Published online Dec 26, 2024. doi: 10.4252/wjsc.v16.i12.1110
Revised: October 17, 2024
Accepted: December 3, 2024
Published online: December 26, 2024
Processing time: 79 Days and 22.4 Hours
This article discusses the interplay between colorectal cancer (CRC) stem cells, tumor microenvironment (TME), and gut microbiota, emphasizing their dynamic roles in cancer progression and treatment resistance. It highlights the adaptability of CRC stem cells, the bidirectional influence of TME, and the multifaceted impact of gut microbiota on CRC. The manuscript proposes innovative therapeutic st
Core Tip: The core objective of this article is to build upon the compelling arguments presented in the review, highlighting emerging trends, and introducing fresh pers
- Citation: Zhang H, Xu BT, Luo DP, He TF. Interplay and therapeutic implications of colorectal cancer stem cells, tumor microenvironment, and gut microbiota. World J Stem Cells 2024; 16(12): 1110-1114
- URL: https://www.wjgnet.com/1948-0210/full/v16/i12/1110.htm
- DOI: https://dx.doi.org/10.4252/wjsc.v16.i12.1110
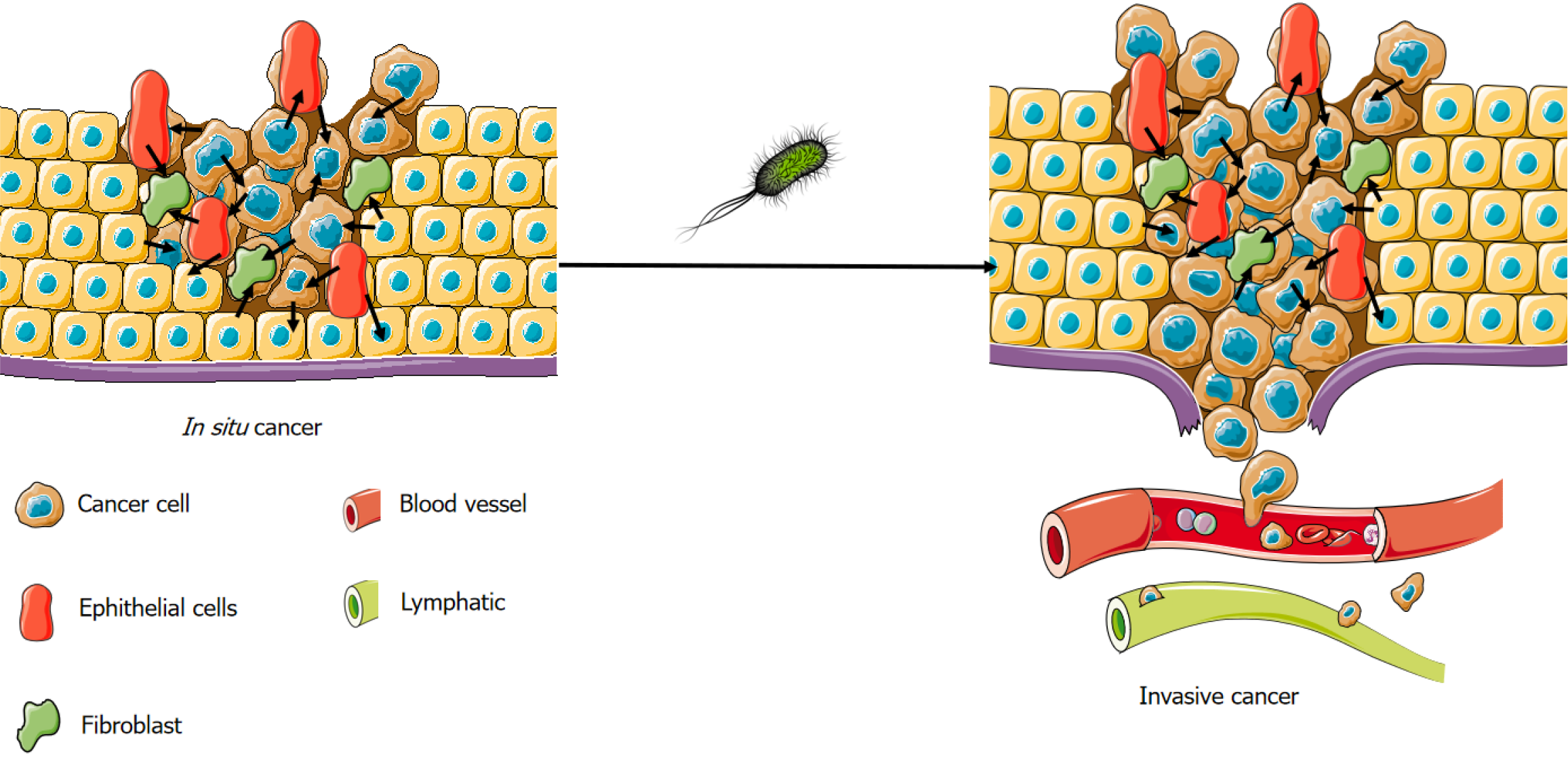
The recent review in the World Journal of Stem Cells on the complex interplay among the tumor microenvironment (TME), gut microbiota, and the stemness of colorectal cancer (CRC) cells offers a profound insight into the evolving landscape of CRC research[1]. This article aims to build upon the compelling narrative presented in the review, highlighting emerging trends and offering novel perspectives that could further elucidate the intricate relationship between these factors. Understanding this interplay is crucial, as it holds the key to unlocking new therapeutic strategies and improving patient outcomes in CRC (Figure 1).
In addressing the complex nature of CRC stem cells, it’s crucial to emphasize their dynamic role in tumor progression and therapy resistance. Recent studies have unveiled that CRC stem cells are not a static entity but are highly influenced by their microenvironment[2-4]. This adaptive nature complicates treatment strategies, as these cells can evade conventional therapies by altering their characteristics. Moreover, the heterogeneity within the stem cell population in CRC presents an additional layer of complexity, necessitating a more nuanced approach in therapeutic development. Understanding the molecular mechanisms driving this plasticity is paramount for devising effective treatments that can target these elusive cells[5-7].
The TME plays a pivotal role in the pathogenesis of CRC[8], particularly in how it shapes the behavior of CRC stem cells. The interplay between CRC stem cells and their microenvironment is a dance of mutual influence, where TME not only nurtures stem cells but also undergoes modifications instigated by these cells[9]. This bidirectional communication is key in understanding tumor progression and resistance to therapy. Recent research highlights the importance of TME components such as immune cells, stromal cells, and extracellular matrix in modulating CRC stem cell functions[10]. These insights offer new avenues for therapeutic intervention, targeting not just the cancer cells but also their supporting environment.
The gut microbiota’s role in CRC has gained significant attention, providing new insights into cancer biology and potential treatment modalities[11-13]. Research indicates that the gut microbiota influences CRC development and progression, possibly through interactions with CRC stem cells and the TME[14]. These microorganisms can affect immune responses, metabolic pathways, and even genetic stability within the tumor, suggesting a multifaceted impact on cancer dynamics. Understanding these mechanisms is crucial for developing innovative treatments that could leverage gut microbiota modulation, potentially offering a novel and less invasive approach to combating CRC.
The evolving understanding of the interplay between CRC stem cells, the TME, and gut microbiota opens up new frontiers for therapeutic strategies. Targeting these complex interactions offers a promising avenue for more effective treatments. Recent advancements propose interventions like microbiota modulation, immune therapy tailored to the TME, and drugs targeting stem cell pathways[6,15,16]. These strategies represent a paradigm shift from conventional chemotherapy to more personalized and holistic approaches, aiming to disrupt the supportive network that fuels CRC progression. The potential of these therapies lies not just in their direct anticancer effects, but also in their ability to reshape the tumor landscape, making it less conducive to cancer growth and resistance.
In conclusion, the intricate interplay between CRC stem cells, the TME, and gut microbiota presents a fascinating yet complex puzzle in the realm of CRC research. The insights gleaned from recent studies not only enhance our under
| Microbial species | Role in colorectal cancer | Associated molecular pathways | Expression in CRC patients (↑/↓) | Potential therapeutic target (yes/no) | Notes | Ref. |
| Bacteroides fragilis | Promotes tumor growth and inflammation | TLR4/NF-κB signaling pathway | ↑ | Yes | Enterotoxigenic strains significant | [17] |
| Fusobacterium nucleatum | Increases cell proliferation and adhesion | FadA adhesion, E-cadherin activation | ↑ | Yes | Linked to tumor progression | [18] |
| Escherichia coli (pks+) | Induces genotoxicity and DNA damage | Colibactin production, DNA double-strand breaks | ↑ | No | pks+ strains carry colibactin genes | [19] |
| Faecalibacterium prausnitzii | Exhibits anti-inflammatory properties | Short-chain fatty acid production, IL-10 stimulation | ↓ | Yes | Protective role in gut health | [20] |
| Streptococcus gallolyticus | Associated with tumor development | Unknown | ↑ | No | Correlation with CRC, mechanisms unclear | [21] |
| Enterococcus faecalis | Promotes oxidative stress and DNA damage | Production of extracellular superoxide | ↑ | Yes | Role in early-stage CRC | [22] |
We would like to thank our colleagues Zhi-Peng Li, Yu-Hua Ruan, and Peng Wang for their review of this manuscript and their support with the data. We also appreciate the reviewers and editorial staff members who contributed to the review and editing of this manuscript.
| 1. | Novoa Díaz MB, Carriere P, Gentili C. How the interplay among the tumor microenvironment and the gut microbiota influences the stemness of colorectal cancer cells. World J Stem Cells. 2023;15:281-301. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 14] [Cited by in RCA: 15] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 2. | Ahamad N, Sun Y, Nascimento Da Conceicao V, Xavier Paul Ezhilan CRD, Natarajan M, Singh BB. Differential activation of Ca(2+) influx channels modulate stem cell potency, their proliferation/viability and tissue regeneration. NPJ Regen Med. 2021;6:67. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in RCA: 11] [Reference Citation Analysis (0)] |
| 3. | Wang H, Tian T, Zhang J. Tumor-Associated Macrophages (TAMs) in Colorectal Cancer (CRC): From Mechanism to Therapy and Prognosis. Int J Mol Sci. 2021;22. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 307] [Cited by in RCA: 292] [Article Influence: 58.4] [Reference Citation Analysis (5)] |
| 4. | Shin AE, Giancotti FG, Rustgi AK. Metastatic colorectal cancer: mechanisms and emerging therapeutics. Trends Pharmacol Sci. 2023;44:222-236. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 522] [Cited by in RCA: 481] [Article Influence: 160.3] [Reference Citation Analysis (0)] |
| 5. | Li J, Ma X, Chakravarti D, Shalapour S, DePinho RA. Genetic and biological hallmarks of colorectal cancer. Genes Dev. 2021;35:787-820. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 461] [Cited by in RCA: 405] [Article Influence: 81.0] [Reference Citation Analysis (7)] |
| 6. | Zheng X, Ma Y, Bai Y, Huang T, Lv X, Deng J, Wang Z, Lian W, Tong Y, Zhang X, Yue M, Zhang Y, Li L, Peng M. Identification and validation of immunotherapy for four novel clusters of colorectal cancer based on the tumor microenvironment. Front Immunol. 2022;13:984480. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 52] [Cited by in RCA: 51] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 7. | Xie S, Cai Y, Chen D, Xiang Y, Cai W, Mao J, Ye J. Single-cell transcriptome analysis reveals heterogeneity and convergence of the tumor microenvironment in colorectal cancer. Front Immunol. 2022;13:1003419. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 5] [Cited by in RCA: 29] [Article Influence: 9.7] [Reference Citation Analysis (0)] |
| 8. | Yadav VK, Huang YJ, George TA, Wei PL, Sumitra MR, Ho CL, Chang TH, Wu ATH, Huang HS. Preclinical Evaluation of the Novel Small-Molecule MSI-N1014 for Treating Drug-Resistant Colon Cancer via the LGR5/β-catenin/miR-142-3p Network and Reducing Cancer-Associated Fibroblast Transformation. Cancers (Basel). 2020;12. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 28] [Cited by in RCA: 31] [Article Influence: 5.2] [Reference Citation Analysis (0)] |
| 9. | Petitprez F, Meylan M, de Reyniès A, Sautès-Fridman C, Fridman WH. The Tumor Microenvironment in the Response to Immune Checkpoint Blockade Therapies. Front Immunol. 2020;11:784. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 127] [Cited by in RCA: 441] [Article Influence: 73.5] [Reference Citation Analysis (0)] |
| 10. | Xu C, Yan L, Guan X, Wang Z, Wu J, Lv A, Liu D, Liu F, Dong B, Zhao M, Jia L, Tian X, Hao C. Tsp2 Facilitates Tumor-associated Fibroblasts Formation and Promotes Tumor Progression in Retroperitoneal Liposarcoma. Int J Biol Sci. 2022;18:5038-5055. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 15] [Article Influence: 3.8] [Reference Citation Analysis (0)] |
| 11. | Qu R, Zhang Y, Ma Y, Zhou X, Sun L, Jiang C, Zhang Z, Fu W. Role of the Gut Microbiota and Its Metabolites in Tumorigenesis or Development of Colorectal Cancer. Adv Sci (Weinh). 2023;10:e2205563. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 212] [Cited by in RCA: 166] [Article Influence: 55.3] [Reference Citation Analysis (1)] |
| 12. | Wong CC, Yu J. Gut microbiota in colorectal cancer development and therapy. Nat Rev Clin Oncol. 2023;20:429-452. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 536] [Cited by in RCA: 456] [Article Influence: 152.0] [Reference Citation Analysis (4)] |
| 13. | Rebersek M. Gut microbiome and its role in colorectal cancer. BMC Cancer. 2021;21:1325. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 300] [Cited by in RCA: 239] [Article Influence: 47.8] [Reference Citation Analysis (4)] |
| 14. | Francescangeli F, De Angelis ML, Zeuner A. Dietary Factors in the Control of Gut Homeostasis, Intestinal Stem Cells, and Colorectal Cancer. Nutrients. 2019;11:2936. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 15] [Cited by in RCA: 35] [Article Influence: 5.0] [Reference Citation Analysis (0)] |
| 15. | Han JX, Tao ZH, Wang JL, Zhang L, Yu CY, Kang ZR, Xie Y, Li J, Lu S, Cui Y, Xu J, Zhao E, Wang M, Chen J, Wang Z, Liu Q, Chen HM, Su W, Zou TH, Zhou CB, Hong J, Chen H, Xiong H, Chen YX, Fang JY. Microbiota-derived tryptophan catabolites mediate the chemopreventive effects of statins on colorectal cancer. Nat Microbiol. 2023;8:919-933. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 178] [Cited by in RCA: 158] [Article Influence: 52.7] [Reference Citation Analysis (1)] |
| 16. | Zhang Q, Zhao Q, Li T, Lu L, Wang F, Zhang H, Liu Z, Ma H, Zhu Q, Wang J, Zhang X, Pei Y, Liu Q, Xu Y, Qie J, Luan X, Hu Z, Liu X. Lactobacillus plantarum-derived indole-3-lactic acid ameliorates colorectal tumorigenesis via epigenetic regulation of CD8(+) T cell immunity. Cell Metab. 2023;35:943-960.e9. [RCA] [PubMed] [DOI] [Full Text] [Cited by in Crossref: 368] [Cited by in RCA: 295] [Article Influence: 98.3] [Reference Citation Analysis (4)] |
| 17. | Sears CL, Garrett WS. Microbes, microbiota, and colon cancer. Cell Host Microbe. 2014;15:317-328. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 710] [Cited by in RCA: 632] [Article Influence: 52.7] [Reference Citation Analysis (11)] |
| 18. | Kostic AD, Chun E, Robertson L, Glickman JN, Gallini CA, Michaud M, Clancy TE, Chung DC, Lochhead P, Hold GL, El-Omar EM, Brenner D, Fuchs CS, Meyerson M, Garrett WS. Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe. 2013;14:207-215. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 2314] [Cited by in RCA: 2074] [Article Influence: 159.5] [Reference Citation Analysis (14)] |
| 19. | Arthur JC, Gharaibeh RZ, Mühlbauer M, Perez-Chanona E, Uronis JM, McCafferty J, Fodor AA, Jobin C. Microbial genomic analysis reveals the essential role of inflammation in bacteria-induced colorectal cancer. Nat Commun. 2014;5:4724. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 348] [Cited by in RCA: 301] [Article Influence: 25.1] [Reference Citation Analysis (4)] |
| 20. | Sokol H, Leducq V, Aschard H, Pham HP, Jegou S, Landman C, Cohen D, Liguori G, Bourrier A, Nion-Larmurier I, Cosnes J, Seksik P, Langella P, Skurnik D, Richard ML, Beaugerie L. Fungal microbiota dysbiosis in IBD. Gut. 2017;66:1039-1048. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 1094] [Cited by in RCA: 973] [Article Influence: 108.1] [Reference Citation Analysis (4)] |
| 21. | Pasquereau-Kotula E, Martins M, Aymeric L, Dramsi S. Significance of Streptococcus gallolyticus subsp. gallolyticus Association With Colorectal Cancer. Front Microbiol. 2018;9:614. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 137] [Cited by in RCA: 102] [Article Influence: 12.8] [Reference Citation Analysis (0)] |
| 22. | Strickertsson JA, Desler C, Martin-Bertelsen T, Machado AM, Wadstrøm T, Winther O, Rasmussen LJ, Friis-Hansen L. Enterococcus faecalis infection causes inflammation, intracellular oxphos-independent ROS production, and DNA damage in human gastric cancer cells. PLoS One. 2013;8:e63147. [RCA] [PubMed] [DOI] [Full Text] [Full Text (PDF)] [Cited by in Crossref: 51] [Cited by in RCA: 75] [Article Influence: 5.8] [Reference Citation Analysis (0)] |
Open-Access: This article is an open-access article that was selected by an in-house editor and fully peer-reviewed by external reviewers. It is distributed in accordance with the Creative Commons Attribution NonCommercial (CC BY-NC 4.0) license, which permits others to distribute, remix, adapt, build upon this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See: https://creativecommons.org/Licenses/by-nc/4.0/