Published online Mar 16, 2026. doi: 10.4253/wjge.v18.i3.116381
Revised: December 10, 2025
Accepted: January 20, 2026
Published online: March 16, 2026
Processing time: 123 Days and 4.9 Hours
Colorectal cancer remains a major global health burden. Accurate real-time char
To determine the diagnostic accuracy of AI in predicting colorectal polyp histo
A comprehensive literature search was conducted in accordance with Preferred Reporting Items for Systematic reviews and Meta-Analyses guidelines and pro
The meta-analysis incorporated 3245 patients encompassing 4752 polyps. Pooled analysis demonstrated that AI achieved an overall diagnostic accuracy of 93%, compared to 82% for human experts (relative risk = 1.13; 95% confidence interval: 1.07-1.20; P < 0.0001). AI consistently outperformed human endoscopists, particularly in cohorts involving less experienced operators or suboptimal imaging conditions. Substantial heterogeneity was observed (I2 = 74.3%), attributed to methodological differences in imaging modalities, AI architectures, and operator proficiency.
AI demonstrates high diagnostic accuracy for real-time colorectal polyp histology and may enhance clinical decision-making, although expert oversight remains essential for atypical or high-risk lesions.
Core Tip: Artificial intelligence enables accurate, real-time differentiation between neoplastic and non-neoplastic colorectal polyps, fulfilling the optical biopsy criteria. By supporting “resect-and-discard” strategies for diminutive lesions and providing decision assistance to less experienced endoscopists, artificial intelligence can streamline colonoscopy workflows, mitigate pathology workloads, and enhance colorectal cancer prevention programs.
- Citation: Curlej P, Soldera J. Artificial intelligence in predicting colorectal polyp histology: Systematic review and meta-analysis of diagnostic accuracy in real-time procedures. World J Gastrointest Endosc 2026; 18(3): 116381
- URL: https://www.wjgnet.com/1948-5190/full/v18/i3/116381.htm
- DOI: https://dx.doi.org/10.4253/wjge.v18.i3.116381
Colorectal cancer (CRC) remains a major global health challenge[1]. It is the third most common cancer worldwide and a leading cause of cancer-related mortality, with approximately 1.93 million new cases and 935000 deaths each year[2,3]. Incidence rates vary widely across regions developed countries have historically high CRC prevalence, while developing regions are experiencing rising incidence reflecting the interplay of genetic predispositions, lifestyle and dietary factors, and healthcare disparities[4]. This growing burden underscores an urgent need for effective prevention strategies and early diagnostic interventions[3].
One cornerstone of CRC prevention is colonoscopy, the gold-standard screening procedure that allows both detection and removal of premalignant polyps, thereby interrupting the adenoma-carcinoma sequence[5]. A critical goal during colonoscopy is to accurately distinguish neoplastic polyps (adenomas and sessile serrated lesions) from non-neoplastic lesions (such as hyperplastic polyps) in order to guide appropriate management and surveillance intervals[6]. Definitive classification, however, traditionally relies on histopathological examination of excised polyps; a process that is invasive, resource-intensive, costly, and slow. Waiting days for pathology results can cause patient anxiety and impedes immediate clinical decision-making, highlighting the drawbacks of our current “resect first, diagnose later” approach[7].
Diagnostic accuracy in polyp assessment also varies notably among endoscopists and pathologists, leading to in
At the same time, healthcare systems are under increasing pressure from aging populations and constrained resources, emphasizing the importance of more efficient approaches to CRC screening and surveillance[11]. An ideal solution would be an accurate, real-time diagnostic tool that can reliably characterize polyp histology in vivo during the endoscopic procedure. Such a tool could streamline decision-making by enabling immediate determination of whether a polyp is benign or precancerous, thereby reducing unnecessary removals, biopsies, and associated costs and risks[12]. In this context, artificial intelligence (AI) has garnered significant attention as a potential game-changer for meeting these clinical needs.
In recent years, rapid advancements in AI, particularly in machine learning and deep learning have yielded sophisticated prognostication models and image analysis tools capable of interpreting medical images with high speed and accuracy[13-17]. In gastroenterology, deep convolutional neural networks have been developed to automatically analyses endoscopic images, allowing precise real-time identification and characterization of colorectal polyps[18,19]. Modern AI-assisted colonoscopy systems can also enhance visualization using advanced imaging modalities such as narrow-band imaging (NBI), blue laser imaging, and ultra-magnification endocytoscopy, which provide greater mucosal detail than conventional white-light endoscopy and improve diagnostic accuracy for differentiating polyp types[20,21]. Preliminary clinical studies have been promising: For example, AI-based endoscopic analysis has successfully distinguished neop
Building on these advances, a growing body of clinical evidence indicates that AI can enhance polyp detection and characterization during colonoscopy. The COACH trial by Renner et al[24], for instance, reported that an AI system outperformed expert endoscopists in classifying colorectal neoplasms, achieving sensitivity rates above 90%. Similarly, Wang et al[25] showed that real-time AI feedback significantly reduced adenoma miss-rates in a tandem colonoscopy study, and Kudo et al[26] demonstrated that an AI-assisted NBI platform could diagnose polyp histology with higher accuracy than experienced endoscopists. These improvements have been consistently observed in studies across various clinical settings, including multicenter trials in different patient populations[24,27,28], suggesting that AI’s benefits are generalizable and not limited to single-center expertise. Professional guidelines are beginning to reflect this progress: The British Society of Gastroenterology (BSG) now encourages the adoption of new technologies such as AI to help endo
Despite its promise, AI integration into colonoscopy is not without challenges. Many AI models are trained on specific image datasets under controlled conditions, and differences in algorithm design or training data can lead to variability in performance when these systems are applied in new settings[21,29]. This raises concerns about generalizability and reproducibility: An AI tool that performs well in one hospital or patient group may not instantly translate to all popu
Several knowledge gaps in the literature remain to be addressed. Most studies to date have been relatively small or single-center, and they often evaluate AI based on per-polyp diagnostic accuracy rather than patient-level outcomes. It therefore remains unclear how the routine use of AI in colonoscopy might affect long-term patient outcomes, such as reducing interval CRC rates or improving overall surveillance efficiency[30]. Additionally, comparative studies between AI systems and human experts have yielded mixed results, with AI showing greater benefit in some contexts than others; notably, the incremental advantage of AI may depend on the baseline skill and adenoma detection rate of endoscopists in a given setting[24,29]. A lack of standardization in study methodologies and reporting further complicates the picture researchers have used different definitions, performance metrics, and validation protocols, making it difficult to directly compare results across studies or to formulate uniform guidelines for AI implementation[31]. These gaps highlight the need for a comprehensive synthesis of the available evidence to determine where AI truly stands in augmenting colore
The aim of this study was to systematically review and meta-analyses prospective evidence on real-time AI-assisted diagnosis of colorectal polyps, comparing its histologic accuracy with expert endoscopists. The review sought to det
This systematic review and meta-analysis were designed to evaluate the diagnostic accuracy of AI systems in predicting colorectal polyp histology during real-time endoscopic assessment. The study was conducted in accordance with the Preferred Reporting Items for Systematic reviews and Meta-Analyses[32] statement and was prospectively registered in the International Prospective Register of Systematic Reviews (No. CRD420251012404). The review was undertaken to address a key evidence gap in endoscopic imaging, evaluating whether AI can improve the accuracy of optical diagnosis, reduce reliance on histopathology, and enhance clinical decision-making and workflow efficiency in CRC prevention.
The primary objective was to determine the rate of correct histological diagnoses made by AI-assisted colonoscopy systems compared to conventional histopathology and human expert endoscopists. A structured research question was formulated using the PICO (Population, Intervention, Comparison, Outcome framework; Table 1)[33]. Additional criteria: (1) Peer-reviewed journal articles published in English; and (2) Studies published within the last 10 years (2014-2024).
| Component | Definition | Criteria for inclusion | Criteria for exclusion |
| Research question | What is the diagnostic accuracy and clinical impact of AI-based real-time histology prediction for colorectal polyps compared to conventional histopathology and endoscopists? | Studies must specifically investigate AI assisted systems for real time polyp histology prediction | Studies that do not focus on real-time AI applications in live colonoscopy settings |
| Population | Patients undergoing colonoscopy with colorectal polyp detection and histology prediction | (1) Adults (≥ 18 years) undergoing colonoscopy; (2) Studies involving patients with colorectal polyps (adenomatous, hyperplastic, sessile serrated); and (3) Human subjects (no in vitro or animal studies) | (1) Studies focusing on animal models, in vitro, or simulation-based research; and (2) Pediatric studies (patients < 18 years) |
| Intervention | AI-based systems for real-time histology prediction of colorectal polyps | (1) AI assisted colonoscopy systems for polyp detection and classification; (2) Machine learning and deep learning models (e.g., convolutional neural networks); and (3) AI enhanced imaging techniques (e.g., narrow-band imaging, endocytoscopy) | AI models used only for retrospective analysis (not real-time) |
| Comparison | Standard histopathological methods or expert endoscopists’ assessments | (1) Histopathological examination as the gold standard; (2) Comparison with experienced endoscopists’ accuracy; and (3) Conventional endoscopy methods without AI assistance | AI models compared only with other AI models (without human or histological reference) |
| Outcome | Diagnostic accuracy of AI systems in polyp histology prediction | Primary outcomes: (1) Sensitivity, specificity, accuracy, and negative predictive value of AI models; and (2) Adenoma detection rate. Secondary outcomes: (1) Reduction in unnecessary polypectomies; (2) Interobserver variability between AI and human experts; and (3) Time efficiency and cost-effectiveness of AI-assisted endoscopy | (1) Studies with incomplete or insufficient clinical validation of AI performance; and (2) Studies that do not report key diagnostic accuracy metrics (e.g., missing sensitivity, specificity, or adenoma detection rate) |
A comprehensive and systematic search strategy was developed using Medical Subject Headings (MeSH) and Boolean operators to identify all relevant studies evaluating AI systems for real-time colorectal polyp histology prediction. The following structured search query was executed in PubMed: [“Artificial Intelligence”(Mesh) OR “Machine Lear
Data extraction was systematically conducted using a standardized extraction form designed to capture essential study information, including author details, year of publication, country, study design, and clinical setting. Patient characteristics such as the number enrolled, demographics, and polyp attributes were documented alongside detailed descriptions of the AI systems employed, specifying the model type, algorithm design, and real-time deployment modalities (standard white-light colonoscopy, NBI, or endocytoscopy). Information regarding comparison groups, including expert endo
Quality assessment and risk of bias evaluation were conducted using the Quality Assessment of Prognostic Accuracy Studies (QUADAS) 2 tool, assessing four domains: Patient selection, index test conduct, reference standard, and patient flow and timing. This comprehensive quality appraisal ensures robustness in interpreting and generalizing the review findings.
Meta-analysis was performed to quantitatively compare the diagnostic accuracy defined as the rate of correct histological diagnoses of AI systems relative to expert human endoscopists during real-time colorectal polyp assessment. The primary aim was to evaluate the comparative effectiveness of AI across multiple clinical studies included in the systematic review. Data extracted from each eligible study included the number of correctly diagnosed lesions and the total number of lesions assessed by both the AI systems and human comparator groups. Using these data, proportions of correct dia
To directly compare AI performance with human expert performance, relative risk (RR) was selected as the primary effect measure. The RR represents the ratio of correct diagnoses by AI compared to those by human endoscopists. An RR greater than 1 indicated that AI provided superior diagnostic accuracy compared to humans, whereas an RR less than 1 would suggest superior human performance. An RR value of exactly 1 represented equivalent diagnostic accuracy between the two groups. The meta-analysis employed the inverse variance method to pool the RRs across studies. This method assigns weights to individual studies according to the precision of their estimates, reflecting differences in sample size and variability in diagnostic outcomes across studies[34].
Given anticipated methodological and clinical heterogeneity stemming from variations in AI systems, patient populations, lesion characteristics, and clinical setting, a random-effects model was used as the primary analytical approach. This model assumes inherent variability across studies and incorporates between-study variation into the pooled estimates, providing more conservative and clinically realistic results. Additionally, common-effect model was calculated for comparative purposes to assess the consistency and robustness of the pooled effect estimates.
To evaluate heterogeneity among the included studies, two widely accepted statistical methods were employed: (1) Cochran’s Q-test was performed to determine whether the observed variation among study estimates exceeded chance alone. Statistical significance indicating heterogeneity was defined as a P-value less than 0.05; and (2) The I2 statistic quantified the extent of observed heterogeneity, expressed as the proportion of total variability attributable to true differences between studies rather than random sampling error. The interpretation of I2 values followed conventional guidelines: 0%-25% indicating low heterogeneity, 26%-50% moderate heterogeneity, and greater than 50% substantial heterogeneity.
Additionally, τ2 and τ were calculated as measures of between-study variance and standard deviation, respectively, providing further insight into the magnitude and clinical significance of the observed heterogeneity.
To assess the potential impact of publication bias, Deeks’ funnel plot asymmetry test was performed, specifically recommended for diagnostic accuracy meta-analyses[35]. Funnel plots were visually inspected for asymmetry, and the statistical significance of asymmetry was tested using a regression based method, with a P-value less than 0.10 indicating significant publication bias or small-study effects.
Results of the meta-analysis were graphically displayed using forest plots, which depicted individual study-level RR estimates, their corresponding 95% confidence intervals (CI), and the pooled RR estimate from the random-effects model. These plots facilitated easy interpretation and comparison of the diagnostic accuracy of AI systems relative to human experts across the included studies. Additionally, sensitivity analyses were performed by sequentially excluding indi
All analyses were conducted using R statistical software meta package, a widely validated tool specifically designed for meta-analysis[37]. Statistical significance for all analyses was determined at a P-value threshold of less than 0.05, unless otherwise stated.
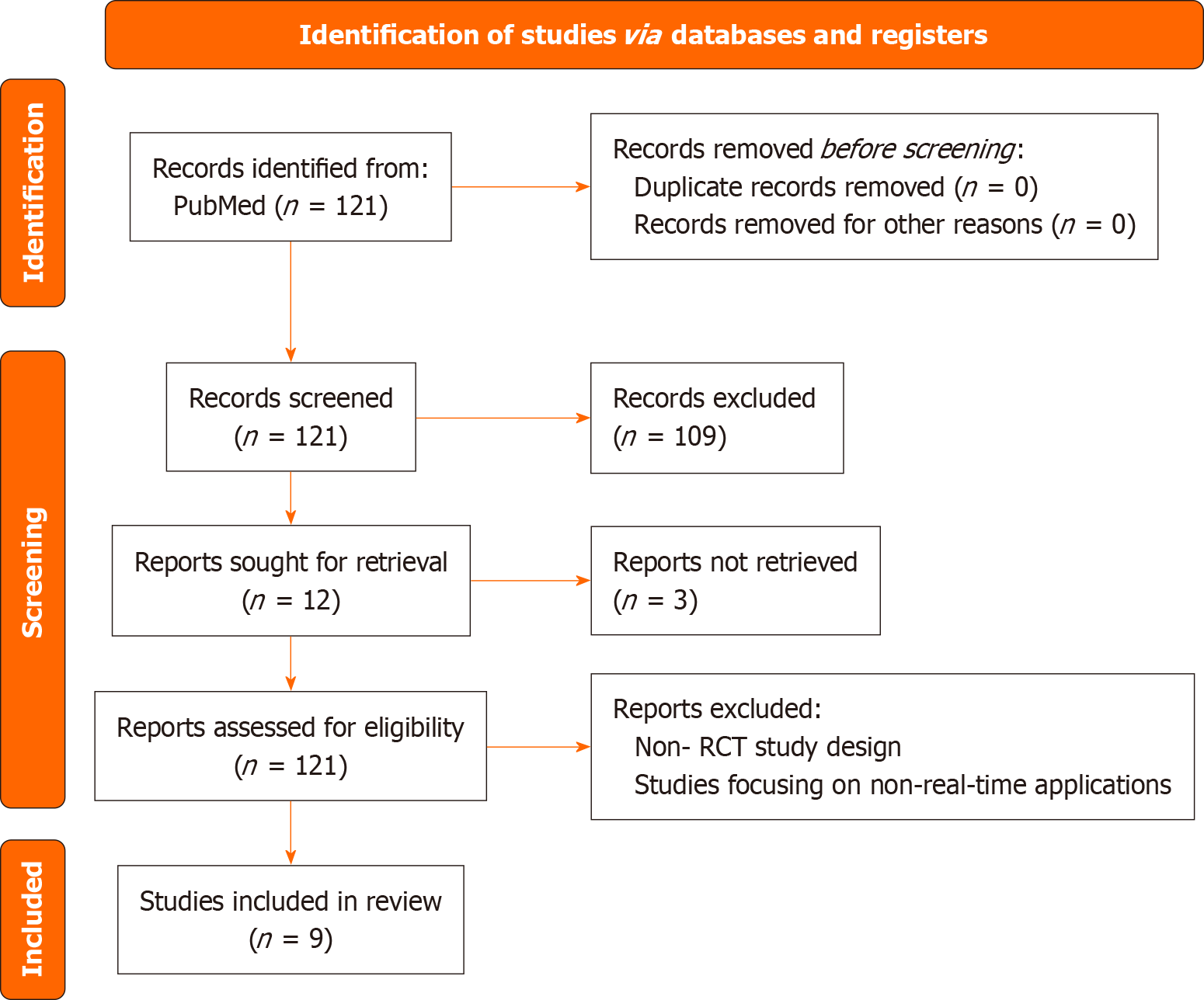
The initial systematic search yielded 121 articles. After applying eligibility criteria (peer-reviewed articles, English language, published within the last 10 years) and removing duplicates, 45 studies remained. These underwent a full-text review, resulting in the exclusion of 33 studies due to criteria including non-real-time AI applications, insufficient diagnostic data, or incompatible study designs. Manual screening of reference lists revealed no additional eligible studies.
Ultimately, 12 studies met eligibility criteria after full-text screening. However, full-text access could not be obtained for 3 studies, despite attempts to contact authors. Consequently, these studies were excluded due to the inability to ens
The nine studies included in this systematic review and meta-analysis collectively evaluated 3245 patients, accounting for a total of 4752 colorectal lesions. These studies encompassed diverse populations from multiple countries, including Germany, Japan, China, the United Kingdom, Canada, the Netherlands, France, and Norway, reflecting a broad geo
Study designs varied significantly, including single-center observational studies[23,24], prospective observational studies[26,38], prospective randomized trials[25,39], and randomized controlled and multicenter studies[27,28,38]. Patient populations across these studies included adults undergoing either screening or diagnostic colonoscopy, contributing to the applicability of the findings across different clinical scenarios.
AI interventions consistently employed advanced deep-learning algorithms, primarily convolutional neural networks, designed for real-time polyp histology classification during endoscopic procedures. Diagnostic modalities included standard high-definition white-light colonoscopy, NBI, and magnification-based technologies such as endocytoscopy. Comparator groups typically comprised experienced human endoscopists. Diagnostic outcomes focused on the number of correctly classified polyps, with clearly documented comparisons between AI systems and human experts. A concise summary of these core characteristics is provided in Table 2.
| Ref. | Country | Study design and setting | Total lesion | Patient population description | AI intervention description | Comparison group | AI correct diagnoses | Human correct diagnoses |
| Barua et al[38] | United Kingdom | Randomized controlled trial | 359 | Screening colonoscopy in general practice | AI-assisted white-light and narrow band imaging | Experienced endoscopists | 325/359 | 317/359 |
| Chino et al[39] | Japan | Prospective observational | 556 | Diagnostic colonoscopy patients | AI system for polyp detection/classification | Human experts | 542/556 | Not reported |
| Djinbachian et al[40] | Canada | Prospective randomized trial | 52 | Screening colonoscopy patients | Autonomous AI system | Human experts | 48/52 | 44/52 (expert 1), 40/52 (expert 2) |
| Kudo et al[26] | Japan | Prospective observational | 194 | Patients with colorectal polyps | AI with magnifying narrow band imaging (EndoBRAIN system) | Human experts | 188/194 | 161/194 |
| Mori et al[23] | Japan | Prospective single-center | 287 | Patients undergoing magnifying endoscopy | AI-assisted endocytoscopy (stained mode) | Human experts | 263/287 | Not reported |
| Renner et al[24] | Germany | Single-center observational | 52 | Adults with colorectal polyps | AI-assisted histology prediction | Two expert readers | 48/52 | 44/52 (expert 1), 40/52 (expert 2) |
| Sato et al[27] | Netherlands | Prospective multicenter | 217 | Patients undergoing colonoscopy | AI using magnifying BLI | Human endoscopists | 214/217 | 177/217 |
| van der Zander et al[28] | Netherlands | Prospective multicenter | 100 | Patients with colorectal polyps | AI system with heatmapping and imaging | Two expert readers | 92/100 | 84/100 (expert 1), 77/100 (expert 2) |
| Wang et al[25] | China | Prospective randomized tandem | 144 | Screening colonoscopy patients | Deep learning CNN (EndoScreener) | Human endoscopists | 124/144 | 96/144 |
This table facilitates an understanding of the heterogeneity and methodological approaches of the included evidence base and serves as a foundation for interpreting the pooled diagnostic accuracy and RR analyses presented later. It highlights key differences in AI systems, clinical settings, and human comparator expertise, which are important considerations when assessing AI’s diagnostic utility in routine clinical practice (Table 2).
The quality assessment of each study was rigorously conducted using the QUADAS-2 tool. Overall, the methodological quality of included studies was high, though minor concerns were identified in certain domains: Mori et al[23] showed low overall risk of bias, with minor limitations related to incomplete reporting of the human comparator’s results. Renner et al[24] demonstrated overall low risk of bias, although minor concerns existed regarding clarity in patient selection methods. Wang et al[25], a prospective randomized study, exhibited low risk of bias, particularly in domains related to index test conduct and patient selection. Kudo et al[26] showed low risk of bias across all assessed domains, with robust methodology clearly outlined. Sato et al[27] exhibited high methodological quality and low risk of bias with clear, well-defined patient selection and diagnostic criteria. van der Zander et al[28] was similarly robust, with slight concerns regarding variability between human experts but low overall risk of bias. Barua et al[38] was methodologically robust, exhibiting minimal bias due to its randomized controlled design and clearly defined inclusion criteria. Chino et al[39] had overall low risk of bias, although minor concerns arose from incomplete reporting of human comparator data. Djinbachian et al[40] was methodologically sound as a randomized controlled trial, though small sample size posed minor concerns about precision.
These minor concerns did not significantly compromise the overall quality or validity of the pooled results. Reco
Diagnostic accuracy, defined as the proportion of correctly classified colorectal lesions by AI systems, was evaluated across the nine included studies. A total of 1961 lesions were analyzed, with AI correctly diagnosing 1844 lesions.
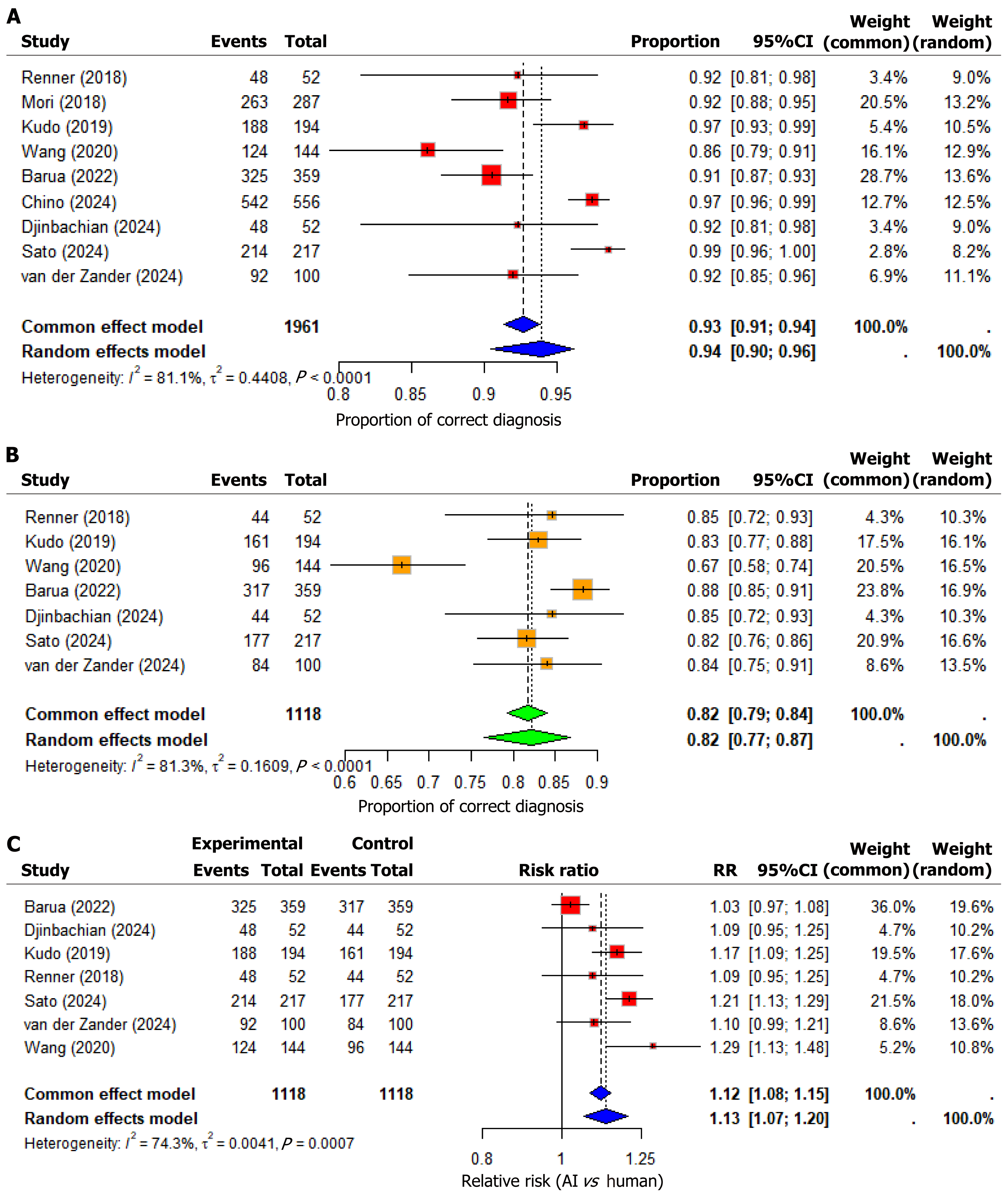
The pooled diagnostic accuracy for AI systems using a random-effects model was 93.97% (95%CI: 90.46%-96.24%), while the common-effect model provided a similar estimate of 92.71% (95%CI: 91.32%-93.89%). This indicates high and consistent performance across diverse clinical settings and imaging modalities (see forest plot for AI accuracy, Figure 2A). However, substantial heterogeneity was observed among these studies, quantified with an I2 value of 81.1% (95%CI: 65.2%-89.8%, Q = 42.42, P < 0.0001), highlighting methodological and clinical variations, such as differences in imaging methods and patient characteristics, influencing diagnostic outcomes.
Human diagnostic accuracy was assessed in seven studies, encompassing a total of 1118 colorectal lesions. Across these, human experts correctly diagnosed 923 lesions. Notably, several studies involved more than one expert reader, revealing inter-observer variability. Renner et al[24], expert 1 correctly classified 44 lesions while expert 2 diagnosed 40 out of 52. A similar pattern appeared in van der Zander et al[28] (84 vs 77 correct) and Djinbachian et al[40] (44 vs 40 correct diag
The meta-analysis revealed that the pooled diagnostic accuracy for human experts was lower than that of AI systems. Using a random-effects model, the estimated correct diagnosis rate for human experts was 82.20% (95%CI: 76.51%-86.75%), while the common-effect model yielded a nearly identical estimate of 81.79% (95%CI: 79.33%-84.01%). These pooled results, though relatively high, were consistently outperformed by AI in most direct comparisons.
Substantial heterogeneity was also noted among human expert assessments, with an I2 value of 81.3% (95%CI: 62.2%-90.7%), a Q statistic of 32.01, and a P < 0.0001, indicating statistically significant inconsistency across studies. This heterogeneity may be attributed to variability in operator experience, classification criteria, image quality, polyp characteristics, and real-time decision-making environments (see forest plot for human rate of correct diagnosis, Figure 2B). Variability in human performance likely relates to differences in expertise levels, training standards, and clinical practice environments.
A direct comparison of diagnostic accuracy between AI and humans was performed through meta-analysis of RRs from seven comparative studies (n = 2236 observations). RR is a particularly effective measure in this context, as it quantifies the ratio of correct diagnoses made by AI systems to those made by human experts. An RR greater than 1 suggests sup
The pooled RR using a random-effects model indicated that AI systems were 13.20% more likely to correctly diagnose colorectal polyps (RR = 1.1320; 95%CI: 1.0659-1.2023; P < 0.0001) compared to human experts. The common-effect model supported this finding, yielding a pooled RR of 1.1153 (95%CI: 1.0821-1.1495; P < 0.0001). These results demonstrate a robust and statistically significant advantage for AI (Figure 2C).
Renner et al[24], van der Zander et al[28], and Djinbachian et al[40]: These studies reported RRs ranging from 1.09 to 1.10, but with CIs that included 1, indicating that the results were not individually statistically significant. Still, their consistent trend in favor of AI contributes to the overall positive pooled effect. Wang et al[25]: RR = 1.2917 (95%CI: 1.1310-1.4751), the highest observed RR, signifying that AI outperformed humans by nearly 29.17% a highly significant result. Kudo et al[26]: RR = 1.1677 (95%CI: 1.0904-1.2505), suggesting that AI was 16.77% more likely to correctly diagnose histology. The CI excludes 1, indicating statistical significance. Sato et al[27]: RR = 1.2090 (95%CI: 1.1327-1.2905), demonstrating an even greater advantage of 20.90%, with a strongly significant CI. Barua et al[38]: RR = 1.0252 (95%CI: 0.9749-1.0782), indicating only a 2.52% improvement by AI. However, the CI includes 1, so the result is not statistically significant, suggesting equi
Substantial heterogeneity was detected in the comparative analysis of AI vs humans, with an I2 value of 74.3% (95%CI: 45.2%-88.0%), Q = 23.35, and P = 0.0007. This indicates that variability across studies was greater than expected by chance alone.
Additional metrics τ2 = 0.0041 and τ = 0.0644 quantify the between-study variance. This heterogeneity is likely due to differences in AI model design, operator skill level, endoscopic imaging modalities, patient demographics, and clinical practice environments. Importantly, such heterogeneity underscores the context-specific nature of AI’s performance.
To explore potential publication bias, Deeks’ funnel plot asymmetry test was performed. Visual and statistical assessment indicated no significant evidence of publication bias (P > 0.10). This supports confidence in the meta-analysis findings and suggests that the observed pooled estimates accurately reflect the published evidence.
The present meta-analysis shows that AI systems meaningfully outperform human endoscopists in the real-time optical diagnosis of colorectal polyp histology. Pooling nine studies and nearly two thousand lesions, AI reached an overall correct-diagnosis rate of approximately 94% (random-effects proportion 0.9397; 95%CI: 0.9046-0.9624), while human end
Patterns across individual studies help explain the pooled result and its heterogeneity. In trials that leveraged adv
The broader literature supports this gradient of benefit. Early prospective work demonstrated that AI can meet international thresholds for “optical biopsy”, including the ASGE’s PIVI benchmark of ≥ 90% NPV for diminutive rectosigmoid hyperplastic polyps, an essential prerequisite for “resect-and-discard” and “diagnose-and-leave” strategies[40]. Subse
Although our review focused on histology prediction (computer-aided diagnosis), it sits atop a complementary success story in polyp detection (computer-aided detection). Tandem and randomized trials show that computer-aided detection reduces adenoma miss rates and improves adenoma detection rate across experience levels, particularly helping lower-detectors approach expert performance[25]. This matters because characterization can only improve outcomes if detection first ensures the lesion is seen. High-sensitivity computer-aided detection (e.g., Chino et al’s system reporting[39] approximately 97.5% sensitivity for detection) enlarges the funnel of polyps that reach the “optical biopsy” step, where com
These studies differ because of three recurring themes. First, imaging modality is pivotal. Magnifying NBI/blue laser imaging and endocytoscopy yield richer pit-vascular detail, enabling both humans and algorithms to perform better; unsurprisingly, AI trained and deployed with magnification shows larger effect sizes than AI working from standard white-light views. Trials without magnification, such as Barua et al[38] and Djinbachian et al[40] generally report lower absolute accuracies for both groups and narrower AI-human gaps. Second, operator expertise shapes headroom for improvement. Where experts already achieve high performance, AI adds confidence and consistency more than raw accuracy; where performance is variable (general practice, trainees, or visually subtle lesions), AI frequently raises both sensitivity and specificity. Third, dataset provenance and algorithm generalizability matter. Models trained on narrow distributions may falter on different scopes, patient populations, or lesion spectra; multicenter trials help, but heterogeneity remains an unavoidable feature during diffusion of innovation.
Clinically, the implications are considerable. AI augments real-time decision-making, offering a “second pair of eyes” and a disciplined application of pre-specified criteria. When AI predicts a small rectosigmoid lesion is hyperplastic with high confidence, immediate “diagnose-and-leave” becomes plausible under Patient-Centered Outcomes Research Institute - Endoscopy Valuation Initiative/European Society for Gastrointestinal Endoscopy thresholds; when AI flags neoplastic features, the endoscopist can ensure complete resection, retrieval, or opt for advanced en bloc techniques. Gains in confidence and standardization are not trivial: Barua et al[38] increase in high-confidence calls illustrates how AI can expand the proportion of clinically actionable optical diagnoses even without dramatic accuracy differences. Health-economic modelling suggests that implementing discard/Leave in situ strategies under validated AI support could reduce pathology submissions and procedural time, yielding meaningful cost offsets at scale (e.g., estimates in Mori et al[23] and subsequent analyses) while maintaining safety[22]. In real-world practice, these potential gains must be balanced against acquisition and maintenance costs, requirements for regulatory approval and local governance, and the need for ongoing surveillance of model performance over time, including monitoring for model drift. The policy climate is also evolving: Societies such as the BSG endorse adopting technologies that help meet quality standards, for example ade
A recurrent question is whether AI should augment or replace human judgement. The evidence and current practice favor augmentation: AI is best conceived as a vigilant, tireless assistant that reduces oversight errors and narrows performance variability, while the endoscopist retains contextual reasoning and accountability. Even the autonomous computer-aided diagnosis arm in Djinbachian et al[40] matched average human performance rather than surpassing expert level, and absolute accuracies of approximately 75%-80% in pragmatic diminutive polyp cohorts indicate that neither party is infallible; in our pooled estimates, approximately 6%-9% of AI diagnoses were still incorrect, emphasizing the ongoing value of a clinician in the loop. Limited “replacement” may be acceptable for specific tasks, for example foregoing histology on diminutive distal hyperplastic lesions when validated AI provides ≥ 90% NPV with high confidence, but even in that context, the endoscopist adjudicates uncertainties, weighs comorbidity risks, and integrates patient preferences[22]. Over the near term, the highest-value model remains partnership: Computer-aided detection to ensure few lesions are missed; computer-aided diagnosis to standardize and accelerate optical biopsy; and clinicians to manage exceptions, complications, and shared decisions. Practical implementation will therefore require not only technical validation but also structured training, workflow integration, and clear medical-legal frameworks.
This synthesis has limitations that mirror the field’s growing pains. Between-study heterogeneity was substantial (I2 = 74%-81%, P < 0.001), reflecting differences in imaging platforms, operator expertise, lesion spectra, and study design; we addressed this with random-effects modelling, but residual variability means that the pooled average may not uniformly apply to every setting. Publication bias is a concern in a rapidly advancing area, given the preferential visibility of posi
Despite these caveats, the weight of evidence points to a consistent conclusion. AI-assisted characterization is at least as good as, and usually better than, expert optical diagnosis across diverse settings, with the largest gains seen when baseline human performance is modest, imaging is enhanced, and outputs are integrated thoughtfully into workflow. Coupled with the well-established improvements in detection (lower adenoma miss rates and better adenoma detection rates across skill levels)[37], the characterization gains suggest that AI can raise both the floor and the ceiling of colonoscopy quality. The path to routine, safe implementation runs through multicenter pragmatic trials, standardized reporting, human-factors-aware interfaces that display interpretable, well-calibrated outputs, and realistic assessments of cost and resource implications. As societies such as the BSG and ASGE refine guidance on discard/Leave in situ policies, carefully governed AI can help endoscopy units deliver more consistent care, reduce unnecessary pathology and procedures, and direct resources where they have the greatest clinical impact[9,22,23]. In that sense, AI is less a rep
The evolution toward AI-supported endoscopy also parallels broader shifts in how gastroenterologists acquire and apply expertise. As shown in a recent systematic review, immersive technologies such as virtual reality have already proven effective in accelerating endoscopic skill acquisition, reducing procedural time, and improving accuracy among trainees. When combined with AI-guided systems, these training modalities can create a feedback-rich environment in which performance improvement becomes continuous and quantifiable[41]. At the same time, translational work in colorectal oncology underscores the biological complexity that persists beyond the visual field, illustrating how tumor micro-environmental factors such as macrophage polarization continue to shape disease behavior and prognosis[42]. Complementing these technological and biological insights, comparative diagnostic studies in endoscopy - such as the meta-analysis by Woods and Soldera[43] demonstrating the non-inferiority of colon capsule endoscopy to conventional colonoscopy for polyp detection - reinforce how innovation can preserve diagnostic quality while expanding accessibility and patient acceptance. Together, these advances highlight that AI integration is not merely a technical enhancement but part of a larger redefinition of precision gastroenterology - one that couples machine vision with biological insight and human expertise to achieve more individualized, evidence-driven care.
This review has several limitations. First, the primary search was conducted in PubMed/MEDLINE, which, although broadly representative of biomedical and endoscopy research, does not capture all AI-related publications. Important studies indexed exclusively in EMBASE, Scopus, Web of Science, or Cochrane Central Register of Controlled Trials may therefore have been missed, introducing potential database-related selection bias. Second, three eligible studies could not be included because full texts were unobtainable despite attempts to contact corresponding authors and institutions. This may have affected the precision of our pooled estimates, particularly given the modest number of included trials. Third, several studies lacked reporting of human-reader accuracy or comparator data. In accordance with standards for diagnostic test-accuracy meta-analysis, these studies were included only in pooled estimates of AI performance and excluded from the AI-vs-human comparisons; however, incomplete reporting limits the depth of comparative analysis. Fourth, although substantial heterogeneity is expected in imaging and AI literature, the limited and inconsistently reported methodological details across studies prevented meaningful subgroup analyses (e.g., by imaging modality, operator expertise, AI architecture, or study design). As more standardized and higher-quality trials become available, future updates of this work will be able to explore these sources of heterogeneity more robustly. Together, these factors should be considered when interpreting the results.
AI is redefining the standard of optical diagnosis in colonoscopy. The evidence synthesized in this review demonstrates that AI can deliver a level of diagnostic accuracy once achievable only by highly experienced endoscopists, while main
Looking forward, the adoption of validated AI systems should be viewed not as experimental but as an essential step in modern CRC prevention. Their use can rationalize resource allocation, reduce procedural inefficiencies, and expand access to high-quality diagnosis in both expert and community settings. For policymakers and professional societies, the challenge now lies not in proving AI’s value but in defining frameworks for its safe deployment, auditing, and con
We extend our appreciation to the Faculty of Life Sciences and Education at the University of South Wales in association with Learna Ltd. for the Master of Science in Gastroenterology program and their invaluable support in our work. We sincerely acknowledge the efforts of the University of South Wales and commend them for their commitment to pro
| 1. | Vabi BW, Gibbs JF, Parker GS. Implications of the growing incidence of global colorectal cancer. J Gastrointest Oncol. 2021;12:S387-S398. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 31] [Cited by in RCA: 27] [Article Influence: 5.4] [Reference Citation Analysis (0)] |
| 2. | World Health Organization. Global Cancer Observatory: Cancer Today. [cited 15 October 2025]. Available from: https://gco.iarc.fr/today. |
| 3. | Morgan E, Arnold M, Gini A, Lorenzoni V, Cabasag CJ, Laversanne M, Vignat J, Ferlay J, Murphy N, Bray F. Global burden of colorectal cancer in 2020 and 2040: incidence and mortality estimates from GLOBOCAN. Gut. 2023;72:338-344. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1885] [Cited by in RCA: 1528] [Article Influence: 509.3] [Reference Citation Analysis (8)] |
| 4. | Arnold M, Sierra MS, Laversanne M, Soerjomataram I, Jemal A, Bray F. Global patterns and trends in colorectal cancer incidence and mortality. Gut. 2017;66:683-691. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3978] [Cited by in RCA: 3573] [Article Influence: 397.0] [Reference Citation Analysis (8)] |
| 5. | Brenner H, Stock C, Hoffmeister M. Effect of screening sigmoidoscopy and screening colonoscopy on colorectal cancer incidence and mortality: systematic review and meta-analysis of randomised controlled trials and observational studies. BMJ. 2014;348:g2467. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 707] [Cited by in RCA: 642] [Article Influence: 53.5] [Reference Citation Analysis (6)] |
| 6. | Gupta S, Lieberman D, Anderson JC, Burke CA, Dominitz JA, Kaltenbach T, Robertson DJ, Shaukat A, Syngal S, Rex DK. Recommendations for Follow-Up After Colonoscopy and Polypectomy: A Consensus Update by the US Multi-Society Task Force on Colorectal Cancer. Am J Gastroenterol. 2020;115:415-434. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 168] [Cited by in RCA: 154] [Article Influence: 25.7] [Reference Citation Analysis (1)] |
| 7. | Yang P, Teng F, Bai S, Xia Y, Xie Z, Cheng Z, Li J, Lei Z, Wang K, Zhang B, Yang T, Wan X, Yin H, Shen H, Pawlik TM, Lau WY, Fu Z, Shen F. Liver resection versus liver transplantation for hepatocellular carcinoma within the Milan criteria based on estimated microvascular invasion risks. Gastroenterol Rep (Oxf). 2023;11:goad035. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 11] [Article Influence: 3.7] [Reference Citation Analysis (1)] |
| 8. | Rastogi A, Keighley J, Singh V, Callahan P, Bansal A, Wani S, Sharma P. High accuracy of narrow band imaging without magnification for the real-time characterization of polyp histology and its comparison with high-definition white light colonoscopy: a prospective study. Am J Gastroenterol. 2009;104:2422-2430. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 150] [Cited by in RCA: 134] [Article Influence: 7.9] [Reference Citation Analysis (2)] |
| 9. | Rutter MD, East J, Rees CJ, Cripps N, Docherty J, Dolwani S, Kaye PV, Monahan KJ, Novelli MR, Plumb A, Saunders BP, Thomas-Gibson S, Tolan DJM, Whyte S, Bonnington S, Scope A, Wong R, Hibbert B, Marsh J, Moores B, Cross A, Sharp L. British Society of Gastroenterology/Association of Coloproctology of Great Britain and Ireland/Public Health England post-polypectomy and post-colorectal cancer resection surveillance guidelines. Gut. 2020;69:201-223. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 335] [Cited by in RCA: 294] [Article Influence: 49.0] [Reference Citation Analysis (6)] |
| 10. | Heldwein W, Dollhopf M, Rösch T, Meining A, Schmidtsdorff G, Hasford J, Hermanek P, Burlefinger R, Birkner B, Schmitt W; Munich Gastroenterology Group. The Munich Polypectomy Study (MUPS): prospective analysis of complications and risk factors in 4000 colonic snare polypectomies. Endoscopy. 2005;37:1116-1122. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 213] [Cited by in RCA: 212] [Article Influence: 10.1] [Reference Citation Analysis (1)] |
| 11. | Kruk ME, Gage AD, Arsenault C, Jordan K, Leslie HH, Roder-DeWan S, Adeyi O, Barker P, Daelmans B, Doubova SV, English M, García-Elorrio E, Guanais F, Gureje O, Hirschhorn LR, Jiang L, Kelley E, Lemango ET, Liljestrand J, Malata A, Marchant T, Matsoso MP, Meara JG, Mohanan M, Ndiaye Y, Norheim OF, Reddy KS, Rowe AK, Salomon JA, Thapa G, Twum-Danso NAY, Pate M. High-quality health systems in the Sustainable Development Goals era: time for a revolution. Lancet Glob Health. 2018;6:e1196-e1252. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1897] [Cited by in RCA: 2219] [Article Influence: 277.4] [Reference Citation Analysis (5)] |
| 12. | Byrne MF, Chapados N, Soudan F, Oertel C, Linares Pérez M, Kelly R, Iqbal N, Chandelier F, Rex DK. Real-time differentiation of adenomatous and hyperplastic diminutive colorectal polyps during analysis of unaltered videos of standard colonoscopy using a deep learning model. Gut. 2019;68:94-100. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 535] [Cited by in RCA: 438] [Article Influence: 62.6] [Reference Citation Analysis (7)] |
| 13. | Topol EJ. High-performance medicine: the convergence of human and artificial intelligence. Nat Med. 2019;25:44-56. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 6739] [Cited by in RCA: 4017] [Article Influence: 573.9] [Reference Citation Analysis (7)] |
| 14. | Chongo G, Soldera J. Use of machine learning models for the prognostication of liver transplantation: A systematic review. World J Transplant. 2024;14:88891. [PubMed] [DOI] [Full Text] |
| 15. | Soldera J, Tomé F, Corso LL, Rech MM, Ferrazza AD, Terres AZ, Cini BT, Eberhardt LZ, Balensiefer JIL, Balbinot RS, Muscope ALF, Longen ML, Schena B, Rost GL Jr, Furlan RG, Balbinot RA, Balbinot SS. Use of a machine learning algorithm to predict rebleeding and mortality for oesophageal variceal bleeding in cirrhotic patients. EMJ Gastroenterol. 2020;9:46-48. |
| 16. | Soldera J, Corso LL, Rech MM, Ballotin VR, Bigarella LG, Tomé F, Moraes N, Balbinot RS, Rodriguez S, Brandão ABM, Hochhegger B. Predicting major adverse cardiovascular events after orthotopic liver transplantation using a supervised machine learning model: A cohort study. World J Hepatol. 2024;16:193-210. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 13] [Cited by in RCA: 14] [Article Influence: 7.0] [Reference Citation Analysis (1)] |
| 17. | Ballotin VR, Bigarella LG, Soldera J, Soldera J. Deep learning applied to the imaging diagnosis of hepatocellular carcinoma. Artif Intell Gastrointest Endosc. 2021;2:127-135. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 3] [Cited by in RCA: 5] [Article Influence: 1.0] [Reference Citation Analysis (0)] |
| 18. | Abut S, Okut H, Kallail KJ. Paradigm shift from Artificial Neural Networks (ANNs) to deep Convolutional Neural Networks (DCNNs) in the field of medical image processing. Expert Syst Appl. 2024;244:122983. [DOI] [Full Text] |
| 19. | Esteva A, Robicquet A, Ramsundar B, Kuleshov V, DePristo M, Chou K, Cui C, Corrado G, Thrun S, Dean J. A guide to deep learning in healthcare. Nat Med. 2019;25:24-29. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3854] [Cited by in RCA: 1898] [Article Influence: 271.1] [Reference Citation Analysis (8)] |
| 20. | Misawa M, Kudo SE, Mori Y, Cho T, Kataoka S, Yamauchi A, Ogawa Y, Maeda Y, Takeda K, Ichimasa K, Nakamura H, Yagawa Y, Toyoshima N, Ogata N, Kudo T, Hisayuki T, Hayashi T, Wakamura K, Baba T, Ishida F, Itoh H, Roth H, Oda M, Mori K. Artificial Intelligence-Assisted Polyp Detection for Colonoscopy: Initial Experience. Gastroenterology. 2018;154:2027-2029.e3. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 355] [Cited by in RCA: 283] [Article Influence: 35.4] [Reference Citation Analysis (4)] |
| 21. | Lou S, Du F, Song W, Xia Y, Yue X, Yang D, Cui B, Liu Y, Han P. Artificial intelligence for colorectal neoplasia detection during colonoscopy: a systematic review and meta-analysis of randomized clinical trials. EClinicalMedicine. 2023;66:102341. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 65] [Cited by in RCA: 51] [Article Influence: 17.0] [Reference Citation Analysis (1)] |
| 22. | Rex DK, Anderson JC, Butterly LF, Day LW, Dominitz JA, Kaltenbach T, Ladabaum U, Levin TR, Shaukat A, Achkar JP, Farraye FA, Kane SV, Shaheen NJ. Quality Indicators for Colonoscopy. Am J Gastroenterol. 2024;119:1754-1780. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 55] [Cited by in RCA: 52] [Article Influence: 26.0] [Reference Citation Analysis (1)] |
| 23. | Mori Y, Kudo SE, Misawa M, Saito Y, Ikematsu H, Hotta K, Ohtsuka K, Urushibara F, Kataoka S, Ogawa Y, Maeda Y, Takeda K, Nakamura H, Ichimasa K, Kudo T, Hayashi T, Wakamura K, Ishida F, Inoue H, Itoh H, Oda M, Mori K. Real-Time Use of Artificial Intelligence in Identification of Diminutive Polyps During Colonoscopy: A Prospective Study. Ann Intern Med. 2018;169:357-366. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 464] [Cited by in RCA: 380] [Article Influence: 47.5] [Reference Citation Analysis (4)] |
| 24. | Renner J, Phlipsen H, Haller B, Navarro-Avila F, Saint-Hill-Febles Y, Mateus D, Ponchon T, Poszler A, Abdelhafez M, Schmid RM, von Delius S, Klare P. Optical classification of neoplastic colorectal polyps - a computer-assisted approach (the COACH study). Scand J Gastroenterol. 2018;53:1100-1106. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 43] [Cited by in RCA: 37] [Article Influence: 4.6] [Reference Citation Analysis (1)] |
| 25. | Wang P, Liu P, Glissen Brown JR, Berzin TM, Zhou G, Lei S, Liu X, Li L, Xiao X. Lower Adenoma Miss Rate of Computer-Aided Detection-Assisted Colonoscopy vs Routine White-Light Colonoscopy in a Prospective Tandem Study. Gastroenterology. 2020;159:1252-1261.e5. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 198] [Cited by in RCA: 168] [Article Influence: 28.0] [Reference Citation Analysis (5)] |
| 26. | Kudo SE, Misawa M, Mori Y, Hotta K, Ohtsuka K, Ikematsu H, Saito Y, Takeda K, Nakamura H, Ichimasa K, Ishigaki T, Toyoshima N, Kudo T, Hayashi T, Wakamura K, Baba T, Ishida F, Inoue H, Itoh H, Oda M, Mori K. Artificial Intelligence-assisted System Improves Endoscopic Identification of Colorectal Neoplasms. Clin Gastroenterol Hepatol. 2020;18:1874-1881.e2. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 209] [Cited by in RCA: 166] [Article Influence: 27.7] [Reference Citation Analysis (10)] |
| 27. | Sato K, Kuramochi M, Tsuchiya A, Yamaguchi A, Hosoda Y, Yamaguchi N, Nakamura N, Itoi Y, Hashimoto Y, Kasuga K, Tanaka H, Kuribayashi S, Takeuchi Y, Uraoka T. Multicentre study to assess the performance of an artificial intelligence instrument to support qualitative diagnosis of colorectal polyps. BMJ Open Gastroenterol. 2024;11:e001553. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (1)] |
| 28. | van der Zander QEW, Roumans R, Kusters CHJ, Dehghani N, Masclee AAM, de With PHN, van der Sommen F, Snijders CCP, Schoon EJ. Appropriate trust in artificial intelligence for the optical diagnosis of colorectal polyps: the role of human/artificial intelligence interaction. Gastrointest Endosc. 2024;100:1070-1078.e10. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 23] [Cited by in RCA: 17] [Article Influence: 8.5] [Reference Citation Analysis (0)] |
| 29. | Wang P, Berzin TM, Glissen Brown JR, Bharadwaj S, Becq A, Xiao X, Liu P, Li L, Song Y, Zhang D, Li Y, Xu G, Tu M, Liu X. Real-time automatic detection system increases colonoscopic polyp and adenoma detection rates: a prospective randomised controlled study. Gut. 2019;68:1813-1819. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 721] [Cited by in RCA: 603] [Article Influence: 86.1] [Reference Citation Analysis (8)] |
| 30. | Choudhury A, Asan O. Role of Artificial Intelligence in Patient Safety Outcomes: Systematic Literature Review. JMIR Med Inform. 2020;8:e18599. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 115] [Cited by in RCA: 188] [Article Influence: 31.3] [Reference Citation Analysis (1)] |
| 31. | Andersen ES, Birk-Korch JB, Hansen RS, Fly LH, Röttger R, Arcani DMC, Brasen CL, Brandslund I, Madsen JS. Monitoring performance of clinical artificial intelligence in health care: a scoping review. JBI Evid Synth. 2024;22:2423-2446. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 27] [Reference Citation Analysis (1)] |
| 32. | Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, Shamseer L, Tetzlaff JM, Akl EA, Brennan SE, Chou R, Glanville J, Grimshaw JM, Hróbjartsson A, Lalu MM, Li T, Loder EW, Mayo-Wilson E, McDonald S, McGuinness LA, Stewart LA, Thomas J, Tricco AC, Welch VA, Whiting P, Moher D. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. 2021;372:n71. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9803] [Reference Citation Analysis (0)] |
| 33. | Frandsen TF, Bruun Nielsen MF, Lindhardt CL, Eriksen MB. Using the full PICO model as a search tool for systematic reviews resulted in lower recall for some PICO elements. J Clin Epidemiol. 2020;127:69-75. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 20] [Cited by in RCA: 126] [Article Influence: 21.0] [Reference Citation Analysis (1)] |
| 34. | Deeks JJ, Higgins JP, Altman DG, McKenzie JE, Veroniki AA. Chapter 10: Analysing data and undertaking meta-analyses. In: Higgins J, Thomas J, Chandler J, Cumpston M, Li T, Page MJ, Welch V, editors. Cochrane Handbook for Systematic Reviews of Interventions version 6.5. London: Cochrane, 2024. |
| 35. | Song F, Khan KS, Dinnes J, Sutton AJ. Asymmetric funnel plots and publication bias in meta-analyses of diagnostic accuracy. Int J Epidemiol. 2002;31:88-95. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 319] [Cited by in RCA: 355] [Article Influence: 14.8] [Reference Citation Analysis (1)] |
| 36. | Thabane L, Mbuagbaw L, Zhang S, Samaan Z, Marcucci M, Ye C, Thabane M, Giangregorio L, Dennis B, Kosa D, Borg Debono V, Dillenburg R, Fruci V, Bawor M, Lee J, Wells G, Goldsmith CH. A tutorial on sensitivity analyses in clinical trials: the what, why, when and how. BMC Med Res Methodol. 2013;13:92. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 520] [Cited by in RCA: 599] [Article Influence: 46.1] [Reference Citation Analysis (1)] |
| 37. | Balduzzi S, Rücker G, Schwarzer G. How to perform a meta-analysis with R: a practical tutorial. Evid Based Ment Health. 2019;22:153-160. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 4492] [Cited by in RCA: 3941] [Article Influence: 563.0] [Reference Citation Analysis (5)] |
| 38. | Barua I, Wieszczy P, Kudo SE, Misawa M, Holme Ø, Gulati S, Williams S, Mori K, Itoh H, Takishima K, Mochizuki K, Miyata Y, Mochida K, Akimoto Y, Kuroki T, Morita Y, Shiina O, Kato S, Nemoto T, Hayee B, Patel M, Gunasingam N, Kent A, Emmanuel A, Munck C, Nilsen JA, Hvattum SA, Frigstad SO, Tandberg P, Løberg M, Kalager M, Haji A, Bretthauer M, Mori Y. Real-Time Artificial Intelligence-Based Optical Diagnosis of Neoplastic Polyps during Colonoscopy. NEJM Evid. 2022;1:EVIDoa2200003. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 108] [Cited by in RCA: 85] [Article Influence: 21.3] [Reference Citation Analysis (1)] |
| 39. | Chino A, Ide D, Abe S, Yoshinaga S, Ichimasa K, Kudo T, Ninomiya Y, Oka S, Tanaka S, Igarashi M. Performance evaluation of a computer-aided polyp detection system with artificial intelligence for colonoscopy. Dig Endosc. 2024;36:185-194. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 5] [Article Influence: 2.5] [Reference Citation Analysis (1)] |
| 40. | Djinbachian R, Haumesser C, Taghiakbari M, Pohl H, Barkun A, Sidani S, Liu Chen Kiow J, Panzini B, Bouchard S, Deslandres E, Alj A, von Renteln D. Autonomous Artificial Intelligence vs Artificial Intelligence-Assisted Human Optical Diagnosis of Colorectal Polyps: A Randomized Controlled Trial. Gastroenterology. 2024;167:392-399.e2. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 62] [Cited by in RCA: 48] [Article Influence: 24.0] [Reference Citation Analysis (1)] |
| 41. | Dương TQ, Soldera J. Virtual reality tools for training in gastrointestinal endoscopy: A systematic review. Artif Intell Gastrointest Endosc. 2024;5:92090. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 6] [Reference Citation Analysis (6)] |
| 42. | Brambilla E, Brambilla DJF, Tregnago AC, Riva F, Pasqualotto FF, Soldera J. Exploring macrophage polarization as a prognostic indicator for colorectal cancer: Unveiling the impact of metalloproteinase mutations. World J Clin Cases. 2025;13:105011. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 3] [Reference Citation Analysis (6)] |
| 43. | Woods M, Soldera J. Colon capsule endoscopy polyp detection rate vs colonoscopy polyp detection rate: Systematic review and meta-analysis. World J Meta-Anal. 2024;12:100726. [RCA] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 1] [Reference Citation Analysis (2)] |