Published online May 26, 2024. doi: 10.4252/wjsc.v16.i5.604
Revised: March 6, 2024
Accepted: April 19, 2024
Published online: May 26, 2024
Processing time: 152 Days and 15.5 Hours
Gliomas pose a significant challenge to effective treatment despite advancements in chemotherapy and radiotherapy. Glioma stem cells (GSCs), a subset within tumors, contribute to resistance, tumor heterogeneity, and plasticity. Recent studi
To analyze targeted therapies against GSC-mediated resistance to radio- and che
A systematic search was conducted across major medical databases (PubMed, Embase, and Cochrane Library) up to September 30, 2023. The search strategy utilized relevant Medical Subject Heading terms and keywords related to in
In a comprehensive review of 66 studies on stem cell therapies for SCI, 452 papers were initially identified, with 203 chosen for full-text analysis. Among them, 201 were deemed eligible after excluding 168 for various reasons. The temporal breakdown of studies illustrates this trend: 2005-2010 (33.3%), 2011-2015 (36.4%), and 2016-2022 (30.3%). Key GSC models, particularly U87 (33.3%), U251 (15.2%), and T98G (15.2%), emerge as significant in research, reflecting their representativeness of glioma characteristics. Pathway analysis indicates a focus on phos
GSCs play a complex role in mediating radioresistance and chemoresistance, emphasizing the necessity for pre
Core Tip: The challenge of treating gliomas persists despite advancements in chemotherapy and radiotherapy, with glioma stem cells (GSCs) contributing to resistance and tumor heterogeneity. This systematic literature review, covering 66 studies, underscores the intricate role of GSCs in therapeutic resistance, particularly highlighting their involvement in DNA repair mechanisms and dynamic cellular state transitions. Targeted therapies face challenges due to GSCs’ high plasticity, and the review emphasizes the need for precision treatments that account for GSC population heterogeneity and the tumor microenvironment’s dynamic nature. Versatile therapeutic agents, including RNA inhibitor/short hairpin RNA, inhibitors (e.g., LY294002, NVP-BEZ235), and monoclonal antibodies (e.g., cetuximab), demonstrate efficacy in overcoming GSC-mediated resistance. Notably, the most common effect on the chemo- and radiotherapy response is a reduction in temozolomide resistance, highlighting the potential for improved outcomes by disrupting GSC-mediated resistance mechanisms in glioblastoma patients.
- Citation: Agosti E, Zeppieri M, Ghidoni M, Ius T, Tel A, Fontanella MM, Panciani PP. Role of glioma stem cells in promoting tumor chemo- and radioresistance: A systematic review of potential targeted treatments. World J Stem Cells 2024; 16(5): 604-614
- URL: https://www.wjgnet.com/1948-0210/full/v16/i5/604.htm
- DOI: https://dx.doi.org/10.4252/wjsc.v16.i5.604
Gliomas, one of the most aggressive and prevalent form of primary brain tumors, continues to present a formidable challenge to effective therapeutic intervention[1]. Despite advancements in treatment modalities such as radiotherapy resistance (RTR) and chemotherapy resistance (CTR) with temozolomide (TMZ), the prognosis for glioma patients remains bleak, with a median overall survival of around 15 months[2-4]. A major obstacle in achieving better outcomes is the intrinsic resistance of gliomas to conventional adjuvant treatments[5]. In recent years, the focus has shifted towards understanding the role of glioma stem cells (GSCs) in fostering resistance to adjuvant therapies, unraveling a complex interplay between cancer stem cells and treatment response[4,6].
Central to the enigma of glioma resistance is the discovery that GSCs, a subpopulation within the heterogeneous tumor mass, contribute not only to tumor initiation and maintenance but also to tumor heterogeneity and plasticity. It is now known that GSCs, characterized by their self-renewal capacity and multilineage differentiation potential, are emerging as key players in the intricate landscape of glioma progression[4,5]. The ability of GSCs to generate different cellular states, driven by intrinsic and extrinsic factors, results in a dynamic equilibrium of heterogeneous cell populations within the tumor microenvironment[2,3].
Therapeutic resistance in gliomas is a multifaceted challenge. Gliomas possess an innate ability to adapt to therapies, and, as postulated by recent studies, this resistance is attributed in part to GSCs[5]. The traditional understanding of resistance mechanisms involves the activation of DNA repair mechanisms, especially in the context of TMZ and ionizing radiation[6]. However, recent studies proposed a spectrum of recurrence patterns, suggesting that resistance can stem from preexisting chemo-resistant clones or treatment-induced changes in cell populations[5]. Indeed, GSCs, equipped with efficient DNA damage repair systems, exhibit heightened resistance to chemotherapy, particularly TMZ, and RTR[2]. The cytotoxic effects of RTR and TMZ, mediated through O6-methylguanine-methyltransferase, are mitigated in GSCs due to increased expression of O6-methylguanine-methyltransferase and other anti-apoptotic proteins. This resistance is further compounded by the ability of GSCs to transition between different cellular states, confounding the efficacy of conventional therapies[4,5].
Despite significant strides in understanding the role of GSCs in therapeutic resistance, there is a lack of consensus on targeted therapeutic approaches[6]. The high plasticity of GSCs and their dynamic transitions between cellular states present a formidable challenge for devising curative strategies[5]. TMZ exposure, while stimulating the conversion of differentiated tumor cells into GSCs, underscores the need for tailored therapeutic interventions that specifically target GSCs[2,3].
In this context, a systematic literature review becomes imperative to consolidate the diverse findings and discern a common line of treatment against GSC-mediated resistance. This systematic literature review aims to analyze the current landscape of targeted therapies addressing chemo- and radioresistance in gliomas, with a specific focus on the mecha
In accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analysis (PRISMA) criteria[7], a systematic evaluation of targeted therapeutics targeting the molecular mechanism of GSC-mediated resistance to adjuvant therapy was conducted[8]. A methodical and thorough literature search of the PubMed, Web of Science, Cochrane, Reference Citation Analysis (RCA) (https://www.referencecitationanalysis.com) and Embase databases was conducted by two authors (Agosti E and Ghidoni M). On September 30, 2023, the initial literature search was carried out; on December 10, 2023, the search was updated. To create a search strategy, many keyword searches were conducted. Combinations of AND and OR were employed to find results for the search terms “glioma stem cells”, “radiotherapy”, “chemotherapy”, “resistance”, and “targeted therapies”. The Medical Subject Heading terms and Boolean operators (“glioma stem cells” OR “GSC” OR “cancer stem cells” OR “CSC”) AND (“chemotherapy” OR “radiotherapy” OR “temozolomide” OR “adjuvant treatments”) AND (“resistance” OR “resilience”) AND (“targeted therapy” OR “targeted treatment” OR “targeted strategy”). These terms were used to retrieve studies. A reference analysis of a few chosen publications revealed other relevant articles. A search filter was applied to display only articles from the specified time frame of 2000-2023. Following that, a further search on https://clinicaltrials.gov/ was conducted to find active clinical studies that target the biological mechanism underlying GSC-mediated resistance to adjuvant therapy.
The selection of papers was based on the following inclusion criteria: (1) English language; (2) Studies on targeted therapies against the molecular mechanism of GSC-mediated resistance to RTR; and (3) Studies on targeted therapies against the molecular mechanism of GSC-mediated resistance to CTR. The following exclusion criteria were applied: (1) Case series, case reports, editorials, meta-analyses, literature reviews, and cohort studies; (2) Studies in which the methods and/or results were imprecisely stated or poorly defined; (3) Studies that left out details regarding certain target treatments; (4) Repeatedly published research; and (5) unavailability of the full text.
Before the qualified studies were imported into Endnote X9, duplicates were removed. Two separate researchers, Panciani PP and Agosti E analyzed the data in compliance with the conditions for inclusion and exclusion. A third reviewer, Zeppieri M, resolved all disputes. Afterwards, the qualifying articles underwent full-text screening.
The data extracted for each study included: Authors, year of publication, glioma cell lines studies, GSCs pathway, therapeutic target and agents, molecular effects and impact on radio- and chemoresistance.
The molecular mechanism of GSC-mediated resistance to radiation therapy and/or chemotherapy, as well as targeted therapeutics that target the molecular mechanism of GSC-mediated resistance to adjuvant treatments, were our primary outcomes.
The Newcastle-Ottawa Scale was used to assess the quality of the listed studies[8]. The quality was conducted by analyzing the study’s outcome evaluation, comparability, and selection criteria. The ideal score was nine. Higher scores corresponded with higher-quality studies. Studies with a seven or above were considered to be of very high caliber. Panciani PP and Agosti E carried out the quality evaluation independently. When discrepancies surfaced, the third author went back and reviewed the papers (Figure 1).
The offered descriptive data contained ranges and percentages. All statistical analyses were performed using R statistical software, version 3.4.1 (http://www.r-project.org).
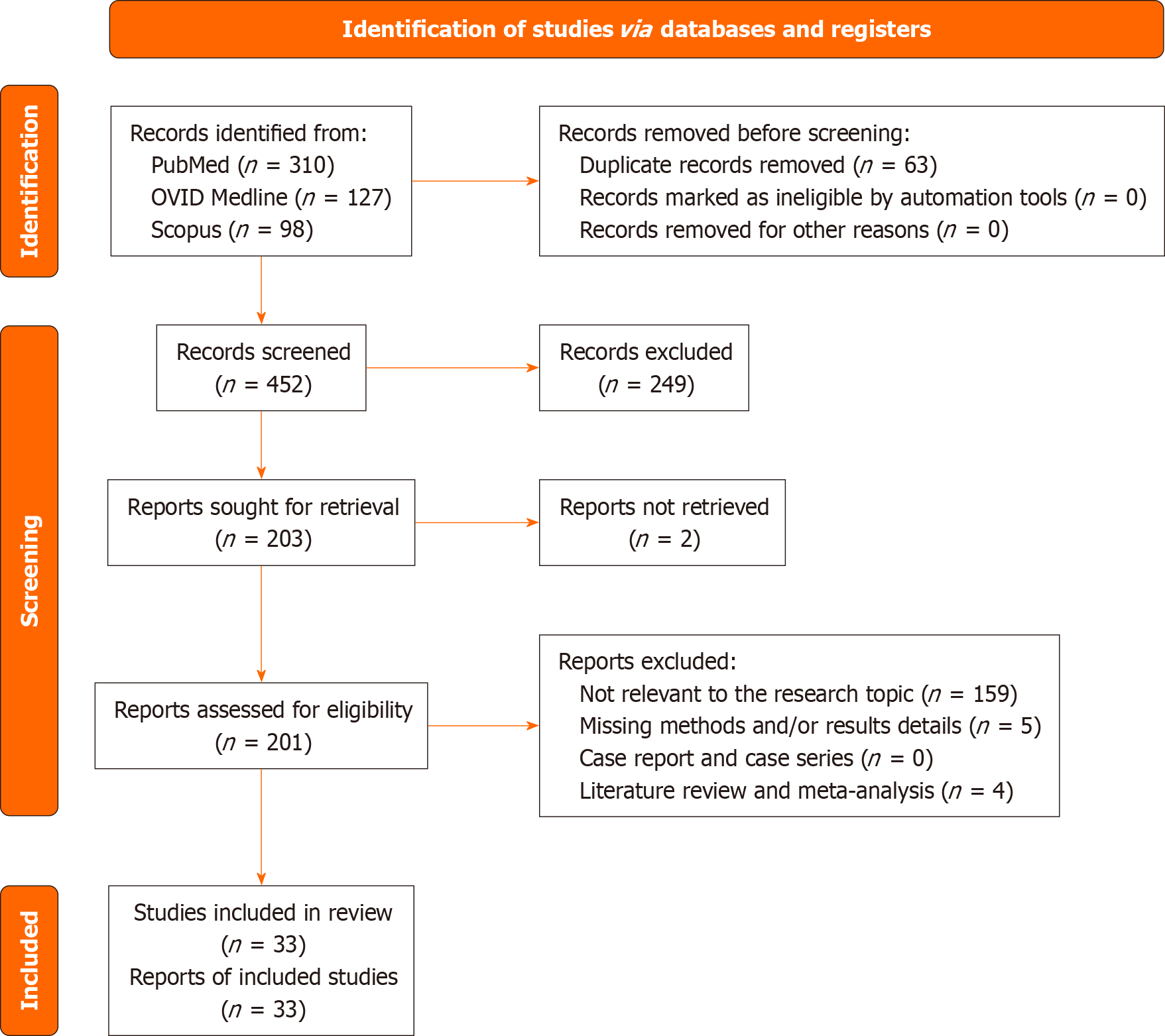
A total of 452 publications were found after duplicates were removed. 203 publications were identified for full-text analysis after the abstracts and titles were analyzed. Qualified articles included 201 papers. A total of 168 items were excluded based on the following criteria: Meta-analysis or systematic literature review (4 papers), lack of details about results and/or results (5 articles), and studies unrelated to the research issue (159 papers). All the studies included in the analysis had one or more outcome measures available for each of the patient categories under consideration. Figure 2 shows the flow chart for the PRISMA statement.
The systematic literature review focused on targeted therapies against GSCs to address resistance to RTR and CTR. The analysis of data from Table 1 provides a comprehensive understanding of the trends and frequencies associated with key parameters, including the year of publication, target GSCs models, GSCs pathways, therapeutic targets, therapeutic agents, molecular effects, and the impact on adjuvant treatment responses.
| Ref. | GCLs | GSCs pathway | Therapeutic target | Therapeutic agent | Effects | |
| Molecular | Adjuvant treatments response | |||||
| Eller et al[9], 2005 | Ros57, Jon52, Mor56 | RAS/RAF/MAPK | EGFR | Cetuximab | EGFR inhibition | RTR reduction |
| Bao et al[1], 2006 | A172 | ATR/Chk1/p53, ATM/Chk2/p53 | Chk1, Chk2 | DBH | Chk1/2 inhibition | RTR reduction |
| Clement et al[10], 2007 | U87 | HH-GLI | GLI1, GLI2, GLI3R | Cyclopamine | GLI1,2 inhibition and GLI3R activation | CTR to TMZ reduction |
| Bleau et al[11], 2009 | U87 | PI3K/AKT/mTOR | PI3Kα/δ, Akt PHD | LY294002, perifosine | ABCG2 inhibition | CTR to TMZ reduction |
| Li et al[12], 2009 | U373MG | NF-κB | LRRFIP1 | miR-21 | LRRFIP1 inhibition | CTR to VM-26 reduction |
| Wang et al[13], 2010 | T3359, T3691, T4105, T4202, T4592 | Notch | NOTCH1, NOTCH2 | GSIs, Notch1/2-specific shRNA | NOTCH1/2 inhibition | RTR reduction |
| Facchino et al[14], 2010 | Not specified | PRC1 | BMI1 | RNAi (shBMI1) | BMI1 inhibition | RTR reduction |
| Li et al[15], 2010 | Not specified | PI3K/Akt/mTOR | ABCG2 | miRNA-328 | ABCG2 inhibition | CTR to TMZ reduction |
| Ulasov et al[16], 2011 | U87MG | Notch, SHH | NOTCH1, SMO | GSI-1, cyclopamine | Notch and SHH inhibition | CTR to TMZ reduction |
| Wu et al[17], 2010 | Not specified | ATR/CHK1/p53 | CHK1 | RNAi (shCHK1) | CHK1 inhibition | RTR reduction |
| Squatrito et al[18], 2012 | Not applicable | ATM/CHK2/p53 | CHK2 | RCAS/PDGF | CHK2 inhibition | RTR reduction |
| Zhu et al[19], 2011 | HSR-GBM1/2/3 | Notch | JAG1, DLL4 | RNAi (shJAG1, shDLL4) | JAG1 and DLL4 inhibition | CTR to TMZ and RTR reduction |
| Nadkarni et al[20], 2012 | U251, U87 | PIKK | ATM | KU-55933 | ATM inhibition | CTR to TMZ reduction |
| Nabissi et al[21], 2013 | U87MG | PI3K/Akt/mTOR | TRPV2 | CBD | TRPV2 inhibition | CTR to TMZ, BCNU and DOXO reduction |
| Wang et al[22], 2013 | SU-2 | PI3K/Akt/mTOR | mTORC1/2 | NVP-BEZ235 | mTORC1/2 inhibition | RTR reduction |
| Martín et al[23], 2013 | A172, U87, U373 | ABC | ABCG2/BCRP | Melatonin | ABCG2/BCRP inhibition | CTR to TMZ reduction |
| Bhat et al[24], 2013 | Not specified | TNF-α/NF-κB | NF-κB | IkB-SR | NF-κB inhibition | RTR reduction |
| Aldea et al[25], 2014 | Not specified | PI3K/AKT/mTOR RAS/RAF/MAPK | mTOR, RAF | Metformin, sorafenib | mTOR and RAF, inhibition | CTR to TMZ reduction |
| Nabissi et al[26], 2015 | Not specified | PI3K/AKT/mTOR TRPV2 | Aml-1a | CBD | Aml-1a inhibition | CTR to BCNU reduction |
| Yu et al[27], 2015 | U87, U251, T98G, SHG44 | PI3K/AKT/mTOR | mTORC1/2 | NVP-BEZ235 | mTORC1/2 inhibition | CTR to TMZ reduction |
| Natsumeda et al[28], 2016 | HSR-GBM1, JHH520 | Notch | HES1, HES5, HEY1, autophagy | MRK003 + CQ | Notch inhibition | CTR reduction |
| Venugopal et al[29], 2015 | Not specified | Wnt/β-catenin | CK1α | Pyrvinium | CK1α inhibition | CTR to TMZ and RTR reduction |
| Yi et al[30], 2016 | U87 | Autophagic PCD | Autophagy, Bcl-2, Caspase-3 | CQ | Bcl-2 inhibition and Caspase-3 activation | RTR reduction |
| Marampon et al[31], 2017 | U87MG, U251MG | HDAC | HDAC4, HDAC6 | shRNAs (HDAC4-shRNA, HDAC6-shRNA) | HDAC4/6 inhibition | RTR reduction |
| Huang et al[32], 2017 | Not specified | MST4/ATG4B | ATG4B | NSC185058 | ATG4B inhibition | RTR reduction |
| Dai et al[33], 2017 | T98G, U87, U251, U343, MGR2, Hs683 | AKT/GSK3β/β-catenin | SCD1 | siRNA (SCD1-siRNA) | SCD1 inhibition | CTR to TMZ reduction |
| Minata et al[34], 2019 | U87, TS528 | PTEN/PI3K/AKT | pY240-PTEN | PD173074 | FGFR2 inhibition | CTR to TMZ and RTR reduction |
| Yuan et al[35], 2019 | T98G-R, U118-R | FUS/MDM2 | FUS domains (RRM, Znf_BP2) | ADAMTS9-AS2 | FUS domains inhibition | CTR to TMZ reduction |
| Moon et al[36], 2020 | T98G, LN229, U118MG, U87MG, U251MG | CK1A/BTRCP/MBD3/NuRD | CK1A | Pyr-Pam | MBD3 pathway inhibition by activating CK1A | CTR to TMZ reduction |
| Huang et al[37], 2021 | Not specified | CK2α/PRMT6/RCC1 | PRMT6 | EPZ020411 | PRMT6 inhibition | RTR reduction |
| Chen et al[38], 2022 | T98G, LN229 | PI3K/AKT/mTOR, NF-κB | EPHB3, TNFAIP3 | YTHDF2 | EPHB3 and TNFAIP3, inhibition | CTR to TMZ reduction |
| Chang et al[39], 2023 | Not specified | USP36/ALKBH5 | USP36 | shRNA | Chk1 inhibition | RTR reduction |
The studies span from 2005 to 2022, showcasing a gradual increase in research over time. Notably, there is a clustering of publications in recent years, suggesting a heightened interest and focus on developing targeted therapies against GSC-mediated resistance in glioma. The distribution of publications is as follows: (1) 2005-2010: 11 studies (33.3%); (2) 2011-2015: 12 studies (36.4%); and (3) 2016-2022: 10 studies (30.3%). This breakdown provides a temporal perspective on the evolving landscape of research in this field.
The most frequently used GSC models are crucial indicators of their relevance in research. The distribution of target GSC models in descending order of frequency is as follows: (1) U87: 11 studies (33.3%); (2) U251: 5 studies (15.2%); (3) T98G: 5 studies (15.2%); and (4) Other cell lines: 6 studies (18.2%). In some studies, the GCLs used were not specified (11, 33.3%). The prominence of U87, U251, and T98G underscores their significance in GSC research, likely due to their representativeness of glioma characteristics.
The pathways involved in GSC-mediated resistance exhibit distinct frequencies, highlighting the emphasis on specific molecular targets. The distribution of GSCs pathways, from most to least frequent, is as follows: (1) Phosphoinositide 3-kinase/protein kinase B/mammalian target of rapamycin (PI3K/AKT/mTOR): 9 studies (27.3%); (2) Notch: 4 studies (12.1%); (3) Ataxia telangiectasia and Rad3-related/checkpoint kinase 1/protein 53 (ATR/Chk1/p53) or ataxia telan
The targeted molecules across various studies demonstrate a diverse range of therapeutic approaches. The distribution of therapeutic targets, from most to least frequent, is as follows: mTOR: 6 studies (18.2%); CHK1/2: 5 studies (15.2%); ATP binding cassette G2 (ABCG2): 4 studies (12.1%); epidermal growth factor receptor (EGFR): 2 studies (6.1%); and others: 19 studies (57.6%). The high frequency of mTOR and CHK1/2 as therapeutic targets underscores their significance in overcoming GSC-mediated resistance.
Different classes of therapeutic agents exhibit varying frequencies, indicating preferences and efficacies. The distribution of therapeutic agents, from most to least frequent, is as follows: RNA inhibitor (RNAi)/short hairpin RNA (shRNA): 9 studies (27.3%); inhibitors (e.g., LY294002, NVP-BEZ235): 8 studies (24.2%); cannabidiol (CBD): 3 studies (9.1%); monoclonal antibodies (e.g., cetuximab): 3 studies (9.1%); and others: 8 studies (24.2%). The prevalence of RNAi/shRNA and inhibitors highlights the versatility of these agents in targeted therapies against GSC-mediated resistance.
The outcomes at the molecular level show distinct frequencies, indicating the varied impacts of therapeutic interventions. The distribution of molecular effects, from most to least frequent, is as follows: Inhibition of key signaling molecules: 32 studies (97.0%); modulation of cellular processes (e.g., autophagy): 3 studies (9.1%); and others: 2 studies (6.1%). The impact of therapeutic agents on adjuvant treatment responses is crucial for assessing overall efficacy. The distribution of effects on CTR response (in total described in 20 studies, 60.6%), from most to least frequent, is as follows: Reduction in TMZ resistance: 17 studies (51.5%); reduction in carmustine (BCNU) resistance: 3 studies (9.1%); and reduction in doxorubicin (DOXO) resistance: 1 study (3.0%). The reduction of resistance to RTR was reported in 14 studies (42.4%).
The effectiveness of therapeutic agents in reducing resistance to specific adjuvant treatments provides valuable insights into their clinical potential[1,9-39] (Table 1). A summary of the ongoing clinical trials focusing on targeted therapies against the molecular mechanism of GSC-mediated resistance to adjuvant treatments is available in Table 2.
| Trial name | Year | Title | Trial phase | Therapeutic agent |
| NCT00916409 | 2014 | A prospective, multicenter trial of NovoTTF-100 A together with TMZ compared to TMZ alone in patients with newly diagnosed GBM | III | NovoTTF-100A device |
| NCT01474239 | 2016 | Randomized noncomparative phase II trial with bevacizumab and FOT in the treatment of recurrent GBM | II | Bevacizumab |
| NCT02698280 | 2018 | Phase II study of bevacizumab and ACNU in patients with recurrent high-grade glioma | II | Bevacizumab |
| NCT04396860 | 2020 | Randomized noncomparative phase II trial with bevacizumab and FOT in the treatment of recurrent GBM | II/III | Ipilimumab, nivolumab, NovoTTF-100 A device |
| NCT01310868 | 2021 | An evaluation of the tolerability and feasibility of combining 5-ALA with BCNU wafers (Gliadel®) in the surgical management of primary GBM | II | 5-ALA Gliadel® wafers |
| NCT01290939 | Ongoing | Phase III trial exploring the combination of bvacizumab and CCNU in patients with first recurrence of a GBM | III | Bevacizumab |
| NCT00884741 | Ongoing | Phase III double-blind placebo-controlled trial of conventional concurrent chemoradiation and adjuvant TMZ plus bevacizumab versus conventional concurrent chemoradiation and adjuvant TMZ in patients with newly diagnosed GBM | III | Bevacizumab |
| NCT02017717 | Ongoing | A randomized phase III open-label study of nivolumab versus bevacizumab and multiple phase I safety cohorts of nivolumab or nivolumab in combination with ipilimumab across different lines of GBM | III | Nivolumab, bevacizumab, ipilimumab |
| NCT00753246 | Ongoing | Phase III study of standard radiotherapy plus concomitant and adjuvant OSAG 101 (Theraloc®) plus TMZ versus standard radiotherapy plus concomitant and adjuvant TMZ patient with newly diagnosed, histologically confirmed GBM multiforme grade IV | III | Nimotuzumab |
The radioresistance and chemoresistance promoted by GSCs poses a significant challenge in the effective treatment of gliomas, and understanding the molecular mechanisms underlying this resistance is crucial for developing targeted therapies[2,3,5,6]. Several studies have identified key signaling pathways implicated in GSC-mediated resistance. For instance, Eller et al[9] highlighted the involvement of the RAS/RAF/MAPK pathway and its therapeutic targeting through EGFR inhibition with Cetuximab. This finding aligns with other studies, such as that by Bao et al[1] emphasizing the importance of ATR/Chk1 and ATM/Chk2 pathways in GSC resistance, which was effectively targeted by Chk1/2 inhibition[5]. Moreover, the PI3K/AKT/mTOR pathway has been implicated in GSC-mediated resistance[11,12], and its inhibition, as demonstrated by Nabissi et al[21] led to a reduction in TMZ resistance[5]. These results collectively underscore the complexity of GSC-associated signaling networks and the need for a multifaceted therapeutic approach[2,3].
Beyond signaling pathways, studies have explored the role of specific molecules and proteins in GSC-mediated re
Different proteins and factors can be implicated in GSC-mediated resistance. For instance, Wang et al[22] highlighted the role of transient receptor potential cation channel subfamily V member 2 in resistance to TMZ, BCNU, and DOXO, which was effectively targeted using CBD. Similarly, the inhibition of ABCG2/breast cancer resistance protein by Melatonin, as demonstrated by Martín et al[23] resulted in reduced resistance to TMZ. These studies shed light on the potential of targeting specific proteins to overcome GSC-mediated resistance[4,5].
The modulation of autophagy has emerged as a promising therapeutic avenue in overcoming GSC resistance. Venugopal et al[29] demonstrated that inhibiting Notch through MRK003 + chloroquine effectively reduced GSC resistance, highlighting the crosstalk between autophagy and GSC-mediated resistance. Additionally, Yi et al[30] targeted CK1α to reduce resistance to TMZ and radiotherapy, emphasizing the intricate network of pathways involved in GSC-mediated resistance. The study by Alhaddad et al[40] delves into the role of autophagy induction in promoting M2-like macrophage polarization, thereby contributing to radioresistance[41]. This contrasts with Li et al[42] where PI3Kγ inhibition suppresses microglia/TAM accumulation in the glioma microenvironment by inhibiting autophagy, ultimately promoting an exceptional response to TMZ[2]. The dual role of autophagy in GSC-mediated resistance mechanisms high
The inclusion of studies by Zhang et al[43] and Wu et al[44] in the systematic review underscores the relevance of long non-coding RNAs (lncRNAs) in GSC-mediated chemoresistance. Zhang et al[43] shed light on the exosomal transfer of lncRNA SBF2-AS1, enhancing chemoresistance to TMZ[6]. In parallel, Wu et al[44] demonstrate the role of lnc-TALC in promoting O6-methylguanine-DNA methyltransferase expression, a key contributor to TMZ resistance[17]. These findings resonate with the growing body of evidence highlighting the regulatory role of lncRNAs in glioma progression[42].
The role of epigenetic regulators in GSC resistance emerged. Huang et al[32] targeted HDAC4/6 using shRNAs, re
The contribution of hypoxia-driven mechanisms to GSC-mediated radioresistance, as discussed by Alhaddad et al[40] resonates with the findings of Hsieh et al[45] NADPH oxidase subunit 4, identified as a mediator of cycling hypoxia-promoted radiation resistance in glioblastoma multiforme, underscores the intricate relationship between the tumor microenvironment and resistance mechanisms. The role of hypoxia in shaping the GSC phenotype adds a layer of complexity to the design of targeted therapies, necessitating strategies that account for the dynamic nature of the tumor microenvironment[2,6,46].
Moreover, the manuscript the interplay between GSCs and the tumor microenvironment seems to play a pivotal role in the development of glioma resistance. Moon et al[36] targeted CK1A/BTRCP/MBD3/NuRD pathway using pyrvinium pamoate, illustrating the complex interaction between GSCs and their microenvironment in mediating resistance. The study by Huang et al[37] further emphasized the role of CK2α/PRMT6/RCC1 pathway, which was targeted using EPZ020411, resulting in a reduction in radioresistance. The study by Alhaddad et al[40] provides insights into GSC-mediated reprogramming of the tumor microenvironment, specifically in the context of macrophages and microglia. The creation of GSC niches at the tumor border, as reported by Hide et al[47] emphasizes the dynamic interplay between oligodendrocyte progenitor cells, macrophages/microglia, and the GSC microenvironment. This aligns with the findings of Li et al[42] where PI3Kγ inhibition suppresses microglia/TAM accumulation in the glioblastoma microenvironment. The intercellular crosstalk between GSCs and immune components suggests a complex landscape that contributes to therapy resistance. The study by Alhaddad et al[40] emphasizes the broader influence of GSCs on the tumor microenvironment and therapy response. The creation of GSC niches at the tumor border, as demonstrated by Hide et al[47] signifies the role of GSCs in orchestrating the cellular composition of the microenvironment. This resonates with the findings of Liu et al[48], where ADAM8 causes tumor infiltration of tumor-associated macrophages and overcomes TMZ chemosensitization. The interplay between GSCs and the microenvironment emerges as a critical determinant of therapy response and poses challenges for designing targeted interventions[49,50]. These findings highlight the need for a holistic approach considering the tumor microenvironment in developing therapeutic strategies[2,5,6].
In conclusion, the role of GSCs in mediating radioresistance and chemoresistance is a complex and multifaceted phenomenon. Versatile therapeutic agents, including RNAi/shRNA, inhibitors (e.g., LY294002, NVP-BEZ235), and monoclonal antibodies (e.g., cetuximab), demonstrate efficacy in overcoming GSC-mediated resistance. The diverse mechanisms discussed in the literature highlight the need for precision therapies that account for the heterogeneity within the GSC population and the dynamic nature of the tumor microenvironment. As research in this field progresses, a deeper understanding of the molecular intricacies will pave the way for targeted interventions that can disrupt GSC-mediated resistance mechanisms, ultimately improving the outcomes for glioblastoma patients.
| 1. | Bao S, Wu Q, McLendon RE, Hao Y, Shi Q, Hjelmeland AB, Dewhirst MW, Bigner DD, Rich JN. Glioma stem cells promote radioresistance by preferential activation of the DNA damage response. Nature. 2006;444:756-760. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 5276] [Cited by in RCA: 4798] [Article Influence: 239.9] [Reference Citation Analysis (0)] |
| 2. | Eckerdt F, Platanias LC. Emerging Role of Glioma Stem Cells in Mechanisms of Therapy Resistance. Cancers (Basel). 2023;15. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 78] [Cited by in RCA: 52] [Article Influence: 17.3] [Reference Citation Analysis (0)] |
| 3. | Mattei V, Santilli F, Martellucci S, Delle Monache S, Fabrizi J, Colapietro A, Angelucci A, Festuccia C. The Importance of Tumor Stem Cells in Glioblastoma Resistance to Therapy. Int J Mol Sci. 2021;22. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 63] [Cited by in RCA: 51] [Article Influence: 10.2] [Reference Citation Analysis (0)] |
| 4. | Auffinger B, Spencer D, Pytel P, Ahmed AU, Lesniak MS. The role of glioma stem cells in chemotherapy resistance and glioblastoma multiforme recurrence. Expert Rev Neurother. 2015;15:741-752. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 267] [Cited by in RCA: 236] [Article Influence: 21.5] [Reference Citation Analysis (0)] |
| 5. | Chen B, Zhou X, Yang L, Zhou H, Meng M, Wu H, Liu Z, Zhang L, Li C. Glioma stem cell signature predicts the prognosis and the response to tumor treating fields treatment. CNS Neurosci Ther. 2022;28:2148-2162. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 27] [Cited by in RCA: 22] [Article Influence: 5.5] [Reference Citation Analysis (0)] |
| 6. | Ahmed AU, Auffinger B, Lesniak MS. Understanding glioma stem cells: rationale, clinical relevance and therapeutic strategies. Expert Rev Neurother. 2013;13:545-555. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 75] [Cited by in RCA: 67] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 7. | Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, Shamseer L, Tetzlaff JM, Akl EA, Brennan SE, Chou R, Glanville J, Grimshaw JM, Hróbjartsson A, Lalu MM, Li T, Loder EW, Mayo-Wilson E, McDonald S, McGuinness LA, Stewart LA, Thomas J, Tricco AC, Welch VA, Whiting P, Moher D. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. 2021;372:n71. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 9803] [Reference Citation Analysis (0)] |
| 8. | Wells G, Shea BJ, O’Connell D, Peterson J, Welch V, Losos M, Tugwell P. The Newcastle-Ottawa Scale (NOS) for Assessing the Quality of Non-Randomized Studies in Meta-Analysis. [cited 19 July 2023]. Available from: https://www.researchgate.net/publication/261773681_The_Newcastle-Ottawa_Scale_NOS_for_Assessing_the_Quality_of_Non-Randomized_Studies_in_Meta-Analysis. |
| 9. | Eller JL, Longo SL, Kyle MM, Bassano D, Hicklin DJ, Canute GW. Anti-epidermal growth factor receptor monoclonal antibody cetuximab augments radiation effects in glioblastoma multiforme in vitro and in vivo. Neurosurgery. 2005;56:155-62; discussion 162. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 85] [Cited by in RCA: 88] [Article Influence: 4.2] [Reference Citation Analysis (1)] |
| 10. | Clement V, Sanchez P, de Tribolet N, Radovanovic I, Ruiz i Altaba A. HEDGEHOG-GLI1 signaling regulates human glioma growth, cancer stem cell self-renewal, and tumorigenicity. Curr Biol. 2007;17:165-172. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 869] [Cited by in RCA: 828] [Article Influence: 43.6] [Reference Citation Analysis (0)] |
| 11. | Bleau AM, Hambardzumyan D, Ozawa T, Fomchenko EI, Huse JT, Brennan CW, Holland EC. PTEN/PI3K/Akt pathway regulates the side population phenotype and ABCG2 activity in glioma tumor stem-like cells. Cell Stem Cell. 2009;4:226-235. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 676] [Cited by in RCA: 623] [Article Influence: 36.6] [Reference Citation Analysis (0)] |
| 12. | Li Z, Bao S, Wu Q, Wang H, Eyler C, Sathornsumetee S, Shi Q, Cao Y, Lathia J, McLendon RE, Hjelmeland AB, Rich JN. Hypoxia-inducible factors regulate tumorigenic capacity of glioma stem cells. Cancer Cell. 2009;15:501-513. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1146] [Cited by in RCA: 1052] [Article Influence: 61.9] [Reference Citation Analysis (0)] |
| 13. | Wang J, Wakeman TP, Lathia JD, Hjelmeland AB, Wang XF, White RR, Rich JN, Sullenger BA. Notch promotes radioresistance of glioma stem cells. Stem Cells. 2010;28:17-28. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 447] [Cited by in RCA: 431] [Article Influence: 26.9] [Reference Citation Analysis (0)] |
| 14. | Facchino S, Abdouh M, Chatoo W, Bernier G. BMI1 confers radioresistance to normal and cancerous neural stem cells through recruitment of the DNA damage response machinery. J Neurosci. 2010;30:10096-10111. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 230] [Cited by in RCA: 218] [Article Influence: 13.6] [Reference Citation Analysis (0)] |
| 15. | Li P, Lu X, Wang Y, Sun L, Qian C, Yan W, Liu N, You Y, Fu Z. MiR-181b suppresses proliferation of and reduces chemoresistance to temozolomide in U87 glioma stem cells. J Biomed Res. 2010;24:436-443. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 32] [Cited by in RCA: 36] [Article Influence: 2.3] [Reference Citation Analysis (0)] |
| 16. | Ulasov IV, Nandi S, Dey M, Sonabend AM, Lesniak MS. Inhibition of Sonic hedgehog and Notch pathways enhances sensitivity of CD133(+) glioma stem cells to temozolomide therapy. Mol Med. 2011;17:103-112. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 156] [Cited by in RCA: 143] [Article Influence: 9.5] [Reference Citation Analysis (0)] |
| 17. | Wu A, Wei J, Kong LY, Wang Y, Priebe W, Qiao W, Sawaya R, Heimberger AB. Glioma cancer stem cells induce immunosuppressive macrophages/microglia. Neuro Oncol. 2010;12:1113-1125. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 556] [Cited by in RCA: 535] [Article Influence: 33.4] [Reference Citation Analysis (0)] |
| 18. | Squatrito M, Vanoli F, Schultz N, Jasin M, Holland EC. 53BP1 is a haploinsufficient tumor suppressor and protects cells from radiation response in glioma. Cancer Res. 2012;72:5250-5260. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 33] [Cited by in RCA: 30] [Article Influence: 2.1] [Reference Citation Analysis (0)] |
| 19. | Zhu TS, Costello MA, Talsma CE, Flack CG, Crowley JG, Hamm LL, He X, Hervey-Jumper SL, Heth JA, Muraszko KM, DiMeco F, Vescovi AL, Fan X. Endothelial cells create a stem cell niche in glioblastoma by providing NOTCH ligands that nurture self-renewal of cancer stem-like cells. Cancer Res. 2011;71:6061-6072. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 319] [Cited by in RCA: 287] [Article Influence: 19.1] [Reference Citation Analysis (0)] |
| 20. | Nadkarni A, Shrivastav M, Mladek AC, Schwingler PM, Grogan PT, Chen J, Sarkaria JN. ATM inhibitor KU-55933 increases the TMZ responsiveness of only inherently TMZ sensitive GBM cells. J Neurooncol. 2012;110:349-357. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 60] [Cited by in RCA: 59] [Article Influence: 4.2] [Reference Citation Analysis (0)] |
| 21. | Nabissi M, Morelli MB, Santoni M, Santoni G. Triggering of the TRPV2 channel by cannabidiol sensitizes glioblastoma cells to cytotoxic chemotherapeutic agents. Carcinogenesis. 2013;34:48-57. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 208] [Cited by in RCA: 193] [Article Influence: 14.8] [Reference Citation Analysis (0)] |
| 22. | Wang J, Ma Y, Cooper MK. Cancer stem cells in glioma: challenges and opportunities. Transl Cancer Res. 2013;2:429-441. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 20] [Reference Citation Analysis (0)] |
| 23. | Martín V, Sanchez-Sanchez AM, Herrera F, Gomez-Manzano C, Fueyo J, Alvarez-Vega MA, Antolín I, Rodriguez C. Melatonin-induced methylation of the ABCG2/BCRP promoter as a novel mechanism to overcome multidrug resistance in brain tumour stem cells. Br J Cancer. 2013;108:2005-2012. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 119] [Cited by in RCA: 103] [Article Influence: 7.9] [Reference Citation Analysis (0)] |
| 24. | Bhat KPL, Balasubramaniyan V, Vaillant B, Ezhilarasan R, Hummelink K, Hollingsworth F, Wani K, Heathcock L, James JD, Goodman LD, Conroy S, Long L, Lelic N, Wang S, Gumin J, Raj D, Kodama Y, Raghunathan A, Olar A, Joshi K, Pelloski CE, Heimberger A, Kim SH, Cahill DP, Rao G, Den Dunnen WFA, Boddeke HWGM, Phillips HS, Nakano I, Lang FF, Colman H, Sulman EP, Aldape K. Mesenchymal differentiation mediated by NF-κB promotes radiation resistance in glioblastoma. Cancer Cell. 2013;24:331-346. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 944] [Cited by in RCA: 882] [Article Influence: 67.8] [Reference Citation Analysis (0)] |
| 25. | Aldea MD, Petrushev B, Soritau O, Tomuleasa CI, Berindan-Neagoe I, Filip AG, Chereches G, Cenariu M, Craciun L, Tatomir C, Florian IS, Crivii CB, Kacso G. Metformin plus sorafenib highly impacts temozolomide resistant glioblastoma stem-like cells. J BUON. 2014;19:502-511. [PubMed] |
| 26. | Nabissi M, Morelli MB, Amantini C, Liberati S, Santoni M, Ricci-Vitiani L, Pallini R, Santoni G. Cannabidiol stimulates Aml-1a-dependent glial differentiation and inhibits glioma stem-like cells proliferation by inducing autophagy in a TRPV2-dependent manner. Int J Cancer. 2015;137:1855-1869. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 137] [Cited by in RCA: 129] [Article Influence: 11.7] [Reference Citation Analysis (0)] |
| 27. | Yu Z, Zhao G, Xie G, Zhao L, Chen Y, Yu H, Zhang Z, Li C, Li Y. Metformin and temozolomide act synergistically to inhibit growth of glioma cells and glioma stem cells in vitro and in vivo. Oncotarget. 2015;6:32930-32943. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 92] [Cited by in RCA: 86] [Article Influence: 7.8] [Reference Citation Analysis (0)] |
| 28. | Natsumeda M, Maitani K, Liu Y, Miyahara H, Kaur H, Chu Q, Zhang H, Kahlert UD, Eberhart CG. Targeting Notch Signaling and Autophagy Increases Cytotoxicity in Glioblastoma Neurospheres. Brain Pathol. 2016;26:713-723. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 44] [Cited by in RCA: 39] [Article Influence: 3.9] [Reference Citation Analysis (0)] |
| 29. | Venugopal C, Hallett R, Vora P, Manoranjan B, Mahendram S, Qazi MA, McFarlane N, Subapanditha M, Nolte SM, Singh M, Bakhshinyan D, Garg N, Vijayakumar T, Lach B, Provias JP, Reddy K, Murty NK, Doble BW, Bhatia M, Hassell JA, Singh SK. Pyrvinium Targets CD133 in Human Glioblastoma Brain Tumor-Initiating Cells. Clin Cancer Res. 2015;21:5324-5337. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 49] [Cited by in RCA: 48] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 30. | Yi Y, Hsieh IY, Huang X, Li J, Zhao W. Glioblastoma Stem-Like Cells: Characteristics, Microenvironment, and Therapy. Front Pharmacol. 2016;7:477. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 97] [Cited by in RCA: 84] [Article Influence: 8.4] [Reference Citation Analysis (0)] |
| 31. | Marampon F, Megiorni F, Camero S, Crescioli C, McDowell HP, Sferra R, Vetuschi A, Pompili S, Ventura L, De Felice F, Tombolini V, Dominici C, Maggio R, Festuccia C, Gravina GL. HDAC4 and HDAC6 sustain DNA double strand break repair and stem-like phenotype by promoting radioresistance in glioblastoma cells. Cancer Lett. 2017;397:1-11. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 91] [Cited by in RCA: 82] [Article Influence: 9.1] [Reference Citation Analysis (0)] |
| 32. | Huang TY, Piunti A, Lulla RR, Qi J, Horbinski CM, Tomita T, James CD, Shilatifard A, Saratsis AM. Detection of Histone H3 mutations in cerebrospinal fluid-derived tumor DNA from children with diffuse midline glioma. Acta Neuropathol Commun. 2017;5:28. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 146] [Cited by in RCA: 137] [Article Influence: 15.2] [Reference Citation Analysis (0)] |
| 33. | Dai S, Yan Y, Xu Z, Zeng S, Qian L, Huo L, Li X, Sun L, Gong Z. SCD1 Confers Temozolomide Resistance to Human Glioma Cells via the Akt/GSK3β/β-Catenin Signaling Axis. Front Pharmacol. 2017;8:960. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 72] [Cited by in RCA: 74] [Article Influence: 9.3] [Reference Citation Analysis (0)] |
| 34. | Minata M, Audia A, Shi J, Lu S, Bernstock J, Pavlyukov MS, Das A, Kim SH, Shin YJ, Lee Y, Koo H, Snigdha K, Waghmare I, Guo X, Mohyeldin A, Gallego-Perez D, Wang J, Chen D, Cheng P, Mukheef F, Contreras M, Reyes JF, Vaillant B, Sulman EP, Cheng SY, Markert JM, Tannous BA, Lu X, Kango-Singh M, Lee LJ, Nam DH, Nakano I, Bhat KP. Phenotypic Plasticity of Invasive Edge Glioma Stem-like Cells in Response to Ionizing Radiation. Cell Rep. 2019;26:1893-1905.e7. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 194] [Cited by in RCA: 181] [Article Influence: 25.9] [Reference Citation Analysis (0)] |
| 35. | Yuan Y, Yan Z, Miao J, Cai R, Zhang M, Wang Y, Wang L, Dang W, Wang D, Xiang D, Zhang P, Cui Y, Bian X, Ma Q. Autofluorescence of NADH is a new biomarker for sorting and characterizing cancer stem cells in human glioma. Stem Cell Res Ther. 2019;10:330. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 35] [Cited by in RCA: 31] [Article Influence: 4.4] [Reference Citation Analysis (0)] |
| 36. | Moon BS, Cai M, Lee G, Zhao T, Song X, Giannotta SL, Attenello FJ, Yu M, Lu W. Epigenetic modulator inhibition overcomes temozolomide chemoresistance and antagonizes tumor recurrence of glioblastoma. J Clin Invest. 2020;130:5782-5799. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 25] [Cited by in RCA: 22] [Article Influence: 3.7] [Reference Citation Analysis (0)] |
| 37. | Huang K, Yue X, Zheng Y, Zhang Z, Cheng M, Li L, Chen Z, Yang Z, Bian E, Zhao B. Development and Validation of an Mesenchymal-Related Long Non-Coding RNA Prognostic Model in Glioma. Front Oncol. 2021;11:726745. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 10] [Cited by in RCA: 10] [Article Influence: 2.0] [Reference Citation Analysis (0)] |
| 38. | Chen J, Liu G, Wang X, Hong H, Li T, Li L, Wang H, Xie J, Li B, Lu D, Zhang Y, Zhao H, Yao C, Wen K, Chen J, Wu S, He K, Zhang WN, Zhao J, Wang N, Han Q, Xia Q, Qi J, Zhou T, Man J, Zhang XM, Li AL, Pan X. Glioblastoma stem cell-specific histamine secretion drives pro-angiogenic tumor microenvironment remodeling. Cell Stem Cell. 2022;29:1531-1546.e7. [RCA] [PubMed] [DOI] [Full Text] [Cited by in RCA: 52] [Reference Citation Analysis (0)] |
| 39. | Chang G, Xie GS, Ma L, Li P, Li L, Richard HT. USP36 promotes tumorigenesis and drug sensitivity of glioblastoma by deubiquitinating and stabilizing ALKBH5. Neuro Oncol. 2023;25:841-853. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 46] [Cited by in RCA: 40] [Article Influence: 13.3] [Reference Citation Analysis (0)] |
| 40. | Alhaddad L, Osipov AN, Leonov S. The Molecular and Cellular Strategies of Glioblastoma and Non-Small-Cell Lung Cancer Cells Conferring Radioresistance. Int J Mol Sci. 2022;23. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 16] [Cited by in RCA: 13] [Article Influence: 3.3] [Reference Citation Analysis (0)] |
| 41. | Xu S, Tang L, Liu Z, Yang K, Cheng Q. Bioinformatic Analyses Identify a Prognostic Autophagy-Related Long Non-coding RNA Signature Associated With Immune Microenvironment in Diffuse Gliomas. Front Cell Dev Biol. 2021;9:694633. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 12] [Cited by in RCA: 13] [Article Influence: 2.6] [Reference Citation Analysis (0)] |
| 42. | Li Z, Li M, Xia P, Wang L, Lu Z. LncRNA FOXD3-AS1 Promotes Tumorigenesis of Glioma via Targeting miR-128-3p/SZRD1 Axis. Cancer Manag Res. 2021;13:9037-9048. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 2] [Cited by in RCA: 2] [Article Influence: 0.4] [Reference Citation Analysis (0)] |
| 43. | Zhang N, Wei L, Ye M, Kang C, You H. Treatment Progress of Immune Checkpoint Blockade Therapy for Glioblastoma. Front Immunol. 2020;11:592612. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 45] [Cited by in RCA: 39] [Article Influence: 6.5] [Reference Citation Analysis (0)] |
| 44. | Wu C, Xu Q, Chen X, Liu J. Delivery luteolin with folacin-modified nanoparticle for glioma therapy. Int J Nanomedicine. 2019;14:7515-7531. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 67] [Cited by in RCA: 56] [Article Influence: 8.0] [Reference Citation Analysis (0)] |
| 45. | Hsieh CH, Wu CP, Lee HT, Liang JA, Yu CY, Lin YJ. NADPH oxidase subunit 4 mediates cycling hypoxia-promoted radiation resistance in glioblastoma multiforme. Free Radic Biol Med. 2012;53:649-658. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 54] [Cited by in RCA: 50] [Article Influence: 3.6] [Reference Citation Analysis (0)] |
| 46. | Han X, Li H, Zhang Y, Qin J, Yang Q, Wang L, Yuan M, Xia C. Brain lipid-binding protein promotes proliferation and modulates cell cycle in C6 rat glioma cells. Int J Oncol. 2017;51:1439-1448. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 5] [Article Influence: 0.6] [Reference Citation Analysis (0)] |
| 47. | Hide T, Komohara Y, Miyasato Y, Nakamura H, Makino K, Takeya M, Kuratsu JI, Mukasa A, Yano S. Oligodendrocyte Progenitor Cells and Macrophages/Microglia Produce Glioma Stem Cell Niches at the Tumor Border. EBioMedicine. 2018;30:94-104. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 106] [Cited by in RCA: 99] [Article Influence: 12.4] [Reference Citation Analysis (0)] |
| 48. | Liu K, Jiang L, Shi Y, Liu B, He Y, Shen Q, Jiang X, Nie Z, Pu J, Yang C, Chen Y. Hypoxia-induced GLT8D1 promotes glioma stem cell maintenance by inhibiting CD133 degradation through N-linked glycosylation. Cell Death Differ. 2022;29:1834-1849. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 59] [Cited by in RCA: 53] [Article Influence: 13.3] [Reference Citation Analysis (0)] |
| 49. | Agosti E, Zeppieri M, De Maria L, Tedeschi C, Fontanella MM, Panciani PP, Ius T. Glioblastoma Immunotherapy: A Systematic Review of the Present Strategies and Prospects for Advancements. Int J Mol Sci. 2023;24. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 82] [Cited by in RCA: 66] [Article Influence: 22.0] [Reference Citation Analysis (0)] |
| 50. | Agosti E, Panciani PP, Zeppieri M, De Maria L, Pasqualetti F, Tel A, Zanin L, Fontanella MM, Ius T. Tumor Microenvironment and Glioblastoma Cell Interplay as Promoters of Therapeutic Resistance. Biology (Basel). 2023;12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 20] [Cited by in RCA: 18] [Article Influence: 6.0] [Reference Citation Analysis (0)] |
Open-Access: This article is an open-access article that was selected by an in-house editor and fully peer-reviewed by external reviewers. It is distributed in accordance with the Creative Commons Attribution NonCommercial (CC BY-NC 4.0) license, which permits others to distribute, remix, adapt, build upon this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See: https://creativecommons.org/Licenses/by-nc/4.0/
Provenance and peer review: Invited article; Externally peer reviewed.
Peer-review model: Single blind
Specialty type: Cell and tissue engineering
Country/Territory of origin: Italy
Peer-review report’s classification
Scientific Quality: Grade B, Grade B
Novelty: Grade B
Creativity or Innovation: Grade B
Scientific Significance: Grade B
P-Reviewer: Li Z, China; Ventura C, Italy S-Editor: Wang JJ L-Editor: A P-Editor: Zheng XM