Published online Mar 27, 2026. doi: 10.4254/wjh.v18.i3.112850
Revised: September 29, 2025
Accepted: December 24, 2025
Published online: March 27, 2026
Processing time: 230 Days and 14.8 Hours
The human leukocyte antigen (HLA) system represents one of the most genetically diverse and densely packed genomic regions, playing a fundamental role in orchestrating immune responses. Among molecular markers linked to viral pathogenesis and immune modulation are insulin-like growth factor 2 (IGF-2) and intestinal fatty-acid-binding protein (I-FABP).
To investigate the influence of the HLA variant rs1131500, together with IGF-2 and I-FABP, on susceptibility to hepatitis C virus (HCV), coronavirus disease 2019 (COVID-19), and their co-occurrence, to identify predictive biomarkers and potential therapeutic targets.
The study involved quantifying circulating levels of IGF-2, I-FABP, and interferon-gamma (IFN-γ) and genotyping HLA rs1131500 using real-time polymerase chain reaction. Participants provided nasopharyngeal swabs for detection of severe acute respiratory syndrome coronavirus 2 RNA and blood samples for HCV RNA analysis and biomarker assessment.
Levels of IGF-2 and I-FABP were notably higher in all patient categories, with the highest values observed in co-infected individuals (P < 0.0001). A moderate positive correlation was observed between I-FABP and IFN-γ in COVID-19 cases (r = 0.261, P = 0.05). IGF-2 had the most substantial predictive value (odds ratio [OR]: 4.5-5.5), while IFN-γ showed a protective trend (OR < 1) when combined with IGF-2 and I-FABP.
IGF-2 emerged as the most consistent and reliable biomarker across all patient groups, particularly in COVID-19 and co-infections. I-FABP was a strong marker for co-infection, less so for COVID-19, and ineffective for HCV alone. Genetically, the CC genotype was more common in the HCV and Control groups, whereas the TT genotype was associated with COVID-19 and co-infection, suggesting potential diagnostic value.
Core Tip: This study identifies insulin-like growth factor 2 as a robust molecular biomarker for distinguishing between coronavirus disease 2019 (COVID-19), hepatitis C virus, and co-infection, with the highest diagnostic performance in co-infected individuals. Intestinal fatty-acid-binding protein showed selective utility in coinfections, while interferon-gamma displayed a protective association. Genotyping of human leukocyte antigen rs1131500 revealed the TT genotype as more frequent in COVID-19 and coinfection cases, suggesting a potential immunogenetic risk factor. These findings provide new insight into host immune responses and offer clinically relevant biomarkers for the diagnosis and personalized management of viral infections.
- Citation: Mohamed AA, Abdel-Hamid RM, Amin A, A Gawad EAH, Saed SM, Gaber DA, Ezz Al Arab M, El-Agawy W, AbuRahma MZA, Elosaily HH, Shaaban EE, Ismail SA, Sakr AA. Immunogenetic and molecular biomarkers in COVID-19 and hepatitis C virus infection: Evaluating the diagnostic potential of HLA (rs1131500), IGF-2, and I-FABP. World J Hepatol 2026; 18(3): 112850
- URL: https://www.wjgnet.com/1948-5182/full/v18/i3/112850.htm
- DOI: https://dx.doi.org/10.4254/wjh.v18.i3.112850
The emergence of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has had profound global health consequences, prompting extensive investigation into host immune responses during coronavirus disease 2019 (COVID-19). Among these responses, interferons (IFNs) play a central role in antiviral defense and regulation of disease pro
Unlike in SARS-CoV-2 infection, IFN secretion during hepatitis C virus (HCV) infection does not necessarily require viral replication within macrophages or dendritic cells. These innate immune cells can detect viral particles or debris containing viral components, maintaining vigilance and initiating cytokine responses independently.
IFN signaling remains a cornerstone of antiviral immunity, though its regulation in HCV infection and the down
The liver serves not only as a metabolic hub but also as a source of endocrine mediators, particularly in insulin-regulated pathways. It synthesizes insulin-like growth factors such as insulin-like growth factor (IGF)-1 and IGF-2, as well as IGF-binding proteins (IGFBPs), all of which are critical to metabolic homeostasis and cellular proliferation[26-28]. While circulating IGF-s contribute to systemic development, their release from hepatic tissue is influenced by various physiological factors. The IGF regulatory system comprises not only IGFs and IGFBPs but also a complex network of receptors and proteolytic enzymes[29,30].
Increasing evidence highlights endocrine abnormalities in COVID-19, including disruptions in the thyroid, adrenal, pancreatic, and gonadal axes. Clinical studies have measured serum IGF-1 and growth hormone concentrations in both critically and non-critically ill patients with COVID-19 to examine their prognostic relevance[31]. Additional research using large datasets, such as the United Kingdom Biobank, has also explored pre-diagnostic IGF-1 levels in relation to COVID-19 mortality[32]. Targeted proteomic studies have similarly investigated plasma biomarkers in small patient cohorts[33].
Despite growing interest, relatively few investigations have examined biomarkers of intestinal damage in critical care settings. I-FABP and citrulline have been proposed as indicators of enterocyte injury in severe COVID-19, though their diagnostic value remains to be fully elucidated[34-39]. One cross-sectional study on patients with cirrhosis demonstrated that I-FABP concentrations reflect hepatic function and complications, suggesting its utility as a potential liver injury marker[40].
The SNP rs1131500, located within the HLA-B gene—a critical element of the major histocompatibility complex class I locus—plays an essential role in presenting viral peptides to cytotoxic T cells. Given the high polymorphism of this region, it has been extensively studied in association with viral diseases, including COVID-19 and HCV. The rs1131500 SNP exhibits three allelic configurations: CC (reference homozygous), CT (heterozygous), and TT (alternate homo
Although direct investigations of rs1131500 in COVID-19 are limited, broader research implicates HLA-B alleles in susceptibility to severe infection and mortality risk. One study identified specific HLA genotypes significantly associated with worse outcomes in SARS-CoV-2-infected individuals lacking comorbidities[41]. Additional research has shown that polymorphic HLA regions influence T cell epitope recognition and effectiveness in clearing viral infections[42]. Within HCV research, HLA variants have been linked to either chronic persistence or spontaneous viral clearance[43]. Genome-wide association studies in Japanese populations have further identified SNPs such as rs1989972, rs2293766, and rs1877033 as predictive of cirrhosis development in HCV-infected individuals[44].
While previous studies have explored genetic and molecular factors in either COVID-19 or HCV, few have examined their combined effects, particularly in cases of coinfection. Our study is unique in simultaneously investigating HLA rs1131500 and key markers (IGF-2, I-FABP, IFN-γ), offering a more integrated view of host susceptibility and immune response. This approach fills a gap in the literature and may help identify shared risk factors and potential therapeutic targets across these infections.
A prospective cross-sectional study was conducted at the National Hepatology and Tropical Medicine Research Institute in Cairo, Egypt, involving a total of 228 participants. Before enrollment, all participants received comprehensive in
Participants were equally distributed into four groups (n = 57 per group): Patients coinfected with HCV and SARS-CoV-2 (COVID-19), patients with COVID-19 only, patients with HCV infection only, and healthy control individuals. Patients were required to be aged ≥ 21 years to be eligible for enrolment in the study. Patients in the HCV-positive groups were required to have detectable HCV RNA levels (>12 IU/mL) persisting for at least 6 months and no prior antiviral treatment or diagnosis of hepatocellular carcinoma. The control group comprised healthy individuals with confirmed negative results for both HCV and SARS-CoV-2.
The exclusion criteria encompassed chronic inflammatory diseases, cardiovascular disorders, thyroid dysfunction, diabetes mellitus, alcohol consumption, active schistosomiasis, hepatitis B virus co-infection, and previous IFN therapy.
This study was conducted as an exploratory investigation using consecutive eligible patients recruited during the study period. No formal a priori sample size calculation was performed owing to the limited availability of well-characterized coinfected patients at our center. We enrolled 57 participants per group, reflecting recruitment feasibility. We acknowledge this as a limitation and note that larger, independent cohorts will be important to validate our findings.
Biological specimens were collected for molecular and serological testing. Nasopharyngeal swabs were obtained from all participants to detect SARS-CoV-2 RNA via real-time reverse transcription polymerase chain reaction (RT-PCR). If the initial test was negative, a second sample was collected within 24-48 hours to confirm the result.
Additionally, 5 mL of venous blood was drawn from each participant after an overnight fast. These samples were used for biochemical analyses and molecular testing.
Comprehensive clinical data and laboratory test results were obtained from patient medical records to assist in diagnostic classification and analysis. Complete blood count was performed using the Swelab Alfa Basic Cell Counter (Boule Medical AB, Sweden), and liver and kidney function tests were conducted using the Beckman CX4 Chemical Analyzer (Beckman Coulter Inc., Brea, CA, United States). HCV RNA was quantified using RT-PCR with the HCV RG RT-PCR Kit (Qiagen, Hilden, Germany) on the Rotor-Gene Q 5plex platform. Serum levels of IGF-2, I-FABP, and IFN-γ were mea
Genomic DNA was extracted from whole blood using the QIAamp DNA Blood Mini Kit (Qiagen), according to the manufacturer’s protocol. The concentration and purity of the DNA were assessed using a NanoDrop® 2000 spectrophotometer. Genotyping of the single-nucleotide polymorphism (SNP) rs1131500 in the HLA gene was performed using the Applied Biosystems® StepOne Real-Time PCR System. TaqMan® SNP Genotyping Assays and Genotyping Master Mix were employed for allelic discrimination.
A comprehensive statistical analysis was performed to evaluate data distribution and the relationships among variables. Continuous variables were summarized using descriptive statistics such as mean values and standard deviations. The Shapiro-Wilk test was applied to assess normality, while heatmaps and bar graphs were used for data visualization. For parametric variables, Student’s t-test was used, and for non-parametric data, the Mann-Whitney U test was used. Pearson’s correlation coefficient was used to evaluate the strength and direction of associations between continuous variables. To determine the diagnostic value of selected biochemical markers, receiver operating characteristic (ROC) curve analysis was performed, and the area under the curve (AUC) was calculated along with 95% confidence intervals (CIs). The Youden index was used to define optimal cutoff thresholds for marker performance, and sensitivity, specificity, and likelihood ratios were computed accordingly. Both univariate and multivariate logistic regression models were constructed to investigate the associations between IGF-2, I-FABP, and IFN-γ with the risk of HCV, COVID-19, and co-infection. All statistical analyses and graphical outputs were conducted using R programming, incorporating packages such as heatmap and ggplot2 to ensure reproducibility and visual clarity in the presentation of biomarker-based findings.
The study included 57 participants in each group (Control, HCV, COVID-19, and HCV-COVID-19 co-infection). The mean participant ages were comparable across groups (range: 45.6-48.2 years, P > 0.05), and sex distribution was balanced (48%-52% male, P > 0.05), as shown in Table 1. No significant differences were observed in baseline nutritional status or major comorbidities between the groups. These results suggest that the groups were generally comparable at baseline, thereby reducing the risk of confounding by demographic factors.
| Characteristic | Control (n = 57) | HCV (n = 57) | COVID-19 (n = 57) | HCV-COVID-19 (n = 57) | P |
| Age (years) | 46.2 ± 10.5 | 45.6 ± 11.2 | 47.8 ± 9.8 | 48.2 ± 10.1 | > 0.05 |
| Male | 28 (49.1) | 29 (50.9) | 27 (47.4) | 30 (52.6) | > 0.05 |
| Female | 29 (50.9) | 28 (49.1) | 30 (52.6) | 27 (47.4) | > 0.05 |
| Major comorbidities | 6 (10.5) | 7 (12.3) | 8 (14.0) | 9 (15.8) | > 0.05 |
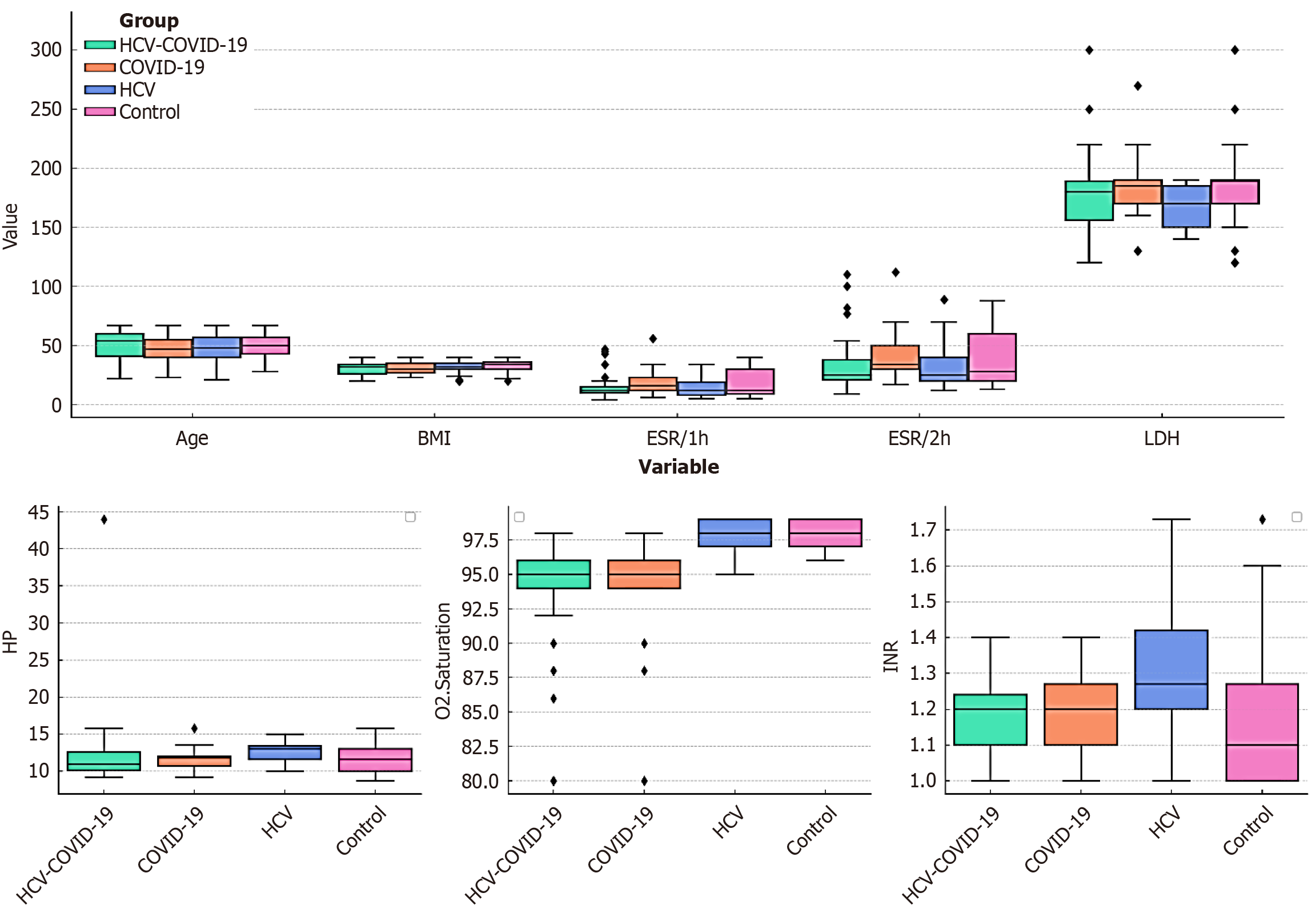
The box plots in Figure 1 demonstrate that sex, age, erythrocyte sedimentation rate (ESR) at 1 h, and ESR at 2 h showed no significant differences across all groups, indicating that these variables are unlikely to be confounding factors. All patient groups exhibited a lower body mass index compared with the control group. The HCV group showed increased hemoglobin (Hb) levels and a significantly higher international normalized ratio (INR) compared with the control group (P = 0.001), indicating impaired blood clotting, likely due to liver dysfunction. No significant differences in INR were observed in the COVID-19 or dual HCV-COVID-19 groups, suggesting that HCV affects clotting factors more pro
Additionally, the HCV group had significantly lower lactate dehydrogenase levels than the control group, which, as a marker of tissue damage and inflammation, may indicate a distinct metabolic or inflammatory response. No significant differences were observed in ESR at 1 h or ESR at 2 h across patient groups. However, the COVID-19 and dual HCV-COVID-19 groups showed significantly lower O2 saturation compared with the control group (P < 0.0001), indicating respiratory impairment, whereas the HCV group did not show a significant difference in O2 saturation. Table 2 presents the mean values and P values for comparisons between each patient group and the control group. Unlike IFN-γ in the HCV group (P = 0.315), all biomarkers were significantly different in all patient groups.
| Markers | HCV-COVID-19 group | COVID group | HCV group | Control group | |||
| mean ± SD | P | mean ± SD | P | mean ± SD | P | mean ± SD | |
| IGF-2 (ng/L) | 121 ± 12.7 | < 0.0001 | 98.46 ± 10.25 | < 0.0001 | 77.91 ± 12.04 | < 0.0001 | 32.67 ± 10.62 |
| I-FABP (ng/L) | 132.42 ± 31 | < 0.0001 | 45.21 ± 8.46 | < 0.0001 | 33.16 ± 9 | < 0.0001 | 31.46 ± 9.04 |
| IFN-γ (ng/L) | 166.18 ± 41.86 | < 0.0001 | 116.81 ± 18.61 | < 0.0001 | 101 ± 14.41 | 0.315 | 95.23 ± 12.92 |
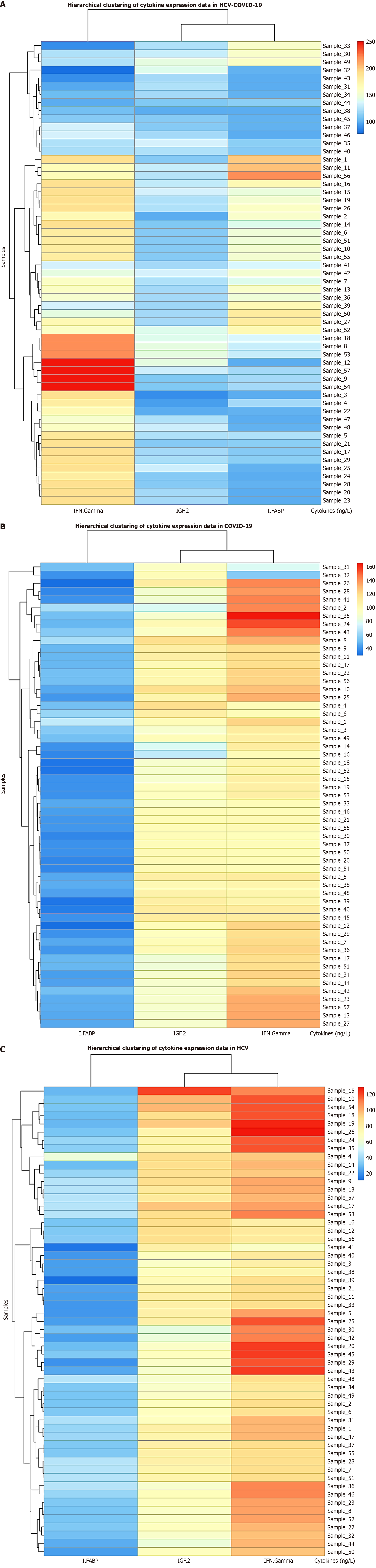
Figure 2 shows a heatmap visualization of biomarker expression in the patient groups generated using R programming packages. The accompanying dendrogram suggests distinct clustering patterns, indicating that certain patients exhibit similar biomarker expression profiles. This clustering may correspond to the different patient groups (HCV, COVID-19, and coinfected), reinforcing the ability of these biomarkers to differentiate disease states.
IGF-2 was significantly elevated in all patient groups, especially in patients coinfected with HCV and COVID-19, indicating a potential role in liver dysfunction and systemic disease progression. IFABP was markedly increased in patients with HCV and COVID-19, suggesting that gut barrier dysfunction is the most severe in this group. IFN-γ levels were highest in the COVID-19 and dual HCV-COVID-19 groups, reflecting strong immune activation. In contrast, HCV alone did not significantly increase IFN-γ, implying that chronic infection may lead to immune tolerance rather than immune activation.
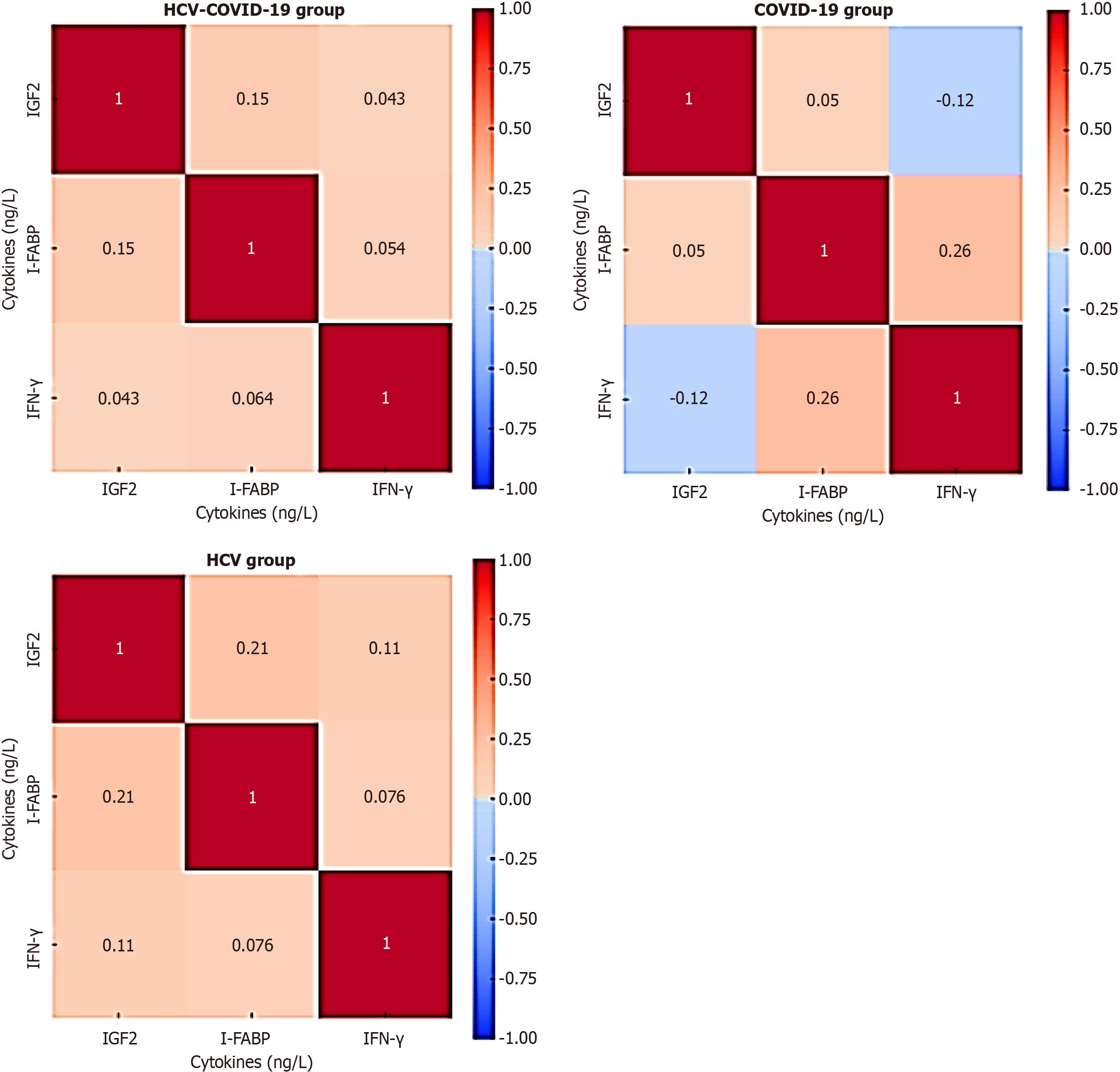
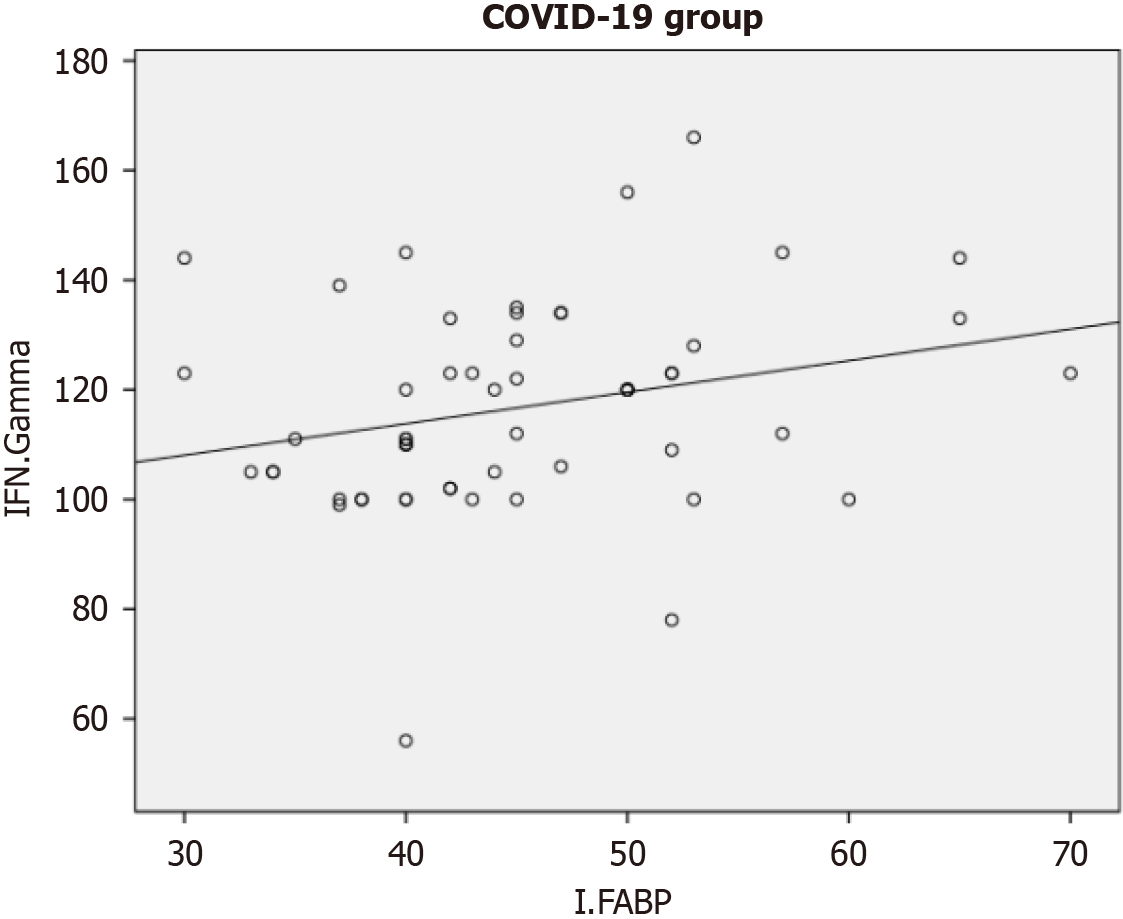
The heatmap in Figure 3 presents Pearson correlation coefficients among three inflammatory biomarkers: IGF-2, I-FABP, and IFN-γ. In the HCV-COVID-19 co-infected group, no statistically significant correlations were observed among these biomarkers, with weak relationships noted for IGF2 vs I-FABP (r = 0.149, P = 0.267), IGF2 vs IFN-γ (r = 0.043, P = 0.754), and I-FABP vs IFN-γ (r = 0.064, P = 0.637). These findings suggest a lack of coordinated interaction between metabolic, intestinal, and immunological pathways in coinfected individuals, possibly reflecting the complex and dysregulated immune response associated with dual viral infections. In the COVID-19 group, a weak but borderline significant positive correlation was detected between I-FABP and IFN-γ (r = 0.261, P = 0.050), indicating a potential link between intestinal barrier damage and immune activation, which may be related to the systemic inflammation and disruption of the gut-lung axis reported in COVID-19. However, IGF2 remained weakly correlated with both I-FABP (r = 0.050, P = 0.713) and IFN-γ (r = −0.119, P = 0.377), suggesting that IGF-2 may not play a central role in the inflammatory profile of COVID-19. In the HCV group, all correlations were also weak and not statistically significant: IGF2 vs I-FABP (r = 0.211, P = 0.115), IGF2 vs IFN-γ (r = 0.106, P = 0.431), and I-FABP vs IFN-γ (r = 0.076, P = 0.573). These results imply that in chronic HCV infection, metabolic and intestinal biomarkers show minimal interaction with immune activation markers such as IFN-γ, possibly owing to immune tolerance or compartmentalized inflammation. Overall, the only potentially meaningful association was found in the COVID-19 group, where the correlation between I-FABP and IFN-γ pointed toward a disease-specific immune-gastrointestinal interplay. In patients with COVID-19, the I-FABP and IFN-γ biomarkers showed a moderate positive correlation (r = 0.261, P = 0.05), as shown in Figure 4.
The logistic regression results presented in Tables 3, 4, 5 reveal distinct biomarker patterns across the study groups. In patients coinfected with HCV and COVID-19, both IGF-2 (odds ratio [OR] = 3.924, P = 0.041) and IFN-γ (OR = 1.107, P < 0.001) were significant predictors, while sex, age, and I-FABP showed no predictive value. In patients with COVID-19, IGF-2 (OR = 4.489, P = 0.037), I-FABP (OR = 1.213, P < 0.001), and IFN-γ (OR = 1.107, P < 0.001) all contributed significantly to disease discrimination, indicating the combined importance of metabolic, intestinal, and immune pathways. In contrast, in HCV infection, IGF-2 (OR = 1.425, P = 0.005) and IFN-γ (OR = 1.032, P = 0.030) were significant, whereas I-FABP was not. Notably, while IFN-γ was consistently significant across groups, its OR values (1.032-1.107) suggest only a modest effect size compared with IGF-2, which showed stronger predictive power. This indicates that IFN-γ may act more as a supportive marker of immune activation rather than a strong independent discriminator.
| Variables | B | SE | P | OR | 95%CI for OR | |
| Lower | Upper | |||||
| Sex | 0.292 | 0.383 | 0.446 | 1.339 | 0.632 | 2.837 |
| Age (years) | 0.004 | 0.018 | 0.833 | 1.004 | 0.97 | 1.039 |
| IGF-2 | 0.367 | 0.472 | 0.041 | 3.924 | 1.56 | 4.736 |
| I-FABP | 0.847 | 87.117 | 0.992 | 2.332 | 0.154 | 3.328 |
| IFN-γ | 0.101 | 0.024 | < 0.001 | 1.107 | 1.056 | 1.16 |
| Variables | B | SE | P | OR | 95%CI for OR | |
| Lower | Upper | |||||
| Sex | 0.221 | 0.384 | 0.566 | 1.247 | 0.588 | 2.646 |
| Age | -0.027 | 0.018 | 0.138 | 0.973 | 0.939 | 1.009 |
| IGF-2 | 0.41 | 0.586 | 0.037 | 4.489 | 1.48 | 5.772 |
| I-FABP | 0.193 | 0.037 | < 0.001 | 1.213 | 1.129 | 1.303 |
| IFN-γ | 0.102 | 0.021 | < 0.001 | 1.107 | 1.062 | 1.154 |
| Variables | B | SE | Sig. | OR | 95%CI for OR | |
| Lower | Upper | |||||
| Sex | 0.148 | 0.385 | 0.7 | 1.160 | 0.545 | 2.467 |
| Age | -0.012 | 0.017 | 0.484 | 0.988 | 0.956 | 1.022 |
| IGF-2 | 0.354 | 0.125 | 0.005 | 1.425 | 1.114 | 1.822 |
| I-FABP | 0.021 | 0.021 | 0.313 | 1.022 | 0.98 | 1.065 |
| IFN-γ | 0.031 | 0.014 | 0.03 | 1.032 | 1.003 | 1.061 |
A limitation of the present study is the sample size. While each group included 57 participants, a post hoc power analysis in Table 6 indicated that this number provided excellent statistical power (> 0.98) for detecting strong effects, such as those observed for IGF-2 and IFN-γ in HCV-COVID-19 co-infection and COVID-19 groups. Moderate power (0.81) was achieved for IGF-2 in the HCV infection group. However, weaker associations, particularly for I-FABP in HCV and HCV-COVID-19 groups, were substantially underpowered, meaning that a large number of participants would be needed in each group to achieve 80% power.
| Groups | Variables | SE | Z | Achieved power | Approx. N per group for 80% power |
| HCV-COVID-19 | IGF-2 | 0.283 | 4.826 | 0.998 | 20 |
| I-FABP | 0.024 | 4.242 | 0.989 | 25 | |
| IFN-γ | 0.936 | 0.919 | 0.16 | 456 | |
| COVID-19 | IGF-2 | 0.347 | 4.325 | 0.991 | 24 |
| I-FABP | 0.037 | 5.281 | 0.999 | 17 | |
| IFN-γ | 0.021 | 4.796 | 0.998 | 20 | |
| HCV | IGF-2 | 0.126 | 2.822 | 0.806 | 57 |
| I-FABP | 0.014 | 2.196 | 0.593 | 93 | |
| IFN-γ | 0.021 | 1.026 | 0.175 | 426 |
These results suggest that while our study was adequately powered for robust biomarker signals, findings involving smaller effect sizes should be interpreted with caution. In addition, potential confounding factors, such as nutritional status, comorbidities, and clinical severity, were not controlled for in the present analysis and may have influenced biomarker variability. Larger, independent cohorts with adjustment for these covariates are necessary to validate our findings and improve their generalizability.
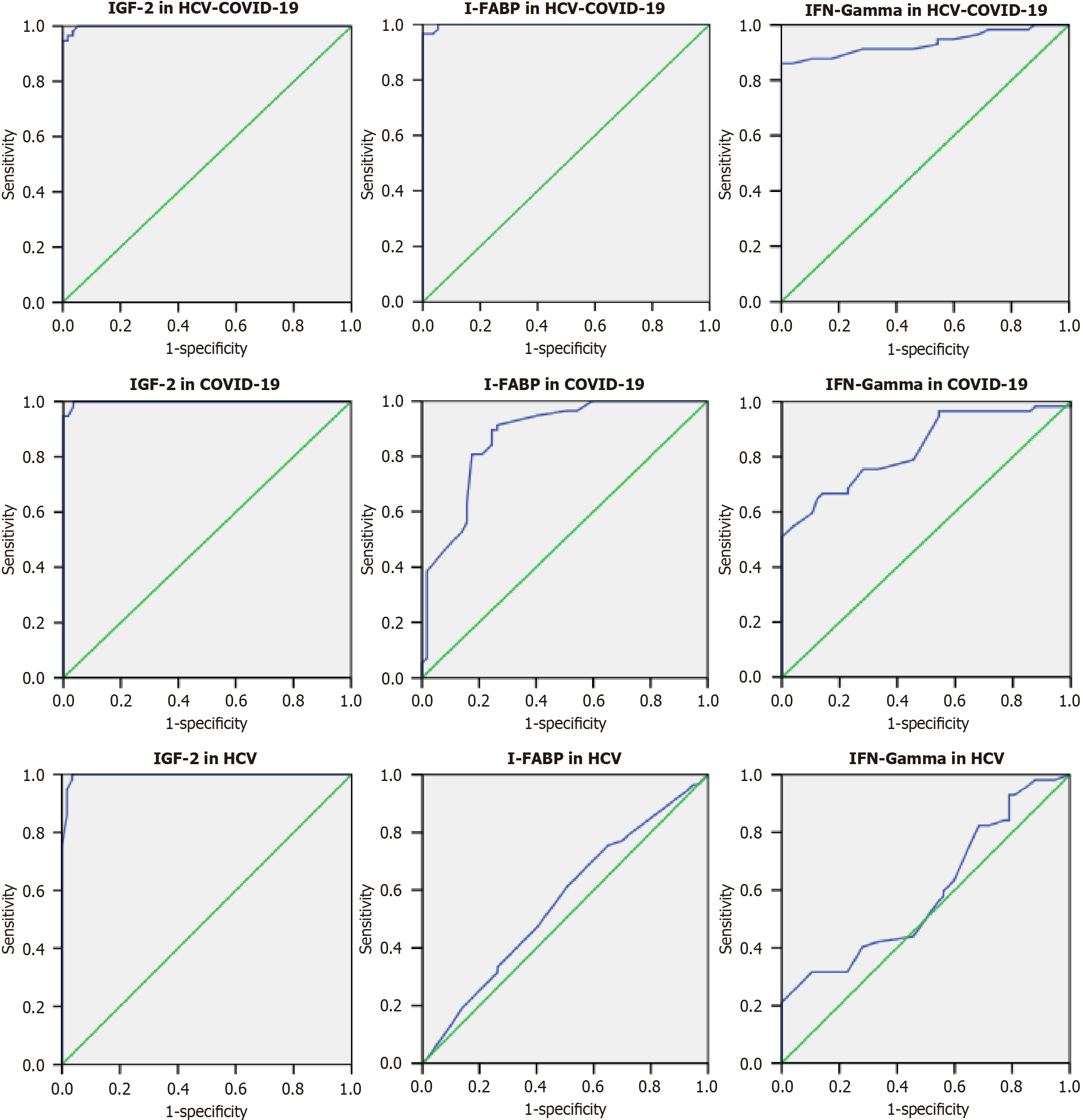
As shown in Figure 5, ROC curve analyses were conducted to evaluate the diagnostic performance of IGF-2, I-FABP, and IFN-γ in distinguishing patient groups from controls. In the HCV-COVID-19 co-infected group, IGF-2 and I-FABP achieved excellent discriminatory ability, each achieving an AUC of 0.998 (95%CI: 0.995-1), while IFN-γ showed strong performance with a slightly lower AUC of 0.934 (95%CI: 0.883-0.985). In the COVID-19 group, IGF-2 retained a high AUC of 0.998 (95%CI: 0.995-1.000), whereas I-FABP and IFN-γ displayed good diagnostic performance, with AUCs of 0.873 (95%CI: 0.808-0.938) and 0.830 (95%CI: 0.756-0.905), respectively. For HCV alone, IGF-2 maintained high discriminatory ability, with an AUC of 0.996 (95%CI: 0.989-1.000), while I-FABP and IFN-γ exhibited more modest diagnostic value, with AUCs of 0.559 (95%CI: 0.453-0.665) and 0.591 (95%CI: 0.486-0.696), respectively.
The traditional academic grading system was used as a benchmark for classifying the accuracy of diagnostic tests[45]. ROC analyses revealed important differences in the diagnostic potential of the studied biomarkers across groups. IGF-2 consistently demonstrated excellent discriminative ability, with AUC values approaching 1.0 in HCV, COVID-19, and HCV-COVID-19 co-infection. This strong performance suggests that IGF-2 may serve as a robust biomarker of virus-related immune dysregulation. In contrast, I-FABP and IFN-γ displayed more variable diagnostic accuracy. In co-infected patients, both biomarkers reached high AUC values, indicating potential synergistic effects of dual viral infection on intestinal and immunological pathways. However, in HCV alone, I-FABP and IFN-γ showed only modest performance, reflecting the limited role of intestinal permeability and systemic inflammatory activation in isolated HCV infection. In COVID-19, both markers achieved intermediate AUC values, supporting their partial but not exclusive contribution to disease discrimination. Collectively, these findings emphasize that while IGF-2 is a consistent predictor across disease contexts, I-FABP and IFN-γ provide additional but context-dependent diagnostic information.
An important limitation of our ROC analyses is that no class balancing, resampling, or cross-validation was performed, and no independent validation cohort was available. As a result, although IGF-2 achieved excellent diagnostic per
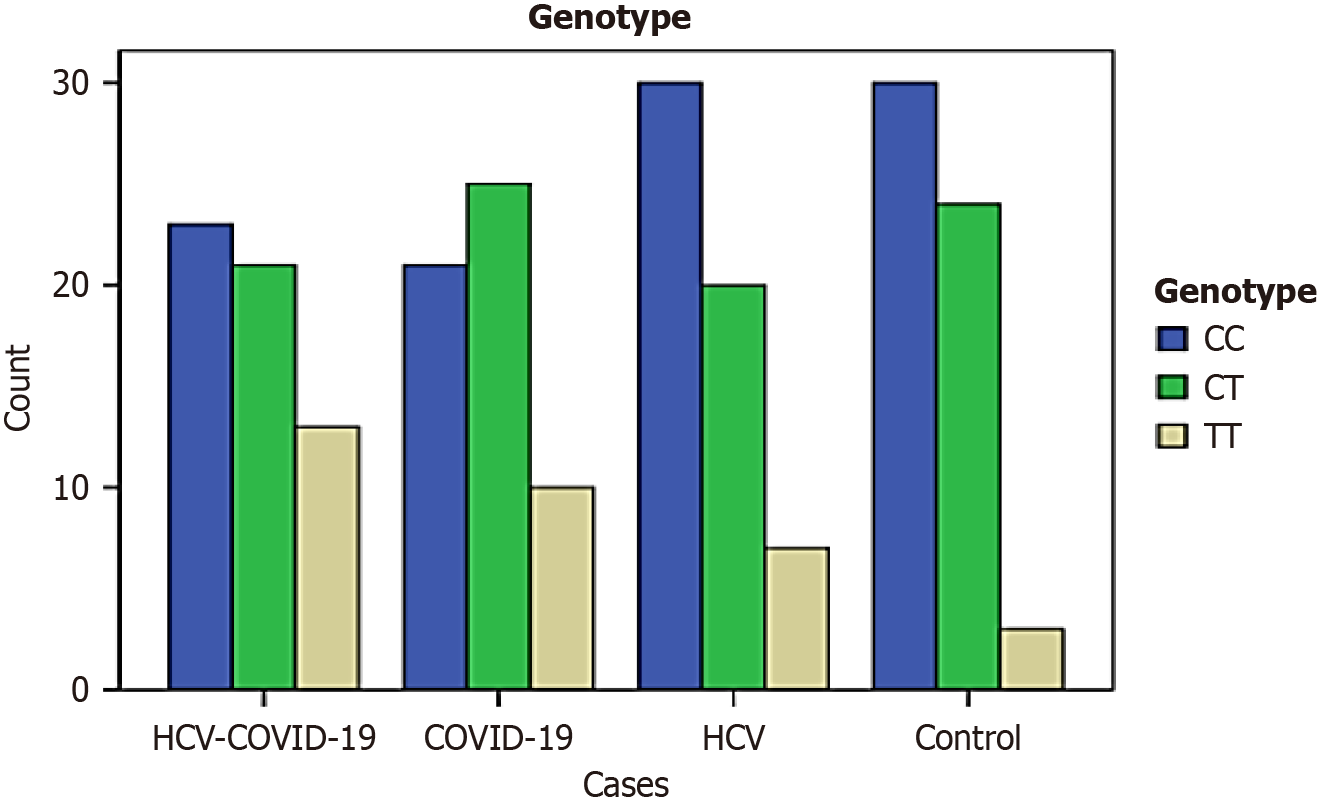
The bar graph in Figure 6 visually represents the genotype distribution (CC, CT, CC) across the four groups: HCV-COVID-19, COVID-19, HCV, and Control. The rs1131500 genotype distribution shows that the CC genotype is more frequent in the HCV and control groups. In contrast, the TT genotype appears more frequently in COVID-19 and co-infected cases, suggesting a possible link between the TT genotype and COVID-19 susceptibility. The CT genotype remains relatively stable across groups, indicating it may not be a strong diagnostic marker on its own.
The biomarkers IGF-2, I-FABP, IFN-γ, and the HLA rs1131500 variant were selected based on prior evidence of their involvement in viral pathogenesis and immune regulation. IGF-2 has been associated with liver disease progression and viral infections[26,41]. I-FABP serves as a sensitive marker of intestinal barrier dysfunction and systemic inflammation[46], while IFN-γ is a central antiviral cytokine with diagnostic and prognostic value in both HCV and COVID-19[47,48]. In addition, these biomarkers showed significant differences between patient groups and controls in our dataset, sup
Our findings not only confirmed the important role of IFN-γ in viral infections but also revealed its limited diagnostic value in HCV. Although type I IFN enhances NK cell cytotoxicity against HCV-infected cells[46-48], NK cells are not directly activated by HCV virions[49]. Instead, IFN-γ production depends on type I IFN signaling and plasmacytoid dendritic cells[50]. This may explain why IFN-γ plays a recognized role in HCV immunity[51,52], yet in our study, it provided strong discrimination in COVID-19 while showing only weak performance in HCV (AUC = 0.591). In contrast, in COVID-19, both experimental and clinical studies highlight IFN-γ as an antiviral mediator, with recombinant IFN-γ reducing SARS-CoV-2 infection and disease progression in experimental models[53-55].
IGF-2 emerged as the most robust biomarker across all groups, with the highest levels observed in coinfection. This agrees with reports that elements of the IGF axis, including IGFBP-2, are upregulated in severe COVID-19 and associated with poor outcomes[56]. Although studies directly addressing IGF-2 in HCV are limited, its elevation in our cohort suggests a role in HCV pathophysiology and warrants further research.
I-FABP levels were significantly elevated in all patient groups, particularly among coinfected individuals. This aligns with studies showing I-FABP elevation in critically ill patients with COVID-19, where it correlated with severity and mortality. Other reports, however, describe decreased levels in some COVID-19 cohorts, suggesting that I-FABP may vary with disease stage or host factors. In HCV, elevated I-FABP has also been documented, especially in coinfected patients, reflecting virus-induced intestinal mucosal damage[38,57].
Taken together, our results suggest that IGF-2 is the most consistent biomarker across groups, while I-FABP con
We also observed a higher frequency of the TT genotype of rs1131500 in patients with COVID-19 and those coinfected with COVID-19 and HCV, whereas the CC genotype predominated in those with HCV and controls. This suggests a possible genetic predisposition influencing susceptibility to COVID-19. Although direct studies on rs1131500 are lacking, extensive work on HLA polymorphisms supports links between genetic variants and viral susceptibility[41].
Nevertheless, our findings must be interpreted with caution. The sample size was modest, statistical power may be limited, and prior evidence is scarce. Therefore, the observed associations should be considered preliminary and require validation in larger, ethnically diverse cohorts.
The present study reveals that IGF-2 and I-FABP levels are markedly increased in individuals with HCV, COVID-19, and coinfections compared with healthy controls, suggesting their potential utility as diagnostic biomarkers for differentiating these conditions. In contrast, IFN-γ exhibited significant elevation only in the COVID-19 and HCV-COVID-19 groups, with no notable difference observed between the HCV group and the control population. This study identified IGF-2 as a strong independent predictor of COVID-19 infection, with I-FABP contributing moderately and IFN-γ potentially offering a protective effect. In contrast, none of the markers were strongly associated with HCV infection alone. These findings suggest IGF-2 may serve as a valuable biomarker for diagnosing COVID-19 and co-infections, warranting further vali
The diagnostic evaluation of IGF-2, I-FABP, and IFN-γ revealed that IGF-2 exhibited outstanding diagnostic per
Our findings suggest a potential association between the TT genotype of rs1131500 and COVID-19 infection, particularly in coinfected individuals, indicating its possible role as a genetic susceptibility marker. In contrast, the CC genotype may be linked to reduced susceptibility or a neutral role in disease presence, especially among HCV-infected and healthy individuals. However, these associations should be interpreted with caution, as our sample size was modest and prior evidence on rs1131500 in viral infections remains limited. Further validation in larger and more diverse populations is essential to confirm the relevance of this variant.
The authors thank all participants who generously participated in this research. The authors also wish to acknowledge Dr. Badria M Alshehri, Dr. Nashaat S Abd El-Gawaad, and Dr. Zehbah A Al-Ahmed for their valuable support and contributions to this work. Finally, the authors are grateful to the referees for their constructive and insightful comments, which greatly helped in improving the manuscript and shaping it into its final form.
| 1. | van der Made CI, Simons A, Schuurs-Hoeijmakers J, van den Heuvel G, Mantere T, Kersten S, van Deuren RC, Steehouwer M, van Reijmersdal SV, Jaeger M, Hofste T, Astuti G, Corominas Galbany J, van der Schoot V, van der Hoeven H, Hagmolen Of Ten Have W, Klijn E, van den Meer C, Fiddelaers J, de Mast Q, Bleeker-Rovers CP, Joosten LAB, Yntema HG, Gilissen C, Nelen M, van der Meer JWM, Brunner HG, Netea MG, van de Veerdonk FL, Hoischen A. Presence of Genetic Variants Among Young Men With Severe COVID-19. JAMA. 2020;324:663-673. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 700] [Cited by in RCA: 584] [Article Influence: 97.3] [Reference Citation Analysis (1)] |
| 2. | Abers MS, Rosen LB, Delmonte OM, Shaw E, Bastard P, Imberti L, Quaresima V, Biondi A, Bonfanti P, Castagnoli R, Casanova JL, Su HC, Notarangelo LD, Holland SM, Lionakis MS. Neutralizing type-I interferon autoantibodies are associated with delayed viral clearance and intensive care unit admission in patients with COVID-19. Immunol Cell Biol. 2021;99:917-921. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 92] [Cited by in RCA: 87] [Article Influence: 17.4] [Reference Citation Analysis (1)] |
| 3. | Asano T, Boisson B, Onodi F, Matuozzo D, Moncada-Velez M, Maglorius Renkilaraj MRL, Zhang P, Meertens L, Bolze A, Materna M, Korniotis S, Gervais A, Talouarn E, Bigio B, Seeleuthner Y, Bilguvar K, Zhang Y, Neehus AL, Ogishi M, Pelham SJ, Le Voyer T, Rosain J, Philippot Q, Soler-Palacín P, Colobran R, Martin-Nalda A, Rivière JG, Tandjaoui-Lambiotte Y, Chaïbi K, Shahrooei M, Darazam IA, Olyaei NA, Mansouri D, Hatipoğlu N, Palabiyik F, Ozcelik T, Novelli G, Novelli A, Casari G, Aiuti A, Carrera P, Bondesan S, Barzaghi F, Rovere-Querini P, Tresoldi C, Franco JL, Rojas J, Reyes LF, Bustos IG, Arias AA, Morelle G, Christèle K, Troya J, Planas-Serra L, Schlüter A, Gut M, Pujol A, Allende LM, Rodriguez-Gallego C, Flores C, Cabrera-Marante O, Pleguezuelo DE, de Diego RP, Keles S, Aytekin G, Akcan OM, Bryceson YT, Bergman P, Brodin P, Smole D, Smith CIE, Norlin AC, Campbell TM, Covill LE, Hammarström L, Pan-Hammarström Q, Abolhassani H, Mane S, Marr N, Ata M, Al Ali F, Khan T, Spaan AN, Dalgard CL, Bonfanti P, Biondi A, Tubiana S, Burdet C, Nussbaum R, Kahn-Kirby A, Snow AL; COVID Human Genetic Effort; COVID-STORM Clinicians; COVID Clinicians; Imagine COVID Group; French COVID Cohort Study Group; CoV-Contact Cohort; Amsterdam UMC Covid-; Biobank; NIAID-USUHS COVID Study Group, Bustamante J, Puel A, Boisson-Dupuis S, Zhang SY, Béziat V, Lifton RP, Bastard P, Notarangelo LD, Abel L, Su HC, Jouanguy E, Amara A, Soumelis V, Cobat A, Zhang Q, Casanova JL. X-linked recessive TLR7 deficiency in ~1% of men under 60 years old with life-threatening COVID-19. Sci Immunol. 2021;6:eabl4348. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 372] [Cited by in RCA: 321] [Article Influence: 64.2] [Reference Citation Analysis (1)] |
| 4. | Bastard P, Gervais A, Le Voyer T, Rosain J, Philippot Q, Manry J, Michailidis E, Hoffmann HH, Eto S, Garcia-Prat M, Bizien L, Parra-Martínez A, Yang R, Haljasmägi L, Migaud M, Särekannu K, Maslovskaja J, de Prost N, Tandjaoui-Lambiotte Y, Luyt CE, Amador-Borrero B, Gaudet A, Poissy J, Morel P, Richard P, Cognasse F, Troya J, Trouillet-Assant S, Belot A, Saker K, Garçon P, Rivière JG, Lagier JC, Gentile S, Rosen LB, Shaw E, Morio T, Tanaka J, Dalmau D, Tharaux PL, Sene D, Stepanian A, Megarbane B, Triantafyllia V, Fekkar A, Heath JR, Franco JL, Anaya JM, Solé-Violán J, Imberti L, Biondi A, Bonfanti P, Castagnoli R, Delmonte OM, Zhang Y, Snow AL, Holland SM, Biggs C, Moncada-Vélez M, Arias AA, Lorenzo L, Boucherit S, Coulibaly B, Anglicheau D, Planas AM, Haerynck F, Duvlis S, Nussbaum RL, Ozcelik T, Keles S, Bousfiha AA, El Bakkouri J, Ramirez-Santana C, Paul S, Pan-Hammarström Q, Hammarström L, Dupont A, Kurolap A, Metz CN, Aiuti A, Casari G, Lampasona V, Ciceri F, Barreiros LA, Dominguez-Garrido E, Vidigal M, Zatz M, van de Beek D, Sahanic S, Tancevski I, Stepanovskyy Y, Boyarchuk O, Nukui Y, Tsumura M, Vidaur L, Tangye SG, Burrel S, Duffy D, Quintana-Murci L, Klocperk A, Kann NY, Shcherbina A, Lau YL, Leung D, Coulongeat M, Marlet J, Koning R, Reyes LF, Chauvineau-Grenier A, Venet F, Monneret G, Nussenzweig MC, Arrestier R, Boudhabhay I, Baris-Feldman H, Hagin D, Wauters J, Meyts I, Dyer AH, Kennelly SP, Bourke NM, Halwani R, Sharif-Askari NS, Dorgham K, Sallette J, Sedkaoui SM, AlKhater S, Rigo-Bonnin R, Morandeira F, Roussel L, Vinh DC, Ostrowski SR, Condino-Neto A, Prando C, Bonradenko A, Spaan AN, Gilardin L, Fellay J, Lyonnet S, Bilguvar K, Lifton RP, Mane S; HGID Lab; COVID Clinicians; COVID-STORM Clinicians; NIAID Immune Response to COVID Group; NH-COVAIR Study Group; Danish CHGE; Danish Blood Donor Study; St. James's Hospital; SARS CoV2 Interest group; French COVID Cohort Study Group; Imagine COVID-Group; Milieu Intérieur Consortium; CoV-Contact Cohort; Amsterdam UMC Covid-19; Biobank Investigators; COVID Human Genetic Effort; CONSTANCES cohort; 3C-Dijon Study; Cerba Health-Care; Etablissement du Sang study group, Anderson MS, Boisson B, Béziat V, Zhang SY, Vandreakos E, Hermine O, Pujol A, Peterson P, Mogensen TH, Rowen L, Mond J, Debette S, de Lamballerie X, Duval X, Mentré F, Zins M, Soler-Palacin P, Colobran R, Gorochov G, Solanich X, Susen S, Martinez-Picado J, Raoult D, Vasse M, Gregersen PK, Piemonti L, Rodríguez-Gallego C, Notarangelo LD, Su HC, Kisand K, Okada S, Puel A, Jouanguy E, Rice CM, Tiberghien P, Zhang Q, Cobat A, Abel L, Casanova JL. Autoantibodies neutralizing type I IFNs are present in ~4% of uninfected individuals over 70 years old and account for ~20% of COVID-19 deaths. Sci Immunol. 2021;6:eabl4340. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 519] [Cited by in RCA: 454] [Article Influence: 90.8] [Reference Citation Analysis (1)] |
| 5. | Bastard P, Lévy R, Henriquez S, Bodemer C, Szwebel TA, Casanova JL. Interferon-β Therapy in a Patient with Incontinentia Pigmenti and Autoantibodies against Type I IFNs Infected with SARS-CoV-2. J Clin Immunol. 2021;41:931-933. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 50] [Cited by in RCA: 50] [Article Influence: 10.0] [Reference Citation Analysis (1)] |
| 6. | Chauvineau-Grenier A, Bastard P, Servajean A, Gervais A, Rosain J, Jouanguy E, Cobat A, Casanova JL, Rossi B. Autoantibodies neutralizing type I interferons in 20% of COVID-19 deaths in a French hospital. Res Sq. 2021;rs.3.rs-r915062. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 17] [Article Influence: 3.4] [Reference Citation Analysis (1)] |
| 7. | Goncalves D, Mezidi M, Bastard P, Perret M, Saker K, Fabien N, Pescarmona R, Lombard C, Walzer T, Casanova JL, Belot A, Richard JC, Trouillet-Assant S. Antibodies against type I interferon: detection and association with severe clinical outcome in COVID-19 patients. Clin Transl Immunology. 2021;10:e1327. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 108] [Cited by in RCA: 94] [Article Influence: 18.8] [Reference Citation Analysis (1)] |
| 8. | Koning R, Bastard P, Casanova JL, Brouwer MC, van de Beek D; with the Amsterdam U. M.C. COVID-19 Biobank Investigators. Autoantibodies against type I interferons are associated with multi-organ failure in COVID-19 patients. Intensive Care Med. 2021;47:704-706. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 114] [Cited by in RCA: 104] [Article Influence: 20.8] [Reference Citation Analysis (1)] |
| 9. | Solanich X, Rigo-Bonnin R, Gumucio VD, Bastard P, Rosain J, Philippot Q, Perez-Fernandez XL, Fuset-Cabanes MP, Gordillo-Benitez MÁ, Suarez-Cuartin G, Boza-Hernandez E, Riera-Mestre A, Parra-Martínez A, Colobran R, Antolí A, Navarro S, Rocamora-Blanch G, Framil M, Calatayud L, Corbella X, Casanova JL, Morandeira F, Sabater-Riera J. Pre-existing Autoantibodies Neutralizing High Concentrations of Type I Interferons in Almost 10% of COVID-19 Patients Admitted to Intensive Care in Barcelona. J Clin Immunol. 2021;41:1733-1744. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 94] [Cited by in RCA: 85] [Article Influence: 17.0] [Reference Citation Analysis (1)] |
| 10. | Troya J, Bastard P, Planas-Serra L, Ryan P, Ruiz M, de Carranza M, Torres J, Martínez A, Abel L, Casanova JL, Pujol A. Neutralizing Autoantibodies to Type I IFNs in >10% of Patients with Severe COVID-19 Pneumonia Hospitalized in Madrid, Spain. J Clin Immunol. 2021;41:914-922. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 122] [Cited by in RCA: 110] [Article Influence: 22.0] [Reference Citation Analysis (1)] |
| 11. | Vazquez SE, Bastard P, Kelly K, Gervais A, Norris PJ, Dumont LJ, Casanova JL, Anderson MS, DeRisi JL. Neutralizing Autoantibodies to Type I Interferons in COVID-19 Convalescent Donor Plasma. J Clin Immunol. 2021;41:1169-1171. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 67] [Cited by in RCA: 66] [Article Influence: 13.2] [Reference Citation Analysis (1)] |
| 12. | Solanich X, Vargas-Parra G, van der Made CI, Simons A, Schuurs-Hoeijmakers J, Antolí A, Del Valle J, Rocamora-Blanch G, Setién F, Esteller M, van Reijmersdal SV, Riera-Mestre A, Sabater-Riera J, Capellá G, van de Veerdonk FL, van der Hoven B, Corbella X, Hoischen A, Lázaro C. Genetic Screening for TLR7 Variants in Young and Previously Healthy Men With Severe COVID-19. Front Immunol. 2021;12:719115. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 100] [Cited by in RCA: 88] [Article Influence: 17.6] [Reference Citation Analysis (1)] |
| 13. | Acosta-Ampudia Y, Monsalve DM, Rojas M, Rodríguez Y, Gallo JE, Salazar-Uribe JC, Santander MJ, Cala MP, Zapata W, Zapata MI, Manrique R, Pardo-Oviedo JM, Camacho B, Ramírez-Santana C, Anaya JM; CP-COVID-19 group. COVID-19 convalescent plasma composition and immunological effects in severe patients. J Autoimmun. 2021;118:102598. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 115] [Cited by in RCA: 100] [Article Influence: 20.0] [Reference Citation Analysis (0)] |
| 14. | Chang SE, Feng A, Meng W, Apostolidis SA, Mack E, Artandi M, Barman L, Bennett K, Chakraborty S, Chang I, Cheung P, Chinthrajah S, Dhingra S, Do E, Finck A, Gaano A, Geßner R, Giannini HM, Gonzalez J, Greib S, Gündisch M, Hsu AR, Kuo A, Manohar M, Mao R, Neeli I, Neubauer A, Oniyide O, Powell AE, Puri R, Renz H, Schapiro J, Weidenbacher PA, Wittman R, Ahuja N, Chung HR, Jagannathan P, James JA, Kim PS, Meyer NJ, Nadeau KC, Radic M, Robinson WH, Singh U, Wang TT, Wherry EJ, Skevaki C, Luning Prak ET, Utz PJ. New-onset IgG autoantibodies in hospitalized patients with COVID-19. Nat Commun. 2021;12:5417. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 395] [Cited by in RCA: 325] [Article Influence: 65.0] [Reference Citation Analysis (1)] |
| 15. | Wang EY, Mao T, Klein J, Dai Y, Huck JD, Jaycox JR, Liu F, Zhou T, Israelow B, Wong P, Coppi A, Lucas C, Silva J, Oh JE, Song E, Perotti ES, Zheng NS, Fischer S, Campbell M, Fournier JB, Wyllie AL, Vogels CBF, Ott IM, Kalinich CC, Petrone ME, Watkins AE; Yale IMPACT Team, Dela Cruz C, Farhadian SF, Schulz WL, Ma S, Grubaugh ND, Ko AI, Iwasaki A, Ring AM. Diverse functional autoantibodies in patients with COVID-19. Nature. 2021;595:283-288. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 763] [Cited by in RCA: 662] [Article Influence: 132.4] [Reference Citation Analysis (1)] |
| 16. | Ziegler CGK, Miao VN, Owings AH, Navia AW, Tang Y, Bromley JD, Lotfy P, Sloan M, Laird H, Williams HB, George M, Drake RS, Christian T, Parker A, Sindel CB, Burger MW, Pride Y, Hasan M, Abraham GE 3rd, Senitko M, Robinson TO, Shalek AK, Glover SC, Horwitz BH, Ordovas-Montanes J. Impaired local intrinsic immunity to SARS-CoV-2 infection in severe COVID-19. Cell. 2021;184:4713-4733.e22. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 307] [Cited by in RCA: 256] [Article Influence: 51.2] [Reference Citation Analysis (1)] |
| 17. | Shaw ER, Rosen LB, Cheng A, Dobbs K, Delmonte OM, Ferré EMN, Schmitt MM, Imberti L, Quaresima V, Lionakis MS, Notarangelo LD, Holland SM, Su HC. Temporal Dynamics of Anti-Type 1 Interferon Autoantibodies in Patients With Coronavirus Disease 2019. Clin Infect Dis. 2022;75:e1192-e1194. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 38] [Cited by in RCA: 32] [Article Influence: 8.0] [Reference Citation Analysis (1)] |
| 18. | Lucas C, Wong P, Klein J, Castro TBR, Silva J, Sundaram M, Ellingson MK, Mao T, Oh JE, Israelow B, Takahashi T, Tokuyama M, Lu P, Venkataraman A, Park A, Mohanty S, Wang H, Wyllie AL, Vogels CBF, Earnest R, Lapidus S, Ott IM, Moore AJ, Muenker MC, Fournier JB, Campbell M, Odio CD, Casanovas-Massana A; Yale IMPACT Team, Herbst R, Shaw AC, Medzhitov R, Schulz WL, Grubaugh ND, Dela Cruz C, Farhadian S, Ko AI, Omer SB, Iwasaki A. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature. 2020;584:463-469. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1913] [Cited by in RCA: 1651] [Article Influence: 275.2] [Reference Citation Analysis (5)] |
| 19. | Galani IE, Rovina N, Lampropoulou V, Triantafyllia V, Manioudaki M, Pavlos E, Koukaki E, Fragkou PC, Panou V, Rapti V, Koltsida O, Mentis A, Koulouris N, Tsiodras S, Koutsoukou A, Andreakos E. Untuned antiviral immunity in COVID-19 revealed by temporal type I/III interferon patterns and flu comparison. Nat Immunol. 2021;22:32-40. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 479] [Cited by in RCA: 406] [Article Influence: 81.2] [Reference Citation Analysis (1)] |
| 20. | Sposito B, Broggi A, Pandolfi L, Crotta S, Clementi N, Ferrarese R, Sisti S, Criscuolo E, Spreafico R, Long JM, Ambrosi A, Liu E, Frangipane V, Saracino L, Bozzini S, Marongiu L, Facchini FA, Bottazzi A, Fossali T, Colombo R, Clementi M, Tagliabue E, Chou J, Pontiroli AE, Meloni F, Wack A, Mancini N, Zanoni I. The interferon landscape along the respiratory tract impacts the severity of COVID-19. Cell. 2021;184:4953-4968.e16. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 242] [Cited by in RCA: 209] [Article Influence: 41.8] [Reference Citation Analysis (1)] |
| 21. | Lee JS, Park S, Jeong HW, Ahn JY, Choi SJ, Lee H, Choi B, Nam SK, Sa M, Kwon JS, Jeong SJ, Lee HK, Park SH, Park SH, Choi JY, Kim SH, Jung I, Shin EC. Immunophenotyping of COVID-19 and influenza highlights the role of type I interferons in development of severe COVID-19. Sci Immunol. 2020;5:eabd1554. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 793] [Cited by in RCA: 676] [Article Influence: 112.7] [Reference Citation Analysis (1)] |
| 22. | Dorgham K, Quentric P, Gökkaya M, Marot S, Parizot C, Sauce D, Guihot A, Luyt CE, Schmidt M, Mayaux J, Beurton A, Le Guennec L, Demeret S, Ben Salah E, Mathian A, Yssel H, Combadiere B, Combadiere C, Traidl-Hoffmann C, Burrel S, Marcelin AG, Amoura Z, Voiriot G, Neumann AU, Gorochov G. Distinct cytokine profiles associated with COVID-19 severity and mortality. J Allergy Clin Immunol. 2021;147:2098-2107. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 66] [Cited by in RCA: 58] [Article Influence: 11.6] [Reference Citation Analysis (1)] |
| 23. | Zhang S, Wang L, Cheng G. The battle between host and SARS-CoV-2: Innate immunity and viral evasion strategies. Mol Ther. 2022;30:1869-1884. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 63] [Cited by in RCA: 55] [Article Influence: 13.8] [Reference Citation Analysis (1)] |
| 24. | Huang M, Jiang JD, Peng Z. Recent advances in the anti-HCV mechanisms of interferon. Acta Pharm Sin B. 2014;4:241-247. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 24] [Article Influence: 2.0] [Reference Citation Analysis (2)] |
| 25. | Aggarwal NR, King LS, D'Alessio FR. Diverse macrophage populations mediate acute lung inflammation and resolution. Am J Physiol Lung Cell Mol Physiol. 2014;306:L709-L725. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 511] [Cited by in RCA: 481] [Article Influence: 40.1] [Reference Citation Analysis (1)] |
| 26. | Postic C, Dentin R, Girard J. Role of the liver in the control of carbohydrate and lipid homeostasis. Diabetes Metab. 2004;30:398-408. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 356] [Cited by in RCA: 315] [Article Influence: 14.3] [Reference Citation Analysis (1)] |
| 27. | Liu Y, Guo W, Pu Z, Li X, Lei X, Yao J, Yang X. Developmental changes of Insulin-like growth factors in the liver and muscle of chick embryos. Poult Sci. 2016;95:1396-1402. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 20] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 28. | Voci A, Arvigo M, Massajoli M, Garrone S, Bottazzi C, Demori I, Gallo G. IGF-I production by adult rat hepatocytes is stimulated by transforming growth factor-alpha and transforming growth factor-beta1. Eur J Endocrinol. 1999;140:577-582. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 13] [Cited by in RCA: 10] [Article Influence: 0.4] [Reference Citation Analysis (1)] |
| 29. | Rosen CJ. Serum insulin-like growth factors and insulin-like growth factor-binding proteins: clinical implications. Clin Chem. 1999;45:1384-1390. [PubMed] [DOI] [Full Text] |
| 30. | Shin M, Kang HS, Park JH, Bae JH, Song DK, Im SS. Recent Insights into Insulin-Like Growth Factor Binding Protein 2 Transcriptional Regulation. Endocrinol Metab (Seoul). 2017;32:11-17. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 31] [Article Influence: 3.4] [Reference Citation Analysis (1)] |
| 31. | Ilias I, Diamantopoulos A, Botoula E, Athanasiou N, Zacharis A, Tsipilis S, Jahaj E, Vassiliou AG, Vassiliadi DA, Kotanidou A, Tsagarakis S, Dimopoulou I. Covid-19 and Growth Hormone/Insulin-Like Growth Factor 1: Study in Critically and Non-Critically Ill Patients. Front Endocrinol (Lausanne). 2021;12:644055. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 29] [Cited by in RCA: 23] [Article Influence: 4.6] [Reference Citation Analysis (1)] |
| 32. | Fan X, Yin C, Wang J, Yang M, Ma H, Jin G, Song M, Hu Z, Shen H, Hang D. Pre-diagnostic circulating concentrations of insulin-like growth factor-1 and risk of COVID-19 mortality: results from UK Biobank. Eur J Epidemiol. 2021;36:311-318. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 17] [Article Influence: 3.4] [Reference Citation Analysis (1)] |
| 33. | Fraser DD, Cepinskas G, Patterson EK, Slessarev M, Martin C, Daley M, Patel MA, Miller MR, O'Gorman DB, Gill SE, Pare G, Prassas I, Diamandis E. Novel Outcome Biomarkers Identified With Targeted Proteomic Analyses of Plasma From Critically Ill Coronavirus Disease 2019 Patients. Crit Care Explor. 2020;2:e0189. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 38] [Cited by in RCA: 43] [Article Influence: 7.2] [Reference Citation Analysis (1)] |
| 34. | Giron LB, Dweep H, Yin X, Wang H, Damra M, Goldman AR, Gorman N, Palmer CS, Tang HY, Shaikh MW, Forsyth CB, Balk RA, Zilberstein NF, Liu Q, Kossenkov A, Keshavarzian A, Landay A, Abdel-Mohsen M. Corrigendum: Plasma Markers of Disrupted Gut Permeability in Severe COVID-19 Patients. Front Immunol. 2021;12:779064. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 17] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 35. | Piton G, Belon F, Cypriani B, Regnard J, Puyraveau M, Manzon C, Navellou JC, Capellier G. Enterocyte damage in critically ill patients is associated with shock condition and 28-day mortality. Crit Care Med. 2013;41:2169-2176. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 106] [Cited by in RCA: 95] [Article Influence: 7.3] [Reference Citation Analysis (1)] |
| 36. | Guedj K, Uzzan M, Soudan D, Trichet C, Nicoletti A, Weiss E, Manceau H, Nuzzo A, Corcos O, Treton X, Peoc'h K. I-FABP is decreased in COVID-19 patients, independently of the prognosis. PLoS One. 2021;16:e0249799. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 18] [Cited by in RCA: 14] [Article Influence: 2.8] [Reference Citation Analysis (1)] |
| 37. | Piton G, Manzon C, Monnet E, Cypriani B, Barbot O, Navellou JC, Carbonnel F, Capellier G. Plasma citrulline kinetics and prognostic value in critically ill patients. Intensive Care Med. 2010;36:702-706. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 78] [Cited by in RCA: 72] [Article Influence: 4.5] [Reference Citation Analysis (4)] |
| 38. | Tyszko M, Lipińska-Gediga M, Lemańska-Perek A, Kobylińska K, Gozdzik W, Adamik B. Intestinal Fatty Acid Binding Protein (I-FABP) as a Prognostic Marker in Critically Ill COVID-19 Patients. Pathogens. 2022;11:1526. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 14] [Article Influence: 3.5] [Reference Citation Analysis (1)] |
| 39. | Mester P, Räth U, Schmid S, Amend P, Keller D, Krautbauer S, Bondarenko S, Müller M, Buechler C, Pavel V. Serum Insulin-like Growth Factor-Binding Protein-2 as a Prognostic Factor for COVID-19 Severity. Biomedicines. 2024;12:125. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 6] [Article Influence: 3.0] [Reference Citation Analysis (0)] |
| 40. | Miuma S, Miyaaki H, Taura N, Kanda Y, Matsuo S, Tajima K, Takahashi K, Nakao Y, Fukushima M, Haraguchi M, Sasaki R, Ozawa E, Ichikawa T, Nakao K. Elevated intestinal fatty acid-binding protein levels as a marker of portal hypertension and gastroesophageal varices in cirrhosis. Sci Rep. 2024;14:25003. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 2.5] [Reference Citation Analysis (1)] |
| 41. | Cheranev V, Bulusheva I, Vechorko V, Korostin D, Rebrikov D. The Search of Association of HLA Class I and Class II Alleles with COVID-19 Mortality in the Russian Cohort. Int J Mol Sci. 2023;24:3068. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 7] [Cited by in RCA: 6] [Article Influence: 2.0] [Reference Citation Analysis (1)] |
| 42. | Weingarten-Gabbay S, Klaeger S, Sarkizova S, Pearlman LR, Chen DY, Gallagher KME, Bauer MR, Taylor HB, Dunn WA, Tarr C, Sidney J, Rachimi S, Conway HL, Katsis K, Wang Y, Leistritz-Edwards D, Durkin MR, Tomkins-Tinch CH, Finkel Y, Nachshon A, Gentili M, Rivera KD, Carulli IP, Chea VA, Chandrashekar A, Bozkus CC, Carrington M; MGH COVID-19 Collection & Processing Team, Bhardwaj N, Barouch DH, Sette A, Maus MV, Rice CM, Clauser KR, Keskin DB, Pregibon DC, Hacohen N, Carr SA, Abelin JG, Saeed M, Sabeti PC. Profiling SARS-CoV-2 HLA-I peptidome reveals T cell epitopes from out-of-frame ORFs. Cell. 2021;184:3962-3980.e17. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 141] [Cited by in RCA: 116] [Article Influence: 23.2] [Reference Citation Analysis (1)] |
| 43. | Sroka S, Bartelheimer N, Winter A, Heukelbach J, Ariza L, Ribeiro H, Oliveira FA, Queiroz AJ, Alencar C Jr, Liesenfeld O. Prevalence and risk factors of toxoplasmosis among pregnant women in Fortaleza, Northeastern Brazil. Am J Trop Med Hyg. 2010;83:528-533. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 59] [Cited by in RCA: 54] [Article Influence: 3.4] [Reference Citation Analysis (1)] |
| 44. | Kamimura S, Tamura A, Ishii T, Kanda T, Moriyama M. Association of Genetic Polymorphisms With Hepatitis C Virus-related Liver Cirrhosis in Japan. In Vivo. 2020;34:3309-3313. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 1] [Article Influence: 0.2] [Reference Citation Analysis (1)] |
| 45. | Hosmer DW, Lemeshow S. Applied Logistic Regression. John Wiley and Sons, 2000. [DOI] [Full Text] |
| 46. | Golden-Mason L, Cox AL, Randall JA, Cheng L, Rosen HR. Increased natural killer cell cytotoxicity and NKp30 expression protects against hepatitis C virus infection in high-risk individuals and inhibits replication in vitro. Hepatology. 2010;52:1581-1589. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 98] [Cited by in RCA: 94] [Article Influence: 5.9] [Reference Citation Analysis (1)] |
| 47. | Hilligan KL, Namasivayam S, Sher A. BCG mediated protection of the lung against experimental SARS-CoV-2 infection. Front Immunol. 2023;14:1232764. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 9] [Article Influence: 3.0] [Reference Citation Analysis (1)] |
| 48. | Ovsyannikova IG, Haralambieva IH, Crooke SN, Poland GA, Kennedy RB. The role of host genetics in the immune response to SARS-CoV-2 and COVID-19 susceptibility and severity. Immunol Rev. 2020;296:205-219. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 211] [Cited by in RCA: 177] [Article Influence: 29.5] [Reference Citation Analysis (1)] |
| 49. | Ahlenstiel G, Titerence RH, Koh C, Edlich B, Feld JJ, Rotman Y, Ghany MG, Hoofnagle JH, Liang TJ, Heller T, Rehermann B. Natural killer cells are polarized toward cytotoxicity in chronic hepatitis C in an interferon-alfa-dependent manner. Gastroenterology. 2010;138:325-35.e1. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 215] [Cited by in RCA: 213] [Article Influence: 13.3] [Reference Citation Analysis (1)] |
| 50. | Sakr AA, Mohamed AA, Ahmed AE, Abdelhaleem AA, Samir HH, Elkady MA, Hasona NA. Biochemical implication of acetylcholine, histamine, IL-18, and interferon-alpha as diagnostic biomarkers in hepatitis C virus, coronavirus disease 2019, and dual hepatitis C virus-coronavirus disease 2019 patients. J Med Virol. 2024;96:e29857. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 3] [Cited by in RCA: 2] [Article Influence: 1.0] [Reference Citation Analysis (1)] |
| 51. | Yoon JC, Shiina M, Ahlenstiel G, Rehermann B. Natural killer cell function is intact after direct exposure to infectious hepatitis C virions. Hepatology. 2009;49:12-21. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 85] [Cited by in RCA: 83] [Article Influence: 4.9] [Reference Citation Analysis (1)] |
| 52. | Zhang S, Saha B, Kodys K, Szabo G. IFN-γ production by human natural killer cells in response to HCV-infected hepatoma cells is dependent on accessory cells. J Hepatol. 2013;59:442-449. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 42] [Cited by in RCA: 45] [Article Influence: 3.5] [Reference Citation Analysis (1)] |
| 53. | Thimme R, Oldach D, Chang KM, Steiger C, Ray SC, Chisari FV. Determinants of viral clearance and persistence during acute hepatitis C virus infection. J Exp Med. 2001;194:1395-1406. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 937] [Cited by in RCA: 885] [Article Influence: 35.4] [Reference Citation Analysis (1)] |
| 54. | Flynn JK, Dore GJ, Hellard M, Yeung B, Rawlinson WD, White PA, Kaldor JM, Lloyd AR, Ffrench RA; ATAHC Study Group. Early IL-10 predominant responses are associated with progression to chronic hepatitis C virus infection in injecting drug users. J Viral Hepat. 2011;18:549-561. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 53] [Cited by in RCA: 53] [Article Influence: 3.5] [Reference Citation Analysis (1)] |
| 55. | Hilligan KL, Namasivayam S, Clancy CS, Baker PJ, Old SI, Peluf V, Amaral EP, Oland SD, O'Mard D, Laux J, Cohen M, Garza NL, Lafont BAP, Johnson RF, Feng CG, Jankovic D, Lamiable O, Mayer-Barber KD, Sher A. Bacterial-induced or passively administered interferon gamma conditions the lung for early control of SARS-CoV-2. Nat Commun. 2023;14:8229. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 42] [Article Influence: 14.0] [Reference Citation Analysis (1)] |
| 56. | Hilligan KL, Namasivayam S, Clancy CS, O'Mard D, Oland SD, Robertson SJ, Baker PJ, Castro E, Garza NL, Lafont BAP, Johnson R, Ronchese F, Mayer-Barber KD, Best SM, Sher A. Intravenous administration of BCG protects mice against lethal SARS-CoV-2 challenge. J Exp Med. 2022;219:e20211862. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 102] [Cited by in RCA: 86] [Article Influence: 21.5] [Reference Citation Analysis (1)] |
| 57. | Wang W, Yu K, Zhao SY, Mo DG, Liu JH, Han LJ, Li T, Yao HC. The impact of circulating IGF-1 and IGFBP-2 on cardiovascular prognosis in patients with acute coronary syndrome. Front Cardiovasc Med. 2023;10:1126093. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 23] [Cited by in RCA: 16] [Article Influence: 5.3] [Reference Citation Analysis (1)] |