Published online Apr 26, 2026. doi: 10.4252/wjsc.v18.i4.115218
Revised: December 14, 2025
Accepted: February 24, 2026
Published online: April 26, 2026
Processing time: 170 Days and 23.9 Hours
Tendon stem/progenitor cells (TSPCs) are a novel type of stem cell. TSPCs share common characteristics with stem cells, including their proliferation, pluripo
To explore how circ_PVT1 regulates the proliferation and differentiation of TSPCs.
Mouse TSPCs were isolated from the Achilles tendon of 12 male C57Bl/6 mice at postnatal day 30, and 5 ng/mL of transforming growth factor (TGF)-β1 was used to induce the tenogenic differentiation in TSPCs. Picro-Sirius red staining was used to detect the collagen expression of TSPCs. A Cell Counting Kit-8, Transwell assays, and flow cytometry were used to assess the proliferation, migration, and apoptosis of TSPCs. Biochemical kits were used to determine the levels of reactive oxygen species, malondialdehyde, glutathione, superoxide dismutase, and ATP. Then, N6-methyladenosine (m6A) dot blot and methylated RNA immunoprecipitation polymerase chain reaction (RIP-PCR) were used to examine the m6A levels. Moreover, RNA pulldown and RIP-PCR were performed to analyze the interaction between WTAP and circ_PVT1.
TGF-β1 treatment induced tenogenic differentiation of TSPCs. Circ_PVT1 knockdown reversed the increase of tendon-specific protein, cell proliferation, and migration that was induced by TGF-β1 treatment. In addition, circ_PVT1 inhibition promoted oxidative stress and mitochondrial damage in the TGF-β1-induced TSPCs. The m6A modification level of circ_PVT1 was upregulated in the TGF-β1-induced TSPCs. Furthermore, RNA pulldown and RIP-PCR showed that circ_PVT1 interacted with WTAP in TSPCs, and the WTAP-mediated m6A modification of circ_PVT1 regulated the differentiation of TSPCs.
WTAP-mediated m6A modification of circ_PVT1 promotes the proliferation and tenogenic differentiation of TSPCs, thereby indicating a promising therapeutic strategy for tendon repair.
Core Tip: Tendon stem/progenitor cells share common characteristics with stem cells, such as proliferation, pluripotency, and self-renewal ability. Under suitable conditions, they can differentiate into tendon cells and play a crucial role in maintaining the stability of tendons and in repair after injury. We found that circular RNA plasmacytoma variant translocation 1 deficiency could inhibit tenogenic differentiation. In addition, WTAP-mediated N6-methyladenosine modification of circular RNA plasmacytoma variant translocation 1 promotes the proliferation and tenogenic differentiation of tendon stem/progenitor cells. These results suggest a promising therapeutic strategy for tendon diseases.
- Citation: Chen Y, Hu X, Jia XT, Gao RZ, Hou QS, Zhou SG. WTAP-mediated N6-methyladenosine modification of circ_PVT1 promotes proliferation and tenogenic differentiation of tendon stem/progenitor cells. World J Stem Cells 2026; 18(4): 115218
- URL: https://www.wjgnet.com/1948-0210/full/v18/i4/115218.htm
- DOI: https://dx.doi.org/10.4252/wjsc.v18.i4.115218
Tendons are fibrous tissues that connect bones and muscles, and a tendon injury is damage to this fibrous tissue due to trauma, overuse, or strain[1]. Current clinical treatments for tendon injuries primarily include physical therapy, surgical treatment, and non-steroidal drug injection, which is limited to pain or symptom relief, based on the degree of injury[2,3]. However, multiple studies have considered stem cell-based treatment as a promising approach for tendon repair. Tendon stem/progenitor cells (TSPCs) have similar characteristics to bone marrow mesenchymal stem cells[4]. Studies have shown that TSPCs share common stem cell characteristics such as proliferation, pluripotency, and self-renewal ability[5,6]. Under suitable conditions, they can differentiate into tendon cells and play a pivotal role in maintaining tendons’ stability and repair after injury[7].
N6-methyladenosine (m6A) is the most prevalent internal modification in eukaryotic mRNA and non-coding RNAs, and it functions in regulating various aspects of RNA metabolism, including stability, translation, and degradation[8]. This modification is dynamically installed by “writer” proteins (methyltransferases) such as WTAP. WTAP, as a core component of the m6A methyltransferase complex, has been implicated in regulating the fate of stem cells and tissue development[9]. For example, WTAP-mediated m6A RNA methylation regulates the differentiation of bone marrow mesenchymal stem cells via the miR-29b-3p/HDAC4 axis[10]. However, the role of m6A modification, particularly WTAP-regulated m6A, on circular RNA (circRNA) in the tenogenic differentiation of TSPCs remains unexplored.
CircRNAs have a circular structure, and their expression is highly stable when compared with linear RNA, thus indicating that they may serve as potential therapeutic targets[11]. Moreover, circRNA plasmacytoma variant translocation 1 (circ_PVT1) is expressed from the broader non-coding RNA region of PVT1[12], and it has been confirmed to promote malignant progression or drug resistance of tumors in multiple cancers[13,14]. Recently, a study suggested that a decrease in circ_PVT1 could promote the senescence and inhibit the proliferation of TSPCs[15]. However, the regulatory mechanism of circ_PVT1 in the differentiation and tendon regeneration of TSPCs has not been fully elucidated. Therefore, we investigated the function of circ_PVT1 in the tenogenic differentiation of TSPCs. Circ_PVT1 knockdown inhibited transforming growth factor (TGF)-β1-induced TSPCs proliferation, migration, and tenogenic differentiation. However, circ_PVT1 inhibition also promoted apoptosis, oxidative stress, and mitochondrial damage in TSPCs. Moreover, the function of circ_PVT1 in TSPCs differentiation was regulated by WTAP-mediated m6A modification.
Mouse TSPCs were isolated from the Achilles tendon of 12 male C57Bl/6 mice at postnatal day 30 as previously described[16]. TSPCs were cultured in low-glucose Dulbecco’s modified Eagle’s medium (DMEM; Gibco, NY, United States) supplemented with 10% fetal bovine serum (FBS; Invitrogen, Carlsbad, CA, United States) and 1% penicillin-streptomycin (Gibco, NY, United States). Upon reaching 50% confluence, the TSPCs were maintained in low-glucose DMEM supplemented with 5% (vol/vol) FBS with 5 ng/mL of TGF-β1 (#HY-P7117, MCE, China) for 1 day, 3 days, or 7 days to induce tenogenic differentiation.
The mouse TSPCs were removed from the incubator (95% air and 5% CO2 at 37 °C) and washed three times with phosphate-buffered saline for 5 minutes each time. Then, 1 mL of 4% paraformaldehyde fixative was added and incubated at room temperature for 20 minutes. Subsequently, the fixative was aspirated off and discarded, and the remaining cells were washed three times with phosphate-buffered saline for 5 minutes each time. Then, the Picro-Sirius red (PSR) solution (Sigma-Aldrich, Darmstadt, Germany) was added, and the cells were stained for 15 minutes. The excess staining solution was then discarded, and distilled water was added to wash off the excess staining solution. The average positive cell rate was measured using ImageJ software (NIH, Bethesda, MD, United States).
The short hairpin RNAs (shRNAs) targeting circ_PVT1 or a negative control (NC), and shRNAs targeting WTAP or an NC, were constructed by Shanghai OBiO Technology Co., Ltd. The WTAP overexpression plasmid (WTAP-OE) and NC were inserted into the pcDNA3.1 vector (Invitrogen, Carlsbad, CA, United States) by subcloning to construct the WTAP-OE and control vectors, respectively. Subsequently, the plasmids were transfected into TSPCs cells using Lipofectamine 2000 transfection reagent (Invitrogen, Carlsbad, CA, United States) in accordance with the manufacturer’s instructions. Transfection efficiency was detected by quantitative polymerase chain reaction (qPCR) at 48 hours after transfection.
Total RNA was extracted from TSPCs using TRIzol reagent (Invitrogen, Carlsbad, CA, United States) and subsequently reverse-transcribed into cDNA with the NovoScript® Plus All-in-one 1st Strand cDNA Synthesis SuperMix (#E047-01A, Novoprotein, China). qPCR analysis was performed using NovoStart® SYBR qPCR SuperMix Plus (#E096-01B, Novoprotein, China) as per the manufacturer’s instructions, and GAPDH was used as the endogenous control. The gene expression levels were quantified using the 2-ΔΔCt method. The primer sequences (5’-3’) were designed as follows: Circ_PVT1, F: GCCTGGAGATGTTACAAGTAGC, R: GAAAGAGCAGAGAGCGAATGG; and GAPDH, F: TGGAAAGCTGTGGCGTGATG, R: CACTTGGCAGGTTTCTCCAGG.
The TSPCs were lysed using RIPA buffer (#MA0151, Meilunbio, China), and protein quantification was performed using a BCA assay kit (#MA0082-2, Meilunbio, China). Aliquots of equal amounts of protein were separated via sodium-dodecyl sulfate gel electrophoresis and subsequently transferred to polyvinylidene fluoride membranes (#BSP0160, PALL, NY, United States). After blocking with 5% skim milk, the membranes were incubated overnight at 4 °C with specific primary antibodies against GAPDH (1:50000, #60004-1-lg, Proteintech, IL, United States), scleraxis (SCX, 1:12000, #PA5-115874, Invitrogen, Carlsbad, CA, United States), tenomodulin (TNMD, 1:15000, bs-7525R, Bioss, Beijing, China), and COL3 (#22734-1-AP, 1:15000, Proteintech, IL, United States). This was followed by incubation with the appropriate horseradish peroxidase-conjugated secondary antibodies (#SA00001-2, 1:3000, Proteintech, IL, United States) for 1 hour. The protein bands were visualized using an enhanced chemiluminescence system (#MA0186-1, Meilunbio, China).
TSPCs were plated in 96-well plates at a cell density of 5 × 104 cells/well and cultured for 24 hours. Cell viability was assessed using Cell Counting Kit-8 reagent (#MA0218, Meilune, China), with 10 μL solution added to each well, followed by incubation for 1 hour at 37 °C in 5% CO2. Absorbance measurements at 450 nm were obtained using a microplate reader from three independent experimental replicates.
The TSPCs’ migration capacity was assessed using Corning Transwell chambers (Corning, NY, United States) according to the manufacturer’s protocol. Briefly, TSPCs (2 × 104 cells/200 μL serum-free medium) were added to the upper chamber, while the lower compartment contained DMEM supplemented with 10% FBS as a chemoattractant. The plate was then incubated at 37 °C in a 5% CO2 incubator for 24 hours to allow sufficient time for the TSPCs to migrate. The migrated cells were fixed with 4% paraformaldehyde (#C104188, Aladdin, China) for 15 minutes and then stained with 0.1% crystal violet (#A600331, Sangon, Shanghai, China) for a further 15 minutes. Finally, three random fields were selected using a microscope (#IX73, Olympus Corporation, Tokyo, Japan) for photography to evaluate the TSPCs’ migration ability.
TSPCs were seeded into 6-well plates and incubated overnight to allow a monolayer to form. Then, cells from the different treatment groups were incubated for a further 48 hours. The supernatant was carefully aspirated off and discarded, and cells were then resuspended in 250 μL of binding buffer, with the concentration adjusted to 1 × 106 cells/mL. Subsequently, the TSPCs were incubated with 5 μL of Annexin V-FITC and propidium iodide for 5 minutes. The stained cells were immediately analyzed using a CytoFlex flow cytometer (Beckman Coulter, CA, United States), and the data were analyzed accordingly.
TSPCs were loaded with 10 μM DCFH-DA (#HY-D0940, MedChemExpress, Shanghai, China) that was prepared in serum-free medium (1:1000 dilution). Following incubation at 37 °C for 30 minutes under light-protected conditions with intermittent gentle mixing at 5-minute intervals, cellular fluorescence was quantified via flow cytometry with 488 nm excitation and 525 nm emission parameters.
Malondialdehyde (#A003-1, Jiancheng Bioengineering, China), glutathione (#A006-2-1, Jiancheng Bioengineering, China), superoxide dismutase (#A001-3, Jiancheng Bioengineering, China), and ATP (#S0026, Beyotime, China) kits were used to detect the levels of the corresponding biochemical indicators in mouse TSPCs, following the manufacturers’ instructions.
TSPCs were stained with 5 μmol/L JC-1 (#C2006, Beyotime, China) for 30 minutes under light-protected conditions. After washing and resuspension, fluorescence microscopy was performed to quantify the JC-1 polymers (red) and monomers (green). Mitochondrial membrane potential (MMP)-dependent redistribution of the dye was assessed based on its dual fluorescence properties, where JC-1 forms red fluorescent polymers in mitochondria with intact MMP but shifts to green monomers upon MMP depolarization[17].
RNA samples from TSPCs were quantified and standardized to working concentrations (300 ng/μL and 600 ng/μL), then heat-denatured at 95 °C for 5 minutes using a metal bath, followed by immediate ice quenching for 10 minutes. The denatured RNA was diluted 1:1 with 20 × SSC buffer and applied to nitrocellulose membranes in a dot-blot configuration. Membranes underwent ultraviolet crosslinking for 15 minutes and were briefly immersed in aqua blue protein-binding stain for 2 minutes, followed by three washes with TBS-T for 5 minutes each time before visualization. Su
A Pierce Magnetic RNA Protein Pull-Down Kit (#20164, Thermo Scientific, Waltham, MA, United States) was employed to investigate the potential interaction between circ_PVT1 and WTAP. Biotinylated RNA probes were incubated with the TSPC-derived protein lysates and streptavidin-conjugated magnetic beads for 2 hours at 25 °C under rotational conditions. Following three washes with high-stringency buffer (150 mmol/L NaCl, 0.1% NP-40), the bead-bound protein complexes were eluted and subjected to immunoblotting for WTAP detection.
RNA-protein complexes were isolated using an RNA Immunoprecipitation Kit (#Bes5101, BersinBio, China). The TSPCs lysates were incubated with anti-WTAP antibody (1:200; #10200-1-AP, Proteintech, IL, United States) or IgG control (#A0208, Beyotime, China) that were pre-conjugated to protein A/G agarose beads for 12 hours at 4 °C. After washing four times with RIP buffer, RNA was liberated by proteinase K digestion for 2 hours at 55 °C and then quantified via qPCR.
RNA methylation profiling was performed using the methylated RNA immunoprecipitation PCR (MeRIP) m6A Transcriptome Profiling Kit (#C11051-1, RiBo Bio, Guangzhou, Guangdong Province, China). The TRIzol-extracted RNA (#15596018CN, Invitrogen, Carlsbad, CA, United States) was fragmented at 94 °C for 3 minutes and immunoprecipitated with 5 μg of anti-m6A antibody-bound magnetic beads at 4 °C for 2 hours. Following elution (0.1 M DTT, 1% sodium-dodecyl sulfate), the m6A-enriched RNA fragments were analyzed by qPCR using gene-specific primers.
Data were analyzed in GraphPad Prism 9.0.1 and presented as the mean ± SD. Intergroup differences were assessed using the Student’s t-test (two-group comparisons) or a one-way analysis of variance with Tukey’s post hoc (multi-group comparisons). Statistical significance was defined as P < 0.05.
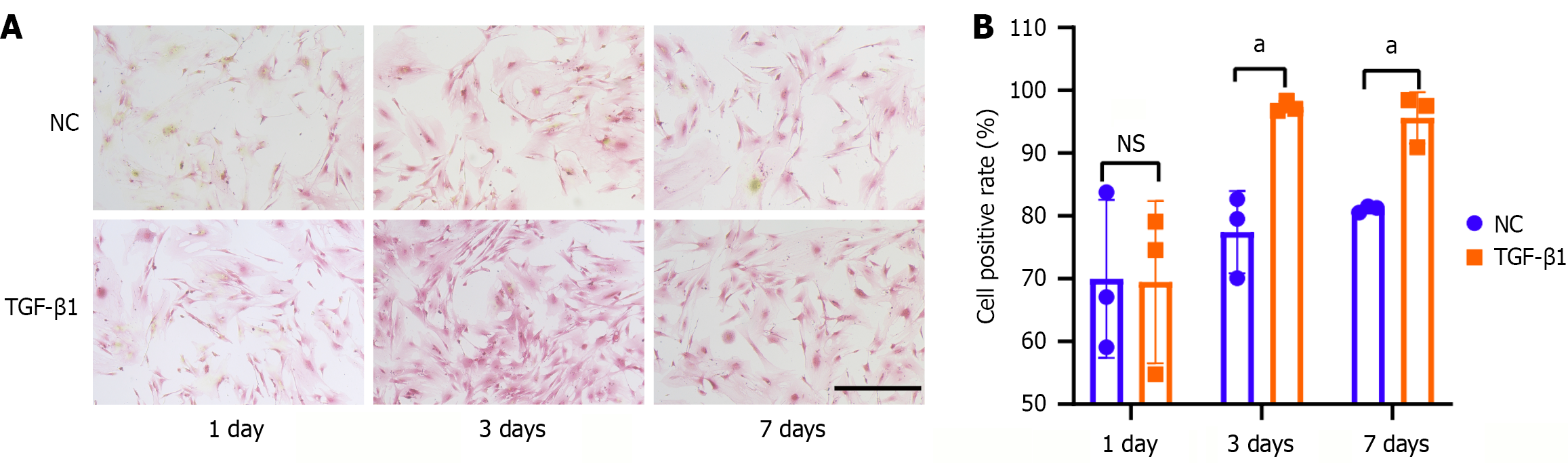
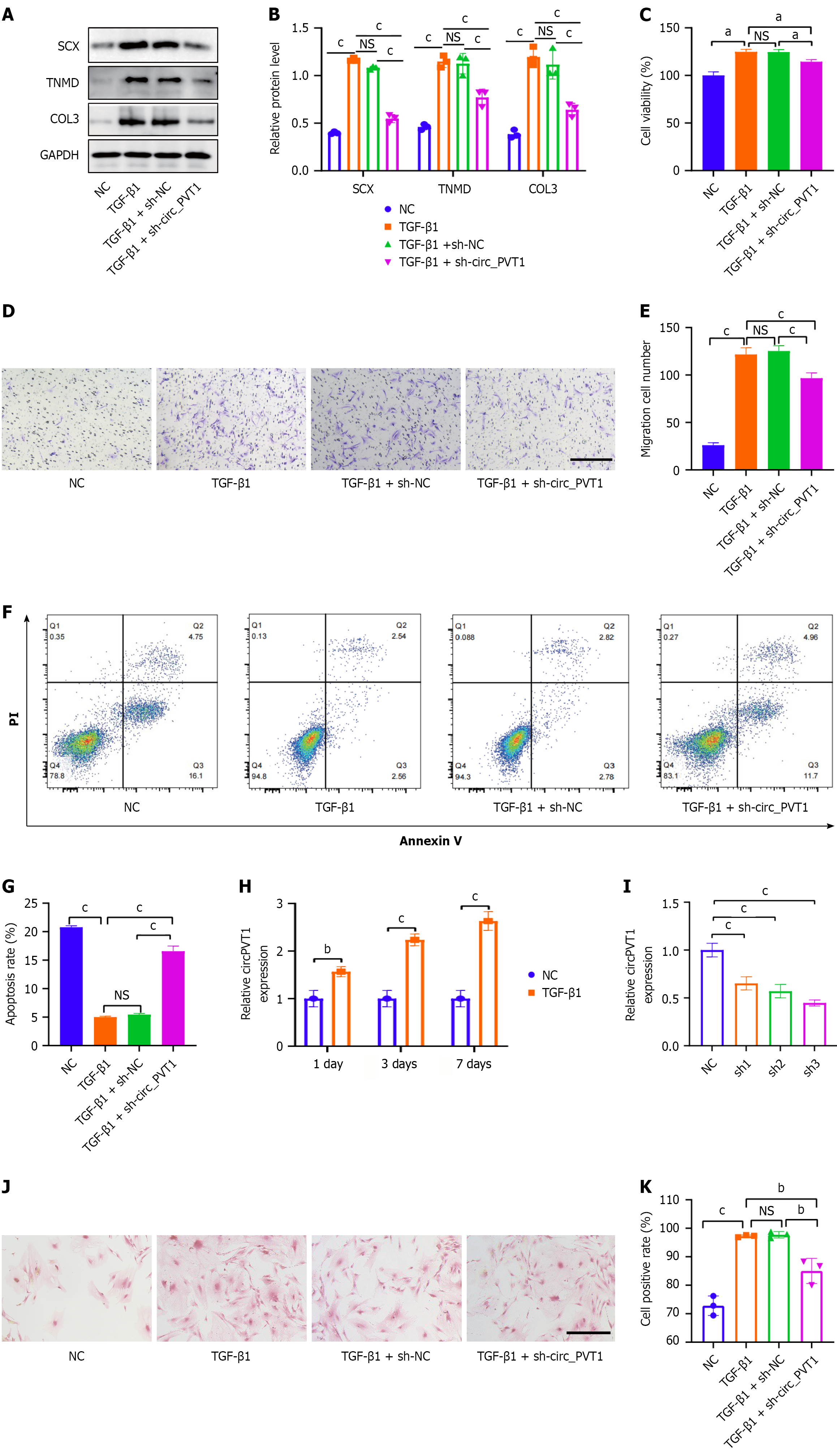
TSPCs were treated with 5 ng/mL of TGF-β1 to induce tendon differentiation. Then, PSR staining was used to detect the TSPCs’ collagen content, as collagen is the main component of tendons. The results showed that the rate of red positive cells was higher than that of the normal control after TGF-β1 treatment at 3 days and 7 days (Figure 1). In addition, the western blot analysis results revealed that the tendon-specific markers SCX, TNMD, and COL3 were elevated after TGF-β1 treatment compared with those of the normal control (Figure 2A and B). The Cell Counting Kit-8 assay exhibited that TGF-β1 treatment promoted the proliferation of TSPCs compared with the normal control (Figure 2C). Moreover, TGF-β1 increased the migration (Figure 2D and E) and decreased the apoptosis (Figure 2F and G) of TSPCs compared to those of the normal control. These results indicate that TGF-β1 treatment induced tenogenic differentiation by promoting the expression of tendon-specific protein markers as well as the proliferation and migration of TSPCs.
After treatment with 5 ng/mL of TGF-β1 for 1 day, 3 days, or 7 days, the expression of TSPCs circ_PVT1 was significantly increased compared with the normal control (Figure 2H). The TGF-β1-induced TSPCs at 3 days were used for the follow-up study. Three shRNAs (shRNA1, shRNA2, and shRNA3) of circ_PVT1 were transfected into the TSPCs, and the expression of circ_PVT1 was decreased compared with the shNC in TSPCs. shRNA3 exhibited the highest knockdown efficiency and was thus used for the functional study of circ_PVT1 (Figure 2I). The transfected TSPCs were then treated with TGF-β1 for 3 days. We found that circ_PVT1 knockdown in the TGF-β1-treated TSPCs decreased the rate of PSR-stained positive cells compared with the TGF-β1 + shNC group (Figure 2J and K). In addition, circ_PVT1 knockdown in the TGF-β1-treated TSPCs reversed the increase of tendon-specific proteins (Figure 2A and B), cell proliferation (Figure 2C), and migration (Figure 2D and E) that were induced by TGF-β1 treatment. In addition, circ_PVT1 knockdown in the TGF-β1-treated TSPCs reversed the decrease in TSPCs apoptosis compared with the TGF-β1 + shNC group (Figure 2F and G).
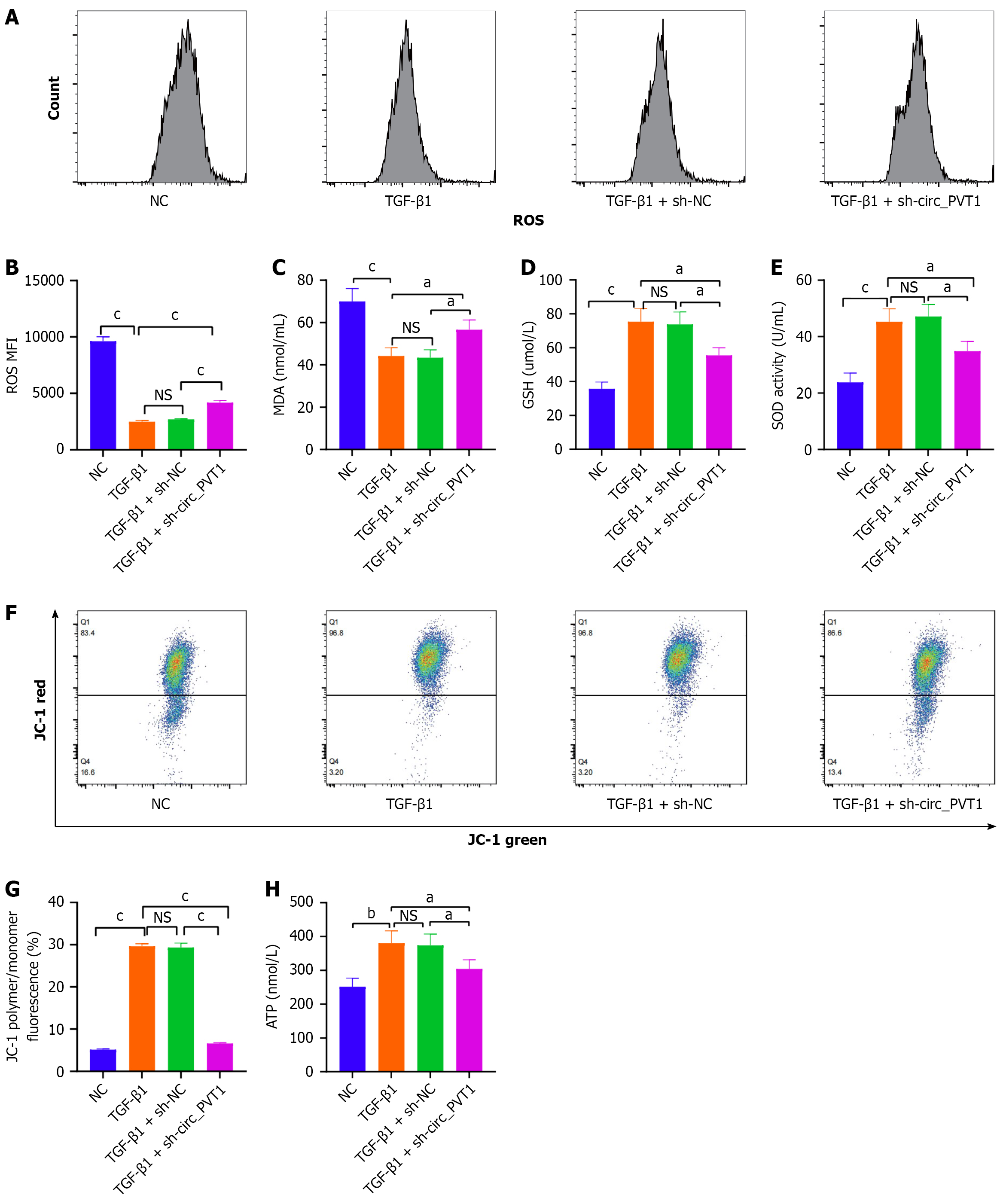
Next, the effect of circ_PVT1 on oxidative stress and mitochondrial damage was investigated. The results revealed that circ_PVT1 knockdown reversed the decrease of reactive oxygen species (Figure 3A and B) and malondialdehyde (Figure 3C), but decreased the glutathione concentration (Figure 3D) and superoxide dismutase activity (Figure 3E) in TGF-β1-induced TSPCs. In addition, circ_PVT1 knockdown significantly changed the MMP. The percentage of JC-1 polymer/monomer fluorescence and ATP production was reversed by sh-circ_PVT1 treatment in TGF-β1-induced TSPCs (Figure 3F-H).
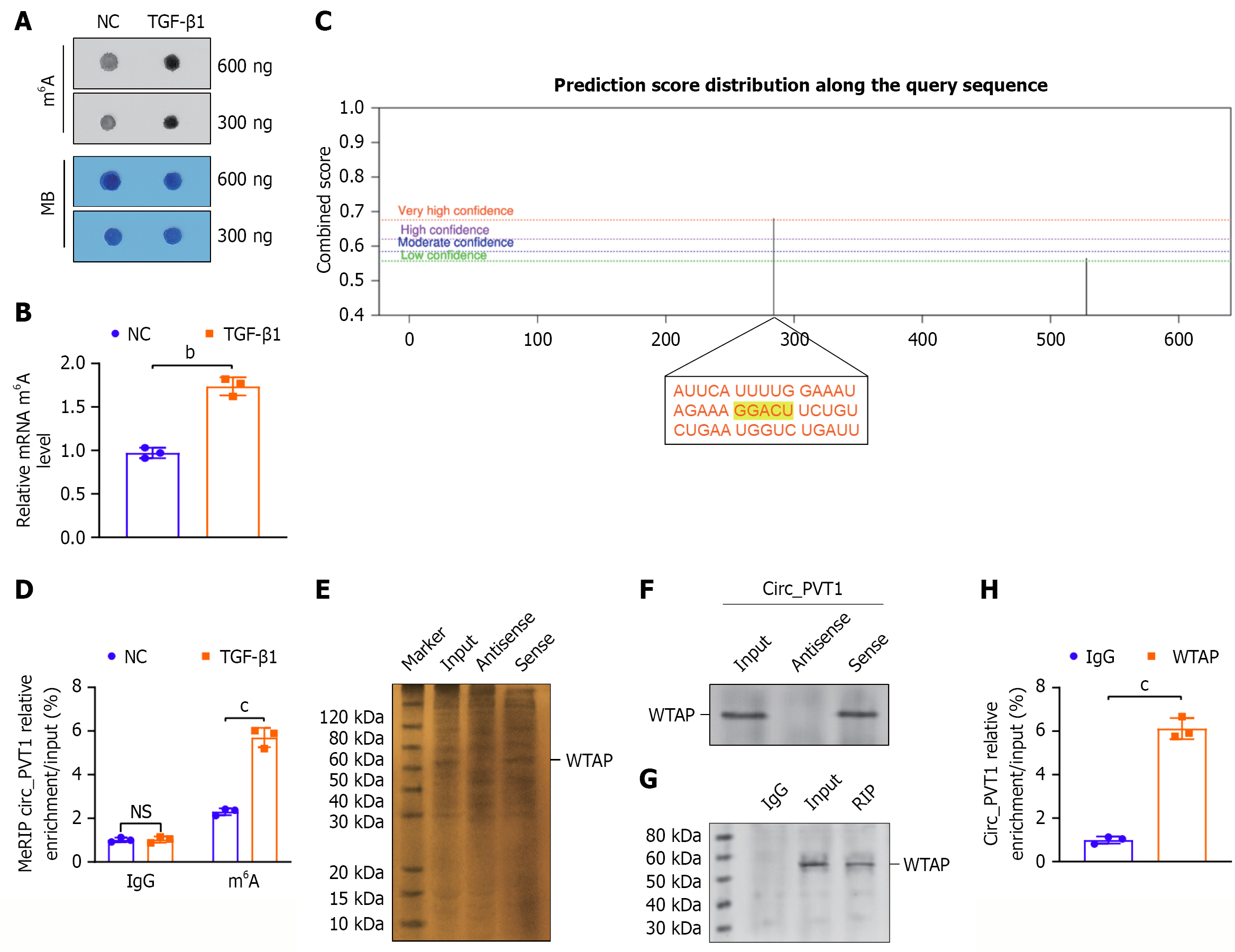
Subsequently, we investigated the potential mechanism of circ_PVT1 in tenogenic differentiation. We analyzed the m6A modification level of circ_PVT1 in TGF-β1-induced TSPCs, and the m6A dot blot assay revealed that the total m6A level was upregulated (Figure 4A and B). The m6A modification site prediction software SRAMP analysis indicated several potential m6A modification sites in the 3’-untranslated region of circ_PVT1, including a GGACU site with very high probability (Figure 4C). In addition, the meRIP-qPCR assay indicated that the m6A level of circ_PVT1 was upregulated in TGF-β1-induced TSPCs (Figure 4D). An RNA pulldown assay was used to analyze the potential m6A regulatory factor, and western blot analysis showed that circ_PVT1 bound to WTAP (Figure 4E and F). Moreover, the RIP-PCR results confirmed that circ_PVT1 was captured by anti-WTAP antibody in TSPCs (Figure 4G and H).
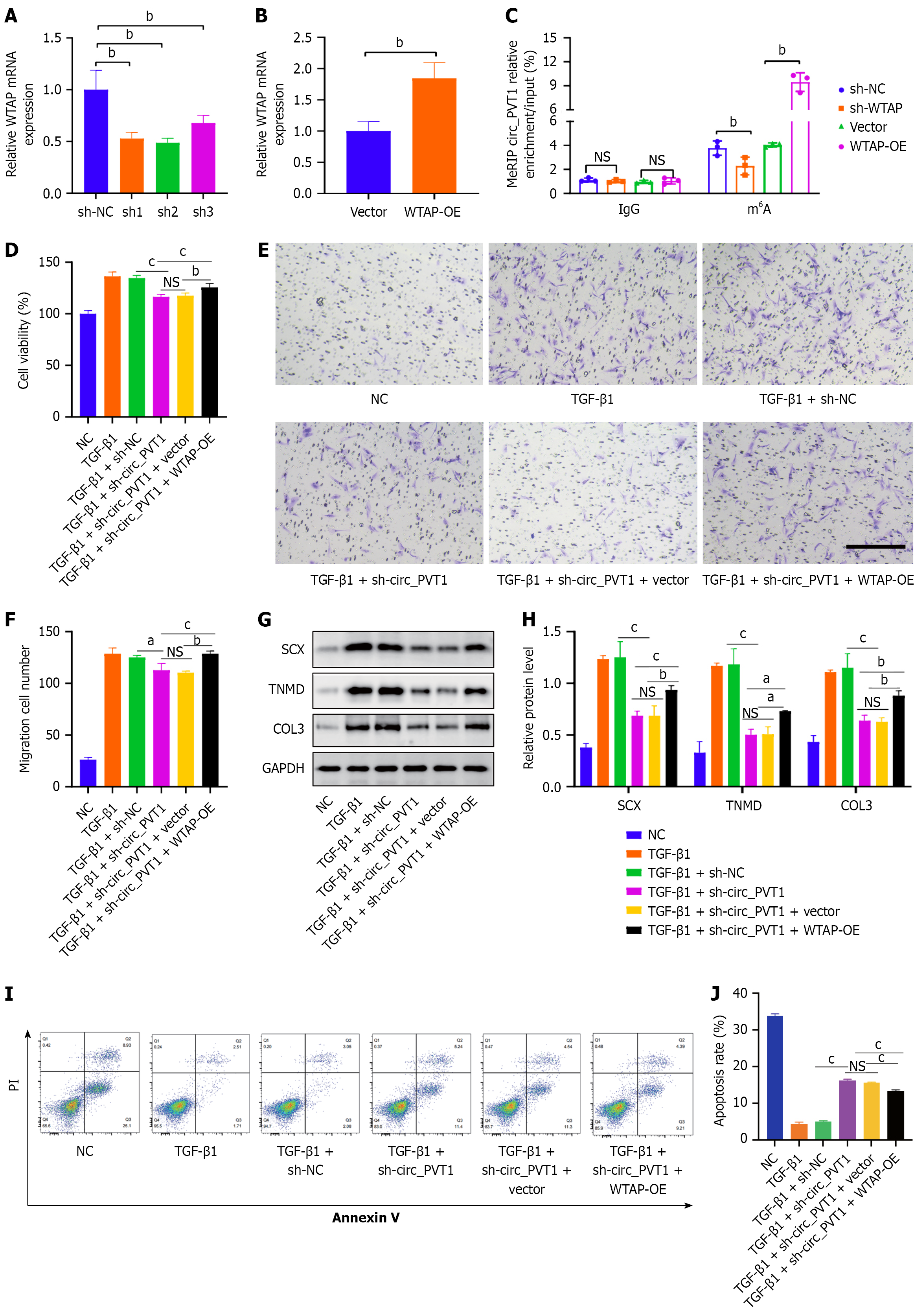
We then explored whether WTAP affects the function of circ_PVT1. Three shRNAs targeting WTAP (sh1WTAP, sh2WTAP, and sh3WTAP) were transfected into the TSPCs, and all decreased the expression of WTAP compared with that of shNC. sh2WTAP exhibited the best efficiency (Figure 5A). Moreover, we confirmed the overexpression efficiency of WTAP-OE (Figure 5B). In addition, the meRIP-qPCR assay showed that the m6A level of circ_PVT1 was upregulated with WTAP-OE and downregulated with WTAP knockdown in TGF-β1-induced TSPCs (Figure 5C). Furthermore, we found that WTAP-OE could reverse the cell proliferation (Figure 5D), cell migration (Figure 5E and F), and reduction of tendon-specific protein markers SCX, TNMD, and COL3 (Figure 5G and H) caused by circ_PVT1 inhibition in TGF-β1-induced TSPCs, as well as reverse the increased apoptosis (Figure 5I and J).
Existing studies have demonstrated that stem cell-based therapy plays an important role in the repair of tendon injuries[18,19]. Stem cells can differentiate into tendon cells and secrete various cytokines that regulate inflammation and tissue healing[20,21]. TSPCs are a novel type of stem cell with the ability to proliferate, self-renew, and differentiate into multiple lines, and are regarded as the ideal cell types for tendon injury regeneration and repair[7,22]. In the repair of tendon injuries, the proliferation, migration, and differentiation of cells are essential for the repair process[23]. Herein, TGF-β1 was used to induce tenogenic differentiation in TSPCs. TGF-β1 treatment induced the proliferation and migration of TSPCs and enhanced the expression of tendon differentiation-specific proteins SCX, TNMD, and COL3, as well as collagen. Thereafter, we found that the expression level of circ_PVT1 increased in the TGF-β1-treated TSPCs. Circ_PVT1 knockdown reversed the increase of tendon-specific proteins, cell proliferation, and migration induced by TGF-β1 treatment.
CircRNAs have been reported to modulate gene expression and promote tissue repair or regeneration. For example, circ_CDK13 can promote wound healing and skin appendage regeneration in type I diabetic rats[24]. Circ_Samd4 induces myocardial tissue repair after myocardial infarction by inhibiting the generation of mitochondria-derived reactive oxygen species[25]. Circ_CDR1as could promote nerve regeneration after spinal cord injury[26]. In addition, circ_0005736 promotes TGF-β1-induced TSPCs tenogenic differentiation via the miR-636/mitogen-activated protein kinases 1 axis[27]. Han et al[15] reported that circ_PVT1 plays a crucial role in inhibiting TSPCs senescence and promoting self-renewal, migration, and tenogenic differentiation. In this study, we found that circ_PVT1 knockdown inhibits the proliferation and migration and decreases the expression of tendon differentiation-specific proteins SCX, TNMD, and COL3, as well as collagen. These results indicated that circ_PVT1 knockdown inhibited tenogenic differentiation by decreasing the expression of tendon-specific protein markers and TSPCs proliferation and migration. These results add a crucial new layer to this narrative by revealing that the functional efficacy of a circRNA can be potentiated by m6A, which is a regulatory axis that has been underexplored in tendon biology.
M6A modification is an important regulatory factor for the production and function of mRNA and non-coding RNA[28]. Previous studies have shown that these circRNAs undergo m6A modification, and the m6A sites have been confirmed to initiate circRNA translation[29]. The inhibition of m6A-regulated circ_ZNF609 improves doxorubicin-induced cardiotoxicity by upregulating FTO[30]. However, the potential m6A modulator regulatory mechanism of circRNAs and the potential function of circRNAs in the mechanism of tenogenic differentiation remain unclear. In this study, we found that the total m6A level and the m6A level of circ_PVT1 were upregulated in the TGF-β1-treated TSPCs. A methyltransferase, WTAP, was confirmed to interact with circ_PVT1. WTAP overexpression reversed the cell proliferation, cell migration, and the reduction of tendon-specific protein markers caused by the circ_PVT1 inhibition of TGF-β1-induced TSPCs, as well as reversed the increase of apoptosis. These results indicate that WTAP is the key “writer” that catalyzes the m6A modification of circ_PVT1 in TSPCs. This finding aligns with the emerging role of WTAP in regulating cell fate determination via m6A deposition on specific RNA targets. For instance, WTAP-mediated m6A modification has been shown to promote the progression of osteosarcoma by regulating circ_0032463[31]. Our results extend this concept by demonstrating that WTAP’s function in modifying circRNAs is also critical in the context of tenogenic differentiation, thereby highlighting a conserved mechanism across different biological processes.
Despite these promising findings, this study has several limitations. First, all experiments were performed using mouse-derived TSPCs. Although these cells share fundamental characteristics with human TSPCs, the absence of validation in human primary tendon cells or tissues may limit the direct clinical applicability of our results. Future studies should incorporate human TSPCs or in vivo humanized models to enhance the translational relevance. Second, the mechanistic insights, while supportive of WTAP-mediated m6A modification in regulating circ_PVT1, do not fully elucidate the downstream signaling pathways through which m6A-modified circ_PVT1 exerts its effects on tenogenic differentiation.
In conclusion, this study demonstrated that circ_PVT1 was upregulated in TGF-β1-induced TSPCs and that circ_PVT1 knockdown could inhibit tenogenic differentiation. Furthermore, the WTAP-mediated m6A modification of circ_PVT1 promotes the proliferation and tenogenic differentiation of TSPCs. These results indicate a promising therapeutic strategy for tendon diseases.
| 1. | Thomopoulos S, Parks WC, Rifkin DB, Derwin KA. Mechanisms of tendon injury and repair. J Orthop Res. 2015;33:832-839. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 299] [Cited by in RCA: 406] [Article Influence: 36.9] [Reference Citation Analysis (0)] |
| 2. | Liu Y, Suen CW, Zhang JF, Li G. Current concepts on tenogenic differentiation and clinical applications. J Orthop Translat. 2017;9:28-42. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 32] [Cited by in RCA: 48] [Article Influence: 5.3] [Reference Citation Analysis (0)] |
| 3. | Malmgaard-Clausen NM, Kjaer M, Dakin SG. Pathological Tendon Histology in Early and Chronic Human Patellar Tendinopathy. Transl Sports Med. 2022;2022:2799665. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 16] [Article Influence: 4.0] [Reference Citation Analysis (0)] |
| 4. | Guo J, Chan KM, Zhang JF, Li G. Tendon-derived stem cells undergo spontaneous tenogenic differentiation. Exp Cell Res. 2016;341:1-7. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 44] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 5. | Walia B, Huang AH. Tendon stem progenitor cells: Understanding the biology to inform therapeutic strategies for tendon repair. J Orthop Res. 2019;37:1270-1280. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 84] [Cited by in RCA: 73] [Article Influence: 10.4] [Reference Citation Analysis (1)] |
| 6. | He Y, Lu S, Chen W, Yang L, Li F, Zhou P, Chen Z, Wan R, Zhang Z, Sun Y, Lin J, Chen Y, Luo Z, Xu C, Chen S. Exosomes derived from tendon stem/progenitor cells enhance tendon-bone interface healing after rotator cuff repair in a rat model. Bioact Mater. 2024;40:484-502. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1] [Cited by in RCA: 21] [Article Influence: 10.5] [Reference Citation Analysis (0)] |
| 7. | Bi Y, Ehirchiou D, Kilts TM, Inkson CA, Embree MC, Sonoyama W, Li L, Leet AI, Seo BM, Zhang L, Shi S, Young MF. Identification of tendon stem/progenitor cells and the role of the extracellular matrix in their niche. Nat Med. 2007;13:1219-1227. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1241] [Cited by in RCA: 1083] [Article Influence: 57.0] [Reference Citation Analysis (0)] |
| 8. | Chen X, Pu S, Lian K, Li L, Jiang X. m6A RNA modification in tumor-associated macrophages: emerging roles in cancer immunity. Front Immunol. 2025;16:1693336. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 7] [Reference Citation Analysis (0)] |
| 9. | Du Y, Huang L, Long M, Wu X, Miao J. WTAP-mediated abnormal m6A modification promotes cancer progression by remodeling the tumor microenvironment: bibliometric and database analyses. Transl Cancer Res. 2024;13:952-974. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 2.5] [Reference Citation Analysis (0)] |
| 10. | Liu J, You Y, Sun Z, Zhang L, Li X, Dai Z, Ma J, Chen Y, Jiao G. WTAP-Mediated m6A RNA Methylation Regulates the Differentiation of Bone Marrow Mesenchymal Stem Cells via the miR-29b-3p/HDAC4 Axis. Stem Cells Transl Med. 2023;12:307-321. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 30] [Reference Citation Analysis (0)] |
| 11. | Liu X, Zhang Y, Zhou S, Dain L, Mei L, Zhu G. Circular RNA: An emerging frontier in RNA therapeutic targets, RNA therapeutics, and mRNA vaccines. J Control Release. 2022;348:84-94. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 120] [Cited by in RCA: 267] [Article Influence: 66.8] [Reference Citation Analysis (0)] |
| 12. | Kun-Peng Z, Xiao-Long M, Chun-Lin Z. Overexpressed circPVT1, a potential new circular RNA biomarker, contributes to doxorubicin and cisplatin resistance of osteosarcoma cells by regulating ABCB1. Int J Biol Sci. 2018;14:321-330. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 169] [Cited by in RCA: 230] [Article Influence: 28.8] [Reference Citation Analysis (17)] |
| 13. | Yi J, Wang L, Hu GS, Zhang YY, Du J, Ding JC, Ji X, Shen HF, Huang HH, Ye F, Liu W. CircPVT1 promotes ER-positive breast tumorigenesis and drug resistance by targeting ESR1 and MAVS. EMBO J. 2023;42:e112408. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 70] [Reference Citation Analysis (1)] |
| 14. | Palcau AC, Canu V, Donzelli S, Strano S, Pulito C, Blandino G. CircPVT1: a pivotal circular node intersecting Long Non-Coding-PVT1 and c-MYC oncogenic signals. Mol Cancer. 2022;21:33. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 6] [Cited by in RCA: 46] [Article Influence: 11.5] [Reference Citation Analysis (0)] |
| 15. | Han W, Tao X, Weng T, Chen L. Circular RNA PVT1 inhibits tendon stem/progenitor cell senescence by sponging microRNA-199a-5p. Toxicol In Vitro. 2022;79:105297. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 4] [Cited by in RCA: 17] [Article Influence: 3.4] [Reference Citation Analysis (0)] |
| 16. | Mienaltowski MJ, Adams SM, Birk DE. Regional differences in stem cell/progenitor cell populations from the mouse achilles tendon. Tissue Eng Part A. 2013;19:199-210. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 112] [Cited by in RCA: 89] [Article Influence: 6.8] [Reference Citation Analysis (0)] |
| 17. | Smiley ST, Reers M, Mottola-Hartshorn C, Lin M, Chen A, Smith TW, Steele GD Jr, Chen LB. Intracellular heterogeneity in mitochondrial membrane potentials revealed by a J-aggregate-forming lipophilic cation JC-1. Proc Natl Acad Sci U S A. 1991;88:3671-3675. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 1162] [Cited by in RCA: 1227] [Article Influence: 35.1] [Reference Citation Analysis (2)] |
| 18. | Ahn SY. Various Strategies of Tendon Stem/Progenitor Cell Reprogramming for Tendon Regeneration. Int J Mol Sci. 2024;25:11745. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (4)] |
| 19. | Yin H, Mao K, Huang Y, Guo A, Shi L. Tendon stem/progenitor cells are promising reparative cell sources for multiple musculoskeletal injuries of concomitant articular cartilage lesions associated with ligament injuries. J Orthop Surg Res. 2023;18:869. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 4] [Reference Citation Analysis (0)] |
| 20. | Migliorini F, Tingart M, Maffulli N. Progress with stem cell therapies for tendon tissue regeneration. Expert Opin Biol Ther. 2020;20:1373-1379. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 16] [Cited by in RCA: 42] [Article Influence: 7.0] [Reference Citation Analysis (0)] |
| 21. | Gaspar D, Spanoudes K, Holladay C, Pandit A, Zeugolis D. Progress in cell-based therapies for tendon repair. Adv Drug Deliv Rev. 2015;84:240-256. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 113] [Cited by in RCA: 141] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 22. | Wu YF, Chen C, Tang JB, Mao WF. Growth and Stem Cell Characteristics of Tendon-Derived Cells with Different Initial Seeding Densities: An In Vitro Study in Mouse Flexor Tendon Cells. Stem Cells Dev. 2020;29:1016-1025. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 8] [Cited by in RCA: 14] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 23. | Chen Q, Liang Q, Zhuang W, Zhou J, Zhang B, Xu P, Ju Y, Morita Y, Luo Q, Song G. Tenocyte proliferation and migration promoted by rat bone marrow mesenchymal stem cell-derived conditioned medium. Biotechnol Lett. 2018;40:215-224. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 21] [Cited by in RCA: 30] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 24. | Huang Q, Chu Z, Wang Z, Li Q, Meng S, Lu Y, Ma K, Cui S, Hu W, Zhang W, Wei Q, Qu Y, Li H, Fu X, Zhang C. circCDK13-loaded small extracellular vesicles accelerate healing in preclinical diabetic wound models. Nat Commun. 2024;15:3904. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 42] [Reference Citation Analysis (0)] |
| 25. | Zheng H, Huang S, Wei G, Sun Y, Li C, Si X, Chen Y, Tang Z, Li X, Chen Y, Liao W, Liao Y, Bin J. CircRNA Samd4 induces cardiac repair after myocardial infarction by blocking mitochondria-derived ROS output. Mol Ther. 2022;30:3477-3498. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 22] [Cited by in RCA: 118] [Article Influence: 29.5] [Reference Citation Analysis (0)] |
| 26. | Wang W, Liu C, He D, Shi G, Song P, Zhang B, Li T, Wei J, Jiang Y, Ma L. CircRNA CDR1as affects functional repair after spinal cord injury and regulates fibrosis through the SMAD pathway. Pharmacol Res. 2024;204:107189. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5] [Cited by in RCA: 15] [Article Influence: 7.5] [Reference Citation Analysis (0)] |
| 27. | Yang G, Chen F, Zhang C, Gu C. Circ_0005736 promotes tenogenic differentiation of tendon-derived stem cells through the miR-636/MAPK1 axis. J Orthop Surg Res. 2023;18:660. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 3] [Reference Citation Analysis (0)] |
| 28. | Chen Y, Lin Y, Shu Y, He J, Gao W. Interaction between N(6)-methyladenosine (m(6)A) modification and noncoding RNAs in cancer. Mol Cancer. 2020;19:94. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 87] [Cited by in RCA: 211] [Article Influence: 35.2] [Reference Citation Analysis (0)] |
| 29. | Yang Y, Fan X, Mao M, Song X, Wu P, Zhang Y, Jin Y, Yang Y, Chen LL, Wang Y, Wong CC, Xiao X, Wang Z. Extensive translation of circular RNAs driven by N(6)-methyladenosine. Cell Res. 2017;27:626-641. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1591] [Cited by in RCA: 1511] [Article Influence: 167.9] [Reference Citation Analysis (4)] |
| 30. | Yu P, Wang J, Xu GE, Zhao X, Cui X, Feng J, Sun J, Wang T, Spanos M, Lehmann HI, Li G, Xu J, Wang L, Xiao J. RNA m(6)A-Regulated circ-ZNF609 Suppression Ameliorates Doxorubicin-Induced Cardiotoxicity by Upregulating FTO. JACC Basic Transl Sci. 2023;8:677-698. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 35] [Reference Citation Analysis (0)] |
| 31. | Huang Z, Chen P, Liu Y. WTAP-mediated m6A modification of circ_0032463 promotes osteosarcoma progression by sponging miR-145-5p and regulating GFRA1 expression. J Biochem Mol Toxicol. 2024;38:e23833. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 5] [Reference Citation Analysis (0)] |