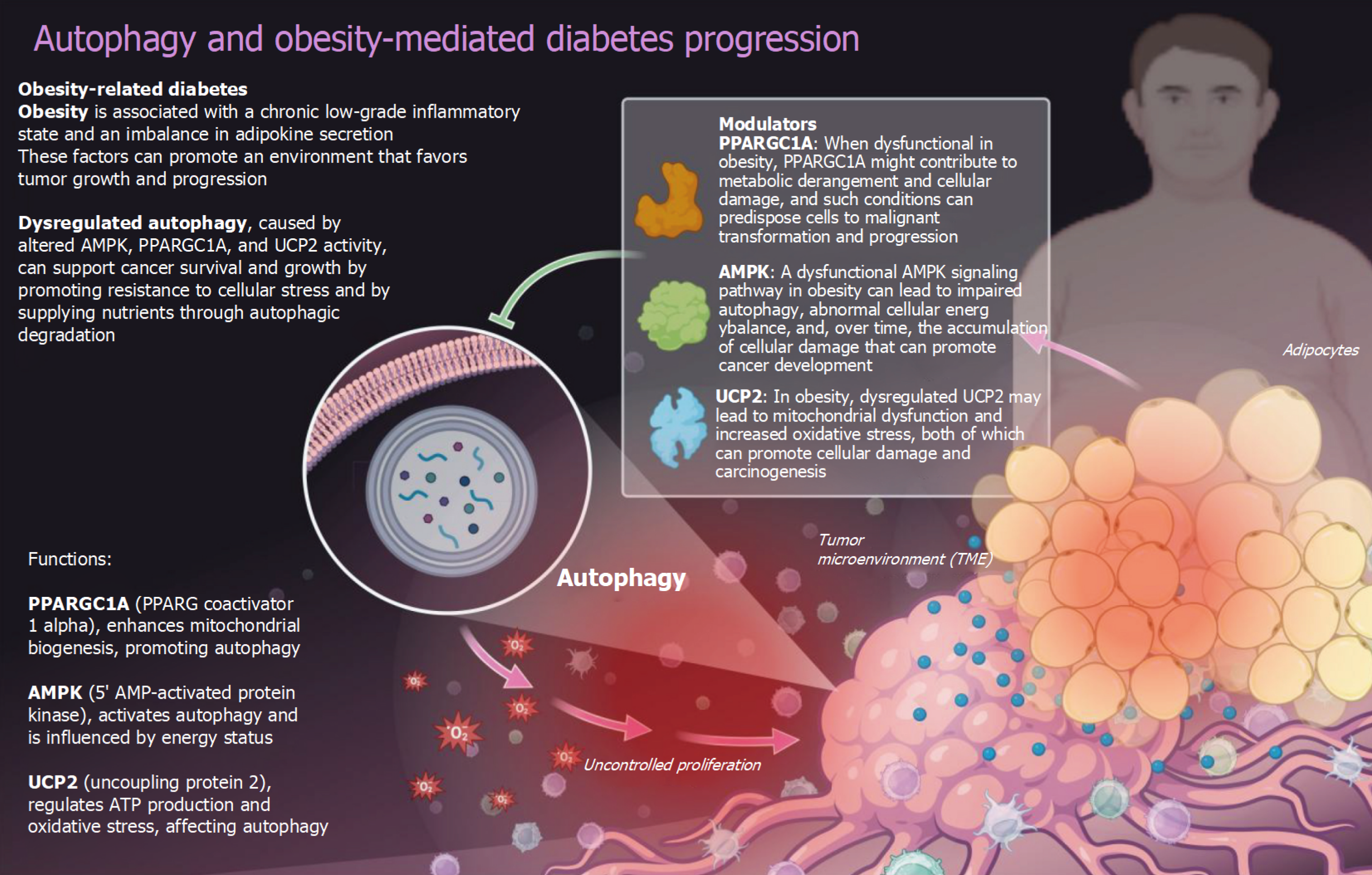
Figure 1 Autophagy- and obesity-mediated diabetes progression.
Illustrations of the relationships among obesity, impaired autophagy, and diabetes development. Under obesity-related conditions, dysregulated autophagy pathways contribute to metabolic dysfunction through multiple mechanisms. The key molecular modulators shown include PPARGC1A (involved in mitochondrial biogenesis and cellular energy metabolism), AMPK (regulating cellular energy status and autophagy initiation), and UCP1 (involved in thermogenesis and energy expenditure). The cellular autophagy machinery, including autophagosome formation and lysosomal degradation processes, becomes compromised in obesity, leading to the accumulation of damaged organelles and metabolic stress. This autophagy dysfunction promotes insulin resistance, inflammation, and ultimately progression to type 2 diabetes mellitus. The figure demonstrates how obesity creates a pathological environment that disrupts normal cellular quality control mechanisms, contributing to the development of diabetic complications. This figure was created by BioRender.com (Supplementary material).
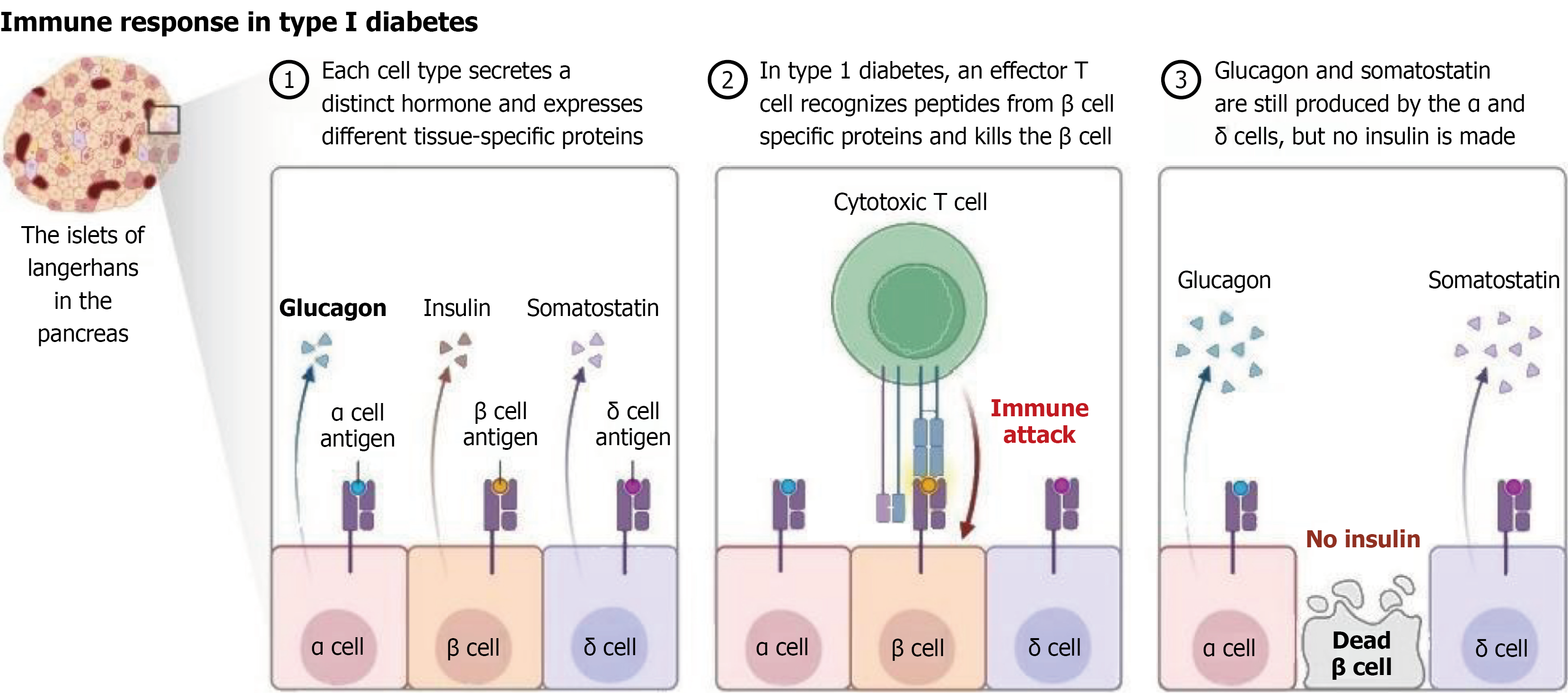
Figure 2 Autoimmune destruction of pancreatic β cells in type 1 diabetes.
The diagram illustrates the pathophysiology of type 1 diabetes mellitus, showing the progressive autoimmune attack on insulin-producing β cells within the pancreatic islets of Langerhans (left panel). The normal pancreatic islet architecture contains α cells (producing glucagon), β cells (producing insulin), and δ cells (producing somatostatin) (middle panel). In type 1 diabetes, autoreactive cytotoxic T cells recognize β cell-specific antigens and mount an immune attack against insulin-producing cells, whereas α and δ cells remain largely unaffected (right panel). The end result of this autoimmune process is the selective destruction of β cells, leading to complete insulin deficiency, whereas glucagon and somatostatin production from α and δ cells continues. This autoimmune-mediated β-cell destruction results in the characteristic hyperglycemia and insulin dependence observed in type 1 diabetes patients. This figure was created by BioRender.com (Supplementary material).
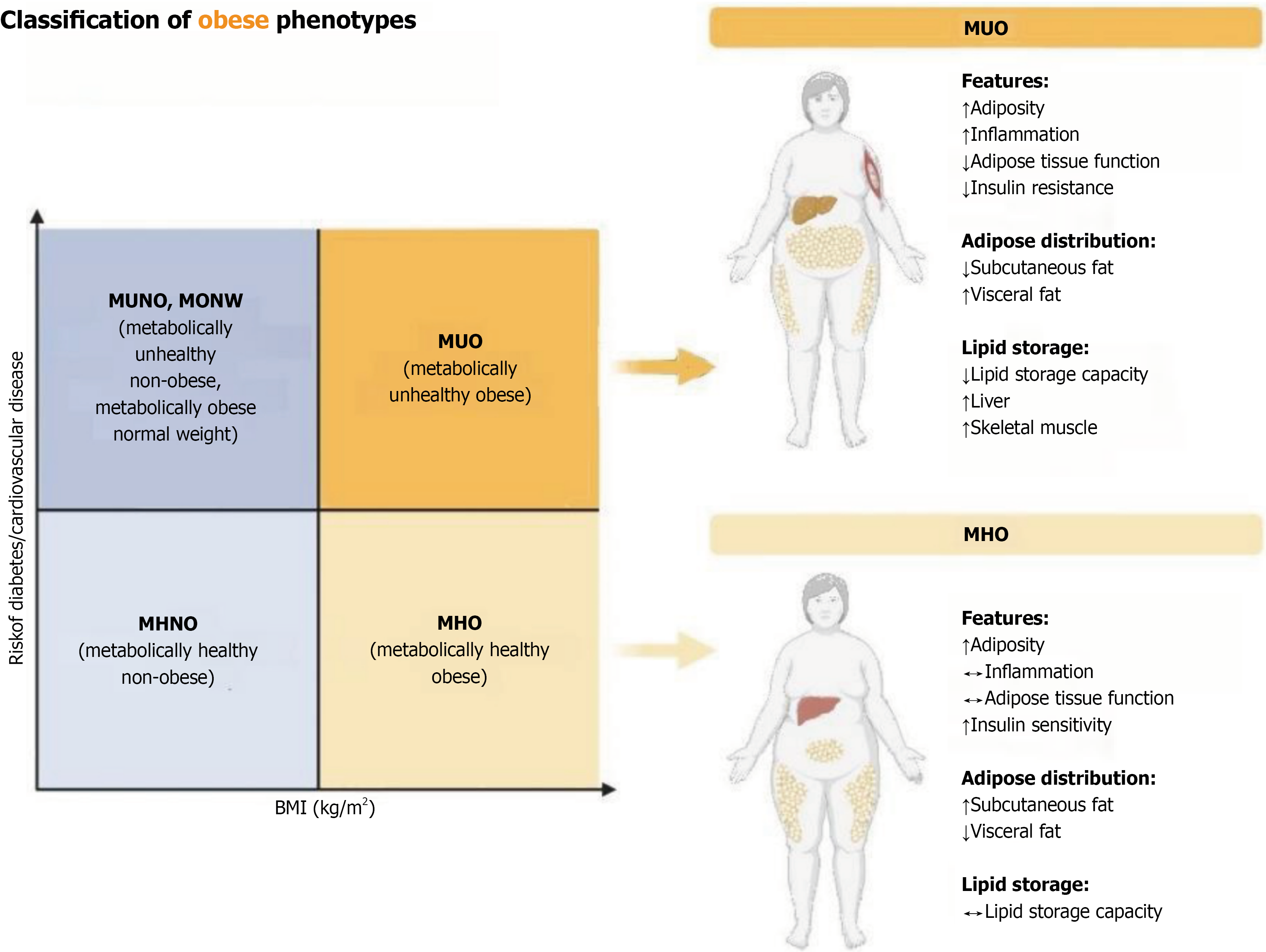
Figure 3 Classification of obesity phenotypes on the basis of metabolic health and body mass index.
The diagram illustrates the heterogeneous nature of obesity by categorizing individuals into four distinct phenotypes on the basis of diabetes/cardiovascular disease risk and body mass index (BMI). The classification includes the following: (1) Metabolically unhealthy obese: Characterized by increased adiposity, inflammation, adipose tissue dysfunction, and insulin resistance, with predominant visceral fat distribution and reduced lipid storage capacity in the liver and skeletal muscle; (2) Metabolically healthy obese: Featuring increased adiposity but preserved metabolic function, predominantly subcutaneous fat distribution, and maintained lipid storage capacity; (3) Metabolically unhealthy normal-weight/metabolically obese normal-weight: Normal BMI individuals with metabolic dysfunction; and (4) Metabolically healthy nonobese: Normal weight individuals with healthy metabolic profiles. This phenotypic classification demonstrated that metabolic health status does not always correlate directly with BMI, highlighting the importance of adipose tissue distribution and function rather than total body weight in determining diabetes and cardiovascular disease risk. MUO: Metabolically unhealthy obese; MHO: Metabolically healthy obese; MUNO/MONW: Metabolically unhealthy normal-weight/metabolically obese normal-weight; MHNO: Metabolically healthy nonobese. This figure was created by BioRender.com (Supplementary material).
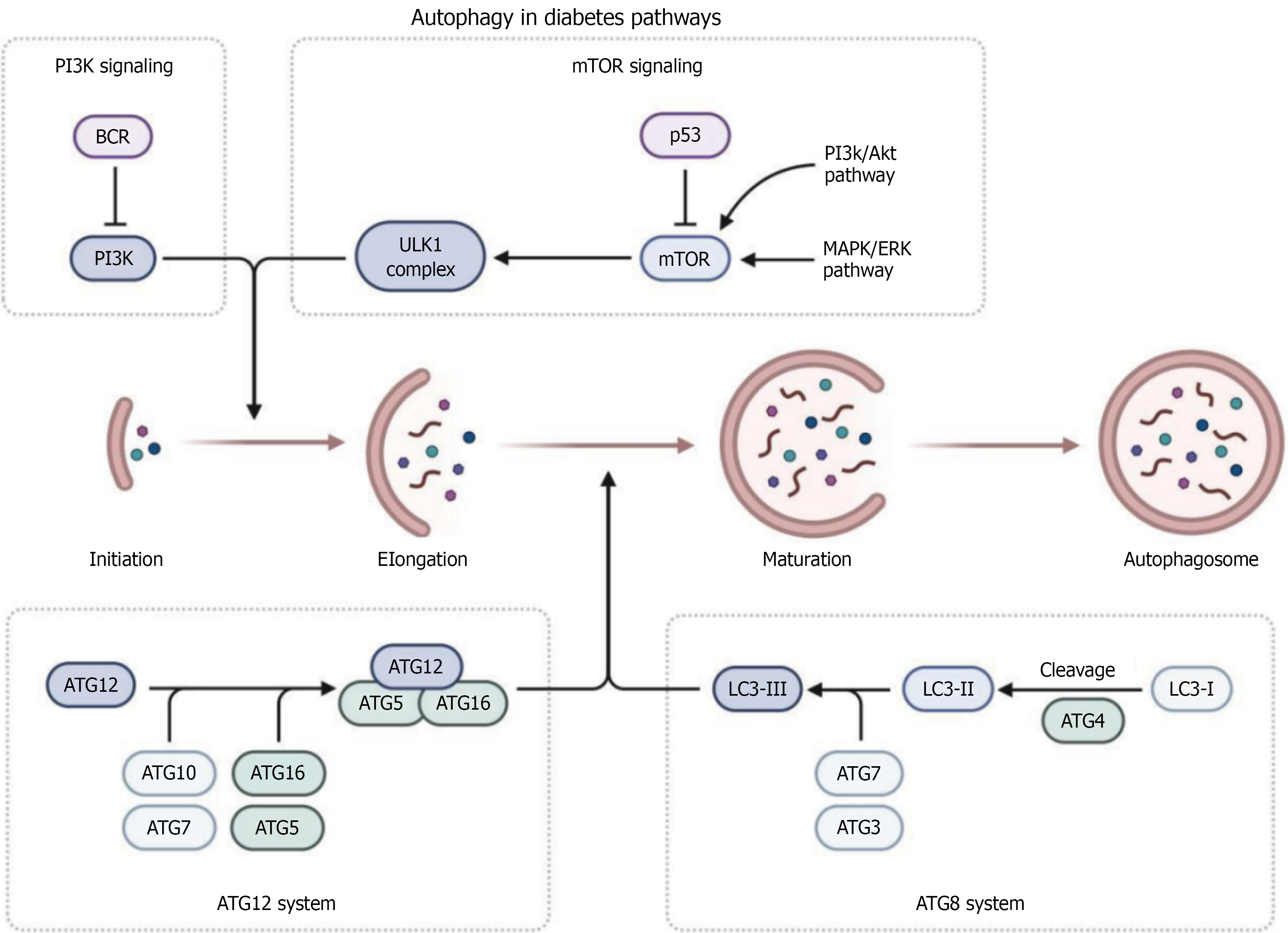
Figure 4 Autophagy signaling pathways and molecular machinery in diabetes.
The diagram illustrates the key regulatory mechanisms controlling autophagy under diabetic conditions (upper panel). Two major signaling pathways regulate autophagy initiation: The PI3K signaling pathway (left), which involves BCR, PI3K, and the ULK1 complex, and the mTOR signaling pathway (right), which involves p53, mTOR, and downstream PI3K/AKT and MAPK/ERK pathways that respond to cellular stress and nutrient availability (middle panel). The autophagy process proceeds through four sequential stages: Initiation of phagophore formation, elongation of the isolation membrane, maturation into a complete autophagosome, and finally, autophagosome formation (lower panel). Two critical molecular systems execute autophagy: The ATG12 system (left), involving ATG12, ATG5, and ATG16 L conjugation through ATG10, ATG16, and ATG7, and the ATG8 system (right), featuring LC3 processing from LC3-I to LC3-II through ATG4-mediated cleavage and ATG7/ATG3-dependent conjugation. This molecular machinery is essential for autophagosome formation and is frequently dysregulated in diabetic conditions, contributing to impaired cellular quality control and metabolic dysfunction. This figure was created by BioRender.com (Supplementary material).
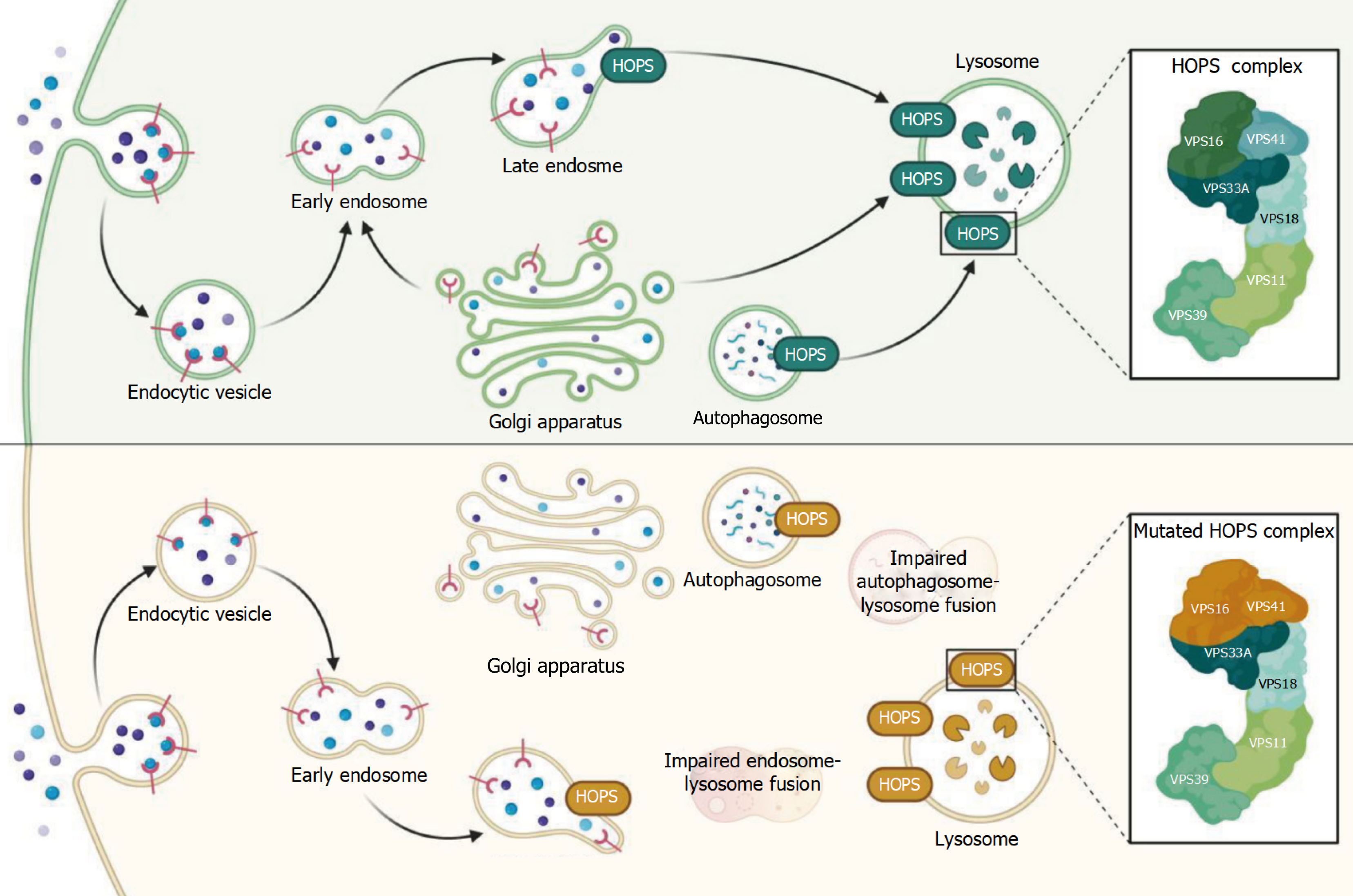
Figure 5 The homotypic fusion and protein sorting complex functions in autophagy-lysosome system fusion and diabetes pathogenesis.
The diagram illustrates the critical role of the homotypic fusion and protein sorting (HOPS) complex in lysosomal fusion processes under normal and pathological conditions (upper panel). The normal HOPS complex facilitates proper vesicle trafficking and fusion events, including endocytic vesicle maturation through early and late endosomes and successful autophagosome-lysosome fusion to form autolysosomes for cargo degradation. The intact HOPS complex (shown in detail on the right) contains multiple subunits that coordinate membrane tethering and fusion (lower panel). Mutating the HOPS complex results in impaired autophagosome-lysosome fusion, leading to the accumulation of undigested autophagosomes and defective lysosomal function. This dysfunction disrupts cellular quality control mechanisms and may contribute to diabetes pathogenesis by affecting pancreatic β-cell autophagy, insulin secretion, and glucose metabolism. HOPS complex mutations represent a potential link between autophagy dysfunction and diabetes development, although the specific connections to neuronal and skin mitophagy in diabetic complications require further investigation. This figure was created by BioRender.com (Supplementary material).
- Citation: Chen X, Zhang LL, Zhou Y, Dong DX, Qian XY. Mitochondrial autophagy in diabetes-related cognitive decline and skin ulcers: Mechanistic insights and therapeutic implications. World J Diabetes 2026; 17(3): 112177
- URL: https://www.wjgnet.com/1948-9358/full/v17/i3/112177.htm
- DOI: https://dx.doi.org/10.4239/wjd.v17.i3.112177