Copyright: ©Author(s) 2026.
World J Stem Cells. Apr 26, 2026; 18(4): 115218
Published online Apr 26, 2026. doi: 10.4252/wjsc.v18.i4.115218
Published online Apr 26, 2026. doi: 10.4252/wjsc.v18.i4.115218
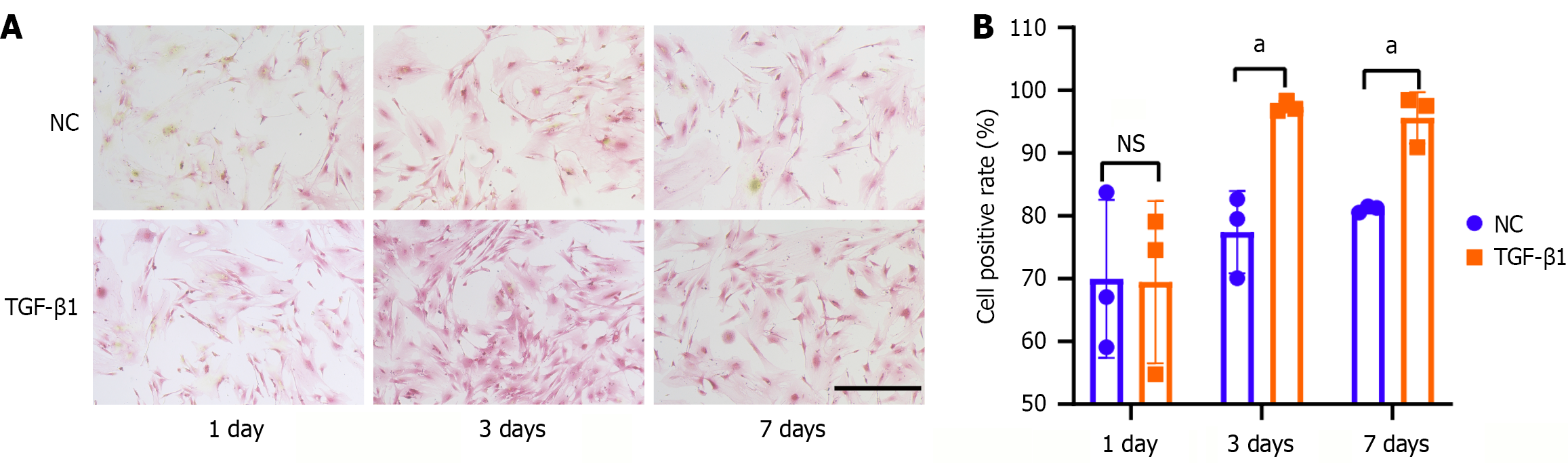
Figure 1 Transforming growth factor-β1 treatment induced collagen accumulation in tendon stem/progenitor cells.
A: Picro-Sirius red staining of tendon stem/progenitor cells after treatment with 5 ng/mL of transforming growth factor-β1 at 1 day, 3 days, and 7 days. Collagen is marked as red. Scale bars = 200 μm; B: The chart shows the percentage of the Picro-Sirius red-stained tendon stem/progenitor cells in different groups. All data are presented as the mean ± SD. aP < 0.05. NS: No significance; NC: Negative control; TGF-β1: Transforming growth factor-β1.
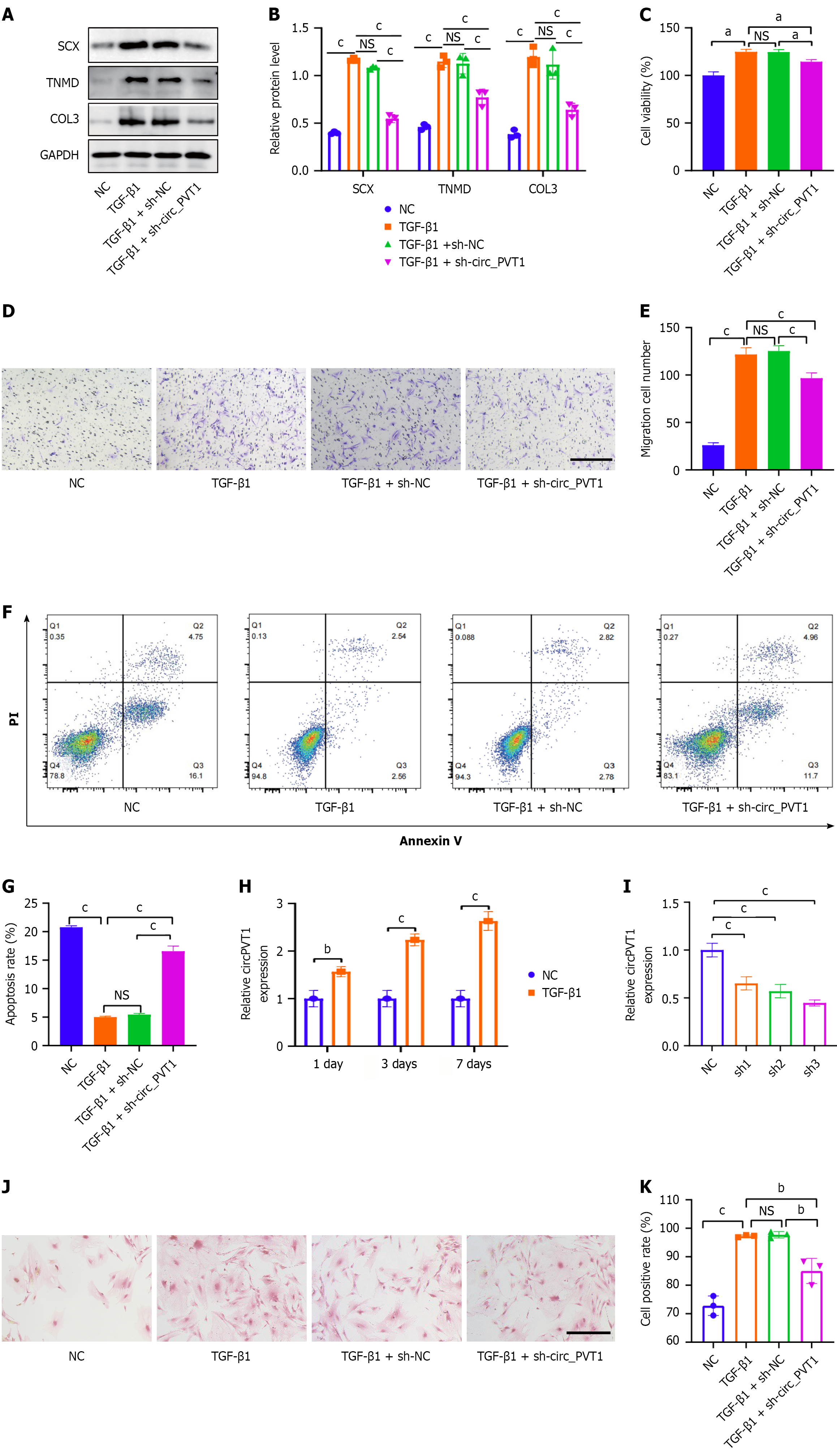
Figure 2 Knockdown of circular RNA plasmacytoma variant translocation 1 reversed the transforming growth factor-β1-induced tenogenic differentiation in tendon stem/progenitor cells.
A and B: Western blot analysis of scleraxis, tenomodulin, and COL3; C: Cell Counting Kit-8 assay for the detection of cell proliferation in tendon stem/progenitor cells (TSPCs) of the negative control (NC), Transforming growth factor (TGF)-β1, TGF-β1 + shNC, and TGF-β1 + circular RNA plasmacytoma variant translocation 1 (circ_PVT1) groups; D and E: Transwell assay for migration analysis of TSPCs in the NC, TGF-β1, TGF-β1 + shNC, and TGF-β1 + circ_PVT1 groups. Scale bars = 200 μm; F and G: Detection and analysis of apoptosis in TSPCs of the NC, TGF-β1, TGF-β1 + shNC, and TGF-β1 + circ_PVT1 groups; H: TGF-β1 treatment promoted the expression of circ_PVT1 in TSPCs. After treatment with 5 ng/mL of TGF-β1 for 1 day, 3 days, or 7 days, the expression of circ_PVT1 was increased compared with that of the normal control; I: Knockdown efficiency of shNC and various shRNAs of circ_PVT1 as determined by quantitative polymerase chain reaction in TGF-β1-induced TSPCs; J and K: The Picro-Sirius red staining of collagen in TSPCs of the NC, TGF-β1, TGF-β1 + shNC, and TGF-β1 + circ_PVT1 groups. Scale bars = 200 μm. All data are presented as the mean ± SD. aP < 0.05, bP < 0.01, cP < 0.001. SCX: Scleraxis; TNMD: Tenomodulin; NS: No significance; NC: Negative control; TGF-β1: Transforming growth factor-β1; circ_PVT1: Circular RNA plasmacytoma variant translocation 1.
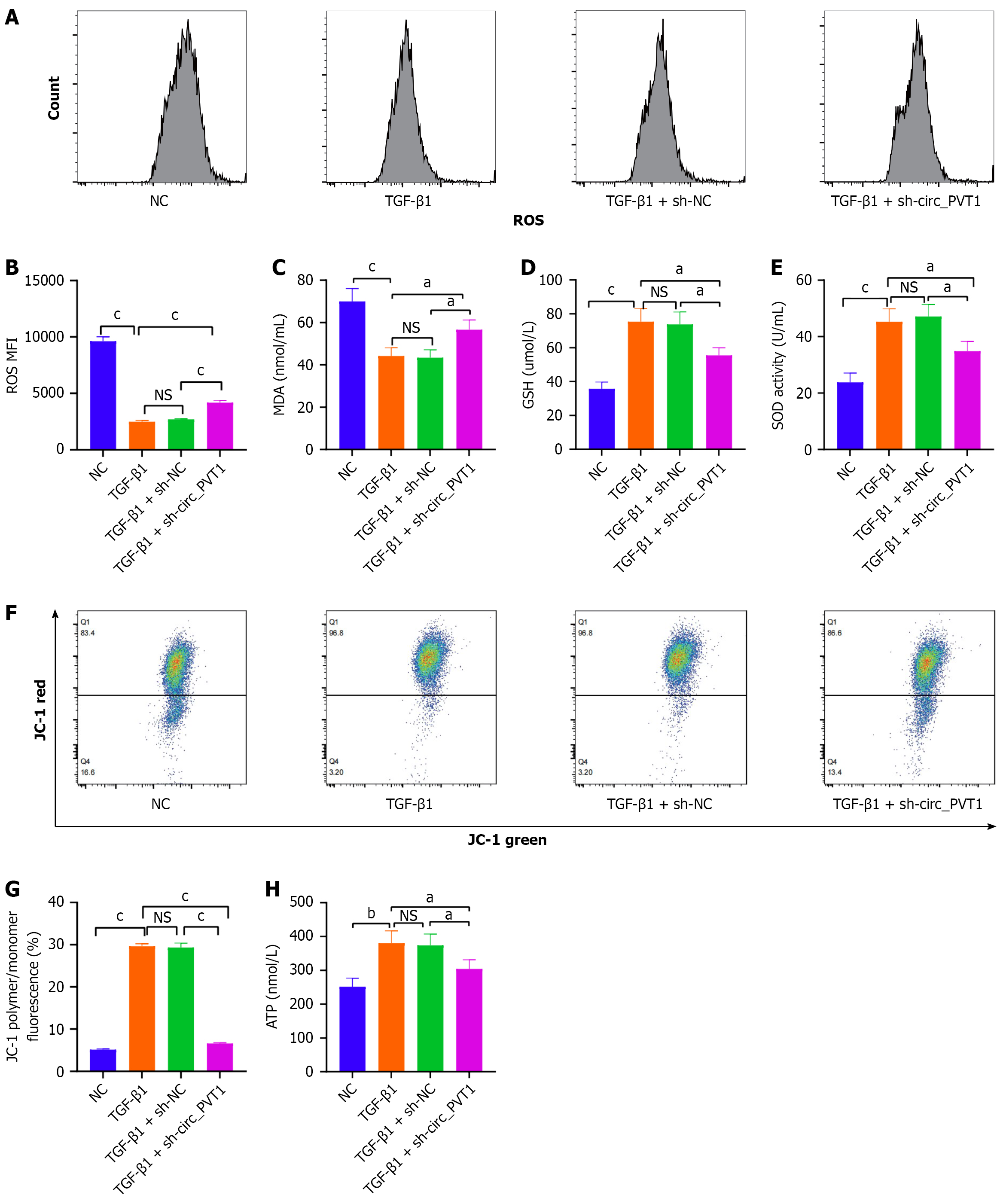
Figure 3 Knockdown of circular RNA plasmacytoma variant translocation 1 promoted oxidative stress and mitochondrial damage in transforming growth factor-β1-induced tendon stem/progenitor cells.
A and B: The DCFH-DA assay was used to detect the content of reactive oxygen species (ROS) generation of tendon stem/progenitor cells (TSPCs) in the negative control (NC), transforming growth factor (TGF)-β1, TGF-β1 + shNC, and TGF-β1 + circular RNA plasmacytoma variant translocation 1 (circ_PVT1) groups; C-E: The detection of malondialdehyde concentration, glutathione concentration, and superoxide dismutase activity in TSPCs of the NC, TGF-β1, TGF-β1 + shNC, and TGF-β1 + circ_PVT1 groups; F and G: The JC-1 assay was performed to evaluate the mitochondrial membrane potential of TSPCs in the NC, TGF-β1, TGF-β1 + shNC, and TGF-β1 + circ_PVT1 groups; H: The ATP production of TSPCs in the NC, TGF-β1, TGF-β1 + shNC, and TGF-β1 + circ_PVT1 groups. All data are presented as the mean ± SD. aP < 0.05, bP < 0.01, cP < 0.001. NS: No significance; NC: Negative control; TGF-β1: Transforming growth factor-β1; circ_PVT1: Circular RNA plasmacytoma variant translocation 1; ROS: Reactive oxygen species; MDA: Malondialdehyde; GSH: Glutathione; SOD: Superoxide dismutase.
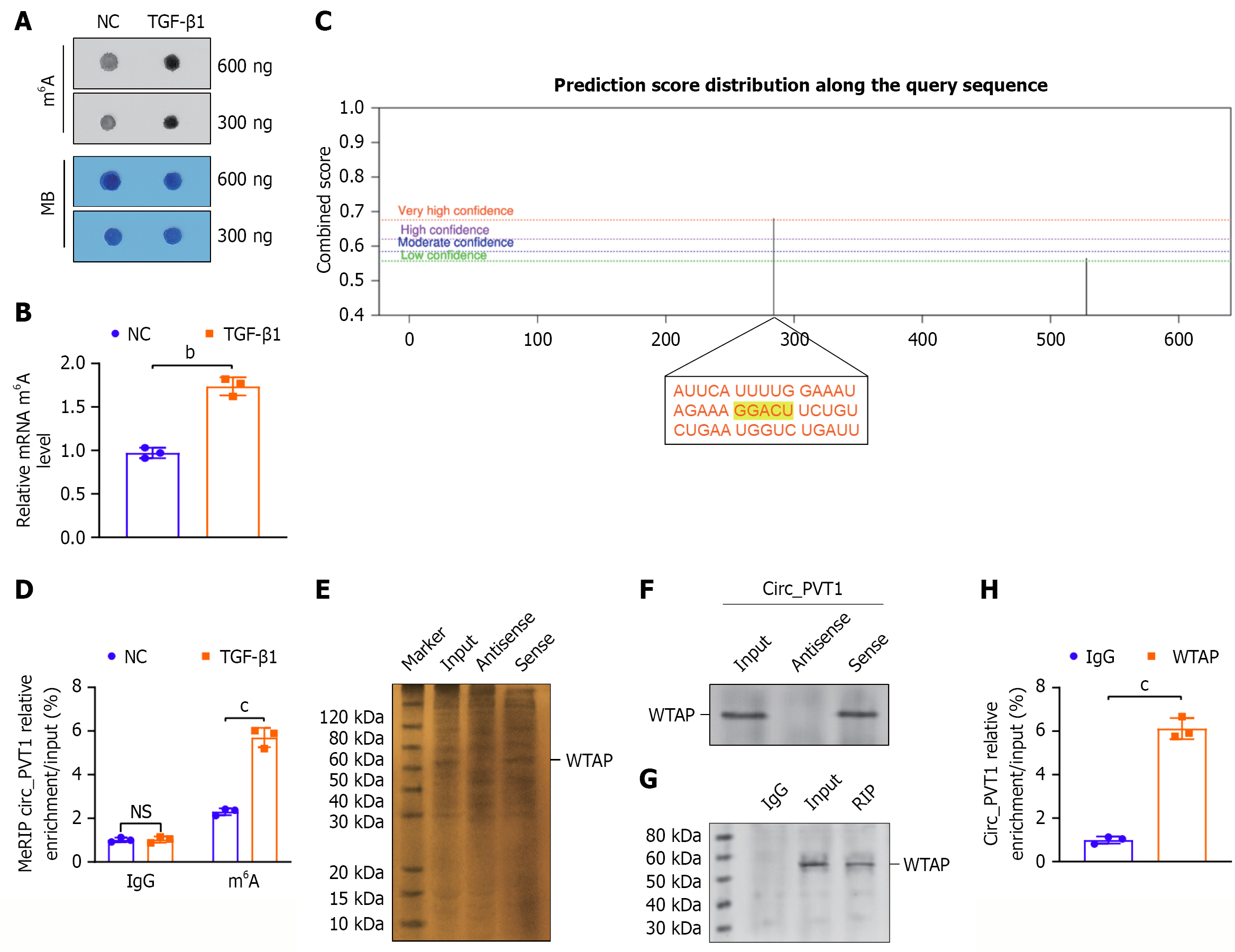
Figure 4 An N6-methyladenosine writer, WTAP, interacted with circular RNA plasmacytoma variant translocation 1.
A and B: An N6-methyladenosine (m6A) dot blot assay showing the m6A levels in the total RNA of mouse tendon stem/progenitor cells (TSPCs); C: SRAMP (http://www.cuilab.cn/sramp) software showing the potential m6A modification sites of circular RNA plasmacytoma variant translocation 1 (circ_PVT1) and the genomic sequence GGACU with very high probability; D: Methylated RNA immunoprecipitation polymerase chain reaction assay indicates that the m6A modification level of circ_PVT1 was upregulated in the transforming growth factor-β1-induced TSPCs; E and F: Silver staining of the gel after sodium-dodecyl sulfate gel electrophoresis separation of the proteins revealed that they immunoprecipitated with the 3’-biotin-labeled probe for circ_PVT1 and with the biotinylated antisense probe in RNA pull-down assays. The line shows the position of WTAP. Western blot analysis following the RNA pull-down assay indicates the interaction between circ_PVT1 and WTAP protein; G and H: Quantitative polymerase chain reaction analysis following the RNA immunoprecipitation assay confirmed the interaction between the WTAP protein and circ_PVT1. All data are presented as the mean ± SD. bP < 0.01, cP < 0.001. NS: No significance; NC: Negative control; TGF-β1: Transforming growth factor-β1; circ_PVT1: Circular RNA plasmacytoma variant translocation 1; m6A: N6-methyladenosine.
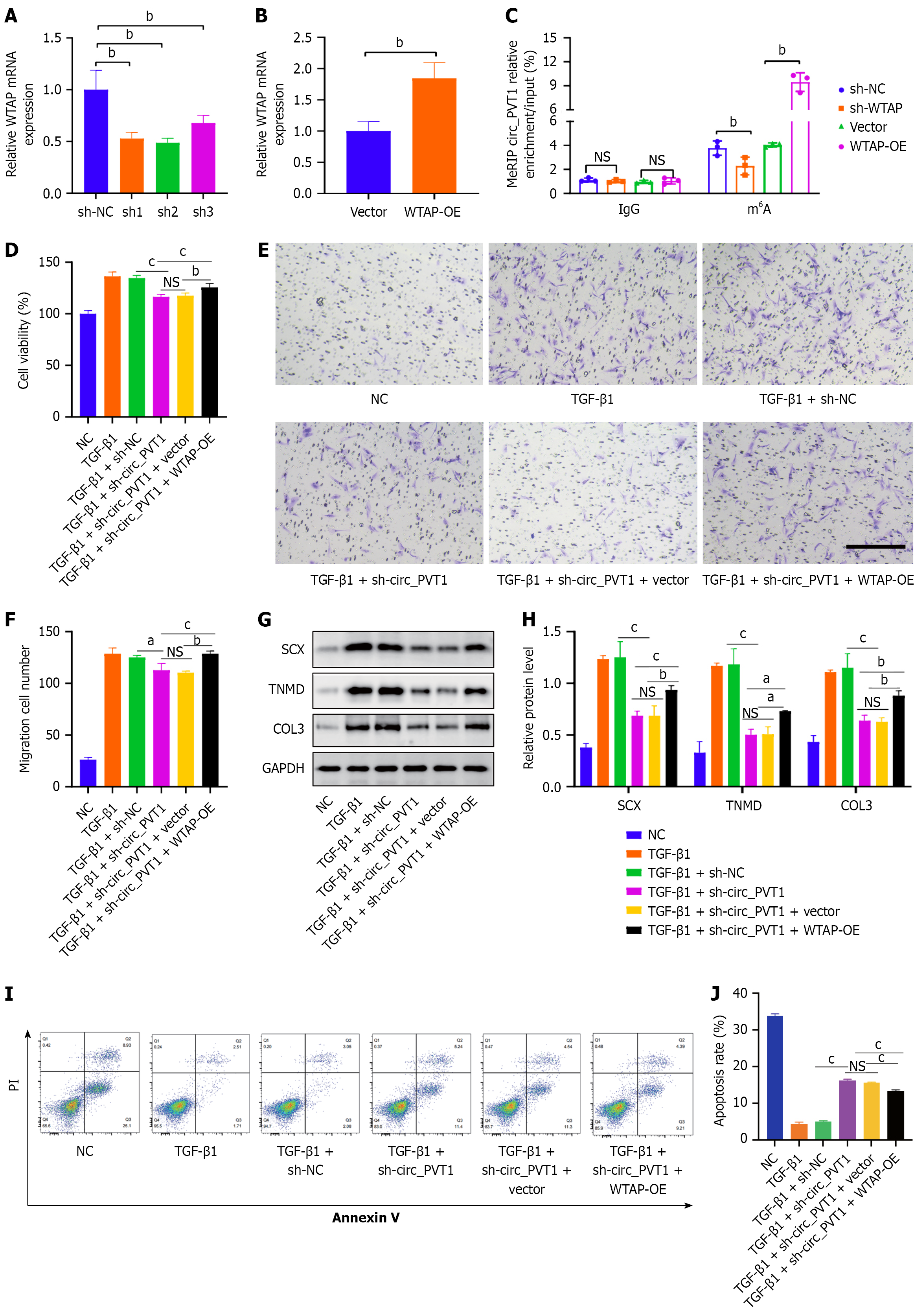
Figure 5 WTAP-mediated N6-methyladenosine modification of circular RNA plasmacytoma variant translocation 1 regulated tendon stem/progenitor cell differentiation.
A and B: The knockdown and overexpression efficiency of shNC, shRNAs, vector, and overexpression of circular RNA plasmacytoma variant translocation 1 (circ_PVT1) by quantitative polymerase chain reaction in transforming growth factor (TGF)-β1-induced tendon stem/progenitor cells (TSPCs); C: Methylated RNA immunoprecipitation polymerase chain reaction assay shows the N6-methyladenosine modification level of TGF-β1-induced TSPCs in circ_PVT1 of the shWTAP and WTAP-overexpression (OE) groups; D-J: Normal TSPCs and TGF-β1-induced TSPCs were transfected with shNC, sh-circ_PVT1, or co-transfected with sh-circ_PVT1 + WTAP-OE. A Cell Counting Kit-8 assay was performed to detect cell proliferation (D). A Transwell assay was performed for TSPC migration analysis (E and F). Western blot analysis was performed to detect the levels of scleraxis, tenomodulin, and COL3 (G and H). Apoptosis in the TSPCs was detected and analyzed (I and J). All data are presented as the mean ± SD. aP < 0.05, bP < 0.01, cP < 0.001. SCX: Scleraxis; TNMD: Tenomodulin; NS: No significance; NC: Negative control; TGF-β1: Transforming growth factor-β1; circ_PVT1: Circular RNA plasmacytoma variant translocation 1.
- Citation: Chen Y, Hu X, Jia XT, Gao RZ, Hou QS, Zhou SG. WTAP-mediated N6-methyladenosine modification of circ_PVT1 promotes proliferation and tenogenic differentiation of tendon stem/progenitor cells. World J Stem Cells 2026; 18(4): 115218
- URL: https://www.wjgnet.com/1948-0210/full/v18/i4/115218.htm
- DOI: https://dx.doi.org/10.4252/wjsc.v18.i4.115218