Published online Mar 15, 2026. doi: 10.4251/wjgo.v18.i3.115028
Revised: November 24, 2025
Accepted: January 14, 2026
Published online: March 15, 2026
Processing time: 158 Days and 3.4 Hours
Colorectal cancer (CRC) poses a substantial public health burden worldwide, consistently exhibiting a notable incidence among malignancies across the globe. Goosecoid (GSC), an evolutionarily conserved homeodomain transcription factor, is recognized as a pivotal organizer gene during embryonic development and has been increasingly associated with oncogenic processes. Despite its established roles in other cancers, the specific functions and molecular mechanisms of GSC in the pathogenesis of CRC remain to be fully elucidated. This study therefore aimed to investigate the contribution of GSC to CRC progression and elucidate the underlying mechanistic pathways.
To investigate the oncogenic properties of GSC and elucidate its mechanism of action in CRC.
We validated the expression levels of GSC in CRC tissues using the UALCAN database, Western blotting, im
Through the UALCAN database, Kaplan-Meier Plotter database, western blotting, immunohistochemistry, and immunofluorescence, we found that GSC was significantly upregulated in CRC. High expression of GSC in CRC was associated with poorer recurrence-free survival and overall survival. Furthermore, in vitro studies showed that knocking down GSC inhibited the epithelial-mesenchymal transition (EMT) process, proliferation, invasion, and metastasis of CRC cells; conversely, overexpression of GSC significantly enhanced the EMT process, proliferation, invasion, and metastasis of CRC cells. Mechanistically, we found that knockdown and overexpression of GSC altered the abundance of target proteins linked to the Wnt/β-catenin signaling cascade.
By activating the Wnt/β-catenin signaling axis, GSC drives CRC progression and EMT, positioning it as a viable diagnostic and therapeutic target.
Core Tip: Goosecoid (GSC) is significantly upregulated in colorectal cancer (CRC) tissues and cell lines, with its elevated expression correlated with unfavorable patient outcomes. Functional analyses demonstrate that GSC knockdown attenuates CRC cell proliferation, migration, and invasion. Mechanistically, these inhibitory effects are mediated through the Wnt/β-catenin signaling pathway, concomitant with the regulation of key epithelial-mesenchymal transition markers. Collectively, our findings establish GSC as a critical promoter of CRC growth and metastatic dissemination.
- Citation: Li BC, Wang BZ, Han L, Zhang ZW, Wang Y, Zhang YJ, Zhang JN. Goosecoid drives colorectal cancer progression by inducing epithelial-mesenchymal transition. World J Gastrointest Oncol 2026; 18(3): 115028
- URL: https://www.wjgnet.com/1948-5204/full/v18/i3/115028.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v18.i3.115028
Colorectal cancer (CRC) ranks as the third most common and second most lethal cancer globally[1]. This high mortality is closely linked to its insidious onset; a consequential 20% of cases are diagnosed at a metastatic stage, and disease recurrence or metastasis develops in a further 40% of patients during their clinical course. The 5-year survival rate for CRC patients without metastasis can reach 90%, while for those with metastasis, it drops to 14%[2]. Therefore, metastasis and recurrence are the main causes of death in patients with advanced CRC, making it crucial to identify indicators associated with the initiation, development, metastasis, and recurrence of CRC.
Goosecoid (GSC) belongs to the homeodomain transcription factor family and is highly conserved across diverse animal species. As a key organizer gene, GSC plays a critical role in early embryonic development in vertebrates. Studies have shown that microinjection of GSC mRNA into blastomeres can induce the formation of ectopic secondary body axes[3]. During otic vesicle development, GSC can induce local epithelial-mesenchymal transition (EMT) and promote the delamination of neural crest cells[4]. Substantial evidence indicates that GSC has a central function in vertebrate gastrulation[5-7]. In Xenopus laevis embryos, GSC orchestrates morphogenetic events in gastrulation and governs cell migration[8], implying that its transcriptional targets may participate in processes such as cell signaling, motility, and adhesion. Notably, GSC can be reactivated during tumor progression. Previous studies have reported that GSC promotes metastasis in human breast and liver cancers by initiating the EMT process[9,10]. The development of metastatic disease is a para
Poor prognosis in CRC patients was linked to the observed upregulation of GSC in our analysis. GSC promotes cell invasion and metastasis and maintains the EMT phenotype through the Wnt/β-catenin pathway in CRC, suggesting that GSC may become a new therapeutic target for CRC.
Leveraging the publicly available The Cancer Genome Atlas data embedded within UALCAN, we determined GSC expression levels as log2(TPM+1) values, with the aim of demonstrating the differential expression profile of GSC between CRC and corresponding normal tissues from The Cancer Genome Atlas colon adenocarcinoma cohort.
This study included a total of 3 pairs of colorectal adenocarcinoma tissues and their corresponding adjacent normal tissue samples. All tissue samples in this study were acquired from individuals undergoing surgery for CRC at the Affiliated Hospital of Hebei University of Engineering spanning January 2024 to September 2025, with a confirmed pathological diagnosis of colorectal adenocarcinoma. No patient included in the study underwent radiotherapy, nor any other form of systemic anti-tumor therapy before surgery. The study was performed pursuant to the approval granted by the Ethics Committee of the Affiliated Hospital of Hebei University of Engineering and in conformity with all applicable guidelines (No. 2021[K]112), and a waiver of patient informed consent was granted.
The immunohistochemistry protocol commenced with the processing of CRC and matched normal tissues through standard formalin fixation, paraffin embedding, and microtomy to generate 4-μm sections. Subsequent steps included dewaxing with xylene and hydration via a descending ethanol gradient. Antigen retrieval was achieved by microwaving the sections in citrate buffer (pH 6.0). Thereafter, endogenous peroxidases were quenched by incubation with 3% H2O2, and non-specific sites were blocked using 5% goat serum. The sections were then exposed to an anti-GSC primary antibody (1:200, Signalway, United States) in a cold room overnight. After a phosphate buffered solution (PBS) wash, a species-appropriate secondary antibody was applied for 30 minutes at ambient temperature. Finally, immunoreactivity was visualized with diaminobenzidine, and cell nuclei were identified by hematoxylin counterstaining. Protein ex
After deparaffinization, rehydration, and microwave antigen retrieval in citrate buffer, paraffin sections were blocked with 3% bovine serum albumin for 30 minutes. Subsequently, they were incubated with anti-GSC primary antibody (1:200, Signalway, United States) overnight at 4 °C, then incubated with a Cy3-conjugated secondary antibody for 1 hour at room temperature. Following DAPI nuclear counterstaining, sections were preserved in an anti-fade medium. Finally, fluorescent images were acquired using a fluorescence microscope, and quantification was performed with ImageJ software to conclude the procedure.
To evaluate the prognostic significance of GSC, this study utilized the Kaplan-Meier Plotter to perform survival analysis for 10-year OS and recurrence-free survival (RFS) in a cohort of CRC cases.
HCT116, HCT8, SW480, DLD-1, and HT29 cell lines were acquired from Wuhan Pricella Biotechnology Co., Ltd., while the normal colonic mucosal cell line NCM460 was acquired from Keycell Biotechnology Co., Ltd. (both in Wuhan, Hubei Province, China). We cultured these cell lines in RPMI 1640 medium containing 10% fetal bovine serum and 1% antibiotic-antimycotic solution, with the exception of the SW480 line, which was maintained in DMEM. All cells were incubated in a 37 °C incubator with 50 mL/L CO2.
We performed protein extraction from cells and tissues through ice-cold homogenization for 30 minutes with a lysis buffer containing a protease inhibitor cocktail. After centrifugation at 14000 rpm for 10 minutes at 4 °C, we collected the supernatants and determined total protein concentration. Following a 2-hour block at ambient temperature using 5% skim milk, the nitrocellulose membranes were subsequently incubated with primary antibodies at 4 °C overnight. After extensive washing with PBS for 30 minutes, membranes were probed with appropriate secondary antibodies for 2 hours at room temperature, and then washed again for 20 minutes. Enhanced chemiluminescence reagent (Amersham) was added, and protein signals were visualized in a dark room. Exposure and development were performed after 5 minutes at room temperature. Image Lab software was used for image analysis. GAPDH or β-actin was employed as an internal reference for normalizing protein expression levels. This investigation utilized primary antibodies against the following proteins: (1) GSC (1:1000, Abcam, United States); (2) β-actin (1:30000, Abways, China); (3) GAPDH (1:40000, Abways, China); (4) E-cadherin (1:2000, Abways, China); (5) N-cadherin (1:1500, Abways, China); (6) Vimentin (1:1000, Abways, China); (7) VEGF-C (1:1000, Proteintech, China); (8) MMP9 (1:1000, Proteintech, China); (9) Claudin 1 (1:1000, Abways, China); (10) Snail (1:800, Abways, China); (11) β-catenin (1:1000, Abways, China); (12) Axin2 (1:1500, Abways, China); (13) c-Myc (1:2000, Abways, China); (14) CyclinD1 (1:1000, Abways, China); and (15) MMP7 (1:800, Abways, China). The secondary antibody was affinity-purified biotinylated rabbit anti-rat IgG, purchased from Abways (1:40000, China).
Following the preparation of a single-cell suspension from cells in the logarithmic phase, 500 cells were seeded per well in 6-well plates and cultured over 14 days, with the medium being refreshed periodically. The resulting monolayers were then washed with PBS, underwent fixation in 4% paraformaldehyde for 20 minutes, and were stained with 0.1% crystal violet for 30 minutes. The final count of colonies exceeding 50 cells was performed with Image J.
A base layer was prepared by mixing 0.6% agarose with an equal volume of 2 × complete medium and plating the mixture into 6-well plates. After solidification, a single-cell suspension was combined with 0.35% top agar and overlaid onto the base layer. We added 1-2 mL of complete medium to every well following solidification to minimize evapo
Cells were seeded in 6-well plates at a density of 5 × 105 cells per well. After forming a confluent monolayer, a linear scratch was created in the cell layer using a sterile 200 μL pipette tip. After removal of floating cells by PBS washing, the cultures were switched to low-serum conditions. Wound closure was monitored by microscopy at 0 hour and 48 hours, with quantitative analysis of migration rates performed based on wound width measurements using Image J.
After harvesting, cells were resuspended in serum-free medium at a density of 2 × 104 cells/mL. A 200 μL aliquot of this suspension was seeded into the upper chamber of an 8 μm pore Transwell insert. The lower compartment was filled with complete medium containing 10% fetal bovine serum. Following a 24-hour incubation at 37 °C and 50 mL/L CO2, non-migratory cells on the upper surface were removed. We fixed and stained the cells that had migrated to the lower membrane using paraformaldehyde and crystal violet, respectively, and rinsed with PBS. The membranes were then imaged using an inverted microscope, and the cells were quantified. For the invasion assay, the protocol was identical except that the upper chamber was coated with 100 μL of Matrigel beforehand.
The human GSC shRNA lentiviral vectors and the negative control lentiviral vector were generated by Shanghai GeneChem Co., Ltd. The sequences of the targeting control shRNA are 5′-CCACCGTAGTATTTATAGT-3′ and 5′-GTTCTCCCGAACGTGTCACGT-3′. For stable knockdown of GSC, four CRC lines (HCT116, HCT8, SW480, and DLD-1) were plated 24 hours before transduction. Upon reaching approximately 50% confluence, cells were exposed to lentivirus at a predetermined multiplicity of infection in fresh medium for 24 hours. The viral medium was then exchanged for standard medium, and cells were cultured for another 24 hours prior to experimental use. The knockdown efficiency of GSC was confirmed by Western blotting analysis. Similarly, stable GSC-overexpressing cell lines were generated using the overexpression vector LV-GSC (LV17-EF1a-GSC-CMV-luciferase-T2A-puro) and its negative control LV-NC (LV17-NC), both obtained from GenePharma (Shanghai, China). The overexpression efficiency was also verified by Western blotting.
Matrigel was used as the extracellular matrix substrate and was handled carefully to avoid repeated freeze-thaw cycles. Prior to the experiment, both the 24-well plates and pipette tips were pre-chilled at 4 °C for 1 hour. After complete thawing on ice, we coated 24-well plates with 300 μL of Matrigel and let it polymerize at 37 °C for 30-60 minutes. Stable transfected cell lines with GSC knockdown or overexpression, along with their corresponding controls, were established in HCT116, HCT8, SW480, and DLD-1 cells. Following transfection, cells were cultured in serum-free medium over 24 hours to produce conditioned medium. Subsequently, HUVECs were plated at 1 × 104-3 × 104 cells/well and treated for 12 hours with the collected conditioned media from respective treatment groups. Subsequent tubular structures were documented by microscopy, with total tube length and branch points quantified through ImageJ analysis.
Data are represented as the mean ± SD. Comparative analyses between two groups were performed using the Student's t-test, whereas one-way analysis of variance was applied for multi-group comparisons. The statistical software packages Statistical Package for the Social Sciences 24.0 and GraphPad Prism 8 were utilized for all data analysis. A P value below 0.05 was deemed statistically significant.
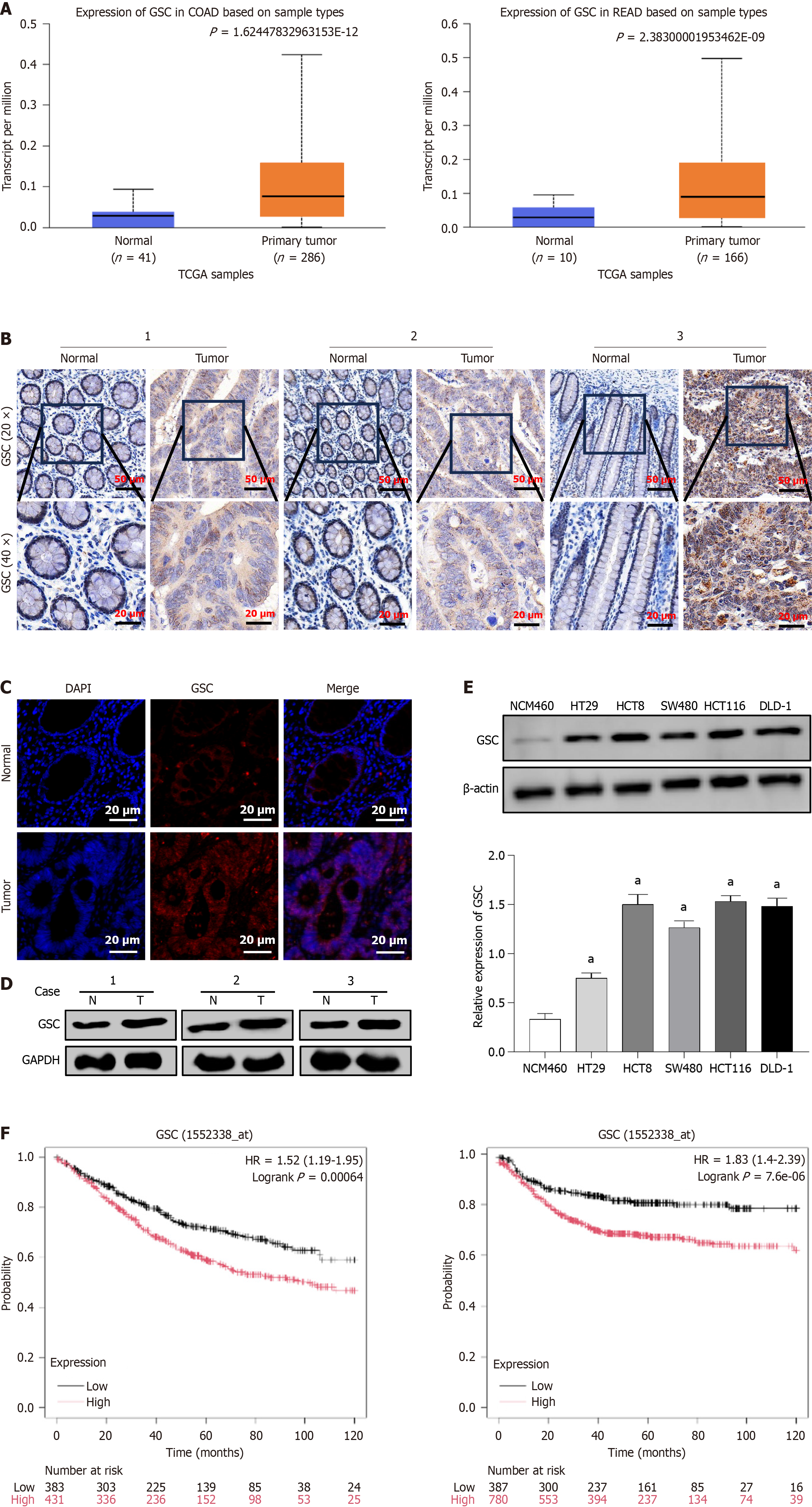
We initially interrogated the UALCAN database, which comprises 41 normal and 286 colon tumor tissues, plus 10 normal and 166 rectal tissues, and found that GSC protein levels were markedly elevated in CRC samples (P < 0.001; Figure 1A). This upregulation was validated in clinical specimens: Immunohistochemistry on tissues from three CRC patients showed stronger GSC staining in tumors than in matched normal mucosa (Figure 1B). Immunofluorescence further delineated the expression pattern, revealing intense nuclear GSC signals in malignant epithelial cells of CRC tissues, whereas normal tissues displayed minimal staining (Figure 1C). Western blotting on patient samples consistently confirmed the pronounced overexpression of GSC protein in CRC relative to adjacent noncancerous tissues (Figure 1D).
A panel of five CRC cell lines (HCT116, HCT8, SW480, DLD-1, and HT29) alongside the normal colonic mucosal line NCM460 were utilized as an in vitro system to assess GSC expression levels. Western blotting results demonstrated that GSC was upregulated across all five CRC cell lines compared to NCM460, with particularly prominent expression in HCT116, HCT8, SW480, and DLD-1 cells (Figure 1E; P < 0.05). To further evaluate the correlation between GSC expression and clinical prognosis, we utilized microarray data from the Kaplan-Meier Plotter database to construct a model associating GSC expression with 10-year survival in CRC patients. Patients with high GSC mRNA levels exhibited an adverse prognosis, characterized by poorer OS (hazard ratio = 1.52, P = 0.00064) and significantly reduced RFS (hazard ratio = 1.83, P = 7.6 × 10-6) (Figure 1F).
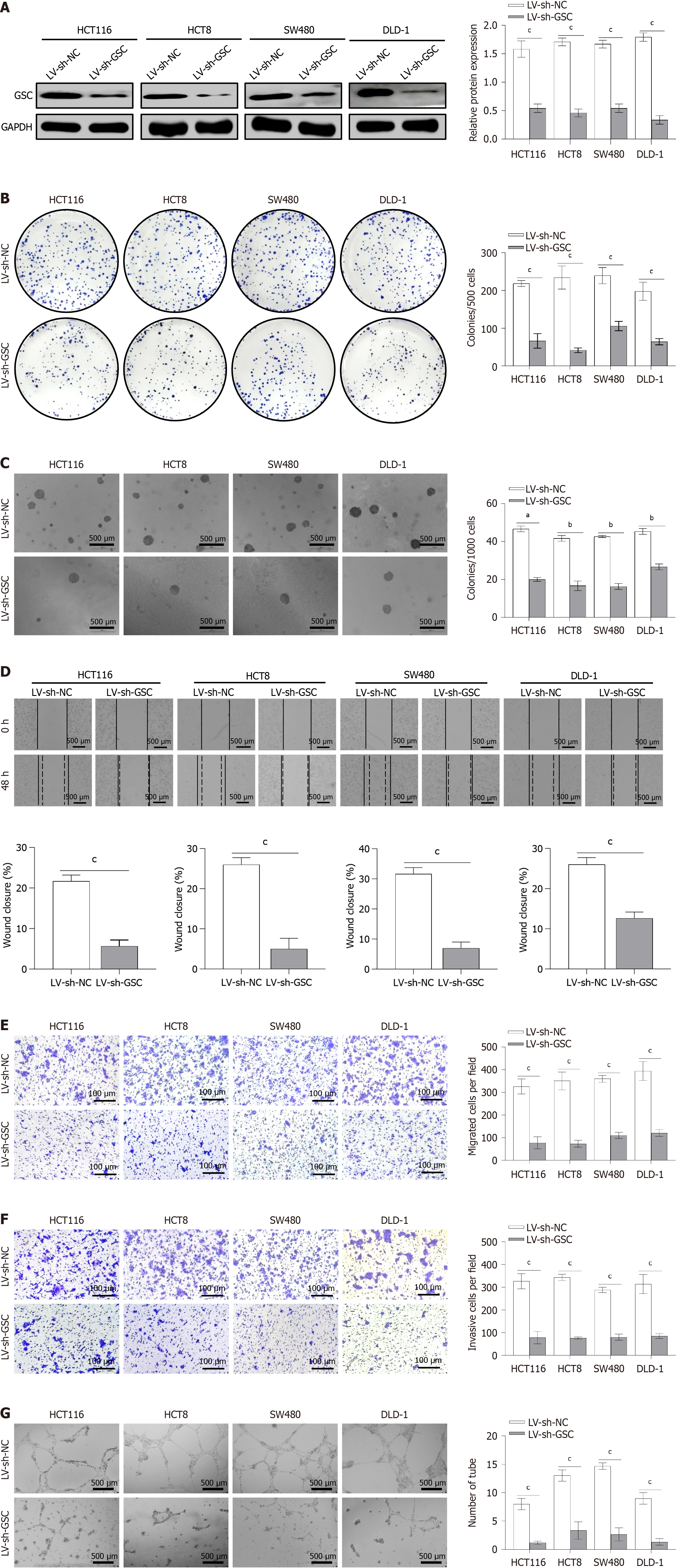
To further investigate the effects of GSC on malignant phenotypes such as cell proliferation, motility, and invasive potential in CRC cells, we silenced GSC expression in four CRC cell lines (HCT116, HCT8, SW480, and DLD-1) via lentivirus-mediated delivery of LV-sh-GSC. Stable knockdown cell lines were selected with 5 μg/mL puromycin. Western blotting analysis confirmed a significant decrease in GSC protein expression in the test group relative to the control group (LV-control shRNA) (P < 0.001; Figure 2A). Using these stable models, we evaluated the effect of GSC on cell proliferation through colony formation and soft agar assays. The results demonstrated that GSC knockdown significantly inhibited the clonogenic ability and anchorage-independent growth of CRC cells (P < 0.01; Figure 2B and C). To further explore the function of GSC in cell motility and invasive ability, we performed wound healing and Transwell experiments. The wound healing assay showed that GSC knockdown noticeably prolonged wound closure in HCT116, HCT8, SW480, and DLD-1 cells, indicating compromised migratory capability (P < 0.001; Figure 2D).
Similarly, the Transwell migration assay revealed a notable decrease in the count of migrated CRC cells in the LV-sh-GSC group in comparison with the control group (P < 0.001; Figure 2E). In the Matrigel-based Transwell invasion assay, GSC knockdown also notably decreased the invasive ability of CRC cells (P < 0.001; Figure 2F). Additionally, to assess whether GSC knockdown inhibits angiogenesis, we performed an in vitro tube formation assay. As shown in Figure 2G, GSC knockdown impaired the ability of cells to form tubular structures (P < 0.001).
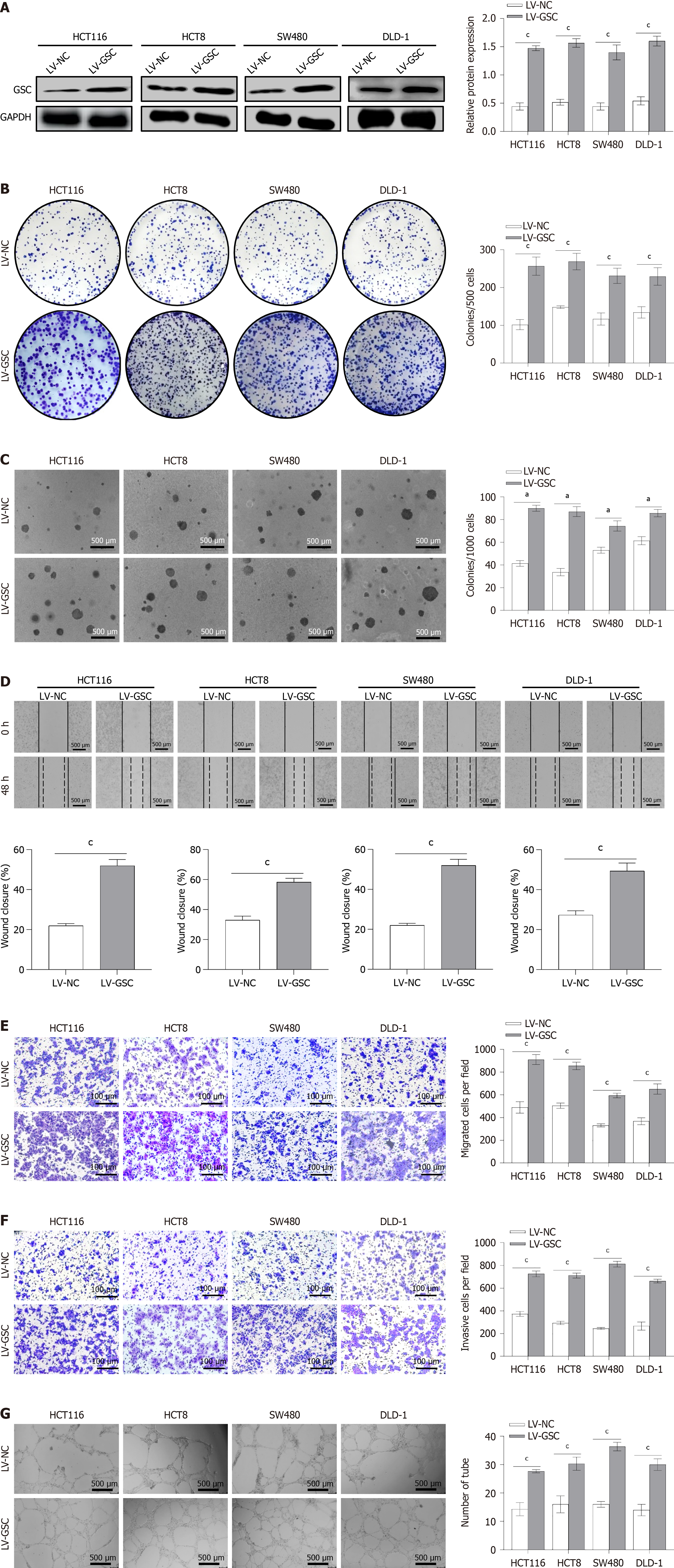
To corroborate the oncogenic function of GSC in CRC, stable GSC-overexpressing lines were generated in HCT116, HCT8, SW480, and DLD-1 cells via lentiviral transduction, using the empty vector LV-NC for control. Subsequent Western blotting verified a pronounced increase in GSC protein levels in LV-GSC cells relative to control cells (P < 0.001; Figure 3A). Using this overexpression system, we evaluated the promotive effects of GSC on malignant phenotypes. Colony formation and soft agar assays demonstrated that GSC overexpression markedly enhanced clonogenic efficiency and anchorage-independent growth in CRC cells (P < 0.01; Figure 3B and C). Furthermore, wound healing assays re
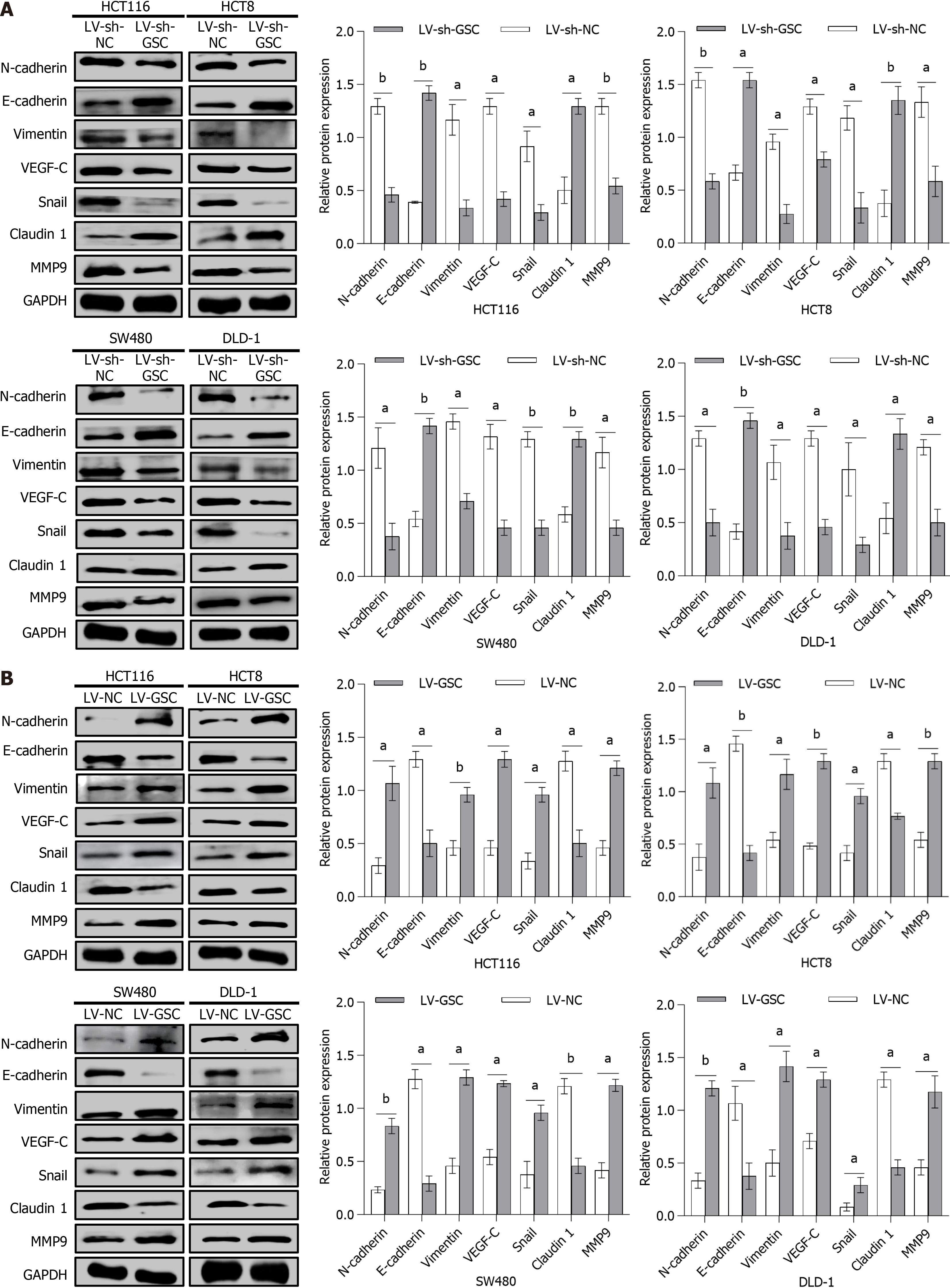
To define GSC’s function in EMT, we systematically analyzed epithelial (E-cadherin and Claudin) and mesenchymal (N-cadherin, Vimentin, Snail, MMP9, and VEGF-C) marker expression in four CRC lines after GSC modulation. Knockdown robustly enhanced epithelial protein expression while reducing mesenchymal marker levels (P < 0.001; Figure 4A). Conversely, GSC overexpression produced opposite effects, with notable downregulation of epithelial markers and concurrent upregulation of mesenchymal proteins (P < 0.001; Figure 4B). These results indicate that GSC modulates EMT-related protein networks to favor mesenchymal transition, suggesting that its metastasis-promoting role in CRC is mediated through EMT activation.
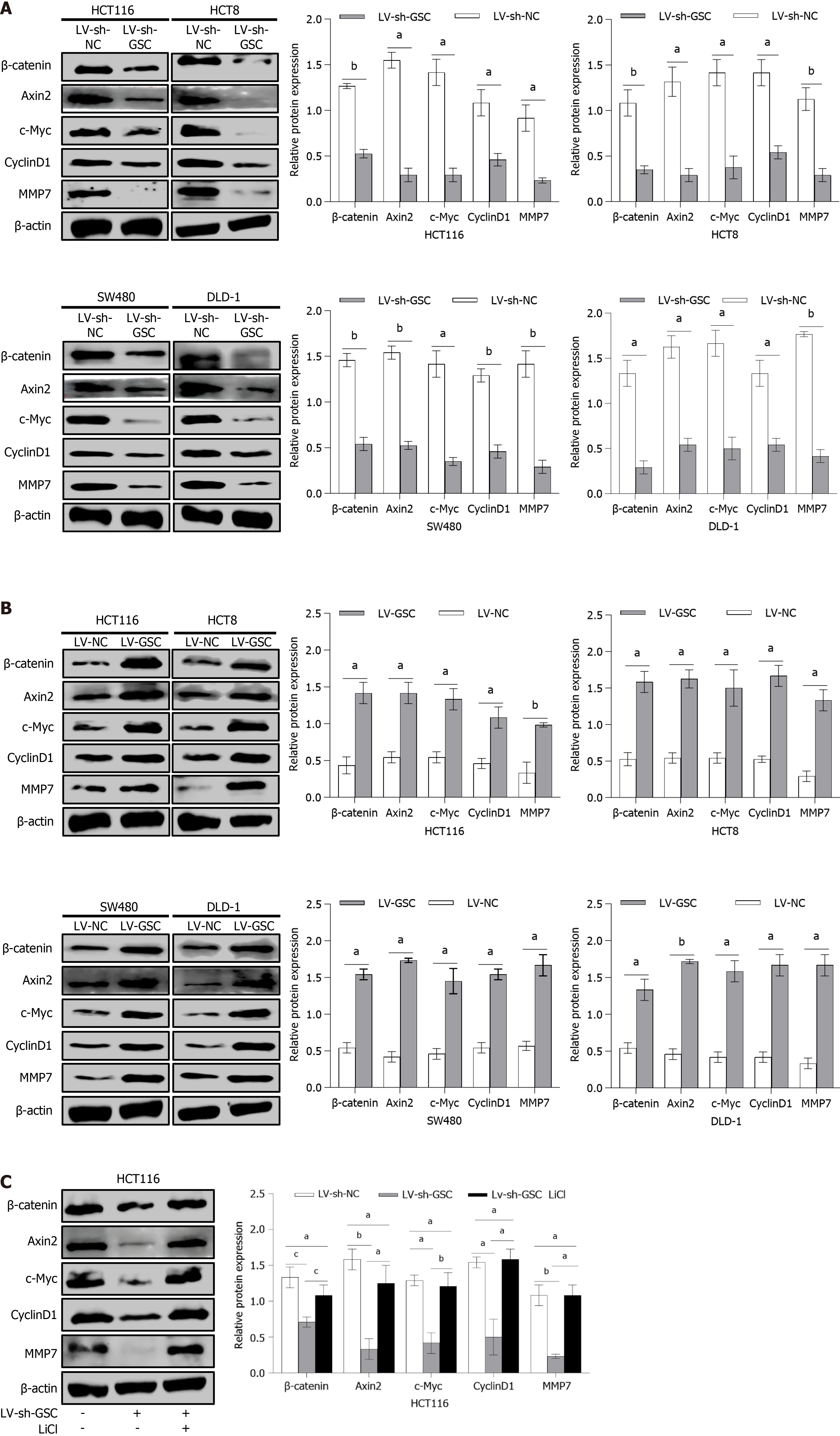
To determine the involvement of the Wnt/β-catenin pathway in GSC-mediated CRC malignant progression, we profiled the expression of its core components following GSC modulation via Western blotting. Silencing GSC in HCT116, HCT8, SW480, and DLD-1 cells substantially suppressed the protein levels of total β-catenin and key downstream targets (c-Myc, Axin2, CyclinD1, and MMP7). In contrast, overexpression of GSC markedly up-regulated the expression of these proteins (P < 0.001; Figure 5A and B). These findings suggest that GSC may positively regulate the Wnt/β-catenin pathway. To more definitively validate the functional involvement of this pathway, we treated GSC-knockdown HCT116 cells with the GSK-3β inhibitor lithium chloride (LiCl) to activate Wnt/β-catenin signaling. The results demonstrated that LiCl treatment reversed the reduced protein expression levels of total β-catenin and its downstream targets (c-Myc, Axin2, Cyclin D1, and MMP7) in GSC-knockdown cells (P < 0.001; Figure 5C). In conclusion, these results indicate that GSC may influence the malignant progression of CRC by modulating the Wnt/β-catenin pathway.
Despite continuous optimization of treatment strategies and improved survival rates in recent years, CRC remains one of the most aggressive malignancies, often characterized by clinical features such as high postoperative recurrence and early metastasis[21]. This study demonstrates that GSC expression is closely associated with the metastatic progression of CRC. To clarify GSC expression levels in CRC, we initially analyzed the UALCAN database and found that GSC expression was significantly elevated in malignant tissues relative to non-cancerous samples. Consistent with this, validation in clinical specimens also revealed markedly higher GSC levels in carcinomas compared with matched non-tumor tissues. Moreover, we consistently observed high GSC expression across multiple CRC cell lines. In addition, we constructed 10-year RFS and OS curves for CRC patients, which indicated that aberrant GSC expression was significantly correlated with poor prognosis in CRC. Subsequent functional experiments demonstrated that ectopic GSC expression markedly promoted the motility and invasiveness of CRC cells, indicating its pivotal contribution to CRC metastasis. Further mechanistic investigations revealed that GSC overexpression induced the EMT process, as evidenced by the suppression of epithelial markers and concurrent elevation of mesenchymal markers. Conversely, knockdown of GSC effectively suppressed the EMT process. These findings suggest that GSC may promote CRC metastasis by regulating EMT.
As cancer treatment has evolved from traditional cytotoxic chemotherapy into the era of precision medicine represented by targeted and immunotherapy, the identification of novel EMT targets capable of overcoming tumor metastasis and drug resistance has emerged as a critical challenge in the field[22]. EMT is a pivotal process wherein tumor cells undergo a transition from an epithelial state to a mesenchymal one, thereby gaining migratory capacity, and is extensively involved in the metastasis and recurrence of various malignancies[23,24]. Its hallmark features include a reduction in epithelial markers like E-cadherin, coupled with an elevation of mesenchymal markers such as N-cadherin and Vimentin, which collectively confer enhanced motility and invasiveness to cells[25-29].
During embryonic development, neural crest cells undergo partial or complete EMT as they delaminate and dissociate from surrounding tissues, subsequently migrating collectively[30,31]. GSC has been implicated in the regulation of neural crest cell function[32]. We therefore hypothesize that GSC may influence tumor cell migration through a similar regulatory mechanism. Furthermore, EMT is not solely dependent on cell-autonomous programs but can also be maintained or reinforced through intercellular interactions within the tumor microenvironment, such as signals derived from extracellular vesicles or cytokines[33]. Building on this, we speculate that GSC expression and its pro-metastatic functions are not purely cell-autonomous events but are likely dynamically modulated by paracrine signaling from stromal components in the tumor microenvironment, such as cancer-associated fibroblasts or immune cells (e.g., tumor-associated macrophages). In summary, the role of GSC in CRC metastasis may represent an integrated outcome of cell-intrinsic molecular characteristics and microenvironment-derived extrinsic signals.
The Wnt/β-catenin signaling pathway is a pivotal regulator of EMT. This pathway activates the EMT process and plays crucial roles in various malignancies, including breast, lung, gastric, and colorectal cancers[21,34-36]. Studies have established that the vast majority of sporadic CRCs display aberrant activation of this signaling axis, which plays a central role in orchestrating fundamental cellular programs including growth, specialization, and motility, thereby critically enabling EMT. Upon Wnt pathway activation, β-catenin escapes phosphorylation and degradation, leading to its stable build-up in the cytosolic compartment. Following its stabilization, β-catenin undergoes nuclear translocation[37,38], where it drives the transcription of EMT-related factors like Slug and Snail1, along with downstream targets including Axin2, c-Myc, and MMP7[21,39], ultimately advancing tumor progression. Our data demonstrate that GSC overexpression elevated total β-catenin protein levels and its key effectors including c-Myc and Cyclin D1. Conversely, GSC suppression reduced their expression, indicating GSC functions as an activator of Wnt/β-catenin signaling. To further validate whether GSC functions through this pathway, we conducted rescue experiments. The results demonstrated that stimulation of the Wnt/β-catenin pathway using the GSK-3β inhibitor LiCl significantly rescued the suppressive effects of GSC knockdown on β-catenin and downstream target gene expression.
Furthermore, cancer cell plasticity has been recognized as a critical determinant of therapy resistance—under stress conditions, tumor cells can undergo transient phenotypic switching independent of canonical EMT transcription factors, thereby generating drug-tolerant subpopulations[40]. Based on our findings, we hypothesize that the GSC-Wnt signaling axis may participate in a broader adaptive regulatory network in CRC cells by modulating the Wnt/β-catenin pathway, consequently influencing phenotypic plasticity and facilitating tumor survival and drug resistance under therapeutic pressure. Meanwhile, emerging high-potential molecular targets such as GSC can be strategically integrated into novel therapeutic frameworks including chimeric antigen receptor T-cell therapy and pathway inhibition. This integration not only leverages the synergistic advantages of new targets and innovative strategies but also transcends the limitations of single-mechanism observations, robustly supporting its potential as a core target for future “precision cure” approaches[22]. Therefore, targeting GSC may not only suppress malignant progression but also potentially enhance treatment sensitivity by disrupting its regulation of cellular plasticity, thereby providing a novel theoretical foundation for combination therapy strategies in CRC.
Although the rescue experiments using LiCl yielded significant results, it should be noted that as a broad-spectrum GSK-3β inhibitor, LiCl may non-specifically activate Wnt signaling through multiple mechanisms. In this study, however, GSC knockdown consistently downregulated all examined key components and downstream targets of the Wnt pathway (β-catenin, CyclinD1, Axin2, c-Myc, and MMP7). This coordinated multi-level molecular phenotype is more likely driven by specific intervention targeting the core machinery of the Wnt pathway rather than by scattered off-target effects. Nevertheless, we acknowledge that the current experimental approach has limitations in conclusively establishing the specificity of this mechanism. Therefore, future work will employ β-catenin degradation activators or specific Wnt inhibitors to functionally validate the GSC-Wnt axis in GSC-overexpressing models, thereby providing more rigorous and direct evidence for the specific regulation of Wnt signaling by GSC. Furthermore, the relatively small sample size included in this study may limit the reliability of our findings. Expanding the cohort should therefore be prioritized in subsequent validation studies. Additionally, in vitro observations may not fully recapitulate the complex regulatory networks operating in vivo, where pathway activity is modulated by multi-tissue and multi-organ crosstalk that cannot be replicated in isolated cell culture systems. Therefore, the physiological relevance of these molecular mechanisms requires final confirmation in vivo models.
Collectively, our findings identify GSC as overexpressed in CRC clinical specimens and cellular models, with its levels correlating with adverse clinical outcomes. Functionally, GSC promotes tumor aggressiveness by coordinately stimulating the Wnt/β-catenin pathway and prompting EMT. These results nominate GSC as both a valuable prognostic indicator and a candidate therapeutic target for CRC.
| 1. | Bray F, Laversanne M, Sung H, Ferlay J, Siegel RL, Soerjomataram I, Jemal A. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2024;74:229-263. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16785] [Cited by in RCA: 14861] [Article Influence: 7430.5] [Reference Citation Analysis (21)] |
| 2. | Siegel RL, Miller KD, Wagle NS, Jemal A. Cancer statistics, 2023. CA Cancer J Clin. 2023;73:17-48. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 12841] [Cited by in RCA: 11251] [Article Influence: 3750.3] [Reference Citation Analysis (4)] |
| 3. | Steinbeisser H, De Robertis EM, Ku M, Kessler DS, Melton DA. Xenopus axis formation: induction of goosecoid by injected Xwnt-8 and activin mRNAs. Development. 1993;118:499-507. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 69] [Cited by in RCA: 55] [Article Influence: 1.7] [Reference Citation Analysis (0)] |
| 4. | Kantarci H, Gerberding A, Riley BB. Spemann organizer gene Goosecoid promotes delamination of neuroblasts from the otic vesicle. Proc Natl Acad Sci U S A. 2016;113:E6840-E6848. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 17] [Cited by in RCA: 14] [Article Influence: 1.4] [Reference Citation Analysis (0)] |
| 5. | Yasuo H, Lemaire P. Role of Goosecoid, Xnot and Wnt antagonists in the maintenance of the notochord genetic programme in Xenopus gastrulae. Development. 2001;128:3783-3793. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 39] [Article Influence: 1.6] [Reference Citation Analysis (0)] |
| 6. | Blum M, Gaunt SJ, Cho KW, Steinbeisser H, Blumberg B, Bittner D, De Robertis EM. Gastrulation in the mouse: the role of the homeobox gene goosecoid. Cell. 1992;69:1097-1106. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 333] [Cited by in RCA: 303] [Article Influence: 8.9] [Reference Citation Analysis (0)] |
| 7. | Luu O, Nagel M, Wacker S, Lemaire P, Winklbauer R. Control of gastrula cell motility by the Goosecoid/Mix.1/ Siamois network: basic patterns and paradoxical effects. Dev Dyn. 2008;237:1307-1320. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 24] [Cited by in RCA: 21] [Article Influence: 1.2] [Reference Citation Analysis (0)] |
| 8. | Niehrs C, Keller R, Cho KW, De Robertis EM. The homeobox gene goosecoid controls cell migration in Xenopus embryos. Cell. 1993;72:491-503. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 161] [Cited by in RCA: 149] [Article Influence: 4.5] [Reference Citation Analysis (0)] |
| 9. | Hartwell KA, Muir B, Reinhardt F, Carpenter AE, Sgroi DC, Weinberg RA. The Spemann organizer gene, Goosecoid, promotes tumor metastasis. Proc Natl Acad Sci U S A. 2006;103:18969-18974. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 180] [Cited by in RCA: 168] [Article Influence: 8.4] [Reference Citation Analysis (3)] |
| 10. | Xue TC, Ge NL, Zhang L, Cui JF, Chen RX, You Y, Ye SL, Ren ZG. Goosecoid promotes the metastasis of hepatocellular carcinoma by modulating the epithelial-mesenchymal transition. PLoS One. 2014;9:e109695. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 32] [Cited by in RCA: 27] [Article Influence: 2.3] [Reference Citation Analysis (4)] |
| 11. | Majidpoor J, Mortezaee K. Steps in metastasis: an updated review. Med Oncol. 2021;38:3. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 226] [Cited by in RCA: 184] [Article Influence: 36.8] [Reference Citation Analysis (0)] |
| 12. | Huang Y, Hong W, Wei X. The molecular mechanisms and therapeutic strategies of EMT in tumor progression and metastasis. J Hematol Oncol. 2022;15:129. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 827] [Cited by in RCA: 745] [Article Influence: 186.3] [Reference Citation Analysis (0)] |
| 13. | Lu J, Kornmann M, Traub B. Role of Epithelial to Mesenchymal Transition in Colorectal Cancer. Int J Mol Sci. 2023;24:14815. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 114] [Cited by in RCA: 100] [Article Influence: 33.3] [Reference Citation Analysis (0)] |
| 14. | Friedl P, Gilmour D. Collective cell migration in morphogenesis, regeneration and cancer. Nat Rev Mol Cell Biol. 2009;10:445-457. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2374] [Cited by in RCA: 1845] [Article Influence: 108.5] [Reference Citation Analysis (0)] |
| 15. | Friedl P, Hegerfeldt Y, Tusch M. Collective cell migration in morphogenesis and cancer. Int J Dev Biol. 2004;48:441-449. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 372] [Cited by in RCA: 307] [Article Influence: 14.0] [Reference Citation Analysis (0)] |
| 16. | Bakir B, Chiarella AM, Pitarresi JR, Rustgi AK. EMT, MET, Plasticity, and Tumor Metastasis. Trends Cell Biol. 2020;30:764-776. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 835] [Cited by in RCA: 778] [Article Influence: 129.7] [Reference Citation Analysis (7)] |
| 17. | Manfioletti G, Fedele M. Epithelial-Mesenchymal Transition (EMT) 2021. Int J Mol Sci. 2022;23:5848. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 91] [Cited by in RCA: 81] [Article Influence: 20.3] [Reference Citation Analysis (2)] |
| 18. | Debnath P, Huirem RS, Dutta P, Palchaudhuri S. Epithelial-mesenchymal transition and its transcription factors. Biosci Rep. 2022;42:BSR20211754. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 258] [Cited by in RCA: 218] [Article Influence: 54.5] [Reference Citation Analysis (3)] |
| 19. | Xu D, Li M, Ran L, Li X, Sun X, Yin T. C5aR1 promotes the progression of colorectal cancer by EMT and activating Wnt/β-catenin pathway. Clin Transl Oncol. 2023;25:440-446. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 19] [Article Influence: 6.3] [Reference Citation Analysis (0)] |
| 20. | Tang Q, Chen J, Di Z, Yuan W, Zhou Z, Liu Z, Han S, Liu Y, Ying G, Shu X, Di M. TM4SF1 promotes EMT and cancer stemness via the Wnt/β-catenin/SOX2 pathway in colorectal cancer. J Exp Clin Cancer Res. 2020;39:232. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 287] [Cited by in RCA: 267] [Article Influence: 44.5] [Reference Citation Analysis (4)] |
| 21. | Yang S, Liu Y, Li MY, Ng CSH, Yang SL, Wang S, Zou C, Dong Y, Du J, Long X, Liu LZ, Wan IYP, Mok T, Underwood MJ, Chen GG. FOXP3 promotes tumor growth and metastasis by activating Wnt/β-catenin signaling pathway and EMT in non-small cell lung cancer. Mol Cancer. 2017;16:124. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 338] [Cited by in RCA: 347] [Article Influence: 38.6] [Reference Citation Analysis (4)] |
| 22. | Sonkin D, Thomas A, Teicher BA. Cancer treatments: Past, present, and future. Cancer Genet. 2024;286-287:18-24. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 437] [Cited by in RCA: 402] [Article Influence: 201.0] [Reference Citation Analysis (0)] |
| 23. | Guarino M. EMT: from embryogenesis, through cancer progression, to the development of carcinosarcoma. Differentiation. 2025;146:100903. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 4] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 24. | Shapiro IM, Cheng AW, Flytzanis NC, Balsamo M, Condeelis JS, Oktay MH, Burge CB, Gertler FB. An EMT-driven alternative splicing program occurs in human breast cancer and modulates cellular phenotype. PLoS Genet. 2011;7:e1002218. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 419] [Cited by in RCA: 386] [Article Influence: 25.7] [Reference Citation Analysis (0)] |
| 25. | Sabouni E, Nejad MM, Mojtabavi S, Khoshduz S, Mojtabavi M, Nadafzadeh N, Nikpanjeh N, Mirzaei S, Hashemi M, Aref AR, Khorrami R, Nabavi N, Ertas YN, Salimimoghadam S, Zandieh MA, Rahmanian P, Taheriazam A, Hushmandi K. Unraveling the function of epithelial-mesenchymal transition (EMT) in colorectal cancer: Metastasis, therapy response, and revisiting molecular pathways. Biomed Pharmacother. 2023;160:114395. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 73] [Cited by in RCA: 61] [Article Influence: 20.3] [Reference Citation Analysis (0)] |
| 26. | Fontana R, Mestre-Farrera A, Yang J. Update on Epithelial-Mesenchymal Plasticity in Cancer Progression. Annu Rev Pathol. 2024;19:133-156. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 155] [Cited by in RCA: 132] [Article Influence: 66.0] [Reference Citation Analysis (0)] |
| 27. | Perelli L, Zhang L, Mangiameli S, Giannese F, Mahadevan KK, Peng F, Citron F, Khan H, Le C, Gurreri E, Carbone F, Russell AJC, Soeung M, Lam TNA, Lundgren S, Marisetty S, Zhu C, Catania D, Mohamed AMT, Feng N, Augustine JJ, Sgambato A, Tortora G, Draetta GF, Tonon G, Futreal A, Giuliani V, Carugo A, Viale A, Kim MP, Heffernan TP, Wang L, Kalluri R, Cittaro D, Chen F, Genovese G. Evolutionary fingerprints of epithelial-to-mesenchymal transition. Nature. 2025;640:1083-1092. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 26] [Cited by in RCA: 20] [Article Influence: 20.0] [Reference Citation Analysis (0)] |
| 28. | Xiao S, Tian L, Gan X, Xu X, Liao M, Song D, Yu Y, Qin W, Zhang R, Lyu H, Guo D, Zhang Q, Chen XZ, Zhou C, Tang J. Role of Ubiquitin-regulated EMT in Cancer Metastasis and Chemoresistance. Int J Biol Sci. 2025;21:6081-6112. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 8] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 29. | Srinivasan D, Balakrishnan R, Chauhan A, Kumar J, Girija DM, Shrestha R, Shrestha R, Subbarayan R. Epithelial-Mesenchymal Transition in Cancer: Insights Into Therapeutic Targets and Clinical Implications. MedComm (2020). 2025;6:e70333. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 15] [Article Influence: 15.0] [Reference Citation Analysis (0)] |
| 30. | Leathers TA, Rogers CD. Time to go: neural crest cell epithelial-to-mesenchymal transition. Development. 2022;149:dev200712. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 35] [Article Influence: 8.8] [Reference Citation Analysis (1)] |
| 31. | Thomas R, Menon V, Mani R, Pruszak J. Glycan Epitope and Integrin Expression Dynamics Characterize Neural Crest Epithelial-to-Mesenchymal Transition (EMT) in Human Pluripotent Stem Cell Differentiation. Stem Cell Rev Rep. 2022;18:2952-2965. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 11] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
| 32. | Parry DA, Logan CV, Stegmann AP, Abdelhamed ZA, Calder A, Khan S, Bonthron DT, Clowes V, Sheridan E, Ghali N, Chudley AE, Dobbie A, Stumpel CT, Johnson CA. SAMS, a syndrome of short stature, auditory-canal atresia, mandibular hypoplasia, and skeletal abnormalities is a unique neurocristopathy caused by mutations in Goosecoid. Am J Hum Genet. 2013;93:1135-1142. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 37] [Cited by in RCA: 32] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 33. | Taki M, Abiko K, Ukita M, Murakami R, Yamanoi K, Yamaguchi K, Hamanishi J, Baba T, Matsumura N, Mandai M. Tumor Immune Microenvironment during Epithelial-Mesenchymal Transition. Clin Cancer Res. 2021;27:4669-4679. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 302] [Cited by in RCA: 277] [Article Influence: 55.4] [Reference Citation Analysis (0)] |
| 34. | Li Q, Lai Q, He C, Fang Y, Yan Q, Zhang Y, Wang X, Gu C, Wang Y, Ye L, Han L, Lin X, Chen J, Cai J, Li A, Liu S. RUNX1 promotes tumour metastasis by activating the Wnt/β-catenin signalling pathway and EMT in colorectal cancer. J Exp Clin Cancer Res. 2019;38:334. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 165] [Cited by in RCA: 161] [Article Influence: 23.0] [Reference Citation Analysis (0)] |
| 35. | Tian S, Peng P, Li J, Deng H, Zhan N, Zeng Z, Dong W. SERPINH1 regulates EMT and gastric cancer metastasis via the Wnt/β-catenin signaling pathway. Aging (Albany NY). 2020;12:3574-3593. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 173] [Cited by in RCA: 163] [Article Influence: 27.2] [Reference Citation Analysis (0)] |
| 36. | Wu Y, Liu H, Gong Y, Zhang B, Chen W. ANKRD22 enhances breast cancer cell malignancy by activating the Wnt/β-catenin pathway via modulating NuSAP1 expression. Bosn J Basic Med Sci. 2021;21:294-304. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 21] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 37. | Liu J, Xiao Q, Xiao J, Niu C, Li Y, Zhang X, Zhou Z, Shu G, Yin G. Wnt/β-catenin signalling: function, biological mechanisms, and therapeutic opportunities. Signal Transduct Target Ther. 2022;7:3. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1990] [Cited by in RCA: 1749] [Article Influence: 437.3] [Reference Citation Analysis (6)] |
| 38. | Zhao H, Ming T, Tang S, Ren S, Yang H, Liu M, Tao Q, Xu H. Wnt signaling in colorectal cancer: pathogenic role and therapeutic target. Mol Cancer. 2022;21:144. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 752] [Cited by in RCA: 625] [Article Influence: 156.3] [Reference Citation Analysis (6)] |
| 39. | Zhu Y, Li X. Advances of Wnt Signalling Pathway in Colorectal Cancer. Cells. 2023;12:447. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 81] [Cited by in RCA: 59] [Article Influence: 19.7] [Reference Citation Analysis (0)] |
| 40. | Bhat GR, Sethi I, Sadida HQ, Rah B, Mir R, Algehainy N, Albalawi IA, Masoodi T, Subbaraj GK, Jamal F, Singh M, Kumar R, Macha MA, Uddin S, Akil ASA, Haris M, Bhat AA. Cancer cell plasticity: from cellular, molecular, and genetic mechanisms to tumor heterogeneity and drug resistance. Cancer Metastasis Rev. 2024;43:197-228. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 211] [Cited by in RCA: 156] [Article Influence: 78.0] [Reference Citation Analysis (1)] |