Published online Nov 15, 2025. doi: 10.4251/wjgo.v17.i11.109481
Revised: August 12, 2025
Accepted: October 20, 2025
Published online: November 15, 2025
Processing time: 142 Days and 17.3 Hours
Colorectal cancer (CRC) is one of the most common causes of cancer mortality worldwide. The transcription factor Myc-associated zinc finger protein (MAZ) has been implicated in cancer progression. However, its precise function and mecha
To investigate the role and mechanism of the MAZ/ubiquitin-like with PHD and RING finger domains 1 (UHRF1)/esophageal cancer-related gene 4 (ECRG4) axis in CRC metastasis.
Western blot, quantitative reverse transcription polymerase chain reaction (PCR) and transwell were performed to evaluate the impact of MAZ knockdown on CRC cell migration and invasion. A xenograft tumor metastasis model was es
MAZ was highly upregulated in CRC cells and promoted CRC migration, inva
MAZ promotes CRC cell migration, invasion, and EMT by transcriptionally activating UHRF1, which downregulates ECRG4 through DNA methylation.
Core Tip: This study elucidated the oncogenic role of Myc-associated zinc finger protein (MAZ) in colorectal cancer (CRC), demonstrating its ability to promote cancer cell migration, invasion, and epithelial-mesenchymal transition by modulating the ubiquitin-like with PHD and RING finger domains 1-esophageal cancer-related gene 4 signaling pathway. MAZ was up-regulated in CRC cell species, and inhibition of its expression significantly inhibited cell invasion, invasion, epithelial-mesenchymal transition and tumor metastasis. Dual-luciferase reporter assay, chromatin immunoprecipitation quantitative polymerase chain reaction and methylation-specific polymerase chain reaction further validated the interaction of the MAZ-ubiquitin-like with PHD and RING finger domains 1-esophageal cancer-related gene 4 axis, providing convincing evidence for MAZ as a potential therapeutic target for CRC.
- Citation: Mao HQ, Yu FC, Hu DQ, Zhang LJ. Myc-associated zinc finger protein drives colorectal cancer metastasis through activating ubiquitin like with ring finger protein one. World J Gastrointest Oncol 2025; 17(11): 109481
- URL: https://www.wjgnet.com/1948-5204/full/v17/i11/109481.htm
- DOI: https://dx.doi.org/10.4251/wjgo.v17.i11.109481
Globally, colorectal cancer (CRC) is a major cause of cancer mortality, largely due to metastasis, which leads to poor prognosis and treatment failure[1]. CRC metastasis is a multifaceted process regulated by various molecular pathways. Studies have shown that epithelial-mesenchymal transition (EMT) enables CRC cells to acquire stronger migratory and invasive abilities, helping cancer cells break through the primary tumor tissue and metastasize to other organs[2]. Although research on CRC has advanced, the poor prognosis associated with its metastasis and the underlying metastatic mechanisms remain incompletely elucidated. Therefore, further identification of key molecules influencing CRC meta
Myc-associated zinc finger protein (MAZ), a transcription factor, has been recognized as a key regulator of gene expression in various cancers[3,4]. In normal physiological conditions, MAZ plays a role in maintaining cellular ho
Ubiquitin-like with PHD and RING finger domains 1 (UHRF1), also referred to as ICBP90, is a multifunctional epigenetic regulator with a pivotal role in maintaining DNA methylation, chromatin remodeling, and gene expression[10]. Research indicates that UHRF1 levels are elevated in CRC tumor tissues, where it drives progression[11,12]. These functional domains, which recognize histones and hemimethylated DNA, are crucial for maintaining CRC-specific DNA methylation[13]. By using the National Center for Biotechnology Information, University of California Santa Cruz, and JASPAR databases, we found multiple MAZ binding sites on the UHRF1 promoter.
Esophageal cancer-related gene 4 (ECRG4) is a newly discovered tumor suppressor. Studies have shown that ECRG4 expression is reduced in CRC tissues and reduces the proliferation of CRC cell lines[14,15]. Genome-wide analysis has revealed hypermethylation of the ECRG4 promoter in CRC. Since DNA methylation is a primary cause of tumor suppressor gene silencing in cancer, this suggests that hypermethylation of ECRG4 may contribute to CRC tumor metastasis[16]. This study aimed to elucidate the mechanism by which MAZ regulates CRC metastasis. By integrating in vivo and in vitro models, we explored the role of MAZ in regulating CRC migration, invasion, and EMT. Mechanistically, our study affirmed the correlation between MAZ, UHRF1, and ECRG4. These findings provide new insights into the molecular mechanisms underlying CRC metastasis.
We used the human normal colon epithelial cell line NCM460 and two CRC cell lines, HCT116 and SW480 (American Type Culture Collection, VA, United States). All cells were cultured at 37 °C in a 5% CO2 environment using Dulbecco’s modified Eagle’s medium or RPMI-1640 medium supplemented with 10% fetal bovine serum (Gibco, NY, United States) and 1% penicillin-streptomycin. Cells were treated with drugs or transfected when they reached 80%-90% confluence. Cell transfection was performed using Lipofectamine 3000 reagent (Invitrogen, CA, United States). Transfection was carried out with short hairpin (sh)-RNA RNA solution, and fresh medium was replaced 48 hours after transfection for subsequent experiments. Transfection efficiency was confirmed by quantitative reverse transcription polymerase chain reaction (qRT-PCR) and western blot.
Total RNA was extracted from HCT116 and SW480 cells or nude mouse tissues using TRIzol reagent (Thermo Fisher Scientific, MA, United States), and RNA concentration was measured using a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, MA, United States). RNA (1 μg) was reverse-transcribed to complementary DNA using the Reverse Transcription System (Thermo Fisher Scientific, MA, United States). qRT-PCR was performed using SYBR Green PCR Master Mix (Applied Biosystems, CA, United States) on a QuantStudio 3 real-time PCR system (Applied Biosystems, CA, United States). The total reaction volume was 20 μL, containing 10 μL SYBR Green mix, 1 μL complementary DNA template, 0.4 μM primers, and water. Glyceraldehyde-3-phosphate dehydrogenase was used as the internal control, and data were analyzed using the 2-ΔΔCt) method. The sequences of the primers were shown in Table 1.
| Gene | Forward | Reverse |
| MAZ | TCTACCACCTGAACCGACAC | GAGCAGTTGTAGGGCTTGTG |
| UHRF1 | AGGTCAATGAGTACGTCGATGC | TTCTCCGGGTAGTCGTCGT |
| ECRG4 | ACTAAGACTAAAGTGGCCGTTG | AATTTCGCTTCGTCAAAGCCC |
| GAPDH | GAGTCAACGGATTTGGTCGT | GACAAGCTTCCCGTTCTCAG |
HCT116 and SW480 cells and lung tumors were lysed using radioimmunoprecipitation assay buffer (Beyotime, Shanghai, China) to extract total protein. Protein concentration was measured using the BCA Protein Assay Kit (Thermo Fisher Scientific, MA, United States). Equal amounts of protein were separated by sodium-dodecyl sulfate gel electrophoresis and transferred to a polyvinylidene difluoride membrane (Millipore, MA, United States). After blocking, the membrane was incubated overnight with primary antibodies, followed by horseradish-peroxidase-conjugated secondary antibodies (Cell Signaling Technology, MA, United States). Protein detection was performed using chemiluminescence (Thermo Fisher Scientific, MA, United States). MAZ (1:1000; 21068-1-AP) was from Proteintech (Wuhan, China), and UHRF1 (1:1000; #12387), E-cadherin (1:1000; #3195), N-cadherin (1:1000; #13116), vimentin (1:1000; #5741), Snail (1:1000; #3879), and glyceraldehyde-3-phosphate dehydrogenase (1:1000; #2118) were from Cell Signaling Technology, MA, United States. ECRG4 (1:1000; PA5-104506) was from Thermo Fisher Scientific, MA, United States. Data were analyzed using ImageJ software, and all experiments were repeated at least three times.
Transfected HCT116 and SW480 cells were seeded into uncoated Transwell chambers (Corning, NY, United States) at 5 × 104 cells/well, with 10% fetal bovine serum-containing medium in the lower chamber as a chemoattractant. After incubating for 24 hours, cells were fixed, and stained with crystal violet (Sigma-Aldrich, MO, United States). Cell mi
The 3’ untranslated region fragment of the UHRF1 gene was cloned into the pMIR-REPORT (Promega, WI, United States) reporter vector and cotransfected with the Renilla luciferase gene into HCT116 and SW480 cells. After 24 hours of transfection, luciferase activity was detected using a luciferase substrate (Promega, WI, United States) and measured with a GloMax Luminometer (Promega, WI, United States). Luciferase activity was determined based on the ratio of Firefly to Renilla luciferase activity, and results were analyzed using GraphPad Prism software.
HCT116 and SW480 cells were crosslinked with 1% formaldehyde and subsequently sonicated to fragment the DNA. DNA fragments were immunoprecipitated and analyzed by quantitative polymerase chain reaction (PCR). Specific antibodies were used for immunoprecipitation, and SYBR Green quantitative PCR was performed for DNA fragment quantification, with immunoglobulin G as a negative control (NC). Data were analyzed using the 2-ΔΔCt method to com
Genomic DNA was extracted from HCT116 and SW480 cells, and methylation-specific or unmethylated-specific primers were used for PCR amplification. Amplified products were separated by agarose gel electrophoresis and stained with ethidium bromide (Sigma-Aldrich, MO, United States). The methylation status was determined by comparing the ampli
Four-week-old female nude mice (SLACOM Company, Shanghai, China) were utilized to develop a tumor metastasis model (n = 6 per group). Stable MAZ knockdown HCT116 cells (106/100 μL saline) were injected into the tail veins of mice to establish a tumor metastasis model. Two groups were formed from the mice: The sh-MAZ group (n = 6) and the sh-NC group (n = 6). The sh-MAZ group was injected via the tail vein with stable MAZ-knockdown HCT116 cells (106/100 mL saline), while the sh-NC group was injected with an equal amount of HCT116 cells transfected with the control plasmid sh-NC. Mouse body weight was monitored regularly, and lung tumor growth and metastasis volume were measured. Six weeks after cell injection, the mice were killed for sample collection and further analysis, using CO2 asphyxiation followed by cervical dislocation, ensuring minimal animal suffering and compliance with ethical guidelines.
Lung tumor specimens were extracted from nude mice, fixed in 4% paraformaldehyde, dehydrated, and embedded in paraffin. Sections of 4 μm thickness were stained with hematoxylin and eosin (Solarbio, Beijing, China). Histological changes of tumors were observed and analyzed under a microscope (Leica DM2500, Germany).
All experimental data were analyzed using GraphPad Prism version 9.5. Between-group comparisons were performed using one-way ANOVA or t test, and data were expressed as mean ± SD. P < 0.05 was considered significant. For mul
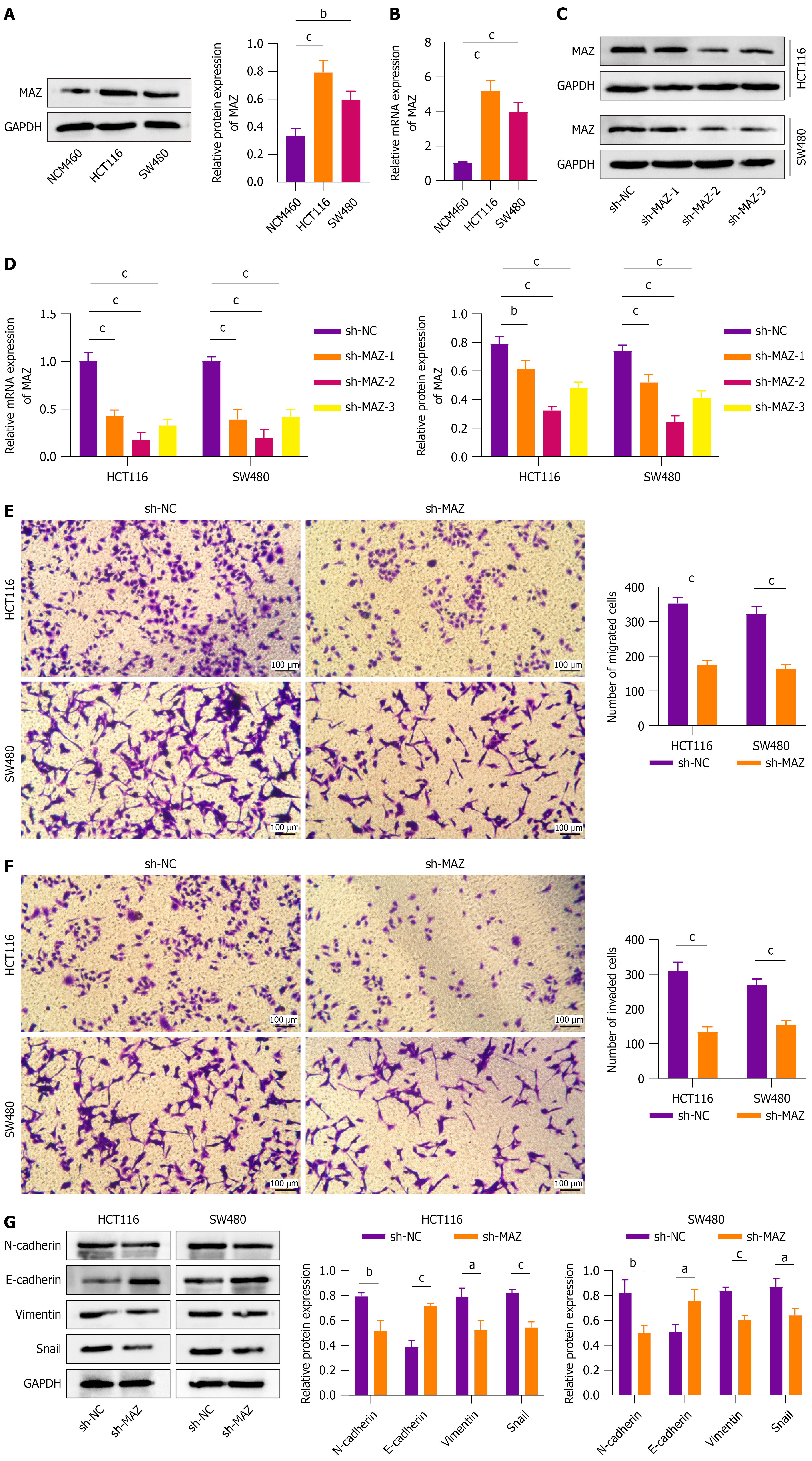
MAZ is an essential transcriptional regulator controlling gene expression under normal physiological conditions but becomes abnormally activated in cancer[17]. To explore whether MAZ participates in the regulation of CRC progression, we conducted a series of experimental validations. Western blot and qRT-PCR (Figure 1A and B) showed that MAZ expression was significantly higher in HCT116 and SW480 cells. Therefore, sh-RNA was used to interfere with MAZ expression in CRC cell lines for further analysis. All three sh-RNAs significantly knocked down MAZ expression, with sh-MAZ-2 exhibiting the most pronounced effect (Figure 1C and D). Therefore, sh-MAZ-2 was selected for subsequent experiments. The transwell assay demonstrated that silencing of MAZ strongly inhibited CRC cell migration and invasion (Figure 1E and F, P < 0.05). To rule out the off-target effect of sh-RNA, we further knocked down MAZ using another two sh-RNAs, and evaluated the migration and invasion of CRC cells (Supplementary Figure 1). Sh-MAZ-1 and sh-MAZ-3 also significantly inhibited migration and invasion of CRC cells (P < 0.05). We assessed EMT-related protein expression in CRC cell lines using western blot. Silencing MAZ upregulated E-cadherin and downregulated N-cadherin, vimentin, and snail levels, indicating inhibition of EMT. These results suggest that MAZ is overexpressed in CRC cells, and facilitates cell migration, invasion, and EMT (Figure 1G).
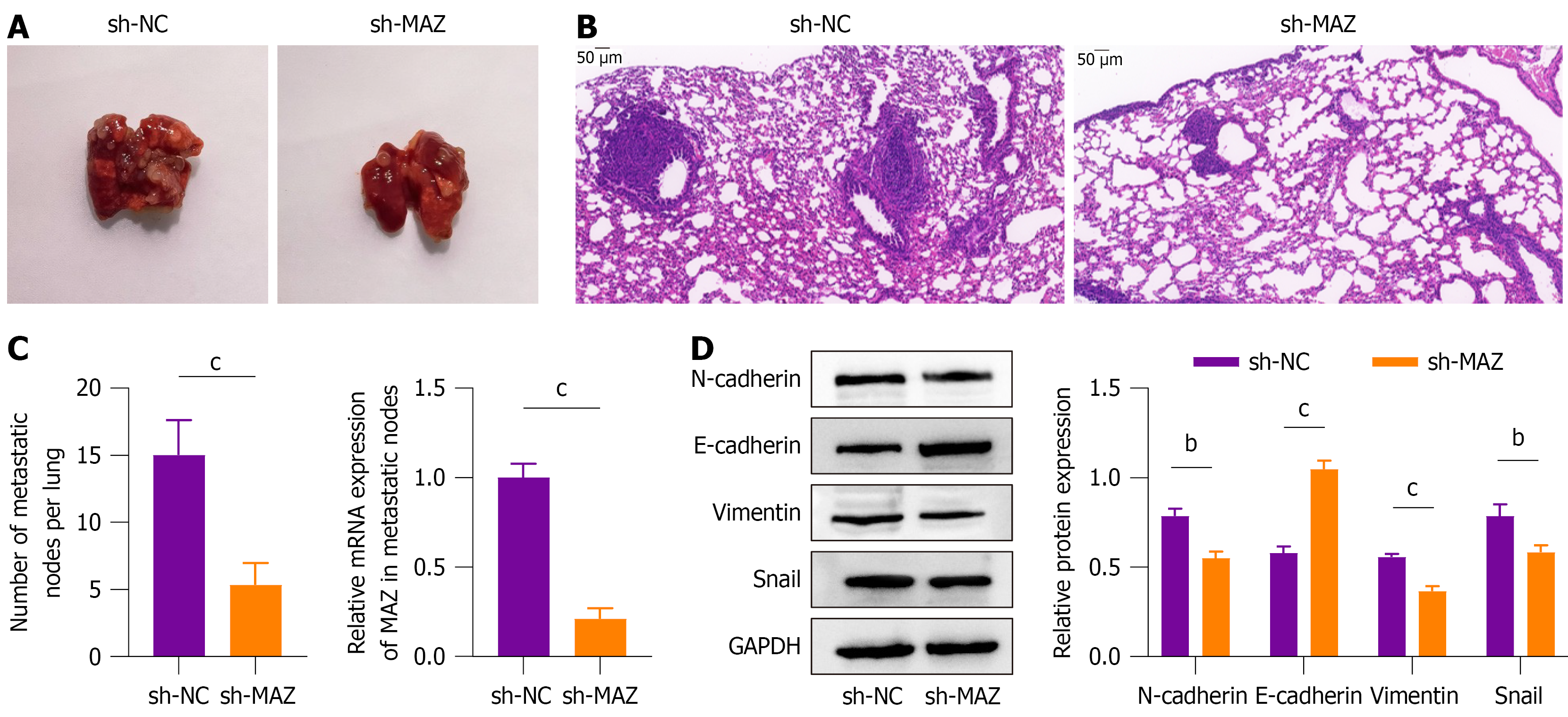
To further investigate the impact of MAZ on CRC tumor metastasis, a xenograft tumor metastasis model was established by injecting sh-NC or sh-MAZ transfected CRC cells into the tail veins of nude mice. After 6 weeks, both groups of nude mice developed lung metastases, and MAZ knockdown significantly reduced metastatic tumor nodules compared to the sh-NC group (Figure 2A). Hematoxylin and eosin staining of lung tissues revealed that MAZ knockdown resulted in a significantly lower number of metastatic nodules (Figure 2B). MAZ mRNA level was significantly decreased in metastatic tumor nodules (Figure 2C). Regarding EMT, western blot demonstrated that MAZ knockdown led to increased E-cadherin expression and decreased levels of N-cadherin, vimentin, and snail, indicating suppression of EMT in metastatic nodules (Figure 2D). The results demonstrated that knockdown of MAZ inhibited CRC tumor metastasis.
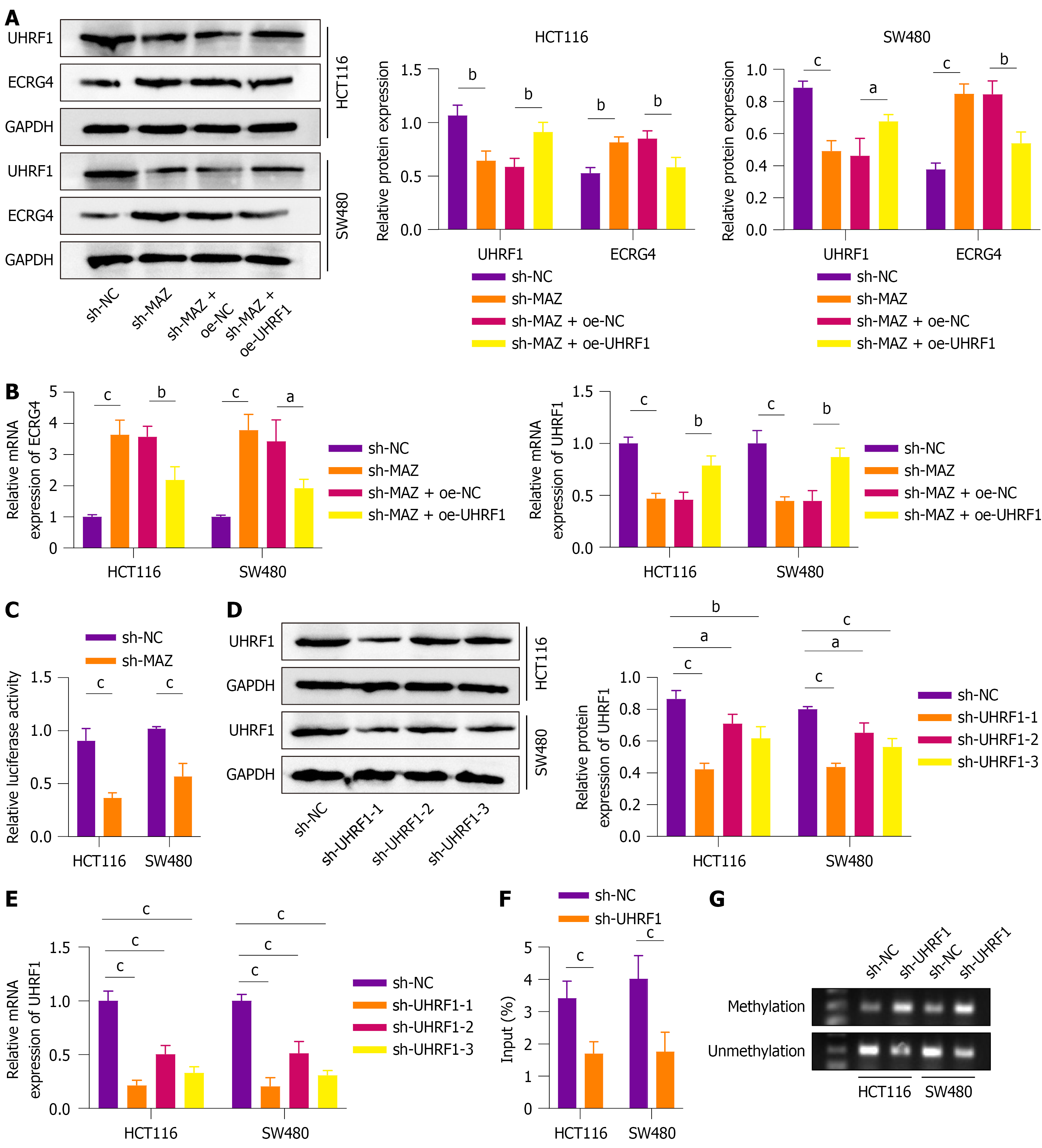
We investigated the downstream mechanisms by which MAZ promotes CRC metastasis. Western blot (Figure 3A) and qRT-PCR (Figure 3B) showed that UHRF1 expression was significantly reduced after knockdown of MAZ, while ECRG4 expression was increased, and ECRG4 expression was decreased after overexpression of UHRF1. MAZ directly bound to the UHRF1 promoter region to activate its transcription, which was confirmed by dual-luciferase reporter assays (Figure 3C). We established UHRF1-knockdown CRC cell lines (HCT116 and SW480) by sh-RNA. Western blot and qRT-PCR (Figure 3D and E) confirmed that all three sh-RNAs significantly downregulated UHRF1 expression, with sh-UHRF1-1 showing the most significant impact. As a result, sh-UHRF1-1 was chosen for further experiments. Moreover, chromatin immunoprecipitation-quantitative PCR assay (Figure 3F) demonstrated decreased enrichment of UHRF1 on the ECRG4 promoter after knockdown of UHRF1. Similarly, methylation-specific PCR results (Figure 3G) showed reduced methylation and increased unmethylation of the ECRG4 promoter after knockdown of UHRF1. These results indicated that in CRC cell lines, MAZ binds to the UHRF1 promoter and activates its transcription. Subsequently, UHRF1 promotes DNA methylation of ECRG4, leading to the downregulation of the tumor suppressor gene ECRG4.
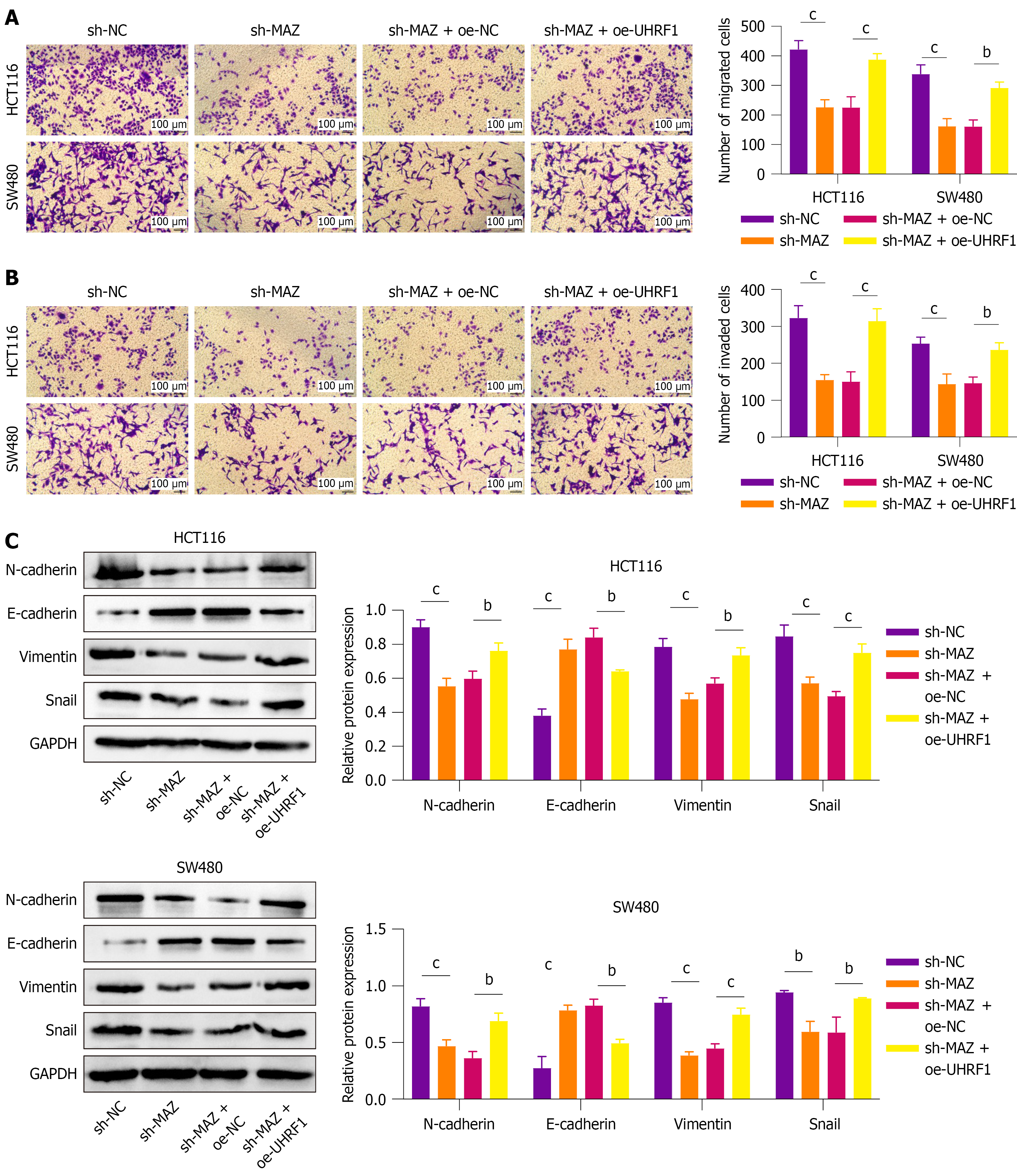
Further functional experiments were conducted to validate the impact of MAZ transcriptional activation of UHRF1 on CRC progression. Silencing MAZ greatly inhibited the migration and invasion of CRC cells; however, UHRF1 overexpression partially restored these effects (Figure 4A and B, P < 0.05). Similarly, UHRF1 overexpression also reversed the effect of sh-MAZ on EMT in CRC cells, with decreased E-cadherin expression and increased expression of N-cadherin, vimentin, and snail (Figure 4C). Our results indicated that in CRC cell lines, MAZ promotes cell migration, invasion, and EMT by transcriptionally activating UHRF1.
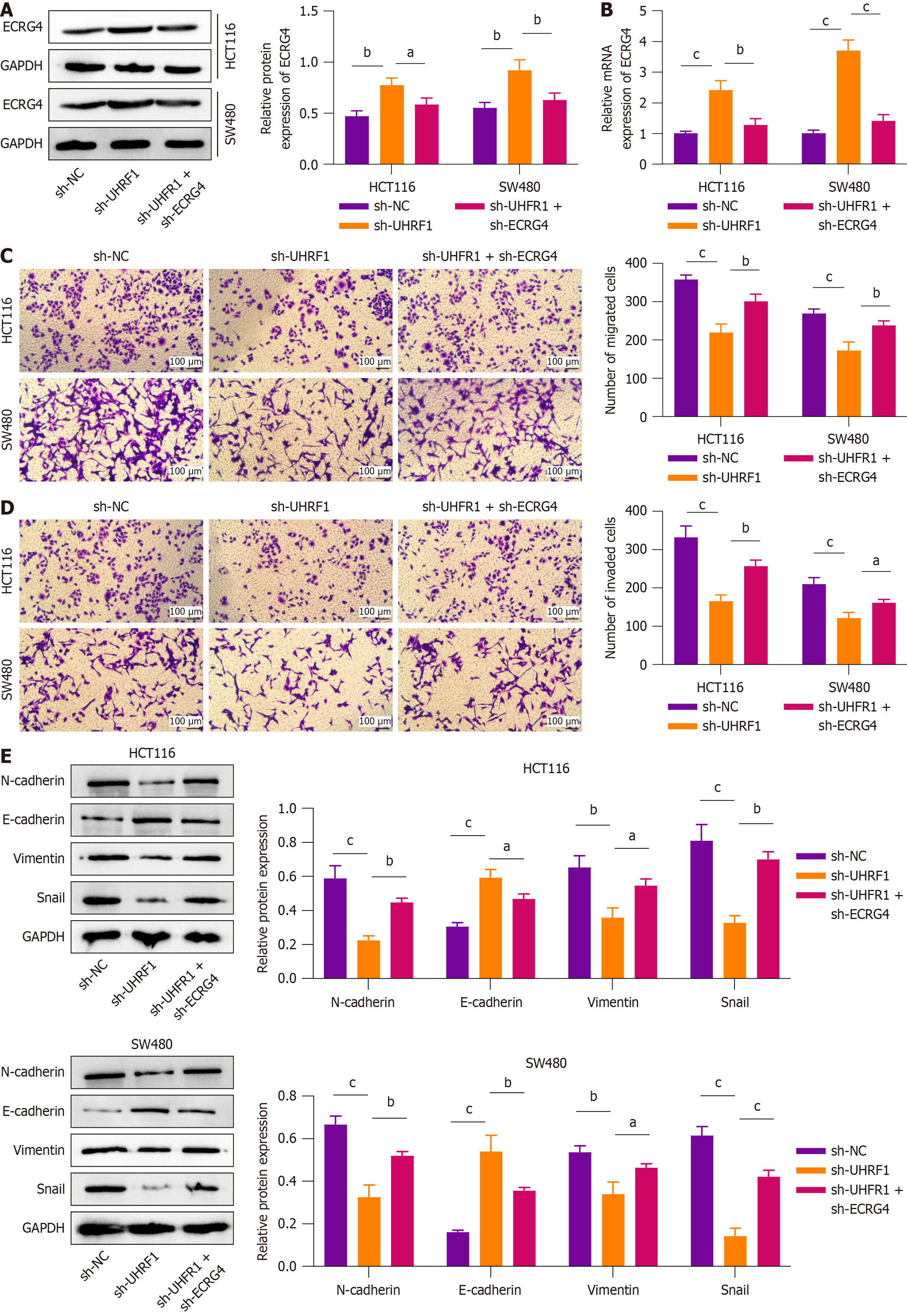
To further explore the impact of UHRF1 in regulating ECRG4 in promoting cell migration, invasion, and EMT in CRC cells, we constructed UHRF1-deficient and ECRG4-deficient CRC cells. Western blot and qRT-PCR (Figure 5A and B) showed that silencing UHRF1 significantly increased ECRG4 expression in CRC cells. Silencing UHRF1 inhibited CRC cell migration and invasion; however, simultaneous knockdown of ECRG4 partially restored these phenotypes (Figure 5C and D, P < 0.05). Western blot (Figure 5E) showed that UHRF1 knockdown increased E-cadherin expression and reduced N-cadherin, vimentin, and snail levels. Simultaneous silencing ECRG4 partially reversed these changes. These results showed that UHRF1 regulates CRC migration, invasion, and EMT by downregulating ECRG4 through DNA methylation in CRC cell lines.
Our study found that MAZ was a crucial transcription factor involved in CRC metastasis. MAZ was upregulated in CRC cells and promoted migration, invasion, and EMT of CRC cells. Our findings provided new insights into the molecular mechanisms underlying CRC progression and suggest the MAZ/UHRF1/ECRG4 axis as a potential therapeutic target. As a transcription factor, MAZ has specific domains for DNA binding and protein-protein interactions[18]. In previous studies, it was found that high expression of MAZ was positively correlated with poor overall survival and bone-metastasis-free survival in prostate cancer patients without bone metastasis[19]. In thyroid cancer patients, those with high MAZ expression have shorter overall survival and disease-free survival[20]. MAZ expression is increased in hepatocellular carcinoma and is associated with distant metastasis[21]. Studies have demonstrated that MAZ drives prostate cancer bone metastasis by upregulating the Kirsten rat sarcoma 2 viral oncogene homolog (KRAS)-dependent Ral guanine nucleotide exchange factors pathway transcriptionally[22]. In addition, MAZ is overexpressed in thyroid cancer and transcriptionally activates TANK binding kinase 1. TANK binding kinase 1 can activate the phosphatidylinositol-3-kinase/protein kinase B/mammalian target of rapamycin signaling pathway, promoting the viability, proliferation, migration, and invasion of thyroid cancer cells[23]. MAZ binds to the promoter of Mitogen-activated protein kinase 2 (MAP2K2), increasing MAP2K2 transcription and thus upregulating its expression. MAP2K2 activates the extracellular signal-regulated kinase signaling pathway, promoting the progression of clear cell renal cell carcinoma. In recent years, studies have found that MAZ is expected to be a potential target for cancer treatment[24]. Our results demonstrated that MAZ expression is significantly upregulated in CRC cells. EMT is a complex and dynamic process, typically marked by the absence of cell polarity, reduced cell-cell adhesion, and enhanced cell migration and invasion capabilities[25,26]. In CRC, we observed that MAZ knockdown effectively reduced cell migration, invasion, and EMT, suggesting its critical role in metastasis.
UHRF1 is a well-known epigenetic regulator[27,28]. UHRF1 knockout selectively impairs growth and induces apoptosis in KRAS-mutant lung cancer cells[29]. Additionally, in lung cancer patients, high UHRF1 expression is nega
ECRG4, a newly identified tumor suppressor gene, has been reported to exhibit reduced expression in CRC tissues due to promoter hypermethylation[16,33]. Research findings indicate that ECRG4, as a tumor suppressor, promotes the expression of osteoglycin by upregulating nuclear factor 1 C-type. Nuclear factor 1 C-type can bind to the promoter region of osteoglycin and regulate its transcription[34]. ECRG4 likely inhibits the malignant phenotype of tongue cancer cells by suppressing the BC200 long non-coding RNA/matrix metalloproteinases signaling pathway[35]. Additionally, ECRG4 has been identified as a novel thrombin-processed monocyte chemoattractant that recruits myeloid cells, promotes their activation, and leads to the blockade of tumor progression[36]. Our study confirmed that ECRG4 exp
Although our study identified a novel regulatory axis in the mechanism of CRC metastasis, it had some limitations. The animal model lacked a functional immune system, which cannot fully recapitulate the role of the tumor microenvironment in MAZ-mediated metastasis. Additionally, we did not correlate the expression levels of MAZ/UHRF1 and the methylation status of ECRG4 with clinicopathological characteristics. In future research, we will attempt to use organoid models to explore the role of MAZ in CRC progression and combine clinical data to further investigate the potential targeting mechanism of MAZ in clinical practice. Additionally, in vivo studies with patient-derived xenograft models would enhance the translational relevance of our findings. Investigating potential small molecule inhibitors targeting MAZ or UHRF1 could also pave the way for novel therapeutic interventions.
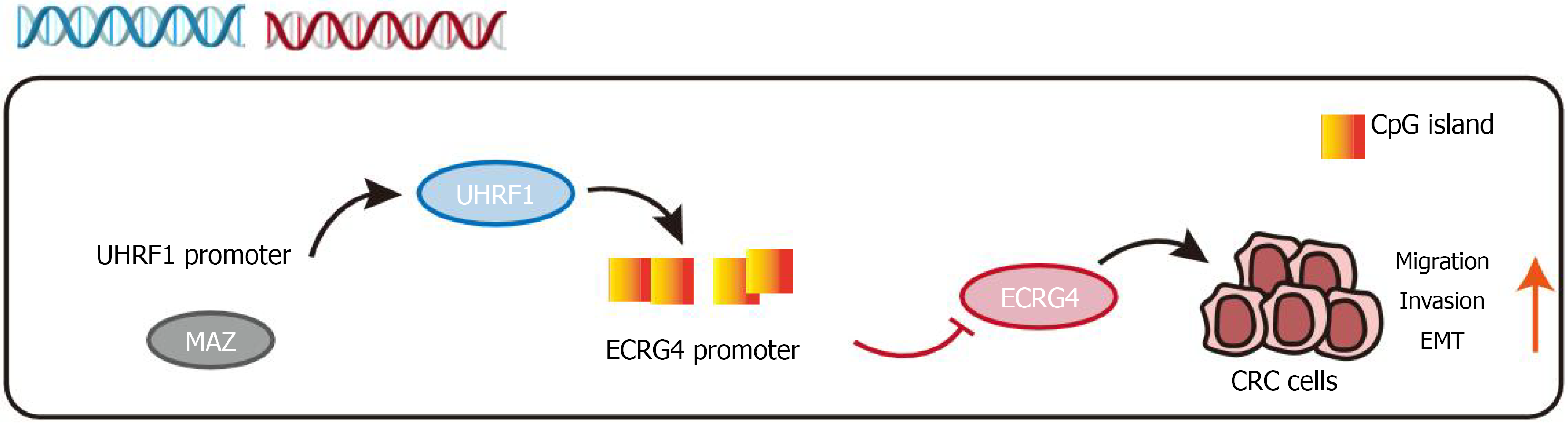
Our findings underscore the pivotal role of MAZ in CRC progression and its downstream regulatory mechanisms. We demonstrated that MAZ promoted CRC migration, invasion, and EMT while transcriptionally activating UHRF1. In turn, UHRF1 epigenetically silenced ECRG4 through DNA methylation, facilitating tumor progression (Figure 6). Targeting the MAZ/UHRF1/ECRG4 signaling axis could provide a novel therapeutic approach for CRC. Inhibiting MAZ, UHRF1, or restoring ECRG4 expression may effectively reactivate tumor suppressor pathways, thereby mitigating tumor metastasis.
Our study identified a novel regulatory axis involving MAZ, UHRF1, and ECRG4 in the progression of CRC, where MAZ transcriptionally activated UHRF1, leading to the epigenetic silencing of the tumor suppressor gene ECRG4 via DNA methylation.
| 1. | Baidoun F, Elshiwy K, Elkeraie Y, Merjaneh Z, Khoudari G, Sarmini MT, Gad M, Al-Husseini M, Saad A. Colorectal Cancer Epidemiology: Recent Trends and Impact on Outcomes. Curr Drug Targets. 2021;22:998-1009. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 240] [Cited by in RCA: 208] [Article Influence: 41.6] [Reference Citation Analysis (2)] |
| 2. | Zhang C, Zheng Z, Wang H, Qi Z, Wang Y, Gao Z, Huang Y, Jin S. Silencing PCCA Suppresses CRC Growth and Spread by Modulating EMT and M1 Macrophage Polarization. Int J Med Sci. 2025;22:87-100. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 3. | Xiao T, Li X, Felsenfeld G. The Myc-associated zinc finger protein epigenetically controls expression of interferon-γ-stimulated genes by recruiting STAT1 to chromatin. Proc Natl Acad Sci U S A. 2024;121:e2320938121. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1] [Cited by in RCA: 12] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
| 4. | Xiao T, Li X, Felsenfeld G. The Myc-associated zinc finger protein (MAZ) works together with CTCF to control cohesin positioning and genome organization. Proc Natl Acad Sci U S A. 2021;118:e2023127118. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 60] [Article Influence: 12.0] [Reference Citation Analysis (0)] |
| 5. | Andersen L, Gülich AF, Alteneder M, Preglej T, Orola MJ, Dhele N, Stolz V, Schebesta A, Hamminger P, Hladik A, Floess S, Krausgruber T, Faux T, Andrabi SBA, Huehn J, Knapp S, Sparwasser T, Bock C, Laiho A, Elo LL, Rasool O, Lahesmaa R, Sakaguchi S, Ellmeier W. The Transcription Factor MAZR/PATZ1 Regulates the Development of FOXP3(+) Regulatory T Cells. Cell Rep. 2019;29:4447-4459.e6. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 19] [Article Influence: 3.2] [Reference Citation Analysis (0)] |
| 6. | Deen D, Butter F, Daniels DE, et al. Identification of the transcription factor MAZ as a regulator of erythropoiesis. Blood Adv. 2021;5(15):3002-3015. Blood Adv. 2022;6:1854. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 7. | Zeng J, Zhang L, Huang L, Yu X, Han L, Zheng Y, Wang T, Zhang N, Yang M. MAZ promotes thyroid cancer progression by driving transcriptional reprogram and enhancing ERK1/2 activation. Cancer Lett. 2024;602:217201. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 20] [Reference Citation Analysis (0)] |
| 8. | Ni G, Sun Y, Jia H, Xiahou Z, Li Y, Zhao F, Zang H. MAZ-mediated tumor progression and immune evasion in hormone receptor-positive breast cancer: Targeting tumor microenvironment and PCLAF+ subtype-specific therapy. Transl Oncol. 2025;52:102280. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 19] [Reference Citation Analysis (0)] |
| 9. | Triner D, Castillo C, Hakim JB, Xue X, Greenson JK, Nuñez G, Chen GY, Colacino JA, Shah YM. Myc-Associated Zinc Finger Protein Regulates the Proinflammatory Response in Colitis and Colon Cancer via STAT3 Signaling. Mol Cell Biol. 2018;38:e00386-e00318. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 48] [Cited by in RCA: 45] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 10. | Oyon D, Lopez-Pascual A, Castello-Uribe B, Uriarte I, Orsi G, Llorente S, Elurbide J, Adan-Villaescusa E, Valbuena-Goiricelaya E, Irigaray-Miramon A, Latasa MU, Martinez-Perez LA, Bonetti LR, Prosper F, Ponz-Sarvise M, Vicent S, Pineda-Lucena A, Ruiz-Clavijo D, Sangro B, Larracoechea UA, Tian TV, Casadei-Gardini A, Amat I, Arechederra M, Berasain C, Urman JM, Avila MA, Fernandez-Barrena MG. Targeting of the G9a, DNMT1 and UHRF1 epigenetic complex as an effective strategy against pancreatic ductal adenocarcinoma. J Exp Clin Cancer Res. 2025;44:13. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 13] [Reference Citation Analysis (0)] |
| 11. | Lin Y, Chen Z, Zheng Y, Liu Y, Gao J, Lin S, Chen S. MiR-506 Targets UHRF1 to Inhibit Colorectal Cancer Proliferation and Invasion via the KISS1/PI3K/NF-κB Signaling Axis. Front Cell Dev Biol. 2019;7:266. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 18] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 12. | Zhu M, Xu Y, Ge M, Gui Z, Yan F. Regulation of UHRF1 by microRNA-9 modulates colorectal cancer cell proliferation and apoptosis. Cancer Sci. 2015;106:833-839. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 43] [Cited by in RCA: 58] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 13. | Kong X, Chen J, Xie W, Brown SM, Cai Y, Wu K, Fan D, Nie Y, Yegnasubramanian S, Tiedemann RL, Tao Y, Chiu Yen RW, Topper MJ, Zahnow CA, Easwaran H, Rothbart SB, Xia L, Baylin SB. Defining UHRF1 Domains that Support Maintenance of Human Colon Cancer DNA Methylation and Oncogenic Properties. Cancer Cell. 2019;35:633-648.e7. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 125] [Article Influence: 17.9] [Reference Citation Analysis (0)] |
| 14. | Cai Z, Liang P, Xuan J, Wan J, Guo H. ECRG4 as a novel tumor suppressor gene inhibits colorectal cancer cell growth in vitro and in vivo. Tumour Biol. 2016;37:9111-9120. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 15] [Cited by in RCA: 21] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 15. | Li X, Wang P, Hao Q, Cao Z, Zhang H, Guo J, Hu S, Bai F. Esophageal cancer-related gene 4 and solid tumors: a brief literature review. J Physiol Pharmacol. 2022;73. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 16. | Wang J, Zhao X, Jin L, Wu G, Yang Y. UBR5 Contributes to Colorectal Cancer Progression by Destabilizing the Tumor Suppressor ECRG4. Dig Dis Sci. 2017;62:2781-2789. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 33] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 17. | Zhang ZH, Yuan CY, Xu M, Wang MF, Feng T, Wang Y, Zheng SF, Zhang HL, Shi GH, Cao DL, Wang ZL, Ye DW. Calycosin inhibits the proliferation and metastasis of renal cell carcinoma through the MAZ/HAS2 signaling pathway. Phytother Res. 2024;38:4774-4791. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 7] [Reference Citation Analysis (0)] |
| 18. | Chen Y, Zhu X, Wang J, Hu J, Zhang J, Zhang X, Han L, Yu H, Hu H, Fei K, Zhang P, Zhang L. MAZ promotes tumor proliferation and immune evasion in lung adenocarcinoma. Oncogene. 2024;43:3619-3632. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 10] [Cited by in RCA: 10] [Article Influence: 5.0] [Reference Citation Analysis (2)] |
| 19. | Zheng C, Wu H, Jin S, Li D, Tan S, Zhu X. Roles of Myc-associated zinc finger protein in malignant tumors. Asia Pac J Clin Oncol. 2022;18:506-514. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 26] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 20. | Zheng C, Wu H, Zhang S, Tian R, Long H, Zhang H, Guo X, Li D, Tan S, Zhu X. Myc-Associated Zinc Finger Protein Promotes Metastasis of Papillary Thyroid Cancer. Front Biosci (Landmark Ed). 2023;28:162. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 10] [Reference Citation Analysis (0)] |
| 21. | Luo W, Zhu X, Liu W, Ren Y, Bei C, Qin L, Miao X, Tang F, Tang G, Tan S. MYC associated zinc finger protein promotes the invasion and metastasis of hepatocellular carcinoma by inducing epithelial mesenchymal transition. Oncotarget. 2016;7:86420-86432. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 21] [Cited by in RCA: 45] [Article Influence: 5.6] [Reference Citation Analysis (0)] |
| 22. | Yang Q, Lang C, Wu Z, Dai Y, He S, Guo W, Huang S, Du H, Ren D, Peng X. MAZ promotes prostate cancer bone metastasis through transcriptionally activating the KRas-dependent RalGEFs pathway. J Exp Clin Cancer Res. 2019;38:391. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 59] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 23. | Jiang Q, Guan Y, Zheng J, Lu H. TBK1 promotes thyroid cancer progress by activating the PI3K/Akt/mTOR signaling pathway. Immun Inflamm Dis. 2023;11:e796. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 11] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 24. | Ren LX, Qi JC, Zhao AN, Shi B, Zhang H, Wang DD, Yang Z. Myc-associated zinc-finger protein promotes clear cell renal cell carcinoma progression through transcriptional activation of the MAP2K2-dependent ERK pathway. Cancer Cell Int. 2021;21:323. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4] [Cited by in RCA: 35] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 25. | Shi G, Holgersson K, Xin Z, Szekely L, Du Q, Jing X. TRIB1 facilitates the proliferation and migration of ovarian cancer cells by inducing EMT progression. Histol Histopathol. 2025;40:1447-1455. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 26. | Glaviano A, Lau HS, Carter LM, Lee EHC, Lam HY, Okina E, Tan DJJ, Tan W, Ang HL, Carbone D, Yee MY, Shanmugam MK, Huang XZ, Sethi G, Tan TZ, Lim LHK, Huang RY, Ungefroren H, Giovannetti E, Tang DG, Bruno TC, Luo P, Andersen MH, Qian BZ, Ishihara J, Radisky DC, Elias S, Yadav S, Kim M, Robert C, Diana P, Schalper KA, Shi T, Merghoub T, Krebs S, Kusumbe AP, Davids MS, Brown JR, Kumar AP. Harnessing the tumor microenvironment: targeted cancer therapies through modulation of epithelial-mesenchymal transition. J Hematol Oncol. 2025;18:6. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 38] [Cited by in RCA: 88] [Article Influence: 88.0] [Reference Citation Analysis (1)] |
| 27. | Wang X, Zhang P, Yan J, Huang J, Shen Y, He H, Dou H. SIRT6 deficiency impairs the deacetylation and ubiquitination of UHRF1 to strengthen glycolysis and lactate secretion in bladder cancer. Cell Biosci. 2024;14:153. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 7] [Reference Citation Analysis (0)] |
| 28. | Bai W, Xu J, Gu W, Wang D, Cui Y, Rong W, Du X, Li X, Xia C, Gan Q, He G, Guo H, Deng J, Wu Y, Yen RC, Yegnasubramanian S, Rothbart SB, Luo C, Wu L, Liu J, Baylin SB, Kong X. Defining ortholog-specific UHRF1 inhibition by STELLA for cancer therapy. Nat Commun. 2025;16:474. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 9] [Article Influence: 9.0] [Reference Citation Analysis (0)] |
| 29. | Kostyrko K, Román M, Lee AG, Simpson DR, Dinh PT, Leung SG, Marini KD, Kelly MR, Broyde J, Califano A, Jackson PK, Sweet-Cordero EA. UHRF1 is a mediator of KRAS driven oncogenesis in lung adenocarcinoma. Nat Commun. 2023;14:3966. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 8] [Cited by in RCA: 24] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 30. | Hou J, Li W, Zhang S, Tan D, Lv K, Zhu Y, Hou Y, Guo H, Jiang L. UHRF1 plays an oncogenic role in small cell lung cancer. Mol Carcinog. 2023;62:385-397. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 9] [Reference Citation Analysis (0)] |
| 31. | Jiao D, Huan Y, Zheng J, Wei M, Zheng G, Han D, Wu J, Xi W, Wei F, Yang AG, Qin W, Wang H, Wen W. UHRF1 promotes renal cell carcinoma progression through epigenetic regulation of TXNIP. Oncogene. 2019;38:5686-5699. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 46] [Article Influence: 6.6] [Reference Citation Analysis (0)] |
| 32. | Song Y, Liu H, Xian Q, Gui C, Xu M, Zhou Y. Mechanistic insights into UHRF1mediated DNA methylation by structurebased functional clarification of UHRF1 domains (Review). Oncol Lett. 2023;26:542. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 4] [Reference Citation Analysis (0)] |
| 33. | Li X, Hu S, Ma M, Wang P, Qi Y, Zhou Y, Zhong Z, Gao H, Bai F. Esophageal cancer-related gene 4 inhibits gastric cancer growth and metastasis by upregulating Krüppel-like factor 2 expression. Ann Transl Med. 2023;11:176. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 1] [Reference Citation Analysis (0)] |
| 34. | Liang X, Gao J, Wang Q, Hou S, Wu C. ECRG4 Represses Cell Proliferation and Invasiveness via NFIC/OGN/NF-κB Signaling Pathway in Bladder Cancer. Front Genet. 2020;11:846. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 30] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 35. | Huang W, Zhou R, Mao L, Deng C, Dang X. Esophageal cancer related gene-4 inhibits the migration and proliferation of oral squamous cell carcinoma through BC200 lncRNA/MMP-9 and -13 signaling pathway. Cell Signal. 2019;62:109327. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 16] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 36. | Lee J, Dang X, Borboa A, Coimbra R, Baird A, Eliceiri BP. Thrombin-processed Ecrg4 recruits myeloid cells and induces antitumorigenic inflammation. Neuro Oncol. 2015;17:685-696. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 34] [Article Influence: 2.8] [Reference Citation Analysis (0)] |
Open Access: This article is an open-access article that was selected by an in-house editor and fully peer-reviewed by external reviewers. It is distributed in accordance with the Creative Commons Attribution NonCommercial (CC BY-NC 4.0) license, which permits others to distribute, remix, adapt, build upon this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See: https://creativecommons.org/Licenses/by-nc/4.0/